Abstract
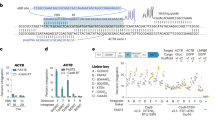
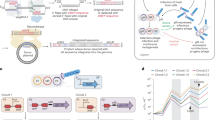
Programmable gene integration technologies are an emerging modality with exciting applications in both basic research and therapeutic development. Programmable addition via site-specific targeting elements (PASTE) is a programmable gene integration approach for precise and efficient programmable integration of large DNA sequences into the genome. PASTE offers improved editing efficiency, purity and programmability compared with previous methods for long insertions into the mammalian genome. By combining the specificity and cargo size capabilities of site-specific integrases with the programmability of prime editing, PASTE can precisely insert cargoes of at least 36 kb with efficiencies of up to 60%. Here we outline best practices for design, execution and analysis of PASTE experiments, with protocols for integration of EGFP at the human NOLC1 and ACTB genomic loci and for readout by next generation sequencing and droplet digital PCR. We provide guidelines for designing and optimizing a custom PASTE experiment for integration of desired payloads at alternative genomic loci, as well as example applications for in-frame protein tagging and multiplexed insertions. To facilitate experimental setup, we include the necessary sequences and plasmids for the delivery of PASTE components to cells via plasmid transfection or in vitro transcribed RNA. Most experiments in this protocol can be performed in as little as 2 weeks, allowing for precise and versatile programmable gene insertion.
Key points
-
Programmable addition via site-specific targeting elements (PASTE) combines the specificity, efficiency and cargo size advantages of site-specific integrases with the programmability of prime editing for precise and efficient integration of large DNA sequences into mammalian genomes.
-
PASTE offers improved editing efficiency, purity and reprogrammability compared with previous methods for long insertions into the mammalian genome.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Sequencing data used in Fig. 6 are deposited at the NCBI Sequence Read Archive (SRA) database under accession PRJNA1101023.
References
Sheridan, C. The world’s first CRISPR therapy is approved: who will receive it? Nat. Biotechnol. 42, 3–4 (2024).
Veit, G. et al. From CFTR biology toward combinatorial pharmacotherapy: expanded classification of cystic fibrosis mutations. Mol. Biol. Cell 27, 424–433 (2016).
Ausländer, S. & Fussenegger, M. Engineering gene circuits for mammalian cell-based applications. Cold Spring Harb. Perspect. Biol. 8, a023895 (2016).
Tou, C. J. & Kleinstiver, B. P. Recent advances in double-strand break-free kilobase-scale genome editing technologies. Biochemistry 62, 3493–3499 (2023).
Suzuki, K. et al. In vivo genome editing via CRISPR–Cas9-mediated homology-independent targeted integration. Nature 540, 144–149 (2016).
Nami, F. et al. Strategies for in vivo genome editing in nondividing cells. Trends Biotechnol. 36, 770–786 (2018).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Cong, L. et al. Multiplex genome engineering using CRISPR–Cas systems. Science 339, 819–823 (2013).
Hsu, P. D., Lander, E. S. & Zhang, F. Development and applications of CRISPR–Cas9 for genome engineering. Cell 157, 1262–1278 (2014).
Wright, A. V., Nuñez, J. K. & Doudna, J. A. Biology and applications of CRISPR systems: harnessing nature’s toolbox for genome engineering. Cell 164, 29–44 (2016).
Chapman, J. R., Taylor, M. R. G. & Boulton, S. J. Playing the end game: DNA double-strand break repair pathway choice. Mol. Cell 47, 497–510 (2012).
Geisinger, J. M. & Stearns, T. CRISPR–Cas9 treatment causes extended TP53-dependent cell cycle arrest in human cells. Nucleic Acids Res. 48, 9067–9081 (2020).
Wang, H. et al. Development of a self-restricting CRISPR–Cas9 system to reduce off-target effects. Mol. Ther. Methods Clin. Dev. 18, 390–401 (2020).
Ihry, R. J. et al. p53 inhibits CRISPR–Cas9 engineering in human pluripotent stem cells. Nat. Med. 24, 939–946 (2018).
Anzalone, A. V. et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576, 149–157 (2019).
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016).
Gaudelli, N. M. et al. Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage. Nature 551, 464–471 (2017).
Kurt, I. C. et al. CRISPR C-to-G base editors for inducing targeted DNA transversions in human cells. Nat. Biotechnol. 39, 41–46 (2021).
Ferreira da Silva, J. et al. Prime editing efficiency and fidelity are enhanced in the absence of mismatch repair. Nat. Commun. 13, 760 (2022).
Chen, P. J. et al. Enhanced prime editing systems by manipulating cellular determinants of editing outcomes. Cell 184, 5635–5652.e29 (2021).
Yan, J. et al. Improving prime editing with an endogenous small RNA-binding protein. Nature https://doi.org/10.1038/s41586-024-07259-6 (2024).
Nelson, J. W. et al. Engineered pegRNAs improve prime editing efficiency. Nat. Biotechnol. 40, 402–410 (2022).
Anzalone, A. V. et al. Programmable deletion, replacement, integration and inversion of large DNA sequences with twin prime editing. Nat. Biotechnol. 40, 731–740 (2022).
Wang, J. et al. Efficient targeted insertion of large DNA fragments without DNA donors. Nat. Methods 19, 331–340 (2022).
Zheng, C. et al. Template-jumping prime editing enables large insertion and exon rewriting in vivo. Nat. Commun. 14, 3369 (2023).
Jiang, T., Zhang, X.-O., Weng, Z. & Xue, W. Deletion and replacement of long genomic sequences using prime editing. Nat. Biotechnol. 40, 227–234 (2022).
Yarnall, M. T. N. et al. Drag-and-drop genome insertion of large sequences without double-strand DNA cleavage using CRISPR-directed integrases. Nat. Biotechnol. https://doi.org/10.1038/s41587-022-01527-4 (2022).
Smith, M. C. M., Brown, W. R. A., McEwan, A. R. & Rowley, P. A. Site-specific recombination by phiC31 integrase and other large serine recombinases. Biochem. Soc. Trans. 38, 388–394 (2010).
Doman, J. L. et al. Phage-assisted evolution and protein engineering yield compact, efficient prime editors. Cell 186, 3983–4002.e26 (2023).
Merrick, C. A., Zhao, J. & Rosser, S. J. Serine integrases: advancing synthetic biology. ACS Synth. Biol. 7, 299–310 (2018).
Meinke, G., Bohm, A., Hauber, J., Pisabarro, M. T. & Buchholz, F. Cre recombinase and other tyrosine recombinases. Chem. Rev. 116, 12785–12820 (2016).
Smith, M. C. M. Phage-encoded serine integrases and other large serine recombinases. Microbiol. Spectr. https://doi.org/10.1128/microbiolspec.MDNA3-0059-2014 (2015).
Choi, J. et al. Precise genomic deletions using paired prime editing. Nat. Biotechnol. 40, 218–226 (2022).
Zhuang, Y. et al. Increasing the efficiency and precision of prime editing with guide RNA pairs. Nat. Chem. Biol. 18, 29–37 (2022).
Tao, R. et al. Bi-PE: bi-directional priming improves CRISPR–Cas9 prime editing in mammalian cells. Nucleic Acids Res. 50, 6423–6434 (2022).
Strecker, J. et al. RNA-guided DNA insertion with CRISPR-associated transposases. Science https://doi.org/10.1126/science.aax9181 (2019).
Klompe, S. E., Vo, P. L. H., Halpin-Healy, T. S. & Sternberg, S. H. Transposon-encoded CRISPR–Cas systems direct RNA-guided DNA integration. Nature 571, 219–225 (2019).
Tou, C. J., Orr, B. & Kleinstiver, B. P. Precise cut-and-paste DNA insertion using engineered type V-K CRISPR-associated transposases. Nat. Biotechnol. 41, 968–979 (2023).
Lampe, G. D. et al. Targeted DNA integration in human cells without double-strand breaks using CRISPR-associated transposases. Nat. Biotechnol. 42, 87–98 (2024).
Durrant, M. G. et al. Systematic discovery of recombinases for efficient integration of large DNA sequences into the human genome. Nat. Biotechnol. 41, 488–499 (2023).
Ghosh, P., Kim, A. I. & Hatfull, G. F. The orientation of mycobacteriophage Bxb1 integration is solely dependent on the central dinucleotide of attP and attB. Mol. Cell 12, 1101–1111 (2003).
Bednarski, C., Tomczak, K., Vom Hövel, B., Weber, W.-M. & Cathomen, T. Targeted integration of a super-exon into the CFTR locus leads to functional correction of a cystic fibrosis cell line model. PLoS ONE 11, e0161072 (2016).
Sun, C. et al. Precise integration of large DNA sequences in plant genomes using PrimeRoot editors. Nat. Biotechnol. 42, 316–327 (2024).
Kim, H. K. et al. Predicting the efficiency of prime editing guide RNAs in human cells. Nat. Biotechnol. 39, 198–206 (2021).
Koeppel, J. et al. Prediction of prime editing insertion efficiencies using sequence features and DNA repair determinants. Nat. Biotechnol. 41, 1446–1456 (2023).
Ferrari, S. et al. Choice of template delivery mitigates the genotoxic risk and adverse impact of editing in human hematopoietic stem cells. Cell Stem Cell 29, 1428–1444.e9 (2022).
Hsu, J. Y. et al. PrimeDesign software for rapid and simplified design of prime editing guide RNAs. Nat. Commun. 12, 1034 (2021).
Hwang, G.-H. et al. PE-Designer and PE-Analyzer: web-based design and analysis tools for CRISPR prime editing. Nucleic Acids Res. 49, W499–W504 (2021).
Anderson, M. V., Haldrup, J., Thomsen, E. A., Wolff, J. H. & Mikkelsen, J. G. pegIT—a web-based design tool for prime editing. Nucleic Acids Res. 49, W505–W509 (2021).
Chow, R. D., Chen, J. S., Shen, J. & Chen, S. A web tool for the design of prime-editing guide RNAs. Nat. Biomed. Eng. 5, 190–194 (2021).
Doman, J. L., Sousa, A. A., Randolph, P. B., Chen, P. J. & Liu, D. R. Designing and executing prime editing experiments in mammalian cells. Nat. Protoc. 17, 2431–2468 (2022).
Yarnall, M. T. N. et al. Drag-and-drop genome insertion of large sequences without double-strand DNA cleavage using CRISPR-directed integrases. Nat. Biotechnol. 41, 500–512 (2023).
Park, S.-J. et al. Targeted mutagenesis in mouse cells and embryos using an enhanced prime editor. Genome Biol. 22, 170 (2021).
Oscorbin, I. P., Wong, P. F., Boyarskikh, U. A., Khrapov, E. A. & Filipenko, M. L. The attachment of a DNA-binding Sso7d-like protein improves processivity and resistance to inhibitors of M-MuLV reverse transcriptase. FEBS Lett. 594, 4338–4356 (2020).
Segura, M. M., Alba, R., Bosch, A. & Chillón, M. Advances in helper-dependent adenoviral vector research. Curr. Gene Ther. 8, 222–235 (2008).
Brunetti-Pierri, N. & Ng, P. Progress towards liver and lung-directed gene therapy with helper-dependent adenoviral vectors. Curr. Gene Ther. 9, 329–340 (2009).
Kay, M. A., He, C.-Y. & Chen, Z.-Y. A robust system for production of minicircle DNA vectors. Nat. Biotechnol. 28, 1287–1289 (2010).
Hendel, A. et al. Chemically modified guide RNAs enhance CRISPR-Cas genome editing in human primary cells. Nat. Biotechnol. 33, 985–989 (2015).
Hu, J. et al. Detecting DNA double-stranded breaks in mammalian genomes by linear amplification-mediated high-throughput genome-wide translocation sequencing. Nat. Protoc. 11, 853–871 (2016).
Giannoukos, G. et al. UDiTaS, a genome editing detection method for indels and genome rearrangements. BMC Genomics 19, 212 (2018).
Tsai, S. Q. et al. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR–Cas nucleases. Nat. Biotechnol. 33, 187–197 (2015).
Lazzarotto, C. R. et al. CHANGE-seq reveals genetic and epigenetic effects on CRISPR–Cas9 genome-wide activity. Nat. Biotechnol. 38, 1317–1327 (2020).
Clement, K. et al. CRISPResso2 provides accurate and rapid genome editing sequence analysis. Nat. Biotechnol. 37, 224–226 (2019).
Luo, J. et al. A protocol for rapid generation of recombinant adenoviruses using the AdEasy system. Nat. Protoc. 2, 1236–1247 (2007).
Suleman, S. et al. Rapid and inexpensive purification of adenovirus vectors using an optimised aqueous two-phase technology. J. Virol. Methods 299, 114305 (2022).
Acknowledgements
C.W.F. is supported by a grant from the Simons Foundation International to the Simons Center for the Social Brain at MIT. C.S.-U. is supported by a Friends of the McGovern fellowship. J.S.G. and O.O.A. are supported by NIH grants 1R21-AI149694, R01-EB031957, 1R01GM148745, R56-HG011857 and R01AG074932; The McGovern Institute Neurotechnology program; the K. Lisa Yang and Hock E. Tan Center for Molecular Therapeutics in Neuroscience; Impetus Grants; the Cystic Fibrosis Foundation Pioneer Grant; Google Ventures; Pivotal Life Sciences; MGB Gene and Cell Therapy Institute; the Yosemite Fund; Harvey Family Foundation; Termeer Foundation; and Winston Fu. We thank the members of the Abudayyeh-Gootenberg labs for support and advice.
Author information
Authors and Affiliations
Contributions
C.W.F. and C.S.-U. equally contributed to writing the introduction and protocol and generating all figures and performing experiments. D.V.T. performed experiments and assisted with protocol writing. O.O.A. and J.S.G supervised research and contributed to writing the manuscript and drafting of the figures. All authors edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
C.W.F., C.S.-U., J.S.G. and O.O.A. are inventors on patent applications related to CRISPR technologies. O.O.A. and J.S.G. are co-founders of Sherlock Biosciences, Doppler Biosciences, Circle Labs and Tome Biosciences.
Peer review
Peer review information
Nature Protocols thanks Rasmus Bak, Shahid Mansoor, Yiping Qi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Key reference
Yarnall, M. T. N. et al. Nat. Biotechnol. 41, 500–512 (2023): https://doi.org/10.1038/s41587-022-01527-4
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Fell, C.W., Schmitt-Ulms, C., Tagliaferri, D.V. et al. Precise kilobase-scale genomic insertions in mammalian cells using PASTE. Nat Protoc 20, 1546–1583 (2025). https://doi.org/10.1038/s41596-024-01090-z
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41596-024-01090-z
This article is cited by
-
Integration of large genetic payloads using prime editing and site-specific integrases
Nature Protocols (2025)