Abstract
The brain moves within the skull, but the drivers and consequences of this motion are not well understood. Here we visualized motion of the dorsal cortex relative to the skull in awake head-fixed mice using high-speed, multiplane two-photon microscopy. Brain motion was directed primarily rostrally and laterally, and was correlated tightly with locomotion, but not with respiration or the cardiac cycle. Specifically, brain motion was driven by abdominal muscle contractions that activate a hydraulic-like vascular connection between the nervous system and the abdominal cavity, and could similarly be induced by pressure applied to the abdomen. Model simulations suggest that brain motion may drive interstitial fluid through and out of the brain into the subarachnoid space, in the opposite direction of fluid flow seen during sleep. These results suggest that the brain is linked mechanically to the abdominal compartment, and that fluid flow in the brain could be coupled to body movements.
Similar content being viewed by others
Main
Brain motion is a ubiquitous but poorly investigated phenomenon1,2. In anesthetized animals and supine humans, brain motion is tied closely to cardiac pulsations and respiration3,4 but in unanesthetized animals, brain motion is usually associated with locomotion and other body movements2,5. In mice, brain motion observed with two-photon microscopy is on the order of a few microns2,6,7 and is primarily within the imaging plane (medial-lateral/rostral-caudal).
Despite the ubiquity of brain motion in the awake animal, its origins are not well understood. A force must be exerted on the brain for it to move, but the central nervous system has been considered to be largely mechanically insulated from the rest of the body by the skull and vertebrae. Despite this partitioning, during locomotion the intracranial pressure of mice rises from a baseline of ~5 mm Hg to more than 20 mm Hg (refs. 8,9), indicating that substantial mechanical forces are applied rapidly to the brain during body movements. The increase in intracranial pressure during locomotion is not due to dilation of blood vessels within the brain, as the hemodynamic response lags both the pressure increase and the onset of locomotion by ~1 s (refs. 10,11). Furthermore, maximally dilating the vessels of the brain does not increase intracranial pressure to the levels seen during locomotion8. These pressure changes are unlikely to be simply an epiphenomenon because brain motion during locomotion excites sensory neurons in the dura12, indicating that the motion of the brain is monitored actively and may serve a physiological role.
One potential physiological purpose for brain motion is to circulate interstitial fluid (ISF) and cerebrospinal fluid (CSF) in the brain. As the brain lacks an internal lymphatic system to remove waste, it depends on mechanical forces exerted on it by pulsation13 and dilation and constrictions14,15,16 of arteries to help circulate fluid though the glymphatic system, along the periarterial space into the parenchyma. During sleep, CSF is driven into the brain along the periarterial spaces of penetrating arteries by slow, alternating dilation and constriction of the vessel17,18,19,20. The patterns of fluid flow in the brain differ markedly from those in the awake animal, where tracers do not enter the cortex21, although the reasons for this difference between sleep and wake CSF flow are not completely understood. The large forces that drive brain motion are also likely to drive movement of CSF, potentially in very different patterns to those seen during sleep. Understanding these fluid flows requires a detailed characterization of the mechanical dynamics of the brain.
Results
We used two-photon (2P) microscopy to quantify brain motion relative to the skull in 24 Swiss Webster mice (12 male) that were head-fixed on a spherical treadmill. We simultaneously imaged brain cells expressing green fluorescent protein22 and fluorescent microspheres attached to a polished and reinforced thinned-skull (PoRTS) window23. This was accomplished by integrating an electrically tunable lens behind the microscope objective to rapidly (39.55 frames per second, 19.78 frames per second per plane) alternate between two focal planes on the skull surface and in the brain (Fig. 1, Extended Data Fig. 1 and Supplementary Fig. 1), separated by ~90 µm. We used the point spread function in z (Extended Data Fig. 1c) to estimate axial motion of the microspheres on the skull and of the brain from the slight changes in fluorescence intensity. We estimate that both brain and skull move ~1 µm from baseline during locomotion (Extended Data Fig. 2a–c), suggesting there is no relative z motion between the skull and brain. Furthermore, tracking of microspheres showed that skull movement was usually less than 1 µm (Extended Data Fig. 3), demonstrating the stability of the head fixation apparatus. The tracking software was tested for stability and accuracy on static paper coated in fluorescein isothiocyanate (FITC) (Extended Data Fig. 2d,e) and microspheres (Extended Data Fig. 2f,g), to simulate the brain and skull environments respectively. The motion of the brain relative to the skull was primarily in the rostral and lateral directions (Fig. 1e) and was correlated strongly with locomotion (Fig. 2d and Supplementary Fig. 2a,b). We found uniform displacements across the field of view (Extended Data Fig. 4 and Supplementary Video 1), indicating that there is minimal strain over the imaged area and that displacement over our field of view can be described as rigid translation.
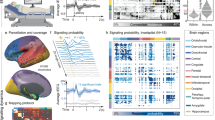
a, Rapid changes in the curvature of the fluid-filled lens move the focal point between the brain and the fluorescent microspheres adhered to the surface of the thinned skull. b, Head-fixed mouse on a treadmill. c, Representative x–z image through a typical thinned-skull window. This describes the general environment of the 134 locations used for brain and skull tracking. The GFP-expressing brain (green) and fluorescent microspheres (magenta) on the thinned skull are separated by the subarachnoid space. d, Images of the brain (green) and microspheres (magenta) during a stationary period (left) and locomotion (right). The outer bounding boxes enclose the search area for the template-matching algorithm; the inner bounding boxes represent the target used to track movement. This was done for each of the 316 recorded trials at 134 unique locations in 24 mice. There is a rostro-lateral shift of the brain during locomotion when compared to rest (visible in the displacement of the inner box) while the skull remains in the same position. e, Example of measured brain motion and treadmill velocity. Locomotion events (gray) drive rostro-lateral motion of the brain (green), whereas the skull (magenta) remains stationary.
a, Net displacement of the brain in each frame (from data in Fig. 3d) plotted as an x–y scatterplot. The displacement vector is taken to be the first principal component of the data, and the magnitude is calculated as the mean of the 80th to 100th percentile of the displacement magnitudes. b, Plot of displacement vectors for different imaging locations on the brain (N = 134 sites in 24 mice). There is a noticeable rostro-lateral brain movement trend in both hemispheres. c, Power spectrums of rostral-caudal brain motion (top) and respiration (middle), showing there is no appreciable brain motion at the respiration frequency. Plotted at the bottom is the coherence between rostral-caudal brain motion and respiration. A lack of overlap in the frequency components of the signals and a low coherence between them (confidence level = 0.319) suggest that the observed motion is not driven by respiration or heartbeat. d, Cross-correlations between the brain motion and locomotion signals from (Fig. 3d). e, Locomotion-triggered rostral-caudal and medial-lateral brain motion. Each colored line represents the locomotion-triggered average for a single trial (N = 153) and the black line is the mean with the shading showing the 90% confidence interval. The brain begins to move rostrally (R) and laterally (L) slightly before locomotion. f, Triggered averages of the cessation of locomotion. Each colored line represents the locomotion-triggered average for a single trial (N = 153) and the black line is the mean with the shading showing the 90% confidence interval. The brain moves caudally (C) and medially (M) to return to baseline following the transition from locomotion to rest.
The brain motion is primarily in the rostral direction and is linked to locomotion
To quantify patterns in the direction of motion, we imaged brain motion during locomotion from 134 sites in frontal, somatosensory and visual cortex and performed principal component analysis on the brain displacement (Fig. 2). The magnitude of each displacement vector was determined by averaging the largest 20% of the displacements from the baseline origin (Fig. 2a). We observed that the motion of the brain during locomotion was primarily in the rostral and lateral directions relative to the resting baseline position (Fig. 2b and Supplementary Videos 2–4). Brain motion amplitude was larger in male than in female mice (Supplementary Fig. 3). When we looked at the power spectrum of the motion, we observed the motion was primarily at low (<0.1 Hz) frequencies (Fig. 2c), and it was correlated strongly to locomotion in both directions (Fig. 2d and Supplementary Fig. 2a,b). Previous work has shown that heartrate-related pulsations are on the submicrometer scale3—much smaller than those evoked during locomotion. We did not observe any appreciable brain movement at respiration or heartrate frequencies (Fig. 2c) in awake mice, or aliasing in lower frequencies. We did not see any evidence of cardiac-related pulsations when imaging at higher frame rates (>100 frames per second; Extended Data Fig. 4), indicating that heartrate pulsations are minimal in awake mice, consistent with previous reports2. However, respiration-linked brain movement was detected under deep isoflurane anesthesia (Extended Data Fig. 5 and Supplementary Video 5).
The skull and brain are separated by the dura—a vascularized membrane surrounding the subarachnoid space24,25. In one instance, we were able to simultaneously record movement of dural vessels labeled with green fluorescent proteins, microspheres on the skull and the brain. This allowed us to determine whether the dura motion more closely resembled brain or skull movement during locomotion. We performed tracking on the three focal planes separately (Supplementary Video 4) and observed that the dura had similar dynamics to the skull. We generated locomotion-triggered averages of brain motion and found a close relationship between locomotion and movement of the brain (Fig. 2e), although in many cases the motion of the brain started before locomotion onset.
These results demonstrate that, in awake mice, locomotion is linked to brain motion, whereas respiration and heartrate are not substantial contributors to brain motion. However, brain motion frequently preceded the onset of locomotion, suggesting that locomotion in and of itself does not cause brain motion within the skull.
Brain motion follows abdominal muscle contractions
The brain motion we observed often slightly preceded locomotion, which indicated that a force was being applied to the brain before locomotion onset. Intracranial pressure in mice increases sharply during locomotion (from 5–10 mm Hg to >25 mm Hg)8, indicating that there are large forces at work on the brain. The increase in intracranial pressure also precedes the onset of locomotion, and this cannot be attributed to vasodilation as it lags locomotion26. Furthermore, brain motion is unlikely to be due to postural changes as these also lag locomotion onset27. We hypothesized that abdominal muscle contractions might contribute to brain motion because movements are preceded by abdominal muscle activation to stiffen the core in anticipation of body motion. We implanted electromyography (EMG) electrodes in the abdominal muscles of 24 mice while simultaneously monitoring brain movement (Fig. 3a). EMG power increased before the onset of locomotion (Fig. 3b,c), and there was a strong correlation between EMG power, which tracks muscle tension, and the motion of the brain (Fig. 3f and Supplementary Fig. 2c,d), with peaks in the cross-correlation at negative lags, indicating the EMG leads. When we aligned brain motion to the onset of locomotion and to the onset of EMG activity, we observed that the motion invariably lagged EMG activity (Fig. 3g,h and Supplementary Fig. 4), but often preceded locomotion, which suggested that abdominal muscle contraction before locomotion drove the displacement of the brain.
a, EMG electrodes were implanted in the abdominal musculature (left), which consist of three muscle layers (right): external oblique (EO), internal oblique and transverse abdominal (TrA). This was performed on all 24 mice used in these experiments, resulting in 316 recorded trials at 134 unique locations. b, Locomotion-triggered abdominal EMG power (orange) from a single representative trial (data in d). Black line: mean; shading: 90% confidence interval; N = 12 EMG events. c, The locomotion-triggered abdominal EMG averages for all trials (orange). Black line: mean; shading: 90% confidence interval; N = 108 EMG trials. The expanded view around the trigger (right) shows that the abdominal EMG increases before the onset of locomotion. d, Representative brain displacement and abdominal EMG. Note the degree of correlation between abdominal muscle contraction and motion of the brain within the skull. e, Two-dimensional histograms of abdominal EMG power and brain displacement in a single trial (data in d). f, Cross-correlation between abdominal muscle EMG power and brain position for data in d. Negative lags indicate EMG leads motion. g, EMG-triggered averages for rostral-caudal and medial-lateral brain motion. Each colored line represents the EMG-triggered average for a single trial and the black line represents the mean with a shaded 90% confidence interval; N = 108 EMG trials. The brain begins to move rostrally and laterally simultaneously with the onset of abdominal muscle activation. h, Triggered averages of the cessation of abdominal muscle activity. Each colored line represents the EMG-triggered average for a single trial and the black line represents the mean with a shaded 90% confidence interval; N = 108 EMG trials. The brain moves caudally and medially to return to baseline around the time that the abdominal muscles relax.
We then tested the relationship between brain motion and recruitment of abdominal musculature in nonlocomotor regimes. Respiration conditionally recruits abdominal musculature: although exhalation does not recruit abdominal musculature at rest, respiratory distress conditionally elicits active expiration through abdominal muscle contraction28. Under deep anesthesia, we observed active expiration as revealed by the onset of abdominal EMG power bursts locked to respiratory rhythm. These EMG bursts were also locked to brain motion (Extended Data Fig. 5b,d and Supplementary Video 5). During periods of shallow, rapid breathing, both EMG power and brain motion were reduced (Extended Data Fig. 5d). Finally, we observed instances of abdominal muscle activation and brain motion in the absence of locomotion (Extended Data Fig. 5e and Supplementary Video 3). These results are consistent with the hypothesis that abdominal muscle activation is responsible for driving brain motion across a wide variety of physiological regimes.
Vertebral venous plexus provides a hydraulic link between abdomen and CNS
How could forces generated by abdominal muscle contraction reach the brain? In humans, abdominal muscle activation drives an increase in intra-abdominal pressure (IAP)29. These increases in IAP are communicated to the brain and spine30 through the vertebral venous plexus (VVP)31—a network of valveless veins that connect the abdomen and spinal canal32,33. The VVP is thought to function like a hydraulic system that provides circulatory regulation during postural changes, in which pressure in one compartment (the abdomen) exerts pressure on another (the spinal column) through the movement of fluid (blood) from higher-pressure regions to lower-pressure regions. However, whether mice possess a VVP was unknown. We filled the vascular system of two mice (one Swiss Webster and one C57bl/6) with a radiopaque tracer, imaged them using micro computed tomography (microCT), and reconstructed the vasculature around each vertebral column (Fig. 4, Extended Data Fig. 6 and Supplementary Video 6).
a, Segmented microCT scan of a C57BL/6 mouse skeleton (gold) and vasculature (red). b, Venous connections from the caudal vena cava are shown to bifurcate before entering the lumbar vertebrae. c, Connections from the caudal vena cava inferior to the L3, L4 and L5 vertebrae penetrate the vertebrae and connect to vasculature surrounding the spinal cord. d, Veins run longitudinally along the interior of the vertebrae (left). The venous bifurcations connect the caudal vena cava and vasculature within the spine. e, Ventral view without and with vasculature shown. Small holes in the ventral surfaces of the lumbar vertebrae provide an entrance for the venous projections to connect to vasculature surrounding the dural sac within the column. f, A semi-transparent view of the vertebrae provides a complete look at the caudal vena cava, the vessels that run the length of the vertebral interior and the connections between them. g, Maximum intensity projection of the lumbar section of a mouse spine in the sagittal plane. The depth of the projection is ~1.7 mm. It is shown both raw (above) and color labeled (below). Note the vessels penetrating the ventral portion of the vertebral bones that connect vasculature within the spine to vessels in the abdominal cavity. h, Segmented microCT scan of a Swiss Webster mouse spine (gold) and vasculature (red). Vessels are present on the outer and inner surfaces of the spine as well as passing through the bone. i, A view of the ventral surface of the mouse spine with vessels (top), semi-transparent vessels (middle) and no vessels (bottom). Holes present along the ventral surface of each lumbar vertebral bone provide a means for blood to flow between the abdominal cavity and spinal interior. j, Mouse lumbar spinal section shown with full bone and vasculature (left) and also shown with semi-transparent bone, penetrating vessels and interior vessels only (right). Note the bifurcation in the ventral penetrating vessels to fill the two holes on the interior ventral surface of each vertebral bone. k, Schematic of the hypothesis that increases in IAP forces blood from the caudal vena cava to the VVP within the vertebral column. The increased blood volume in an enclosed space applies pressure to the dural sac, forcing the cranially directed CSF flow that generates brain motion.
We found the lumbar and sacral vertebrae, but not the thoracic vertebrae, had small ventral foramina that communicate with the spinal canal (Fig. 4e). These foramina were typically in pairs and located on both sides of the vertebral body, although some vertebrae possess only one. Blood vessels were observed clearly to communicate through these holes into a vascular network that lined the walls of the spinal cavity, providing a physical link between the abdominal compartment and the CNS. The diaphragm partitions the thoracic and abdominal cavities while also separating the VVP-connected lumbar and sacral vertebrae from the thoracic vertebrae that lack VVP communication pathways. This separation allows the VVP to transmit abdominal (but not thoracic) pressure changes to the CNS. In humans, intra-abdominal pressures rise drastically (~90 mm Hg) when the abdominal muscles are contracted27. A pressure increase of this magnitude will drive some of the blood in the abdomen into the spinal canal, narrowing the dural sac. This results in cranial CSF flow that raises intracranial pressure and drives brain motion (Fig. 4k).
Brain motion induced by externally applied abdominal pressure
If the mechanical coupling between the abdomen and central nervous system through the VVP drives brain motion, then we reasoned that passively applied pressures to the abdomen should drive similar brain movements. To test this idea, we constructed a pneumatic pressure cuff (Supplementary Fig. 5) to apply controlled pressure to the abdomen of lightly anesthetized (~1% isoflurane in oxygen) mice (Fig. 5). We were careful to apply the pressure only to the abdomen, not the thorax, and used video monitoring to ensure the spine was not being stretched or elevated. We observed that the brain began moving rostrally and sometimes laterally within the skull shortly following the onset of the abdominal compression (Fig. 5e and Supplementary Video 7). Furthermore, the brain began moving back to its baseline position immediately upon relief of the abdominal pressure. This suggests that abdominal pressure can alter the position of the brain within the skull rapidly and substantially.
a, A mouse was lightly anesthetized with isoflurane and wrapped with an inflatable belt around the abdomen. b, Displacement of the brain relative to the skull (green) for a single abdominal compression trial (data in c). The brain was displaced rostrally and slightly laterally. c, Displacements of the brain (green) and skull (magenta) during abdominal compressions delivered to the anesthetized mouse (below). d, Brain displacement during abdominal compression trials across the brain (109 trials spread between 36 locations in six mice, averaged by location). The motion trend is in the rostro-lateral direction, as seen with brain motion during locomotion. Generated using brainrender60. e, Abdominal compression-triggered average of brain motion for each trial in the medial-lateral (green) and rostral-caudal (blue) direction. Black line: mean; shading: 90% confidence interval; N = 109 abdominal compression trials. The brain begins moving immediately upon abdominal pressure application and continues to displace as the compression continues. Upon pressure release, the brain returns quickly to baseline. f, Abdominal compression-triggered skull motion averages for each trial in the medial-lateral (green) and rostral-caudal (blue) direction. Black line: mean; shading, 90% confidence interval; N = 109 abdominal compression trials.
Motion generates fluid flow out of the brain in simulation models with simplified brain geometry
The movement of CSF/ISF into, through and out of the brain through the glymphatic system is important for the clearance of waste13, and recent work has pointed to the mechanical forces generated by the dilation or constrictions of blood vessels in generating this fluid motion14,15,16,34. We hypothesized that the large movements that we see of the entire brain could drive fluid motion of a different sort. However, although fluid flow in the subarachnoid space and ventricles can be visualized in certain instances18,35, the rapid dynamics of any motion-driven fluid flow through the parenchyma and around the brain in the awake animal is not accessible to current imaging techniques in behaving mice. Therefore, we simulated the fluid flow produced by a squeezing action on the spinal cord using a poroelastic model of the brain and spinal cord16,36,37,38,39 (Fig. 6). We used a simplified model geometry as a ‘proof-of-principle’ that brain motion can induce the movement of fluid. This geometry is meant to represent the relevant length scales of the CNS overall and of the subarachnoid space (SAS) in particular. Our axisymmetric model of a brain with simplified geometry incorporated a rostral outflow point corresponding to the cribriform plate and a compliant vascular portion in the brain corresponding to the bridging veins40 to buffer pressure changes (Fig. 6a). As the CNS aqueducts (for example, Luschka’s) cannot be represented in an axisymmetric geometry, our model connects a single internal ventricle with the SAS through the central canal. Although Fig. 6a is not to scale, the geometry used in the simulations is based on the mouse anatomy (cf. Supplementary Table 1). Our mathematical model is provided in the Supplementary Information and is detailed in ref. 39. The model is based on mixture theory whereby several phases can coexist at a point in space. Our mixture consists of only two phases: a solid and a fluid. By ‘poroelastic domain’ we mean a deformable solid skeleton saturated by the fluid. The amount of a phase at a point is described by its volume fraction. The body’s porosity is the fluid’s volume fraction. As the deformation affects local volume changes, the porosity changes accordingly. The deformation of the solid phase and fluid flow are fully coupled: just like in a wet sponge, a change of shape in the body will cause the fluid to be redistributed within the body; vice versa, the fluid flow can affect the shape of the body. We consider a poroelastic domain labeled \({\Omega }_{{\rm{BR}}}\) (Fig. 6a) occupied by brain parenchyma saturated with ISF. The initial porosity of \({\Omega }_{{\rm{BR}}}\) was set to a uniform value of 20%. \({\Omega }_{{\rm{BR}}}\) is surrounded by a poroelastic domain with a uniform initial porosity of 80% representing the SAS, the central canal and a central ventricle. These fluid-rich compartments form a single domain we denoted by \({\Omega }_{{\rm{SAS}}}\) (Fig. 6a). The interface between \({\Omega }_{{\rm{SAS}}}\) and \({\Omega }_{{\rm{BR}}}\), denoted by \(\Gamma\), is assumed to be both permeable and coherent. The permeability of \(\Gamma\) allows the deformation to cause fluid movement from one domain to the other. The coherence of \(\Gamma\) constraints the displacement of the solid phase over \({\Omega }_{{\rm{BR}}}\cup {\Omega }_{{\rm{SAS}}}\) to be continuous across \(\Gamma\). Not only do the domains \({\Omega }_{{\rm{SAS}}}\) and \({\Omega }_{{\rm{BR}}}\) differ in their initial porosity, they also differ in their physical properties (for example, permeability and elastic shear modulus; cf. Supplementary Table 1). To study the fluid exchange between \({\Omega }_{{\rm{SAS}}}\) and \({\Omega }_{{\rm{BR}}}\), the interface \(\Gamma\) has been partitioned as indicated at the bottom of Fig. 6e. Given that the fluid phase can be convected by the solid motion and that the porosity evolves, we describe the fluid’s redistribution induced by deformation through filtration velocity, namely, the velocity of the fluid relative to the solid scaled by the (pointwise) value of the fluid’s porosity. We simulated pressure application to the distal spinal cord to mimic abdominal muscle contraction such that the model gave intracranial pressure changes and brain motion consistent with our experimental observations (Fig. 6b,c). We then used the model to see what the corresponding fluid flows (Fig. 6d,e) were in and around the brain. Deformation immediately following the squeeze redistributed the fluid initially present in \({\Omega }_{{\rm{SAS}}}\) and \({\Omega }_{{\rm{BR}}}\), with a net flow of fluid out of the brain (Fig. 6e) into the subarachnoid space. This flow is quantified by integrating the component of the filtration velocity normal to the interface \(\Gamma\) over its various parts. A positive value of the filtration volumetric flow corresponds to motion out of the brain domain into the other domain. To help localize the quantification of the filtration volumetric flow, we partitioned the interface into four parts (Fig. 6e): \({\Gamma }_{{\rm{BR}}}\), \({\Gamma }_{{\rm{V}}}\), \({\Gamma }_{{\rm{CC}}}\) and \({\Gamma }_{{\rm{SC}}}\) with areas (measured in the revolved geometry of the initial configuration and expressed in mm2) of 312.7, 3.215, 13.80 and 390.1 and \(390.1\), respectively.
The duration of the squeeze pulse is 2 s. The duration of the simulation is 10 s. The simulation is based on equations (1)—(9). The boundary conditions are described in the Supplementary Information. The parameters used in the simulation are found in Supplementary Table 1. As mentioned in the main text, the geometry we consider includes a rostral outflow point corresponding to the cribriform plate and a compliant vascular portion in the brain corresponding to the bridging veins to buffer pressure changes. These two elements are accounted for in the simulation by resistance boundary conditions (see ‘Boundary conditions’ section in the Supplementary Information). These resistances have similar mathematical expressions (Supplementary Table 1) and differ in the value of two resistance scaling parameters, denoted by \({\alpha }_{{\rm{CS}}}\) (for the central sinus) and \({\alpha }_{{\rm{out}}}\) (for the rostral outflow). Here their values are \({\alpha }_{{\rm{CS}}}={10}^{6}\) and \({\alpha }_{{\rm{out}}}={6\times 10}^{8}\). a, Initial geometry (not to scale) detailing model domains and boundaries. \({\Omega }_{{\rm{BR}}}\): brain and spinal cord domain (tan); \({\Omega }_{{\rm{SAS}}}\): CSF-filled domain (cyan); \(\Gamma\): \({\Omega }_{{\rm{BR}}}-{\Omega }_{{\rm{SAS}}}\) interface (red); \({\Gamma }_{{\rm{ext}}}\): external boundary of meningeal layer (blue); \({\Gamma }_{{\rm{SZ}}}\): squeeze zone (orange); \({\Gamma }_{{\rm{out}}}\): outlet boundary representing the cribriform plate CSF outflow pathway (green); \({\Gamma }_{{\rm{CS}}}\): central sinus boundary (purple). b, Average of pore pressure (in mm Hg) over \({\Omega }_{{\rm{BR}}}\) excluding the spinal cord over time. c, Spatial distribution of pore pressure (in mm Hg) over \({\Omega }_{{\rm{BR}}}\cup {\Omega }_{{\rm{SAS}}}\) at \(t=1\,{\rm{s}}\) during the squeeze pulse. d, Streamlines of filtration velocity \({{\boldsymbol{v}}}_{{\rm{flt}}}\) (that is, curves tangent to filtration velocity field; red lines with direction indicated by arrow tips) within \({\Omega }_{{\rm{BR}}}\) excluding the spinal cord, at \(t=1\,{\rm{s}}\) (left, middle of the squeeze pulse) and \(t=3\,{\rm{s}}\) (right, a squeeze pulse time interval from \(t=1\,{\rm{s}}\)). The streamlines overlay the color plot of the filtration velocity magnitude (in \({\rm{nm}}/{\rm{s}}\)), computed as \(\left|{v}_{{\rm{flt}}}\right|=\sqrt{{v}_{{\rm{flt}},r}^{2}+{v}_{{\rm{flt}},z}^{2}}\). The colorbar range is limited to 10 nm s−1 to emphasize the spatial variation of the field. The maximum and minimum values of the field are indicated next to the triangles at the top and bottom of the colorbar, respectively. Because the SAS is extremely thin, the streamlines in this region are not shown. e, Volumetric fluid exchange rate \({Q}_{{\rm{flt}}}\) (in nL s−1) over time across: the brain shell surface \({\Gamma }_{{\rm{BR}}}\) (blue), spinal cord surface \({\Gamma }_{{\rm{SC}}}\) (green), ventricle surface \({\Gamma }_{{\rm{V}}}\) (red) and central canal surface \({\Gamma }_{{\rm{CC}}}\) (light blue). \({Q}_{{\rm{flt}}} > 0\): fluid flow from \({\Omega }_{{\rm{BR}}}\) into \({\Omega }_{{\rm{SAS}}}\). \({Q}_{{\rm{flt}}}\) is computed as the integral of the normal component of filtration velocity over the surfaces indicated. The plot displays four lines, two that are seen easily (blue and green lines), and two that overlap and appear as horizontal lines near zero (red and light blue lines). This is due to the different orders of magnitude of \({Q}_{{\rm{flt}}}\) across the different portions of \(\Gamma\). f, Rostro-caudal (ros-cau) (blue) and medio-lateral (med-lat) (green) motion of point \({P}_{{\rm{br}}}\) on the brain surface (inset) over time caused by the squeeze pulse.
The direction of the fluid flow relative to the solid motion over the top portion of \({\Omega }_{{\rm{BR}}}\) can be deduced from the streamlines of the filtration velocity (Fig. 6d; for an animated GIF of the streamlines for the entire simulation interval, see Supplementary Video 10). The visualization of the same field over \({\Omega }_{{\rm{SAS}}}\) is challenging due to the very small thickness of this domain. Hence, we reported details about the fluid flows across specific cross-sections within both the brain and SAS domains in Extended Data Fig. 7. This brain motion induced flux across the brain surface was large, corresponding to several times the normal CSF production rate (~1 nl s−1)41 (Fig. 6e), meaning that brain-motion-induced fluxes could be the dominant driver of fluid flow in the awake brain. These flows are in the opposite direction of the glymphatic flow seen during sleep20 and consistent with experimental observations that tracers infused into the cisterna magna in awake mice do not enter into the cortex21. Our simulations showed that flows across the cranial and spinal SAS are orders of magnitude larger than those across the ventricle and central canal surfaces (Fig. 6e).
We saw similar patterns of fluid flow out of the brain when we varied the outflow resistance/bridging vein compliance within ranges that produced physiologically realistic intracranial pressure changes and brain motions, suggesting that these results hold generally (Extended Data Figs. 8 and 9). The simulations predicted rostral/medial motion at the rostral tip of the brain (Fig. 6f and Extended Data Fig. 7d). We performed imaging of brain motion dynamics in the corresponding position in the brain, the olfactory bulb and also saw rostral/medial motion (Supplementary Fig. 6 and Supplementary Video 8), indicating that our simple model geometry is capturing the fundamental aspects of brain motion. Because the solid phase in our model has an elastic response, and because the boundary conditions we adopted allow it, it is expected that the system will eventually recover its initial state once the VVP mechanical stimulus is removed. This recovery can be observed in the displacement curves in Fig. 6f. It can also be observed in the trajectories in Extended Data Figs. 7d, 8f and 9f, which also show that the recovery does not happen uniformly over the system. An additional quantity that indicates the recovery is the volume of the central sinus (Supplementary Fig. 7). The relevance of these observations is that the system’s recovery is a physiological feature evident in our empirical data. From a modeling perspective, it is therefore important for our model to allow for a recovery. At this stage of development, our model is too simple to accurately match the dynamics of the observed recovery, also because our model does not include any element of physiological active control. This said, the challenge of matching the observable dynamics offers an opportunity for model improvement and verification.
In toto, our simulations indicate that brain motion could cause fluid flows out of the brain, in the opposite direction of glymphatic flow during sleep, potentially explaining why the quiescence during sleep seems to be required to drive fluid flow through the glymphatic system21.
The parameter values used in the simulations were chosen to provide some agreement with the empirical measurements (brain surface displacement and intracranial pressure). It is important to acknowledge that there is uncertainty in the permeability of brain tissue. For example, Holter and colleagues42 report various values of brain neuropil permeability between 10.3 nm2 and 16.6 nm2. These values are two orders of magnitude smaller than some other measurements (see ref. 16) and, although they are not intended to account for the possibility of bulk flow in brain parenchyma, they do represent an important reference to consider. In previous works from our group (see ref. 16), we had used a value of nominal permeability of \(2\times {10}^{-15}\,{{\rm{m}}}^{2}\). In this paper, we used a nominal value for the brain parenchyma permeability of \(2\times {10}^{-17}\,{{\rm{m}}}^{2}\), which is in line with the values used by Holter et al.42. For the sake of comparison, we also included some results with higher permeability values, namely, \(2\times {10}^{-15}\,{{\rm{m}}}^{2}\) and \(2\times {10}^{-16}\,{{\rm{m}}}^{2}\). These results are combined with those in Fig. 6e and presented in Extended Data Fig. 10.
Discussion
Our work suggests that the brain is not isolated mechanically from the body, but rather is coupled very closely to the abdominal cavity, probably through the VVP. In humans, the VVP is thought to help buffer intracranial pressure31, but its role in rodents is puzzling as the hydrostatic pressure gradients in a mouse will be much smaller than those in a human, both overall and relative to their respective arterial pressures. This hydraulic system could generate brain motion within the skull and drive CSF flow out of the brain into the subarachnoid space. Simulations suggest that brain motion driven by this mechanical coupling pushes fluid out of the brain, potentially explaining why injected tracers do not enter into the cortex in awake animals, but do so readily during sleep21. Tension by spinal nerves43 during the motor act of locomoting are unlikely to have generated brain motion in this experiment because we observed brain motion in the absence of changes in body configuration, and we see respiration-linked brain motion when abdominal muscles are engaged during deep breathing under deep anesthesia. Our simulations with a simplified brain geometry show that brain motion of the observed magnitude and could be induced by the force exerted by the VVP on the spinal cord (Extended Data Fig. 7f).
There are several caveats to our study. One is that the mice were head-fixed, preventing the normal forces generated by head motion from acting on the brain. However, the forces created by head movement in mice are much smaller than those generated by IAP and intracranial pressure changes. Measurements in freely behaving mice show self-generated accelerations of the order of 1 g44, resulting in a force of ~4 mN (9.8 m s−2 × 0.4 g brain mass). The forces generated by a 10 mm Hg anterior-posterior pressure change8 on the ~30 mm2 coronal cross-sectional area of the mouse brain will be substantially larger than those generated by head motion, on the order of ~40 mN (1,333 N m−2 × 30 × 10−6 m2). In contrast, head motion-generated forces will be greater in humans where the brain mass is several orders of magnitude larger, though intracranial pressure changes are also greater in humans than in mice45. We only imaged the dorsal portion of the brain, and there is almost certainly movement and/or strain in subcortical areas and potentially in the region around the sagittal sinus. Because we only image a small field of view, we would not be able to detect any small strains that integrated over the entire brain could contribute to the observed motion. We estimated ~1 µm of z motion in both the brain and skull (Extended Data Fig. 2) whose most parsimonious explanation is that brain and skull are both moving together in the same direction, but there could be a z motion of up to ~2 µm of the brain relative to the skull. Finally, our model has a simplified, cylindrically symmetric anatomy in which the brain and spinal tissue is homogenous. Although the model has fluid communication between the ventricles and subarachnoid space down the central canal, it lacks the connection of the ventricles through the Foramens of Magendie and Luschka. Future work with more anatomically detailed models could potentially reveal more complicated patterns of fluid movement. Another limitation of our current model is the lack of any active physiological regulation of the CNS. As discussed earlier, our model does allow the system to recover its initial state after the VVP squeeze. However, we cannot claim that the temporal dynamics of the recovery is physiologically accurate because the dynamics of the current model is governed by physical properties such as the fluid viscosity and the elastic shear modulus of the solid phase that are fundamentally passive. We believe that this limitation is acceptable in this study because the focus of our mathematical model was an estimation of the fluid flow resulting from the fast mechanical stimulus promoted by the VVP.
Our results indicate a potential link between the brain and viscera state, mediated by abdominal pressure. This work builds on previous work on brain motion driven in humans46. In addition to brain motion driven by cardiac pulsations4, respiration47 (which causes changes in IAP due to the diaphragm pushing on the viscera), and the Valsalva maneuver48,49 (which engages the abdominal muscles), can both drive brain motion. The amplitude and nature of brain motion can be changed by pathologies such as intracranial hypertension, meningioma and Chiari malformations46,50,51,52, suggesting that brain motion could be used as a biomarker46. Obesity53 elevates IAP, which could disrupt the normal flow of blood between the abdominal cavity and spinal canal and/or lead to remodeling of the VVP. Alteration of blood flow and pressure gradients between the abdomen and spinal canal could reduce the movement of the brain and CSF circulation, contributing to the adverse effects of obesity on cognitive function54. Reduction of abdominal pressure though voiding or defecation55 may partly contribute to their impacts on cognition56,57. Mechanical coupling between the abdomen and the brain is especially interesting considering the functional mechanosensitive channels in CNS neurons58 and glia59, as the forces that cause brain motion could also activate mechanosensitive channels in the brain. In addition to interoceptive pathways in the viscera, the direct signaling through mechanical forces to the brain may play a role in communicating internal states to the brain.
The simulations also indicate the importance of accounting for the deformation of vascular compartments, such as the central sinus. This observation adds to considerations coming from existing literature on the glymphatic system, emphasizing the importance of capturing the interaction between vascular dynamics and brain motion in the understanding of brain waste clearance.
Methods
All experiments were done in accordance with National Institutes of Health (NIH) guidelines and approved by the Penn State Institutional Animal Care and Use Committee (protocol no. 201042827). We imaged 30 (15 male) Swiss Webster (Charles River, cat. no. 024CFW) mice. We chose Swiss Webster mice as the dorsal skull is substantially flatter than other mouse strains, their skull bones are fused and their larger size made it easier to implant abdominal muscle EMG electrodes. Qualitatively, the brain motion we observe is similar in magnitude to previous reports in C57Bl/6 mice2. For microCT studies we used one C57BL/6 mouse (male, 27 weeks) and one Swiss Webster mouse (female, 8 weeks). Confocal imaging of histological sections was done in the Huck Institutes’ Microscopy Core Facility (RRID: SCR_024457) with a Leica SP8 DIVE Multiphoton Microscope. No animals were excluded. No statistical methods were used to predetermine sample sizes but our sample sizes are similar to those reported in previous publications17,61. Data distribution was assumed to be normal but this was not tested formally. Data collection and analysis were not performed blind to the conditions of the experiments. Animals were not randomized.
One month before window implantation, expression of GFP across brain cells22 was induced using retroorbital injection of 10 μl AAV (Addgene, cat. no. 37825-PHPeB, 1 × 1013 vector genomes (vg) ml−1) in 90 μl H2O (Supplementary Fig. 1b). We implanted a PoRTS window, with the additional step that fluorescent microspheres were applied to the surface of the skull (Fig. 1c and Supplementary Fig. 1a). In all mice, EMG electrodes were implanted in the abdominal muscles. Mice were then habituated to head fixation over several days before imaging.
Window and abdominal EMG surgery
Mice were anesthetized with isoflurane (5% induction, 2% maintenance) in oxygen throughout the surgical procedure. The scalp was shaved, and an incision was made from just rostral of the olfactory bulbs to the neck muscles, which was opened to expose the skull. A custom 1.65-mm thick titanium head bar was adhered to the skull using cyanoacrylate glue (Vibra-Tite, cat. no. 32402) and dental cement. To assist with head bar stabilization, two small self-tapping screws (J.I. Morris, cat. no. F000CE094) were inserted in the frontal bone without penetrating the subarachnoid space and were connected to the head bar with dental cement. A PoRTS window was then created over both hemispheres23. Windows typically spanned an area from lambda to rostral of bregma and were up to 0.5-cm wide, spanning across somatosensory and visual cortex. This allowed for maximum viewable brain surface. The skull was thinned and polished, and 1-μm diameter fluorescent microspheres (Invitrogen, cat. no. T7282) were spread across the surface of the thinned-skull areas (2 µl, 910,000 particles µl−1) and allowed to dry. The beads tended to form rings, similar to the patterns produced by drying coffee62. They were then covered with cyanoacrylate glue and a 0.1-mm-thick borosilicate glass piece (Electrode Microscopy Sciences, cat. no. 72198) cut to the size of the window. The position of bregma was marked with a fluorescent marker for positional reference, allowing for anatomical mapping across mice and a method of returning to imaging locations across several recording sessions.
To implant abdominal EMG electrodes, an incision 1 cm long was made in the skin below the rib cage to expose the oblique abdominal muscle. A small guide tube was then inserted into this incision and tunneled subcutaneously it reached the open scalp. Two coated stainless steel electrode wires (A-M Systems, cat. no. 790500) were inserted through the tube until the ends were exposed though both incisions, allowing the tube to be removed while the wires remained embedded under the skin. Two gold header pins (Mill-Max Manufacturing Corporation, cat. no. 0145-0-15-15-30-27-04-0) were adhered to the head bar with cyanoacrylate glue and the exposed wires between the header and neck incision were covered with silicone to prevent damage. Each wire exiting the abdominal incision was stripped of a section of coating and threaded through the muscle ~2 mm parallel from each other to allow for a bipolar abdominal EMG recording63. A biocompatible silicone adhesive (World Precision Instruments, KWIK-SIL) was used to cover the entry and exit of the muscle by the wires for implantation stability. The incision was then closed with a series of silk sutures (Fine Science Tools, cat. no. 18020-50) and Vetbond (3M, cat. no. 1469).
Multiplane imaging
To switch the focal plane rapidly between the brain and the skull, we integrated an electrically tunable lens (ETL) (Optotune, cat. no. EL-16-40-TC-VIS-5D-C) into the laser path (Extended Data Fig. 1a). The ETL was placed adjacent to and parallel with the back aperture of the microscope objective (Nikon, cat no. CFI75 LWD 16X W) to maximize axial range, avoid vignetting64 and remove gravitational effects on the fluid-filled lens that could alter focal plane depth or cause image distortion65. An ETL controller (Gardasoft, cat. no. TR-CL180) was used to control the liquid lens curvature. Preprogrammed steps in the curvature created rapid focal plane changes that were synchronized with image acquisition using transistor-to-transistor logic (TTL) pulses from the microscope. A microcontroller board (Arduino, Arduino Uno Rev3) was programmed to pass the first TTL pulse of every rapid stack to the ETL controller, which triggered a program that changed the lens curvature at predefined intervals (Extended Data Fig. 1b). The parameters of these steps were based on the frame rate, axial depth and number of images within the stack and were chosen to ensure the transitions of the lens’ curvature were done between the last raster scans of a frame and the beginning scans of the subsequent frame. The ability to trigger each rapid image stack independently using the microscope ensured consistent synchronization of the ETL and two-photon microscope even over long periods of data collection.
ETL calibration
We calibrated the ETL-induced changes in focal plane against those induced by translating the objective along the z axis (Extended Data Fig. 1). To generate a three-dimensional structure for calibration, strands of cotton were saturated with a solution of fluorescein isothiocyanate and placed in a 1.75-mm slide cavity (Carolina Biological Supply Company, cat. no. 632255). These cotton fibers were then suspended in optical adhesive (Norland Products, cat. no. NOA 133), covered with a glass cover slip, and cured with ultraviolet light (Extended Data Fig. 1d,e). At baseline, an ETL diopter input value of 0.23 was used as baseline as this generated a working distance closest to what would occur without an ETL. The objective was then stepped physically in the axial direction for 400 μm up and down in 5-μm steps, spanning 800 μm axially. The objective was then moved to the center of the stack and the diopter values were changed from −1.27 to 1.73 in 0.1 diopter steps while the objective was stationary, averaging 100 frames at each diopter value to obtain an image stack. The spatial cross-correlation between a single frame of the diopter stack and each frame of the objective movement stack were calculated to determine the change in focus location for each diopter value. This procedure was performed at three separate locations on the suspended fluorescein isothiocyanate cotton (Extended Data Fig. 1f). We performed calibrations of the magnitude across the usable range of ETL diopter values. Although the difference in micrometers per pixel scaling relative to the baseline focal values was large across extremes in ETL-induced axial focal plane shift, the typical range used for imaging the brains of mice (<100 μm) had a negligible effect (~0.01 µm per pixel) (Extended Data Fig. 1g).
To account for distortions of the image within the focal plane, we imaged a fine mesh copper grid (SPI Supplies, cat. no. 2145C-XA) (Supplementary Fig. 8). This square grid had 1,000 lines per inch (19-µm hole width, 6-µm bar width). These values were used to determine the micrometers per pixel in the center of each hole in both the x and y direction. This allowed us to generate two three-dimensional plots of x, y and micrometers per pixel points that were then fitted with a surface plot for distance calculations. The residuals of this fit were very small (nearly all <0.025 µm per pixel, Supplementary Fig. 8), showing that we can account for nearly all the distortion.
EMG, locomotion and respiration signals
EMG signals from oblique abdominal muscles were amplified and band pass-filtered between 300 Hz and 3 kHz (World Precision Instruments, cat. no. SYS-DAM80). Thermocouple (Omega Engineering, cat. no. 5SRTC-TT-K-20-36) signals were amplified and filtered between 2 Hz and 40 Hz (Dagan Corporation, cat. no. EX4-400 Quad Differential Amplifier)11. The treadmill velocity was obtained from a rotary encoder (US Digital, cat. no. E5-720-118-NE-S-H-D-B). Analog signals were captured at 10 kHz (Sutter Instrument, MScan).
The analog signal collected from the rotary encoder on the ball treadmill was smoothed with a Gaussian window (MATLAB function: gausswin, σ = 0.98 ms). Locomotion event onset was determined using a threshold between 0.05 and 0.1 m s−1 depending on mouse activity levels. EMG signal recorded from the oblique abdominal muscles from the mouse were filtered between 300 Hz and 3,000 Hz using a fifth-order Butterworth filter (MATLAB functions: butter, zp2sos, filtfilt) before squaring and smoothing (MATLAB function: gausswin, σ = 0.98 ms) the signal to convert voltage to power. Abdominal EMG contraction onset was determined using a threshold of ~100, or one order of magnitude increase from baseline power levels. The thermocouple signal was filtered between 2 Hz and 40 Hz using a fifth-order Butterworth filter (MATLAB functions: butter, zp2sos, filtfilt) and smoothed with a Gaussian kernel (MATLAB function: gausswin, σ = 0.98 ms).
Abdominal pressure application
A custom-made pneumatically inflatable belt (Supplementary Fig. 5a) was fabricated to directly apply pressure to the abdomen of mice. It consisted of three plastic bladders that were fully wrapped around the mouse abdomen. The belt was positioned between the rib cage and the hip bones of the mouse to ensure that pressure was applied only to the abdominal compartment between the diaphragm and pelvic floor. The belt was inflated with 7 psi of pressure to apply a steady squeeze for 2 s with 30 s of rest between squeezes to allow for a return to baseline (Supplementary Fig. 5b). The abdominal compression belt was oriented so that inflating portion was underneath the mouse so that no compression or tension was imparted to the spine longitudinally, as this could affect the results by pushing or pulling on the spine itself. Mice were observed with a behavioral video camera during imaging to check for potential compression-induced body positional changes and to monitor respiration.
Motion tracking
Brain and fluorescent skull bead frames were deinterleaved. Each frame was then processed with a two-dimensional spatial median filter (3 × 3, MATLAB function: medfilt2). Occasionally, a spatial Gaussian filter (ImageJ function: Gaussian Blur) and contrast alterations (ImageJ function: Brightness/Contrast) were also applied before the median filter if the signal to noise ratio of the images resulted in poor tracking analysis.
At least three locations within the image sequence were chosen as targets for tracking. These template targets were regions of high spatial contrast (for example cell bodies) selected manually and then averaged by pixel intensity across 100 frames during a period without brain motion to reduce noise for a robust matching template. Following the target template selection, a larger rectangular region of interest enclosing the template area was selected manually (MATLAB function: getpts) to restrict the search spatially (Fig. 1d and Extended Data Fig. 2d,f).
For tracking, a MATLAB object was created (MATLAB object: vision.TemplateMatcher). A three-step search method was typically deployed at this step to increase computational speed for long image sequences. The sum of absolute differences between overlapping pixel intensities was calculated between the target and search windows, and the minimum value was chosen as the target position within the image. To monitor motion tracking, a displacement vector was then calculated that showed the motion in pixels between the current and previous image frames which was used to translate each image into a stabilized video sequence (MATLAB function: imtranslate). For visualization, a stabilized image was displayed alongside the target box displacement in the original image (MATLAB object: vision.VideoPlayer) to aid in checking manually for tracking failure.
Once the displacement in pixels was calculated for each target in a frame, the matrix of these values was searched for unique rows (MATLAB function: unique) to determine the number of unique target locations within the image. We then calculated the corresponding real distance between each unique location and the midlines of the image. A line was drawn between the image midline and the pixel location of the target. Then the calibration surface plot that depicts the calibration value in micrometers per pixel at each pixel for both x and y directions was integrated across this line (Supplementary Fig. 8c, MATLAB function: trapz) to determine the distance in micrometers from the vertical and horizontal midline of the image. The real distance traveled between sequential frames was then calculated using these references by finding the difference of the target distances from the center of each frame. Performing the unique integrations first greatly increased the speed of processing the data. Motion was averaged across targets filtered with a Savitzky–Golay filter (MATLAB function: sgolayfilt) with an order of 3 and a frame length of 13 (at a nominal frame rate of 19.78 frames s−1)66. This filter was not applied to the position data in Fig. 2c before analyzing the frequency spectrum of the brain motion and its coherence with the thermocouple signal. The s.e.m. was calculated among the targets for each frame as well as the 90% probability intervals of the t-distribution (MATLAB function: tinv). The 90% confidence interval of the average object position in x and y was then calculated using the s.e.m. and the probability intervals for the three signals at each frame (Extended Data Fig. 4b). The displacement of the fluorescent microspheres on the skull was then subtracted from the displacement of the brain to obtain a measurement of the motion of the brain relative to the skull.
Motion direction quantification
We used principal component analysis to find the primary direction of brain motion. Displacement data was first centered around the mean, then the covariance matrix of the positional data was calculated (MATLAB function: cov). The eigenvectors of this covariance matrix were then calculated (MATLAB function: eig) to determine the direction of the calculated principal components. To determine the magnitude of the vector, we took the mean of the largest 20% of the displacements from the origin (MATLAB function: maxk) (Fig. 2a). We took the top 20% rather than the amplitude of the leading eigenvalue of the principal component because the eigenvalue is sensitive to locomotion amount. The amplitude of the leading principal component increases not just with the displacement amplitude, but also with the amount of time spent locomoting, so, for the same amount of locomotion-related displacement, the leading eigenvalue will be larger if there is more locomotion. To avoid this interpretation issue with the leading eigenvalue, we averaged the top 20% of the displacements, which proved robust to locomotion amount. This was done for each of the 316 recorded trials at 134 unique locations in 24 mice, where each trial is a continuous 10-min recording. For locations with several trials, motion vectors were averaged to produce a single vector (Figs. 2b and 5d and Extended Data Fig. 3b).
MicroCT and vascular segmentation
A C57BL/6 mouse (male, 27 weeks) and Swiss Webster mouse (female, 8 weeks) were anesthetized with 5% isoflurane in oxygen and perfused a radiopaque compound (MICROFIL, MV-120) to label the vasculature. The mice were then scanned with a microCT scanner (GE v|tome|x L300) at the PSU Center for Quantitative Imaging core (RRID: SCR_026734). The C57BL/6 mouse was imaged from the nose to the base of the tail, covering 99.36 mm separated into 8,280 slices with an isotropic pixel resolution of 12 μm. The Swiss Webster mouse was imaged from the diaphragm to the caudal edge of the hip bones, covering 49.73 mm separated into 4,144 slices with an isotropic pixel resolution of 12 μm (Supplementary Video 9). Images were collected using 75 kV and 180 μΑ with aluminum filters for best contrast of tissue densities. Segmentation was done with Slicer 3D52,67. Thresholding (3D Slicer function: thresholding) was first used to isolate the bone, and all voxels above a manually chosen intensity threshold were retained. Voxels that were preserved by the threshold tool but not required for the segmentation were removed within user-defined projected volumes (3D Slicer function: scissors). The result was a high-resolution reconstruction of the skull, ribs, vertebrae, hips and other small bones along the length of the mouse that retained their inner cavities. Segmentation of the vasculature surrounding the spine and skull was more difficult than isolating the bone because of the overlap in voxel intensity between the small vessels and the surrounding bone and tissues. The contrast agent also filled other organs (for example, liver) with a similar intensity, so a simple threshold could not be used for the vasculature. We separated the vessels by using a freeform drawing tool (3D Slicer function: draw) to encapsulate the desired segmentation area for a single slice in two dimensions while ignoring unwanted similar contrast tissues. This process was repeated along the spine with a spacing of ~100 to 200 slices between labeled transverse areas. Once enough transverse freeform slices were created, they were used to create a volume by connecting the outer edges of consecutive drawn areas (3D Slicer function: fill between slices). This served as a mask that required all segmentation tools used to focus only on the voxels within the defined volume and ignore all others. The initial segmentation of the vasculature was created using a flood filling tool (3D Slicer function: flood filling). This tool labels vessels that are clearly connected within and across slices to quickly segment large branches of the network. The masking volume was used here to ignore connections to vessels or organs outside of the wanted space. The flood fill tool did not detect some connecting vessels, particularly ones located near the inner and outer surfaces of the vertebrae. In these instances, we utilized a segmentation tool that finds areas within a slice that shares the same pixel intensity around the entire edge (3D Slicer function: level tracing) to fill these gaps. In comparison to the bone, the three-dimensional reconstruction of the vessels was not smooth as they were smaller and had much more voxel intensity overlap with surrounding tissues and spaces. Thus, the segmentation was processed with a series of slight dilation operations that were followed by a matched erosion (3D Slicer function: margin). This technique of growing and shrinking the object repeatedly smoothed the surface and linked gaps between vessels. A smoothing tool was then used for final polishing of the vasculature (3D Slicer function: smoothing).
Brain motion simulations
We aimed for our calculations to serve as a proof-of-concept of the ability of induced brain motion to drive fluid flow. Thus, we selected an extremely simple geometric representation of the mouse CNS (Fig. 6). The brain and spinal cord (in tan) are surrounded by communicating fluid-filled spaces (in cyan). These consist of a central spherical ventricle internal to the brain and the SAS on the outside of both brain and spinal cord. The SAS is connected to the ventricle by a straight central canal. In the center of the brain, above the ventricle, we placed a cavity meant to model the presence of the central sinus. In addition, we placed an outlet at the top of the skull to account for the fluid leakage out of the system through structures such as the cribriform plate. The dimensions for system’s geometry in the reference (initial) state are reported in Supplementary Table 1. As in Kedarasetti et al.16, both brain and fluid-filled spaces are modeled as poroelastic domains: each consists of a deformable solid elastic skeleton through which fluid can flow. The two domains, which can exchange fluid, differ in the values of their constitutive parameters, the latter being discontinuous across the interface that separates said domains. All constitutive and model parameters adopted in our simulations are listed in Supplementary Table 1.
The governing equations have been obtained using mixture-theory16,36,68 along with Hamilton’s principle37, following the variational approach demonstrated in ref. 38. Our formulation, detailed in ref. 39, differs from that in ref. 38 in that (1) each constituent herein is assumed to be incompressible in its pure form, and (2) the test functions for the fluid velocity across the brain/SAS interface are those consistent with choosing independent pore pressure and fluid velocity fields over the brain and SAS, respectively. Hence, the overall pore pressure and fluid velocity fields can be discontinuous across the brain/SAS interface. The Hamilton’s principle approach allowed us to obtain consistent relations both in the brain and SAS interiors as well as across the brain–SAS interface. In addition, this approach yielded a corresponding weak formulation for the purpose of numerical solutions using the finite element method (FEM)69.
By \({\Omega }_{{\rm{BR}}}(t)\) we denote the domain occupied by the cerebrum and spinal cord at time \(t\). Similarly, by \({\Omega }_{{\rm{SAS}}}(t)\) we denote all fluid-filled domain, that is, the SAS in a strict sense along with the central canal and the ventricle, again at time \(t\). These domains are time dependent. We denote the interface between \({\Omega }_{{\rm{BR}}}(t)\) and \({\Omega }_{{\rm{SAS}}}(t)\) by \(\Gamma ({t})\). The unit vector \({\boldsymbol{m}}\) is taken to be normal to \(\Gamma ({t})\) pointing from \({\Omega }_{{\rm{BR}}}(t)\) into \({\Omega }_{{\rm{SAS}}}(t)\). To avoid a proliferation of symbols, we omit the notation ‘\((t)\)’ to indicate a domain at the initial time, that is, for \(t=0\), so that, for example, \({\Omega }_{{\rm{BR}}}={\Omega }_{{\rm{BR}}}\left(t\right){|t}=0\). We denote the domain occupied by the system (that is, the mixture of the fluid and solid phases) by \(\Omega \left(t\right)\). This domain is the union of \({\Omega }_{{\rm{BR}}}(t)\) and \({\Omega }_{{\rm{SAS}}}(t)\). As we have adopted a mixture theory framework, we posit the coexistence of a fluid phase and a solid phase at every point in \(\Omega (t)\). Although it is possible to conceive these phases to have different reference configurations, for convenience but without loss of generality, here we assume that the reference configurations of each phase coincide with \(\Omega\). Subscripts s and f denote quantities for the solid and fluid phases, respectively. In their pure forms, each phase is assumed incompressible with constant mass densities \({\rho }_{{\rm{s}}}^{* }\) and \({\rho }_{{\rm{f}}}^{* }\). Then, denoting the volume fractions by \({\phi }_{{\rm{s}}}\) and \({\phi }_{{\rm{f}}}\), for which we enforce the saturation condition \({\phi }_{{\rm{s}}}+{\phi }_{{\rm{f}}}=1\), the mass densities of the phases in the mixture are \({\rho }_{{\rm{s}}}={\phi }_{s}{\rho }_{s}^{* }\) and \({\rho }_{{\rm{f}}}={\phi }_{{\rm{f}}}{\rho }_{{\rm{f}}}^{* }\). The symbols \({\boldsymbol{u}}\), \({\boldsymbol{v}}\) and \({\bf{T}}\) (each with the appropriate subscript), denote the displacement, velocity and Cauchy stress fields, respectively. The quantity \({{\boldsymbol{v}}}_{{\rm{flt}}}={\phi }_{{\rm{f}}}({{\boldsymbol{v}}}_{{\rm{f}}}-{{\boldsymbol{v}}}_{{\rm{s}}})\) is the filtration velocity. The pore pressure, denoted by \(p\), serves as a multiplier enforcing the balance of mass under the constraint that each pure phase is incompressible. To enforce the jump condition of the balance of mass across \(\Gamma ({t})\), we introduce a second multiplier, denoted ℘. The notation \([\kern-2pt[ a]\kern-2pt]\) indicates the jump of \(a\) across \(\Gamma ({t})\). We choose the solid’s displacement field so that \([\kern-2pt[ {{\boldsymbol{u}}}_{{\rm{s}}}]\kern-2pt] =\boldsymbol{0}\) (that is, \({{\boldsymbol{u}}}_{{\rm{s}}}\) is globally continuous). Formally, \({{\boldsymbol{v}}}_{{\rm{f}}}\) and \(p\) need not be continuous across \(\Gamma ({t})\). Possible discontinuities in these fields have been the subject of extensive study in the literature (cf., for example, refs. 38,70,71) and there are various models to control their behavior (for example, often \({{\boldsymbol{v}}}_{{\rm{flt}}}\) and \({p}_{{\rm{f}}}\) are constrained to be continuous71). We select discontinuous functional spaces for \(p\) and \({{\boldsymbol{v}}}_{{\rm{f}}}\) and we control their behavior by building an interface dissipation term in the Rayleigh pseudopotential in our application of Hamilton’s principle (similarly to ref. 38). This dissipation can be interpreted as a penalty term for the discontinuity of the filtration velocity. Before presenting the governing equations, we introduce the following two quantities: \({k}_{{\rm{f}}}=\left(1/2\right){\rho }_{{\rm{f}}}{{\boldsymbol{v}}}_{{\rm{f}}}\cdot {{\boldsymbol{v}}}_{{\rm{f}}}\) (kinetic energy of the fluid per unit volume of the current configuration) and \(d={\rho }_{{\rm{f}}}\left({{\boldsymbol{v}}}_{{\rm{f}}}-{{\boldsymbol{v}}}_{{\rm{s}}}\right)\cdot {\boldsymbol{m}}\), which the jump condition of the balance of mass requires to be continuous across \(\Gamma ({t})\).
The strong form of the governing equations, expressed in the system’s current configuration (Eulerian or spatial form; ref. 72) are as follows:
where \({{\boldsymbol{a}}}_{{\rm{s}}}\) and \({{\boldsymbol{a}}}_{{\rm{f}}}\) are material accelerations, the superscript ± refers to limits approaching each side of the interface, \({\mu }_{s}\) is a viscosity like parameter (with dimensions of velocity per unit volume) characterizing the dissipative nature of the interface and where the terms \({{\bf{T}}}_{{\rm{s}}}\), \({{\bf{T}}}_{{\rm{f}}}\) and \({{\boldsymbol{p}}}_{{\rm{sf}}}\) are governed by the following constitutive relations
where \({\Psi }_{{\rm{s}}}\) is the strain energy of the solid phase per unit volume of its reference configuration, \({{\bf{F}}}_{{\rm{s}}}={\bf{I}}+{\nabla }_{{\rm{s}}}{{\boldsymbol{u}}}_{{\rm{s}}}\) is the deformation gradient with \({\nabla }_{{\rm{s}}}\) denoting the gradient relative to position in the solid’s reference configuration, \({{\bf{C}}}_{{\rm{s}}}={{\bf{F}}}_{{\rm{s}}}^{{\rm{T}}}{{\bf{F}}}_{{\rm{s}}}\), \({\mu }_{B}\) is the Brinkmann dynamic viscosity, \({{\bf{D}}}_{{\rm{s}}}={\left(\nabla {{\boldsymbol{v}}}_{{\rm{s}}}\right)}_{{\rm{sym}}}\), \({{\bf{D}}}_{{\rm{f}}}={\left(\nabla {{\boldsymbol{v}}}_{{\rm{f}}}\right)}_{{\rm{sym}}}\), \({\left(\nabla {\boldsymbol{v}}\right)}_{{\rm{sym}}}\) denoting the symmetric part of \(\nabla {\boldsymbol{v}}\), \({\mu }_{{\rm{f}}}\) is the traditional dynamic viscosity of the fluid phase, \({\mu }_{D}\) is the Darcy viscosity and \({\kappa }_{{\rm{s}}}\) is the solid’s permeability. For \({\Psi }_{{\rm{s}}}\) we choose a simple isochoric neo-Hookean model: \(\Psi =({\mu }_{{\rm{s}}}^{e}/2)({J}^{-2/3}{\bf{I}}:{{\bf{C}}}_{{\rm{s}}}-3)\), where \(J=\det {{\bf{F}}}_{{\rm{s}}}\) and \({\mu }_{{\rm{s}}}^{e}\) is the elastic shear modulus of the pure solid phase. It is understood that the constitutive parameters in \({\Omega }_{{\rm{BR}}}(t)\) are different from those in \({\Omega }_{{\rm{SAS}}}(t)\).
The details of the boundary conditions and of the finite element formulation are provided in the Supplementary Information. Here we limit ourselves to state that the problem is solved by using the motion of the solid as the underlying map of an otherwise Lagrangian–Eulerian formulation for which the reference configuration of the solid phase serves as the computational domain. The loading imposed on the system consists of a displacement over a portion of the dural sac of the spinal cord we denote as the squeeze zone, meant to simulate a squeezing pulse provided by the VVP. This displacement is controlled so that a prescribed nominal uniform squeezing pressure is applied to said zone. Flow resistance boundary conditions are enforced at the outlet at the top of the skull, and a resistance to deformation is also imposed on the walls of the central sinus.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All the data presented in the paper are available at https://github.com/DrewLab/Garborg_2025_Figures.
Code availability
The simulation code is available at https://github.com/DrewLab/Garborg_2025_Figures. Code used for experimental data analysis is available at https://github.com/DrewLab/Garborg_2025_Code.
References
Fee, M. S. Active stabilization of electrodes for intracellular recording in awake behaving animals. Neuron 27, 461–468 (2000).
Dombeck, D. A., Khabbaz, A. N., Collman, F., Adelman, T. L. & Tank, D. W. Imaging large-scale neural activity with cellular resolution in awake, mobile mice. Neuron 56, 43–57 (2007).
Paukert, M. & Bergles, D. E. Reduction of motion artifacts during in vivo two-photon imaging of brain through heartbeat triggered scanning. J. Physiol. 590, 2955–2963 (2012).
Poncelet, B. P., Wedeen, V. J., Weisskoff, R. M. & Cohen, M. S. Brain parenchyma motion: measurement with cine echo-planar MR imaging. Radiology 185, 645–651 (1992).
Andermann, M. L., Kerlin, A. M. & Reid, R. C. Chronic cellular imaging of mouse visual cortex during operant behavior and passive viewing. Front. Cell. Neurosci. 4, 3 (2010).
Nimmerjahn, A., Mukamel, E. A. & Schnitzer, M. J. Motor behavior activates Bergmann glial networks. Neuron 62, 400–412 (2009).
Kong, L., Little, J. P. & Cui, M. Motion quantification during multi-photon functional imaging in behaving animals. Biomed. Opt. Express 7, 3686 (2016).
Gao, Y. R. & Drew, P. J. Effects of voluntary locomotion and calcitonin gene-related peptide on the dynamics of single dural vessels in awake mice. J. Neurosci. 36, 2503–2516 (2016).
Norwood, J. N. et al. Anatomical basis and physiological role of cerebrospinal fluid transport through the murine cribriform plate. Elife 8, e44278 (2019).
Echagarruga, C. T., Gheres, K. W., Norwood, J. N. & Drew, P. J. nNOS-expressing interneurons control basal and behaviorally evoked arterial dilation in somatosensory cortex of mice. Elife 9, e60533 (2020).
Zhang, Q. et al. Cerebral oxygenation during locomotion is modulated by respiration. Nat. Commun. 10, 5515 (2019).
Blaeser, A. S. et al. Trigeminal afferents sense locomotion-related meningeal deformations. Cell Rep. 41, 111648 (2022).
Rasmussen, M. K., Mestre, H. & Nedergaard, M. Fluid transport in the brain. Physiol. Rev. 102, 1025–1151 (2022).
van Veluw, S. J. et al. Vasomotion as a driving force for paravascular clearance in the awake mouse brain. Neuron 105, 549–561 (2020).
Holstein-Ronsbo, S. et al. Glymphatic influx and clearance are accelerated by neurovascular coupling. Nat. Neurosci. 26, 1042–1053 (2023).
Kedarasetti, R. T., Drew, P. J. & Costanzo, F. Arterial vasodilation drives convective fluid flow in the brain: a poroelastic model. Fluids Barriers CNS 19, 34 (2022).
Turner, K. L., Gheres, K. W., Proctor, E. A. & Drew, P. J. Neurovascular coupling and bilateral connectivity during NREM and REM sleep. Elife 9, e62071 (2020).
Fultz, N. E. et al. Coupled electrophysiological, hemodynamic, and cerebrospinal fluid oscillations in human sleep. Science 366, 628–631 (2019).
Bojarskaite, L. et al. Sleep cycle-dependent vascular dynamics in male mice and the predicted effects on perivascular cerebrospinal fluid flow and solute transport. Nat. Commun. 14, 953 (2023).
Xie, L. et al. Sleep drives metabolite clearance from the adult brain. Science 342, 373–377 (2013).
Miyakoshi, L. M. The state of brain activity modulates cerebrospinal fluid transport. Prog. Neurobiol. 229, 102512 (2023).
Chan, K. Y. et al. Engineered AAVs for efficient noninvasive gene delivery to the central and peripheral nervous systems. Nat. Neurosci. 20, 1172–1179 (2017).
Drew, P. J. et al. Chronic optical access through a polished and reinforced thinned skull. Nat. Methods 7, 981–984 (2010).
Coles, J. A., Myburgh, E., Brewer, J. M. & McMenamin, P. G. Where are we? The anatomy of the murine cortical meninges revisited for intravital imaging, immunology, and clearance of waste from the brain. Prog. Neurobiol. 156, 107–148 (2017).
Levy, D. & Moskowitz, M. A. Meningeal mechanisms and the migraine connection. Annu. Rev. Neurosci. 46, 39–58 (2023).
Huo, B. X., Smith, J. B. & Drew, P. J. Neurovascular coupling and decoupling in the cortex during voluntary locomotion. J. Neurosci. 34, 10975–10981 (2014).
Cresswell, A. G., Grundstrom, H. & Thorstensson, A. Observations on intra-abdominal pressure and patterns of abdominal intra-muscular activity in man. Acta Physiol. Scand. 144, 409–418 (1992).
Del Negro, C. A., Funk, G. D. & Feldman, J. L. Breathing matters. Nat. Rev. Neurosci. 19, 351–367 (2018).
Grillner, S., Nilsson, J. & Thorstensson a. Intra-abdominal pressure changes during natural movements in man. Acta Physiol. Scand. 103, 275–283 (1978).
Depauw, P. et al. The significance of intra-abdominal pressure in neurosurgery and neurological diseases: a narrative review and a conceptual proposal. Acta Neurochir. (Wien) 161, 855–864 (2019).
Nathoo, N., Caris, E. C., Wiener, J. A. & Mendel, E. History of the vertebral venous plexus and the significant contributions of Breschet and Batson. Neurosurgery 69, 1007–1014 (2011).
Batson, O. V. The function of the vertebral veins and their role in the spread of metastases. Ann. Surg. 112, 138–149 (1940).
Carpenter, K. et al. Revisiting the vertebral venous plexus—a comprehensive review of the literature. World Neurosurg 145, 381–395 (2021).
Williams, S. D. et al. Neural activity induced by sensory stimulation can drive large-scale cerebrospinal fluid flow during wakefulness in humans. PLoS Biol. 21, e3002035 (2023).
Mestre, H. et al. Flow of cerebrospinal fluid is driven by arterial pulsations and is reduced in hypertension. Nat. Commun. 9, 4878 (2018).
Costanzo, F. & Miller, S. T. An arbitrary Lagrangian–Eulerian finite element formulation for a poroelasticity problem stemming from mixture theory. Comput. Meth. Appl. Mech. Eng. 323, 64–97 (2017).
Bedford, A. Hamilton’s Principle in Continuumm Mechanics (Springer, 2021).
dell’Isola, F., Madeo, A. & Seppecher, P. Boundary conditions at fluid-permeable interfaces in porous media: a variational approach. Int. J. Solids Struct. 46, 3150–3164 (2009).
Costanzo, F., Jannesari, M. & Ghitti, B. Poroelastic flow across a permeable interface: a Hamilton’s principle approach and its finite element implementation. Math. Models Methods Appl. Sci. https://doi.org/10.1142/S0218202526500259 (2026).
Auer, L. M., Ishiyama, N., Hodde, K. C., Kleinert, R. & Pucher, R. Effect of intracranial pressure on bridging veins in rats. J. Neurosurg. 67, 263–268 (1987).
Liu, G. et al. Direct measurement of cerebrospinal fluid production in mice. Cell Rep. 33, 108524 (2020).
Holter, K. E. et al. Interstitial solute transport in 3D reconstructed neuropil occurs by diffusion rather than bulk flow. Proc. Natl Acad. Sci. USA 114, 9894–9899 (2017).
Kwan, M. K., Wall, E. J., Massie, J. & Garfin, S. R. Strain, stress and stretch of peripheral nerve. Rabbit experiments in vitro and in vivo. Acta Orthop. Scand. 63, 267–272 (1992).
Meyer, A. F., Poort, J., O’Keefe, J., Sahani, M. & Linden, J. F. A head-mounted camera system integrates detailed behavioral monitoring with multichannel electrophysiology in freely moving mice. Neuron 100, 46–60 (2018).
Hamilton, W. F., Woodbury, R. A. & Harper, H. T. Arterial, cerebrospinal and venous pressures in man during cough and strain. Am. J. Physiol. 141, 42–50 (1944).
Almudayni, A. et al. Magnetic resonance imaging of the pulsing brain: a systematic review. MAGMA 36, 3–14 (2023).
Maier, S. E., Hardy, C. J. & Jolesz, F. A. Brain and cerebrospinal fluid motion: real-time quantification with M-mode MR imaging. Radiology 193, 477–483 (1994).
Ertl-Wagner, B. B., Lienemann, A., Reith, W. & Reiser, M. F. Demonstration of periventricular brain motion during a Valsalva maneuver: description of technique, evaluation in healthy volunteers and first results in hydrocephalic patients. Eur. Radiol. 11, 1998–2003 (2001).
Mousavi, S. R. et al. Measurement of in vivo cerebral volumetric strain induced by the Valsalva maneuver. J. Biomech. 47, 1652–1657 (2014).
Saindane, A. M. et al. Noninvasive assessment of intracranial pressure status in idiopathic intracranial hypertension using displacement encoding with stimulated echoes (DENSE) MRI: a prospective patient study with contemporaneous CSF pressure correlation. AJNR Am. J. Neuroradiol. 39, 311–316 (2018).
Leung, V., Magnussen, J. S., Stoodley, M. A. & Bilston, L. E. Cerebellar and hindbrain motion in Chiari malformation with and without syringomyelia. J. Neurosurg. Spine 24, 546–555 (2016).
Taoka, T. et al. Brain surface motion imaging to predict adhesions between meningiomas and the brain surface. Neuroradiology 52, 1003–1010 (2010).
Cobb, W. S. et al. Normal intraabdominal pressure in healthy adults. J. Surg. Res. 129, 231–235 (2005).
Balasubramanian, P. et al. Obesity-induced cognitive impairment in older adults: a microvascular perspective. Am. J. Physiol. Heart Circ. Physiol. 320, H740–H761 (2021).
Niwa, M., Muramatsu, K. & Sasaki, S. Discharge patterns of abdominal and pudendal nerves during induced defecation in anesthetized cats. J. Physiol. Sci. 65, 223–231 (2015).
Mamet, D. Glengarry Glen Ross (Grove Press, 1983).
Wei, C. C. et al. Defecation after magnesium supplementation enhances cognitive performance in triathletes. Sports Med. Health Sci. 7, 102–108 (2025).
Jammal Salameh, L., Bitzenhofer, S. H., Hanganu-Opatz, I. L., Dutschmann, M. & Egger, V. Blood pressure pulsations modulate central neuronal activity via mechanosensitive ion channels. Science 383, eadk8511 (2024).
Chi, S. et al. Astrocytic Piezo1-mediated mechanotransduction determines adult neurogenesis and cognitive functions. Neuron 110, 2984–2999 (2022).
Claudi, F. et al. Visualizing anatomically registered data with brainrender. Elife 10, e65751 (2021).
Winder, A. T., Echagarruga, C., Zhang, Q. & Drew, P. J. Weak correlations between hemodynamic signals and ongoing neural activity during the resting state. Nat. Neurosci. 20, 1761–1769 (2017).
Deegan, R. D. et al. Capillary flow as the cause of ring stains from dried liquid drops. Nature 389, 827–829 (1997).
Pearson, K. G., Acharya, H. & Fouad, K. A new electrode configuration for recording electromyographic activity in behaving mice. J. Neurosci. Methods 148, 36–42 (2005).
Grewe, B. F., Voigt, F. F., van ‘t Hoff, M. & Helmchen, F. Fast two-layer two-photon imaging of neuronal cell populations using an electrically tunable lens. Biomed. Opt. Express 2, 2035–2046 (2011).
Ito, K. N., Isobe, K. & Osakada, F. Fast z-focus controlling and multiplexing strategies for multiplane two-photon imaging of neural dynamics. Neurosci. Res. 179, 15–23 (2022).
Savitzky, A. & Golay, M. J. E. Smoothing and differentiation of data by simplified least squares procedures. Anal. Chem. https://doi.org/10.1021/ac60214a047 (2002).
Fedorov, A. et al. 3D Slicer as an image computing platform for the Quantitative Imaging Network. Magn. Reson. Imaging 30, 1323–1341 (2012).
Bowen, R. Theory of mixtures (AC Eringen, 1976).
Brenner, S. & Scott, L. The Mathematical Theory of Finite Element Methods (Springer-Verlag, 2008).
Hou, J. S., Holmes, M. H., Lai, W. M. & Mow, V. C. Boundary conditions at the cartilage-synovial fluid interface for joint lubrication and theoretical verifications. J. Biomech. Eng. 111, 78–87 (1989).
Shim, J. J. & Ateshian, G. A. A hybrid biphasic mixture formulation for modeling dynamics in porous deformable biological tissues. Arch. Appl. Mech. 92, 491–511 (2022).
Gurtin, M., Fried, E. & Anand, L. The Mechanics and Thermodynamics of Continua (Cambridge Univ. Press, 2010).
Acknowledgements
We thank M. Tribone for mouse illustrations, and Sutter Instruments for assistance in the design and machining the ETL mount. This work is supported by R01NS078168 and U19NS128613 to P.J.D., Pennsylvania Department of Health Award no. 4100095613 to F.C. and P.J.D., F31ES036154 to D.I.G. and AHA Career Development Award no. 935961 to Q.Z. D.I.G. was also supported by T32GM108563.
Author information
Authors and Affiliations
Contributions
P.J.D., F.C. and C.S.G. conceived and designed the study. C.S.G. performed two-photon imaging experiments and associated data analysis, and microCT reconstructions. B.G. and F.C. performed simulations. J.M.R. analyzed data. C.S.G. and N.F. developed the pneumatic squeezer. S.J.M. developed the microCT scanning methodology. Q.Z., D.I.G., K.L.T., M.M., R.T.K. and H.L. performed experiments and developed methodology. P.J.D., F.C. and C.S.G. wrote the paper with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Neuroscience thanks Jeff Tithof and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Axial calibration of electrically-tunable lens.
a. A change in the current input to the lens generates a curvature change in the lens, which alters the focus. b. Synchronization of ETL focus change with microscope scanning. A TTL pulse is generated at the beginning of each frame from the PCIe board in the computer controlling the microscope (top left). An Arduino Uno was programmed to filter all pulses besides the first of the stack (middle left). This pulse was then sent to the ETL controller to prompt a predetermined set of current steps that were sent to the ETL (bottom left). These current changes created a rapid stack with each depth captured as a single frame (right). c. The point spread function in the X (left) and Z (right) directions of the two-photon microscope created with a 0.17 µm fluorescent microsphere and a 0.8 NA N16XLWD-PF 16x Nikon objective. The ETL obscures part of the back aperture, resulting in a lower effective NA. d. Calibration of the ETL focal range. To provide a fluorescent three-dimensional structure, cotton stands were dipped in a solution of fluorescein isothiocyanate and suspended in optical adhesive within a concave slide. e. Three-dimensional segmentations created using fluorescent cotton strands from three locations (left to right). f. Calibration of the ETL diopter shifts to focal plane shifts. Three locations in the cotton (shown in e) were imaged by shifting the ETL focus and by translating the object in Z and aligned by correlational matching of images (top). These averages are plotted for each location in colored lines with the shaded standard deviation. The linear regression is also plotted as a solid blue line, with zero µm being the focus neutral diopter value (bottom). g. From top to bottom, change in X and Y scaling and laser power as a function of diopter value. Changing the diopter of the ETL had negligible changes in magnification and laser power output in the typical imaging range.
Extended Data Fig. 2 Motion tracking is robust with minimal z-axis displacement of the brain and skull during locomotion.
a. Typical tracking locations were selected for fluorescent microspheres (magenta) and GFP-expressing brain tissue (green) as well as a baseline location with no visible fluorescence (brown). b. The mean fluorescence in both locations was ~9% lower during locomotion than the mean fluorescence when the animal was still in n = 1 mouse. c. Using the axial point spread function of the two-photon microscope, a displacement in z of ~1 µm will drive a fluorescence decrease of ~9%. It is likely that the brain and skull are shifting in the same direction in z as the calculated values are so similar, possibly due to slight movements of the whole head and/or head fixation apparatus. d. Static microspheres on a slide were imaged to demonstrate error in the tracking software. e. The tracking software detected no motion from the static fluorescent microspheres. f. Static FITC-stained filter paper on a slide was imaged to demonstrate to simulate brain tissue. g. The tracking software detected no motion from the static FITC paper.
Extended Data Fig. 3 Negligible skull motion during locomotion.
a. ‘Worst-case’ skull motion in a 55 g mouse. A fluorescent marker on the skull at bregma was imaged due to its large distance from the implanted head bar (implanted caudally of lambda) to maximize the ability for the skull to displace during locomotion. b. A plot of skull displacement, calculated from the same trials as the brain motion (N = 316 trials across 134 sites in 24 mice). Note the small size and lack of clear direction. c. Locomotion-triggered average skull motion for each trial. The black line shows the mean, and the shaded portion denotes 90 percent confidence interval. N = 153 locomotion events. d. Locomotion cessation-triggered average skull motion. The black line shows the mean, and the shaded portion denotes 90 percent confidence interval. N = 153 locomotion events. e. EMG-triggered average skull motion. The black line shows the mean, and the shaded portion denotes 90 percent confidence interval. N = 108 EMG events. f. EMG cessation-triggered average skull motion. The black line shows the mean, and the shaded portion denotes 90 percent confidence interval. N = 108 EMG events.
Extended Data Fig. 4 Template-matching algorithm used to track the brain is robust across the field of view.
a. An image of the GFP-expressing parenchyma. Each of the eight bounding boxes (white) represents a tracking template area for the matching algorithm to follow. b. The targets were tracked at each of the eight locations and the mean and 90 percent confidence interval (shading) were calculated and plotted. N = 1 recording location in one mouse. The tight confidence interval bounds highlight the confidence in tracking different structures at various locations within the image as well as a lack of brain distortion within the field of view, indicating rigid translation. Frame rate was 106.89 frames/sec. Note the lack of appreciable motion at the cardiac or respiratory frequencies. c. Images of the brain (from a) when the mouse is at rest (top) and during a locomotion event (bottom). The neuron seen in the bounding box (gray) displaces rostrally and laterally during locomotion when compared to its resting position.
Extended Data Fig. 5 Respiration-driven brain motion is only observed under deep anesthesia in mice when abdominal muscles are engaged.
a. The respiration of the mouse anesthetized with isoflurane and instrumented with abdominal EMG electrode was monitored using a behavioral camera by measuring the mean pixel intensity of a box drawn across the edge of the body (box in bottom left image). Inset shows brain visualized under the two-photon microscope (bottom right). b. EMG-triggered motion during period of deep anesthesia respiration trial in the medial/lateral (top) and rostral-caudal (bottom) directions. N = 1 recording at one location in one mouse, shown in d. The black line shows the mean with a shaded 90 percent confidence interval. c. EMG-triggered skull motion. N = 1 recording at one location in one mouse, shown in d. The black line shows the mean with a shaded 90 percent confidence interval. d. Brain (green) and skull (magenta), abdominal EMG power (orange) and behavioral camera respiration (brown) during varying levels of anesthesia. Isoflurane concentration began at 0.5% at 5 seconds, then was increased to 5% for 120 seconds, after which it was reduced to 0.5% once again. Light anesthesia is characterized by shallow breaths with minimal abdominal muscle contraction and produced no detectable pattern of brain motion within the skull (lower left). Deep anesthesia, characterized by slower and deeper breaths, resulted in increased abdominal muscle activation and brain motion within the skull (lower right). e. The same location was imaged again in the same mouse on a subsequent day. Locomotion drove larger brain (green) and skull (magenta) displacements compared to respiration-induced brain motion under anesthesia (shown in d). The abdominal muscle power (orange) also shows much stronger abdominal muscle contractions during locomotion events (gold). Abdominal muscle activation without a locomotion event (lower left) still resulted in a rostro-lateral brain displacement within the skull.
Extended Data Fig. 6 MicroCT imaging of spine and associated vasculature.
a. The skeleton and spinal vasculature. b. A view of the vessels within the rib cage. The gap observed between the caudal vena cava and both cranial vena cava is occupied by the heart, which was not included. Note the lack of connections between the caudal vena cava and the vertebrae within the rib cage. c. A view of the L3, L4, and L5 vertebrae showing connections between the caudal vena cava and VVP within the vertebrae. d. The internal VVP is shown both with bone (left) and without bone (right). e. Two holes are present on the internal ventral surface of the vertebrae. These may act as pathways for veins in the abdomen to connect to the VVP within the lumbar section of the vertebral column. f. Visualization of the veins and vertebrae with a focus on the internal ventral holes in the bone. Veins occupy the holes in the vertebrae, which can be seen both when the bone is opaque (left) and semi-transparent (right). g. A view of the lumbar vertebral bones of a Swiss Webster mouse. Holes can be seen along the interior ventral surface of the vertebral bones, which are also present in the C57BL/6 mouse. h. A transverse view of the lumbar vessels both with bone (left) and without bone (right). A complex network of veins are present along the outer and inner surfaces of the vertebrae with projections through the ventral section of bone that connect the inner and outer networks.
Extended Data Fig. 7 Ancillary results to those presented in Fig. 6 from the same finite element simulation of the flow induced by a squeeze of intensity p0 = 20 mmHg applied over the SZ.
The duration of the squeeze pulse is \(2\,{\rm{s}}\). The duration of the simulation is \(10\,{\rm{s}}\). The simulation is based on Equations (1)—(9). The boundary conditions are described in the Supplementary Material. The parameters used in the simulation are found in Supplementary Table 1. As mentioned in the main text, the geometry we consider includes a rostral outflow point corresponding to the cribriform plate and a compliant vascular portion in the brain corresponding to the bridging veins to buffer pressure changes. These two elements are accounted for in the simulation by resistance boundary conditions (cf. the Boundary conditions section in the Supplementary Information). These resistances have similar mathematical expressions (cf. Supplementary Table 1) and differ in the value of two resistance scaling parameters, denoted by \({\alpha }_{{\rm{CS}}}\)(for the central sinus) and \({\alpha }_{{\rm{out}}}\) (for the rostral outflow). Here their values are \({\alpha }_{{\rm{CS}}}={10}^{6}\) and \({\alpha }_{{\rm{out}}}={6\times 10}^{8}\). a. Average of normal filtration velocity (in \({\rm{\mu }}{\rm{m}}/{\rm{s}})\) over each of the cranial and spinal SAS sections (shown in the inset) over time. The plot displays 4 lines, with the blue one appearing as horizontal line near zero –due to the different orders of magnitude of the filtration velocity across the different SAS sections. The unit normal vector to the sections points in the rostral direction. b. Average of normal filtration velocity (in \({\rm{nm}}/{\rm{s}})\) over each of the brain and spinal cord sections (shown in the inset) over time. The plot displays 3 lines, with the orange one appearing as horizontal line near zero –due to the different orders of magnitude of the filtration velocity across \({\Omega }_{{\rm{BR}}}\). The unit normal vector to the sections points in the rostral direction. c. Average of normal filtration velocity (in \({\rm{\mu }}{\rm{m}}/{\rm{s}})\) over each of the central canal sections (shown in the inset) over time. The unit normal vector to the sections points in the rostral direction. d. Trajectories of points P1–P6 (shown in the inset) on the surface of the brain: traces of the points indicated in the inset over the time interval \(0 < t < 10\,{\rm{s}}\). e. Volumetric fluid outflow \({Q}_{{\rm{out}}}\) (in \({\rm{nL}}/{\rm{s}})\) through the outlet boundary \({\Gamma }_{{\rm{out}}}\) over time. \({Q}_{{\rm{out}}} > 0\): fluid flow out of \({\Omega }_{{\rm{SAS}}}\). \({Q}_{{\rm{out}}}\) is computed as the integral of the normal component of filtration velocity over the surface indicated. f. Average force \(F\) (in \({\rm{N}}\)) exerted by CSF over time onto brain and spinal cord during the squeeze. \(F(t)\) is computed as the integral average of \(({\boldsymbol{m}}\cdot {\bf{T}}{\boldsymbol{m}})\) over the surface \({\Gamma }_{{\rm{BR}}}\cup {\Gamma }_{{\rm{SC}}}\), where \({\bf{T}}\) is the total Cauchy stress acting on the mixture in the SAS and \({\boldsymbol{m}}\) is the outward unit normal to the surface indicated.
Extended Data Fig. 8 Simulation of the flow induced by a squeeze of intensity p0 = 20 mmHg applied over the SZ identical to that reported in Fig. 6 except for the resistance parameters on the central sinus and the outflow boundary.
The duration of the squeeze pulse is \(2\,{\rm{s}}\). The duration of the simulation is \(10\,{\rm{s}}\). The simulation is based on Equations (1)—(9). The boundary conditions are described in the Supplementary Information. The parameters used in the simulation are found in Supplementary Table 1. As mentioned in the main text, the geometry we consider includes a rostral outflow point corresponding to the cribriform plate and a compliant vascular portion in the brain corresponding to the bridging veins to buffer pressure changes. These two elements are accounted for in the simulation by resistance boundary conditions (cf. the Boundary conditions section in the Supplementary Information). These resistances have similar mathematical expressions (cf. Supplementary Table 1) and differ in the value of two resistance scaling parameters, denoted by \({\alpha }_{{\rm{CS}}}\)(for the central sinus) and \({\alpha }_{{\rm{out}}}\) (for the rostral outflow). Here their values are \({\alpha }_{{\rm{CS}}}={10}^{8}\) and \({\alpha }_{{\rm{out}}}={6\times 10}^{6}\). a. Initial geometry (not to scale) detailing model domains and boundaries. \({\Omega }_{{\rm{BR}}}\): brain and spinal cord domain (tan); \({\Omega }_{{\rm{SAS}}}\): CSF-filled domain (cyan); \(\Gamma\): \({\Omega }_{{\rm{BR}}}-{\Omega }_{{\rm{SAS}}}\) interface (red); \({\Gamma }_{{\rm{ext}}}\): external boundary of meningeal layer (blue); \({\Gamma }_{{\rm{SZ}}}\): squeeze zone (orange); \({\Gamma }_{{\rm{out}}}\): outlet boundary representing the cribriform plate CSF outflow pathway (green); \({\Gamma }_{{\rm{CS}}}\): central sinus boundary (purple). b. Average of pore pressure (in \({\rm{mmHg}}\)) over \({\Omega }_{{\rm{BR}}}\) excluding the spinal cord over time. c. Spatial distribution of pore pressure (in \({\rm{mmHg}}\)) over \({\Omega }_{{\rm{BR}}}\cup {\Omega }_{{\rm{SAS}}}\) at \(t=1\,{\rm{s}}\) during the squeeze pulse. d. Streamlines of filtration velocity \({{\boldsymbol{v}}}_{{\rm{flt}}}\) (that is, curves tangent to filtration velocity field; red lines with direction indicated by arrow tips) within \({\Omega }_{{\rm{BR}}}\) excluding the spinal cord, at \(t=1\,{\rm{s}}\) (left, middle of the squeeze pulse) and \(t=3\,{\rm{s}}\) (right, a squeeze pulse time interval from \(t=1\,{\rm{s}}\)). The streamlines overlay the color plot of the filtration velocity magnitude (in \({\rm{nm}}/{\rm{s}}\)), computed as \(\left|{v}_{{\rm{flt}}}\right|=\sqrt{{v}_{{\rm{flt}},r}^{2}+{v}_{{\rm{flt}},z}^{2}}\). The colorbar range is limited to \(10\,{\rm{nm}}/{\rm{s}}\) to emphasize the spatial variation of the field. The maximum and minimum values of the field are indicated next to the triangles at the top and bottom of the colorbar, respectively. Because the SAS is extremely thin, the streamlines in this region are not shown. e. Volumetric fluid exchange rate \({Q}_{{\rm{flt}}}\) (in \({\rm{nL}}/{\rm{s}})\) over time across: the brain shell surface \({\Gamma }_{{\rm{BR}}}\) (blue), spinal cord surface \({\Gamma }_{{\rm{SC}}}\) (green), ventricle surface \({\Gamma }_{{\rm{V}}}\) (red), and central canal surface \({\Gamma }_{{\rm{CC}}}\) (light blue). \({Q}_{{\rm{flt}}} > 0\): fluid flow from \({\Omega }_{{\rm{BR}}}\) into \({\Omega }_{{\rm{SAS}}}\). \({Q}_{{\rm{flt}}}\) is computed as the integral of the normal component of filtration velocity over the surface indicated. The plot displays 4 lines, two that are easily seen (blue and green lines), and two that overlap and appear as horizontal lines near zero (red and light blue lines). This is due to the different orders of magnitude of \({Q}_{{\rm{flt}}}\) across the different portions of \(\Gamma\). f. Trajectories of points P1–P6 (shown in the inset) on the surface of the brain: traces of the points indicated in the inset over the time interval \(0 < t < 10\,{\rm{s}}\).
Extended Data Fig. 9 Repeat of the simulation presented in Extended Data Fig. 8 of a finite element simulation of the flow induced by a squeeze of intensity p0 = 20 mmHg applied over the SZ, but with a different set of resistance parameters on the central sinus and the outflow boundary.
The duration of the squeeze pulse is \(2{\rm{s}}\). The duration of the simulation is \(10{\rm{s}}\). The simulation is based on Equations (1)—(9). The boundary conditions are described in the Supplementary Information. The parameters used in the simulation are found in Supplementary Table 1. As mentioned in the main text, the geometry we consider includes a rostral outflow point corresponding to the cribriform plate and a compliant vascular portion in the brain corresponding to the bridging veins to buffer pressure changes. These two elements are accounted for in the simulation by resistance boundary conditions (cf. the Boundary conditions section in the Supplementary Information). These resistances have similar mathematical expressions (cf. Supplementary Table 1) and differ in the value of two resistance scaling parameters, denoted by \({\alpha }_{\mathrm{CS}}\)(for the central sinus) and \({\alpha }_{{out}}\) (for the rostral outflow). Here their values are \({\alpha }_{{\rm{CS}}}={10}^{8}\) and \({\alpha }_{{\rm{out}}}={6\times 10}^{8}\). a. Initial geometry (not to scale) detailing model domains and boundaries. \({\Omega }_{{\rm{BR}}}\): brain and spinal cord domain (tan); \({\Omega }_{{\rm{SAS}}}\): CSF-filled domain (cyan); \(\Gamma\): \({\Omega }_{{\rm{BR}}}-{\Omega }_{{\rm{SAS}}}\) interface (red); \({\Gamma }_{{\rm{ext}}}\): external boundary of meningeal layer (blue); \({\Gamma }_{{\rm{SZ}}}\): squeeze zone (orange); \({\Gamma }_{{\rm{out}}}\): outlet boundary representing the cribriform plate CSF outflow pathway (green); \({\Gamma }_{{\rm{CS}}}\): central sinus boundary (purple). b. Average of pore pressure (in \({\rm{mmHg}}\)) over \({\Omega }_{{\rm{BR}}}\) excluding the spinal cord over time. c. Spatial distribution of pore pressure (in \({\rm{mmHg}}\)) over \({\Omega }_{{\rm{BR}}}\cup {\Omega }_{{\rm{SAS}}}\) at \(t=1\,{\rm{s}}\) during the squeeze pulse. d. Streamlines of filtration velocity \({{\boldsymbol{v}}}_{{\rm{flt}}}\) (that is, curves tangent to filtration velocity field; red lines with direction indicated by arrow tips) within \({\Omega }_{{\rm{BR}}}\) excluding the spinal cord, at \(t=1\,{\rm{s}}\) (left, middle of the squeeze pulse) and \(t=3\,{\rm{s}}\) (right, a squeeze pulse time interval from \(t=1\,{\rm{s}}\)). The streamlines overlay the color plot of the filtration velocity magnitude (in \({\rm{nm}}/{\rm{s}}\)), computed as \(\left|{v}_{{\rm{flt}}}\right|=\sqrt{{v}_{{\rm{flt}},r}^{2}+{v}_{{\rm{flt}},z}^{2}}\). The colorbar range is limited to \(10{\rm{nm}}/{\rm{s}}\) to emphasize the spatial variation of the field. The maximum and minimum values of the field are indicated next to the triangles at the top and bottom of the colorbar, respectively. Because the SAS is extremely thin, the streamlines in this region are not shown. e. Volumetric fluid exchange rate \({Q}_{{\rm{flt}}}\) (in \({\rm{nL}}/{\rm{s}})\) over time across: the brain shell surface \({\Gamma }_{{\rm{BR}}}\) (blue), spinal cord surface \({\Gamma }_{{\rm{SC}}}\) (green), ventricle surface \({\Gamma }_{{\rm{V}}}\) (red), and central canal surface \({\Gamma }_{{\rm{CC}}}\) (light blue). \({Q}_{{\rm{flt}}} > 0\): fluid flow from \({\Omega }_{{\rm{BR}}}\) into \({\Omega }_{{\rm{SAS}}}\). \({Q}_{{\rm{flt}}}\) is computed as the integral of the normal component of filtration velocity over the surface indicated. The plot displays 4 lines, two that are easily seen (blue and green lines), and two that overlap and appear as horizontal lines near zero (red and light blue lines). This is due to the different orders of magnitude of \({Q}_{{\rm{flt}}}\) across the different portions of \(\Gamma\). f. Trajectories of points P1–P6 (shown in the inset) on the surface of the brain: traces of the points indicated in the inset over the time interval \(0 < t < 10\,{\rm{s}}\).
Extended Data Fig. 10 Exchange flow rates vary with brain permeability.
a. Exchange flow rates across the spinal cord \({\Gamma }_{{\rm{SC}}}\) for different values of brain permeability. b. Exchange flow rates across the brain interface \({\Gamma }_{{\rm{BR}}}\) for different values of brain permeability. Except for the values of the brain permeability, all other conditions used in this simulation are identical to those used in the simulation reported in Fig. 6. In fact, the values reported here include the results presented in Fig. 6e to facilitate comparison. As a reminder, these results pertain to a simulation of the flow induced by a \(2\,{\rm{s}}\) pressure pulse \({p}_{0}=20\,{\rm{mmHg}}\) applied over \({\Gamma }_{{\rm{SZ}}}\). The nonlinear character of the model and the dependence of the volume fractions of the system’s constituents on the deformation contribute to the fact that the exchange flow rates are not directly proportional to the value of the permeability.
Supplementary information
Supplementary Information (download PDF )
Supplementary Tables 1 and 2, Text—finite element formulation and boundary conditions, Figs. 1–12 and References.
Supplementary Video 1 (download MP4 )
Brain motion is rigid. A single-plane video of a mouse brain through a cranial window was tracked in eight locations. The high degree of similarity between the calculated movement at each location demonstrates the robustness of the template-matching tracking program, the accuracy of the two-dimensional distortion calibration, and a rigid shift of the parenchyma within the imaging plane. These data can be found plotted in Supplementary Fig. 6 with the 90% confidence interval surrounding the mean position of the eight locations.
Supplementary Video 2 (download MP4 )
Relationship of brain motion to abdominal muscle EMG activity, locomotion and respiration. The brain moves rostro-laterally in response to abdominal muscle contractions before and during locomotion events. Respiration does not drive brain motion during the resting phase in the awake state. Two and three-dimensional figures are included to demonstrate the cranial window environment used to capture the brain and skull motion in vivo.
Supplementary Video 3 (download MP4 )
Brain motion without locomotion (hunching). Before locomotion, the mouse exhibits a hunching behavior that changes its posture and invokes abdominal muscle contraction. This results in rostro-lateral motion of the brain without the presence of locomotion activity. Shortly after, the mouse begins a locomotion event that shows a higher degree of abdominal muscle contraction and increased rostro-lateral brain motion. These data show that although locomotion events can predict motion of the brain within the skull, it is not required for brain motion.
Supplementary Video 4 (download MP4 )
Simultaneous skull, dura and brain tracking. The ETL was programmed for simultaneous capture of three layers to track skull, dura and brain motion. As seen in the three-dimensional reconstruction of the cranial window, the dural vessel (white) is much closer to the parenchyma surface (green) than the fluorescent microspheres on the window (magenta) as it resides on the internal surface of the skull. The data demonstrate that, when the brain moves, the dura remains stationary with the skull despite the small size of the subarachnoid space.
Supplementary Video 5 (download MP4 )
Respiration-linked brain motion during anesthesia. Respiration-driven brain motion was not observed during the awake and behaving state in mice. However, brain motion was detected occasionally during periods of deep anesthesia. In this example, the mouse exhibits very little brain motion when anesthetized with 1% isoflurane in oxygen in the first 20 s of data collection. The isoflurane was then increased to 5% in oxygen to generate deeper and slower respiration. In this state, the abdominal muscles are recruited more strongly in each breath and the brain exhibits a rostro-lateral shift within the skull. Reducing the isoflurane to 1% in oxygen at the end of the dataset resulted in reduced abdominal muscle contraction force and less brain motion. These results suggest that brain motion can be driven by respiration in mice only when the abdominal muscle contractions associated with each breath generate sufficient pressure changes within the abdomen.
Supplementary Video 6 (download MP4 )
microCT of vertebrae and vertebral venous plexus. This three-dimensional segmentation of a mouse microCT shows how vasculature inside and outside of the vertebral bones are oriented. Furthermore, it demonstrates how the vessels connect through the ventral surface of the individual vertebrae. The bone transitions between opaque and transparent to display the entirety of the vertebral venous plexus.
Supplementary Video 7 (download MP4 )
Brain motion induced by abdominal compression. A pressure cuff wrapped around the abdomen of a lightly anesthetized mouse was used to induce an increase in intra-abdominal pressure for 2 s. These pressure increases resulted in rostro-lateral motion of the brain in the skull for the duration of the compression and a return to baseline position following pressure release. These results suggest that externally applied intra-abdominal pressure changes can drive brain motion when controlling for behavior in a mouse model.
Supplementary Video 8 (download MP4 )
Olfactory bulb motion. The olfactory bulb moves rostrally within its skull compartment during locomotion, similar to the cortex. However, the olfactory bulbs shift medially, in contrast to the lateral motion seen in the cortex. Following locomotion, the olfactory bulb begins to move laterally to return to its baseline position but also overshoots its resting position caudally before slowly returning rostrally. This movement pattern is unique to the olfactory bulbs and suggests a difference in brain motion mechanics between the olfactory bulbs and the cortical hemispheres.
Supplementary Video 9 (download MP4 )
Sagittal sweep of lumbar spine. A sweep with a depth of ~1.7 mm through the spine of a Swiss Webster mouse in the sagittal plane of a microCT image stack. Vessels (bright white) can be seen both in networks within the vertebral bones (gray) and passing through the bones to connect to external vessels.
Supplementary Video 10 (download GIF )
Filtration velocity streamlines over \({\Omega }_{{\bf{BR}}}\). In relation to the results presented in Fig. 6, this is a GIF sequence of the streamlines of the filtration velocity field over the domain \({\Omega }_{{\rm{BR}}}\). The streamlines are plotted as red lines and the flow direction is indicated by arrow tips. The streamlines are plotted over a background whose color represents the magnitude of the filtration velocity. Images were output every 0.1 s. The entire 10 s of simulation is represented for a total of 101 frames. The color scale is limited to represent magnitudes ranging from 0 to 10 nm s−1. However, the minimum and maximum values of the magnitude of the filtration velocity are indicated for reference.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Garborg, C.S., Ghitti, B., Zhang, Q. et al. Brain motion is driven by mechanical coupling with the abdomen. Nat Neurosci (2026). https://doi.org/10.1038/s41593-026-02279-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41593-026-02279-z