Abstract
Disease progression is a substantial challenge in patients with non-Hodgkin lymphoma (NHL) undergoing chimeric antigen receptor T cell (CAR-T) therapy. Here we present InflaMix (INFLAmmation MIXture Model), an unsupervised quantitative model integrating 14 pre-CAR-T infusion laboratory and cytokine measures capturing inflammation and end-organ function. Developed using a cohort of 149 patients with NHL, InflaMix revealed an inflammatory signature associated with a high risk of CAR-T treatment failure, including increased hazard of death or relapse (hazard ratio, 2.98; 95% confidence interval, 1.60–4.91; P < 0.001). Three independent cohorts comprising 688 patients with NHL from diverse treatment centers were used to validate our approach. InflaMix consistently and reproducibly identified patients with a higher likelihood of disease relapse and mortality, and it provided supplementary predictive value beyond established prognostic markers, including tumor burden. Moreover, InflaMix exhibited robust performance in cases with missing data, maintaining accuracy when considering only six readily available laboratory measures. These findings show that InflaMix is a valuable tool for point-of-care clinical decision-making in patients with NHL undergoing CAR-T therapy.
Similar content being viewed by others
Main
CD19-directed chimeric antigen receptor T cell (CAR-T) therapy brought about a paradigm shift1,2,3,4 in the treatment of relapsed or refractory large B cell lymphoma (LBCL), improving overall survival (OS) and prolonging event-free survival and progression-free survival (PFS) nearly fourfold compared to standard second-line therapy5,6,7. CAR-T therapies have also been approved for treating other variants of non-Hodgkin lymphomas (NHLs), including mantle cell lymphoma (MCL) and follicular lymphoma (FL)8,9.
Despite these advances, CAR-T treatment failure remains a substantial challenge in LBCL. Over 50% of patients with LBCL develop disease relapse or progression within the first 6 months after CAR-T therapy, and those patients have a median OS of 6 months4,10,11,12. The benefits of CAR-T treatment must also be balanced against the substantial risk of toxicities, including cytokine release syndrome (CRS), neurotoxicity, infectious complications, prolonged cytopenias and death13,14,15,16. There is a clinical need for predictive tools that can be implemented at different decision points (Supplementary Fig. 1) to identify patients at high-risk of CAR-T treatment failure.
Poor clinical outcomes have been correlated with factors such as tumor TP53 mutation, higher disease burden, elevated inflammatory markers, lower CAR-T cell expansion, and product CCR7+CD45RA+ T cell enrichment before infusion17,18,19,20. Although genomic, radiomic and CAR-T cell immunophenotyping evaluations are not widely available in clinical practice, laboratory, and cytokine measures of systemic inflammation via routine blood tests have been studied as accessible prognostic biomarkers. Prelymphodepletion levels of inflammatory markers such as interleukin-6 (IL-6), ferritin, and lactate dehydrogenase (LDH) have been inversely associated with durable response, CAR-T expansion and immunotoxicity17. These markers are surrogates for myeloid immune activation, tumor metabolic activity or cellular turnover21,22,23,24. Moreover, IL-6 and IL-10 exhibit pleiotropic effects within the lymphoma tumor microenvironment, including upregulation of regulatory T cells, inhibition of myeloid effectors and promotion of T cell exhaustion25,26. Inflammation has also been linked with toxicity indices such as the modified EASIX (Endothelial Activation and Stress Index) and CAR-HEMATOTOX, which combine inflammatory markers, such as C-reactive protein (CRP), and blood cell counts to predict cytopenias, CRS and immune-effector cell-associated neurotoxicity syndrome (ICANS)19,27. However, there are no validated biomarkers designed to predict the likelihood of relapse or disease progression following CAR-T cell therapy.
In this study, we present InflaMix (INFLAmmation MIXture Model), an unsupervised Gaussian mixture model. It defines a preinfusion laboratory and cytokine profile, evaluated at the preinfusion timepoint, that strongly correlates with and predicts poor disease response and survival after CD19-directed CAR-T therapy in NHL across multiple patient cohorts (Extended Data Fig. 1). This point-of-care tool, requiring only a single blood test, offers an unbiased quantitative assessment of 14 blood markers, 11 of which are routinely assayed for patients with lymphoma. It can also be implemented when only six specific measures (hemoglobin (Hgb), LDH, CRP, aspartate aminotransferase (AST), alkaline phosphatase (ALP) and albumin) are available.
Results
Correlated laboratory data provide complementary information
Given prior evidence that blood counts and inflammatory markers are informative of CAR-T cell therapy outcomes17,18,19,27, we explored correlative patterns among preinfusion laboratory measurements (labs; Supplementary Fig. 2) of end-organ function (creatinine, ALP, AST, alanine aminotransferase (ALT), albumin, total bilirubin (Tbili), white blood cell (WBC) count, Hgb, platelets (Plt)), tumor burden (LDH) and inflammation (CRP, ferritin, D-dimer, IL-6, IL-10 and tumor necrosis factor alpha (TNF)). We used blood tests that are part of routine clinical care at Memorial Sloan Kettering Cancer Center (MSK) in a 16-lab panel. All results were from ≤2 days before infusion. Our model-derivation cohort included 149 patients with LBCL treated with CD19 CAR-T at MSK (Table 1 and Extended Data Fig. 2).
Creatinine and ALT did not correlate with any inflammatory markers (IL-6, CRP, ferritin, LDH, IL-10 and TNF) and were discounted from further analysis (Supplementary Fig. 3). Partially overlapping groups of correlated laboratory values included measures of inflammation such as CRP, ferritin and IL-6, which were correlated among themselves and with LDH and inversely correlated with albumin and Hgb (Fig. 1a). This finding is broadly consistent with clinical intuition characterizing acute-phase reactants of systemic inflammation (for example, ferritin) and negative acute-phase reactants (for example, albumin). In contrast, IL-6 and LDH did not correlate with IL-10, a pleiotropic cytokine that regulates pro-inflammatory cytokines by negative feedback28. IL-10 correlated with Tbili, ALP and TNF and inversely correlated with albumin and Hgb. IL-10 and TNF have known associations with liver injury, where IL-10 has a hepatoprotective role29. Collectively, these findings suggest that laboratory tests of inflammation and organ function provide both partially redundant and orthogonal information.
a, Pearson correlation matrix of center-scaled values of 14 labs and cytokines normalized by ULN in the model derivation cohort (n = 149), with FDR-corrected P values of correlation tests against the null of zero correlation. Ferritin and CRP values are log-10 transformed. b, Heatmap of median scaled lab values by cluster. The text describes unscaled values with IQR. c, Scaled lab values projected in UMAP space, colored by cluster (inflammatory, n = 39; noninflammatory, n = 108). The sizes of the dots represent cluster membership probabilities in percentage corresponding to the assigned cluster. d–f, Comparing measures of tumor burden between clusters in the derivation cohort. Inferences by FDR-corrected Wilcoxon tests for LDH (d), MTV (e) and (f) SUVmax (f). g, Variable importance of labs in predicting cluster assignment, derived across 100 independent runs of cross-validated random forest models. Boxplots depict the median bounded by the first and third quartile values. Boxplot whiskers depict 1.5 times the IQR beyond the boxplot hinges. h–j, Rates of grade 2–4 CRS (h), grade 2–4 ICANS (i) and CR by day 100 (j) by cluster. Odds ratios for no CR by day 100 were estimated with 95% CI using logistic regression adjusted for age, primary refractory disease, costimulatory domain and prelymphodepletion LDH elevated above ULN. k,l, Kaplan-Meier survival estimates for PFS (k) and OS (l) stratified by cluster. HRs were estimated with 95% CI using Cox proportional hazard regression adjusted for age, primary refractory disease, costimulatory domain, and prelymphodepletion LDH elevated above ULN. Significance of cluster associations with clinical outcomes was determined by the Wald test. All tests were two sided with a significance level of 0.05. Adj., adjusted; CI, confidence interval; CR, complete response; FDR, false discovery rate; HR, hazard ratio; Infl., inflammatory cluster; IQR, interquartile range; MTV, metabolic tumor volume; Non-Infl., noninflammatory cluster; OR, odds ratio; SUV, standardized uptake value; ULN, upper limit of normal; UMAP, Uniform Manifold Approximation and Projection.
InflaMix establishes an unbiased inflammatory signature
To identify unique peri-infusion CAR-T patient subgroups using unsupervised learning in our model-derivation cohort, we built a Gaussian mixture model based on all 14 laboratory markers. This approach considered the dependency structure among different laboratory features and allowed for probabilistic classification30. Various configurations of mixture models were generated, and we selected a two-cluster model that maximized integrated complete likelihood while accounting for feature covariance (Extended Data Fig. 3a and Supplementary Notes)30. We named this model InflaMix. It identified two distinct clusters of patients, which additionally segregated well in UMAP (Uniform Manifold Approximation and Projection) space (entropy = 0.99; Methods), each patient assigned with probability > 0.88) (Fig. 1b,c). These included an ‘inflammatory’ cluster (n = 39 (26%), orange) and a ‘noninflammatory’ cluster (n = 110 (74%), blue). The inflammatory cluster enriched for patients with elevated inflammatory markers and cytokines (Fig. 1b). From a clinical perspective, the inflammatory cluster was also enriched for patients with higher rates of primary refractory disease (64% versus 23%, P < 0.001) (Extended Data Table 1). Baseline measures of disease burden, including LDH and radiomic features of lymphoma at the most recent PET-CT assessment before CAR-T (metabolic tumor volume (MTV) and maximum standardized uptake values (SUVmax)), were higher in the inflammatory cluster compared with the noninflammatory cluster (Fig. 1d–f).
To understand which features mattered most for InflaMix cluster assignment, we developed 100 iterations of a cross-validated random forest model trained to predict cluster assignment from the same features. Variable importance distributions were compared by laboratory feature (Fig. 1g). Features with both high median importance (for example, IL-6, CRP and LDH) and low median importance (for example, Hgb and Tbili) had significant discriminating power between the two clusters (P < 0.01) (Extended Data Fig. 3b–g). The discriminating power of low-importance features is likely owed to their strong correlations with high-importance features (Fig. 1a). Because InflaMix accounts for laboratory feature covariance, even low-importance features affected cluster assignment and their absence from model derivation affected cluster relationships among high-importance features (Extended Data Fig. 3h,i). Thus, all laboratory features are important for unsupervised model performance.
InflaMix is an unsupervised model, trained without knowledge of clinical outcomes. To determine whether cluster assignments were prognostic in the derivation cohort, we evaluated their predictive ability in multivariable regression models, adjusted for patient features (age), product features (costimulatory domain) and disease features (primary refractory disease and elevated prelymphodepletion LDH as a widely available surrogate for baseline disease burden). In the derivation cohort, cluster assignment was not associated with increased odds of CRS (P = 0.2) or ICANS (P = 0.2) (Fig. 1h,i). Assignment to the inflammatory cluster was associated with increased odds of not achieving a complete response (CR) by day 100 (odds ratio (OR) 4.76; 95% confidence interval (CI), 1.04–8.38; P < 0.001) (Fig. 1j), reduced PFS (increased hazard of death or relapse; hazard ratio (HR), 2.98; 95% CI, 1.60–4.91; P < 0.001) (Fig. 1k) and reduced OS (increased hazard of death; HR, 2.90; 95% CI, 1.75–5.08; P < 0.001) (Fig. 1l). In conclusion, InflaMix provides a summative and quantitative approach for patient subgrouping. Despite being an unsupervised method, InflaMix contributes additional value compared to established prognostic markers (for example, LDH) in effectively stratifying the risk of disease response and survival for patients with LBCL undergoing CAR-T therapy.
InflaMix maintains reliable clustering despite missing data
A benefit of mixture modeling is the ability to learn how features correlate during model training and leverage this information even when there are missing variables (Methods). Therefore, we hypothesized that InflaMix cluster assignments remain reliable with missing laboratory features and would still be concordant with assignments made using all 14 laboratory measures. This is a valuable property, as several measures used in InflaMix are not routinely collected. However, they are significantly correlated with readily available labs (Fig. 1a) and therefore inform clustering in new patients despite their absence. Among patients across three independent validation cohorts (Table 1), most had up to five missing labs (most commonly IL-6, IL-10, TNF and D-dimer) (Supplementary Fig. 4a,b).
We first evaluated InflaMix assignments in all MSK patients with NHL who had complete laboratory data (n = 288). These assignments were compared to those made using varying levels of simulated missing data. Even with up to five randomly missing laboratory values, we observed high consistency in cluster assignments (97% agreement and a Lin’s concordance correlation coefficient (CCC)31,32,33 of 0.93 for assignment probability; Supplementary Fig. 4c). CCC values greater than 0.81, between 0.61 and 0.80 and between 0.41 and 0.60 are typically considered ‘excellent’, ‘good’ and ‘moderate’, respectively33. Next, we evaluated InflaMix assignments using a minimum set of six core laboratory features (albumin, Hgb, AST, ALP, CRP, and LDH). These assays were selected because they are commonly available (Supplementary Fig. 4b) and had at least moderate correlation (Pearson r ≥ 0.4) with measures of higher variable importance for cluster assignment (Fig. 1g). Using this limited panel, InflaMix clustering remained highly concordant (91% agreement, CCC = 0.76) with clustering assignments derived from complete laboratory panels.
InflaMix clusters are robust pre-CAR-T properties
Validating cluster assignment by unsupervised models is challenging due to the absence of a ground truth defining an inflammatory cluster. To assess the robustness of cluster assignment, we constructed de novo, variant Gaussian mixture models in multiple bootstrapped populations from either the derivation cohort (n = 149) or an independent cohort comprised of MSK patients with NHL that have complete laboratory data (n = 139). Similarity of cluster assignments between bootstrapped mixture model variants and the original InflaMix model were then compared across all patients in both cohorts. Median agreement in cluster assignments ranged between 0.86 and 0.93 with well-calibrated assignment probabilities (Extended Data Fig. 4 and Supplementary Table 1). These comparisons included instances where mixture model variants were applied with all 14 labs and InflaMix was challenged with simulated missing values or the limited six-lab panel described above (Extended Data Fig. 4b,c,e,f).
Our findings demonstrate that inflammatory clustering, as defined by InflaMix, is highly reproducible, likely representing a fundamental biological process. We conclude that InflaMix cluster assignment is robust, even in the presence of missing informative laboratory components. Moreover, it can be reliably implemented using a core set of six widely available laboratory measurements, supporting its practicality as a point-of-care clinical tool that addresses real-world barriers to prognostication34.
InflaMix reproducibly stratifies risk across centers
We validated the association between clinical outcomes and inflammatory cluster assignment by InflaMix using two independent LBCL validation cohorts, adjusting for age, costimulatory domain, baseline elevated LDH and primary refractory disease (Table 1). The first validation cohort (MSK LBCL validation) included patients from the same center as the model-derivation cohort (MSK) with the same disease (LBCL) but who were excluded from the derivation cohort either because they had missing laboratory data or were treated after 1 January 2022 (the cutoff used to generate this same-center validation cohort). The second validation cohort included patients with LBCL from different treatment centers (Sheba Medical Center (SMC), Ramat Gan, Israel), and Hackensack Meridian Health (HMH), Hackensack, NJ); SMC + HMH LBCL validation). Inflammatory cluster assignment reproducibly identified patients with elevated inflammatory markers (Extended Data Fig. 5a–c). This assignment was associated with reduced probability of disease response and survival in both cohorts after multivariable adjustment (Fig. 2).
a–i, Kaplan-Meier survival estimates of PFS and OS and rates of CR by day 100 by InflaMix clustering across all three validation cohorts. Odds ratios of no CR by day 100 and HRs estimated with 95% CI using regression models adjusted for age, primary refractory disease, costimulatory domain and prelymphodepletion LDH elevated above ULN. Estimates for PFS (a), OS (b) and rates of CR by day 100 (c) in the MSK cohort (Cohort II); and PFS (d), OS (e) and rates of CR by day 100 (f) in the SMC + HMH LBCL cohort (Cohort III). Estimates for PFS (g), OS (h) and rates of CR by day 100 (i) in the MCL and FL cohort (Cohort IV). Regression models used for Cohort IV adjusted for age, primary refractory disease, lymphoma subtype (MCL versus FL), and prelymphodepletion LDH elevated above ULN. Significance of cluster associations with clinical outcomes was determined by the Wald test. All tests were two sided with a significance level of 0.05. HMH, John Theurer Cancer Center of Hackensack Meridian Health.
To better understand the extent to which this reproducible signature was driven by tumor burden, we evaluated risk conferred by cluster assignment after adjusting for baseline MTV instead of LDH as well as its interaction with cluster assignment across all patients at MSK who had PET radiomic assessments. Inflammatory cluster assignment remained significantly associated with increased risk of CAR-T treatment failure, as were MTV and their interaction (P < 0.001), suggesting that tumor burden and systemic inflammation are independent risk factors for CAR-T treatment failure (Supplementary Table 2). This association was consistently observed in subgroup analyses of patients with LBCL and low or high tumor burden by MTV (Extended Data Fig. 6).
InflaMix is predictive and improves clinical decision-making
A biomarker is considered predictive if its inclusion in prediction models enhances discrimination between meaningful outcomes, improves risk calibration, and aids clinical decision-making. To assess the added predictive value of InflaMix, we trained PFS prediction models using the InflaMix-derivation cohort, incorporating known clinical factors influencing CAR-T efficacy and InflaMix cluster assignment probability. InflaMix-informed prediction models were benchmarked against alternative modeling approaches in an independent validation cohort of patients with LBCL (Cohorts II and III) using key metrics including area under the receiver-operator curve (AUROC)35, calibration curves and decision curve analysis36,37. For decision curve analysis, we considered whether to pursue consolidation therapy with bispecific T cell engager therapy or autologous hematopoietic cell transplantation (auto-HCT) in patients achieving early partial response (PR) 1 month after CAR-T therapy. In this setting, the current standard approach is observation, as many will convert to CR38,39. Our approach to decision curve analysis is further explained in Methods.
We first benchmarked InflaMix clustering against an alternative modeling approach using all 14 laboratory features with regularization for dimensionality reduction instead of mixture model clustering. InflaMix-informed prediction of PFS at 6 months conferred a 9% improvement in AUROC (0.73 versus 0.64; P = 0.029) over regularized models of 14 laboratory features. Unlike InflaMix-informed models, the regularized models require availability of unconventional cytokines such as TNF and IL-10, limiting their utility in real-world settings. Next, we benchmarked InflaMix-informed prediction against models trained with known clinical drivers of CAR-T outcomes with or without CRP, the current standard biomarker of systemic inflammation. InflaMix-informed prediction of PFS at 6 months again conferred a significantly improved AUROC (P < 0.01) over both alternative models (InflaMix 0.74, CRP 0.67, Base 0.68). In all benchmark comparisons, InflaMix-informed models consistently demonstrated better calibrations and provided greater net benefit in clinical decision-making across all relevant threshold probabilities (Fig. 3). This advantage was lost if unconventional cytokines (IL-6, TNF and IL-10) were excluded from model derivation (Extended Data Fig. 7).
All models were trained using the InflaMix model-derivation cohort (Cohort I). The InflaMix model uses base clinical features (age, costimulatory domain, primary refractory disease, elevated prelymphodepletion LDH) and InflaMix score (log-transformed cluster assignment probability). Conventional model benchmarks include: Base, base clinical features only; CRP, base clinical features and prelymphodepletion CRP; Lab14Reg, regularized regression model of base clinical features and all 14 analytes used to develop InflaMix. All models were trained using Cox proportional hazards regression in the same MSK cohort InflaMix was derived from. Prediction performance was assessed using an independent validation cohort of patients with LBCL treated at MSK, SMC, or HMH. For each set of model comparisons, the validation cohort was divided into a group used to recalibrate the original model and an independent test group, repeated 100 times with twofold cross-validation for an unbiased assessment. Calibration curves, density plots, and net benefit here are evaluated using risk estimates aggregated across all repeated validation folds. A positive event here is defined as disease progression, relapse, or death by 6 months. a,b, Calibration curves of InflaMix-informed models compared to those of conventionally trained models (Lab14Reg (a), Base and CRP (b)). c,d, Decision curve analyses comparing net benefit conferred by InflaMix-informed models against conventional models (Lab14Reg (c), Base and CRP (d)) in patients who obtain a PR by day +30 after CAR-T infusion. Net benefit is evaluated for consolidation therapies across low (20%–30%) and high (30%–40%) probability threshold ranges for patients and clinicians who have lower or higher risk aversions to consolidation therapy toxicity, respectively. We suggest that bispecific T cell engager (that is, CD3xCD20 bispecific antibody) and auto-HCT consolidation should be evaluated over high probability thresholds but recognize others may consider lower threshold probabilities for bispecific antibody therapy depending on individual preferences. Auto-HCT, autologous hematopoietic cell transplantation; PR, partial response.
Although the derivation of InflaMix was unsupervised, it is a predictive biomarker that consistently outperforms alternative dimensionality-reduction methods and conventional benchmarks of known risk factors. Furthermore, InflaMix enhances the net benefit of prediction models for clinical decision-making compared to alternative approaches. Its distinct advantage in prediction stems from both our mixture modeling approach and the use of unconventional cytokines in model development.
InflaMix cluster assignments stratify risk in MCL and FL
Given that InflaMix was not derived from any disease-specific features, we hypothesized that its cluster assignment would inform disease response and survival in other lymphomas. The third validation cohort included patients from all three treatment centers (MSK, SMC and HMH) with other types of NHL, specifically MCL and FL. In this cohort, inflammatory cluster assignment was associated with lower CR rates and shorter PFS, but not shorter OS (Fig. 2g–i). The loss of association with OS might reflect the number of patients in this cohort with a more indolent disease course. Cluster laboratory profiles were again similar to those identified in the LBCL cohorts, with the inflammatory cluster enriched for patients with elevated inflammatory markers (Extended Data Fig. 5d).
To assess whether the inflammatory signature’s association with poor clinical outcomes depends on the CAR-T cell costimulatory domain, we evaluated InflaMix clustering in patients with LBCL, MCL and FL across all treatment centers, stratified by CD28- or 41BB-based CAR-T products. InflaMix assignment to the inflammatory cluster was significantly associated with decreased survival and disease response in both groups (Extended Data Fig. 8), validating its role as a disease- and product-agnostic risk stratification model in NHL.
Clustering is reliable with a simplified six-lab panel
Laboratory assay availability and limited clinician time can hinder the broad use of risk stratification tools34. To address this, we applied InflaMix using a simplified set of readily available laboratory tests (albumin, Hgb, AST, ALP, CRP and LDH), aiming to create a more accessible bedside tool. As noted above, InflaMix could be reliably applied using this six-lab panel (Extended Data Fig. 4c,f), as the model was informed by correlation with key inflammatory cytokines in the development phase. Notably, inflammatory cluster assignments with the simplified panel consistently correlated with reduced disease response and survival (Fig. 4).
a–i, Kaplan-Meier survival estimates of PFS and OS and rates of CR by day 100 by InflaMix clustering with the six-lab panel across all three validation cohorts. Odds ratios of no CR by day 100 and HRs estimated with 95% CI using regression models adjusted for age, primary refractory disease, costimulatory domain, and prelymphodepletion LDH elevated above ULN. Estimates for PFS (a), OS (b) and rates of CR by day 100 (c) in the MSK LBCL cohort (Cohort II); PFS (d), OS (e) and rates of CR by day 100 (f) in the SMC + HMH LBCL cohort (Cohort III). Estimates for PFS (g), OS (h) and rates of CR by day 100 (i) in the MCL and FL cohort (Cohort IV). Regression models used for Cohort IV adjusted by age, primary refractory disease, disease (MCL versus FL), and prelymphodepletion LDH elevated above ULN. Significance of cluster associations with clinical outcomes was determined by the Wald test. All tests were two sided with a significance level of 0.05.
We developed an online calculator for bedside application of InflaMix. The calculator is available via GitHub (https://github.com/vdblab/InflaMix). For optimal results, users enter as many of the 14 labs as are available, giving precedence to the six laboratory measurements from the limited panel.
Transition in InflaMix clusters informs clinical outcome
Systemic inflammation is a potentially modifiable property40,41. Therefore, we asked whether transition in InflaMix cluster assignments across key time points (preapheresis, prelymphodepletion and preinfusion; Fig. 5a and Extended Data Table 2) correspond with a change in clinical outcome. Notably, assignment to the inflammatory cluster by InflaMix at the preapheresis and prelymphodepletion time points still captures an inflamed phenotype (Fig. 5b and Supplementary Fig. 5). Most patients maintained their cluster assignment between preapheresis and preinfusion. Among patients in the inflammatory cluster at apheresis (n = 255), 54% transitioned to the noninflammatory cluster by CAR-T infusion and 107 of 137 received bridging antineoplastic therapy (Fig. 5a).
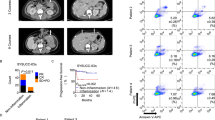
Patients across all cohorts are included in these analyses. a, Alluvial plot showing patient transitions between cluster assignments across apheresis, lymphodepletion and CAR-T infusion. Alluvia are colored by whether patients were treated with bridging therapy after apheresis. b, Heatmaps of median normalized preapheresis and prelymphodepletion laboratory values scaled by distributions of ULN-normalized preinfusion lab values in the model derivation cohort46,47. Text reports unscaled, nonnormalized medians with IQR. NA signifies laboratory measures not available at the preapheresis timepoint. c,e, Estimates of PFS (c) and OS (e) in patients who transition cluster assignments between apheresis and infusion (inflammatory cluster at apheresis to noninflammatory cluster at infusion (I. → NI.; dark orange dashed curve), noninflammatory cluster at apheresis to inflammatory cluster at infusion (N → I.; dark blue dashed curve), and no change (I.→ I.; bright orange solid curve, NI. → NI.; bright blue solid curve)). For patients who are NI. at apheresis, we report HRs of transitioning to I. at infusion versus not transitioning clusters. For patients who are I. at apheresis, we report HRs of transitioning to NI. at infusion versus not transitioning clusters. d,f, Estimates of PFS (d) and OS (f) for transitions between clusters as described above, except between lymphodepletion and infusion. HRs estimated with 95% CIs using regression models adjusted for age, primary refractory disease, costimulatory domain, bridging therapy, disease and prelymphodepletion LDH elevated above ULN. Censor marks are omitted from Kaplan-Meier curves for the sake of visual clarity. These associations remain significant if all cluster assignments are performed using only a six-lab panel (Supplementary Table 2). Significance of cluster transition associations with clinical outcomes was determined by the Wald test. All tests were two sided with a significance level of 0.05. Units of measure: albumin and Hgb, grams per deciliter; ALP, AST and LDH, units per liter; CRP and Tbili, milligrams per deciliter; D-dimer, micrograms per milliliter; ferritin, nanograms per milliliter; IL-6, IL-10 and TNF, picograms per milliliter; Plt and WBC, thousands of cells per microliter blood.
Patients initially assigned to the inflammatory cluster at preapheresis who transitioned to the noninflammatory cluster by preinfusion showed significantly better survival and disease response rates compared to those who remained in the inflammatory cluster, after adjusting for several clinical risk factors including bridging therapy (Fig. 5c,e and Supplementary Fig. 6a). A similar improvement was observed in patients who transitioned out of the inflammatory cluster between prelymphodepletion and preinfusion (Fig. 5d,f and Supplementary Fig. 6b). Conversely, patients in the noninflammatory cluster at earlier time points who shifted to the inflammatory cluster by preinfusion had worse outcomes. Our findings suggest that CAR-T treatment failure risk is not fixed by earlier cluster assignments, as patients who resolved systemic inflammation by preinfusion experienced a substantial reduction in treatment failure risk.
Discussion
CAR-T treatment failure remains a major challenge in managing refractory NHL, with approximately half of the patients experiencing relapse, depending on line of therapy11. Using an unsupervised computational approach blind to clinical outcomes, we develop InflaMix, a predictive model, designed as a point-of-care clinical tool utilizing blood test markers. InflaMix reproducibly identifies CAR-T recipients with a preinfusion inflammatory profile indicative of high risk for CAR-T treatment failure. Beyond establishing a strong association between cluster assignment and clinical outcomes in the derivation cohort, we validated this association across three independent cohorts totaling 688 patients, each differing in clinical characteristics, geography, NHL subtypes and CAR-T products. Prior studies have linked inflammatory markers and toxicity19,27,42. Our findings extend previous work17,26,27,43 by emphasizing a role for inflammation in CAR-T treatment failure, and introducing a robust, easily implementable bedside tool for risk stratification.
The mixture modeling approach used the joint distributions of all laboratory variables. Covariance between uncommonly assessed cytokines directly related to immune activation (that is, IL-6 and IL-10)25,26 and commonly accessible measures such as albumin, Hgb, and LDH informed cluster assignment and outcome prediction. These features collectively refine an inflammatory signature beyond individual surrogate assays like CRP (Extended Data Fig. 7). This is highlighted both by the successful validation of InflaMix in an independent cohort of patients from different treatment centers where cytokine measures (IL-6, IL-10 and TNF) are not assessed and by the model’s ability to accurately assign clusters using a simplified six-lab panel.
Validation and implementation are important for risk stratification models and are often hindered by model overfitting to the training data. To reduce the risk of overfitting, we used an unsupervised approach guided by prior evidence17,19,25,27. We ensured the derivation cohort data were kept separate during scaling and applied a frugal parameterization of the mixture model (Methods). This strategy facilitated derivation of the InflaMix signature and explains its predictive capacity for poor clinical outcomes beyond known drivers of CAR-T treatment failure, such as individual inflammatory markers and disease burden. InflaMix-informed prediction resulted in greater discrimination and improved calibration compared to rigorous benchmarks. Most importantly, InflaMix improved net benefit of prediction models for clinically relevant decision-making compared to alternative modeling approaches and conventional clinical risk factors. This further established the value of our mixture modeling approach using unconventional cytokine assays and careful selection of the other laboratory features.
Our work builds on the link between inflammation and poor CAR-T outcomes17,19,26,27,40,44. Previous studies have demonstrated that tumor burden and myeloid-derived inflammatory markers, such as IL-6 and monocytic-myeloid-derived suppressor cells, are associated with reduced probability of durable response, suggesting that myeloid-derived inflammation stunts CAR-T activation and expansion17,26. Scholler et al.43, however, reported that tumor microenvironment immune contextures in LBCL associated with CAR-T treatment failure did not correlate with myeloid cell densities, suggesting a more complicated role for myeloid-derived inflammation. InflaMix cluster assignment reproducibly identified a role for IL-6, CRP, and ferritin in CAR-T treatment failure, but also characterized an additional axis beyond tumor burden and these individual myeloid-derived inflammatory markers to improve associative and predictive strength for disease response. Further mechanistic studies are needed to explore associations between InflaMix cluster assignment and the tumor immune environment, as well as CAR-T function.
To determine the effect of a changing inflammatory environment on CAR-T outcomes, we evaluated InflaMix cluster assignment at apheresis and lymphodepletion. We observed an association between resolution of inflammation by preinfusion and improved outcomes, which were nearly identical to those in patients without inflammatory cluster assignment at earlier time points. This finding suggests that preapheresis inflammation is not associated with irreversible, diminished CAR-T functionality or exhaustion and points to the value of intervening on the preinfusion inflammatory cytokine milieu and tumor microenvironment. Most patients who remained in the inflammatory cluster by infusion had already undergone bridging therapy and lymphodepletion. Our observations suggest that targeting residual inflammation after lymphodepletion via anti-inflammatory treatments before CAR-T infusion may improve outcomes, although this requires further study.
InflaMix defines an inflammatory signature that is reproducibly associated with poor clinical outcomes in multiple contexts. Nonetheless, this study has important limitations. InflaMix was derived from a retrospective, single-center cohort. Prospective and mechanistic studies of InflaMix are needed to mitigate bias, evaluate its real-time utility in risk stratification, and determine if there is a causal link between preinfusion inflammation and poor clinical outcomes. Additionally, InflaMix does not consider all the factors contributing to CAR-T efficacy.
The predictive capacity of InflaMix may be augmented in multimodal models considering clinical, tumor genomic and radiomic features18,20,45. It might also be useful in other contexts: InflaMix is applied before CAR-T infusion, suggesting that its inflammatory signature is not necessarily specific to the CAR-T context and may have broader utility across T cell immunotherapies including immune checkpoint blockade and stem cell transplant.
InflaMix uses a novel, unbiased approach to characterize a preinfusion inflammatory signature, encouraging new mechanistic hypotheses for CAR-T resistance in NHL. With further prospective validation, we envision InflaMix being implemented in clinical practice and trial design to identify patients at high risk of treatment failure and support informed risk-benefit discussions for prophylactic or consolidative therapies. The predictive capacity of InflaMix complements existing toxicity prediction tools and may enhance multimodal prediction of CAR-T outcomes. Due to its robust performance with incomplete data, InflaMix can also be used effectively with a limited six-lab panel (albumin, AST, ALP, Hgb, LDH and CRP) and is easily implemented in clinical settings via an online calculator https://github.com/vdblab/InflaMix), reducing barriers to point-of-care use that often hinder other prognostic tools.
Methods
This study was conducted in accordance with the principles outlined in the Declaration of Helsinki. Ethical approval was obtained from the institutional review boards (IRBs) of all participating institutions, including MSK, SMC and HMH. The study involved retrospective data collection from medical records, and as such, the requirement for informed consent was waived by the IRBs at all institutions. All patient data were handled in compliance with applicable privacy and confidentiality regulations.
Patient characteristics
This was a multicenter observational study of patients with NHL (age ≥18 years) treated with autologous, commercially available CD19 CAR-T therapy, including a derivation cohort and three validation cohorts (Table 1 and Extended Data Figs. 1a and 2). Patient clinical data were manually reviewed and entered into a REDCap database48. Laboratory values were collected for 16 assays: Hgb, Plt, WBC, ALP, Tbili, IL-10, TNF, LDH, D-dimer, ferritin, CRP, AST, IL-6, creatinine, fibrinogen and albumin. These values were obtained from the electronic medical record at specified time points (days (d)) before or after apheresis (d−10, d+1 (preapheresis), lymphodepletion (d−2, d1 (prelymphodepletion) and cell infusion (d−2, d0) (preinfusion)). The study was approved by the IRBs of each institution (MSK, SMC and HMH).
MSK derivation cohort
We first defined a cohort of 149 patients with LBCL who were treated with autologous CD19 CAR-T infusion (58% axicabtagene ciloleucel (axicel), 31% tisagenlecleucel (tisacel) and 11% lisocabtagene maraleucel (lisocel)) at MSK, New York, NY) between 1 April 2016 and 1 January 2022. These patients had no missing laboratory features from our laboratory panel (Table 1, Cohort I). Except lisocel infusions, which were performed before 2021 as part of the TRANSCEND NHL 001 study (NCT02631044)3, CAR-T products were administered as standard therapy.
Validation cohorts
All patients in the validation cohorts were treated with CAR-T infusion between 1 April 2016 and 1 April 2024. Missing laboratory data were allowed. Cohort II (MSK LBCL validation) included 186 patients with LBCL treated at MSK with CAR-T (47% axicel, 13% tisacel and 40% lisocel) and not in the derivation cohort (Cohort I). Cohort III (SMC + HMH LBCL validation) included 243 patients with LBCL treated with CD19 CAR-T. SMC does not treat many patients with lisocel and has its own unique CAR-T construct. Cohort III patients were treated with 37% SMC-specific point-of-care CD28-costimulatory domain-based CAR-T49 (38% axicel, 20% tisacel and 5% lisocel) at either SMC (73%) or HMH (27%). Cohort IV (MCL and FL validation) included 110 patients with MCL (55%) or FL (45%) treated with CAR-T (27% SMC-specific point-of-care CD28-costimulatory domain-based CAR-T49; 29% brexucabtagene autoleucel, 17% axicel, 22% lisocel and 5% tisacel) at MSK (61%), SMC (27%) or HMH (12%) (Table 1).
Definitions
Day 100 CR to CAR-T was defined by a best response of CR ≤ 100 days after infusion, according to the Lugano criteria50. Disease status before CAR-T infusion was defined by the most recent disease assessment before infusion. Stage at apheresis was defined by Ann Arbor staging51. CRS and ICANS were graded using the American Society for Transplantation and Cellular Therapy grading criteria52. OS and PFS were measured from time of CAR-T infusion until death and until progression or death, respectively. CAR-T treatment failure was defined as disease progression, relapse or death after CAR-T therapy.
Laboratory data normalization and scaling
To account for variability in laboratory assays used both within and across different institutions, all values were normalized by the associated upper limit of normal (ULN). If a measurement was reported as either less than or greater than a limit of detection, those limit values were used instead. For example, for a ferritin measure reported as <3 ng ml−1 (lower limit of normal 7, ULN 245), the value was normalized as 3/245 = 0.012. The distributions of ULN-normalized preinfusion labs from the model derivation cohort showed that most feature distributions were skewed (Supplementary Fig. 7). To avoid arbitrary and superfluous data transformation, we only applied log-10 transformations when skew was >1 and was reduced by ≥90% after transformation (CRP and ferritin). ULN-normalized preinfusion values in the derivation cohort were then centered and scaled to mean 0 and variance 1. Laboratory values across all model-derivation and validation cohorts across all time points were also normalized by their corresponding ULN53,54. Ferritin and CRP were log-10 transformed. Finally, all values were scaled and centered by the mean and variance of the feature distributions of Cohort I (derivation cohort) at preinfusion. This approach avoided influencing model development by validation cohort data parameters or over- or underscaling extreme laboratory values from uncommon assays, as well as providing a rigorous framework for normalizing and scaling individual patient data for a point-of-care tool.
InflaMix derivation
Using normalized and scaled preinfusion laboratory data from the derivation cohort, we generated Gaussian mixture models using the mclust package in R statistical software (R Foundation for Statistical Computing, version 4.4.1)30. The best two-cluster model that also allowed for flexible parameterization of feature covariance (model VVV) was selected based on the integrated, complete-data likelihood Bayesian information criterion metric (Extended Data Fig. 3a, Supplementary Notes and Supplementary Fig. 8)30. The various combinations of parameterizations (for example, VVI, VVV) are described in Scrucca et al.30. We named this mixture model InflaMix. The distributions of feature importance in the mixture model were evaluated by 100 independent random forest models, where the mixture model cluster assignments were treated as outcome while normalized laboratory measurements were used as features. For each individual random forest model, the corresponding hyperparameters were determined by a fivefold cross-validation.
A custom approach was applied to calculate the cluster membership probability for patients with partially available laboratory data. For a patient with a vector of values \(x\), the class-specific joint density of \(x\) for the ith cluster, \({f}_{i}\left({x;}{\hat{\mu }}_{i},{\hat{\Sigma }}_{i}\right)=\frac{1}{\sqrt{{\left(2\pi \right)}^{d}\det \left({\hat{\Sigma }}_{i}\right)}}\exp \{-\frac{1}{2}{\left(x-{\hat{\mu }}_{i}\right)}^{T}{\hat{\Sigma }}_{i}\left(x-{\hat{\mu }}_{i}\right)\}\), was calculated using the mvtnorm package in R. Here, \({\hat{\mu }}_{i}\) and \({\hat{\Sigma }}_{i}\) were the estimated class-specific mean vector and covariance matrix, respectively, for the ith cluster, where i = 1, 2. If any missing values from \(x\), the corresponding values and entries were removed from \({\mu }_{i}\) and \({\Sigma }_{i}\) and only the dim=d available labs were included. The posterior probability of assignment to cluster i is then given by \(\frac{{\hat{p}}_{i}\,{f}_{i}\left({x;}{\hat{\mu }}_{i},{\hat{\Sigma }}_{i}\right)}{\mathop{\sum }\nolimits_{k=1}^{2}{\hat{p}}_{k}{f}_{k}\left({x;}{\hat{\mu }}_{k},{\hat{\Sigma }}_{k}\right)}\), where \({\hat{p}}_{i}\) is the estimated marginal relative frequency of cluster i. Entropy of clustering is given by \(1+\frac{{\sum }_{i=1}^{2}{\sum }_{n=1}^{N}{\tau }_{n,i}\mathrm{ln}{\tau }_{n,i}}{N\mathrm{ln}2}\), where N is the number of patients being clustered and \({\tau }_{n,{i}}\) is the posterior probability of assigning patient n to cluster i. Cluster assignment is defined when the probability of belonging to a given cluster is greater than 50%. To evaluate the agreement between cluster assignment by InflaMix with or without missing values (Supplementary Fig. 4c), we considered all 288 patients with complete laboratory data. We then simulated n randomly missing labs from each patient across 100 different iterations of up to seven missing labs (700 iterations total). Agreement was measured either by the proportion of matching cluster assignments or by calculating Lin’s CCC31,32 between cluster assignment probabilities assigned with and without missing laboratory features.
Cluster assignment consistency
To assess the robustness of cluster assignment, we built alternative Gaussian mixture models (which we will refer to as mixture model variants) using 100 bootstrapped populations from the derivation cohort (Cohort I) and compared similarity of cluster assignment with InflaMix in the derivation cohort by agreement in cluster assignments, adjusted Rand Index55 and by CCC between the cluster assignment probabilities averaged across all bootstraps. To increase the rigor of this analysis, we pursued a similar approach in an additional validation cohort (n = 139) of patients with NHL with fully available laboratory data, which is needed for mixture model development. We named this cohort the cluster assignment validation (CAV) cohort. We then built mixture model variants from 100 bootstrapped populations from the CAV cohort and compared similarity of cluster assignment with InflaMix in the CAV cohort (Extended Data Fig. 4a,d and Supplementary Table 1).
Finally, because InflaMix can be applied towards patients with missing laboratory data, we repeated the same analyses noted above over multiple iterations with various combinations of simulated missing laboratory values. Cluster assignments by InflaMix were made with missing laboratory values to generate the predicted probabilities, but the approximated ‘true’ probabilities of cluster assignment were generated by mixture model-variant mixture models using all 14 laboratory values. We evaluated cluster assignment concordance and calibration as defined above when InflaMix was applied with up to seven missing laboratory values (Extended Data Fig. 4b,e and Supplementary Table 1) and with only the six-lab panel (albumin, Hgb, CRP, AST, LDH and ALP; Extended Data Fig. 4c,f and Supplementary Table 1). Except in the case of the six-lab panel, laboratory data missingness was simulated with random sampling informed by the rate of missingness for each laboratory assay across all cohorts. This was repeated ten times for each patient and InflaMix assignment probabilities were averaged across the repeats. Given that InflaMix is trained by complete laboratory data, cluster assignments obtained by partially available data are expected to be robust to some extent, but not as effective as using complete data. Therefore, compared to the concordance level (for example, CCC) using full data, a lower concordance level using data with more severe missingness is also well expected.
Radiomic features
We performed 18F-FDG PET/CT on various in-house and outside scanners, either shortly before apheresis and/or after apheresis or bridging therapy, but before CAR-T cell infusion. At MSK, PET/CT was performed on Discovery 690 and Discovery 710 scanners (GE Healthcare) 1 h after intravenous injection of 444 MBq ± 10% of 18F-FDG. A low-dose, non-contrast-enhanced CT scan from skull base to upper thighs was used for attenuation correction. A heavy z-axis filter and Gaussian transaxial filter with 6.4 mm cutoff was used. Blood glucose levels were <180 mg dl−1 prior to PET. Using the Beth-Israel PET/CT viewer plugin (v.4.14) and the International Biomarker Standardization Initiative compliant PyRadiomics plugin (v.2.2.0) for FIJI (v.1.52 g)56,57, MTV was constructed semiautomatically by a board-certified radiologist as previously described58; the reader had access to current, previous and follow-up imaging data and reports. An SUV threshold of 4–200 was used. Maximum lesion diameter50 was measured on CT transaxial or coronal planes59 in the Hermes Viewer software (v.6.1.4; Hermes Medical Solutions).
Statistical analysis
Clinical outcome associations
Multivariable Cox proportional hazards models and multivariable logistic regression models were fitted to obtain the corresponding HRs and ORs. To account for the propagated uncertainty of parameter estimates from the Gaussian mixture model to subsequent regression models, the regression models were fitted for the pseudo-observations for both classes weighted by the cluster membership probabilities60, while the inferences were conducted by bootstrap with 100 resamples. For validation cohorts, CIs and the P values were calculated via an analytical approach assuming that the membership probability weights were observed quantities. All tests were two sided with a significance level of 0.05. Two sets of regression models were fitted to evaluate the risk of CAR-T treatment failure with cluster transitions either between apheresis and infusion or lymphodepletion and infusion. The transitions were determined by changes in assigned cluster labels based on the posterior probability. One set of models evaluated odds of not achieving CR by day 100 and hazard of death or disease progression, if transitioning from the inflammatory cluster at apheresis or lymphodepletion to the noninflammatory cluster at infusion compared with not transitioning. Another set evaluated the converse transition. Covariates included in each multivariable model are reported in Supplementary Table 2. The bridging therapy variable was stratified as systemic therapy, radiation therapy, or no bridging.
InflaMix cluster properties
Wilcoxon rank-sum tests were performed for comparisons of continuous variables across clusters with FDR correction for multiple hypotheses61. Pearson chi-squared tests were performed for binary variables. Pearson correlation was used to assess correlation between different laboratory features, where the P values of the corresponding tests against zero correlation were also FDR corrected. All tests were two sided with a significance level of 0.05.
Prediction modeling
An InflaMix-informed Cox proportional hazards regression model for PFS was trained using the model derivation cohort using age, costimulatory domain, primary refractory disease and elevated prelymphodepletion LDH as base clinical variables and the log-transformed (base 10) value of inflammatory cluster assignment probability. Alternative benchmark models were trained on the same cohort using the same 4 base clinical features and either (1) all 14 individual laboratory features used to derive InflaMix subjected to regularization, (2) prelymphodepletion CRP, (3) no other features, or (4) inflammatory cluster assignment probability by a two-cluster, VVV parameterized Gaussian mixture model30 derived without IL-6, TNF and IL-10. Regularization was achieved by the least absolute shrinkage and selection operator (LASSO) with a penalty parameter tuned to reach the minimum mean cross-validated error (\({\lambda }_{\min }\)) using the glmnet R package. Model performance was compared using an independent validation cohort composed of patients with LBCL (Cohorts II and III). The validation cohort subset for each comparative analysis was constrained by laboratory data availability required of all models being compared but was the same within each comparison. AUROCs were calculated for predicting PFS at 6 months and compared using the Wald test35. Systemic miscalibration was expected given the stark differences in our temporally defined training and validation cohorts. We used a parsimonious updating method where the validation cohort was divided into a group used to recalibrate the original models and an independent test group, repeated 100 times with twofold cross-validation for an unbiased assessment62. The recalibrated risk estimates were then used for decision curve analysis.
Decision curve analysis
Decision curve analysis helps interpret model predictions in the context of outcome prevalence, weigh the benefits and risks of a specific clinical intervention and estimate net benefit36,37,63. Net benefit is evaluated over a range of relevant threshold probabilities. The threshold probability can be interpreted as patient or clinician preference to balance aversion to the toxicity of the intervention against the perceived risk of relapse. The threshold probability of relapse or death should be high for pursuing a more toxic intervention. We compared the utility of all prediction models to decide on consolidation with bispecific T cell engager or auto-HCT therapy 1 month after CAR-T in patients achieving PR at the standard first disease response assessment. This clinical scenario is an area of active investigation64,65 and ideal for applying prediction modeling, as there is equipoise between balancing toxicity and benefit in preventing relapse66. In the early post-CAR-T setting, these immunotherapies can compound immune, hematologic and infectious toxicities and would warrant high threshold probabilities. In our estimation, this would be roughly 20% to 30% for bispecific T cell engager therapy and 30% to 40% for auto-HCT.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Data requests for patient-related laboratory measurements or clinical outcomes will be reviewed by the corresponding author in consultation with coauthors from HMH and SMC. Any data and materials that can be shared will be released via data transfer agreement. Laboratory values and their corresponding upper limits of normal for the InflaMix-derivation cohort of MSK patients are provided in the Source data. Source data are provided with this paper.
Code availability
All statistical and computational analyses were performed using R v.4.4.1. The code, all packages used, the InflaMix model, and detailed instructions for its implementation can be accessed via GitHub (https://github.com/vdblab/InflaMix). The clinical calculator application for implementing InflaMix can be accessed via this GitHub link as well.
References
Neelapu, S. S. et al. Axicabtagene ciloleucel CAR T-cell therapy in refractory large B-cell lymphoma. N. Engl. J. Med. 377, 2531–2544 (2017).
Schuster, S. J. et al. Tisagenlecleucel in adult relapsed or refractory diffuse large B-cell lymphoma. N. Engl. J. Med. 380, 45–56 (2019).
Abramson, J. S. et al. Lisocabtagene maraleucel for patients with relapsed or refractory large B-cell lymphomas (TRANSCEND NHL 001): a multicentre seamless design study. Lancet 396, 839–852 (2020).
Locke, F. L. et al. Long-term safety and activity of axicabtagene ciloleucel in refractory large B-cell lymphoma (ZUMA-1): a single-arm, multicentre, phase 1-2 trial. Lancet Oncol. 20, 31–42 (2019).
Locke, F. L. et al. Axicabtagene ciloleucel as second-line therapy for large B-cell lymphoma. N. Engl. J. Med. 386, 640–654 (2022).
Kamdar, M. et al. Lisocabtagene maraleucel versus standard of care with salvage chemotherapy followed by autologous stem cell transplantation as second-line treatment in patients with relapsed or refractory large B-cell lymphoma (TRANSFORM): results from an interim analysis of an open-label, randomised, phase 3 trial. Lancet 399, 2294–2308 (2022).
Westin, J. R. et al. Survival with axicabtagene ciloleucel in large B-cell lymphoma. N. Engl. J. Med. 389, 148–157 (2023).
Jacobson, C. A. et al. Axicabtagene ciloleucel in relapsed or refractory indolent non-Hodgkin lymphoma (ZUMA-5): a single-arm, multicentre, phase 2 trial. Lancet Oncol. 23, 91–103 (2022).
Wang, M. et al. KTE-X19 CAR T-cell therapy in relapsed or refractory mantle-cell lymphoma. N. Engl. J. Med. 382, 1331–1342 (2020).
Nastoupil, L. J. et al. Standard-of-care axicabtagene ciloleucel for relapsed or refractory large B-cell lymphoma: results from the US Lymphoma CAR T Consortium. J. Clin. Oncol. 38, 3119–3128 (2020).
Alarcon Tomas, A. et al. Outcomes of first therapy after CD19-CAR-T treatment failure in large B-cell lymphoma. Leukemia 37, 154–163 (2023).
Spiegel, J. Y. et al. Outcomes of patients with large B-cell lymphoma progressing after axicabtagene ciloleucel therapy. Blood 137, 1832–1835 (2021).
Pennisi, M. et al. Comparing CAR T-cell toxicity grading systems: application of the ASTCT grading system and implications for management. Blood Adv. 4, 676–686 (2020).
Wudhikarn, K. et al. DLBCL patients treated with CD19 CAR T cells experience a high burden of organ toxicities but low nonrelapse mortality. Blood Adv. 4, 3024–3033 (2020).
Wudhikarn, K. et al. Infection during the first year in patients treated with CD19 CAR T cells for diffuse large B cell lymphoma. Blood Cancer J. 10, 79 (2020).
Jain, T. et al. Hematopoietic recovery in patients receiving chimeric antigen receptor T-cell therapy for hematologic malignancies. Blood Adv. 4, 3776–3787 (2020).
Locke, F. L. et al. Tumor burden, inflammation, and product attributes determine outcomes of axicabtagene ciloleucel in large B-cell lymphoma. Blood Adv. 4, 4898–4911 (2020).
Vercellino, L. et al. Predictive factors of early progression after CAR T-cell therapy in relapsed/refractory diffuse large B-cell lymphoma. Blood Adv. 4, 5607–5615 (2020).
Pennisi, M. et al. Modified EASIX predicts severe cytokine release syndrome and neurotoxicity after chimeric antigen receptor T cells. Blood Adv. 5, 3397–3406 (2021).
Shouval, R. et al. Impact of TP53 genomic alterations in large B-cell lymphoma treated with CD19-chimeric antigen receptor T-cell therapy. J. Clin. Oncol. 40, 369–381 (2022).
Jagannath, S. et al. Tumor burden assessment and its implication for a prognostic model in advanced diffuse large-cell lymphoma. J. Clin. Oncol. 4, 859–865 (1986).
Ferraris, A. M., Giuntini, P. & Gaetani, G. F. Serum lactic dehydrogenase as a prognostic tool for non-Hodgkin lymphomas. Blood 54, 928–932 (1979).
International Non-Hodgkin’s Lymphoma Prognostic Factors Project A predictive model for aggressive non-Hodgkin’s lymphoma. N. Engl. J. Med. 329, 987–994 (1993).
Kernan, K. F. & Carcillo, J. A. Hyperferritinemia and inflammation. Int. Immunol. 29, 401–409 (2017).
Gholiha, A. R. et al. Revisiting IL-6 expression in the tumor microenvironment of classical Hodgkin lymphoma. Blood Adv. 5, 1671–1681 (2021).
Jain, M. D. et al. Tumor interferon signaling and suppressive myeloid cells are associated with CAR T-cell failure in large B-cell lymphoma. Blood 137, 2621–2633 (2021).
Rejeski, K. et al. CAR-HEMATOTOX: a model for CAR T-cell-related hematologic toxicity in relapsed/refractory large B-cell lymphoma. Blood 138, 2499–2513 (2021).
Saraiva, M. & O’Garra, A. The regulation of IL-10 production by immune cells. Nat. Rev. Immunol. 10, 170–181 (2010).
Nagaki, M. et al. High levels of serum interleukin-10 and tumor necrosis factor-alpha are associated with fatality in fulminant hepatitis. J. Infect. Dis. 182, 1103–1108 (2000).
Scrucca, L., Fop, M., Murphy, T. B. & Raftery, A. E. mclust 5: clustering, classification and density estimation using Gaussian finite mixture models. R J. 8, 289–317 (2016).
Lin, L., Hedayat, A. S. & Wu, W. A unified approach for assessing agreement for continuous and categorical data. J. Biopharm. Stat. 17, 629–652 (2007).
Lin, L. I. A concordance correlation coefficient to evaluate reproducibility. Biometrics 45, 255–268 (1989).
Kocks, J. W. et al. Health status in routine clinical practice: validity of the clinical COPD questionnaire at the individual patient level. Health Qual. Life Outcomes 8, 135 (2010).
Tuzzio, L. et al. Barriers to implementing cardiovascular risk calculation in primary care: alignment with the consolidated framework for implementation research. Am. J. Prev. Med. 60, 250–257 (2021).
Harrell, F. E. Jr, Califf, R. M., Pryor, D. B., Lee, K. L. & Rosati, R. A. Evaluating the yield of medical tests. JAMA 247, 2543–2546 (1982).
Vickers, A. J., Cronin, A. M., Elkin, E. B. & Gonen, M. Extensions to decision curve analysis, a novel method for evaluating diagnostic tests, prediction models and molecular markers. BMC Med. Inform. Decis. Mak. 8, 53 (2008).
Vickers, A. J. & Elkin, E. B. Decision curve analysis: a novel method for evaluating prediction models. Med. Decis. Making 26, 565–574 (2006).
Crombie, J. L. et al. Prognostic value of early PET in patients with aggressive non-Hodgkin lymphoma treated with anti-CD19 CAR T-cell therapy. Blood 138, 886 (2021).
Kuhnl, A. et al. Early FDG-PET response predicts CAR-T failure in large B-cell lymphoma. Blood Adv. 6, 321–326 (2022).
Oluwole, O. O. et al. Prophylactic corticosteroid use in patients receiving axicabtagene ciloleucel for large B-cell lymphoma. Br. J. Haematol. 194, 690–700 (2021).
Itacitinib pre-modulation in DLBCL receiving CAR T cell therapy. ClinicalTrials.gov identifier: NCT05757219. ClinicalTrials.gov https://clinicaltrials.gov/study/NCT05757219 (2024).
Rejeski, K. et al. The CAR-HEMATOTOX risk-stratifies patients for severe infections and disease progression after CD19 CAR-T in R/R LBCL. J. Immunother. Cancer 10, e004475 (2022).
Scholler, N. et al. Tumor immune contexture is a determinant of anti-CD19 CAR T cell efficacy in large B cell lymphoma. Nat. Med. 28, 1872–1882 (2022).
Strati, P. et al. Prognostic impact of corticosteroids on efficacy of chimeric antigen receptor T-cell therapy in large B-cell lymphoma. Blood 137, 3272–3276 (2021).
Jain, M. D. et al. Whole-genome sequencing reveals complex genomic features underlying anti-CD19 CAR T-cell treatment failures in lymphoma. Blood 140, 491–503 (2022).
Gu, Z., Eils, R. & Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32, 2847–2849 (2016).
Gu, Z., Gu, L., Eils, R., Schlesner, M. & Brors, B. circlize implements and enhances circular visualization in R. Bioinformatics 30, 2811–2812 (2014).
Harris, P. A. et al. The REDCap consortium: Building an international community of software platform partners. J. Biomed. Inform. 95, 103208 (2019).
Kedmi, M. et al. Point-of-care anti-CD19 CAR T-cells for treatment of relapsed and refractory aggressive B-cell lymphoma. Transplant. Cell Ther. 28, 251–257 (2022).
Cheson, B. D. et al. Recommendations for initial evaluation, staging, and response assessment of Hodgkin and non-Hodgkin lymphoma: the Lugano classification. J. Clin. Oncol. 32, 3059–3068 (2014).
Carbone, P. P., Kaplan, H. S., Musshoff, K., Smithers, D. W. & Tubiana, M. Report of the Committee on Hodgkin’s Disease Staging Classification. Cancer Res. 31, 1860–1861 (1971).
Lee, D. W. et al. ASTCT consensus grading for cytokine release syndrome and neurologic toxicity associated with immune effector cells. Biol. Blood Marrow Transplant. 25, 625–638 (2019).
Karvanen, J. The statistical basis of laboratory data normalization. Drug Inf. J. 37, 101–107 (2003).
Chuang-Stein, C. Some issues concerning the normalization of laboratory data based on reference ranges. Drug Inf. J. 35, 153–156 (2001).
Hubert, L. & Arabie, P. Comparing partitions. J. Classif. 2, 193–218 (1985).
Kanoun, S. et al. Influence of software tool and methodological aspects of total metabolic tumor volume calculation on baseline (18F)FDG PET to predict survival in Hodgkin lymphoma. PLoS ONE 10, e0140830 (2015).
Zwanenburg, A. et al. The Image Biomarker Standardization Initiative: standardized quantitative radiomics for high-throughput image-based phenotyping. Radiology 295, 328–338 (2020).
Leithner, D. et al. Conventional and novel ((18)F)FDG PET/CT features as predictors of CAR-T cell therapy outcome in large B-cell lymphoma. J. Hematol. Oncol. 17, 21 (2024).
Kumar, A. et al. Definition of bulky disease in early stage Hodgkin lymphoma in computed tomography era: prognostic significance of measurements in the coronal and transverse planes. Haematologica 101, 1237–1243 (2016).
Cox, D. R. Regression models and life-tables. J. R. Stat. Soc. 34, 187–202 (1972).
Hochberg, Y. & Benjamini, Y. More powerful procedures for multiple significance testing. Stat. Med. 9, 811–818 (1990).
Steyerberg, E. W., Borsboom, G. J., van Houwelingen, H. C., Eijkemans, M. J. & Habbema, J. D. Validation and updating of predictive logistic regression models: a study on sample size and shrinkage. Stat. Med. 23, 2567–2586 (2004).
Vickers, A. J., van Calster, B. & Steyerberg, E. W. A simple, step-by-step guide to interpreting decision curve analysis. Diagn. Progn. Res. 3, 18 (2019).
Epcoritamab compared to observation for treating B-cell lymphoma patients not in complete remission after CD19-directed CAR-T therapy. ClinicalTrials.gov identifier: NCT06238648. https://clinicaltrials.gov/study/NCT06238648 (2024).
Testing drug treatments after CAR T-cell therapy in Patients with relapsed/refractory diffuse large B-cell lymphoma. ClinicalTrials.gov identifier: NCT05633615. ClinicalTrials.gov https://clinicaltrials.gov/study/NCT05633615 (2024).
Iacoboni, G. et al. Treatment outcomes in patients with large B-cell lymphoma after progression to chimeric antigen receptor T-cell therapy. Hemasphere 8, e62 (2024).
Acknowledgements
The reported research was supported in part by the National Institutes of Health/National Cancer Institute (NIH/NCI) award number P01-CA023766 and MSK Support Grant (P30 CA008748). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. Editorial support in the preparation of the paper was provided by H. Rice at MSK. S.S.R. was supported by the American Society for Transplantation and Cellular Therapy (ASTCT) New Investigator Award, the Louis V. Gerstner, Jr. Physician Scholars Award and the Weill Cornell Medicine Clinical and Translational Science Center Grant 2UL1-TR-2384. R.S. received grant support from an NIH-NCI K08CA282987, the Long Island Sound Chapter, Swim Across America, the Robert Hirschhorn Award, Comedy vs. Cancer and the MSK Steven Greenberg Lymphoma Research. M.R.M.v.d.B. was supported by the NCI (R01-CA228358, R01-CA228308 and P01-CA023766); National Heart, Lung, and Blood Institute (NHLBI; R01-HL123340 and R01-HL147584); National Institute on Aging (NIA; P01-AG052359) and Tri-Institutional Stem Cell Initiative. Additional funding was received from the Lymphoma Foundation, the Susan and Peter Solomon Family Fund, the Solomon Microbiome Nutrition and Cancer Program, Cycle for Survival, Parker Institute for Cancer Immunotherapy, Paula and Rodger Riney Multiple Myeloma Research Initiative, Starr Cancer Consortium and Seres Therapeutics. J.U.P. was supported by NHLBI NIH Award K08HL143189. M.C. was supported by a grant from the Alfonso Martin Escudero Foundation.
Author information
Authors and Affiliations
Contributions
S.S.R. and R.S. defined the research goals and objectives. S.S.R., T. Fei, R.S., T. Funnell and A.J.V. developed the experimental and analytical methods. S.S.R., T. Fei and R.S. contributed to formal analysis. S.S.R., T. Fei and R.S. analyzed and interpreted data. J.A.F., L.A.L., A.A.T., M.C., I.D., Z.E.-P., I.L., L.P., N.O., I.S. and N.S.-T. collected and curated data. H.S., D.L., S.S.R., T. Fei and R.S. planned, acquired and interpreted PET-CT scans. S.S.R. deployed a website for InflaMix implementation. S.S.R., T. Fei and R.S. wrote the initial draft of the paper. All authors revised the paper critically for intellectual content. M.-A.P., M.R.M.v.d.B. and R.S. provided oversight and leadership in the planning and execution of research. A.P., A. Saldia and J.S. managed and coordinated research activities. M.-A.P., M.R.M.v.d.B., J.H.P., M.L.P. and R.S. secured financial support for the project. A.I., M.K., A. Shimoni, H.C.S., A.N. and A.A. designed and managed external or validation cohorts.
Corresponding author
Ethics declarations
Competing interests
R.S. reports speaker honorarium from Incyte. A.A. reports honoraria from AbbVie and has consulting and advisory roles with Takeda, Gilead, Novartis, Roche and Bristol Myers Squibb. L.A.L. served as a consultant and/or speaker bureau for Kite/Gilead, Beigene, Pharmacyclics, AbbVie, Genmab, SeaGen, Janssen, AstraZeneca, Eli Lilly, Epizyme, TG Therapeutics, Merck and ADC Therapeutics. A.I. owns stock and has ownership interests in Cota Healthcare. He also reports honoraria from MJH Life Sciences and Pfizer. He served as a consultant and/or speaker bureau for TG Therapeutics, Secura Bio, AstraZeneca and Seattle Genetics. Z.E.-P. serves on Genmab advisory board. J.U.P. reports research funding, intellectual property fees and travel reimbursement from Seres Therapeutics and consulting fees from Da Volterra, CSL Behring and MaaT Pharma. He serves on an advisory board of and holds equity in Postbiotics Plus Research. He has filed intellectual property applications related to the microbiome (reference numbers #62/843,849, #62/977,908 and #15/756,845). R.J.L. has served as a consultant for Kite. G.L.S. has received research funding from Janssen, Amgen, BMS, Beyond Spring, GPCR and Recordati and serves on the DSMB for Arcellx. M.S. served as a paid consultant for McKinsey & Company, Angiocrine Bioscience, Inc., and Omeros Corporation; received research funding from Angiocrine Bioscience, Inc., Omeros Corporation, Amgen Inc., Bristol Myers Squibb, and Sanofi; served on ad hoc advisory boards for Kite – A Gilead Company, and Miltenyi Biotec; and received honoraria from i3Health, Medscape, CancerNetwork, Intellisphere LLC, Curio Science LLC, and IDEOlogy. S.A.G. receives research funding from Amgen, Johnson & Johnson, Takeda, Celgene, Actinium, Sanofi, Miltenyi, Kite and EUSA. He is on the advisory boards of Amgen, Johnson & Johnson, Takeda, Celgene, Actinium, Sanofi, Miltenyi, Novartis, Kite, Jazz, BMS, Spectrum Pharma and EUSA Omeros. J.H.P. received consulting fees from Affyimmune Therapeutics, Amgen, Autolus, Be Biopharma, Beigene, Bright Pharmaceutical Services, Curocel, Kite, Medpace, Minerva Biotechnologies, Pfizer, Servier, Sobi and Takeda; received honoraria from OncLive, Physician Education Resource and MJH Life Sciences; serves on scientific advisory board of Allogene Therapeutics and Artiva Biotherapeutics; and received institutional research funding from Autolus, Genentech, Fate Therapeutics, Incyte, Servier and Takeda. M.L.P. has served as a consultant for Novartis, Cellectar, Synthekine, Kite, Seres, Magenta, WindMIL, Rheos, Nektar, Notch, Priothera, Ceramedix, Lygenesis and Pluto. G.S. has received in the last 12 months financial compensation for participating in advisory boards or consulting from AbbVie, Atbtherapeutics, Beigene, BMS/Celgene, Debiopharm, Genentech/Roche, Genmab, Incyte, Ipsen, Janssen, Kite/Gilead, Loxo/Lilly, Merck, Molecular Partners, Nordic Nanovector, Novartis, Nurix and Orna. He has also received research support managed by his institution from Genentech, Janssen and Ipsen. He is a shareholder of Owkin. B.D.G. has received honoraria for speaking engagements from Merck, Bristol Myers Squibb and Chugai Pharmaceuticals; has received research funding from Bristol Myers Squibb and Merck; and has been a compensated consultant for Darwin Health, Merck, PMV Pharma, Shennon Biotechnologies and Rome Therapeutics, of which he is a co-founder. He additionally has intellectual property rights with Rome Therapeutics and the Icahn School of Medicine at Mount Sinai. He has served in an advisory role at Merck Sharpe and Dohme and Darwin Health. M.-A.P. reports honoraria from Adicet, Allogene, Allovir, Caribou Biosciences, Celgene, Bristol Myers Squibb, Equilium, Exevir, ImmPACT Bio, Incyte, Karyopharm, Kite/Gilead, Merck, Miltenyi Biotec, MorphoSys, Nektar Therapeutics, Novartis, Omeros, OrcaBio, Syncopation, VectivBio AG and Vor Biopharma. He serves on DSMBs for Cidara Therapeutics, Medigene and Sellas Life Sciences and the scientific advisory board of NexImmune. He has ownership interests in NexImmune, Omeros and OrcaBio. He has received institutional research support for clinical trials from Allogene, Incyte, Kite/Gilead, Miltenyi Biotec, Nektar Therapeutics, and Novartis. M.R.M.v.d.B. has received research support and stock options from Seres Therapeutics and stock options from Notch Therapeutics and Pluto Therapeutics; he has received royalties from Wolters Kluwer; has consulted, received honorarium from or participated in advisory boards for Seres Therapeutics, Vor Biopharma, Rheos Medicines, Frazier Healthcare Partners, Nektar Therapeutics, Notch Therapeutics, Ceramedix, Lygenesis, Pluto Therapeutics, GlaxoSmithKline, Da Volterra, Thymofox, Garuda, Novartis (spouse), Synthekine (spouse), Beigene (spouse) and Kite (spouse); he has IP licensing with Seres Therapeutics and Juno Therapeutics; and holds a fiduciary role on the Foundation Board of DKMS (a nonprofit organization). Memorial Sloan Kettering Cancer Center has institutional financial interests relative to Seres Therapeutics. The other authors declare no competing interests. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Peer review
Peer review information
Nature Medicine thanks Jordan Gauthier, Lana Garmire, Sattva Neelapu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Ulrike Harjes, in collaboration with the Nature Medicine team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Workflow visual abstract.
(a) Our workflow involved 1) using 14 preinfusion laboratory measures in a model derivation cohort from MSK to build a Gaussian mixture model we named InflaMix. InflaMix clustered patients into two groups, one with an inflammatory blood profile and another with a noninflammatory profile. 2) We then evaluated the association between inflammatory cluster assignment by InflaMix and clinical outcomes after CAR-T therapy. 3). Finally, we then used InflaMix to predict patient cluster assignment in three independent validation cohorts (Cohort 2–MSK patients with LBCL, Cohort 3–SMC and HMH patients with LBCL, and Cohort 4–patients from all 3 centers with either FL or MCL) and evaluate their associations with clinical outcomes. (b) In developing InflaMix from the derivation cohort (Cohort I), we 1) normalized every laboratory value by their upper limit of normal to standardize measurements across different assays. 2) We then systemically log transformed any laboratory measures with distribution skew > 1 that improved > 90% by log transformation. 3) Laboratory values for patients across all cohorts were then scaled by the mean and standard deviation of laboratory values in the derivation cohort. These normalized, log-transformed, and scaled values are then used for cluster assignment by mixture modeling. These cluster assignments were then used as predictors in fitted regression models of disease response and survival to evaluate their associations with clinical outcomes. FL, follicular lymphoma; HMH, Hackensack Meridian Health; MCL, mantle cell lymphoma; MSK, Memorial Sloan Kettering Cancer Center; SMC, Sheba Medical Center.
Extended Data Fig. 2 Consort diagram.
Multicenter observational study with 4 independent cohorts. A Gaussian mixture model of 14 preinfusion laboratory and cytokine assays (InflaMix) defined a cluster enriched for patients with elevated inflammatory markers in the MSK model derivation cohort (Cohort I). This “inflammatory cluster” reproducibly associated with and was predictive of poor clinical outcomes across all 4 cohorts. Three independent validation cohorts included: 1) MSK LBCL cohort (Cohort II); 2) SMC + HMH LBCL cohort (Cohort III); 3) MCL and FL cohort (Cohort IV, all centers). FL, follicular lymphoma; HMH, John Theurer Cancer Center of Hackensack Meridian Health; lisocel, lisocabtagene maraleucel; MCL, mantle cell lymphoma; MSK, Memorial Sloan Kettering Cancer Center; PMBCL, primary mediastinal large B cell lymphoma; SMC, Sheba Medical Center.
Extended Data Fig. 3 InflaMix is a 2-cluster Gaussian mixture model of pre-CAR-T laboratory and cytokine measurements that jointly considers covariance across lab features and optimizes cluster separation and entropy.
(a) Integrated complete likelihood criteria of various Gaussian mixture models with varying numbers of clusters built from pre-CAR-T labs in the derivation cohort. Each 3-letter combination (for example, VVI) represents a different parameterization approach described in Scrucca et. al30. InflaMix is a VVV model, which means that different means, variances, and covariances can be estimated for each multivariate mixture (cluster) distribution. b-g, Comparing lab measures between inflammatory (n = 39) and noninflammatory (n = 110) clusters in the derivation cohort. Inferences by FDR-corrected Wilcoxon tests for (b) IL-6, (c) CRP, (d) LDH, (e) Hgb, (f) WBC, and (g) Tbili. Boxplots depict the median bounded by the 1st and 3rd quartile values. Boxplot whiskers depict 1.5 times the IQR beyond the boxplot hinges. All tests were 2-sided with a significance level of 0.05. InflaMix cluster assignments viewed through AST and CRP dimensions in the derivation cohort when (h) all 14 lab features are used to generate the model and (i) lab features with lowest variable importance (WBC, Plt, Tbili) by random forest prediction of cluster assignment are removed from model generation. Hgb, hemoglobin; Plt, platelets; Tbili, total bilirubin.
Extended Data Fig. 4 InflaMix reliably estimates cluster assignment probabilities.
InflaMix performance in consistent cluster assignment was benchmarked against mixture model variants derived from bootstrapped populations from the (a-c) derivation cohort (Cohort I) and an (d-f) independent Cluster Assignment Validation (CAV) cohort of patients with NHL. Calibrations by linear regression fits of inflammatory cluster assignment (ICA) probabilities by InflaMix to averaged ICA probabilities conferred by all mixture model variants for a given patient are plotted when InflaMix predictions are made with (a, d) no missing laboratory values, (b, e) up to 7 missing laboratory values (simulated over multiple iterations), and (c, f) only the limited 6-lab InflaMix panel (albumin, Hgb, AST, alkaline phosphatase, LDH, and CRP). 95% confidence intervals are provided for the regression fits. The black lines represent ideal calibration, and the gray shaded boxes overlap with concordant cluster assignments. Both Pearson correlation coefficients and Lin’s CCC31,32 are reported. Intercepts and slopes for the least squares regression line fits, as well as the adjusted Rand indices55 for cluster assignment agreement are provided in Supplementary Table 1. AST, aspartate aminotransferase; CCC, Concordance correlation coefficient; CRP, C-reactive protein; Hgb, hemoglobin; LDH, lactate dehydrogenase; NHL, non-Hodgkin lymphoma; PCC, Pearson correlation coefficient.
Extended Data Fig. 5 InflaMix cluster assignment identifies similar laboratory profiles across 4 independent cohorts.
Heatmaps of normalized preinfusion laboratory values in (a) the model derivation cohort of patients with LBCL treated at MSK (Cohort I), (b) the MSK LBCL validation cohort (Cohort II), (c) the SMC + HMH LBCL validation cohort (Cohort III), and (d) the MCL and FL validation cohort (Cohort IV; all centers) scaled by distributions of ULN-normalized preinfusion lab values in the model derivation cohort46,47. Patients (columns) are ordered by probability of cluster assignment. Flanking heatmaps are colored by median scaled lab values in each cluster, including unscaled medians with IQR. Units of measure: albumin, Hgb (g/dL); ALP, AST, LDH (U/L); CRP, Tbili (mg/dL); D-dimer (mcg/mL); ferritin (ng/mL); IL-6, IL-10, TNFα (pg/mL); Plt, WBC (K/mcL). ALP, alkaline phosphatase; FL, follicular lymphoma; g, grams; Hgb, hemoglobin; HMH, John Theurer Cancer Center of Hackensack Meridian Health; K, 1000 cells; mcL, microliter; MCL, mantle cell lymphoma; mg, milligram; MSK, Memorial Sloan Kettering Cancer Center; ng, nanograms; NHL, non-Hodgkin lymphoma; pg, picogram; Plt, platelets; SMC, Sheba Medical Center; Tbili, total bilirubin; U/L, units per liter; ULN, upper limit of normal.
Extended Data Fig. 6 InflaMix-assigned clusters associate with clinical outcomes independently of tumor burden.
Kaplan-Meier survival estimates of PFS and OS, and rates of CR by day 100 by InflaMix clustering across all patients across all patients with LBCL at MSK who had PET radiomic assessments with either high or low tumor burden by MTV. Odds ratios of no CR by day 100 and hazard ratios estimated with 95% CI using regression models adjusted for age, primary refractory disease, and costimulatory domain. Estimates for (a) PFS, (b) OS by day 100, and (c) rates of CR by day 100 in patients with MTV greater than upper tercile MTV value (83.45 mm3); and (d) PFS, (e) OS, (f) rates of CR by day 100 in patients with MTV lower than the upper tercile value. In a Cox proportional hazards regression model adjusted for age, primary refractory disease, costimulatory domain, MTV as a continuous variable, and an interaction term between MTV and inflammatory cluster assignment, inflammatory clustering was still significantly associated with outcomes (Supplementary Table 2). Significance of cluster associations with clinical outcomes was determined by the Wald test. All tests were 2-sided with a significance level of 0.05. Adj. adjusted; CI, confidence interval; CR, complete response; FL, follicular lymphoma; HR, hazard ratio; Infl., Inflammatory Cluster; MTV, metabolic tumor volume; Non-Infl., Non-Inflammatory Cluster; PET, positron emission tomography; PFS, progression-free survival; OR, odds ratio; OS, overall survival; ULN, upper limit of normal.
Extended Data Fig. 7 InflaMix-informed prediction models for PFS at 6 months outperform models trained with conventional biomarkers or alternative mixture models trained without unconventional cytokine measurements.
The InflaMix model uses base clinical features (age, costimulatory domain, primary refractory disease, elevated prelymphodepletion LDH) and InflaMix score (log-transformed cluster assignment probability). Conventional model benchmarks include: Base - base clinical features only, CRP–base clinical features and prelymphodepletion CRP, NoCytoMM–base clinical features and log-transformed probability of inflammatory clustering assigned by an alternative Gaussian mixture model trained without IL-6, IL-10, or TNFα, Lab11Reg–Regularized regression model of base clinical features and all 11 analytes used to develop the alternative Gaussian mixture model. All models were trained using Cox proportional hazards regression. Prediction performance was assessed using an independent validation cohort of patients with LBCL. For each set of model comparisons, the validation cohort was divided into a group used to recalibrate the original model and an independent test group, repeated 100 times with 2-fold cross-validation for an unbiased assessment. InflaMix-informed prediction of PFS at 6 months conferred a significantly improved AUROC (Wald test, p < 0.01) over all alternative models: (InflaMix 0.72, NoCytoMM 0.62, Lab11Reg 0.61 and InflaMix 0.74, NoCytoMM 0.68, CRP 0.67, Base 0.68). Calibration curves, density plots, and net benefit here are evaluated using risk estimates aggregated across all repeated validation folds. A positive event here is defined as disease progression, relapse, or death by 6 months. (a, b) Calibration curves of InflaMix-informed models compared to those of conventionally trained models ((a) Lab11Reg and NoCytoMM (b) Base, NoCytoMM, and CRP) (c, d) Decision curve analyses comparing net benefit conferred by InflaMix-informed models against conventional models ((c) Lab11Reg and NoCytoMM, (d) Base, NoCytoMM, and CRP) in patients who obtain a PR by day +30 after CAR-T infusion. The net benefit is evaluated for consolidation therapies across relatively low (20-30%) and high (30-40%) probability threshold ranges. Auto-HCT, autologous hematopoietic cell transplantation; PFS, progression-free survival; PR, partial response.
Extended Data Fig. 8 InflaMix-assigned clusters associate with clinical outcomes agnostic of CAR-T product used.
Kaplan-Meier survival estimates of PFS and OS, and rates of CR by day 100 by InflaMix clustering across all patients in the study (all cohorts) treated by either CD28- or 41BB-costimulatory domain CAR-T products. Odds ratios of no CR by day 100 and hazard ratios estimated with 95% CI using regression models adjusted for age, primary refractory disease, costimulatory domain, prelymphodepletion LDH elevated above ULN, disease (MCL, FL, or LBCL), and CAR-T product. Estimates for (a) PFS, (b) OS, and (c) rates of CR by day 100 in patients treated with CD28-costimulatory domain products; and (d) PFS, (e) OS, and (f) rates of CR by day 100 in patients treated with 41BB-costimulatory domain products. Significance of cluster associations with clinical outcomes was determined by the Wald test. All tests were 2-sided with a significance level of 0.05. Adj. adjusted; CI, confidence interval; CR, complete response; FL, follicular lymphoma; HR, hazard ratio; Infl., Inflammatory Cluster; MCL, mantle cell lymphoma; Non-Infl., Non-Inflammatory Cluster; PFS, progression-free survival; OR, odds ratio; OS, overall survival; ULN, upper limit of normal.
Supplementary information
Supplementary Information (download PDF )
Supplementary Notes, Tables 1 and 2, Figs. 1–8, references and TRIPOD Checklist.
Source data
Source Data Fig. 1 (download CSV )
Complete laboratory data needed for InflaMix model derivation from the model development cohort.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Raj, S.S., Fei, T., Fried, S. et al. An inflammatory biomarker signature of response to CAR-T cell therapy in non-Hodgkin lymphoma. Nat Med 31, 1183–1194 (2025). https://doi.org/10.1038/s41591-025-03532-x
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41591-025-03532-x
This article is cited by
-
Late hematologic toxicity after CAR T-cell therapy in large B-cell lymphoma: incidence, risk factors, and clinical impact
Bone Marrow Transplantation (2026)
-
Blood proteomics for quantitative biomarkers of cellular therapies
Biomarker Research (2025)
-
Optimization and validation of the international metabolic prognostic index for CD19 CAR-T in large B-cell lymphoma
Blood Cancer Journal (2025)