Abstract
The field of computational pathology has been transformed with recent advances in foundation models that encode histopathology region-of-interests (ROIs) into versatile and transferable feature representations via self-supervised learning. However, translating these advancements to address complex clinical challenges at the patient and slide level remains constrained by limited clinical data in disease-specific cohorts, especially for rare clinical conditions. We propose Transformer-based pathology Image and Text Alignment Network (TITAN), a multimodal whole-slide foundation model pretrained using 335,645 whole-slide images via visual self-supervised learning and vision-language alignment with corresponding pathology reports and 423,122 synthetic captions generated from a multimodal generative AI copilot for pathology. Without any fine-tuning or requiring clinical labels, TITAN can extract general-purpose slide representations and generate pathology reports that generalize to resource-limited clinical scenarios such as rare disease retrieval and cancer prognosis. We evaluate TITAN on diverse clinical tasks and find that it outperforms both ROI and slide foundation models across machine learning settings, including linear probing, few-shot and zero-shot classification, rare cancer retrieval, cross-modal retrieval and pathology report generation.
Similar content being viewed by others
Main
Foundation models are transforming computational pathology by accelerating the development of AI tools for diagnosis, prognosis and biomarker prediction from digitized tissue sections1. Developed using self-supervised learning (SSL) on millions of histology image patches (or regions of interests), these models capture morphological patterns in histology patch embeddings, such as tissue organization and cellular structure2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17. These representations serve as a ‘foundation’ for models that predict clinical endpoints from whole-slide images (WSIs), such as diagnosis or biomarker status18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38. However, translating the capabilities of current patch-based foundation models to address patient- and slide-level clinical challenges still remains complex due to the immense scale of gigapixel WSIs, compounded by the small size of patient cohorts in real-world evidence39,40,41,42, especially for rare diseases with limited training data43,44,45. Similarly, given a diagnostically challenging WSI, retrieving a similar WSI via slide search5,46,47,48,49,50,51,52,53 or pathology reports through cross-modal report search10,54,55,56 typically requires specialized algorithms to bridge the gap between fine-grained patch embeddings and slide-level information, introducing hurdles to clinical adoption.
To overcome these limitations, new types of foundation models have recently been proposed for encoding entire WSIs into slide-level general-purpose feature representations57,58,59,60,61,62,63,64,65,66,67,68,69,70,71,72. Instead of training an additional model on top of patch embeddings from scratch34,73,74,75,76,77,78,79,80, these whole-slide representation models can be pretrained to distill pathology-specific knowledge from large WSI collections, simplifying clinical endpoint prediction with their off-the-shelf application. The outstanding challenge then becomes developing whole-slide foundation models that faithfully encode the tissue microenvironment based on a set of patch embeddings while also handling arbitrarily large WSIs. Although relatively underexplored, slide-level self-supervision can be performed with vision-only pretraining, either through masked image reconstruction58 or intraslide contrastive learning59,60,81, or in a multimodal fashion involving pathology reports, bulk transcriptomics, or immunohistochemistry (IHC)61,62,63,64,66,67,82. Furthermore, long-range context modeling can either be neglected, essentially treating a WSI as a bag of independent features59,62,63,64,72,83, or explicitly modeled using Transformers57,58,60,61. With efforts to learn general-purpose slide representations intensifying, we believe that adapting successful patch-level recipes to the entire WSI would lead to powerful general-purpose slide representations.
Despite their widespread application potential, previous works on pretraining slide foundation models have several shortcomings. First, these models are predominantly pretrained using vision-only modeling57,59,60, which neglects not only rich supervisory signals found in pathology reports but also precludes multimodal capabilities such as zero-shot visual-language understanding and cross-model retrieval—a fundamental hallmark in foundation models84,85. Second, whereas current patch foundation models are trained with millions of histology image patches, slide foundation models are developed with orders of magnitude fewer samples and limited optimization of SSL recipes, leading to slide representations with restricted generalization capability58,62,82,83. Even with multimodal techniques such as vision-language pretraining that augment the pretraining dataset with pathology reports, current slide foundation models still require end-to-end training or fine-tuning and lack the capability of learning transferable slide representations for challenging clinical scenarios58,82,83. Finally, the current models are nascent in transforming pathology AI model development due to their limited evaluations in diagnostically relevant settings, such as few-shot learning or slide retrieval.
Here, we introduce Transformer-based pathology Image and Text Alignment Network (TITAN), a multimodal whole-slide vision-language model designed for general-purpose slide representation learning in histopathology. Building on the success of knowledge distillation and masked image modeling86,87 for patch encoder pretraining21,22, TITAN introduces a large-scale pretraining paradigm that leverages millions of high-resolution region-of-interests (ROIs; at 8,192 × 8,192 pixels at 20× magnification) for scalable WSI encoding. Trained using 336k WSIs across 20 organ types, vision-only TITAN produces general-purpose slide representations that can readily be applied to slide-level tasks such as cancer subtyping, biomarker prediction, outcome prognosis and slide retrieval tasks, outperforming supervised baselines and existing multimodal slide foundation models. To augment TITAN with language capabilities, we further fine-tune it by contrasting with 423k synthetic fine-grained ROI captions generated using PathChat88, a multimodal generative AI copilot for pathology and with 183k pathology reports at the slide level. By leveraging free-text morphological descriptions, TITAN gains the ability to generate pathology reports, perform zero-shot classification and enable cross-modal retrieval between histology slides and clinical reports. Pretraining TITAN on an extensive repository of multimodal pathology data unlocks higher levels of performance compared to existing slide foundation models, particularly in low-data regimes, language-guided zero-shot classification and rare cancer retrieval. Additionally, we demonstrate the utility of pretraining with synthetic fine-grained morphological descriptions, suggesting the scaling potential of TITAN pretraining with synthetic data89,90,91. Through comprehensive evaluation across a large range of clinical tasks, including the application to rare cancer retrieval, we demonstrate the efficacy of our vision-language pretraining approach, showcasing the general-purpose capability of our slide foundation model.
Results
Scaling SSL from histology patches to whole-slide images (WSIs)
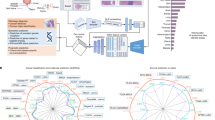
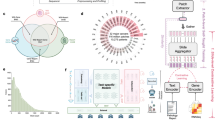
TITAN is a Vision Transformer (ViT)92 that creates a general-purpose slide representation readily deployable in diverse clinical settings. It is pretrained on an internal dataset (termed Mass-340K) consisting of 335,645 WSIs and 182,862 medical reports (Fig. 1a). To ensure the diversity of the pretraining dataset, which has proven to be a key factor in successful patch encoders21, Mass-340K is distributed across 20 organs, different stains, diverse tissue types and scanned with various scanner types (Fig. 1a and Supplementary Table 1). The pretraining strategy consists of three distinct stages to ensure that the resulting slide-level representations capture histomorphological semantics both at the ROI-level (4 × 4 mm2) and at the WSI-level with the help of visual and language supervisory signals—stage 1, vision-only unimodal pretraining with Mass-340K on ROI crops (Fig. 1b,c), stage 2, cross-modal alignment of generated morphological descriptions at ROI-level (423k pairs of 8k × 8k ROIs and captions) and stage 3, cross-modal alignment at WSI-level (183k pairs of WSIs and clinical reports; Fig. 1d; see Methods for more details). For ease of notation, we refer to the model pretrained with vision-only in stage 1 as TITANV and to the full model after all three stages of pretraining as TITAN.
a, Tissue site distribution of Mass-340K used for TITANV pretraining (stage 1). Mass-340K includes 335,645 WSIs across 20 organs with a mix of tissue sections stained with H&E (89.7%), IHC (7.9%), special stains (2.3%) and others (0.1%) or a mix of neoplastic (70.0%), tissue damage response (8.4%), normal (4.7%), inflammatory (3.4%) and others (13.5%) scanned with diverse scanner types. TITAN pretraining (stages 2 and 3) uses a subset of Mass-340K with paired captions and medical reports. b–d, Block diagram of TITANV pretraining. b, TITAN uses a ViT to encode a WSI into a slide embedding. c, TITANV (stage 1) is pretrained using SSL with student–teacher knowledge distillation. d, TITAN (stage 2 and 3) is pretrained using vision-language modeling, first by aligning the slide embedding with synthetic captions (stage 2) and then with medical reports (stage 3). e, UMAP visualization of TCGA slide embeddings obtained with TITAN, color-coded by organ. UMAP, uniform manifold approximation and projection; px, pixel.
The cornerstone of our approach is emulating the patch encoder designed for input patch images at the slide level. Instead of using tokens from a partitioned image patch, the slide encoder takes a sequence of patch features encoded by powerful histology patch encoders4,7,8,9,10,11,12,13,14,58. Consequently, all of TITAN pretraining stages occur in the embedding space based on pre-extracted patch features, with the patch encoder assuming the role of the ‘patch embedding layer’ in a conventional ViT (Fig. 1b). To preserve the spatial context of each patch and consequently enable the use of positional encoding in the embedding space, the patch features are spatially arranged in a two-dimansional (2D) feature grid replicating the positions of the corresponding patches within the tissue (Fig. 1c). Following the success of masked image modeling and knowledge distillation in patch encoders21, we apply the iBOT86 framework for vision-only pretraining of TITAN on the 2D feature grid.
While the conceptual transition to slide-level is simple, this presents a new set of model design and pretraining challenges as follows: (1) handling long and variable input sequences (>104 tokens at slide-level versus 196 to 256 tokens at the patch-level), (2) creating multiple views of one sample for SSL and (3) ambiguity over positional encoding schemes that capture local and global context in the tissue microenvironment. First, to tame the computational complexity caused by long input sequences, we construct the input embedding space by dividing each WSI into nonoverlapping patches of 512 × 512 pixels (instead of widely used 256 × 256 pixels) at ×20 magnification, followed by the extraction of 768-dimensional features for each patch with CONCHv1.5, the extended version of CONCH10. To address the issue of large and irregularly shaped WSIs, we create views of a WSI by randomly cropping the 2D feature grid (Fig. 1c). Specifically, a region crop of 16 × 16 features covering a region of 8,192 × 8,192 pixels is randomly sampled from the WSI feature grid. From this region crop, two random global (14 × 14) and ten local (6 × 6) crops are sampled for iBOT pretraining. We further augment these feature crops with vertical and horizontal flipping, followed by posterization feature augmentation93. Finally, to ensure that the limited context pretraining translates to slide-level tasks, we use attention with linear bias (ALiBi) for long-context extrapolation of TITAN at inference time94. Originally proposed for long-context inference in large language models, we extended ALiBi to 2D, where the linear bias is based on the relative Euclidean distance between features in the feature grid, which reflects the actual distances between patches in the tissue (Supplementary Tables 2 and 3; see Methods for more details).
To equip our model with language capabilities, we implement two additional multimodal and multiscale pretraining strategies (stages 2 and 3) using a subset of Mass-340K (Fig. 1d). These stages leverage language descriptions that exist at multiple morphological scales, from fine-grained descriptions in pathologist annotations or textbooks at the patch- or region-level (stage 2) to high-level descriptions in pathology reports at the slide-level (stage 3). For both stages, we use contrastive captioners (CoCa)95 as the pretraining strategy that aligns ROI and slide representations with the corresponding captions and reports, while generating accurate descriptions at ROI-level or reports at slide level, respectively. The slide encoder (weights initialized with TITANV), the text encoder and the multimodal decoder are all finetuned as part of the pretraining. In stage 2, we pretrain TITANV with 423,122 pairs of 8,192 × 8,192 pixels ROIs and synthetic captions generated by the vision-language copilot PathChat88. In stage 3, we further pretrain the model with 182,862 pairs of WSIs and corresponding pathology reports, resulting in our final model TITAN (see Methods for more details; Supplementary Tables 4–10).
TITAN improves region and slide-level diagnostic capabilities
We evaluate TITAN, TITANV and existing slide encoders on a large set of diverse slide-level tasks, including morphological subtyping and molecular classification by linear probing on the frozen slide embeddings. For tasks with multiple cohorts available, we perform cross-validation on one cohort, for example, from The Cancer Genome Atlas (TCGA)96,97, and use the remaining cohorts, for example, from Clinical Proteomic Tumor Analysis Consortium (CPTAC)98,99 or Dartmouth-Hitchcock Medical Center (DHMC)100,101, as an external test cohort. As baselines, we compare to recent publicly available slide foundation models, PRISM62, GigaPath63 and CHIEF83. These models employ different slide-level pretraining strategies (PRISM, WSI-report contrastive pretraining; GigaPath, masked image reconstruction pretraining; CHIEF, supervised contrastive learning of cancerous versus noncancerous WSIs), different patch-level encoders (PRISM and GigaPath, 256 × 256 pixels at ×20 magnification; CHIEF, 256 × 256 pixels at ×10 magnification) and a varying number of WSIs for pretraining (PRISM, 1.7×; GigaPath, 0.49×; CHIEF, 0.18× the WSIs used for TITAN pretraining). Except for CHIEF, the pretraining datasets of TITAN (Mass-340K), PRISM and GigaPath do not include TCGA and PANDA, which allows us to use these two datasets as benchmarking tasks without concern for data leakage102. Additionally, we compare our approach with mean pooling using the same CONCHv1.5 patch encoder as TITAN, a simple yet powerful unsupervised slide representation framework65,66,103.
Furthermore, for a comprehensive evaluation of the baselines, we introduce two tumor classification tasks based on the publicly available repository TCGA with the following two different context lengths: (1) main cancer type classification on ROIs (TCGA-Uniform-Tumor-8K or TCGA-UT-8K), a ROI-level cancer subtyping task with 32 classes, where we manually curated 25,495 tumor-containing regions of 8,192 × 8,192 pixels at ×20 magnification (~4 × 4 mm2) across TCGA, covering the same tissue context as the region crops in TITANV pretraining (Extended Data Fig. 1 and Supplementary Table 11) and (2) a slide-level pan-cancer classification (TCGA-OncoTree or TCGA-OT) task of OncoTree codes104 with 46 classes, consisting of 11,186 formalin-fixed paraffin-embedded (FFPE) WSIs from TCGA (Supplementary Table 12; see Methods for more details).
We first assess how the pretraining data scale affects the downstream performance of TITANV, focusing on the four subtyping tasks—TCGA-UT-8K, TCGA-OT, OT108 and EBRAINS. The purpose of these multiclass classification tasks is to assess the generalizability and richness of feature representations across diverse diagnostic classes. We observe that the performance increases on all four tasks as more pretraining data is used, where TITANV with full Mass-340K exhibits an average increase of 3.65%, 3.21% and 1.21%, compared to 12.5%, 25% and 50%, respectively, of Mass-340K, where the same distribution across the organs was maintained (Fig. 2a and Supplementary Tables 13–16). Despite the difference in pretraining recipes, we observe the same general trend for the three other slide encoders, where PRISM outperforms GigaPath and CHIEF by 9.01% and 20.1% on average, having 3.4 times and 9.7 times the number of pretraining WSIs, respectively. Furthermore, we observe that TITAN and TITANV, with 48.5 million and 42.1 million parameters, outperform heavier slide encoders PRISM and GigaPath, with 99.0 million and 86.3 million parameters, demonstrating superior parameter efficiency of our model (Fig. 2b).
a, Impact of pretraining data size on TITANV and baselines across four challenging subtyping tasks. TITANV is pretrained with 12.5%, 25%, 50% and 100% of Mass-340K. b, The average performance of the four tasks against the number of parameters. c, Linear probe evaluation of TITAN and baselines on morphological classification, molecular status and survival prediction tasks. The mean pooling baseline uses the same patch encoder as TITAN (CONCHv1.5). Multiclass tasks are evaluated with balanced accuracy, binary tasks with AUROC and survival tasks with the concordance index. For external cohorts (DHMC, CPTAC), the classifier is trained on the corresponding TCGA cohort. All error bars represent s.d. based on bootstrapping (n = 1,000) or k-fold evaluation (k = 5). d, Ablation for positional encoding, number of transformer layers and inclusion of vision-pretraining stage. The performance is averaged across the four subtyping tasks. e, Change in performance of slide encoders averaged across the four subtyping tasks for different learning paradigms. For mean pooling and ABMIL, the respective patch encoder for each framework is used. PRISM fine-tuning is not evaluated as the fine-tuning recipes are not provided. f, Linear probe few-shot performance using K shots, K ∈ {1, 2, 4, 8, 16}, comparing baselines and ABMIL with CONCHv1.5. For each setting, 50 runs were performed. The center of each box plot (horizontal line) represents the median, with whiskers extending to data points within 1.5× the interquartile range. Statistical significance was assessed by fitting generalized linear mixed-effects model and two-sided Wald z test on the fitted model. Significance shown with respect to TITAN. P values for nonsignificant results are shown. **P ≤ 0.01, ***P ≤ 0.001, ****P ≤ 0.0001. C, number of classes; Ft., fine-tune.
We next evaluate TITAN on a range of clinically relevant tasks that span morphological classification (14 tasks), grading (3 tasks), molecular classification (39 tasks) and survival prediction (6 tasks; Supplementary Tables 17–21). On average, we observe that TITAN and TITANV outperform other slide encoders (Fig. 2c), demonstrating the superior slide representation quality of our models. In particular, TITAN significantly outperforms all existing slide encoders in morphological subtyping tasks across the entire spectrum of diagnostic complexities, including fine-grade pan-cancer classification (challenging morphological classification tasks, as shown in Fig. 2c) and noncancerous tasks, such as cardiac allograft assessment (cellular-mediated rejection) and renal allograft assessment (antibody and cellular-mediated rejection). TITAN and TITANV achieve an average of +8.4% and 6.7%, respectively, in performance on multiclass (balanced accuracy) and binary subtyping tasks (area under the receiver operator curve (AUROC)) over the next best-performing model, PRISM (Fig. 2c and Supplementary Tables 22–33). In particular, TITANV (and TITAN) not only outperforms others on TCGA-UT-8K with 8k × 8k context that the model was trained on (+6% and 7.5% over PRISM) but also on WSI-level tasks that involve the entire tissue context, where TITANV benefits from the long-context extrapolation via ALiBi, for example, TCGA-OT (+7% and 9.5% over PRISM), OT108 (+10% and 16% over PRISM) and EBRAINS (+9% and 9.1% over PRISM). Even with other nonparametric evaluations with prototyping105,106 and 20 nearest-neighbor evaluation, which predicts each WSI’s label based on the proximity to other WSI embeddings in the embedding space, we observe that TITAN and TITANV maintain superior performance Supplementary Tables 22–33. On grading tasks, TITAN outperforms the next best models CHIEF on average by +3.2% and PRISM by +4% in quadratic-weighted Cohen’s κ, where the high performance of CHIEF can be attributed to including the dataset PANDA in pretraining (Supplementary Tables 34–36). To evaluate the molecular classification performance, we tested the model on tasks from public datasets (BCNB and MUT-HET) and internal–external paired public datasets (TCGA, CPTAC and EBRAINS), on IHC tasks, and MGB internal molecular tasks (Fig. 2c, Extended Data Fig. 2 and Supplementary Tables 37–63). Averaged across all molecular tasks, TITAN significantly outperforms its mean baseline on CONCHv1.5 features, GigaPath and CHIEF (P < 0.0001).
On survival prediction tasks, we observe that TITAN and TITANV are generally the best-performing baselines, outperforming the next best-performing model CHIEF by +3.62% and +2.90%, respectively, on concordance index for disease-specific survival97 although CHIEF was pretrained on TCGA slides (Supplementary Table 64). Interestingly, the mean pooling baseline shows competitive performance, suggesting that the proportion of different morphological phenotypes is an important prognostic factor65,103.
To further understand how the slide embedding space is organized and consequently affects the downstream performance, we visualize UMAP embeddings of WSIs from TCGA-OT colored by organ type, showing that TITAN and TITANV form distinct organ clusters (for example, breast further separated from bladder, stomach and lung) better separated than with other slide encoders (Fig. 1e and Extended Data Fig. 3). To investigate the robustness of TITAN to nonmorphology-related effects (that is, batch effects), we evaluate how well slide representations from TCGA-OT cluster according to tumor type, organ and TCGA submission site. Both TITAN and TITANV mix submission sites well, while performing best in clustering them by biological factors, which suggests superior generalization capabilities (Extended Data Fig. 4). For interpretability analysis, we visualize attention heatmaps derived from the Transformer attention heads. The heatmaps indicate that different heads focus on distinct morphological regions such as dense tumor, tumor-adjacent stroma and nontumor regions, with majority of the heads focusing on dense tumor, consistently across multiple inputs (Extended Data Fig. 5).
To assess the calibration and the confidence of predictions from TITAN, we implement the expected calibration error (ECE)107 and entropy-based confidence score, and average both metrics across the four challenging subtyping tasks. Again, we observe that TITAN and TITANV achieve the best calibration and the highest confidence prediction (Supplementary Table 65).
Finally, to better understand how our model choices affect the downstream performance, we perform ablation experiments on the following four design choices of TITAN: the positional encoding, the number of transformer layers in TITANV, the inclusion of vision pretraining and the region size in vision-only pretraining (Fig. 2d and Supplementary Tables 66–77; see Methods for more details). Averaged across the four challenging subtyping tasks, our results demonstrate that ALiBi positional encoding outperforms original absolute positional encoding92 by +1.89%, six transformer layers provide best performance compared to 12 layers (+1.16%) and 4 layers (+1.74%), vision pretraining improves results by +2% over vision-language alignment alone, and a region size of 8,1922 achieves the best balance between performance (+3.6% over smaller regions of 4,0962) and computational efficiency.
Comparison with different learning paradigms for slide encoding
To further assess the quality of the slide embeddings and how application settings affect downstream performance, we evaluate different learning paradigms by comparing the linear probe performance of each slide encoder against other MIL models comprised of mean pooling, that is, averaging the patch embeddings, attention-based MIL (ABMIL)73 and task-specific fine-tuning of the slide encoder from random or pretrained weights. For mean pooling and ABMIL, we use respective patch encoders for each slide encoder. This allows us to gauge whether the pretrained slide encoders have learned meaningful slide representations and outperform the simple yet powerful unsupervised (mean pooling) and supervised (ABMIL) baselines, neither of which involves large-scale pretraining.
We observe several trends with TITAN (Fig. 2e, Extended Data Fig. 6 and Supplementary Tables 78–81). First, ABMIL outperforms mean pooling, as expected, since ABMIL is supervised and equivalent to weighted averaging of the patch features. Next, the linear probe outperforms ABMIL, demonstrating that multimodal self-supervised pretraining of TITAN and TITANV effectively captures the contextual and semantic morphological details of the slide. This further suggests that our task-agnostic slide embeddings are better equipped for downstream tasks than task-specific supervised slide embeddings. Finally, we observe that task-specific fine-tuning of TITAN leads mostly to performance improvement over linear probe of TITAN and TITANV while fine-tuning the slide encoder from randomly initialized weights yields lower performance (-3.63% on average). This suggests that the pretrained weights of TITANV can serve as a good initialization for task-specific training, in line with previous works62,64. One exception is OT108, which could be attributed to the small number of samples for each class (ranging from 4 to 42), which may lead to overfitting. However, we observe that other slide encoders do not necessarily follow such important trends, possibly suggesting suboptimal model pretraining and lack of generalizability.
Few-shot learning for low-data regime
We also evaluate the data-constrained setting of few-shot learning, where only a few samples for each category are provided within the linear probe setting (Fig. 2f; see Methods for more details). We observe that TITAN significantly outperforms all other encoders across different tasks and the number of shots (P < 0.0001), demonstrating strong generalizability. TITANV is the second-best-performing model, again underscoring that vision-language alignment benefits the downstream task performance. Notably, TITAN and TITANV exhibit especially high performance in one-shot learning, on par with other slide encoders trained on more shots (Supplementary Tables 82–85). Specifically, TITAN and TITANV outperform CHIEF by 22.4% and 13.5% (TCGA-UT-8K) and 18.7% and 6.8% (TCGA-OT), respectively, on 16 shots, although CHIEF has been pretrained on TCGA slides.
Interestingly, both TITAN and TITANV also outperform ABMIL with the same patch encoder across all settings, particularly in lower-shot settings. The largest gap for 1-shot is observed in the OT108 task, where TITAN outperforms ABMIL by 56.7%, with similar trends in prototyping evaluation (Supplementary Tables 86–89). Such superior data efficiency suggests that TITANV can excel in rare cancer settings with a limited number of samples, such as OT108 in our benchmark, without the need for task-specific fine-tuning.
Language-aligned TITAN enables cross-modal capabilities
We further assess the language capabilities of TITAN by aligning the slide representations of TITANV to language-based morphological descriptions. Specifically with TITAN, we assess the cross-modal zero-shot classification55,56,108 and report-generation capabilities and study the effect of stage 2 pretraining for caption alignment with fine-grained morphological descriptions and stage 3 pretraining with coarse clinical reports of relevant microscopic findings.
To evaluate the quality of vision-language alignment, we first perform cross-modal zero-shot experimentation on 13 subtyping tasks of varying difficulties comparing with PRISM, also equipped with cross-modal capabilities (Fig. 3a). In zero-shot classification, the diagnostic labels expressed as text prompts (Supplementary Tables 90–96) are encoded with the text encoder. The diagnostic prediction of the query slide is decided by the closest label embedding to the TITAN-encoded slide embedding, based on ℓ2 distance in the embedding space. We observe that TITAN performs the best across these tasks, significantly outperforming PRISM by a large margin on multiclass classification tasks (balanced accuracy +56.52%) and binary subtyping tasks (AUROC +13.8%), for both cancer subtyping tasks and noncancerous tasks (Fig. 3b and Supplementary Tables 97–109). The performance gap between TITAN and PRISM is the widest on the 30-class EBRAINS subtyping task, where the balanced accuracy of TITAN is more than double that of PRISM (balanced accuracy of +121.9%).
a, A schematic for zero-shot evaluation. The query slide is classified by identifying the closest text prompt embedding in the slide embedding space. b, Zero-shot performance of TITAN and PRISM. All multiclass tasks are evaluated with balanced accuracy and binary tasks are evaluated with AUROC. All error bars represent s.d. based on bootstrapping (n = 1,000). Dashed lines represent average performance for respective models (red, TITAN; teal, PRISM) c, Ablation study comparing different pretraining strategies, and assessed with zero-shot performance averaged across TCGA-UT-8K, TCGA-OT, OT108 and EBRAINS. Evaluations are based on the percentage changes of balanced accuracy from the reference zero-shot performance of TITAN. d, Report-generation evaluation on TCGA-Slide-Reports, and evaluated using METEOR, ROUGE and BLEU. All error bars represent s.d. based on bootstrapping (n = 1,000). e, TCGA examples of generated reports of TITAN and PRISM, with the corresponding clinical reports. Additional examples of generated reports are available in Extended Data Fig. 7. Statistical significance was assessed by fitting a generalized linear mixed-effects model and performing a two-sided Wald z test on the fitted model. Significance shown with respect to TITAN. ****P ≤ 0.0001.
To understand how different design considerations affect the zero-shot performance, we ablate over pretraining stages and the slide encoder architecture (Fig. 3c). In total, we experiment with four variations of TITAN and present the average performance over four challenging subtyping tasks, TCGA-UT-8K, TCGA-OT, OT108 and EBRAINS (individual results can be found in Supplementary Tables 110–113). We observe that TITAN maintains the best overall zero-shot performance. Of the three pretraining stages, stage 1 vision pretraining contributes the least (balanced accuracy of −0.4% against TITAN), followed by stage 2 ROI caption alignment (−3.6% against TITAN) and stage 3 slide-report alignment (−7.3% against TITAN). This underscores the importance of aligning vision and language at both fine-grained and global levels, thereby combining the insights independently derived at patch-level7,10 and slide-level58,62,72, which is lacking in report-only aligned baselines such as PRISM and GigaPath. Finally, a multiheaded ABMIL (MH-ABMIL) network, serving as the vision backbone with vision-language alignment pretraining, lags behind TITAN with and without vision pretraining by 1.94% and 1.54%, respectively. This indicates that the ViT architecture, incorporating self-attention and ALiBi, provides better downstream performance than attention-based alternatives.
Finally, we assess TITAN’s capabilities of generating pathological reports, using the text decoder trained during CoCa pretraining. To this end, we introduce a report-generation task on TCGA, TCGA-Slide-Reports, consisting of 10,108 FFPE WSIs with paired slide-level reports parsed from 9,523 patient-level TCGA reports released by a previous study109 (see Methods for more details). We evaluate the models using three metrics METEOR110, ROUGE111 and BLEU112. We observe that TITAN outperforms PRISM by a large margin, on average by 161% across the three metrics (Fig. 3d). Examples of the generated reports for TITAN considered high-quality by the pathologists are shown in Fig. 3e, often capable of correctly capturing key attributes such as tissue site, diagnosis and tumor grade as well as key representative morphology (Extended Data Fig. 7).
TITAN enables rare cancer retrieval and cross-modal retrieval
Consulting cases with similar morphological features and diagnoses is essential for pathologists to make informed decisions, especially when dealing with complex or rare cases5,17,47,48,50,51,53,113,114. Retrieving similar histology slides or pathology reports facilitates the identification of relevant cases from large archival databases, and has become an essential clinical decision support in digital pathology workflows. This is especially beneficial for rare cancers that affect fewer than 15 individuals per 100,000 annually43,44,45, for which pathologists can identify nonspecific malignancies based on similar WSIs and their corresponding pathology reports. Slide foundation models readily provide WSI representations for vector database indexing, fundamentally simplifying the task of histology slide retrieval.
Given a query slide and a set of support slides with diagnostic labels (indexed by a slide foundation model), histology slide search is evaluated by assessing the accuracy in retrieving identically labeled slides from the support set. Specifically, we test whether the K-closest neighbors of a query slide in the embedding space, determined using Euclidean distance with K = {1, 3, 5}, include slides of the same diagnostic label as the query (see Methods for more details).
We design three variations of the rare cancer retrieval task, Rare-Cancer, Rare-Cancer-Public and Rare-Cancer-External, to assess generalization in different scenarios (see Methods for more details). For Rare-Cancer, we curate a large database of 186 cancer types with 19,626 WSIs by combining a ‘rare cancer set’ of 43 cancer types (3,039 WSIs) with the ‘common cancer set’ of 143 more common cancer types (16,587 WSIs) from TCGA, EBRAINS and MGB internal data (Fig. 4a and Supplementary Table 114). This emulates the real-world setting of clinicians interacting with an extensive cancer database encompassing a diverse mix of rare and common cancer types. A query set is the subset of the ‘rare cancer set’, ensuring representation of all 43 rare cancer types, and a support set contains all remaining WSIs of the ‘rare cancer set and the common cancer set’, ensuring representation of all 186 cancer types. For Rare-Cancer-Public, we curate a public version with 127 cancer types and 14,062 WSIs using the data from TCGA and EBRAINS, resulting in 29 rare cancer types (1,982 WSIs) and 98 common cancer types with lower diversity (12,080 WSIs; Supplementary Table 115). Finally, we curate Rare-Cancer-External for external validation, comprised of 39 WSIs covering 12 challenging rare ovary and soft tissue cancers from Kanagawa Cancer Center Hospital, Japan (Supplementary Table 116).
a, Slide retrieval results on rare cancer retrieval tasks assessed with Accuracy@K, with K = {1, 3, 5}. Rare-Cancer (internal rare cancer cohort) consists of TCGA, EBRAINS and the MGB internal cohort, with 43 rare and 143 common cancer types for a total of 186 classes. Rare-Cancer-Public (public rare cancer cohort) consists of TCGA and EBRAINS only, with 29 rare and 98 common cancer types for a total of 127 classes. Rare-Cancer-External consists of 12 rare cancer types for the ovary and soft tissue, curated at Kanagawa Cancer Center Hospital, Japan. b, Example of rare cancer retrieval on Rare-Cancer with the query slide and four representative retrieved slides. The number indicates the cosine similarity between the query and the retrieved slide. Additional examples of rare cancer retrieval are available in Extended Data Fig. 8. c, Slide retrieval results on five subtyping tasks. Mean represents the average performance across three shots. d, Report-to-slide and slide-to-report cross-modal retrieval performance assessed with Recall@K, with K = {1, 3, 5, 10} on TCGA cohort of 10,108 pairs of WSIs and reports for TITAN and PRISM. Mean represents the average performance across four shots. All error bars represent s.d. based on bootstrapping (n = 1,000). Statistical significance was assessed using TITAN by the fitting of a generalized linear mixed-effects model and a two-sided Wald z test on the fitted model. Significance shown with respect to TITAN. P values for nonsignificant results are shown. **P ≤ 0.01, ***P ≤ 0.001, ****P ≤ 0.0001.
We observe that TITAN significantly outperforms other slide encoders on average with +14.8% in Accuracy@K and +18.1% in MVAcc@K to the next best model PRISM (Supplementary Table 117). On Rare-Cancer-External, we observe that our slide encoder is significantly more robust to the domain shift to the external institution than other slide encoders with +30.8% and +41.5% in Accuracy@K and +31.2% and +26.7% in MVAcc@K for TITAN and TITANV to the next best model GigaPath (P < 0.0001; Supplementary Table 118). The trends in performance are preserved on Rare-Cancer-Public with slightly higher performance levels as the task is easier with a support set containing fewer cancer types (Supplementary Table 119). An example of rare cancer retrieval is demonstrated in Fig. 4b, where the closest slide to the paraganglioma query is also of paraganglioma with a high similarity of 0.794 and less similar slides are of different cancer types (haemangioma from brain, similarity of 0.341). One of the retrieved slides is pheochromocytoma with a high similarity of 0.651, agreeing with the clinical understanding that both are morphologically tightly connected as rare neuroendocrine tumors115 (additional examples in Extended Data Fig. 8). With multiclass cancer subtyping tasks of varying difficulties, we also observe that both TITAN and TITANV significantly outperform other slide encoders (P < 0.0001; Fig. 4c and Supplementary Tables 120–124).
We further investigate the cross-modal retrieval performance of TITAN, as the slide and report embedding spaces are already aligned (see Methods for more details). We perform the cross-modal experiments on TCGA-Slide-Reports, our proposed dataset for report generation with 10,108 slide-report pairs (Supplementary Table 125). We observe that TITAN significantly outperforms PRISM on both retrieval tasks across all K retrievals, as measured with Recall@K for K = {1, 3, 5, 10}, with +10.5% and +20.5% on average for report-to-slide and slide-to-report retrieval tasks, respectively (Fig. 4d and Supplementary Tables 126–127). The strong performance of TITAN even with a single report (0.75) hints at the clinical potential. For a diagnostically challenging query slide, clinicians can benefit from sifting through retrieved past reports with similar diagnoses.
Discussion
We introduce TITAN, a multimodal whole-slide foundation model for pathology, which combines and elevates successful SSL recipes from the patch level to the slide level. Methodologically, TITAN employs histology knowledge distillation in the feature space (vision-only) and contrastive learning by aligning ROIs with synthetic captions and WSIs with reports (vision-language). Pretrained on 336k WSIs, TITAN, a ViT architecture equipped with ALiBi positional encoding for long-context extrapolation, produces powerful general-purpose slide representations for a large variety of downstream tasks even without task-specific fine-tuning. From cancer subtyping to molecular classification, TITAN consistently outperforms other state-of-the-art slide encoders, such as PRISM62, GigaPath58 and CHIEF83. This superiority is maintained in data-constrained settings such as rare disease classification and histology slide retrieval, which underscores the representation quality of TITAN. Further aligning the vision-pretrained TITAN with 423k ROI-level captions generated by PathChat and 183k pathology reports equips the model with multimodal capabilities such as zero-shot diagnosis, slide-report retrieval and report generation. We observe that aligning the slide embedding with both the fine-grained (ROI captions) and coarse-level (pathology reports) descriptions is crucial for handling the multiscale information inherent in tissue slides.
Detailed ablation analyses reveal further insights into TITAN. We observe that stage 1 unimodal pretraining of TITANV captures morphological concepts already with much less data than existing slide encoders. In particular, TITANV consistently outperforms its mean pooling and task-specific attention-based pooling baselines that use the same patch encoder as TITANV, proving that unimodal pretraining effectively captures the context of patch features in contrast to existing unimodal slide encoders. Next, in addition to unlocking language-related capabilities, we observe that the vision-language alignment further enhances the representation quality of our vision-only model. Specifically, TITAN outperforms TITANV for slide-level tasks, with the strongest improvements observed in nonparametric evaluation settings. While slide embeddings from pretrained TITAN are already promising, especially in the low-data regime, task-specific fine-tuning of the pretrained model can further enhance the downstream performance for tasks with a large enough patient cohort, pointing to the flexibility of TITAN when applied to diverse clinical and data settings. We conjecture that some of these insights can be readily translated into other domains of pathology foundation models, such as hematopathology116, spatial transcriptomics117, 3D pathology118 and multiplex imaging119.
Providing multimodal slide embedding off-the-shelf presents immediate clinical potential to assist clinicians in their routine diagnostic workflows85. Presented with diagnostically challenging tissue slides, pathologists and oncologists can greatly benefit from being able to retrieve and analyze diagnostically similar slides or clinical reports51. This would lead to a reduction in patient misdiagnosis and interobserver variability. TITAN can accurately retrieve similar diagnostic slides and reports for challenging scenarios from a large number of cancer types (>100), as well as rare cancer types45 where the corresponding slides have scarce representation in the database. That all of these tasks could be performed off-the-shelf with TITAN without a dedicated algorithm for each underscores both the generalizability of TITAN slide embeddings and the simplicity of slide-level tasks with the pretrained slide encoders.
Despite the encouraging performance of TITAN, our framework has a few shortcomings. First, pretraining on 8k × 8k region crops and extrapolating with ALiBi to the entire WSI may still not capture the full contextual information. Other positional encodings for extrapolation could address this limitation. Next, despite our best efforts to curate a diverse pretraining dataset, patch foundation models and, consequently, slide foundation models are susceptible to encoding nonbiological features, such as tissue processing sites and scanners, which may compromise their translational impact120,121,122,123. We believe that systematic investigations similar to our robustness analysis and insights discovered124,125, combined with ongoing efforts to curate larger and multi-institutional pretraining datasets can mitigate the issue. Next, clinical reports processing still poses a challenge for vision-language alignment. Incorporating comprehensive clinical information conducive to contrastive learning, while ensuring that it is linked to morphology to some degree, involves substantial manual tuning even with the automated processing pipelines. Restructuring the reports into distinctive morphology and molecular characteristics could facilitate more effective learning. Finally, Mass-340K contains fewer slides compared to other pretraining datasets used for patch encoders12,13,126 and slide encoders62,72. We believe that the already strong performance of TITAN, merged with efforts to expand Mass-340K, will further improve performance.
Promisingly, TITAN can be scaled up in terms of data and architecture. WSIs and corresponding medical reports are routinely available and stored. The synthetic region-level captions can easily be generated with the generative AI model to provide a wealth of text guidance88,127. Combining the additional data and a heavier architecture can potentially improve the performance, as demonstrated with patch encoders12,13,126. Additionally, improved patch representation quality is likely to enhance the quality of the downstream slide encoder.
In conclusion, we envision TITAN and its future iterations being incorporated into practitioners’ everyday toolkits for routine application and comparison with other task-specific supervised frameworks, together reaching higher levels of performance in clinically important tasks.
Methods
Ethics statement
The retrospective analysis of internal pathology images and associated reports used in this study received approval from the Mass General Brigham Institutional Review Board. Before the computational analysis and model development, all internal digital data, including WSIs, pathology reports and electronic medical records, were anonymized. Since the study did not involve direct patient participation or recruitment, informed consent was waived for the analysis of archival pathology slides.
Pretraining dataset
For large-scale visual pretraining, we curated Mass-340K, a diverse dataset consisting of 335,645 WSIs across 20 organs, with 89.7% hematoxylin and eosin (H&E), 7.9% IHC, 2.3% special stains and 0.1% others, across different tissue types (neoplastic 70.0%, tissue damage response 8.4%, normal 4.7%, inflammatory 3.4% and others 13.5%), sourced from the combination of in-house histology slides at Mass General Brigham (MGB), consult slides sent to MGB and the GTEx consortium128,129. Sourced from several sites, Mass-340K covers a wide range of tissue preprocessing protocols with diverse scanners and stainers.
Scanner type
Setting aside the publicly available GTEx cohort for which the scanner type information is not available, we confirm that Mass-340K uses 16 different scanners from seven different manufacturers. Detailed data breakdown along scanner types can be found in Supplementary Table 1.
Stainer type
For the internally-curated cohort at MGB, the following stainers were used: Leica HistoCore Spectra (H&E), Agilent DAKO CoverStainer (H&E), Leica Bond III (molecular), Leica BOND PRIME (IHC), Agilent Dako AutoStainer Link 48 (IHC) and Agilent Dako Artisan Link Pro (special stain).
Stains
The 27k IHC slides in Mass-340K span 100+ unique stains, without focus on particular biomarkers. The goal of IHC curation was to ensure that TITAN is exposed to a large set of slides with diverse tissue appearances during the pretraining process. For example, these stains include proliferation markers (Ki-67), lymphoid and hematopoietic markers (CD4, CD20) and oncogenes and tumor markers (MYC, BRAF, human epidermal growth factor receptor 2 (HER2)). In addition, Mass-340K contains 50+ unique special stains, such as Masson’s trichrome and Congo red.
To explore the effects of data scale at the pretraining stage, we formed three additional partitions of Mass-340K, containing 12.5%, 25% and 50% of the original dataset. These partitions were sampled to maintain the ratio of different data sources and preserve organ distribution.
Synthetic caption generation using PathChat
For the initial stage of vision-language alignment (stage 2 of TITAN), we used synthetic captions generated by PathChat, a state-of-the-art multimodal large language model designed for pathology88. To go beyond the typically brief clinical reports focused on the final diagnosis, we prompted PathChat to generate detailed morphological descriptions of ROIs, providing important training data for models to capture complex pathological features. Using PathChat, we generated synthetic captions for 423,122 diverse ROIs of 8,192 × 8,192 pixels sampled from Mass-340K. Since PathChat cannot process inputs of size 8,192 × 8,192 pixels directly, we divide each ROI into 64 patches of size 1,024 × 1,024 pixels. To retain the most representative morphological features, we applied K-means clustering with K = 16 to the 64 patches and then randomly sampled one patch from each cluster. The resulting 16 morphologically representative 1,024 × 1,024 patches were subsequently fed to PathChat. To further enhance the diversity of these captions, we used Qwen2-7B-Instruct130 to rewrite the generated captions, ensuring varied language structures and expressions. Detailed prompts for both PathChat and Qwen2, along with examples of generated and diversified captions, are provided in Supplementary Tables 4 and 5.
Curation of slide-report dataset
For the second stage of vision-language alignment (stage 3 of TITAN), we curated a dataset of 182,862 slide-report pairs from a combination of in-house clinical reports and pathology notes from the GTEx consortium129. However, clinical reports are often noisy and are typically organized at the patient level, hence contain information on multiple slides from the same patient, complicating the slide-report alignment. To address this, we used a locally served Qwen2-7B-Instruct130 model to extract slide-specific descriptions and remove sensitive information unrelated to pathological diagnosis, such as gross descriptions, hospital and doctor names and patient clinical history. Additionally, we applied the same rewriting strategy used for synthetic captions to diversify the report text. Example prompts used for report cleaning and rewriting can be found in Supplementary Tables 6–8.
Unimodal visual pretraining
Preprocessing
Similar to previous studies9,10,74, WSIs were preprocessed by tissue segmentation, tiling, and feature extraction using a pretrained patch encoder. We used the CLAM toolbox74 for tissue segmentation and tiling. Tissues were segmented by binary thresholding of the saturation channel in HSV color space at a low resolution. Following this, we applied median blurring, morphological closing and filtering of contours below a minimum area to smooth tissue contours and eliminate artifacts. Nonoverlapping 512 × 512 pixel patches were then extracted from the segmented tissue regions of each WSI at ×20 magnification. For feature extraction, we used CONCHv1.5, an extended version of CONCH10, which was trained with 1.26 million image-caption pairs using the CoCa training objective for 20 epochs. The choice of CONCHv1.5 for feature extraction was due to the fact that the model was pretrained on histology regions with diverse stains and tissue types, including FFPE, frozen tissue and IHC, thereby yielding region features that are robust against diverse tissue processing protocols. By increasing the patch size from the widely used 256 × 256 pixels, we effectively reduce the sequence length by four without impacting the representation quality due to higher resolution patch input, leveraging the robustness of the patch-level foundation models in generalizing to higher resolutions9,10,87.
Refer to Supplementary Table 2 for detailed hyperparameters of the patch encoder.
To enhance the effectiveness of the ROI sampling strategy during stage 1 training of TITANV, an additional preprocessing step was performed to group the segmented tissue contours based on their spatial proximity within the slide. This addresses the challenging cases where multiple tissue regions are interspersed with background areas, particularly for biopsy samples where tissue fragments are often widely dispersed and for samples with multiple slices placed on the same slide. Specifically, we grouped tissue contours into clusters based on their coordinates, resulting in tissue groups that contain densely packed tissue regions with minimal background regions between them. Furthermore, tissue groups that contained fewer than 16 patches were filtered out. This grouping operation produced a total of 345,782 tissue groups from Mass-340K.
Pretraining protocol
For training TITANV on Mass-340K, we use iBOT, a state-of-the-art SSL method that combines student–teacher knowledge distillation and masked image modeling86. As iBOT is applied in the patch embedding space, instead of the typical use case of the raw image space, we adapt the pretraining recipes as follows.
View generation
During training, we create region crops randomly sampled from the tissue groups, each of which corresponds to a feature grid of size 16 × 16, corresponding to a field of view of 8,192 × 8,192 pixels at ×20 magnification (Fig. 1b). The random sampling of region crops, instead of precomputing fixed regions, increases the diversity of the training set and effectively acts as an additional data augmentation, as the model encounters different parts of the same WSI at each training epoch. A region crop contains 256 features, which is equivalent in length to training on images of 256 × 256 pixels with a token size of 16 × 16 in the typical natural image setting. From this region crop, two global views (14 × 14 crops) and ten local views (6 × 6 crops) are generated by cropping within the region crop without scaling or interpolation and subsequently fed to iBOT training. The 2D feature grid setup allows us to directly apply student–teacher knowledge distillation approaches, which typically require square crop inputs.
To achieve realistic augmentations in the embedding space, existing methods have employed offline image augmentations in the pixel space34,59 by extracting multiple patch features from different views of a given patch. While effective, this approach limits the number of additional views and becomes computationally infeasible for large training datasets. Additionally, choosing color space augmentations tailored to histopathology that go beyond standard color transformations introduces additional computational overhead. A few recent approaches addressed the difficulty with training generative networks on the feature space to transform the features131,132, but also introduced additional computational cost for training. Instead, we apply frozen feature augmentations, which have been shown to work well for a few-shot classification task in the feature space of pretrained ViTs93.
Positional encoding
Traditional multiple instance learning methods consider the patches to be permutation-invariant within the slide. Despite the promising results, this approach ignores the tissue context, which can be essential for capturing the interaction in the tumor microenvironments and can thus affect the model’s performance133. In this context, for TITAN, we employ positional encodings in the patch embedding space to break permutation invariance and encode tissue context. Furthermore, TITAN adopts the strategy of ‘train short, test long’ to ease the computational burden, which also requires positional information via positional encodings. Trained at the region crops (ROIs) of 8,192 × 8,192 pixels (train short), we directly apply TITAN on the whole slide during inference (test long). We used ALiBi, a method originally proposed for 1D sequence in large language models94. Absolute positional encoding, another popular alternative that works well for images at training sizes, was shown to have weak extrapolation abilities94. Unlike other positional encodings applied to the input features, ALiBi adds a bias to the query-key dot product during the computation of attention scores. ALiBi effectively penalizes the attention score for tokens that are further apart from each other. Formally, let \({q}_{i}\in {{\mathbb{R}}}^{d}\) and \({k}_{j}\in {{\mathbb{R}}}^{d}\) represent the i-th query and j-th key, respectively. The attention score, which is typically computed as \({\rm{softmax}}\,({q}_{i}{k}_{j}^{\,\text{T}})\), is modified with 1D ALiBi as \({\rm{softmax}}\,({q}_{i}{k}_{j}^{T}-m| i-j|)\), where m is a predefined slope specific to each attention head. Since the feature grids and the resulting views are of 2D grid structure, we extend ALiBi to 2D by incorporating the Euclidean distance between the patches i and j. The 2D ALiBi can be written as
where ix, iy and jx, jy are the 2D grid coordinates of patches i and j. The x and y coordinates are defined as the 2D patch coordinates (at magnification ×20) divided by the patch size of 512.
Network architecture and training details
For the slide encoder, we use a ViT92 with six transformer layers, 12 attention heads of dimension 64, resulting in an embedding dimension of 768 and a hidden dimension of 3,072. This smaller architecture, compared to typical ViTs used in patch encoders, is chosen based on previous studies57, which suggest that a compact network suffices for slide representation learning in the embedding space, especially given the limited data scale of WSIs compared to histology patch datasets, which are on the scale of billions. The patch embedding layer is replaced by an MLP to process the feature inputs. We train the model for 270 epochs (equivalent to 91,260 iterations), distributed across four NVIDIA A100 80GB graphics processing units (GPUs) with a local batch size of 256 per GPU. For all training hyperparameters, refer to Supplementary Table 3.
Vision-language continual pretraining
To enhance the unimodal capabilities of TITANV, we further explored the multimodal vision-language alignment of TITANV with clinical text. Training a multimodal foundation model, however, faces several limitations related to data and compute. First, paired slide-report data are scarce compared to the scale of millions of image-caption pairs for patches. Additionally, real-world clinical reports typically contain only brief diagnostic information, unlike the detailed morphological descriptions in educational captions for histology ROI images. Finally, contrastive learning-based cross-modal training typically requires a large batch size, which is computationally infeasible for WSIs.
To address these issues, we propose a two-stage continual pretraining approach (referred to as stage 2 and stage 3 for TITAN) that progressively aligns the model with increasing context. We first align synthetic captions for ROIs of 8,192 × 8,192 pixels, followed by real clinical reports for WSIs. With emphasis on detailed morphological descriptions, the first vision-language alignment stage allows the model to learn fine-grained pathological concepts using a large batch size. In the next stage, we further augment the model’s understanding of diagnostic terminology and reasoning, targeted to enhance its zero-shot understanding in downstream tasks. The second stage also serves as a ‘high-resolution fine-tuning’ phase, adapting the model from the local contexts of ROIs to the full-scale global context of WSIs. Altogether, these two stages are designed to gradually build the model’s ability to comprehend and generate meaningful vision-language representations for WSIs.
Network architecture and training details
Following the success of previous studies10, we use CoCa95, a state-of-the-art visual-language foundation model pretraining method, for both stages of vision-language alignment. The model consists of an image encoder, a text encoder and a multimodal text decoder. Using our unimodal TITANV as the image backbone, we add two attentional pooler components on top. The first attentional pooler uses a single query (contrastive query) to pool a single global representation of the feature grids and enable cross-modal contrastive learning with text embeddings. This global WSI representation can then be used for zero-shot or unsupervised evaluation of TITAN on downstream tasks. The second attentional pooler uses n = 128 queries (reconstruction queries) to generate a set of 128 image tokens designed for interacting with the multimodal text decoder for caption generation. We use the pretrained text encoders and multimodal decoders of CONCHv1.5 (ref. 10), each consisting of 12 transformer layers with an embedding dimension of 768 and a hidden dimension of 3,072.
For both stages, we used eight NVIDIA A100 80GB GPUs. During stage 2 vision-caption pretraining, we used a local batch size of 196 per GPU, with gradient accumulation of 2, resulting in an effective batch size of 3,136. For stage 3 vision-report pretraining, we randomly crop the WSIs to 64 × 64 feature grids, allowing for larger batch sizes while maintaining a large field of view, corresponding to 32,768 × 32,768 pixels, which already covers most slides in our pretraining dataset. We used a local batch size of 16 per GPU, with a gradient accumulation of 2 to achieve an effective batch size of 256. To avoid deteriorating the quality of the pretrained vision encoder, we used a smaller learning rate and weight decay, as well as a slow warm-up strategy for the vision backbone, following previous work134. For all hyperparameters, refer to Supplementary Tables 9 and 10.
Evaluation setting
Baselines
We compare TITANV against (1) unsupervised baselines with four other slide encoders, Prov-GigaPath (referred to as GigaPath)58, PRISM62, CHIEF83, and the mean pooling baselines with features from the respective patch encoders, (2) supervised baselines and (3) our vision-language model TITAN against zero-shot baseline PRISM.
Unsupervised baselines
GigaPath uses LongNet architecture as the slide encoder, a ViT92 in the ‘base configuration’, replacing the vanilla dense attention with dilated attention. It was trained on 171,189 in-house WSIs from Providence via masked autoencoder135. As a patch encoder, GigaPath uses ViT-G/14 pretrained with DINOv2 (ref. 87) on the same in-house dataset. While GigaPath further performed continual vision-language pretraining, we only assess the unimodal model, as the multimodal model is not publicly available. For performance analysis, we use the output of the Transformer layer 11 as slide representation, which yields the best results on downstream tasks and also agrees with the provided fine-tuning recipe. PRISM62 uses the Perceiver architecture136 as the slide encoder, incorporating CoCa-based vision and language alignment95 on 195,344 specimen-report pairs, which comprise a total of 587,196 WSIs, each containing one or more WSIs. As for the patch encoder, PRISM uses Virchow11, a ViT-H/14 pretrained with DINOv2 (ref. 87) on an in-house dataset. CHIEF83 applies attention-based feature aggregation, trained via slide-level contrastive learning and anatomic site information. The patch encoder is based on CTransPath4, a self-supervised SwinTransformer137 trained on 15 million patches. In addition to the pretrained slide encoders, we evaluate mean pooling as a baseline, where the patch features are averaged within each slide, as it serves as a strong unsupervised baseline despite its simplicity64,65,66. While we mainly compare with mean pooling based on CONCHv1.5 patch features, we also provide results for mean pooling with the corresponding patch encoders of each slide encoder for a subset of analyses.
Supervised baselines
We compare TITAN against ABMIL73,74 and the fine-tuning of the pretrained slide encoders. For ABMIL, the model was trained with a batch size of 1 using the AdamW optimizer with weight decay 10−5 and a Cosine annealing learning rate scheduler with peak learning rate 10−4 over 20 epochs. The patch encoders were selected accordingly for each analysis. For GigaPath fine-tuning, we used the publicly available code, which uses a batch size of 1, AdamW optimizer with weight decay 0.05 and Cosine annealing learning rate scheduler with warm-up and base learning rate 2 × 10−3 over five epochs. For CHIEF fine-tuning, we also used the publicly available fine-tuning code. For tasks with a validation set, the best model is chosen based on the validation loss.
Cross-modal baselines
For cross-modal zero-shot retrieval and clinical report generation, we compare TITAN against PRISM62.
Linear and k-nearest neighbor (k-NN) probe evaluation
To evaluate the transfer capabilities and representation quality of slide encoders, we adopt recent work in representation learning with self-supervised frameworks and perform linear (logistic regression) and k-NN probing. For linear probing, we minimize cross-entropy loss using the scikit-learn L-BFGS solver with ℓ2 regularization, selecting ℓ2 from 45 logarithmically spaced values between 10−6 and 105 based on the validation loss. The maximum number of L-BFGS iterations is set to 500. For datasets without a validation set, such as small datasets or few-shot experiments, we use the default values of ℓ2 = 1 with 1,000 iterations. We additionally evaluated with k-NN probing, a nonparametrized measure to quantify the representation quality of fixed embeddings. We apply it in the following two settings: first, we follow SimpleShot to create a prototypical class representation by averaging all slide embeddings per diagnostic class105; second, we use the scikit-learn implementation of k-NN with k = 20 following stability observations from SSL literature87,138. In both settings, Euclidean distance is used as the distance metric based on the centered and normalized slide embeddings.
Slide retrieval
To further evaluate the representation quality of different slide encoders, we perform content-based slide retrieval using slide-level classification datasets, where we retrieve slides with the same class label as a given query slide. Specifically, we extract slide features for all WSIs. The training and validation sets are combined to serve as the database of candidate slides (keys), and we treat each slide in the test set as a query slide. Before retrieval, we preprocess both keys and queries by centering the slide embeddings, which involves subtracting their Euclidean centroid, followed by ℓ2 normalization. The similarity between the query and each candidate in the database is computed using the ℓ2 distance metric, where a smaller distance indicates a higher similarity. The retrieved slides are then sorted based on their similarities to the query. The class labels are used to evaluate the retrieval performance using Acc@K for K ∈ {1, 3, 5}, which measures whether at least one of the top K retrieved slides shared the same class label as the query, and MVAcc@5, which considers the majority class label among the top five retrieved slides. Detailed descriptions of these metrics are provided in ‘Evaluation metrics’.
Cross-modal retrieval
Leveraging the vision-language aligned embedding space, we also evaluate cross-modal retrieval performance on TCGA-Slide-Reports. Specifically, we assess both slide-to-report and report-to-slide retrieval tasks. All slides and reports are embedded into a shared space using the vision and the text encoders, respectively, followed by ℓ2 normalization. Retrieval is performed by calculating pairwise cosine similarity between the slide and report embeddings. Our class-based approach mirrors the unimodal slide retrieval, where retrieval is successful if the retrieved slide or report belongs to the same diagnostic class as the query. Performance is quantified using Recall@K for K ∈ {1, 3, 5, 10} for the class-based approach, which measures the proportion of queries for which the correct result appears among the top K retrieved items. Additionally, we report the mean recall, computed as the average of the Recall@K values across the four K levels. Further details on these metrics can be found in ‘Evaluation metrics’.
Few-shot slide classification
We evaluate few-shot classification by varying the number of shots K in {1, 2, 4, 8, 16, 32}. For each K, we select K shots per class or all samples per class if the class has less than K samples. We follow previous studies that used the SimpleShot105 framework for evaluation of the few-shot learning performance of self-supervised models9. SimpleShot computes a prototypical representation per class by averaging all samples within that class. The distances to the class prototypes are then computed on the test set. All embeddings are centered and normalized based on the few-shot samples. To make the evaluation more comparable to supervised baselines, such as ABMIL, we also assess few-shot classification with linear probing. As no validation set is available in few-shot experiments, we use the default scikit-learn recipe with regularization strength ℓ2 = 1 and up to 1,000 iterations of the L-BFGS solver. To mitigate sampling bias, we aggregate the results across 50 different runs, using random samples for training while keeping the test set fixed.
Survival analysis
For survival analysis, we employed the linear Cox proportional hazards model on the disease-specific survival clinical endpoint. We note that this differs from typical MIL survival prediction with negative log likelihood65,139, as we deal with a single embedding for the slide (as opposed to a bag of patch embeddings), and patients can be batched (as opposed to the single patient per batch due to memory usage). To reduce the impact of batch effects, we performed a five-fold site-preserved stratification140. Due to the small cohort size for reliable survival prediction modeling, we used four folds for training and the remaining fold for evaluation, without employing the validation fold. A hyperparameter α was searched over 25 logarithmically spaced values between 101 and 105, with the ℓ2 coefficient defined as C = α. For each combination of encoder and cancer type, we chose C that yielded the best average test metric across the five folds. For fitting and testing the Cox model, we used the scikit-surv package.
Zero-shot slide classification
For zero-shot slide classification, we adopted the method described in CLIP108 to use the similarities between a given slide and the text prompts of each class as its prediction logits. Specifically, for a class c ∈ {1, 2, …, C}, we first created the text prompts for each class, followed by extracting their ℓ2-normalized text embeddings vc using the text encoder. Since the model could be sensitive to the specific choice of text prompts, we created an ensemble of prompts for each class. The complete set of prompt ensembles are provided in Supplementary Table 103. For each WSI, we similarly computed a ℓ2-normalized embedding ui using the slide encoder. We then calculated the cosine similarity between the slide embedding and each class text embedding. The predicted class for a slide was the one with the highest cosine similarity score:
Report generation
Slide captioning provides concise and interpretable summaries of visual findings in pathology, potentially enhancing clinical workflows. The generative objective of CoCa enabled the model’s capabilities of generating pathological reports, which we explored on 10,108 slide-report pairs from TCGA. We performed zero-shot captioning using TITAN and compared the quality of the generated report against PRISM62. Specifically, we use a beam search decoding strategy with 5 beams and 1 beam group, where the model explores five potential sequences at each step and retains only the most likely sequence within a single group to maximize quality while minimizing redundancy.
Evaluation metrics
We report balanced accuracy and weighted F1-score for all classification tasks with more than two classes. For ordinal multiclass classification tasks, we report balanced accuracy and quadratic-weighted Cohen’s κ. For binary classification tasks, we report balanced accuracy and AUROC. For survival tasks, we report the concordance index (c-index), which measures the agreement between the model’s predicted risks and the actual survival times. The expected calibration error (ECE)107 measures whether the model’s predicted probabilities match the actual frequencies of each diagnostic label, with the lower value indicating that the model’s confidence estimates are well-calibrated. We use a multiclass variant of the original ECE, with one-versus-all binarization of the labels with respect to a given diagnostic label computed and averaged across all labels. The entropy score measures the uncertainty of predictions, with a lower value indicating that the model has higher confidence in its predictions. The entropy of the predicted probabilities was computed.
For slide retrieval tasks, we report Acc@K for K ∈ {1, 3, 5}, which measures if at least one slide among the top K retrieved slides has the same class label as the query. We also report MVAcc@5, which is a stricter metric that considers whether the majority vote of the top 5 retrieved slides is in the same class as the query. For cross-modal retrieval tasks, we report Recall@K for K ∈ {1, 3, 5, 10}, which measures the proportion of queries for which the correct result appears in the top K retrieved items. We also report mean recall, which is calculated as the average of the four Recall@K values. For report generation, we compare the generated reports with the ground truth pathological reports using METEOR, ROUGE and BLEU. METEOR110 is a metric that evaluates text quality through unigram matching by considering both precision and recall while also accounting for synonyms, stemming and word order between the candidate and reference texts. ROUGE111 compares the overlap of n-gram, word sequences and word pairs between the generated and reference texts, focusing on recall. We use ROUGE-1, which specifically measures the overlap of unigrams. BLEU112 measures the quality of generated text based on unigram overlap, focusing on precision. We use BLEU-1, which evaluates the extent of word-level matches between the generated and reference texts.
Statistical analysis
For the datasets with five-fold splits, where we employ five-fold cross-validation, we report the mean performance and the s.d. across all folds. For the datasets with a single split, we use nonparametric bootstrapping with 1,000 samples to calculate the mean and s.d.
To compare the performance of multiple methods across different datasets, we used a hierarchical generalized linear mixed-effects model (GLMM). A GLMM is a statistical model that enables analysis of the data with both fixed and random effects. Specifically, we are interested in estimating the effect of each method (fixed effects) while accounting for variability across datasets (random effects). The hierarchical structure captures the fact that datasets differ in their overall performance levels, while the mixed-effects framework ensures that method comparisons are made after adjusting for these dataset-specific effects. Since the performance metric is bounded between 0 and 1, we used a β distribution, parameterized in terms of a mean μij and a precision parameter ϕ. The expected value of the metric for method j on dataset i is modeled as:
where the mean μij was linked to the predictors using a logit transformation, with
-
α is the overall intercept,
-
βj is the fixed effect of method j,
-
bi is a random intercept for dataset i modeled with Gaussian distribution, that is, bi ~ n(0, σ2).
This approach accounts for the possibility that some datasets may consistently produce higher or lower performance scores, preventing these systematic differences from being misattributed to the methods themselves. We assume that, while absolute performance scores vary across datasets, the relative ranking of methods remains approximately consistent (for example, if Method A tends to outperform Method B, it is likely to do so across most datasets). Parameters were fitted using the maximum likelihood estimation, and model fit was assessed through diagnostic checks of residual distributions and variance components. To compare methods, we compute estimated marginal means—the predicted average performance for each method adjusted for dataset-level variability. Pairwise comparisons of these means are conducted using two-sided Wald z tests, with the Tukey correction applied to control for multiple comparisons and ensure robust inference.
We also evaluate few-shot learning performance, where methods are compared with limited training examples (K = 1, 2, 4, 8, 16). For a given task (or dataset), to isolate the effect of method choice, we include the number of training examples as the random effect. We use a hierarchical GLMM with a β distribution and compute estimated marginal means, with correction for multiple hypothesis testing, to assess whether substantial performance differences exist between models. For the retrieval tasks, we follow a similar approach to the few-shot by treating different numbers of retrieved samples as the random effect.
Downstream evaluation datasets
For the evaluation of TITAN on a diverse set of downstream tasks (Supplementary Tables 18–21), we re-arrange the pre-extracted CONCHv1.5 features from patches of 512 × 512 pixels to feature grids cropped around the tissue regions of the WSIs. Additionally, background masks are created to mask out features corresponding to background patches. Each WSI is then one single input image to TITAN. For downstream tasks with patient-level annotations, we create patient embeddings by averaging all slide embeddings of TITAN corresponding to a single patient. In the following, we detail all datasets used in our downstream evaluations, including splits and targets. We first describe the six datasets that we introduce in our study, TCGA-UniformTumor-8K, TCGA-OncoTree, TCGA-Slide-Reports, Rare-Cancer, Rare-Cancer-Public and Rare-Cancer-External, followed by existing datasets in alphabetical order. To mitigate the impact of batch effects, all datasets based on TCGA are split into label-stratified and site-preserving folds such that slides from one clinical site only occur in one fold following140.
TCGA-UniformTumor-8K (TCGA-UT-8K)
The TCGA-UT-8K dataset is a region-level pan-cancer subtyping resource comprising 25,495 ROIs of 8,192 × 8,192 pixels. These regions were extracted from 9,662 H&E-stained FFPE diagnostic histopathology WSIs sourced from TCGA. The tumor regions were manually annotated by two expert pathologists, with slide exclusion due to poor staining, poor focus, lacking cancerous regions and incorrect cancer types. Approximately three representative tumor regions per WSI were annotated with pixel-level contours. For each contour, we center-cropped an image region of 8,192 × 8,192 pixels to encompass both the dense tumor and its surrounding tissue context. We split the regions into train-validation-test split (train-val-test; 13,853:3,434:8,208 slides), preserving the source site. Refer to Supplementary Table 11 for a detailed overview of all classes contained in this dataset.
TCGA-OncoTree (TCGA-OT)
The TCGA-OT is a pan-cancer subtyping dataset of 11,186 H&E FFPE diagnostic histopathology WSIs from TCGA96. All WSIs are classified into 46 classes according to the OncoTree classification system, such that every class is represented by at least 50 samples. We select all diagnostic H&E FFPE WSIs from TCGA with primary tumors. Concretely, we exclude frozen tissue slides, slides without magnification information, metastatic or recurrent tumor slides, slides without tumor tissue and IHC slides. For training and evaluation, we split the dataset into training-validation-test folds of 8,226:1,612:1,348 samples while preserving the source sites; that is, all slides from one source site are in one split. Refer to Supplementary Table 12 for a detailed overview of all classes.
TCGA-Slide-Reports
The TCGA-Slide-Reports is a pan-cancer slide-report dataset of H&E FFPE diagnostic histopathology WSIs from TCGA96. The dataset consists of 10,108 WSIs with paired pathological reports at the slide level. The dataset is built on the TCGA-Reports dataset, which consists of 9,523 patient-level reports released by a previous study109. The dataset TCGA-Reports was created using 11,108 pathology report PDFs, corresponding to 11,010 patients, available on the TCGA data portal. The raw reports were preprocessed by removing 82 patients with multiple reports, 399 patients with nonprimary tumors, 72 patients with no survival data, 381 ‘missing pathology’ reports and 212 ‘TCGA Pathologic Diagnosis Discrepancy Form’ reports, resulting in 9,850 reports. Optical character recognition was then performed to extract text from the PDFs, followed by the removal of ‘Consolidated Diagnostic Pathology Form’ reports, ‘Synoptic Translated’ forms, within-report TCGA metadata insertions and clinically irrelevant reports, resulting in 9,523 patient-level reports. While these reports are clean and clinically relevant, they often contain descriptions of multiple tissue blocks per patient. This lack of one-to-one mapping between slides and reports poses a challenge for slide-level report generation and cross-modal retrieval, which require distinct slide-to-report alignment. Since block IDs are unavailable in TCGA metadata, we used the slide-level diagnoses to map diagnoses in each tissue block description. Specifically, if a block’s diagnosis matched the slide-level diagnosis, we designated it as corresponding to the slide. This process was automated using GPT4o-mini, resulting in a final set of 10,108 slide-report pairs. These paired slides are all H&E FFPE WSIs from primary tumors adhering to the same exclusion criteria as mentioned for TCGA-OT. We excluded all frozen tissue slides, slides without magnification information, metastatic or recurrent tumor slides, slides without tumor tissue and IHC slides. Refer to Supplementary Table 125 for a detailed overview of the diagnosis distribution.
Rare-Cancer-Public
The Rare-Cancer-Public is a pan-cancer dataset of H&E FFPE diagnostic WSIs from TCGA96. The dataset consists of 1,982 WSIs, with 1,548 WSIs from TCGA and 434 WSIs from EBRAINS, representing 29 rare cancer types. According to the National Institute of Health, rare cancers are defined as those occurring in fewer than 15 individuals per 100,000 annually44. The OncoTree codes of WSIs from TCGA and EBRAINS were manually curated for this criterion by two expert pathologists (A.K. and D.F.K.W.). EBRAINS provides more granular diagnostic classifications than the OncoTree codes, enabling the dataset to include finer distinctions for rare brain tumors. The dataset was divided into five patient-level folds. To assess retrieval performance for rare cancers within a clinically representative dataset, we use one fold of the rare cancer dataset as the query set and the remaining folds combined with the common cancer types as a support set. In total, the support and query datasets contain 14,062 slides, including 11,646 WSIs from TCGA and 2,416 from EBRAINS.
Rare-Cancer
The Rare-Cancer is an in-house extension of the public dataset Rare-Cancer-Public with MGB internal cases. This dataset comprises 43 rare cancer types and 3,039 H&E FFPE diagnostic histopathology WSIs, where 1,056 additional cases were added from Brigham and Women’s Hospital (BWH). The entire dataset, including common cancer types, comprises 19,626 WSIs, with 5,564 WSIs from BWH, covering 186 OncoTree codes.
Rare-Cancer-External
The Rare-Cancer-External is an external testing cohort for rare cancer cases collected from the Department of Pathology, Kanagawa Cancer Center Hospital, Japan. This dataset consists of 39 H&E FFPE diagnostic WSIs from 12 rare ovarian and soft tissue cancers. The slides were stained using SAKURA TISSUE-TEK PRISMA 6130 Slide Stainer, and scanned by Leica Aperio AT2 at ×20 magnification. Detailed breakdown of the cohort can be found in Supplementary Table 116.
BCNB
The BCNB consists of 1,058 H&E FFPE WSIs of early breast cancer core-needle biopsies141. All cases are annotated with estrogen receptor (ER; WT, 227; MUT, 831), progesterone receptor (PR; WT, 268; MUT, 790) and HER2 (WT, 781; MUT, 277) expressions. We split the dataset label-stratified by a ratio of 60:20:20 (676:170:212 slides).
BRACS
The BRACS consists of 547 H&E FFPE WSIs of benign (including normal), atypical and malignant breast tumors from 189 patients142. The cases are annotated in coarse and fine-grained subtypes of three classes (benign tumors, 265; atypical tumors, 89; malignant tumors, 193) and six classes (atypical ductal hyperplasia, 48; ductal carcinoma in situ, 61; flat epithelial atypia, 41; invasive carcinoma, 132; normal, 44; pathological benign, 147; usual ductal hyperplasia, 74). We split the dataset label-stratified at the patient level into five splits, with a ratio of 60:20:20 (approximately 302:94:151 slides).
Cardiac allograft rejection
The cardiac allograft rejection consists of 5,021 H&E FFPE WSIs of 1,688 patient biopsies collected from BWH24. Each biopsy is labeled for the presence of cardiac rejection, characterized by acute cellular rejection (no rejection, 866 patients; rejection, 822 patients). We split the dataset label-stratified on the patient level into train, val and test splits by a ratio of 70:10:20 (3547:484:990 slides).
DHMC-LUAD
The DHMC-LUAD consists of 143 H&E FFPE WSIs of lung adenocarcinoma (LUAD) from the Department of Pathology and Laboratory Medicine at DHMC100. All WSIs are labeled into five classes of the predominant patterns of LUAD (acinar, 59; lepidic, 19; micropapillary, 9; papillary, 5; solid, 51). Given the limited size of the dataset, we use it exclusively for evaluation in a zero-shot setting, where we use the entire dataset as test set.
DHMC-RCC
The DHMC-RCC consists of 563 H&E FFPE WSIs of renal cell carcinoma (RCC) from DHMC101. All slides are labeled into the four predominant patterns of RCC, including one benign class (renal oncocytoma, chromophobe RCC, clear cell RCC, papillary RCC). We use the three RCC subtypes as an external test set for the three-class subtyping task, TCGA RCC.
EBRAINS
The EBRAINS consists of 2,319 H&E FFPE diagnostic histopathology WSIs from the EBRAINS Digital Tumor Atlas sourced from the University of Vienna143. Due to the small sample size, we exclude two classes and predict a fine-grained 30-class brain tumor subtyping task. All brain tumors in these tasks are designated as rare cancers by the RARECARE project and the NCI-SEER program. For training and evaluation, we approximately label-stratified the dataset into a train-val-test fold with a ratio of 50:25:25 (1,151:595:573 slides). Additionally, we use 873 samples with annotations for isocitrate dehydrogenase 1 (IDH1) mutation as an external test set for IDH1 mutation prediction on the TCGA-Glioblastoma Multiforme and Lower-Grade Glioma (GBMLGG) cohort.
IMP-CRC
The IMP-CRC consists of 5,333 H&E FFPE colorectal biopsy and polypectomy WSIs retrieved from the data archive of IMP Diagnostics laboratory, Portugal144,145,146. All cases are classified into one of the following three categories: non-neoplastic (847 slides), low-grade lesions (2,847 slides), which include conventional adenomas with low-grade dysplasia, and high-grade lesions (1,639 slides), which include conventional adenomas with high-grade dysplasia, intramucosal carcinomas and invasive adenocarcinomas. We split the dataset label-stratified by a ratio of 60:20:20 into train-val-test set (3546:887:900 slides).
MGB-BRCA
The MGB-BRCA consists of 1,264 H&E FFPE WSIs of biopsies and resections of invasive breast cancers (BRCA) from BWH66,124. Each case is annotated with the following three IHC status prediction tasks: ER status prediction (negative, 261; positive, 613), PR status prediction (negative, 37; positive, 504) and HER2 status prediction (negative, 665; positive, 151), where ER, PR and HER2 status were manually extracted from pathology reports.
MGB-LUAD
The MGB-LUAD consists of 1,939 H&E FFPE WSIs of LUAD from BWH66,124. The WSIs are annotated by five molecular tasks with ground truth from IHC—protein 40 (P40) status prediction (negative, 113; positive, 72), protein 63 (P63) status prediction (negative, 72; positive, 81), Napsin A status prediction (negative, 60; positive, 66), caudal type homeobox 2 (CDX2) status prediction (negative, 55; positive, 24) and cytokeratin 5 and 6 (CK-5&6) status prediction (negative, 29; positive, 29).
MGH-BRCA
The MGH-BRCA consists of 1,071 IHC FFPE WSIs of invasive breast carcinoma from Mass General Hospital66. The cases contain annotations for IHC quantification in six expression levels of ER abundance (levels 1–6 with counts—168, 169, 219, 170, 175 and 169, respectively) and PR abundance (levels 1–6 with counts—2,603, 2,397, 1,209, 1,118, 1,124 and 1,101, respectively).
MUT-HET
The MUT-HET consists of 1,291 H&E FFPE WSIs of clear cell RCC, each representing a single patient treated at the Mayo Clinic147,148. All cases are labeled with the following mutations, determined from matched IHC slides—BAP1 mutation (WT, 1,130; MUT, 162), PBRM1 mutation (WT, 622; MUT, 670) and SETD2 mutation (WT, 943; MUT, 349). We split the dataset into five splits with train-val-test ratio of 60:20:20 (774:258:259 slides) in each split.
OT108
The OT108 is an in-house pan-cancer subtyping dataset consisting of 5,564 H&E FFPE diagnostic WSIs from BWH classified into 108 classes according to the OncoTree classification104. We split the dataset into train-val-test (3,164:780:1,620 slides). The test set is balanced across the classes and contains 15 slides per class.
PANDA
The PANDA consists of 10,616 H&E FFPE diagnostic histopathology WSIs of core-needle biopsies of prostate cancer sourced from the Radboud University Medical Center and the Karolinska Institute. Each slide is assigned a score recommended by the International Society of Urological Pathology (ISUP) that defines prostate cancer grade (six-class grading task). For quality control, we follow prior work149 in excluding slides that were erroneously annotated or had noisy labels, resulting in an overall 9,555 slides (grade 0, 2,603; grade 1, 2,399; grade 2, 1,209; grade 3, 1,118; grade 4, 1,124; grade 5, 1,102). For training and evaluation, we label-stratified PANDA into 80:10:10 train-val-test folds (7,645:954:953 slides).
PD-L1
The PD-L1 consists of 234 IHC FFPE diagnostic histopathology WSIs from 217 patients with stage IV nonsmall cell lung cancer (NSCLC) who initiated treatment with anti-PD-(L)1 blockade therapy between 2014 and 2019 at Memorial Sloan Kettering Cancer Center150. Patients who received chemotherapy concurrently with immunotherapy were not included. We used the clinical PD-L1 assessments as labels and substituted these labels by pathologist re-annotations on 157 slides when available. Following the original study, we created three levels of PD-L1 expression (<1%, 62; 1–50%, 49; ≥50%, 123) as target predictions. We split the dataset into five splits with train-val-test ratio of 60:20:20 (129:44:44 slides) in each split.
Renal allograft rejection
The renal allograft rejection consists of 4,847 H&E FFPE WSIs of renal allograft biopsies from 1,118 patients collected at BWH between 2013 and 2022. Each case has associated labels for antibody-mediated rejection (AMR) status (AMR, 286 patients; no AMR, 832 patients), cellular-mediated rejection (cellular rejection, 341; no cellular rejection, 777) and interstitial fibrosis and tubular atrophy (IFTA) status (advanced IFTA, 162 patients; mild IFTA, 706 patients; moderate IFTA, 250 patients). We split the dataset into a label-stratified train-val-test set (3002:376:824 slides).
TCGA-BRCA
The TCGA-BRCA consists of 1,049 invasive breast carcinoma (BRCA) H&E FFPE diagnostic histopathology WSIs from TCGA. The WSIs are classified into the following two classes: invasive ductal carcinoma and invasive lobular carcinoma.
TCGA-NSCLC
The TCGA-NSCLC consists of 1,043 H&E FFPE diagnostic histopathology WSIs from TCGA of 946 patients with NSCLC. The WSIs are classified into the following two classes: LUAD (531 slides) and lung squamous cell carcinoma (512 slides). We split the dataset into fivefold cross-validation, stratified by labels with a ratio of 60:20:20 (for example, 659:191:193 for fold 0). CPTAC-NSCLC serves as an external dataset with 1,091 H&E FFPE diagnostic histopathology WSIs from CPTAC of 422 patients with NSCLC.
TCGA-LUAD
The TCGA-LUAD consists of 524 H&E FFPE diagnostic histopathology WSIs from TCGA of 462 patients with LUAD. We predict the mutations in the genes EGFR (wild type (WT), 404 patients; mutated (MUT), 58 patients), KRAS (WT, 317; MUT, 145), STK11 (WT, 391; MUT, 71) and TP53 (WT, 222; MUT, 240). We split the dataset into fivefold cross-validation, stratified by labels with a ratio of 60:20:20 (for example, 659:191:193 for fold 0). CPTAC-LUAD serves as an external dataset with 324 H&E FFPE diagnostic histopathology WSIs from CPTAC of 108 patients with LUAD.
TCGA-CRC
The TCGA-CRC consists of 549 H&E FFPE diagnostic histopathology WSIs from TCGA of 543 patients with colorectal cancer (CRC). We predict microsatellite instability (61 patients) and microsatellite stable (353 patients), mutations in the genes BRAF (WT, 429 patients; MUT, 58 patients) and KRAS (WT, 286 patients; MUT, 201 patients), and tumor staging (T1, 16 slides; T2, 97 slides; T3, 372 slides; T4, 64 slides). CPTAC-COAD with 107 H&E FFPE diagnostic histopathology WSIs from 103 patients with colon adenocarcinoma serves as external validation dataset for all tasks (microsatellite instability, 24 patients; microsatellite stable, 79 patients; BRAF WT, 16 patients; BRAF MUT, 87 patients; KRAS WT, 36 patients; KRAS MUT, 58 patients; T2, 17 slides; T2, 77 slides; T4, 13 slides).
TCGA-GBMLGG
The TCGA-GBMLGG consists of 1,123 H&E FFPE diagnostic histopathology WSIs from TCGA of 558 patients with gliomas, more specifically GBMLGG. The WSIs are classified into the following two classes: IDH1 mutation (425 slides) and no IDH1 mutation (698 slides). EBRAINS serves as an external cohort for this task (IDH1 MUT, 333 slides; IDH1 WT, 540 slides).
Computing software and hardware
We used Python (version 3.9.16) for all experiments and analyses in the study, which can be replicated using open-source libraries as outlined below. We used PyTorch (version 2.0.1, CUDA 11.8) for training and inference of our deep learning model. To train TITANV and TITAN, we modified the public implementation of iBOT (http://github.com/bytedance/ibot) and CoCa (http://github.com/mlfoundations/open_clip). We used 4× and 8× 80GB NVIDIA A100 GPUs configured for multi-GPU training using distributed data parallelism for TITANV and TITAN training, respectively. All downstream experiments were conducted on a single 24GB NVIDIA 3090 GPUs. All WSI processing was supported by OpenSlide (version 4.3.1), openslide-python (version 1.2.0) and CLAM (http://github.com/mahmoodlab/CLAM). We used Scikit-learn (version 1.2.2) for its implementation of k-NN, and the logistic regression implementation and SimpleShot implementation provided by the LGSSL codebase (http://github.com/mbanani/lgssl). For survival tasks, we used scikit-survival (Version 0.23.1). Implementations of other slide encoders benchmarked in the study are found at the following links: GigaPath (http://github.com/prov-gigapath/prov-gigapath), PRISM (https://huggingface.co/paige-ai/Prism) and CHIEF (http://github.com/hms-dbmi/CHIEF). For training weakly-supervised ABMIL models, we adapted the training scaffold code from the CLAM codebase (http://github.com/mahmoodlab/CLAM). Matplotlib (version 3.8.4) and Seaborn (version 0.13.2) were used to create plots in Figs. 1–4. Usage of other miscellaneous Python libraries is listed in the Reporting summary.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
GTEx data used in pretraining can be accessed through the GTEx portal (https://www.gtexportal.org/home/). For benchmarks, TCGA and CPTAC data can be accessed through the NIH genomic data commons (https://portal.gdc.cancer.gov) and proteomics data commons (https://proteomic.datacommons.cancer.gov), respectively. Coordinates and labels of TCGA-UniformTumor-8K dataset is made publicly available in the TITAN GitHub repository. All other publicly available datasets benchmarked in this work can be can accessed in their respective data portals: EBRAINS (https://doi.org/10.25493/WQ48-ZGX), DHMC RCC (https://bmirds.github.io/KidneyCancer), DHMC LUAD (https://bmirds.github.io/LungCancer/), BRACS (https://bracs.icar.cnr.it), PANDA (https://panda.grand-challenge.org), IMP (https://rdm.inesctec.pt/dataset/nis-2023-008), BCNB (https://bupt-ai-cz.github.io/BCNB/), MUT-HET-RCC (https://aacrjournals.org/cancerres/article/82/15/2792/707325/Intratumoral-Resolution-of-Driver-Gene-Mutation). Links for all public datasets are also presented in Supplementary Table 17. Following institution policies, all requests for data collected or curated in-house will be evaluated on a case-by-case basis to determine whether the data requested is compliant with intellectual property and patient privacy obligations. Data can only be shared for academic research purposes and will require a material transfer agreement.
Code availability
Code and model weights for loading both TITAN and TITANV can be accessed for academic research purposes at https://github.com/mahmoodlab/TITAN.
References
Song, A. H. et al. Artificial intelligence for digital and computational pathology. Nat. Rev. Bioeng. 1, 930–949 (2023).
Riasatian, A. et al. Fine-tuning and training of DenseNet for histopathology image representation using TCGA diagnostic slides. Med. Image Anal. 70, 102032 (2021).
Ciga, O., Xu, T. & Martel, A. L. Self supervised contrastive learning for digital histopathology. Mach. Learn. Appl. 7, 100198 (2022).
Wang, X. et al. Transformer-based unsupervised contrastive learning for histopathological image classification. Med. Image Anal. 81, 102559 (2022).
Wang, X. et al. RetCCL: clustering-guided contrastive learning for whole-slide image retrieval. Med. Image Anal. 83, 102645 (2023).
Kang, M., Song, H., Park, S., Yoo, D. & Pereira, S. Benchmarking self-supervised learning on diverse pathology datasets. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) 3344–3354 (IEEE, 2023).
Huang, Z., Bianchi, F., Yuksekgonul, M., Montine, T. J. & Zou, J. A visual–language foundation model for pathology image analysis using medical Twitter. Nat. Med. 29, 2307–2316 (2023).
Azizi, S. et al. Robust and data-efficient generalization of self-supervised machine learning for diagnostic imaging. Nat. Biomed. Eng. 7, 756–779 (2023).
Chen, R. J. et al. Towards a general-purpose foundation model for computational pathology. Nat. Med. 30, 850–862 (2024).
Lu, M. Y. et al. A visual-language foundation model for computational pathology. Nat. Med. 30, 863–874 (2024).
Vorontsov, E. et al. A foundation model for clinical-grade computational pathology and rare cancers detection. Nat. Med. 30, 2924–2935 (2024).
Zimmermann, E. et al. Virchow2: scaling self-supervised mixed magnification models in pathology. Preprint at https://arxiv.org/abs/2408.00738 (2024).
Saillard, C. et al. bioptimus/releases. GitHub http://github.com/bioptimus/releases/tree/main/models/h-optimus/v0 (2024).
Filiot, A. et al. Scaling self-supervised learning for histopathology with masked image modeling. Preprint at medRxiv https://doi.org/10.1101/2023.07.21.23292757 (2023).
Filiot, A., Jacob, P., Kain, A. M. & Saillard, C. Phikon-v2, a large and public feature extractor for biomarker prediction. Preprint at https://arxiv.org/abs/2409.09173 (2024).
Juyal, D. et al. Pluto: pathology-universal transformer. In ICML 2024 Workshop on Foundation Models in the Wild. https://openreview.net/forum?id=7yn50e6tVX (OpenReview, 2024).
Dippel, J. et al. Rudolfv: a foundation model by pathologists for pathologists. Preprint at https://arxiv.org/abs/2401.04079 (2024).
Bera, K., Schalper, K. A., Rimm, D. L., Velcheti, V. & Madabhushi, A. Artificial intelligence in digital pathology—new tools for diagnosis and precision oncology. Nat. Rev. Clin. Oncol. 16, 703–715 (2019).
Echle, A. et al. Deep learning in cancer pathology: a new generation of clinical biomarkers. Br. J. Cancer 124, 686–696 (2021).
Shmatko, A., Ghaffari Laleh, N., Gerstung, M. & Kather, J. N. Artificial intelligence in histopathology: enhancing cancer research and clinical oncology. Nat. Cancer 3, 1026–1038 (2022).
Campanella, G. et al. A clinical benchmark of public self-supervised pathology foundation models. Nat. Commun. 16, 3640 (2025).
Neidlinger, P. et al. Benchmarking foundation models as feature extractors for weakly-supervised computational pathology. Preprint at https://arxiv.org/abs/2408.15823 (2024).
Yu, K.-H. et al. Predicting non-small cell lung cancer prognosis by fully automated microscopic pathology image features. Nat. Commun. 7, 12474 (2016).
Lipkova, J. et al. Deep learning-enabled assessment of cardiac allograft rejection from endomyocardial biopsies. Nat. Med. 28, 575–582 (2022).
Coudray, N. et al. Classification and mutation prediction from non-small cell lung cancer histopathology images using deep learning. Nat. Med. 24, 1559–1567 (2018).
Campanella, G. et al. Clinical-grade computational pathology using weakly supervised deep learning on whole slide images. Nat. Med. 25, 1301–1309 (2019).
Kather, J. N. et al. Deep learning can predict microsatellite instability directly from histology in gastrointestinal cancer. Nat. Med. 25, 1054–1056 (2019).
Lu, M. Y. et al. AI-based pathology predicts origins for cancers of unknown primary. Nature 594, 106–110 (2021).
Fu, Y. et al. Pan-cancer computational histopathology reveals mutations, tumor composition and prognosis. Nat. Cancer 1, 800–810 (2020).
Bulten, W. et al. Artificial intelligence for diagnosis and Gleason grading of prostate cancer: the PANDA Challenge. Nat. Med. 28, 154–163 (2022).
Zheng, Y. et al. A graph-transformer for whole slide image classification. IEEE Trans. Med. Imaging 41, 3003–3015 (2022).
Skrede, O.-J. et al. Deep learning for prediction of colorectal cancer outcome: a discovery and validation study. Lancet 395, 350–360 (2020).
Bejnordi, B. E. et al. Diagnostic assessment of deep learning algorithms for detection of lymph node metastases in women with breast cancer. JAMA 318, 2199–2210 (2017).
Wagner, S. J. et al. Transformer-based biomarker prediction from colorectal cancer histology: a large-scale multicentric study. Cancer Cell 41, 1650–1661 (2023).
Foersch, S. et al. Multistain deep learning for prediction of prognosis and therapy response in colorectal cancer. Nat. Med. 29, 430–439 (2023).
Courtiol, P. et al. Deep learning-based classification of mesothelioma improves prediction of patient outcome. Nat. Med. 25, 1519–1525 (2019).
Lee, Y. et al. Derivation of prognostic contextual histopathological features from whole-slide images of tumours via graph deep learning. Nat. Biomed. Eng. 6, 1452–1466 (2022).
Niehues J. M. et al. Generalizable biomarker prediction from cancer pathology slides with self-supervised deep learning: a retrospective multi-centric study Cell Rep. Med. 4 100980 (2023).
Yu, K.-H., Beam, A. L. & Kohane, I. S. Artificial intelligence in healthcare. Nat. Biomed. Eng. 2, 719–731 (2018).
Krishnan, R., Rajpurkar, P. & Topol, E. J. Self-supervised learning in medicine and healthcare. Nat. Biomed. Eng. 6, 1346–1352 (2022).
Campanella, G. et al. Real-world deployment of a fine-tuned pathology foundation model for lung cancer biomarker detection. Nat. Med. 31, 3002–3010 (2025).
Van der Laak, J., Litjens, G. & Ciompi, F. Deep learning in histopathology: the path to the clinic. Nat. Med. 27, 775–784 (2021).
Gatta, G. et al. Burden and centralised treatment in Europe of rare tumours: results of RARECAREnet—a population-based study. Lancet Oncol. 18, 1022–1039 (2017).
NCI Dictionary of Cancer Terms: rare cancer. National Cancer Institute https://www.cancer.gov/publications/dictionaries/cancer-terms/def/rare-cancer (accessed 20 November 2024).
Surveillance, Epidemiology, and End Results Program Rare cancer classification. National Cancer Institute https://seer.cancer.gov/seerstat/variables/seer/raresiterecode/ (accessed 21 November 2024).
Lew, M. S., Sebe, N., Djeraba, C. & Jain, R. Content-based multimedia information retrieval: State of the art and challenges. ACM Trans. Multimed. Comput. Commun. Appl. 2, 1–19 (2006).
Cruz-Roa, A., Caicedo, J. C. & González, F. A. Visual pattern mining in histology image collections using bag of features. Artif. Intell. Med. 52, 91–106 (2011).
Caicedo, J. C., Cruz, A. & Gonzalez, F. A. Histopathology image classification using bag of features and kernel functions. In Proc. 12th Conference on Artificial Intelligence in Medicine (eds Combi, C., et al.) 126–135 (Springer, 2009).
Sridhar, A., Doyle, S. & Madabhushi, A. Content-based image retrieval of digitized histopathology in boosted spectrally embedded spaces. J. Pathol. Inform. 6, 41 (2015).
Kalra, S. et al. Yottixel—an image search engine for large archives of histopathology whole slide images. Med. Image Anal. 65, 101757 (2020).
Chen, C. et al. Fast and scalable search of whole-slide images via self-supervised deep learning. Nat. Biomed. Eng. 6, 1420–1434 (2022).
Zheng, Y. et al. Histopathological whole slide image analysis using context-based CBIR. IEEE Trans. Med. Imaging 37, 1641–1652 (2018).
Shang, H. H. et al. Histopathology slide indexing and search—are we there yet? NEJM AI 1, AIcs2300019 (2024).
Zhang, Z., Xie, Y., Xing, F., McGough, M. & Yang, L. Mdnet: a semantically and visually interpretable medical image diagnosis network. In Proc. IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 6428–6436 (IEEE, 2017).
Lu, M. Y. et al. Visual language pretrained multiple instance zero-shot transfer for histopathology images. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) 19764–19775 (IEEE, 2023).
Ikezogwo W. et al. Quilt-1m: one million image–text pairs for histopathology Adv. Neural Inf. Process. Syst. 36 37995–38017 (2024).
Chen, R. J. et al. Scaling vision transformers to gigapixel images via hierarchical self-supervised learning. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) 16144–16155 (IEEE, 2022).
Xu H. et al. A whole-slide foundation model for digital pathology from real-world data Nature 630 181–188 (2024).
Lazard, T., Lerousseau, M., Decencière, E. & Walter, T. Giga-SSL: self-supervised learning for gigapixel images. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) 4304–4313 (IEEE, 2023).
Hou, X. et al. A self-supervised framework for learning whole slide representations. Preprint at https://arxiv.org/abs/2402.06188 (2024).
Tran, M. et al. Generating dermatopathology reports from gigapixel whole slide images with Histogpt. Nat. Commun. 16, 4886 (2025).
Shaikovski, G. et al. PRISM: a multi-modal generative foundation model for slide-level histopathology. Preprint at https://arxiv.org/abs/2405.10254 (2024).
Xu, Y. et al. A multimodal knowledge-enhanced whole-slide pathology foundation model. Preprint at https://arxiv.org/abs/2407.15362 (2024).
Jaume, G. et al. Transcriptomics-guided slide representation learning in computational pathology. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) 9632–9644 (IEEE, 2024).
Song, A. H. et al. Morphological prototyping for unsupervised slide representation learning in computational pathology. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) 11566–11578 (IEEE, 2024).
Jaume, G. Multistain pretraining for slide representation learning in pathology. In Proc. European Conference on Computer Vision—ECCV 2024 (eds Leonardis, A. et al.) 19–37 (Springer, 2024).
Vaidya, A. et al. Molecular-driven foundation model for oncologic pathology. Preprint at https://arxiv.org/abs/2501.16652 (2025).
Shao, D. et al. Do multiple instance learning models transfer? In Proc. 42nd International Conference on Machine Learning, (eds Singh, A. et al.) 54219–54238 (PMLR, 2025).
Pyeon, M. et al. Exaone path 2.0: Pathology foundation model with end-to-end supervision. Preprint at https://arxiv.org/abs/2507.06639 (2025).
Lenz, T. et al. Unsupervised foundation model-agnostic slide-level representation learning. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) 30807–30817 (IEEE, 2025).
Neidlinger, P. et al. A deep learning framework for efficient pathology image analysis. Preprint at https://arxiv.org/abs/2502.13027 (2025).
Shaikovski, G. et al. Prism2: unlocking multi-modal general pathology AI with clinical dialogue. Preprint at https://arxiv.org/abs/2506.13063 (2025).
Ilse, M., Tomczak, J. M. & Welling, M. Attention-based deep multiple instance learning. In Proc. 35th International Conference on Machine Learning (eds Dy, J. & Krause, A.) 2127–2136 (PMLR, 2018).
Lu, M. Y. et al. Data-efficient and weakly supervised computational pathology on whole-slide images. Nat. Biomed. Eng. 5, 555–570 (2021).
Li, B., Li, Y. & Eliceiri, K. W. Dual-stream multiple instance learning network for whole slide image classification with self-supervised contrastive learning. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) 14318–14328 (IEEE, 2021).
Chen, R. J. et al. Whole slide images are 2D point clouds: context-aware survival prediction using patch-based graph convolutional networks. In Proc. International Conference on Medical Image Computing and Computer Assisted Intervention—MICCAI 2021 (eds de Bruijne, M. et al.) 339–349 (Springer, 2021).
Shao, Z. et al. Transmil: transformer based correlated multiple instance learning for whole slide image classification. Adv. Neural Inf. Process. Syst. 34, 2136–2147 (2021).
Carmichael, I. et al. Incorporating intratumoral heterogeneity into weakly-supervised deep learning models via variance pooling. In Proc. International Conference on Medical Image Computing and Computer-Assisted Intervention—MICCAI 2022 (eds Wang, L. et al.) 387–397 (Springer, 2022).
Xiang, J. & Zhang, J. Exploring low-rank property in multiple instance learning for whole slide image classification. Proceedings of the 11th International Conference on Learning Representations https://openreview.net/forum?id=01KmhBsEPFO (OpenReview, 2023).
Yang, S., Wang, Y. & Chen, H. Mambamil: enhancing long sequence modeling with sequence reordering in computational pathology. In Proc. International Conference on Medical Image Computing and Computer-Assisted Intervention—MICCAI 2024 (eds Linguraru, M.G. et al.) 296–306 (Springer, 2024).
Kondepudi A. et al. Foundation models for fast, label-free detection of glioma infiltration Nature 637 439–445 (2025).
Ahmed, F. et al. PathAlign: a vision-language model for whole slide images in histopathology. In Proc. MICCAI Workshop on Computational Pathology (eds Ciompi, F. et al.) 72–108 (PMLR, 2024).
Wang X. et al. A pathology foundation model for cancer diagnosis and prognosis prediction Nature 634 970–978 (2024).
Bommasani, R. et al. On the opportunities and risks of foundation models. Preprint at https://arxiv.org/abs/2108.07258 (2021).
Moor, M. et al. Foundation models for generalist medical artificial intelligence. Nature 616, 259–265 (2023).
Zhou, J. et al. iBOT: image BERT pre-training with online tokenizer. In Proc. International Conference on Learning Representations https://openreview.net/forum?id=ydopy-e6Dg (OpenReview, 2022).
Oquab, M. et al. Dinov2: learning robust visual features without supervision. Trans. Mach. Learn. Res. https://openreview.net/forum?id=a68SUt6zFt (2024).
Lu M. Y. et al. A multimodal generative AI copilot for human pathology Nature 634 466–473 (2024).
Chen, R. J., Lu, M. Y., Chen, T. Y., Williamson, D. F. & Mahmood, F. Synthetic data in machine learning for medicine and healthcare. Nat. Biomed. Eng. 5, 493–497 (2021).
Kokosi, T. & Harron, K. Synthetic data in medical research. BMJ Med. 1, e000167 (2022).
Carrillo-Perez F. et al. Generation of synthetic whole-slide image tiles of tumours from RNA-sequencing data via cascaded diffusion models Nat. Biomed. Eng. 9 320–332 (2025).
Dosovitskiy, A. et al. An image is worth 16 × 16 words: transformers for image recognition at scale. In Proc. International Conference on Learning Representations https://openreview.net/forum?id=YicbFdNTTy (OpenReview, 2021).
Bär, A., Houlsby, N., Dehghani, M. & Kumar, M. Frozen feature augmentation for few-shot image classification. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) 16046–16057 (IEEE, 2024).
Press, O., Smith, N. & Lewis, M. Train short, test long: attention with linear biases enables input length extrapolation. In Proc. International Conference on Learning Representations https://openreview.net/forum?id=R8sQPpGCv0 (OpenReview, 2022).
Yu J. et al. CoCa: contrastive captioners are image-text foundation models Trans. Mach. Learn. Res. https://openreview.net/forum?id=Ee277P3AYC (2022).
Weinstein, J. N. et al. The Cancer Genome Atlas pan-cancer analysis project. Nat. Genet. 45, 1113–1120 (2013).
Liu, J. et al. An integrated TCGA pan-cancer clinical data resource to drive high-quality survival outcome analytics. Cell 173, 400–416 (2018).
Edwards, N. J. et al. The CPTAC data portal: a resource for cancer proteomics research. J. Proteome Res. 14, 2707–2713 (2015).
Thangudu R. R. et al. Abstract LB-242: proteomic data commons: a resource for proteogenomic analysis Cancer Res. 80 LB-242 (2020).
Wei J. W et al. Pathologist-level classification of histologic patterns on resected lung adenocarcinoma slides with deep neural networks Sci. Rep. 9 3358 (2019).
Zhu M et al. Development and evaluation of a deep neural network for histologic classification of renal cell carcinoma on biopsy and surgical resection slides Sci. Rep. 11 7080 (2021).
Kapoor S., & Narayanan A. Leakage and the reproducibility crisis in machine-learning-based science Patterns (N Y) 4 100804 (2023).
Song, A. H. et al. Multimodal prototyping for cancer survival prediction. In Proc. 41st International Conference on Machine Learning (eds Salakhutdinov, R. et al.) 46050–46073 (PMLR, 2024).
Kundra, R. et al. Oncotree: a cancer classification system for precision oncology. JCO Clin. Cancer Inform. 5, 221–230 (2021).
Wang, Y., Chao, W.-L., Weinberger, K. Q. & Van Der Maaten, L. Simpleshot: revisiting nearest-neighbor classification for few-shot learning. Preprint at arXiv https://arxiv.org/abs/1911.04623 (2019).
Snell, J., Swersky, K. & Zemel, R. Prototypical networks for few-shot learning. In Proc. 31st International Conference on Neural Information Processing Systems (eds Guyon, I. et al.) 4080–4090 (Curran Associates, 2017).
Naeini, M. P., Cooper, G. & Hauskrecht, M. Obtaining well calibrated probabilities using Bayesian binning. In Proc. 29th AAAI Conference on Artificial Intelligence 2901–2907 (AAAI, 2015).
Radford, A. et al. Learning transferable visual models from natural language supervision. In Proc. 38th International Conference on Machine Learning (eds Meila, M. & Zhang, T.) 8748–8763 (PMLR, 2021).
Kefeli J., & Tatonetti N. TCGA-Reports: a machine-readable pathology report resource for benchmarking text-based AI models Patterns (N Y) 5 100933 (2024).
Banerjee, S. & Lavie, A. METEOR: an automatic metric for MT evaluation with improved correlation with human judgments. In Proc. ACL Workshop on Intrinsic and Extrinsic Evaluation Measures for Machine Translation and/or Summarization (ed Goldstein, J. et al) 65–72 (Association for Computational Linguistics, 2005).
Lin, C.-Y. Rouge: a package for automatic evaluation of summaries. In Text Summarization Branches Out: Proceedings of the ACL-04 Workshop 74–81 (Association for Computational Linguistics, 2004).
Papineni, K., Roukos, S., Ward, T. & Zhu, W.-J. Bleu: a method for automatic evaluation of machine translation. In Proc. 40th Annual Meeting of the Association for Computational Linguistics 311–318 (Association for Computational Linguistics, 2002).
Komura, D. & Ishikawa, S. Machine learning methods for histopathological image analysis. Comput. Struct. Biotechnol. J. 16, 34–42 (2018).
Hegde, N. et al. Similar image search for histopathology: SMILY. NPJ Digit. Med. 2, 56 (2019).
Neumann, H. P., Young Jr, W. F. & Eng, C. Pheochromocytoma and paraganglioma. N. Engl. J. Med. 381, 552–565 (2019).
Koch, V. et al. DinoBloom: a foundation model for generalizable cell embeddings in hematology. In Proc. International Conference on Medical Image Computing and Computer-Assisted Intervention (eds Linguraru, M. G. et al.) 520–530 (Springer, 2024).
Jaume, G. et al. Hest-1k: A dataset for spatial transcriptomics and histology image analysis. Adv. Neural Inf. Process. Syst. 37, 53798–53833 (2024).
Song, A. H. et al. Analysis of 3D pathology samples using weakly supervised AI. Cell 187, 2502–2520 (2024).
Shaban, M. et al. A foundation model for spatial proteomics. Preprint at https://arxiv.org/abs/2506.03373 (2025).
De Jong, E. D., Marcus, E. & Teuwen, J. Current pathology foundation models are unrobust to medical center differences. Preprint at https://arxiv.org/abs/2501.18055 (2025).
Filiot, A. et al. Distilling foundation models for robust and efficient models in digital pathology. Preprint at https://arxiv.org/abs/2501.16239 (2025).
Kömen, J. et al. Towards robust foundation models for digital pathology. Preprint at https://arxiv.org/abs/2507.17845 (2025).
Gindra, R. H. et al. Image analysis: understanding and mitigating batch effects in histopathology. Trillium Pathol. 4, 20–25 (2025).
Vaidya, A. et al. Demographic bias in misdiagnosis by computational pathology models. Nat. Med. 30, 1174–1190 (2024).
Liu, J. et al. Hasd: hierarchical adaption for pathology slide-level domain-shift. Preprint at https://arxiv.org/abs/2506.23673 (2025).
Nechaev, D., Pchelnikov, A. & Ivanova, E. Hibou: a family of foundational vision transformers for pathology. Preprint at https://arxiv.org/abs/2406.05074 (2024).
Brodsky, V. et al. Generative artificial intelligence in anatomic pathology. Arch. Pathol. Lab. Med. 149, 298–318 (2025).
Lonsdale, J. et al. The genotype-tissue expression (GTEx) project. Nat. Genet. 45, 580–585 (2013).
The GTEx Consortium et al. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015).
Yang, A. et al. Qwen2 technical report. Preprint at https://arxiv.org/abs/2407.10671 (2024).
Zaffar, I., Jaume, G., Rajpoot, N. & Mahmood, F. Embedding space augmentation for weakly supervised learning in whole-slide images. In Proc. 2023 IEEE 20th International Symposium on Biomedical Imaging (ISBI) https://doi.org/10.1109/ISBI53787.2023.10230723 (IEEE, 2023).
Shao, Z., Dai, L., Wang, Y., Wang, H. & Zhang, Y. Augdiff: diffusion based feature augmentation for multiple instance learning in whole slide image. IEEE Trans. Artif. Intell. 5, 6617–6628 (2024).
Jaume, G., Song, A. H. & Mahmood, F. Integrating context for superior cancer prognosis. Nat. Biomed. Eng. 6, 1323–1325 (2022).
Beyer, L. et al. PaliGemma: a versatile 3B VLM for transfer. Preprint at https://arxiv.org/abs/2407.07726 (2024).
He, K. et al. Masked autoencoders are scalable vision learners. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) 16000–16009 (IEEE, 2022).
Jaegle, A. et al. Perceiver: general perception with iterative attention. In Proc. 38th International Conference on Machine Learning (eds Meila, M. & Zhang, T.) 4651–4664 (PMLR, 2021).
Liu, Z. et al. Swin transformer: hierarchical vision transformer using shifted windows. In Proc. IEEE/CVF International Conference on Computer Vision 10012–10022 (IEEE, 2021).
Caron, M. et al. Emerging properties in self-supervised vision transformers. In Proc. IEEE/CVF International Conference on Computer Vision 9650–9660 (IEEE, 2021).
Zadeh, S. G. & Schmid, M. Bias in cross-entropy-based training of deep survival networks. IEEE Trans. Pattern Anal. Mach. Intell. 43, 3126–3137 (2020).
Howard, F. M. et al. The impact of site-specific digital histology signatures on deep learning model accuracy and bias. Nat. Commun. 12, 4423 (2021).
Xu F. et al. Predicting axillary lymph node metastasis in early breast cancer using deep learning on primary tumor biopsy slides Front. Oncol. 11 759007 (2021).
Brancati, N. et al. Bracs: a dataset for breast carcinoma subtyping in H&E histology images. Database 2022, baac093 (2022).
Roetzer-Pejrimovsky, T. et al. The Digital Brain Tumour Atlas, an open histopathology resource. Sci. Data 9, 55 (2022).
Oliveira S. P et al. CAD systems for colorectal cancer from WSI are still not ready for clinical acceptance Sci. Rep. 11 14358 (2021).
Neto, P. C. et al. iMIL4PATH: a semi-supervised interpretable approach for colorectal whole-slide images. Cancers 14, 2489 (2022).
Neto, P. C. et al. An interpretable machine learning system for colorectal cancer diagnosis from pathology slides. NPJ Precis. Oncol. 8, 56 (2024).
Joseph, R. W. et al. Clear cell renal cell carcinoma subtypes identified by BAP1 and PBRM1 expression. J. Urol. 195, 180–187 (2016).
Acosta, P. H. et al. Intratumoral resolution of driver gene mutation heterogeneity in renal cancer using deep learning. Cancer Res. 82, 2792–2806 (2022).
Pati, P. et al. Weakly supervised joint whole-slide segmentation and classification in prostate cancer. Med. Image Anal. 89, 102915 (2023).
Vanguri, R. S. et al. Multimodal integration of radiology, pathology and genomics for prediction of response to PD-(L) 1 blockade in patients with non-small cell lung cancer. Nat. Cancer 3, 1151–1164 (2022).
Acknowledgements
This study was funded in part by the BWH President’s Fund, MGH Pathology and by the National Institute of Health (NIH) National Institute of General Medical Sciences (NIGMS; R35GM138216 to F.M.). S.J.W. was supported by the Helmholtz Association under the joint research school ‘Munich School for Data Science—MUDS’ and the Add-on Fellowship of the Joachim Herz Foundation. This work was supported by a fellowship of the German Academic Exchange Service (DAAD). M.Y.L. was supported by the Tau Beta Pi Fellowship and the Siebel Foundation. This work was additionally supported by the AMED Practical Research for Innovative Cancer Control (grant JP 25ck0106873 to S.I.). We thank K. Ono at the Department of Pathology, Kanagawa Cancer Center Hospital, Kanagawa, Japan, for her contribution to the case collection of the Rare Cancer External dataset. The content is solely the responsibility of the authors and does not reflect the official views of the NIH, NIGMS, NCI and DoD.
Author information
Authors and Affiliations
Contributions
T.D., S.J.W., A.H.S., R.J.C. and F.M. conceived the study and designed the experiments. L.P.L., T.D., R.J.C. and B.C. curated the Mass-340K WSIs and corresponding pathology reports. R.J.C., S.J.W., A.H.S., A.J.V., G.J., C.A.-P., P.D. and C.S.C. scanned the WSIs. T.D., S.J.W., R.J.C. and A.H.S. developed the stage 1 vision-only TITAN model. T.D., R.J.C., M.Y.L., S.J.W. and A.H.S. developed the stage 2 and stage 3 vision-language TITAN models. T.D., S.J.W. and M.Y.L. developed the codebase for zero-shot vision-language slide understanding. T.D., S.J.W., A.H.S., A.Z., A.J.V. and G.J. implemented the benchmarking codebase for pretrained slide models. A.K. and D.F.K.W. evaluated the synthetic captions and generated reports, and helped with the study design for slide retrieval. A.K., D.F.K.W. and C.C. curated the rare disease retrieval dataset. R.J.C., D.K., A.K. and S.I. curated and annotated the TCGA-Uniform-8K dataset. H.R. ran the statistical significance tests on all evaluation tasks. S.S. curated the renal allograft rejection dataset. D.K., M.O., S.S., T.K., Y.M. and S.I. curated and processed the Rare Cancer External dataset. A.Z., A.J.V. and G.J. contributed equally. T.D., S.J.W., A.H.S., R.J.C. and F.M. prepared the manuscript. L.P.L. and F.M. supervised the research. All authors contributed to the writing of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
R.J.C., M.Y.L., D.F.K.W., B.C., L.P.L. and F.M. hold equity interests in ModellaAI. The other authors declare no competing interests.
Peer review
Peer review information
Nature Medicine thanks Julien Calderaro, Olivier Elemento and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Examples of TCGA-UT-8K dataset.
Examples of TCGA-UT-8K, which are ROIs of 8,192 × 8,192 pixels selected by the pathologists. The green contours illustrate the cancer region annotations, with the red number indicating the ROI index within a given TCGA slide.
Extended Data Fig. 2 Linear probe results for molecular classification tasks.
(a) Linear models are fitted and evaluated on binary molecular status predictions for BCNB and MUT-HET. We observe that TITAN consistently performs best with + 0.9% on BCNB and MUT-HET, +1.7% on TCGA, and +3.7% on internal molecular classification of BRCA and LUAD, in averaged AUROC scores over the next best model PRISM. (b) Linear models are fitted and evaluated on five-fold splits on TCGA. (c) The same models are evaluated on the corresponding external datasets from CPTAC and EBRAINS. (d) 6-level ER and PR prediction from Mass General Hospital (MGH) and 3-level PD-L1 prediction, all from immunohistochemistry (IHC) slides. (e) Molecular classification tasks for BRCA and LUAD from Mass General Brigham (MGB). All error bars represent standard deviations based on bootstrapping (n = 1,000) or k-fold evaluation (k = 5).
Extended Data Fig. 3 UMAP of slide embedding space for TCGA-OT.
UMAP visualization of slide embeddings in TCGA-OT cohort (n = 11,186) for all slide encoder baselines, including TITAN and TITANV, color-coded by different organs for visual decluttering.
Extended Data Fig. 4 UMAP of TCGA-OT slide representations (n = 11,186) from all slide encoders.
The first row is labeled by OncoTreeCode, the second row by OncoTreeSiteCode, and the third row by submission site. Clustering metrics, mean local diversity (mLD), adjusted rand index (ARI), and normalized mutual information (NMI), are computed for all labels. Note that CHIEF includes TCGA in the pretraining dataset.
Extended Data Fig. 5 Attention heatmaps of TITAN.
Exemplar attention heatmaps for three Transformer attention heads of TITAN (head #4, #10, #11) are shown across three different TCGA WSIs. Out of the 12 attention heads, we find that most attention heads focus on dense tumor regions, with certain attention heads such as head #10 focusing on tumor-adjacent stroma and head #11 focusing on non-tumor areas. Across different cancer types, while head #11 attends to tissue-specific morphologies such as peritumoral stroma in the thymoma WSI and the tumor-adjacent stroma and ducts in the BRCA WSI, we do observe that general morphological patterns such as tumor/non-tumor are conserved across tissue types.
Extended Data Fig. 6 Ablation experiments on different learning paradigms.
Change in balanced accuracy performance for several learning paradigms on four subtyping tasks with respect to the linear probe. The baselines include mean pooling, ABMIL, linear probe, and finetuned from pretrained or randomly initialized weights. The number under each task name indicates the linear probe performance. TITAN-L represents the variation of TITAN without vision-pretraining. For mean pooling and ABMIL, we use the respective patch encoder for each framework, as specified under each slide encoder name. Finetuning results are not provided for PRISM, as the finetuning recipes were not available.
Extended Data Fig. 7 Examples of generated reports.
TCGA examples of generated reports of TITAN and PRISM, with the corresponding clinical reports.
Extended Data Fig. 8 Rare cancer retrieval with TITAN.
(a)–(c) Examples of slide retrieval on Rare-Cancer. The number for each retrieved slide represents the cosine similarity between the query and the retrieved slide. The retrieved slides with high similarity are either of the same diagnostic label or from the same organ as the query slide. (a) Thyroid (THFO) query (b) Pleura (PLBMESO) query (c) Adrenal gland (ACC) query.
Supplementary information
Supplementary Information (download PDF )
Supplementary Tables 1–127.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ding, T., Wagner, S.J., Song, A.H. et al. A multimodal whole-slide foundation model for pathology. Nat Med 31, 3749–3761 (2025). https://doi.org/10.1038/s41591-025-03982-3
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41591-025-03982-3
This article is cited by
-
Leveraging multi-modal foundation models for analysing spatial multi-omic and histopathology data
Nature Biomedical Engineering (2026)
-
Mucin phenotype-based deep learning framework for intestinal metaplasia-carcinogenesis progression prediction
npj Precision Oncology (2025)