Abstract
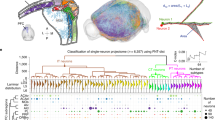
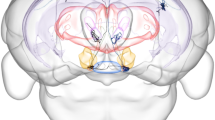
The hypothalamus coordinately regulates physiological homeostasis and innate behaviors, yet the detailed arrangement of hypothalamic axons remains unclear. Here we mapped the whole-brain projections of over 7,000 hypothalamic neurons expressing distinct neuropeptides in male mice, identifying 2 main classes and 31 types using single-neuron projectome analysis. These classes/types exhibited regionally biased soma distribution and specific neuropeptide enrichment. Notably, many projectome types extended long-range axon collaterals to distinct brain regions, allowing single axons to co-regulate multiple targets. We uncovered topographic organization of certain peptidergic axons at specific targets, along with diverse single-neuron projectome patterns in Orexin, Agrp and Pomc populations. Furthermore, hypothalamic peptidergic neurons showed correlated innervation of subdomains in the periaqueductal gray and organized into modular subnetworks within the hypothalamus, providing a structural basis for coordinated outputs. This dataset highlights the complexity of hypothalamic axonal projections and lays a foundation for future investigation of the circuit mechanisms underlying hypothalamic functions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The reconstructed projectome data for the 7,180 neurons are publicly accessible on our websites: https://mouse.digital-brain.cn/projectome/hy and https://www.braindatacenter.cn/datacenter/web/#/dataSet/details?id=1800731789745954817. Source data are provided with this paper.
Code availability
A Python library containing computer code used for feature calculations is available at https://pypi.org/project/pyswcloader/. Additionally, all source code for analyzing these features and for generating each figure can be found at https://zenodo.org/records/14588094 (ref. 68), which is organized according to the panels in each figure. All other data and codes are available from the corresponding authors upon reasonable request.
Change history
02 December 2025
A Correction to this paper has been published: https://doi.org/10.1038/s41593-025-02184-x
References
Biörklund, A., Hökfelt, T. & Swanson, L. W. Integrated Systems of the CNS, Part I: Hypothalamus, Hippocampus, Amygdala, Retina (Elsevier Science, 1987).
Saper, C. B. & Lowell, B. B. The hypothalamus. Curr. Biol. 24, R1111–R1116 (2014).
Simerly, R. B. in The Rat Nervous System 4th edn (ed. Paxinos, G.) Ch. 13 (Elsevier, 2015).
Puelles, L., Martinez-de-la-Torre, M., Bardet, S. & Rubenstein, J. L. R. in The Mouse Nervous System (eds Watson, C., Paxinos, G. & Puelles, L.) Ch. 8 (Academic Press, 2012).
Hoyle, C. H. Neuropeptide families and their receptors: evolutionary perspectives. Brain Res. 848, 1–25 (1999).
Van Den Pol, A. N. Neuropeptide transmission in brain circuits. Neuron 76, 98–115 (2012).
Canteras, N. S., Simerly, R. B. & Swanson, L. W. Projections of the ventral premammillary nucleus. J. Comp. Neurol. 324, 195–212 (1992).
Canteras, N. S., Simerly, R. B. & Swanson, L. W. Organization of projections from the ventromedial nucleus of the hypothalamus: a Phaseolus vulgaris-leucoagglutinin study in the rat. J. Comp. Neurol. 348, 41–79 (1994).
Canteras, N. S. & Swanson, L. W. Projections of the ventral subiculum to the amygdala, septum, and hypothalamus: a PHAL anterograde tract-tracing study in the rat. J. Comp. Neurol. 324, 180–194 (1992).
Goto, M., Canteras, N. S., Burns, G. & Swanson, L. W. Projections from the subfornical region of the lateral hypothalamic area. J. Comp. Neurol. 493, 412–438 (2005).
Risold, P. Y., Canteras, N. S. & Swanson, L. W. Organization of projections from the anterior hypothalamic nucleus: a Phaseolus vulgaris-leucoagglutinin study in the rat. J. Comp. Neurol. 348, 1–40 (1994).
Risold, P. Y., Thompson, R. H. & Swanson, L. W. The structural organization of connections between hypothalamus and cerebral cortex. Brain Res. Rev. 24, 197–254 (1997).
Thompson, R. H., Canteras, N. S. & Swanson, L. W. Organization of projections from the dorsomedial nucleus of the hypothalamus: a PHA-L study in the rat. J. Comp. Neurol. 376, 143–173 (1996).
Thompson, R. H. & Swanson, L. W. Organization of inputs to the dorsomedial nucleus of the hypothalamus: a reexamination with Fluorogold and PHAL in the rat. Brain Res. Brain Res. Rev. 27, 89–118 (1998).
Peyron, C. et al. Neurons containing hypocretin (orexin) project to multiple neuronal systems. J. Neurosci. 18, 9996–10015 (1998).
Wang, D. et al. Whole-brain mapping of the direct inputs and axonal projections of POMC and AgRP neurons. Front. Neuroanat. 9, 40 (2015).
Biglari, N. et al. Functionally distinct POMC-expressing neuron subpopulations in hypothalamus revealed by intersectional targeting. Nat. Neurosci. 24, 913–929 (2021).
Quarta, C. et al. POMC neuronal heterogeneity in energy balance and beyond: an integrated view. Nat. Metab. 3, 299–308 (2021).
Bittencourt, J. C. Anatomical and functional heterogeneity of ‘hypothalamic’ peptidergic neuron populations. Nat. Rev. Endocrinol. 18, 450–450 (2022).
Steuernagel, L. et al. HypoMap—a unified single-cell gene expression atlas of the murine hypothalamus. Nat. Metab. 4, 1402–1419 (2022).
Yao, Z. et al. A high-resolution transcriptomic and spatial atlas of cell types in the whole mouse brain. Nature 624, 317–332 (2023).
Langlieb, J. et al. The molecular cytoarchitecture of the adult mouse brain. Nature 624, 333–342 (2023).
Li, H. et al. Single-neuron projectomes of mouse paraventricular hypothalamic nucleus oxytocin neurons reveal mutually exclusive projection patterns. Neuron 112, 1081–1099 (2024).
Gong, H. et al. High-throughput dual-colour precision imaging for brain-wide connectome with cytoarchitectonic landmarks at the cellular level. Nat. Commun. 7, 12142 (2016).
Qiu, S. et al. Whole-brain spatial organization of hippocampal single-neuron projectomes. Science 383, eadj9198 (2024).
Gao, L. et al. Single-neuron projectome of mouse prefrontal cortex. Nat. Neurosci. 25, 515–529 (2022).
Wang, Q. et al. The Allen Mouse Brain Common Coordinate Framework: a 3D reference atlas. Cell 181, 936–953 (2020).
Sunkin, S. M. et al. Allen Brain Atlas: an integrated spatio-temporal portal for exploring the central nervous system. Nucleic Acids Res. 41, D996–D1008 (2013).
Costa, M., Manton, J. D., Ostrovsky, A. D., Prohaska, S. & Jefferis, G. S. NBLAST: rapid, sensitive comparison of neuronal structure and construction of neuron family databases. Neuron 91, 293–311 (2016).
Peng, H. et al. Morphological diversity of single neurons in molecularly defined cell types. Nature 598, 174–181 (2021).
Mizrahi, A. et al. Comparative analysis of dendritic architecture of identified neurons using the Hausdorff distance metric. J. Comp. Neurol. 422, 415–428 (2000).
Engelmann, M., Landgraf, R. & Wotjak, C. T. The hypothalamic–neurohypophysial system regulates the hypothalamic–pituitary–adrenal axis under stress: an old concept revisited. Front. Neuroendocrinol. 25, 132–149 (2004).
Swaab, D. F., Pool, C. W. & Nijveldt, F. Immunofluorescence of vasopressin and oxytocin in the rat hypothalamo-neurohypophyseal system. J. Neural Transm. 36, 195–215 (1975).
Winnubst, J. et al. Reconstruction of 1,000 projection neurons reveals new cell types and organization of long-range connectivity in the mouse brain. Cell 179, 268–281 (2019).
Xu, S. et al. Behavioral state coding by molecularly defined paraventricular hypothalamic cell type ensembles. Science 370, eabb2494 (2020).
Wang, Y. et al. EASI-FISH for thick tissue defines lateral hypothalamus spatio-molecular organization. Cell 184, 6361–6377 (2021).
Lanciego, J. L., Luquin, N. & Obeso, J. A. Functional neuroanatomy of the basal ganglia. Cold Spring Harb. Perspect. Med. 2, a009621 (2012).
Wallen-Mackenzie, A. et al. Spatio-molecular domains identified in the mouse subthalamic nucleus and neighboring glutamatergic and GABAergic brain structures. Commun. Biol. 3, 338 (2020).
Kita, H. & Kitai, S. T. Efferent projections of the subthalamic nucleus in the rat: light and electron microscopic analysis with the PHA-L method. J. Comp. Neurol. 260, 435–452 (1987).
Sato, F., Parent, M., Levesque, M. & Parent, A. Axonal branching pattern of neurons of the subthalamic nucleus in primates. J. Comp. Neurol. 424, 142–152 (2000).
Fanselow, M. S. & Dong, H. W. Are the dorsal and ventral hippocampus functionally distinct structures? Neuron 65, 7–19 (2010).
Dillingham, C. M., Frizzati, A., Nelson, A. J. & Vann, S. D. How do mammillary body inputs contribute to anterior thalamic function? Neurosci. Biobehav. Rev. 54, 108–119 (2015).
Hou, Y. et al. Topographical organization of mammillary neurogenesis and efferent projections in the mouse brain. Cell Rep. 34, 108712 (2021).
Jbabdi, S., Sotiropoulos, S. N. & Behrens, T. E. The topographic connectome. Curr. Opin. Neurobiol. 23, 207–215 (2013).
Thivierge, J. P. & Marcus, G. F. The topographic brain: from neural connectivity to cognition. Trends Neurosci. 30, 251–259 (2007).
Sakurai, T. The role of orexin in motivated behaviours. Nat. Rev. Neurosci. 15, 719–731 (2014).
Sternson, S. M. & Eiselt, A.-K. Three pillars for the neural control of appetite. Annu. Rev. Physiol. 79, 401–423 (2017).
Toda, C., Santoro, A., Kim, J. D. & Diano, S. POMC neurons: from birth to death. Annu. Rev. Physiol. 79, 209–236 (2017).
Tervo, D. G. et al. A designer AAV variant permits efficient retrograde access to projection neurons. Neuron 92, 372–382 (2016).
Betley, J. N., Cao, Z. F. H., Ritola, K. D. & Sternson, S. M. Parallel, redundant circuit organization for homeostatic control of feeding behavior. Cell 155, 1337–1350 (2013).
Bandler, R. & Shipley, M. T. Columnar organization in the midbrain periaqueductal gray: modules for emotional expression? Trends Neurosci. 17, 379–389 (1994).
Koutsikou, S., Apps, R. & Lumb, B. M. Top down control of spinal sensorimotor circuits essential for survival. J. Physiol. 595, 4151–4158 (2017).
Tovote, P. et al. Midbrain circuits for defensive behaviour. Nature 534, 206–212 (2016).
Han, W. et al. Integrated control of predatory hunting by the central nucleus of the amygdala. Cell 168, 311–324 (2017).
Kohl, J. et al. Functional circuit architecture underlying parental behaviour. Nature 556, 326–331 (2018).
Hahn, J. D., Sporns, O., Watts, A. G. & Swanson, L. W. Macroscale intrinsic network architecture of the hypothalamus. Proc. Natl Acad. Sci. USA 116, 8018–8027 (2019).
Canteras, N. S. The medial hypothalamic defensive system: hodological organization and functional implications. Pharmacol. Biochem. Behav. 71, 481–491 (2002).
Swanson, L. W. Cerebral hemisphere regulation of motivated behavior. Brain Res. 886, 113–164 (2000).
Choi, G. B. et al. Lhx6 delineates a pathway mediating innate reproductive behaviors from the amygdala to the hypothalamus. Neuron 46, 647–660 (2005).
Stagkourakis, S. et al. A neural network for intermale aggression to establish social hierarchy. Nat. Neurosci. 21, 834–842 (2018).
Park, H.-J. & Friston, K. Structural and functional brain networks: from connections to cognition. Science 342, 1238411 (2013).
Anderson, D. J. Circuit modules linking internal states and social behaviour in flies and mice. Nat. Rev. Neurosci. 17, 692–704 (2016).
Sun, P. et al. Highly efficient and super-bright neurocircuit tracing using vector mixing-based virus cocktail. Preprint at bioRxiv https://doi.org/10.1101/705772 (2020).
Li, X.-Y. et al. AGRP neurons project to the medial preoptic area and modulate maternal nest-building. J. Neurosci. 39, 456–471 (2019).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2023).
Rubinov, M. & Sporns, O. Complex network measures of brain connectivity: uses and interpretations. NeuroImage 52, 1059–1069 (2010).
Preibisch, S., Saalfeld, S. & Tomancak, P. Globally optimal stitching of tiled 3D microscopic image acquisitions. Bioinformatics 25, 1463–1465 (2009).
Jiao, Z., Gao, T. & Xu, X. Projectome-based characterization of hypothalamic peptidergic neurons in male mice. Zenodo https://doi.org/10.5281/zenodo.14588094 (2025).
Acknowledgements
We thank Z. Liang for valuable discussions during paper preparation. This work was supported by the National Science and Technology Innovation 2030 Major Program (2021ZD0204400 to Y.-G.S., 2021ZD0203200-03 to X.X., 2021ZD0201000 to H.G., 2021ZD0200100 to Y.S. and 2021ZD0204401 to S.X.); the National Science Foundation of China (32321003 and 32371072 to S.X.); the Shanghai Municipal Science and Technology Major Project (2018SHZDZX05 to X.X.); and the Lingang Laboratory (LG202104-01-01 to Y.-G.S. and LG202104-01-04 to X.X.). The funders had no role in study design, data collection and analysis, preparation of the manuscript or decision to publish.
Author information
Authors and Affiliations
Contributions
Conceptualization: M.-m.P., Y.-G.S. and X.X. Experiments: virus injections, X.D., Z.Y., M.L., M.H., H.Z., X.C., S.L., C.W., E.L. Y.H., Z.T. and H.L.; fMOST imaging, T.J., J.Q., X.J., Z.F., A.L., Q.L. and H.G.; EASI-FISH, A.W. and S.X.; RNAscope, X.D., M.L., M.H., H.Z., X.C., S.L., C.W., E.L., Y.H., Z.T., H.L. and M.G.; and fMOST data curation, Xinran Wang and Y.C. Data analysis: fMOST neurite tracing, L. Gou, B.R., X.S., D.W., L.H., Y.L. and Xiaofei Wang; fMOST data visualization and comparative analysis, Z.J., T.G., L. Gao, N.B., E.E.B., L.S., S.M.S., J.C.B., M.-m.P., D.J.A., J.Y. and X.X.; single-cell transcriptome analysis, Y.M., L.F. and Y.S.; and EASI-FISH data analysis, A.W., Z.J., T.G. and S.X. Website construction: L.Q., C.J., J.H. and W.D. Figure plotting: Z.J., T.G., W.Z., Z.Y. and M.L. Supervision: X.Y., M.X., H.-C.C., Y.Z., H.X., S.X., Y.S., Y.-G.S., X.X., Q.L. and H.G. Paper writing: R.S., M.-m.P. and X.X.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Neuroscience thanks Dayu Lin and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Viral strategies for sparsely labeling hypothalamic peptidergic neurons.
a. To sparsely label neuropeptide-expressing populations listed at the bottom, the indicated virus or virus mixture was injected into wild-type mice to label Orexin-expressing neurons or into Cre-expressing mice to label all other peptidergic populations. b. Representative images showing co-localization of EYFP or GFP signal with in situ or immunohistochemistry signals of the indicated neuropeptide (red). The quantification shows the percentage (%) of EYFP or GFP-labeled cells that co-expressed the neuropeptide. Data are presented as mean ± S.E.M. N indicates the number of biological replicates. The images on the right show magnified regions within the white box on the left. Scale bar, left, 100 μm; right, 20 μm.
Extended Data Fig. 2 Soma distribution of each peptidergic population.
a. 3D view of the soma distribution of neurons across the 16 peptidergic populations within various hypothalamic nuclei (outlined on the contralateral side and color-coded). The number of reconstructed neurons for each peptidergic population is indicated in the parenthesis. b. Bar graphs showing the number (top) and the density (bottom) of reconstructed neurons with somas located in the indicated hypothalamic nuclei.
Extended Data Fig. 3 Transcriptional diversity of hypothalamic peptidergic neuron populations.
Heatmap representation of normalized neuropeptide expression (columns) across 1,451 hypothalamic neuron subtypes (rows), referred to as “cell clusters” in the original manuscript21. The names of the neuropeptides are indicated above each column. Subtypes are color-coded, labeled on the left, and grouped into 31 cell types, which are color-coded with the legend on the right. A neuropeptide is considered enriched in a cell subtype if more than 25% of neurons within that subtype express it. Among the 1,451 neuron subtypes, 368 showed enrichments for at least one of the 16 neuropeptides: 306 subtypes are enriched for only one neuropeptide, 55 are enriched for two neuropeptides, and 7 are enriched for three neuropeptides.
Extended Data Fig. 4 Characteristics of non-midbrain projecting types 1–15.
a. A dot plot depicting selective projections of each type (row) in each brain area (column). The size of the dot indicates the percentage of neurons in each type that project to the indicated brain area, and the color intensity represents the average projection length (in μm) in a heatmap fashion, with the scale on the right. b–j. The total axon projections of neurons of the indicated type are plotted in a 3D brain on the left. Zoom-in views of representative axon projections from three representative neurons (color-coded) from each type in defined subdomains of the indicated target areas are shown on the right. Scale bar, 500 μm.
Extended Data Fig. 5 Characteristics of the midbrain projecting types 16–31.
a. A dot plot depicting selective projections of each type (row) in each brain area (column). The dot circle’s size and color intensity indicate the percentage of neurons in each type that project to the indicated brain area and the average projection length (in μm) in a heatmap fashion, respectively, with the scale on the right. b–i. The total axon projections of neurons of the indicated type are plotted in a 3D brain on the left. Zoom-in views of representative axon projections from three representative neurons (color-coded) from each type in defined subdomains of the indicated target areas are shown on the right. Scale bar, 500 μm. j. Schematics showing the nuclei and column structures of the PAG, which includes anterior PAG (aPAG) and rostral nuclei (INC, Su3, ND, and PRC), as well as four longitudinally organized PAG columns - dorsomedial (dm), dorsolateral (dl), lateral (l), and ventrolateral (vl) PAG. k. A dot plot depicting selective projections of each type (row) in each PAG column/nucleus on the ipsilateral or contralateral side. The dot circle’s size and color intensity indicate the percentage of neurons in each type that project to the indicated brain area and the average projection length (in μm) in each column/nucleus in a heatmap fashion with the scale shown on the right. l. Zoom-in views of representative axon projections of three neurons (color-coded) from the indicated type in a coronal (top) and sagittal (bottom) plane of a PAG section. Scale bar, 500 μm. Dashed lines show the position of the coronal section on the sagittal plane. m. Correlation (corr) analysis of preference index in the ipsilateral (ipsi) and contralateral side (contra) of the indicated PAG column/nucleus for bilaterally PAG-projecting neurons. Representative images on the bottom show a coordinated innervation pattern of a similar column/nucleus in both hemispheres by two bilaterally PAG-projecting neurons. n. Heatmap representation (sorted) of column/nucleus preference score for projections in the ipsilateral PAG, calculated as the fold difference of projection density (total arbor length divided by the volume of targeted area) for a particular column/nucleus relative to the average projection density for all areas, of individual neurons in each type.
Extended Data Fig. 6 Topographic arrangements of axon projections of type 4 neurons expressing Adcyap1 in the ventral CA1.
a. No topographic correspondence between the soma and terminal distribution in dorsal CA1 in non-Penk type 4 neurons. b. Adcyap1-expressing type 4 neurons predominantly projected to ventral CA1 while Penk-expressing type 4 neurons predominantly projected to dorsal CA1. c. Correlation analysis between the soma distribution of Adcyap1-expressing type 4 neurons and their termini in ventral CA1 reveals a topographic correspondence between the soma distribution along the D-V axis and their termini distribution along the M-L axis. A linear regression analysis was conducted in panels a and c to examine the relationship between the soma position and the center position of projections in downstream targets along the specified axis. The “r” values represent the regression coefficient, and their significance (P values) was evaluated using a two-tailed t-test.
Extended Data Fig. 7 Complex topographic arrangements of axon projections of type 22 neurons.
a. Correlation analysis of the soma distribution of various neuropeptide populations (indicated by colors) within MBO type 22 neurons and their axon terminal distribution in ATN reveals intricate topographic arrangements across all three axes. b. Illustration of the collective arrangement of axon collateral projections from all MBO type 22 neurons in and around VTN in reference to the soma locations along the three axes. No discernible topographic correspondence was observed. c. Correlation analysis between the soma distribution of specific neuropeptide populations within MBO type 22 neurons along the Medial-Lateral (M-L) axis and their ATN terminals along the Dorsal-Ventral (D-V) axis shows topographic axon arrangements for Tac2-expressing neurons but not Nts-expressing neurons. A linear regression analysis was conducted in a and c to examine the relationship between the soma position and the center position of projections in downstream targets along the specified axis. The “r” values represent the regression coefficient, and their significance (P values) was evaluated using a two-tailed t-test.
Extended Data Fig. 8 Characteristics of projectome-defined Orexin neuron types.
a. Soma locations of LHA Orexin neurons from a published MERFISH dataset21. These neurons were identified as a single transcriptome-defined type among the 1,451 transcriptionally defined neuronal types found in the hypothalamus. b. Violin plots of individual neurons’ projection length (in μm) in specific cortical areas for the five projectome types. Each circle represents an individual neuron. c. Illustration of the relative distribution of axons from individual Orexin neurons within each type across the four designated projecting routes (color-coded). d. Correlation (corr) analysis of the projection length of individual Orexin neurons in different cortical domains identifies co-innervated areas, as highlighted in red boxes.
Extended Data Fig. 9 Axon projections of arcuate Pomc and Agrp neurons.
a. Schematics showing the strategy to label PAG-projecting arcuate Pomc and Agrp neurons by injecting retroAAVs expressing Cre-GFP into a reporter mouse line expressing Cre-inducible histone-GFP. b. Representative images and quantification showing a higher percentage of Pomc neurons being labeled with GFP with the strategy described in a. The fluorescent images show immunostaining of GFP along with florescent in situ signals of Agrp and Pomc. Regions within the white box are highlighted on the right with zoom-in views. Scale bar, 100 μm. Data are presented as mean ± S.E.M., with individual data points shown as circles. N = 4 for either group. The P value above the bar graphs was determined by two-tailed t-test. c. Correspondence between Pomc and Agrp projectomes types and the 31 projectome types defined based on the entire dataset. d. An upset plot showing the intersection size of Agrp neurons that send axons to multiple brain areas indicated on the left. e. Axon projections of two representative Agrp neurons in 3D brain view on the left and zoom-in views of axon in the indicated target areas for either neuron on the right. Both neurons project to multiple targets.
Extended Data Fig. 10 Prominent PAG projections from hypothalamic Peptidergic Neurons to PAG.
a. The total length (in μm) of axon projections from all reconstructed neurons in various midbrain areas demonstrates the highest projection strength observed in PAG. b. Heatmap depiction illustrating the enrichment of soma location across indicated hypothalamic nuclei (columns) for the newly defined PAG subdomains (rows). Only log2 fold change (LogFC) values reaching significance (p < 0.05) are shown.
Supplementary information
Supplementary Tables 1–6 (download XLSX )
Supplementary Table 1. Details of brain samples. Supplementary Table 2. Nomenclature and abbreviations of brain structures. Supplementary Table 3. Total neuron counts for each projectome type and the percentage expressing each of the 16 neuropeptides. Supplementary Table 4. Correlation coefficients (corr) between axon projection lengths and synaptic terminal counts across brain regions and areas. Correlations across all brain areas are highly significant, with P values approaching zero. Supplementary Table 5. Alignment of 77 MouseLight34 hypothalamic neurons with neurons in the current dataset. Supplementary Table 6. Summary of statistical analyses for all bar graphs in the paper, including sample sizes, statistical tests performed and exact P values.
Source data
Source Data Fig. 1 (download ZIP )
Dissimilarity_matrix.csv: Dissimilarity matrix of the 7,180 hypothalamic neurons. all_neurons_projection_length.csv: Projection length of the 7,180 hypothalamic neurons in each target area.
Source Data Fig. 2 (download XLSX )
Soma location and the type information of the 7,180 hypothalamic neurons.
Source Data Fig. 4 (download CSV )
Terminal information of the type 2, type 4 and type 22 neurons.
Source Data Fig. 5 (download XLSX )
Soma location and the type information of the Orexin neurons.
Source Data Fig. 6 (download XLSX )
Soma location and the type information of the Agrp/Pomc neurons.
Source Data Extended Data Fig. 5 (download XLSX )
Projection length of the 7,180 hypothalamic neurons in the subregion of the PAG.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Jiao, Z., Gao, T., Wang, X. et al. Projectome-based characterization of hypothalamic peptidergic neurons in male mice. Nat Neurosci 28, 1073–1088 (2025). https://doi.org/10.1038/s41593-025-01919-0
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41593-025-01919-0
This article is cited by
-
Hypothalamic control of arousal
Reviews in Endocrine and Metabolic Disorders (2026)
-
Activation of hypothalamic-pontine-spinal pathway promotes locomotor initiation and functional recovery after spinal cord injury in mice
Nature Communications (2025)
-
Brain-wide input-output analysis of tuberal nucleus somatostatin neurons reveals hierarchical circuits for orchestrating feeding behavior
Nature Communications (2025)
-
From identity to function: unveiling the cellular complexity of hypothalamic feeding circuits
Reviews in Endocrine and Metabolic Disorders (2025)