Abstract
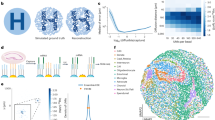
Imaging-based spatial transcriptomics methods allow for the measurement of spatial determinants of cellular phenotypes but are incompatible with random barcode-based clone-tracing methods, preventing the simultaneous detection of clonal and spatial drivers. Here we report SpaceBar, which enables simultaneous clone tracing and spatial gene expression profiling with standard imaging-based spatial transcriptomics pipelines. Our approach uses a library of 96 synthetic barcode sequences that combinatorially labels each cell. Thus, SpaceBar can distinguish between clonal dynamics and environmentally driven transcriptional regulation in complex tissue contexts.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
Processed data are available with the paper and via Github at https://github.com/grantkinsler/SpatialBarcodes.
Raw data, which consist of extremely large files, are available upon request from the authors.
Code availability
Code for analysis and figure generation are available via GitHub at https://github.com/grantkinsler/SpatialBarcodes.
References
Ben-Moshe, S. et al. Spatial sorting enables comprehensive characterization of liver zonation. Nat. Metab. 1, 899–911 (2019).
Moor, A. E. et al. Spatial reconstruction of single enterocytes uncovers broad zonation along the intestinal villus axis. Cell 175, 1156–1167 (2018).
Shaffer, S. M. et al. Memory sequencing reveals heritable single-cell gene expression programs associated with distinct cellular behaviors. Cell 182, 947–959 (2020).
Oren, Y. et al. Cycling cancer persister cells arise from lineages with distinct programs. Nature 596, 576–582 (2021).
Emert, B. L. et al. Variability within rare cell states enables multiple paths toward drug resistance. Nat. Biotechnol. 39, 865–876 (2021).
Biddy, B. A. et al. Single-cell mapping of lineage and identity in direct reprogramming. Nature 564, 219–224 (2018).
Jindal, K. et al. Single-cell lineage capture across genomic modalities with CellTag-multi reveals fate-specific gene regulatory changes. Nat. Biotechnol. 42, 946–959 (2024).
Vickovic, S. et al. High-definition spatial transcriptomics for in situ tissue profiling. Nat. Methods 16, 987–990 (2019).
Ståhl, P. L. et al. Visualization and analysis of gene expression in tissue sections by spatial transcriptomics. Science 353, 78–82 (2016).
Ratz, M. et al. Clonal relations in the mouse brain revealed by single-cell and spatial transcriptomics. Nat. Neurosci. 25, 285–294 (2022).
Jones, M. G. et al. Spatiotemporal lineage tracing reveals the dynamic spatial architecture of tumor growth and metastasis. Preprint at bioRxiv https://doi.org/10.1101/2024.10.21.619529 (2024).
Shah, S., Lubeck, E., Zhou, W. & Cai, L. In situ transcription profiling of single cells reveals spatial organization of cells in the mouse hippocampus. Neuron 92, 342–357 (2016).
Eng, C.-H. L. et al. Transcriptome-scale super-resolved imaging in tissues by RNA seqFISH. Nature 568, 235–239 (2019).
Chen, K. H., Boettiger, A. N., Moffitt, J. R., Wang, S. & Zhuang, X. RNA imaging. Spatially resolved, highly multiplexed RNA profiling in single cells. Science 348, aaa6090 (2015).
Zhang, M. et al. Spatially resolved cell atlas of the mouse primary motor cortex by MERFISH. Nature 598, 137–143 (2021).
Frieda, K. L. et al. Synthetic recording and in situ readout of lineage information in single cells. Nature 541, 107–111 (2017).
Askary, A. et al. In situ readout of DNA barcodes and single base edits facilitated by in vitro transcription. Nat. Biotechnol. 38, 66–75 (2020).
Koblan, L. W. et al. High-resolution spatial mapping of cell state and lineage dynamics in vivo with PEtracer. Science 390, eadx3800 (2025).
Yuan, L., Chen, X., Zhan, H., Henry, G. L. & Zador, A. M. Massive multiplexing of spatially resolved single neuron projections with axonal BARseq. Nat. Commun. 15, 8371 (2024).
Lubeck, E., Coskun, A. F., Zhiyentayev, T., Ahmad, M. & Cai, L. Single-cell in situ RNA profiling by sequential hybridization. Nat. Methods 11, 360–361 (2014).
Stringer, C., Wang, T., Michaelos, M. & Pachitariu, M. Cellpose: a generalist algorithm for cellular segmentation. Nat. Methods 18, 100–106 (2021).
Goyal, Y. et al. Diverse clonal fates emerge upon drug treatment of homogeneous cancer cells. Nature 620, 651–659 (2023).
Boe, R. H., Triandafillou, C. G., Lazcano, R., Wargo, J. A. & Raj, A. Spatial transcriptomics reveals influence of microenvironment on intrinsic fates in melanoma therapy resistance. Preprint at bioRxiv https://doi.org/10.1101/2024.06.30.601416 (2024).
Pérez-Gutiérrez, L. & Ferrara, N. Biology and therapeutic targeting of vascular endothelial growth factor A. Nat. Rev. Mol. Cell Biol. 24, 816–834 (2023).
Hemmerlein, B. et al. Vascular endothelial growth factor expression, angiogenesis, and necrosis in renal cell carcinomas. Virchows Arch. 439, 645–652 (2001).
Fane, M. E. et al. Stromal changes in the aged lung induce an emergence from melanoma dormancy. Nature 606, 396–405 (2022).
Brown, J. M. & Wilson, W. R. Exploiting tumour hypoxia in cancer treatment. Nat. Rev. Cancer 4, 437–447 (2004).
Acknowledgements
We thank C. Triandafillou and members of the Raj laboratory for helpful discussions and feedback on the work. G.K. acknowledges support from NIH Training Grant T32-CA-009140. A.R. acknowledges support from the Samuel Waxman Cancer Research Foundation, the Mark Foundation, the Melanoma Research Alliance and the Templeton Foundation (63532). R.H.B. acknowledges support from the NIH Training Grant in Computational Genomics T32HG000046 and the NIH Medical Scientist Training Program T32GM007170. S.S. is a Bakewell Foundation Innovator of the Damon Runyon Cancer Research Foundation (DRR-81-24). S.S. also acknowledges support from Wistar/Penn Skin Cancer SPORE Career Enhancement Program P50 CA261608 and the American Cancer Society (RSG-23-1152597-01-CDP).
Author information
Authors and Affiliations
Contributions
G.K. and Y.H. designed SpaceBar, performed experiments and conducted analysis. G.K., C.F. and Y.H. performed cloning, validation and initial experiments. H.L., J.K. and M.D. performed mouse experiments, with supervision from M.H. R.V.V. designed and cloned the backbone plasmid, with supervision from S.S. R.B. assisted with panel design and spatial analysis. A.R. provided supervision and editing. G.K. and Y.H. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
A.R. receives royalties related to Stellaris RNA FISH probes. A.R. serves on the scientific advisory board of Spatial Genomics. All other authors declare no competing interests.
Peer review
Peer review information
Nature Methods thanks Alejo Rodriguez-Fraticelli, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Madhura Mukhopadhyay, in collaboration with the Nature Methods team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Detection of barcodes with smFISH.
Microscopy image depicting single-molecule FISH of melanoma cells transduced with a lentiviral pool containing 3 of the SpaceBar barcodes (barcodes 1, 2, and 3). DAPI is shown in gray. Probes targeting barcodes 1, 2, and 3 were coupled to Atto 674n (pseudocolored purple), Cy3 (pseudocolored orange), and Alexa Fluor 594 (pseudocolored green), respectively. Merge of all channels shown in A. Each barcode channel is shown separately in B-D.
Extended Data Fig. 2 Uniqueness of barcode combinations, clustering reduces barcode mis-detection.
A. Top 99th percentile distance between clone cells vs the number of barcodes integrated per cell. We use the value for four barcodes as the distance threshold above which the clone calling would be considered “false clones”. B. “False clone” rate reflected by the frequency of clone with above threshold size. C. Comparing the mis-assignment rate (reflected by neighbor barcode overlap) between clustered and unclustered data vs number of barcodes per cell.
Extended Data Fig. 3 Barcode abundance distribution.
A histogram that depicts the number of cells that express each of the barcodes, measured in in vitro data.
Extended Data Fig. 4 In vitro, cells of the same clone have more similar gene expression than random cells.
Depicts histograms for the mean pairwise distance in PCA space (of the top 10 PCs) of cells that belong to the same clone. The blue histogram shows the main pairwise distance for actual clones. The gray histogram depicts cells randomly assigned to clones (keeping the number of clones and cells per clone as the real distribution).
Extended Data Fig. 5 SpaceBar identifies clonal structure in several xenograft tumors.
Each panel depicts a section of a melanoma xenograft from a different mouse. Each dot depicts a barcode transcript, colored by the barcode identity. A. Tissue section 2 (a different section from the same tumor as tumor section 1 discussed throughout the main text). B-D. Depict tumor sections 3-5, respectively.
Extended Data Fig. 6 Tumor section images.
A. GFP channel from the first hybridization round of the focal xenograft section. Barcoded cells are GFP+ and autofluorescence shows the vascularized interior of the tumor. B. DAPI channel of the section.
Extended Data Fig. 7 SFRP1 clonal structure would be difficult to detect without high resolution.
A. Expression of SFRP1 in the neighborhood of Clone 31 as depicted in Fig. 2, with grids spaced 55µm apart. B. A series of heatmaps that depict the expression pattern of SFRP1 as well as the distribution of spots that label distinct clones in this area of the tumor section. Clones are depicted as the sum of all transcripts for barcodes assigned to that clone. While only clone 31 has true clonal SFRP1 gene expression from our high-resolution data, all nine depicted clones have significant correlations (p < 0.01) with SFRP1 expression when binned at 55µm resolution.
Extended Data Fig. 8 IFIT2 has high expression in nearby cells of the same clone, suggesting transiently heritable expression.
The expression of IFIT2 is shown in the tumor section. Color indicates the expression level per cell; cells with zero or one spot are depicted in gray. The first inset depicts the expression of IFIT2 in the neighborhood of Clone 72 (clone 72 cells outlined in red). The second inset depicts the expression of IFIT2 in the neighborhood of Clone 15 (clone 15 cells outlined in red). IFIT2 expression is high in groups of 3-5 cells within the same clone, suggesting its expression is transiently heritable.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figures, Tables and Notes.
Supplementary Data 1 (download XLSX )
Supplementary data file which includes all of the clone and space scores calculated for the tumor sections.
Supplementary Table 1 (download XLSX )
Supplementary file which includes the barcode sequences, probe oligo sequences for smFISH, and the GenePS panel information.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kinsler, G., Fagan, C., Li, H. et al. SpaceBar enables single-cell-resolution clone tracing with imaging-based spatial transcriptomics. Nat Methods 23, 328–333 (2026). https://doi.org/10.1038/s41592-025-02968-w
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41592-025-02968-w