Abstract
Compared to individuals carrying two copies of the ε4 allele of apolipoprotein E (APOE), ε2 homozygotes have an approximate 99% reduction in late-onset Alzheimer’s disease (AD) risk. Here we develop a knock-in model that allows for an inducible ‘switch’ between risk and protective alleles (APOE4s2). Gene expression and proteomic analyses confirm that APOE4s2 mice synthesize E4 at baseline and E2 after tamoxifen administration. A whole-body allelic switch results in a metabolic profile resembling E2/E2 humans and drives AD-relevant alterations in the lipidome and single-cell transcriptome, particularly in astrocytes. Finally, when crossed to the 5xFAD background, astrocyte-specific E4 to E2 switching improves cognition, decreases amyloid pathology, lowers gliosis and reduces plaque-associated apolipoprotein E. Together, these data show that a short-term transition from APOE4 to APOE2 can broadly affect the cerebral transcriptome and lipidome, and that astrocyte-specific APOE replacement may be a viable strategy for future gene editing approaches to simultaneously reduce multiple AD-associated pathologies.
Similar content being viewed by others
Main
The strongest genetic risk factor for late-onset Alzheimer’s disease (AD) is apolipoprotein E (APOE). In humans, APOE has three common polymorphic alleles that encode three protein isoforms of apolipoprotein E (ApoE): E2, E3 and E4. Compared with E3, E4 confers up to a 13-fold increase in risk for AD in homozygous carriers1. Conversely, E2 is strongly neuroprotective, reducing AD risk by up to 40%, slowing cognitive decline and increasing longevity2,3,4. These striking differences in AD risk have been attributed to several possible pathogenic mechanisms, including E4-driven exacerbation of amyloid and tau pathology, synaptic dysfunction, network connectivity, cerebrovascular impairments, a heightened neuroinflammatory state and alterations in glial glucose and lipid metabolism5.
Given this strong risk profile and multitude of biological effects, targeting APOE4 and ApoE4 presents as a promising therapeutic approach to simultaneously recalibrate multiple AD-associated pathways6. Studies that have successfully targeted APOE using gene editing in induced pluripotent stem cells (iPSCs) provide strong proof of concept that in vitro editing of ε4 to ε3 or ε2 simultaneously ‘corrects’ several phenotypes, including glial transcriptomic profiles, lipid metabolism and other functional measures7,8. Moving in vivo, other studies have explored vector-mediated delivery to supplement the E4 brain with neuroprotective E2 protein, showing that exogenous E2 markedly decreases amyloid burden in mice9,10,11,12. Efficacy of a similar viral-based methodology was validated in nonhuman primates13 and is currently undergoing clinical trials14. Conversely, decreasing E4 protein levels is another promising strategy several groups have demonstrated via genetic, antibody-based or antisense oligonucleotide-based knockdown methods15,16,17, or indirect modulation of ApoE levels by targeting its primary cell surface receptors18.
Together, these studies suggest that an approach where E4 is eliminated and replaced with E2 could be a highly effective therapeutic strategy. Importantly, recent advances in CRISPR–Cas9 and related base-editing technologies make this type of precision medicine a conceivable approach for E4 carriers in the coming years19. To generate a preclinical model in which we could investigate the feasibility and therapeutic potential of a full APOE4 to APOE2 transition in vivo, we developed a knock-in model coined the APOE ‘switch’ mouse (APOE4s2). Using a ubiquitously expressed inducible Cre, we drove a highly efficient transition from E4 to E2 in both the periphery and brain, as confirmed at both the mRNA and protein level. The metabolic profile of post-switch APOE4s2 mice resembled that of humanized APOE2 ‘targeted replacement’ (TR) mice and E2/E2 humans, with elevated ApoE concentrations and increased plasma lipids. Importantly, replacement of E4 with E2 in adult mice resulted in transcriptomic changes across multiple cerebral cell types, particularly within astrocytes and in pathways related to the immune response and metabolism. Interestingly, an astrocyte-only APOE4 to APOE2 switch produced very similar AD-relevant transcriptomic changes as a full-body transition. Finally, when crossed with 5xFAD mice, astrocyte-specific APOE allele switching from E4 to E2 was sufficient to decrease total amyloid levels, reduce gliosis and plaque-associated ApoE, and improve cognition. Collectively, these data suggest that even a short-term, postdevelopmental transition from E4 to E2 broadly affects the cerebral transcriptome and lipidome, and can simultaneously decrease multiple AD pathologies. These findings highlight this inducible system as a promising preclinical model in which to assess the timing, cell-specific contributions and overall therapeutic potential of APOE gene editing.
APOE ‘switch’ mice efficiently transition from E4 to E2
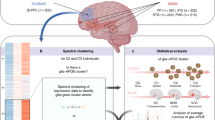
To investigate the in vivo effects of an APOE4 to APOE2 allelic switch, we generated APOE4s2loxP/loxP (APOE4s2) mice that express a floxed coding region (exon 4) of human APOE4 followed by exon 4 of human APOE2 (Fig. 1a). APOE4s2 mice were crossed to a tamoxifen (TAM)-activable strain ROSA26-CreERT1 Cre to generate floxed ROSA26-CreERT1(APOE4s2G), allowing for temporal induction of global (‘G’) APOE allele switching. To confirm that TAM successfully induces a whole-body transition from expression of E4 to E2, 2-month-old APOE4s2G mice were compared to TAM-treated Cre− APOE4s2 (APOE4s2) littermates and human APOE2-targeted and APOE4 TR mice20,21. As brain and liver are the two major sources of ApoE, efficiency of the E4 to E2 switch was assessed using quantitative PCR (qPCR) to determine APOE mRNA expression in these two tissues. Allelic discrimination plots demonstrated an efficient transition from APOE4 to APOE2 mRNA in both the liver and the brain (Fig. 1b,c). Targeted mass spectrometry (MS)-based proteomic analysis confirmed the E4 to E2 transition at the protein level, where 84–93% and 92–99% of the ApoE peptides detected in the brain and plasma respectively from APOE4s2G mice were unique to E2 (Fig. 1e,f, Extended Data Fig. 1). Together, these data demonstrate that APOE4s2G mice synthesize human ApoE4 at baseline, and that TAM induces a highly efficient transition to APOE2 and ApoE2 at the gene and protein level.
a, Schematic depicting the genetic construct for the APOE4 to APOE2 ‘switch’ mouse (APOE4s2) and the experimental design for the experiments performed in APOE4s2 mice crossed to a globally (‘G’) expressed ROSA26-CreERT1 recombinase (APOE4s2G). Homozygous floxed Cre− littermates (APOE4s2) were used as controls. b,c, Allelic discrimination plots depicting a shift from APOE4 to APOE2 mRNA expression (b) in APOE4s2G mice in both the brain (b) and liver (c) compared to Cre− APOE4s2 littermates and ApoE2 and ApoE4 TR controls injected with TAM. d, Methodology used to semiquantitatively detect the ratio of peptides unique to E2 or E4 in the brain and plasma using liquid chromatography (LC)–MS/MS-based proteomic analysis. e,f, Ratio of unique E2 versus E4 peptides detected using LC–MS/MS-based proteomic analysis showing an efficient transition to E2 in both the brain (f) and plasma (g) of APOE4s2G (n = 4), APOE4s2 (n = 4), E2-TR (n = 2) and E4-TR (n = 2) mice. Data are represented as mean values ± s.e.m. RFU, relative fluorescence unit.
Whole-body switch results in an ‘E2-like’ metabolic profile
Given ApoE’s central role in lipid metabolism, we first asked if the global E4 to E2 switch could affect peripheral and cerebral metabolic pathways. E2-TR mice display a plasma lipid profile similar to the type III hyperlipoproteinemia found in ~5–10% of E2/E2 humans, particularly when challenged with a high-fat diet (HFD)20. To determine if APOE4s2G mice phenocopy this hyperlipidemic profile after the E4 to E2 transition, mice were fed a Western diet for 4 months. Blood was collected 3 months into the diet before TAM treatment, and 1 month after the allelic switch.
Before TAM, total plasma triglyceride (TG) levels in APOE4s2 and APOE4s2G mice were indistinguishable from the E4-TR controls (Fig. 2a). However, 1 month after TAM, APOE4s2G mice fed a Western diet displayed a substantial increase in very-low-density lipoprotein (VLDL) TGs and plasma ApoE levels, which rose to E2-TR levels (Fig. 2b–e and Extended Data Fig. 2d). APOE4s2G mice fed a normal chow diet also showed elevations in TGs, cholesterol and phospholipids relative to the APOE4s2 controls, although the increases were more modest (Extended Data Fig. 2e–j). Brain ApoE levels were also elevated in APOE4s2G mice 1 month after switching (Fig. 2f). Colocalization of ApoE with cell-specific markers for astrocytes, microglia and neurons showed that like the E2-TR, E4-TR and APOE4s2 controls, ApoE primarily colocalized with astrocytes (glial fibrillary acidic protein (GFAP)+) in APOE4s2G mice, with measurable, but much lower, colocalization with the microglial marker IBA1 (Fig. 2g,h and Extended Data Fig. 2k–o). While neurons synthesize ApoE under certain conditions, we observed negligible colocalization of ApoE with NeuN (Extended Data Fig. 2k–o).
a, Plasma TGs before the TAM-induced switch (E2-TR, n = 8; E4-TR, n = 8; APOE4s2, n = 8; APOE4s2G, n = 8; one-way analysis of variance (ANOVA) with Tukey’s multiple comparisons test). b, Plasma TGs after TAM (E2-TR, n = 8; E4-TR, n = 8; APOE4s2, n = 7; APOE4s2G n = 7; one-way ANOVA with Tukey’s multiple comparisons test). c, TGs in each lipoprotein fraction after switching (E2-TR, n = 3 pooled; E4-TR, n = 3 pooled; APOE4s2, n = 3 pooled; APOE4s2G, n = 3 pooled). d,e, ELISA quantification of ApoE in plasma before TAM (d) and after the TAM-induced switch (e) (E2-TR, n = 5; E4-TR, n = 5; APOE4s2, n = 6; APOE4s2G, n = 7; one-way ANOVA with Tukey’s multiple comparisons test). f, ELISA quantification of ApoE in whole-brain tissue after TAM (E2-TR, n = 3; E4-TR, n = 3; APOE4s2, n = 5; APOE4s2G, n = 5; one-way ANOVA with Tukey’s multiple comparisons test). g,h, Representative images showing GFAP (green) (g) or IBA1 (green) (h) and ApoE (red) in APOE4s2 controls (top) or APOE4s2G brains (bottom). i, Volcano plot comparing brain lipids altered in APOE4s2G mice versus Cre− APOE4s2 controls (APOE4s2, n = 8; APOE4s2, n = 8; all female). The cutoff value for significance is log10(P) > 1.3. j, Heatmap depicting the top ten most significantly changed lipids between APOE4s2G and APOE4s2 controls. k,l, WGCNA analysis identified lipid modules associated with the switch from E4 to E2. k, Module-trait correlations between modules and APOE expression indicated by the numbers in the table, with P values shown in parentheses. l, Network plot of lipids identified in the blue module. a–f, For detailed statistical information, see Supplementary Table 3. Data represent mean ± s.e.m. of biological replicates. Individual data points have been plotted; in the box plots, the center line indicates the median; the box limits indicate the upper and lower quartiles; and the whiskers indicate the minimum and maximum. ***P < 0.001, ****P < 0.0001. i–l, Twenty-two classes of lipids were detected: CER; cardiolipin (CI); coenzyme (CO); diglyceride (DG); dimethyl phosphotidylehtanolamine (DMEPE); fatty acid (FA); ganglioside (GM); lysodimethylphosphatidylethanolamine (LDMEPE); LPC; lysophosphatidylethanolamine (LPE); lysophosphatidylglycerol (LPG); lysophosphatidylinositol (LPI); lysophosphatidylserine (LPS); (O-acyl)-ω-hydroxy FA (OAHFA); phosphatidic acid (PA); PC; PE; phosphatidylglycerol (PG); phosphatidylinositol (PI); phosphatidylserine (PS); sphingomyelin (SM); and TG. HDL, high-density lipoprotein. LDL, low-density lipoprotein.
We next examined possible changes to the cerebral lipidome using an untargeted LC–MS/MS approach. A comparison of APOE4s2G mice versus Cre− APOE4s2 controls identified 14 lipid species that were significantly different in abundance between the two groups (Fig. 2i,j). In agreement with previous postmortem AD studies22,23, many of these lipids were glycerophospholipids, including many phosphatidylcholine (PC) and ceramide (CER) species. Interestingly, there were nuanced changes within these lipid classes; in both PC and CER classes, some lipid species increased in abundance while others decreased. For reference, a comparison of E2-TR and E4-TR control mice identified 57 lipids of differing abundance, again most of which were glycerophospholipids (Extended Data Fig. 2o,p). We then performed a weighted gene coexpression network analysis (WGCNA) on the cerebral lipidome, where we identified five coexpression modules, including one network that was significantly correlated with the E4 to E2 switch (blue module) and consisted primarily of PC and phosphatidylethanolamine (PE) species (Fig. 2k,l). Together, these data demonstrate that even a short-term transition from E4 to E2 in the adult mouse is sufficient to drive alterations in ApoE concentrations and lipid metabolism, most notably increases in multiple PC species within the brain.
Full-body switching drives AD-relevant transcriptomic changes
To unbiasedly examine how switching from APOE4 to APOE2 globally influences cell-type-specific changes to the brain transcriptome, we performed single-cell RNA sequencing (scRNA-seq) on whole-brain tissue from 6-month-old APOE4s2G mice, Cre− APOE4s2 controls and age-matched and sex-matched E2-TR and E4-TR mice. Dimensionality reduction identified 41 clusters, which were assigned to one of 12 unique cell types, with cell and cluster proportions similarly distributed across all groups (Fig. 3a and Extended Data Fig. 3). Analysis of differentially expressed genes (DEGs) revealed astrocytes, oligodendrocytes, microglia and endothelial cells as the cell types most affected by a 1-month-long APOE4 to APOE2 switch (Fig. 3b, Extended Data Fig. 4 and Supplementary Table 1). Across these cell types, there was notable overlap of DEGs from APOE4s2G mice versus APOE4s2 controls with DEGs from E2-TR versus E4-TR mice (Fig. 3c), suggesting that many genes that distinguish ‘lifetime’ expression of E4 are acutely malleable. Gene Ontology (GO) analysis of switch-related DEGs highlighted a number of pathways related to metabolism, including substrate transport, redox management and cellular respiration (Fig. 3d–g and Supplementary Table 2). Furthermore, many of the genes altered by the E4 to E2 switch fell under one of six primary AD ‘umbrella’ terms—pathways implicated across multiple postmortem AD studies24—in particular those related to cerebral metabolism, inflammation and cytoskeleton/vascular integrity (Fig. 3h). Finally, the DEGs associated with the E4 to E2 switch also strongly reflected several APOE-associated and AD-associated gene lists, particularly the late-onset AD (LOAD) genes identified in and merged from publicly available human transcriptomic datasets (Fig. 3i)25,26,27,28. Collectively, these results demonstrate that, even in the absence of AD pathology, a postdevelopmental transition from expression of E4 to E2 is sufficient to drive transcriptomic changes in pathways previously implicated in AD.
a, Uniform manifold approximation and projection (UMAP) of all 85,701 cells analyzed, classified based on canonical gene expression markers into 12 cell types (APOE4s2G, n = 3; APOE4s2, n = 3; E2-TR, n = 3, E4-TR, n = 2; all females). b, Number of DEGs between APOE4s2G and Cre− APOE4s2 controls according to cell type (Padj = <0.001). c, Venn diagrams revealing commonality of DEGs in the APOE4s2G versus APOE4s2 and E2-TR versus E4-TR datasets. d–g, Upregulated and downregulated GO terms associated with DEGs from astrocytes (d), oligodendrocytes (e), microglia (f) and endothelial cells (g). The cutoff value for significance was set at a false discovery rate (FDR)-adjusted P value of 0.05. h, Pie charts visualizing the categorical distribution of APOE switch-related GO terms into AD-related pathways. Upregulated (left) AD pathways account for 24%, while downregulated (right) AD pathways account for 21% of total downregulated GO terms. i, Overlap of the APOE switch transcriptomics dataset with five relevant gene lists from AD-related transcriptomic studies. DAA, disease-associated astrocyte; OPC, oligodendrocyte precursor cell; Transcription corep. binding, transcription corepressor binding; VLC, vascular leukocyte.
Astrocyte-only switch to E2 drives similar gene expression
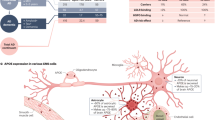
Considering the potential translational concerns associated with germline, whole-body APOE editing, we reasoned that a central nervous system (CNS)-selective APOE4 to APOE2 switch could afford the neuroprotection of E2 while minimizing off-target effects. Because astrocytes synthesize most of CNS ApoE and were the cell type most affected by the global E4 to E2 transition, we generated APOE4s2loxP/loxP/Aldh1l1-CreERT2 APOE switch mice (APOE4s2A) that would express APOE4 in all CNS cell types except for astrocytes, which would selectively transition to APOE2 upon TAM injection29,30 (Fig. 4a). Cell-type specificity was confirmed via colocalization of GFAP with tdTomato expression in APOE4s2A mice crossed to an Ai9 Cre reporter strain (Fig. 4b and Extended Data Fig. 5a). Leveraging the Ai9 cross, we then used cell sorting and qPCR to validate the high efficiency and cell specificity of the astrocyte-specific E4 to E2 switch (Fig. 4c). A similar, albeit more modest, shift toward APOE2 expression was observed in whole-brain tissue, suggesting that most of total brain ApoE is derived from astrocytes, which were now synthesizing E2 (Fig. 4d). While the Aldh1l1-CreERT2 showed high specificity for astrocytes in the brain; plasma lipid profiling and peripheral ApoE measures indicated that there is also notable E2 expression in the periphery (Extended Data Fig. 5d,e).
a, Graphic depicting astrocyte-specific APOE4s2A mice with Aldh1l1-CreERT2. b, Representative images showing colocalization of tdTomato Ai9 Cre reporter (red) and GFAP (green). c,d, Allelic discrimination plots measuring APOE4 versus APOE2 mRNA expression in astrocytes (c) sorted from other cell types in the brain, or from whole-brain homogenates (d), of APOE4s2A mice versus APOE4s2 controls (E2-TR, n = 2; E4-TR, n = 2; APOE4s2A, n = 6; APOE4s2, n = 6). e, Number of DEGs between APOE4s2A and Cre− APOE4s2 controls according to cell type (Padj ≤ 0.001) (APOE4s2A, n = 3; APOE4s2, n = 3). f, Overlap of the APOE switch transcriptomic dataset with five other gene lists from AD-related transcriptomic studies. g, Venn diagrams revealing the commonality of DEGs in the APOE4s2A versus APOE4s2 and APOE4s2G versus APOE4s2 datasets. h–k, GO analysis for downregulated and upregulated terms associated with DEGs from microglia (h), oligodendrocytes (i), astrocytes (j) and endothelial cells (k). The cutoff value for significance was set at an FDR-adjusted P < 0.05 and fold change ≥ 1.3.
We next asked if an astrocyte-specific switch could induce similar transcriptomic changes as the full-body APOE allele switch. Analysis of DEGs showed that the astrocyte-specific APOE4 to APOE2 transition primarily affected similar cell types as observed in the global model (Fig. 4e and Extended Data Fig. 5). Additionally, as seen in the whole-body model, the DEGs associated with an astrocyte-specific E4 to E2 switch also strongly mirrored the LOAD gene lists (Fig. 4f). Interestingly, several cell types that continued expressing E4 in APOE4s2A mice (that is, microglia, oligodendrocytes) still showed a high number of DEGs, suggesting non-cell-autonomous effects of astrocyte-derived E2. There was also substantial overlap (39–60% depending on the cell type) between the specific DEGs associated with a global versus astrocyte-specific E4 to E2 switch (Fig. 4g). Across all cell types, pathway analyses of astrocyte switch DEGs revealed similar terms to those seen in whole-body APOE4s2G mice, particularly in pathways related to neurotransmission, redox status and metabolite transport (Fig. 4h–k). Together, these data suggest that astrocyte-exclusive expression of E2 is sufficient to drive AD-relevant alterations in the glial transcriptome, even when all other cell types continue to express E4 and even in a model that lacks overt AD pathology.
Astrocyte-only E4 to E2 switch decreases amyloid plaque load
We then tested if an astrocyte-specific E4 to E2 switch could directly affect AD pathology. To this end, we crossed APOE4s2A mice to the 5xFAD model of AD (4s2A/FAD) and administered TAM at 6 months of age. At 8 months of age, associative fear conditioning and the Morris water maze were used to assess contextual, cued and spatial memory; brains were subsequently collected for histopathological analyses (Fig. 5a). While the Morris water maze did not show striking differences in spatial learning or memory, 4s2A/FAD mice showed improvements in associative learning and more robust contextual and cued memory, an effect that was strongest in females (Fig. 5b–d and Extended Data Fig. 6a–d).
a, Schematic depicting the APOE4s2A × 5xFAD cross to generate the 4s2A/FAD mice. b–d, Percentage time freezing measured during fear conditioning (learning) (b), and the contextual (memory) (c) and cued memory test (d) tests, in 4s2A/FAD mice compared to Cre− controls 2 months after TAM (4s2A/FAD, n = 16, 4s2/FAD, n = 16; **P < 0.01,***P < 0.001, ****P < 0.0001, using a repeated measures ANOVA). e, Representative images of Amylo-Glo (blue) and nucleus (red) staining in 4s2A/FAD and 4s2/FAD mice. f, Quantification of the total percentage amyloid+ area in 4s2A/FAD mice showed significant decreases in total amyloid measured according to the percentage of the amyloid+ area. The dashed line represents the mean amyloid percentage area in 4s2/FAD mice at the 6-month pre-switch time point (y = 0.118). **P < 0.01, using a two-tailed unpaired t-test. g,h, Heatmap (g) and quantification (h) of the percentage of amyloid+ area according to region in 4s2A/FAD versus 4s2/FAD mice (4s2A/FAD, n = 10; 4s2/FAD, n = 10; two-way ANOVA with multiple comparisons). i–l, ELISA quantification of Aβ40 (i,j) or Aβ42 (k,l) in soluble (PBS) or insoluble (5 M GuHCl) whole-brain fractions from 4s2A/FAD mice compared to 4s2/FAD controls (4s2A/FAD, n = 14; 4s2/FAD, n = 14; *P < 0.05, **P < 0.01, ****P < 0.0001, using a two-tailed unpaired t-test). m, Representative images of mOC31 anti-CAA (green) and CD31 (red) staining in the cortex of 8-month-old 4s2A/FAD and 4s2/FAD mice. n, Quantification of CAA+ vessels normalized to total CD31+ area in 8-month-old 4s2A/FAD mice and 4s2/FAD controls after the switch compared with 6-month-old mice before the switch (4s2A/FAD 8 months, n = 8; 4s2/FAD 8 months, n = 8; 4s2/FAD 6 months, n = 8; *P < 0.05, **P < 0.01, using a two-way ANOVA with multiple comparisons). a–n, For detailed statistical information, see Supplementary Table 3. Data represent mean ± s.e.m. of biological replicates. Individual data points have been plotted. Sex is denoted by the shape of the symbol: females (circle), males (triangle). In the box plots, the center line indicates the median; the box limits indicate the upper and lower quartiles; and the whiskers indicate the minimum and maximum.
Previous work established that human APOE affects amyloid plaque deposition in the 5xFAD brain in a genotype-dependent fashion (E4 > E2)31. To test if a late-stage, astrocyte-only replacement of APOE4 with APOE2 could decrease plaque load, we performed immunohistochemical analyses of amyloid accumulation. Strikingly, after a 2-month astrocyte-specific allelic switch, 4s2A/FAD animals showed significant reductions in total amyloid levels (Fig. 5e,f and Extended Data Fig. 6e–h). When compared to ‘baseline’ amyloid measures at 6 months of age, the reduced burden in 4s2A/FAD mice was due to slowed or delayed amyloid accumulation (Fig. 5f and Extended Data Fig. 7). Anatomical analyses revealed decreases in many regions, with significant reductions in the hippocampus, olfactory area and thalamus primarily driving the decrease in plaque load (Fig. 5g,h). Enzyme-linked immunosorbent assay (ELISA) quantification of Aβ42 and Aβ40 in whole-brain soluble and insoluble fractions revealed reductions in both soluble and insoluble Aβ42 and Aβ40 observed in 4s2A/FAD brains (Fig. 5i–l and Extended Data Fig. 7).
Overall, there were very few sex differences in AD-related pathology. While higher amyloid plaque loads were observed in female compared to male 4s2/FAD Cre− controls, we did not detect any other amyloid-dependent sex differences, nor was this difference reflected in the soluble and insoluble Aβ42 and Aβ40 measures (Extended Data Fig. 7e–h). We did note subtle sex differences in the expression of the postsynaptic density marker PSD95 in the cortex, although this was limited to plaque-distal regions and was not seen in the hippocampus (Extended Data Fig. 7i–m).
Because APOE4 is also a risk factor for cerebral amyloid angiopathy (CAA)31, we measured CAA before and after the inducible, astrocyte-specific transition to APOE2. Interestingly, we observed no differences in CAA between 4s2A/FAD and 4s2/FAD controls at 8 months of age (Fig. 5m,n). Finally, we measured the expression of the tight junction protein ZO1 and found no change after the switch to astrocyte-derived E2 (Extended Data Fig. 7n–r). Together, these results suggest that a relatively late-stage switch to astrocytic expression of E2 is sufficient to substantially lower parenchymal plaque burden, but not CAA, and improve some, but not all, of the cognitive deficits associated with E4.
Astrocyte E4 to E2 replacement lowers plaque-associated gliosis
In addition to the reduction in parenchymal amyloid plaque load, we also observed significantly lower levels of GFAP in 4s2A/FAD mice compared to controls (Fig. 6a,b). Interestingly, although only astrocytes transitioned to the expression of E2 in the 4s2A/FAD mice, we noted a dramatic reduction in microglial IBA1 positivity (Fig. 6c), again suggestive of non-cell-autonomous effects of ApoE2. Like the regional reductions in plaque load, GFAP and IBA1 levels were most substantially decreased in the hippocampus, thalamus and cortical areas (Fig. 6d,e and Extended Data Fig. 8a–c).
a, Representative images of Amylo-Glo (blue), IBA1 (green), GFAP (orange) and nucleus (red) staining in 4s2A/FAD mice and 4s2/FAD controls. b,c, Quantification of the percentage GFAP+ (b) or IBA1+ (c) area in 4s2A/FAD mice versus 4s2FAD controls (4s2A/FAD, n = 16; 4s2/FAD, n = 16; ***P < 0.001, ****P < 0.000, using a two-tailed unpaired t-test). d,e, Quantification of GFAP+ (d) or IBA1+ (e) percentage area according to region in 4s2A/FAD and 4s2/FAD mice (4s2A/FAD n = 10; 4s2/FAD n = 10; *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001, using a two-way ANOVA with multiple comparisons). f, Representative images of Amylo-Glo (blue), IBA1 (green), GFAP (orange) and nucleus (red) proximity and colocalization. g–i, Percentage area of colocalization of Amylo-Glo and GFAP (g), Amylo-Glo and IBA1 (h) or Amylo-Glo with GFAP and IBA1 (i) in 4s2A/FAD and 4s2/FAD mice (4s2A/FAD, n = 16, 4s2/FAD, n = 16; **P < 0.01, ***P < 0.001, using a two-tailed unpaired t-test). j–m, Histograms (j,l) and area under the curve (AUC) quantification of histograms (k,m) from plaque proximity analyses measuring the percentage area of GFA+ (j,k) and IBA1+ (l,m) cells from the plaque center in 4s2A/FAD mice compared to 4s2/FAD controls (4s2A/FAD, n = 8; 4s2/FAD, n = 8). a–m, For detailed statistical information, see Supplementary Table 3. Data represent mean ± s.e.m. of biological replicates. Individual data points have been plotted. Sex is denoted by the shape of the symbol: females (circle), males (triangle). In the box plots, the center line indicates the median; the box limits indicate the upper and lower quartiles; and the whiskers indicate the minimum and maximum. NS, not significant.
Reductions in total GFAP and IBA1 levels were especially pronounced in close proximity to amyloid plaques. 4s2A/FAD mice showed decreased colocalization of amyloid with GFAP+ and IBA1+ puncta (Fig. 6f–i and Extended Data Fig. 8d–g); plaque proximity analyses showed more pronounced decreases in GFAP and IBA1 in 4s2A/FAD mice when close to the plaque center, with distribution of the GFAP+ and IBA1+ signal becoming more similar to controls as the distance from the center increased (Fig. 6j–m and Extended Data Fig. 8h–k). Together, these findings highlight a strong decrease in plaque-associated astro-gliosis and micro-gliosis after an astrocytic E4 to E2 switch.
E4 to E2 switch reduces reactive microglia and plaque ApoE
We next investigated more nuanced changes to microglia by measuring major histocompatibility complex class II (MHC-II), a marker of activated-response microglia (ARM), an Apoe-dependent subpopulation32. 4s2A/FAD mice showed a decrease in total MHC-II+ area and a decrease in colocalization between MHC-II + IBA1+ cells and amyloid plaques (Fig. 7a–d and Extended Data Fig. 9a–f), indicating a reduction in ARMs after the 2-month astrocyte-specific switch to E2. Notably, sex-based analysis showed significantly more cortical plaques with MHC-II+ microglia in female mice than males (Extended Data Fig. 9c). Although there was no change in homeostatic gene expression, decreased expression of several disease-associated microglia (DAM) genes, including Trem2, Clec7a and APOE, was also noted after the astrocyte-specific transition to E2 (Fig. 7e,f and Extended Data Fig. 9g,h).
a, Representative images of Amylo-Glo (blue), IBA1 (green), MHC-II (orange) and nuclei (red) staining. b–d, Quantification of total MHC-II+ percentage area (b), colocalized MHC-II+ and IBA1+ area (c), and colocalization of MHC-II+, IBA1+ and Amylo-Glo+ area (d) in 4s2A/FAD and 4s2/FAD mice in the cortex and hippocampus (4s2A/FAD, n = 16; 4s2/FAD, n = 16; *P < 0.0, **P < 0.01, ***P < 0.001, ****P < 0.0001 using a two-way ANOVA with multiple comparisons). e,f, Fold change of Tmem119, P2ry12 and Cx3cr1 (e) or Clec7a, Trem2 and APOE (f) gene expression normalized to 18S in the whole brain from 4s2A/FAD mice compared to 4s2/FAD controls (4s2A/FAD, n = 14; 4s2/FAD, n = 14; *P < 0.0, **P < 0.01 using a two-tailed unpaired t-test). g, Representative images of Amylo-Glo (blue), ApoE (green) and GFAP (orange) in 4s2A/FAD mice and 4s2/FAD controls. h, Quantification of plaque-associated ApoE in 4s2A/FAD mice relative to 4s2/FAD controls in both the cortex and hippocampus (4s2A/FAD, n = 16, 4s2/FAD, n = 16; *P < 0.05, **P < 0.01, using a two-way ANOVA with multiple comparisons). i,j, ELISA quantification of ApoE within the soluble (PBS) (i) and insoluble (5 M GuHCl) (j) brain fractions of 4s2/FAD and 4s2A/FAD whole-brain tissue (4s2A/FAD, n = 14; 4s2/FAD, n = 14; *P < 0.05, using a two-tailed unpaired t-test). k, Correlation of cognitive measures and neuropathological analyses. The red squares indicate a positive correlation; the blue squares indicate a negative correlation. a–k, For detailed statistical information, see Supplementary Table 3. Data represent mean ± s.e.m. of biological replicates. Individual data points have been plotted. Sex is denoted by the shape of the symbol: females (circle), males (triangle). In the box plots, the center line indicates the median; the box limits indicate the upper and lower quartiles; and the whiskers indicate the minimum and maximum.
While there was no difference in total ApoE protein levels between the 4s2A/FAD and 4s2/FAD mice, we observed a striking difference in plaque-associated ApoE. Control mice expressing E4 in all cell types displayed many bright halos of ApoE staining surrounding the cores of plaques (Fig. 7g,h and Extended Data Fig. 8i–m). However, these intense bursts of plaque-associated ApoE signal were largely absent in 4s2A/FAD mice, highlighting potential differential effects of the E2 versus E4 isoforms on plaque expansion or compaction. ELISA quantification of ApoE supported these findings, with significant reductions of ApoE within the insoluble (plaque-associated) brain fractions from 4s2A/FAD mice, but no differences in soluble ApoE (Fig. 7i,j and Extended Data Fig. 9n,o).
Finally, we linked behavioral and pathological measures from each individual mouse to identify potential significant correlations between cognition and AD-associated pathologies. Several measures were significantly correlated but, paradoxically, positively correlated in 4s2/FAD mice yet negatively correlated in 4s2A/FAD mice (Fig. 7k). Taken together, these results suggest that astrocyte-selective expression of ApoE2 leads to downstream reductions in microglial activation and lower levels of plaque-associated ApoE, and that these various measures of gliosis differentially correlate with memory.
Discussion
In the current study, we describe a new APOE ‘switch’ model (APOE4s2) that allows for inducible, cell-specific, in vivo replacement of the high-risk APOE4 allele with the protective APOE2 variant. We characterize how a global E4 to E2 replacement drives peripheral metabolic changes and AD-relevant alterations in the cerebral transcriptome and lipidome. We further show that an astrocyte-selective replacement of E4 with E2 significantly alters the transcriptomic profiles of multiple CNS cell types, reduces amyloid plaque load, dampens gliosis and improves cognitive outcomes in an amyloidosis model of AD. These findings underscore the dynamic and reversible nature of APOE’s contribution to AD pathology and suggest that targeted APOE allele replacement, particularly in astrocytes, may offer a promising therapeutic avenue to modify multiple AD-relevant pathways simultaneously.
APOE4s2G mice, in which all cell types transition from APOE4 to APOE2, reliably phenocopy the humanized E4-TR mice before the switch and E2-TR mice within 1 month after the switch. Specifically, untargeted lipidomic profiling of the brain revealed that the allelic switch altered multiple PC, CER and lyso-PC (LPC) species, a class of pro-inflammatory lipids that act as ‘find me’ signals for myeloid cells. These results align with our previous work showing decreased PC levels in the aged E4 and E4 5xFAD mouse brain, albeit relative to E3 (ref. 33). They also closely parallel the lipidomic profiling of postmortem AD brains, where several PC species show E4-associated decreases relative to E2 (ref. 22). Interestingly, the most significantly upregulated lipid in our dataset is a CER (Cer(d40:2)) involved in glucosylceramide synthesis, a process downregulated in AD brains34. Alterations to lipid species such as CERs or PCs could have important implications for intracellular and cell–cell communication, membrane composition and apoptosis35.
Across multiple studies, glial cells show APOE4-associated transcriptomic changes in both human E4+ brains and E4-TR mice5. Interestingly, these are the same primary cell types affected by a short-term E4 to E2 transition. Our results showing changes in microglia, oligodendrocyte and endothelial cell transcriptomes after the astrocyte-selective switch also suggest strong non-cell-autonomous effects of astrocyte-derived E2. These findings were mirrored by histopathological reductions in microglial reactivity in 4s2A/FAD mice and are in line with previous studies showing profound downstream effects on microglia after an astrocyte-specific knockout of APOE4 (refs. 5,15,16,17,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36). Direct comparison of the DEGs distinguishing E2-TR from E4-TR mice to the DEGs of APOE4s2G versus APOE4s2 mice showed some commonality (~20%) across multiple cell types, suggesting that E4’s transcriptional signature is not permanently established during development or early life. Rather, many of these DEGs are acutely malleable, at least within the specific parameters tested in this study.
Further analyses of these DEGs revealed that the pathways most affected by the E4 to E2 switch—both whole-body and astrocyte-specific—were pathways centered on cerebral metabolism, inflammation and cytoskeleton/vascular integrity, which are strongly reminiscent of the lipid metabolism, immune and matrisome findings from in vitro comparisons of isogenic APOE glia8,37. Furthermore, with the caveat that comparisons to late-disease postmortem tissue is inherently limiting, several pathways altered by a short-term E4 to E2 transition were cellular pathways that are also perturbed across 17 large-scale human AD snRNA-seq studies24. Importantly, this overlap with AD scRNA-seq datasets occurred in both the global and astrocyte-specific switch models that lack overt AD pathology. Excitingly, this may suggest that the overarching cellular pathways that define AD in the human brain could be altered by transitioning from E4 to E2.
While fully ‘correcting’ the E4 risk allele to the neuroprotective E2 allele could, in principle, simultaneously improve cerebral function across multiple fronts and greatly decrease AD risk, this approach is not without its own risks. An estimated 5–10% of E2/E2 individuals present with type III hyperlipoproteinemia (HLP-III), and E2 carriage has been linked to a higher risk of melanoma38, age-related macular degeneration39 and even some neurological disorders such as posttraumatic stress disorder40 and progressive supranuclear palsy41. In fact, peripheral phenotyping of APOE4s2G mice showed that the in vivo E4 to E2 switch shifted animals to an HLP-III-like plasma lipid profile. Because Sullivan et al.20 reported HLP-III-like plasma lipids in E2-TR mice, the increases in plasma TGs and ApoE observed in APOE4s2G mice ‘after the switch’ were exciting confirmatory results regarding the effectiveness of our model design. On the other hand, the potential of inducing HLP-III or increasing the risk of other disorders after either a germline or universal (that is, ‘whole-body’) allelic switch to E2 raises important translational concerns.
To this end, we reasoned that CNS APOE targeting would hold the greatest therapeutic promise for AD. Strikingly, we show that a 2-month-long astrocyte-specific E4 to E2 switch after pathology onset is sufficient to decrease plaque load, reduce markers of glial reactivity and improve associative, but not spatial, memory in the 5xFAD model. It is possible that reductions in microglial reactivity are merely a consequence of decreased amyloid burden. However, the astrocyte-driven, APOE-dependent alterations to the microglial transcriptome found in our scRNA-seq data, coupled with glia–plaque colocalization data showing less IBA1+ and ARM microglia around plaques, suggest an alternative scenario where astrocyte-derived E2 could be having non-cell-autonomous effects on microglial activation and response to plaque (possibly via isoform-specific differences in ApoE–low-density lipoprotein receptor family interactions)42,43. While these findings may appear in contrast to studies indicating a cell-autonomous pathogenic role of ApoE in microglia, macrophages and neurons, these phenomena are not necessarily mutually exclusive15,36,44,45.
Similarly, findings from a study by Mahan et al.36 suggested a role for astrocyte-derived ApoE in amyloid reduction and associated gliosis. While that study did not report decreases in IBA1 as those seen in this study, they observed a similar decrease in reactive microglia using Clec7a, and a reduction in plaque-associated ApoE. Using an inverse approach, Liu et al.46 showed that astrocyte-specific expression of E4 suppressed amyloid clearance, with particularly strong effects during the amyloid seeding phase. In this study, we observed that expression of astrocytic E2 slowed amyloid deposition during the 2-month period after the allelic switch in 4s2A/FAD mice. To fully tease out the exact mechanisms at play, more nuanced experiments and timing paradigms will be required to examine how cell-specific ApoE contributes to amyloid deposition and the glial response to plaque.
Despite reductions in parenchymal plaque load, we observed similar levels of CAA in the 4s2A/FAD and 4s2/FAD mice. These findings may reflect human AD studies in which both E4 and E2 carriage were associated with higher CAA compared to E3/E3 individuals47,48, and could have translational implications for amyloid-related imaging abnormalities and other APOE-associated vascular disorders. Additionally, microglia cluster around Aβ deposits in CAA49. In this study, we show that short-term expression of astrocyte-derived E2 reduces proximity and colocalization of reactive microglia to plaques. Perhaps, the lack of observed effects on CAA after the astrocytic-E2 switch can be attributed to a dampened microglial response. While the specific role of microglia in CAA still requires more investigation, decreased microglial reactivity could also reduce or impair recruitment of other immune cells, such as perivascular or border-associated macrophages50.
Lastly, we show that a late disease stage ‘switch’ to expression of astrocytic E2 reduces plaque-associated ApoE. ApoE is a major component of plaques51. The literature suggests that there are isoform-dependent differences in ApoE–plaque associations but that several factors, such as ApoE lipidation state, compartment or cell type of origin, and Aβ conformational state, also have important roles in ApoE–Aβ interactions52,53,54,55,56. Our findings, taken together with the literature, suggest that astrocyte-derived E4’s ability to bind Aβ increases Aβ accumulation and ApoE protein aggregation within or surrounding the plaque. These results are consistent with studies implicating ApoE in plaque compaction and reflect a similar loss of function as seen in Apoe knockout mice, which also feature reductions in plaque-associated microgliosis57. Also in line with these results is a recent study, which showed that plaque-associated ApoE attracts microglia to the plaque and stimulates their activation via a vascular cell adhesion molecule 1-based mechanism58.
In summary, the current study leverages targeted APOE allele replacement as a promising therapeutic intervention to mitigate AD risk and describes a model in which we can recapitulate potentially beneficial or deleterious effects of APOE replacement. However, several limitations should be addressed. First, the degree of risk and protection afforded by E4 and E2 respectively vary according to ancestry1,59,60,61,62. Thus, some populations may not appreciably benefit from E4-E2 replacement strategies; future APOE-targeted therapies should take ancestry into account when conducting risk–benefit analyses. Second, the 5xFAD model used in this study provided a model with well-characterized APOE-dependent changes in amyloid pathology55,63,64. However, future studies should consider the roles of other facets of the disease, including tauopathies. Finally, Aldh1l1-CreERT2 is highly selective for astrocytes in the CNS, but Aldh1l1 is also expressed in the liver65, as evidenced by the plasma lipid profile of APOE4s2A mice. Our study suggests that astrocyte-exclusive ‘switching’ from E4 to E2 in the CNS can mitigate AD pathologies; however, future studies should specifically dissect out the role of the peripheral ApoE on the CNS. Ultimately, we hope that this and other inducible APOE models will provide clean experimental systems to test the efficacy of several APOE gene editing strategies. In combination with select Cre strains and other molecular biology approaches, next-generation ‘switch’ models can provide cell-specific and temporally controlled editing among APOE alleles. Using this framework, future studies can help determine optimal cell-specific contributions of ApoE to AD pathogenesis, test for potential off-target effects of APOE manipulation and establish optimal therapeutic windows for APOE-directed clinical strategies.
Methods
Animals
All animal procedures and protocols were approved by the Institutional Animal Care and Use Committee and the University of Kentucky. APOE switch mice (APOE4s2) were generated on a C57BL/6N background at TransViragen. Mouse Apoe exons 2–4 were replaced by human APOE exons 2–4. The coding region of APOE (exon 4) was followed by a stop cassette (3× SV40 poly(A)) and flanked with loxP. After the coding region for APOE4, an additional exon 4 contained the sequence for APOE2. ROSA26-cre/ERT1 (strain no. 004847, research resource identifier (RRID): IMSR_JAX:004847) and Aldh1l1-Cre/ERT2 mice (strain no. 029655, RRID: IMSR_JAX: 029655) were purchased from The Jackson Laboratories and crossed with homozygous APOE4s2loxP/loxP mice to generate ROSA26Cre/+/APOE4s2loxP/loxP (APOE4s2G) or Aldh1l1Cre/+/APOE4s2loxP/loxP (APOE4s2A) mice. APOE4s2A mice were further crossed to 5xFAD mice to generate Aldh1l1Cre/+/APOE4s2loxP/loxP hemizygous for the familial AD mutation (4s2A/FAD). Human APOE TR mice homozygous for APOE2 or APOE4 and Cre− littermates were used as controls37.
Genotyping
Animals were genotyped at 21 days for the APOE switch (APOE4s2) transgene using PCR. Animals were bred to homozygosity for the APOE4s2 allele. Primers specific for the humanized target allele were used to identify whether animals were homozygous for the APOE4s2 allele. Primers for the native and target alleles are as follows: WT-1 forward (5’-AGGGGTTACCTCCAGGAAAGG-3’); WT-1 reverse (5’-TGGATCCGCTGCCAAAA-3’); WT-2 forward (5’-CAAAGGCTGGAATTACAGG-3’); WT-2 reverse (5’-TTGGGCTCCATGAGCTCTGG-3’); Tg-1 forward (5’-AGGGGTTACCTCCAGGAAGG-3’); Tg-1 reverse (5’-GCCGTGTTCCATTTATGAGC-3’); Tg-2 forward (5’-TTCTCCCCGGCCTGCTTGTT-3’); and Tg-2 reverse (5’-TTGGGCTCCATGAGCTCTGG-3’).
TAM administration
TAM (cat. no. T5648, Sigma-Aldrich) was dissolved overnight at 37 C° with gentle rocking in sterile sunflower seed oil (cat. no. S5007, Sigma-Aldrich) at a concentration of 20 mg ml−1 and protected from light. TAM was administered via intraperitoneal (i.p.) injection once every 24 h for five consecutive days. The injection dose of 75 mg kg−1 was determined according to animal body weight.
Gene expression analysis
Brain and liver were collected from APOE4s2G, APOE4s2A, APOE4s2, E2-TR and E4-TR mice switched between 4–6 months of age 72 h, 1 month, 3 months, 6 months or 12 months after TAM administration. RNA was extracted from whole-brain and liver tissue using the RNeasy Mini Kit (cat. no. 74104, QIAGEN) and converted to complementary DNA (cDNA) (cat. no. 4388950, Thermo Fisher Scientific). The APOE genotype was determined using PCR with TaqMan assay primers for the two allele-determining single-nucleotide polymorphisms of APOE: rs7412 and rs429358 (cat. no. 4351379, Thermo Fisher Scientific). Positive human DNA controls for the APOE genotypes E2/E2 and E4/E4 were included with each assay.
To quantify expression of homeostatic and DAM genes in 4s2/FAD and 4s2A/FAD mice, brains were collected 2 months after TAM. RNA was extracted and converted to cDNA as described above. TaqMan Gene Expression Assays (cat. no. 4331182, Thermo Fisher Scientific) were used to measure Cx3cr1 (Mm02620111_s1), Tmem119 (Mm00525305_m1), P2ry12 (Mm01950543_s1), Trem2 (Mm04209422_m1), Clec7a (Mm01183349_m1) and APOE (Hs00171168_m1). Data were normalized to endogenous 18S control (Hs99999901_s1) and the fold change was calculated between groups using \({2}^{-\varDelta \varDelta \,{{\rm{C}}}_{{\rm{t}}}}\) values.
Brain tissue sample processing
Brains were collected from mice who were transcardially perfused with sterile 0.9% saline (cat. no. S9625, Sigma-Aldrich) unless stated otherwise. The brain was dissected into two hemispheres. The left hemisphere was flash-frozen in liquid nitrogen for biochemical analyses while the right hemisphere was drop-fixed in 4% paraformaldehyde for histology. For the biochemical analyses, the left hemisphere was cryopulverized and 25 mg of tissue was homogenized in radioimmunoprecipitation assay buffer supplemented with a cOmplete protease inhibitor (cat. no. 11836153001, Sigma-Aldrich) and PhosSTOP phosphatase inhibitor (cat. no. 4906837001, Sigma-Aldrich) unless specified otherwise. The homogenate was passed through a 20-g needle, agitated for 15 min using a Disrupter Genie (cat. no. 7403-2380, USA Scientific) and centrifuged for 30 min at 15,000 rpm at 4 °C. To extract the soluble and insoluble brain fractions, 25 mg of liquid-nitrogen-pulverized whole-brain tissue was sequentially homogenized with PBS and then 5 M guanidine buffer with protease and phosphatase inhibitors. PBS buffer was added at 20 μl mg−1; then, the homogenate was passed through a 20-g needle, agitated for 15 min and centrifuged for 30 min at 15,000 rpm at 4 °C. The supernatant was collected and saved as a PBS-soluble fraction. The same amount of 5 M guanidine buffer was added to the pellet and agitated for 15 min followed by 1-h rotation at room temperature. The homogenate was centrifuged for 30 min at 15,000 rpm at 4 °C. The supernatant was collected as an insoluble fraction.
Proteomics
Plasma ApoE was concentrated using the AlbuVoid Albumin Depletion Kit (cat. no. AVK10, Biotech Support Group) and filter centrifugation (cat. no. UFC503024, EMD Millipore). Brain homogenates were immunoprecipitated for ApoE using magnetic Dynabeads (cat. no. 10018D, Thermo Fisher Scientific) and incubated with 100 μg of sample for 1 h at room temperature. Samples were eluted from the beads using a denaturing elution with elution buffer and 2× Laemmli loading buffer (cat. no. S3401-10VL, Sigma-Aldrich). Equal amounts of protein were loaded (40 μg) with 2× Laemmli loading buffer and run on a 4–20% gel (cat. no. 5671024, Bio-Rad Laboratories). The gel was stained with SimplyBlue SafeStain (cat. no. LC6060, Thermo Scientific) for 1 h at room temperature.
Protein bands corresponding to ApoE (~36 kDa) were excised from the gel and subjected to a dithiothreitol reduction, iodoacetamide alkylation and in-gel trypsin digestion. Peptides were extracted and concentrated using filtration with a 0.22-μm polyvinylidene fluoride filter and then subjected to LC–MS/MS analysis using the Q Exactive Orbitrap Mass Spectrometer (Thermo Fisher Scientific). The LC–MS/MS datasets were searched in MASCOT against a custom database of exclusively human ApoE2 and ApoE4 peptides, including dynamic modifications of carbamidomethyl (C) and oxidation (M). The overall ratio of E2 to E4 peptides detected in each sample was calculated as the total intensity for specific E4 or E2 peptides.
Plasma lipid quantification
Experimental and control animals were fed either normal chow or a Western diet (cat. no. TD.88137, Envigo) for 4 months to exacerbate hyperlipidemic phenotypes. Blood was collected from fasted animals at 3 months (before the switch), and at the end of the 4-month study (after the switch). Total plasma TG, cholesterol and phospholipid levels were quantified using colorimetric assays (Fujifilm Wako Chemicals).
Plasma lipoprotein fractions were separated using fast protein liquid chromatography. Plasma samples were pooled to a final volume of 100 μl (n = 4 per group) and separations were performed on an ÄKTA pure instrument equipped with Superose 6 (GE Healthcare) columns arranged in series. Plasma was run over the columns at a flow rate of 0.5 ml min−1 in PBS buffer and 0.5-ml fractions were collected. TG, cholesterol and phospholipid levels were quantified in each fraction using colorimetric assays (Fujifilm Wako Chemicals).
ApoE ELISAs
A commercially available sandwich ELISA kit (cat. no. ab108813, Abcam) was used to measure total ApoE in the periphery and brain, and ApoE in soluble and insoluble whole-brain fractions. Plasma was diluted 1:400 and whole-brain homogenate was diluted 1:10. PBS-soluble brain fractions were diluted 1:10 and insoluble fractions were diluted 1:50. The average absorbance of each standard and sample were calculated and corrected by the dilution factor. ApoE concentration was calculated using the equation of the standard curve and normalized by total tissue weight.
Untargeted lipidomics
Untargeted lipidomic analysis was performed on whole-brain homogenate from 6–8-month-old TAM-treated APOE4s2G, APO4s2, E2-TR and E4-TR mice by a commercial company (BGI) using LC–MS/MS. A high-resolution Q Exactive mass spectrometer (Thermo Fisher Scientific) was used for data acquisition in positive-ion and negative-ion mode, respectively, to improve lipid coverage. Data were processed using LipidSearch v.4.1 and the BGI’s statistical software package, metaX. Metabolite abundance averages were calculated within groups and multiple group t-tests were performed on the entire dataset. Volcano plots were made for each condition in the dataset and also according to lipid class.
Brain single-cell suspension, cDNA library and sequencing
Mice were anesthetized using 5.0% isoflurane before exsanguination and transcardial perfusion with ice-cold Dulbecco’s PBS (DPBS) (cat. no. 14040133, Gibco). After perfusion, brains were quickly removed and whole right hemispheres sans brainstem and cerebellum were quickly minced using forceps on top of an ice-chilled Petri dish. Minced tissue from the hemispheres (n = 3 APOE4s2G, n = 3 APOE4s2A, n = 1 APOE4s2, n = 2 E2-TR and n = 1 E4-TR) were immediately transferred into a gentleMACS C-tube (cat. no. 130-093-237, Miltenyi Biotec) containing Adult Brain Dissociation Kit (ABDK) enzymatic digest reagents (cat. no. 130-107-677, Miltenyi Biotec) prepared according to the manufacturer’s protocol. Tissues were dissociated using the ‘37C_ABDK’ protocol on the gentleMACS Octo Dissociator instrument (cat. no. 130-095-937, Miltenyi Biotec) with heaters attached. After tissue digestion, cell suspensions were filtered through 70-μm mesh cell filters to remove debris according to the manufacturer’s suggested ABDK protocol. The resultant suspension was sequentially filtered (×2) using fresh 30-μm mesh filters. Cell viability was checked using the Acridine Orange/Propidium Iodide viability kit (cat. no. LGBD10012, Logos Biosystems). After viability and counting, cells were diluted to achieve a concentration of ~1,000 cells per 100 μl. The diluted cell suspensions were loaded onto the 10x Chromium instrument. Each sample was loaded into a separate channel on the Single Cell 3′ Chip and libraries were prepared using the Chromium v3 Single Cell 3′ Library and Gel Bead Kit (10x Genomics). Final library quantification and quality checking were performed using a Bioanalyzer instrument (Agilent Technologies); 150-bp paired-end sequencing (Novogene) was performed on a NovaSeq 6000 S4 flow cell.
scRNA-seq data processing
After libraries were sequenced and quality control was performed, samples were aligned with STAR aligner to the mm10 mouse reference genome using the Cell Ranger v.6.1.1 pipeline. The expression matrix was loaded into R for further analysis and visualization using Seurat (v.5.0.3). Ambient RNA contamination was removed using the SoupX R package. Doublets were removed using the scDblFinder package. Cells were then filtered to reduce the potential of including low-quality cells with the following criteria: 200 < nGene < 2000; 200 < nCount; and percent.mito <20%. Feature counts were normalized using sctransform and the effects of percent.mito were regressed out. Datasets were integrated using the Seurat standard protocol. Principal component analysis was performed and nearest neighbor analysis was done using the FindNeighbors function. FindClusters with a Louvain algorithm was used to identify clusters, and clustering was performed using a resolution of 1.0. Dimension reduction was performed using UMAP on the top 50 principal components. Forty-five clusters were identified. Four mixed clusters with high hemoglobin were removed along with several clusters with pituitary contamination (high expression of Gh, Prl and Lhx3).
To assign glial cell type identity to each cluster, we manually examined the expression levels of cell-type-specific markers across each cluster using the Partek Flow software to identify clusters containing unique populations of different cell types. Canonical CNS cell type markers were compiled from previous studies66,67,68,69. This process resulted in stringent filtering of cells with ambiguous assignments, leaving a total of 85,701 cells in 41 carefully assigned clusters.
GO analysis of DEGs
The Seurat function FindMarkers with the MAST differential expression test were used to conduct the DEG analysis via grouping for comparison according to clusters and treatment group. DEGs were selected if the Padj < 0.001 and the absolute value of the log fold change was higher than 1.0. Based on the identified DEGs, enrichment analyses of GO terms (Biological Process (BP2021)) were performed via Enrichr or the rWikiPathways R software package70, with cutoff represented by an FDR-adjusted P < 0.05. Bar plots of the top five most significant downregulated and upregulated pathways were generated for the visualizations.
Fluorescence-activated cell sorting
Mice were anesthetized using 5.0% isoflurane before exsanguination and transcardial perfusion with ice-cold DPBS. After perfusion, brains were quickly removed and whole right hemispheres sans brainstem and cerebellum were quickly minced using forceps on top of an ice-chilled Petri dish. Minced tissue from the hemispheres was immediately transferred into a gentleMACS C-tube containing ABDK enzymatic digest reagents prepared according to the manufacturer’s protocol. Tissues were dissociated using the ‘37C_ABDK’ protocol on the gentleMACS Octo Dissociator instrument with heaters attached. After tissue digestion, cell suspensions were filtered through 70-μm mesh cell filters to remove debris according to the manufacturer’s suggested ABDK protocol. The resultant suspension was sequentially filtered (×2) using fresh 30-μm mesh filters. Cell viability was checked using the Acridine Orange/Propidium Iodide viability kit. Cell suspensions were then subjected to fluorescence-activated cell sorting to sort tdTomato expression astrocytes from non-fluorescent Cre− cells in the brain. A total of ~110,000 cells were sorted and collected in DPBS. Cells were subsequently prepped for RNA isolation.
Immunofluorescence
Brains were post-fixed in 4% paraformaldehyde (cat. no. 043368.9M, Thermo Fisher Scientific) for 24 h at 4 °C then stored in a 30% sucrose solution with 0.05% sodium azide (cat. no. 76102-628, VWR). Coronal cryosections (30 μM) were obtained using a Leica Cryostat (CM1520) and serially collected in a cryoprotectant solution (50% 0.1 M PB, 30% ethylene glycol, 20% glycerol). Hippocampus-containing sections were selected for all stains, which were performed free-floating unless stated otherwise. For immunofluorescence detection of ApoE (1:500 dilution, cat. no. 13366S, Cell Signaling Technologies), GFAP (1:500 dilution, cat. no. 13-0300, Invitrogen), IBA1 (1:500 dilution, cat. no. 3NB100-1028, Novus Biologicals; 1:500 dilution, cat. no. ab178846, Abcam), MHC-II (1:200 dilution, cat. no. 14-5321-82, Thermo Fisher Scientific), PSD95 (1:500 dilution, cat. no. 51-6900, Invitrogen), anti-CAA mOC31 (1:100 dilution, cat. no. ab201059, Abcam), ZO1 (1:00 dilution, cat. no. 40-2200, Invitrogen) and CD31 (1:100 dilution, cat. no. ab56299, Abcam), sections were washed three times for 5 min in 1× PBS and permeabilized in 1× PBS + 0.2% Triton X-100 for 10 min. Sections were blocked in 10% goat serum in 1× PBS + 0.2% Triton X-100 (ApoE + GFAP) or 10% donkey serum in 1× PBS + 0.2% Triton X-100 (ApoE + IBA1) for 1 h at room temperature. Primary antibody cocktails were diluted in 3% goat or donkey serum and sections were incubated overnight at 4 °C. Sections were washed three times for 10 min in 1× PBS before applying the corresponding secondary antibodies (Goat anti-Rabbit Alexa Fluor 488, cat. no. A32731, Invitrogen; Goat anti-Rat Alexa Fluor 568, cat. no. A11077, Invitrogen; Donkey anti-Rabbit Alexa Fluor 488, cat. no. A32790, Invitrogen; Donkey anti-Goat Alexa Fluor 555, cat. no. A32816, Invitrogen) at a dilution of 1:250. Sections were incubated in secondary antibodies for 2 h at room temperature, protected from light. Sections were briefly washed in 1× PBS and mounted and coverslipped using a mounting medium with DAPI (cat. no. H-1500, Vector Laboratories) or NucSpot nuclear stain (cat. no. 23016, Biotium). For immunofluorescence detection of amyloid, sections were mounted onto slides after secondary incubation and allowed to dry. Slides were stained for amyloid plaques using Amylo-Glo (cat. no. TR-300-AG, Biosensis) and coverslipped with a mounting medium containing a far-red nuclear stain (cat. no. 23016, Biotium).
Fluorescence microscopy and HALO quantification
Imaging of GFAP, Iba1 and Amylo-Glo tissue-mounted slides was performed using the ZEISS AxioScan 7 to capture full mouse brain coronal sections. Images were uploaded to the Indica Labs HALO image analysis platform. All other tissue was imaged using a Nikon AX/AXR confocal microscope to capture the superior half (including the hippocampus and isocortex) of whole mouse brain coronal sections. All images were uploaded and analyzed with the Indica Labs HALO image analysis platform using the following algorithms: Area Quantification FL (v.2.3.3); Co-localization FL (v.2.1.4); and Proximity analysis (v.2.1.4).
Behavioral assessments
Associative fear conditioning
Associative learning and memory were assessed using the cued and contextual fear conditioning test over the course of 2 days. Mice were placed in a sound-attenuated chamber with grid flooring capable of delivering an electric shock. Freezing was measured with an overhead camera and the EthoVision XT software as a fear response output measure. On day 1, each chamber was prepared with context no. 1 consisting of white walls, gray metal grid flooring and MB-10 cleaning solution scent. Mice were placed in the chamber and were undisturbed for 3 min as baseline freezing was recording. On the third minute, a paired light and tone cue was played for 15 s; this was immediately followed by delivery of a 0.35-mA shock stimulus that lasted for 1 s. This cue and stimulus combination was repeated four times on every 60-s increment. On day 2, 24 h after day 1, mice were again placed in a chamber with context no. 1 for the contextual test. Mice were recorded for 5 min with no cues or stimulus to assess their ability to associate the context of the chamber from the conditioned fear response. After recording, mice were removed from the chambers. All chambers were thoroughly cleaned and prepared with context no. 2, which consisted of vertical black and white striped walls, white floors and a vanilla scent. Two hours later, mice performed the cued test, in which they were placed in chambers with context no. 2. Mice were undisturbed for 3 min, after which three paired light and tone cues were played for 15 s each on every 60-s interval. Percentage freezing was recorded in 60-s intervals.
Morris water maze
The water maze consisted of a circular pool, filled with opaque water (white paint added, 24 °C) divided conceptually into four quadrants. Mice were trained to locate an ‘escape’ platform (Plexiglas circle, 6-cm radius) submerged 2 cm below the surface of the water for five consecutive days during the ‘hidden platform’ trails, which required mice to rely on extra-maze cues for spatial reference and orientation. Extra-maze cues consisted of four large (50 × 50 cm) cues of different shapes and color combinations, positioned at the borders of the four quadrants. The platform was not rotated during the hidden platform trials, remaining in the ‘target’ quadrant. However, the location in which mice were dropped into the pool differed for each session between the four quadrants to avoid procedural bias in task learning. Mice were given two sessions per day (separated by 3 h) consisting of two trials each (separated by 10 min). Spatial memory retention was assessed 72 h after the final ‘hidden platform’ training session. The submerged platform was removed and spatial memory retention assessed during these ‘probe’ trials by analyzing the number of times mice crossed over the location of the platform and the time spent swimming in the target quadrant compared to the time spent in the three non-target quadrants.
Statistics
All values are reported as ± s.e.m. All statistical analyses were conducted in Prism 10 (GraphPad Software) unless noted otherwise; t-tests were used for assessing significance between two groups, while two-way ANOVAs were used to assess significance between more than two groups. Significance was determined as P < 0.05 for all tests unless stated otherwise. Significant P values, the value of n per group and experimental groups can be found in the figure legends.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The scRNA-seq data were deposited into the Gene Expression Omnibus under accession no. GSE300079. We reanalyzed data from the following existing sources: Murdock & Tsai (syn18485175GSE138852, GSE147528; syn21125841, GSE157827, GSE129308, GSE146639, GSE163577, GSE148822; and syn3219045, GSE181279, GSE134578, GSE160936); De Bastiani et al. (GSE149661, GSE145907, GSE28146, GSE29378, GSE36980, GSE48350, GSE84422 and GSE123496); Zhao et al. (syn22307008); Habib et al. (GSE143758); Keren-Shaul et al. (GSE98971); and Krasemann et al. (GSE101689). All other data supporting the findings of this study are available within the paper and its Supplementary Information.
References
Belloy, M. E. et al. APOE genotype and Alzheimer disease risk across age, sex, and population ancestry. JAMA Neurol. 80, 1284–1294 (2023).
Corder, E. et al. Protective effect of apolipoproteinE type 2 allele for late onset Alzheimer disease. Nat. Genet. 7, 180–184 (1994).
Garatachea, N. et al. ApoE gene and exceptional longevity: insights from three independent cohorts. Exp. Gerontol. 53, 16–23 (2014).
Wilson, R., Bienias, J., Berry-Kravis, E., Evans, D. & Bennett, D. The apolipoprotein E ε2 allele and decline in episodic memory. J. Neurol. Neurosurg. Psychiatry 73, 672–677 (2002).
Martens, Y. A. et al. ApoE cascade hypothesis in the pathogenesis of Alzheimer’s disease and related dementias. Neuron 110, 1304–1317 (2022).
Raulin, A.-C. et al. ApoE in Alzheimer’s disease: pathophysiology and therapeutic strategies. Mol. Neurodegener. 17, 72 (2022).
Brookhouser, N., Raman, S., Frisch, C., Srinivasan, G. & Brafman, D. A. APOE2 mitigates disease-related phenotypes in an isogenic hiPSC-based model of Alzheimer’s disease. Mol. Psychiatry 26, 5715–5732 (2021).
Lin, Y. T. et al. APOE4 causes widespread molecular and cellular alterations associated with Alzheimer’s disease phenotypes in human iPSC-derived brain cell types. Neuron 98, 1141–1154 (2018).
Dodart, J.-C. et al. Gene delivery of human apolipoprotein E alters brain Aβ burden in a mouse model of Alzheimer’s disease. Proc. Natl Acad. Sci. USA 102, 1211–1216 (2005).
Hudry, E. et al. Gene transfer of human Apoe isoforms results in differential modulation of amyloid deposition and neurotoxicity in mouse brain. Sci. Transl. Med. 5, 212ra161 (2013).
Zhao, L. et al. Intracerebral adeno-associated virus gene delivery of apolipoprotein E2 markedly reduces brain amyloid pathology in Alzheimer’s disease mouse models. Neurobiol. Aging 44, 159–172 (2016).
Jackson, R. J. APOE2 gene therapy reduces amyloid deposition and improves markers of neuroinflammation and neurodegeneration in a mouse model of Alzheimer disease. Mol. Ther. 32, 1373–1386 (2024).
Rosenburg, J. et al. AAVrh.10-mediated APOE2 central nervous system gene therapy for APOE4-associated Alzheimer’s disease. Hum. Gene Ther. Clin. Dev. 29, 24–47 (2018).
Gene therapy for APOE4 homozygote of Alzheimer's disease https://clinicaltrials.gov/study/NCT03634007 (ClinicalTrials.gov, 2024).
Wang, C. et al. Selective removal of astrocytic APOE4 strongly protects against tau-mediated neurodegeneration and decreases synaptic phagocytosis by microglia. Neuron 109, 1657–1674 (2021).
Huynh, T. V. et al. Age-dependent effects of apoE reduction using antisense oligonucleotides in a model of β-amyloidosis. Neuron 96, 1013–1023 (2017).
Liao, F. et al. Targeting of nonlipidated, aggregated apoE with antibodies inhibits amyloid accumulation. J. Clin. Invest. 128, 2144–2155 (2018).
Shi, Y. et al. Overexpressing low-density lipoprotein receptor reduces tau-associated neurodegeneration in relation to apoE-linked mechanisms. Neuron 109, 2413–2426 (2021).
Salomonsson, S. E. & Clelland, C. D. Building CRISPR gene therapies for the central nervous system: a review. JAMA Neurol. 81, 283–290 (2024).
Sullivan, P., Mezdour, H., Quarfordt, S. & Maeda, N. Type III hyperlipoproteinemia and spontaneous atherosclerosis in mice resulting from gene replacement of mouse Apoe with human Apoe*2. J. Clin. Invest. 102, 130–135 (1998).
Sullivan, P. M., Mace, B. E., Maeda, N. & Schmechel, D. E. Marked regional differences of brain human apolipoprotein E expression in targeted replacement mice. Neuroscience 124, 725–733 (2004).
Lefterov, I. et al. APOE2 orchestrated differences in transcriptomic and lipidomic profiles of postmortem AD brain. Alzheimers Res. Ther. 11, 113 (2019).
Fitz, N. F. et al. Phospholipids of APOE lipoproteins activate microglia in an isoform-specific manner in preclinical models of Alzheimer’s disease. Nat. Commun. 12, 3416 (2021).
Murdock, M. H. & Tsai, L.-H. Insights into Alzheimer’s disease from single-cell genomic approaches. Nat. Neurosci. 26, 181–195 (2023).
De Bastiani, M. A. et al. Cross-species evidence of differential expression of S100A6 and SLC11A1 in the hippocampus of Alzheimer’s disease patients and mouse models. Alzheimers Dement. 19, e060565 (2023).
Zhao, N. et al. Alzheimer’s risk factors age, APOE genotype, and sex drive distinct molecular pathways. Neuron 106, 727–742 (2020).
Habib, N. et al. Disease-associated astrocytes in Alzheimer's disease and aging. Nat. Neurosci. 23, 701–706 (2020).
Keren-Shaul, H. et al. A unique microglia type associated with restricting development of Alzheimer’s disease. Cell 169, 1276–1290 (2017).
Xu, Q. et al. Profile and regulation of apolipoprotein E (ApoE) expression in the CNS in mice with targeting of green fluorescent protein gene to the ApoE locus. J. Neurosci. 26, 4985–4994 (2006).
Srinivasan, R. et al. New transgenic mouse lines for selectively targeting astrocytes and studying calcium signals in astrocyte processes in situ and in vivo. Neuron 92, 1181–1195 (2016).
Castellano, J. M. et al. Human apoE isoforms differentially regulate brain amyloid-β peptide clearance. Sci. Transl. Med. 3, 89ra57 (2011).
Sala Frigerio, C. et al. The major risk factors for Alzheimer’s disease: age, sex, and genes modulate the microglia response to Aβ plaques. Cell Rep. 27, 1293–1306 (2019).
Lee, S. et al. APOE modulates microglial immunometabolism in response to age, amyloid pathology, and inflammatory challenge. Cell Rep. 42, 112196 (2023).
Filippov, V. et al. Increased ceramide in brains with Alzheimer’s and other neurodegenerative diseases. J. Alzheimers Dis. 29, 537–547 (2012).
Exton, J. H. Signaling through phosphatidylcholine breakdown. J. Biol. Chem. 265, 1–4 (1990).
Mahan, T. E. et al. Selective reduction of astrocyte apoE3 and apoE4 strongly reduces Aβ accumulation and plaque-related pathology in a mouse model of amyloidosis. Mol. Neurodegener. 17, 13 (2022).
Tcw, J. et al. Cholesterol and matrisome pathways dysregulated in astrocytes and microglia. Cell 185, 2213–2233 (2022).
Ostendorf, B. N. et al. Common germline variants of the human APOE gene modulate melanoma progression and survival. Nat. Med. 26, 1048–1053 (2020).
Adams, M. K. M. et al. Apolipoprotein E gene associations in age-related macular degeneration: the Melbourne Collaborative Cohort Study. Am. J. Epidemiol. 175, 511–518 (2012).
Johnson, L. A. et al. ApoE2 exaggerates PTSD-related behavioral, cognitive, and neuroendocrine alterations. Neuropsychopharmacology 40, 2443–2453 (2015).
Wang, H. Whole-genome sequencing analysis reveals new susceptibility loci and structural variants associated with progressive supranuclear palsy. Mol. Neurodegener. 19, 61 (2024).
Johnson, L. A. et al. Apolipoprotein E-low density lipoprotein receptor interaction affects spatial memory retention and brain ApoE levels in an isoform-dependent manner. Neurobiol. Dis. 64, 150–162 (2014).
Guo, J. L. et al. Decreased lipidated ApoE-receptor interactions confer protection against pathogenicity of ApoE and its lipid cargoes in lysosomes. Cell 188, 187–206 (2025).
Koutsodendris, N. et al. Neuronal APOE4 removal protects against tau-mediated gliosis, neurodegeneration and myelin deficits. Nat. Aging 3, 275–296 (2023).
Anfray, A. et al. A cell-autonomous role for border-associated macrophages in ApoE4 neurovascular dysfunction and susceptibility to white matter injury. Nat. Neurosci. 27, 2138–2151 (2024).
Liu, C.-C.et al. ApoE4 accelerates early seeding of amyloid pathology. Neuron 96, 1024–1032 (2017).
Boyle, P. A. et al. Cerebral amyloid angiopathy and cognitive outcomes in community-based older persons. Neurology 85, 1930–1936 (2015).
Nelson, P. T. et al. APOE-ε2 and APOE-ε4 correlate with increased amyloid accumulation in cerebral vasculature. J. Neuropathol. Exp. Neurol. 72, 708–715 (2013).
Zabel, M. et al. A shift in microglial β-amyloid binding in Alzheimer’s disease is associated with cerebral amyloid angiopathy. Brain Pathol. 23, 390–401 (2013).
Munsterman, D. et al. Cerebral amyloid angiopathy and the immune system. Alzheimers Dement. 20, 4999–5008 (2024).
Gal, J. et al. Apolipoprotein E proteinopathy is a major dementia-associated pathologic biomarker in individuals with or without the APOE Epsilon 4 Allele. Am. J. Pathol. 192, 564–578 (2022).
Strittmatter, W. J. et al. Binding of human apolipoprotein E to synthetic amyloid beta peptide: isoform-specific effects and implications for late-onset Alzheimer disease. Proc. Natl Acad. Sci. USA 90, 8098–8102 (1993).
LaDu, M. J. et al. Purification of apolipoprotein E attenuates isoform-specific binding to β-amyloid. J. Biol. Chem. 270, 9039–9042 (1995).
LaDu, M. J. et al. Isoform-specific binding of apolipoprotein E to β-amyloid. J. Biol. Chem. 269, 23403–23406 (1994).
Tai, L. M. et al. Levels of soluble apolipoprotein E/amyloid-β (Aβ) complex are reduced and oligomeric Aβ increased with APOE4 and Alzheimer disease in a transgenic mouse model and human samples. J. Biol. Chem. 288, 5914–5926 (2013).
Hashimoto, T. et al. Apolipoprotein E, especially apolipoprotein E4, increases the oligomerization of amyloid β peptide. J. Neurosci. 32, 15181–15192 (2012).
Ulrich, J. D. et al. ApoE facilitates the microglial response to amyloid plaque pathology. J. Exp. Med. 215, 1047–1058 (2018).
Lau, S.-F. et al. The VCAM1–ApoE pathway directs microglial chemotaxis and alleviates Alzheimer’s disease pathology. Nat. Aging 3, 1219–1236 (2023).
Griswold, A. J. et al. Increased APOE ε4 expression is associated with the difference in Alzheimer’s disease risk from diverse ancestral backgrounds. Alzheimers Dement. 17, 1179–1188 (2021).
Huggins, L. K. L. et al. Meta-analysis of variations in association between APOE ɛ4 and Alzheimer’s disease and related dementias across Hispanic regions of origin. J. Alzheimers Dis. 93, 1095–1109 (2023).
& Choi, K. Y. APOE promoter polymorphism-219T/G is an effect modifier of the influence of APOE ε4 on Alzheimer's disease risk in a multiracial sample. J. Clin. Med. 8, 1236 (2019).
Eisenberg, D. T., Kuzawa, C. W. & Hayes, M. G. Worldwide allele frequencies of the human apolipoprotein E gene: climate, local adaptations, and evolutionary history. Am. J. Phys. Anthropol. 143, 100–111 (2010).
Tai, L. M. et al. EFAD transgenic mice as a human APOE relevant preclinical model of Alzheimer’s disease. J. Lipid Res. 58, 1733–1755 (2017).
Balu, D. et al. The role of APOE in transgenic mouse models of AD. Neurosci. Lett. 707, 134285 (2019).
Nemeth, D., Luqman, N., Chen, L. & Quan, N. Aldh1l1-Cre/ER T2 is expressed in unintended cell types of the salivary gland, pancreas, and spleen. MicroPubl. Biol. https://doi.org/10.17912/micropub.biology.000832 (2023).
McKenzie, A. T. et al. Brain cell type specific gene expression and co-expression network architectures. Sci. Rep. 8, 8868 (2018).
Zeisel, A. et al. Molecular architecture of the mouse nervous system. Cell 174, 999–1014 (2018).
Ximerakis, M. et al. Single-cell transcriptomic profiling of the aging mouse brain. Nat. Neurosci. 22, 1696–1708 (2019).
Han, X. et al. Mapping the mouse cell atlas by Microwell-seq. Cell 172, 1091–1107 (2018).
Slenter, D. N. et al. WikiPathways: a multifaceted pathway database bridging metabolomics to other omics research. Nucleic Acids Res. 46, D661–D667 (2018).
Acknowledgements
We thank D. Adreon, C. Friday and H. Williams for their invaluable feedback. We thank the University of Kentucky Proteomics Core for their help with proteomics analysis and the Animal Behavior Core for their assistance with behavioral testing. This work was supported by the National Institute of Health (grant nos. R01AG062550, R01AG081421 and R01AG080589 to L.A.J.; grant nos. R01AG070830 and RF1NS118558 to J.M.M.; grant nos. R01AG068330 and R01AG093847 to S.L.M.; grant no. R01DK133184 to S.M.G.; grant no. T32AG057461 to L.R.G.), the CNS Metabolism COBRE P20 GM148326 (J.M.M., S.L.M. and L.A.J.), BrightFocus Foundation A20201775S (S.L.M.), Coins for Alzheimer’s Research Trust Grant (S.L.M.) and the Alzheimer’s Association (L.A.J., J.M.M., S.M.G.).
Author information
Authors and Affiliations
Contributions
L.A.J., J.M.M., S.M.G., S.L.M. and L.R.G. designed the experiments. L.R.G. and L.A.J. analyzed the data and wrote the manuscript. J.M.M. and L.A.J. supervised the scRNA-seq workflow, with data analysis by K.S, S.L. and S.M.M. C.S. and S.L. contributed to the lipidomics analyses. D.S.S., I.O.S., J.L.F., G.L.N. and A.V.P. assisted with the biochemical and immunohistochemical analyses. G.H. and B.C.F. assisted with experimental design, organization and implementation of mouse studies. J.C. and H.Z. completed the proteomics analyses. S.M.G. supervised the plasma lipid analyses with technical assistance from C.M.W. and C.V. D.O.C. and K.P. designed and generated the transgenic animals with additional design input from L.A.J. All authors read the paper and provided edits.
Corresponding author
Ethics declarations
Competing interests
D.O.C. is employed by, has equity ownership in and serves on the board of directors of TransViragen, the company which has been contracted by UNC-Chapel Hill to manage its Animal Models Core Facility. The other authors declare no competing interests.
Peer review
Peer review information
Nature Neuroscience thanks Guojun Bu, Hui Zheng and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Peaks for unique E2 and unique E4 peptides look similar in TR mice and APOE4s2 and APOE4s2G mice.
a,d,g,j, Extracted-ion chromatogram (XIC) of peaks from unique E4 peptide LGADMEDVR in an E4-TR control sample (a) and an APOE4s2 sample expressing E4 protein (d) or peaks from unique E2 peptide LGADMEDVRGC in and E2-TR control sample (g) or an APOE4s2G sample (j) expressing E2 protein. b,e,h,k, Mass spectra (MS) showing peaks with similar mass to charge ratios of unique E4 peptide LGADMEDVR in an E4-TR control sample (b) and an APOE4s2 sample expressing E4 protein (e) or peaks from unique E2 peptide LGADMEDVRGC in and E2-TR control sample (h) or an APOE4s2G sample (k) expressing E2 protein. c,f,i,l, Tandem mass spectra (MS/MS) showing ion fragment peaks with similar mass to charge ratios of unique E4 peptide LGADMEDVR in an E4-TR control sample (c) and an APOE4s2 sample expressing E4 protein (f) or peaks from unique E2 peptide LGADMEDVRGC in and E2-TR control sample (i) or an APOE4s2G sample (l) expressing E2 protein. m, Peptide sequences unique to ApoE2 or ApoE4 at amino acid residues 112 and 159 following trypsin digestion. Sequences were used to create a custom database of hApoE2 and hApoE4 peptides. LC-MS/MS data sets generated from plasma and brain samples were searched in against these sequences at both amino acid residues to generate a semi-quantitative ratio of E2:E4 in APOE4s2 and APOE4s2G samples.
Extended Data Fig. 2 Peripheral and cerebral metabolic profiling of APOE4s2 and ApoE-TR mice and ApoE colocalization with glia.
a, Allelic discrimination plot of mRNA analysis showing efficient and sustained expression of APOE2 in APOE4s2G mice at 72 h, 3 months, 6 months, and 12 months post switch compared to E2-TR, E4-TR, and APOE4s2 controls (nE2-TR = 6, nE4-TR = 6, nAPOE4s2 = 6, nAPOE4s2G = 6 for all timepoints).). b, Percent change in bodyweight recorded between mice on western diet (APOE4s2G: n = 12 total, n = 6 female, n = 6 male; APOE4s2: n = 13 total, n = 6 females, n = 7 males; E2-TR: n = 18 total, n = 10 females, n = 8 males; E4-TR: n = 12 total, n = 9 females, n = 3 males; no significance; 2way ANOVA with multiple comparisons). c, ApoE quantified in SEC fractions corresponding to VLDL, IDL/LDL and HDL lipoprotein classes. d,f,h, Triglycerides (d), total cholesterol (f), and phospholipids (h) quantified in plasma from E2-TR, E4-TR, APOE4s2, and APOE4s2G mice on normal chow (nE2-TR = 3, nE4-TR = 3, nAPOE4s2 = 5, nAPOE4s2G = 5; *p < 0.05, **p < 0.01, ****p < 0.0001; 2way ANOVA with multiple comparisons). e,g,i, Triglycerides (e), total cholesterol (g), and phospholipids (i) quantified in FPLC eluants corresponding to VLDL, LDL and HDL (nE2-TR = 3 pooled, nE4-TR = 3 pooled, nAPOE4s2 = 4 pooled, nAPOE4s2G = 4 pooled). j, Representative images of ApoE (red), NeuN (green), and DAPI (blue) staining in APOE4s2G and APOE4s2 controls. Scale is 100um. k, Representative images of ApoE (red) and NeuN (green) with DAPI nuclear stain (blue) in E4-TR and E2-TR brains. Scale is 100um. l, Representative images of ApoE (red), IBA1 (green), and DAPI (blue) in E4-TR and E2-TR mice. Scale is 100um. m, Representative images of ApoE (red), GFAP (green), and DAPI (blue) in E4-TR and E2-TR brains. Scale is 100um. n, Quantification of ApoE with astrocytes, microglia, and neurons in the hippocampus. *p < 0.05, ***p < 0.001, 2way ANOVA with multiple comparisons. o, Heatmap depicting top 10 most significantly changed lipids between E2-TR and E4-TR controls. p, Volcano plot showing differentially expressed lipids betweenE4-TR vs. E2-TR mice (nE2-TR = 8, nE4-TR = 8; all females). (FDR adjusted p < 0.05 (log10(P) > 1.3)). For detailed statistical information, see supplementary table 3. Data represent mean ± s.e.m. of biological replicates. Individual data points plotted. Box plots show center line, median; box limits, upper and lower quartiles; whiskers, minimum and maximum.
Extended Data Fig. 3 scRNAseq cell-type assignment.
a, UMAP identifying a total of 41 unique clusters before cell-type assignment. b, UMAP showing relatively even distribution of clusters across genotypes. c, After cell-type assignment, plots show relatively even distribution of cell type across different genotypes. d, Violin plots showing expression of gene markers in each cell type. e, Gene expression in each cell-type cluster of cell-specific gene markers.
Extended Data Fig. 4 Plotted DEGs for all single cell comparisons.
a, Volcano plots depicting differentially expressed genes in astrocytes identified between APOE4s2A vs. APOE4s2, APOE4s2G vs. APOE4s2, and E2-TR vs. E4-TR comparisons. p = 0.001, log2FC = 1.0. b, Volcano plots depicting differentially expressed genes in oligodendrocytes identified between APOE4s2A vs. APOE4s2, APOE4s2G vs. APOE4s2, and E2-TR vs. E4-TR comparisons. p = 0.001, log2FC = 1.0. c, Volcano plots depicting differentially expressed genes in microglia identified between , APOE4s2A vs. APOE4s2, APOE4s2G vs. APOE4s2, and E2-TR vs. E4-TR comparisons. p = 0.001, log2FC = 1.0. d, Volcano plots depicting differentially expressed genes in endothelial cells identified between APOE4s2A vs. APOE4s2, APOE4s2G vs. APOE4s2, and E2-TR vs. E4-TR comparisons. p = 0.001, log2FC = 1.0.
Extended Data Fig. 5 Validation and plasma lipid profiling of astrocyte-selective APOE4s2A mice.
a, Allelic discrimination plot of mRNA analysis from whole brain tissue showing efficient and sustained expression of APOE2 in APOE4s2A mice at 72 h, 3 months, 6 months, and 12 months post switch compared to E2-TR, E4-TR, and APOE4s2 controls (nE2-TR = 6, nE4-TR = 6, nAPOE4s2 = 6, nAPOE4s2A = 6 for all timepoints) b, Ratio of unique E2 vs. E4 peptides detected using mass spec based proteomic analysis (nE2-TR = 2, nE4-TR = 2, nAPOE4s2 = 3, nAPOE4s2A = 3). c, Percent td-tomato Aldh1l1-Cre reporter positive area vs. tdTomato Aldh1l1-Cre reporter colocalized with GFAP staining in APOE4s2A (65%) mice vs. APOE4s2 Cre-negative controls (4%%) (APOE4s2A = 3, nAPOE4s2 = 3). d-e, Allelic discrimination plot from liver tissue (d) with ratios of relative fluorescence from SNPs associated with APOE2 vs. APOE4 showing an average of 81% APOE2 mRNA expression in the liver (e) (nE2-TR = 6, nE4-TR = 6, nAPOE4s2 = 6, nAPOE4s2A = 6). f, ApoE quantified by ELISA in SEC fractions corresponding to VLDL, IDL/LDL and HDL lipoprotein classes. g-i, Triglycerides (g), total cholesterol (h), and phospholipids (i) quantified in plasma from E2-TR, E4-TR, APOE4s2, and APOE4s2A mice on normal chow (nE2-TR = 3, nE4-TR = 3, nAPOE4s2 = 5, nAPOE4s2G = 5; *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001; 2way ANOVA with multiple comparisons). j-l Triglycerides (j), total cholesterol (k), and phospholipids (l) quantified in FPLC eluants corresponding to VLDL, LDL and HDL lipoprotein classes (nE2-TR = 3 pooled, nE4-TR = 3 pooled, nAPOE4s2 = 4 pooled, nAPOE4s2G = 4 pooled). a-i, For detailed statistical information, see supplementary table 3. Data represent mean ± s.e.m. of biological replicates. Individual data points plotted. Box plots show center line, median; box limits, upper and lower quartiles; whiskers, minimum and maximum.
Extended Data Fig. 6 Sex stratified analysis of behavior showing improvements in associative, but not spatial learning and memory.
a-d, Percent time freezing measured during conditioning in 4s2A/FAD vs. 4s2/FAD stratified by sex. Analyses showed only one significant sex difference where 4s2/FAD males froze more than 4s2/FAD females. (n4s2A/FAD = 8 females; 8 males, n4s2/FAD = 8 females; 8 males; *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001; Repeated measures ANOVA with multiple comparisons; symbols denote significance by sex(es). e, Latency to platform measured in 4s2A/FAD compared to tamoxifen treated, Cre-negative 4s2/FAD controls over the 5-day hidden platform training trials (n4s2A/FAD = 10, n4s2/FAD = 10; two-tailed, paired t-test). f, Thigmotaxis (defined by 30 s of swimming around perimeter) in 4s2A/FAD and 4s2/FAD controls during hidden platform trials (n4s2A/FAD = 10, n4s2/FAD = 10; two-tailed, paired t-test). g, Latency to platform by sex showing no sex differences in 4s2A/FAD or 4s2/FAD controls. (n4s2A/FAD = 5 females; 5 males, n4s2/FAD = 5 females; 5 males; 2way ANOVA with repeated measures and multiple comparisons). h, There were no significant sex-based differences in thigmotaxis (n4s2A/FAD = 5 females; 5 males, n4s2/FAD = 5 females; 5 males 2way ANOVA with repeated measures and multiple comparisons). i-k, Number of platform crosses (i), occupancy of target quadrant (j), and occupancy of target quadrant compared to other quadrants (k) in the 72-hour memory probe. (n4s2A/FAD = 5 females; 5 males, n4s2/FAD = 5 females; 5 males; **p < 0.01, ***p < 0.001, ****p < 0.0001; two-tailed, unpaired t-test (I,k); 2way ANOVA with multiple comparisons (k)). l-n, Sex-based analyses of platform crosses (l), time in target quadrant (m), and frequency in quadrants relative to other quadrants (n) showed no sex differences (n4s2A/FAD = 5 females; 5 males, n4s2/FAD = 5 females; 5 males; *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001; 2way ANOVA with multiple comparisons. a-n. For detailed statistical information, see supplementary table 3. Data represent mean ± s.e.m. of biological replicates. Individual data points plotted. Sex is denoted by symbol shape: females (circle), males (triangle). Box plots show center line, median; box limits, upper and lower quartiles; whiskers, minimum and maximum.
Extended Data Fig. 7 Amyloid deposition by sex, plaque toxicity, and tight junction protein quantification.
a, Quantification of total amylo-glo+ area by sex showing significantly higher amyloid+ area in 4s2/FAD females than 4s2/FAD males (n4s2A/FAD = 8 females; 8 males, n4s2/FAD = 8 females; 8 males; **p < 0.01; 2way ANOVA with multiple comparisons). b, Representative image of amylo-Glo (blue) and nuclei (red) staining in 6-month-old (pre-switch) 4s2/FAD mice. Scale is 1 mm. c, Percent amyloid+ area quantified over the 2-month period of astrocytic E2 expression showing a significant increase in amyloid+ percent area in8-month-old 4s2/FAD mice. (n4s2A/FAD-6mo = 8, n4s2/FAD-8mo. = 16, n4s2A/FAD-8mo. = 16; **p < 0.01; 2way ANOVA with multiple comparisons). d, Average plaque size at 8 mo. between 4s2/FAD controls and 4s2A/FAD mice (n4s2/FAD = 7, n4s2A/FAD = 7; two-tailed, unpaired t-test). e-h, Sex based analyses ELISA quantification of soluble and insoluble Ab42 and Ab40 species show no sex differences but suggest decreases in amyloid could be driven by trends within one sex (n4s2A/FAD = 7 females; 7 males, n4s2/FAD = 7 females; 7 males; *p < 0.05, **p < 0.01, ***p < 0.001, 2way ANOVA with multiple comparisons). i, Representative image of proximity analysis used to quantify PSD-95 puncta in 3um increments up to 15um from the plaque. PSD-95 (green), Amylo-Glo (blue), nuclei (red). Scale is 100um. j, l, PSD-95 quantification in the cortex showing decreases in PSD-95 in 4s2A/FAD mice starting at 12um from plaque center (j). Sex based analyses revealed that 4s2/FAD females had significantly more PSD-95 puncta than 4s2/FAD males farther from the plaque (l) (n4s2A/FAD = 6 females; 6 males, n4s2/FAD = 6 females; 6 males; **p < 0.01, ***p < 0.001, ****p < 0.0001; 2way ANOVA with multiple comparisons). k, m, PSD-95 quantification in the hippocampus (n4s2A/FAD = 6 females; 6 males, n4s2/FAD = 6 females; 6 males; 2way ANOVA with multiple comparisons). n, Representative images of CD31 (red), ZO1 (green), and DAPI (blue) staining. Scale is 100um. o, q, Quantification of tight junction protein ZO1 (o) with sex stratified analyses showing that 4s2A/FAD females have higher ZO1 than 4s2A/FAD males (q) (n4s2A/FAD = 8 females; 8 males, n4s2/FAD = 8 females; 8 males; **p < 0.01; 2way ANOVA with multiple comparisons). p, r, Quantification of total % area of CD31+ vessels (n4s2A/FAD = 8 females; 8 males, n4s2/FAD = 8 females; 8 males; 2way ANOVA with multiple comparisons). a-r, Detailed statistical information in supplementary table 3. Data represent mean ± s.e.m. of biological replicates. Individual data points plotted. Sex is denoted by symbol shape: females (circle), males (triangle). Box plots show center line, median; box limits, upper and lower quartiles; whiskers, minimum and maximum.
Extended Data Fig. 8 Amyloid-associated gliosis measures stratified by sex.
a,c, Quantification of total percent GFAP+ (a) or IBA1+ (c) area in 4s2A/FAD and 4s2/FAD controls at 8 months compared to baseline measures of gliosis at 6 months (nAPOE4s2-6mo = 8, nAPOE4s2-8mo. = 16, nAPOE4s2A-8mo. = 16; **p < 0.01, ****p < 0.000, 2way ANOVA with multiple comparisons). b,d, Quantification by sex of total percent GFAP+ (a) or IBA1+ (c) area in 4s2A/FAD and 4s2/FAD controls at 8 months compared to baseline measures of gliosis at 6 months (nAPOE4s2-6mo = 8, nAPOE4s2-8mo. = 8 females; 8 males, nAPOE4s2A-8mo. = 8 females; 8 males; *p < 0.05, **p < 0.01, ****p < 0.0001; 2way ANOVA with multiple comparisons). e-g, Sex stratified analyses for colocalization of GFAP+ astrocytes and plaque (e), IBA1+ microglia and plaque (f), and plaque with both GFAP+ astrocytes and IBA1+ microglia (g). (nAPOE4s2-6mo = 8, nAPOE4s2-8mo. = 8 females; 8 males, nAPOE4s2A-8mo. = 8 females; 8 males; *p < 0.05, **p < 0.01; 2way ANOVA with multiple comparisons). h,j, Representative images exemplifying plaque and glial cell proximity analyses in HALO software for proximity analysis of amyloid plaques and GFAP positive cells (h), or amyloid plaques and IBA1 positive cells (j). Scale is 100um. i-k, Plaque proximity analyses by sex (nAPOE4s2-6mo = 4, nAPOE4s2-8mo. = 4 females; 4 males, nAPOE4s2A-8mo. = 4 females; 4 males; *p < 0.05; 2way ANOVA with multiple comparisons). a-k, Detailed statistical information in supplementary table 3. Data represent mean ± s.e.m. of biological replicates. Individual data points plotted. Sex is denoted by symbol shape: females (circle), males (triangle). Box plots show center line, median; box limits, upper and lower quartiles; whiskers, minimum and maximum.
Extended Data Fig. 9 Sex based analyses of microglia sub-populations and plaque-associated ApoE.
a-c, Sex stratified quantification of cortical MHCII+ percent area (a) MHCII+ and IBA1+ percent area (b) or MHCII + , IBA1 + , and amylo-glo+ percent area (c) (nAPOE4s2-6mo = 8, nAPOE4s2-8mo. = 8 females; 8 males, nAPOE4s2A-8mo. = 8 females; 8 males; *p < 0.05, **p < 0.01, ***p < 0.001. 2way ANOVA with multiple comparisons). d-f, Sex stratified quantification of hippocampal MHCII+ percent area (d) MHCII+ and IBA1+ percent area (e) or MHCII + , IBA1 + , and amylo-glo+ percent area (f) (nAPOE4s2-6mo = 8, nAPOE4s2-8mo. = 8 females; 8 males, nAPOE4s2A-8mo. = 8 females; 8 males; *p < 0.05, **p < 0.01, ***p < 0.001, 2way ANOVA with multiple comparisons). g-h, Sex stratified analysis of homeostatic (g) and DAM (h) gene expression showed no sex differences between groups. i, Representative images of ApoE (green), amylo-glo (blue), and IBA1 (orange) staining. Scale is 100um. j-k, Quantification of percent area of total ApoE in the cortex (j) or cortical plaque-associated ApoE (k) with significant reduction in total cortical ApoE, as well as plaque-associated ApoE, appearing to be driven by 4s2A/FAD females (nAPOE4s2-6mo = 8, nAPOE4s2-8mo. = 8 females; 8 males, nAPOE4s2A-8mo. = 8 females; 8 males; *p < 0.05. 2way ANOVA with multiple comparisons. l-m, Quantification of percent area of total ApoE in the hippocampus (l) or plaque-associated ApoE (m) (nAPOE4s2-6mo = 8, nAPOE4s2-8mo. = 8 females; 8 males, nAPOE4s2A-8mo. = 8 females; 8 males; 2way ANOVA with multiple comparisons). n-o, ELISA quantification of total brain soluble ApoE (n) and insoluble ApoE (o) by sex 4s2/FAD females had higher insoluble ApoE than males. (nAPOE4s2-6mo = 8, nAPOE4s2-8mo. = 8 females; 8 males, nAPOE4s2A-8mo. = 8 females; 8 males; *p < 0.05, **p < 0.01. 2way ANOVA with multiple comparisons. a-o, Detailed statistical information in supplementary table 3. Data represent mean ± s.e.m. of biological replicates. Individual data points plotted. Sex is denoted by symbol shape: females (circle), males (triangle). Box plots show center line, median; box limits, upper and lower quartiles; whiskers, minimum and maximum.
Supplementary information
Supplementary Table 1 (download XLSX )
Differentially expressed genes according to cell type for APOE4s2G versus APOE4s2, APOE4s2A versus APOE4s2, and E2-TR and E4-TR comparisons.
Supplementary Table 2 (download XLSX )
GO terms according to cell type for APOE4s2G versus APOE4s2, APOE4s2A versus APOE4s2, and E2-TR versus E4-TR comparisons.
Supplementary Table 3 (download XLSX )
Detailed statistical test information according to the figures.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Golden, L.R., Siano, D.S., Stephens, I.O. et al. APOE4 to APOE2 allelic switching in mice improves Alzheimer’s disease-related metabolic signatures, neuropathology and cognition. Nat Neurosci 28, 2461–2475 (2025). https://doi.org/10.1038/s41593-025-02094-y
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41593-025-02094-y
This article is cited by
-
Protective mutations associated with APOE in Alzheimer’s disease
Molecular Psychiatry (2026)
-
Flipping APOE to reverse Alzheimer’s disease traits
Nature Neuroscience (2025)