Abstract
This longitudinal Mobile Brain-Body Imaging dataset was acquired during six rehearsal sessions and three public performances of a scene from a play with highly emotional components. Three student actor dyads (N=6), one theatre director (N=1) and three audience members (N=3) participated in this study. The MoBI data recorded includes mobile electroencephalography, electrooculography, blood volume pulse, heart rate, body temperature, electrodermal activity, triaxial arm and head acceleration. The visual data includes five streams of video. This article describes the experimental setup, the multi-modal data streams acquired using a hyperscanning methodology, and an assessment of the data quality.
Similar content being viewed by others
Background & Summary
We present a longitudinal Mobile Brain-Body Imaging (MoBI) dataset consisting of six theatre rehearsals and three theatre performances containing simultaneously recorded physiological and visual data using a hyperscanning approach based on hardware triggers. The dataset has been acquired over the course of a week in which a theatre director staged a theatre scene with three dyads of undergraduate acting students. In two rehearsal sessions, the director deployed a predefined sequence of rehearsal methods identical for each dyad. For all participants, we recorded electroencephalography (EEG), electrooculography (EOG), blood volume pulse (BVP), heart rate, body temperature, electrodermal activity (EDA), triaxial arm and head acceleration, and five streams of video. Additionally, on the day of the performance, three audience members observing the scene had their EEG and video recorded.
Acting has been an emerging area of interest for neuroscience in recent years, as techniques such as functional Near-Infrared Spectroscopy (fNIRS) have been implemented in order to analyze neural, physiological, and behavioral signatures of pairs of actors1. Nevertheless, to our knowledge, this is the first publicly available longitudinal dataset combining MoBI and visual data of theatre student actors during rehearsals and public performances.
It will allow researchers to quantify the synchronization of physiological activity patterns within and across all participating individuals. Furthermore, it will permit the execution of a range of functional connectivity analyses and brain mapping techniques using electroencephalographic (EEG) data. These methodologies can then be integrated with corresponding visual data to enhance the understanding of the neural correlates of theatre acting2. Previous studies have examined acting processes related to cognition and text memorization using fMRI3, a method that investigates actors in an artificial laboratory setting and prevents any physical movement or interaction with other actors. With MoBI technology, we have the opportunity to study brain activity in action and in context, associated with behavioral data in real-world settings.
Methods
Participants
Ten healthy individuals (six males, four females; ages 21-47 years old) with no history of neurological disorder participated in this study. For data collection purposes in the experiment, one individual participated as director, six individuals participated as student actors (three male, three female), and three as audience members. All participants provided written informed consent for their participation in the study, including explicit permission for the open publication of their identities, video footage, and associated data on a credible public data repository. The experimental protocol, informed consent form, and image release documentation were reviewed and signed by each participant. Additionally, all members of the recording crew granted permission for the use of the captured video material. All published data have been de-identified where applicable, except in cases where participants consented to identifiable publication. The protocol forms were approved by the Institutional Review Board of the University of Houston (UH IRB #14415-01: NCS-FO: Assaying neural individuality and variation in freely behaving people based on qEEG). All procedures were performed in accordance with the 45 Code of Federal Regulations (CFR) part 46 (“The Common Rule”), specifically addressing the protection of human study subjects as promulgated by the U.S. Department of Health and Human Services (DHHS).
Instrumentation and Data Collection
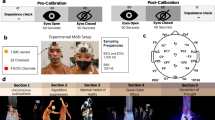
The actors, the director, and the audience members were instrumented with a 32-channel mobile EEG cap (BrainAmpDC with actiCAP and MOVE, Brain Products GmbH, Germany) sampled at 500 Hz. Four electrodes from the EEG cap were used for EOG recording. The actors were fitted with E4 Empatica wristband (Empatica Inc.) used to measure BVP sampled at 64 Hz, heart rate calculated from the BVP at 1 Hz, temperature sampled at 4 Hz, EDA sampled at 4 Hz, and triaxial arm acceleration sampled at 32 Hz4. Head movement was recorded by attaching an Inertial Measurement Unit (IMU) device on each actors’ headset. These units used wireless Magnetic, Angular Rate, and Gravity (MARG) sensors (OPAL, APDM Inc., Portland, OR) with a sampling rate of 128 Hz. Each IMU provided 9-axis data, consisting of three-axis accelerometer, three-axis magnetometer, and three-axis gyroscope measurements. Figure 1 shows the MoBI instrumentation on the participants, and Fig. 2 displays all data simultaneously collected from one of the participants.
(a) EEG setup on the actors of Dyad B. The setup includes a 32-channel EEG head cap connected to a signal transmitter (MOVE system). The transmitter is attached to the actors’ back with a belt around their waist to allow freedom of movement. Actors also wore Empatica E4 wristbands. (b) Channel locations of the 32-channel headset with electrodes around the eyes. (c) Close up view of the electrodes. Front view of the electrodes around the eyes on the acting director (top) and view of the full EEG set up on an audience member (bottom). (d) Stage set for the performance.
All data collected from one participant (P01 - Dyad C) during the performance: EEG data (32 channels: 28 channels for EEG and the remaining 4 channels for EOG), Empatica data for left and right hand (acceleration in x, y and z, BVP, heart rate, EDA and temperature) and IMU sensor data (head and body acceleration in x, y and z). The EEG and EOG signals have a scale of 150 μV, and the signals from Empatica and IMU have a scale of 30 units according to their respective measuring units.
The head circumference of each participant was measured before the experiment to select an EEG cap of appropriate fit5. A 32-channel Ag/AgCl active electrode EEG cap was used to record from the face and scalp. The data were recorded using the BrainVision Recorder software (Brain Products GmbH, Germany). Four EEG electrodes were removed from the cap and used for EOG to capture blinks and eye movements (Fig. 1c). The remaining 28 electrodes were arranged according to the international 10-20 system. The vertical EOG electrodes were positioned 1 cm above (EOG_VA) and below (EOG_VB) to the right eye, while the horizontal EOG electrodes were placed 1 cm on the left (EOG_HL) and right (EOG_HR) temples on the sides of the face. The electrode locations used for recording are shown in Fig. 1b.
The participants were asked to refrain from using products in their hair that may increase the impedance at the scalp electrode interface. Prior to donning the cap, the skin on the face around the eyes, the temples, and the earlobes were gently cleaned with alcohol wipes to remove any dirt and decrease impedance. A conductive electrolyte gel was applied between the electrode tips and the scalp to reduce the interface impedance. The impedance was maintained below 50 kΩ and in most cases reduced to below 20 kΩ before the start of the experiment recording sessions for each participant. The channel impedance values were recorded prior to the start of the experiment and after the end of the experiment.
The video streams were recorded with four tripod-mounted cameras: two Sony EX1, one Sony EX3, and one Canon G1X Mark II. The default camera positions are shown in Fig. 3. For some sequences, one of the four cameras was dismounted from the tripod and used for hand-held recording. The frame rate varied between 24 fps and 30 fps, and resolutions ranged from 720p to 4K. Although the audio is not included in the shared dataset due to copyright restrictions, it was recorded with two Tascam DR-05 field recorders in stereo mode at 48kHz / 24 bit. In post-production - using the AVID Media Composer software - the camera source files were converted to 1080p / 25 fps. The video and audio streams were synchronized using the recorded UTC time screen, audiovisual hand clap cues and - where necessary - manual alignment based on visual cues and audio information from the discarded camera microphones. The aligned video streams were then merged by resizing them into a four-quadrant, single-stream video representing four camera perspectives. The four (2 × 2) audio channels were merged and mixed down to 2-channel stereo, discarding duplicate channels to avoid noise. Still images of the videos are shown in Fig. 4.
(a) Top view of Quintero Theatre located in the University of Houston. (b) Right view of Quintero Theatre with the actors. Section highlighted in blue shows what will be captured by the bottom right camera, as displayed in Fig. 3c. (c) View from the bottom right camera. Two actors, Alice and Dan, act out an emotional scene. The scene takes place in a hotel room set. (d) EEG setup on 3 participants (2 actors and 1 director/audience member). Setup includes a 32 channel electrode cap (M), secure belt (N), and EEG wireless transmitter (O).
MoBI data from all actors was collected from both rehearsals and performance. (Rehearsal I) The actors gave table readings of the script and conducted partial rehearsals under the guidance of the director. (Rehearsal II) The actors improvised and performed Meisner repetition exercises. The director provided feedback between and during partial and full rehearsals. (Performance) The actors performed the scene in front of an audience. MoBI data was also collected from three audience members.
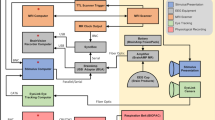
The data streams of the video, the wireless EEG, and head IMU device were time synchronized using a custom hardware trigger and aligned using MATLAB R2019a (The Mathworks Inc., Natick, MA). The video recordings captured a computer screen displaying UTC time as streamed through www.time.gov(The National Institute of Standards and Technology, US Department of Commerce). This time recording allowed the E4 Empatica data to be synchronized by matching the corresponding Unix time stamps in the data stream using MATLAB R2019a.
Experimental Protocol
The experiment was conducted over the course of four days at the Quintero Theater (Kathrine G. McGovern College of the Arts, University of Houston). Each rehearsal session lasted for about one hour per dyad, all led by the same director. The actor dyads did not witness the rehearsal sessions of the other dyads. The full scene itself lasted about 7 minutes. Table 1 shows the itinerary followed during the experiment.
On the rehearsal days, the director staged an emotionally charged scene from Patrick Marber’s stage play “Closer”6 (scene 11) with three dyads (A, B and C) of two acting students each: a female playing the character “Alice”, and a male playing the character “Dan”. The scene progresses from harmony through disagreement to violent altercation. Each rehearsal session was conducted as a sequence of predefined rehearsal methods. These rehearsal methods included: table readings, improvisations, physical blocking, scene rehearsals, and Meisner repetition exercises8. MoBI data was gathered from all six actors during rehearsals and the performance. During rehearsals, MoBI data from the director was also gathered. On the day of the performance, MoBI data was collected from the six acting students, and from three audience members. The performances of the three dyads took place sequentially on the same day, in front of a live audience. On a screen to the side of the stage, out of sight for the actors, a visual representation of the actors’ brain activity was displayed in real time for the audience. After the performance, an open question and answer session between the audience members, the acting students, the director, and the research team ensued.
Each participant wore a 32-channel EEG headset and wireless signal transmitter secured by a belt on their waist as they continued with the experiment. Raw EEG data was compiled for each participant member for each dyad (A, B and C) into one large file for each day (rehearsal-day 1, rehearsal-day 2, performance-day 3). P01 corresponds to the actor playing the character “Dan”, P02 the actress playing the character “Alice”, and P03 the director (in days 1 and 2) and a distinct audience member (in day 3). There are 27 individual datasets corresponding to: each participant (three participants) on each dyad (three dyads) on each day (three days). P01 on Dyads A/B/C included 31 EEG channels (27 EEG, 4 EOG), instead of 32, due to error with the electrode CP6, which had to be removed. P02 and P03 include 32 EEG channels (28 EEG, 4 EOG).
Assessment of Data Quality
Custom MATLAB R2021a software and functions from EEGLAB9 (https://sccn.ucsd.edu/eeglab/index.php) were used to assess the quality of the data. This assessment included impedance check, assessment of motion artifacts, line noise and physiological artifacts such as eye movement and muscle activity. An automated process for the removal of bad channels and timepoints was implemented. Figure 5 shows the pre-processing steps suggested, with their respective parameters, for a de-noising approach for each performance data set.
Suggested EEG-denoising preprocessing pipeline. The steps only apply to EEG data. They can be split into two sections: video plus EEG data visualization, and cleanup of the EEG data.
EEG data was pre-processed using EEGLAB9. By downsampling the EEG data to 240 Hz (EEGLAB pop_resample plugin), video and EEG data simultaneous visualization was integrated as that frequency is a multiple of the video resolution (24 fps) without a significant loss of data.
Noisy data portions were identified visually in the EEG and video, and timestamps were extracted for rejected 1-second segments that were observed with large motion artifacts. The remaining data with clean epochs was concatenated and moved to the preprocessing stage. In order to remove ocular activity obtained with the four EOG channels, EEG was filtered with the H∞ filter considering the parameters of how rapidly the weights vary with time (q = 1e-10), the initial noise covariance matrix (p0 = 0.5), and the maximum bound on H∞ gain (γ= 1.15)10.
EEG data was then re-referenced to the average using robust re-referencing though the PREP pipeline11, and band-pass filtered using a 5th order Butterwoth FIR filter in [0.01 - 50] Hz range.
In addition, for the purpose of removing motion-related noise, a Motion Artifact Filter was implemented at second order (N = 2), using similar parameters to the H∞ filter, including q = 1e-10 and γ = 1.5, with three samples time taps and f = 1 Hz. This frequency was identified from the visualization of the power spectrum density of the gravity-compensated acceleration12. After motion artifact removal, the Artifact Subspace Reconstruction (ASR) algorithm was applied to the data13,14. It was used to reconstruct data periods on each channel that are contaminated by an artifact, which amplitude is higher than κ = 10 standard deviations.
Finally, Extended Infomax Independent Component Analysis (ICA)15 was applied to decompose data into independent sources using the principal component analysis option to compensate for the maximum number of components to the data rank reduction caused by the interpolation and re-referencing. ICA was used to detect independent components (ICs) of artifacts mixed with the EEG signals, such as ocular, muscle, electrocardiography, and power lines artifacts16.
Subsequently, the standard template from the Montreal Neurological Institute (MNI) was used for dipole fitting and improvement of artifactual ICs removal by identifying the scalp projection for each of them. A K-means clustering algorithm within EEGLAB was applied to group all the scalp projections of each IC across all subjects, considering their spatial coordinate and distribution. Components with under 10% residual scalp map variance from the projection of the best-fitting equivalent dipole were kept. The optimal number of clusters, determined as 5 in this case, was identified using the Silhouette method17. ICs from subjects not assigned to any cluster, as well as those with spatial coordinates distant from the clusters centroid and power spectra indicative of non-brain artifacts, were excluded from the EEG data. A group of experienced researchers evaluated the independent components and removed those that they identified as artifacts. An example of the identified artifactual components from the EEG data are shown in Fig. 6. Figure 7 shows a sample output of the EEG pre-processing sequence.
Example of decomposed artifacts that affect EEG signals from P01 - Dyad C during Performance, such as muscle, ocular, and other type of artifacts.
Comparison of EEG signals (P01 - Dyad C during Performance) before (EEG and EOG) and after (EEG only) the pre-processing steps were implemented in the data.
Data Records
All data files are available at FigShare18. The data are archived in a single file set and organized with the following naming convention:
BOA_recordingX_P0X_DyadX_suffix
-
BOA: Brain on Acting
-
recording1: Day 1 of rehearsal, first EEG recording of the experiment (roughly an hour long)
-
recording2: Day 2 of rehearsal, second EEG recording of the experiment (roughly an hour long)
-
recording3: Final performance of scene on Day 4, final EEG recording of the experiment (roughly 7 minutes long)
-
P01: Actor on Dyad X for Dan
-
P02: Actor on Dyad X for Alice
-
P03: Same acting director for all dyads(for recording1 and recording2) or Audience member watching Dyad X (for recording3)
-
DyadA: Data collected for Dyad A
-
DyadB: Data collected for Dyad B
-
DyadC: Data collected for Dyad C
-
_suffix: Type of data
_EEG.mat
The _EEG.mat file contains a 1 × 3 structure (described in Table 2; variable name: EEG) with electroencephalographic data collected by EEG sensors. The first element in the structure corresponds with the recording 1 (day 1 of rehearsal); the second element in the structure corresponds with the recording 2 (day 2 of rehearsal) and the third element corresponds with the recording 3 (final performance). The structure contains multiple fields corresponding to each dyad (A,B and C) and each subject (P01, P02 and P03). Table 2 breaks down the information contained within these fields.
_opals.mat
The _opals.mat file contains a 1 × 2 structure (described in Table 3; variable name: opals) with kinematic data collected by opal sensors. The first element in the structure corresponds with the head sensor; the second element in the structure corresponds with the body sensor. The structure contains multiple fields. Table 3 breaks down the information contained within these fields. In recording 3 for dyad A, there are two minutes of data missing from the end due to a technical error.
_empatica.mat
The _empatica.mat file contains a 1 × 2 structure (described in Table 4; variable name: empatica) with kinematic and bodily data collected by the Empatica E4. The first element in the structure corresponds with the left-hand sensor; the second element in the structure corresponds with the right-hand sensor. The structure contains multiple fields. Table 4 breaks down the information contained within these fields.
CSV Files
CSV versions of the data files are included in the repository, following the same naming convention defined previously. All data have been organized into folders within a ZIP file for easier access and download.
Video records
The video recordings corresponding to each day are included in the repository and are named according to the convention: “DyadX_Y_MMDDYY.mp4”, where X refers to the dyad identifier (A, B, or C), Y indicates the session type ("Rehearsal” or “Performance”), and MMDDYY denotes the date of the recording (month, day, year), as detailed in Table 1. Due to copyright restrictions associated with the script content7, these videos are provided without audio.
Call Sheet and Planning
Refer to supplementary material document A for the call sheet and planning document for each rehearsal and performance.
Annotation Script
The six rehearsal videos and three performance videos were annotated in Microsoft Excel to provide an instant-by-instant account of the activities observed. These annotations facilitate the navigation and filtering of activities and events. The annotation format is also included in the public data repository mentioned in Data Records. To link the annotation file to the datasets, the multi-modal data was synchronized based on a shared timeline. This file includes a dedicated section specifying the data positions, defined as Segmentation, for the start and end of each annotated event. These positions refer to the exact sample numbers for each recorded modality, enabling precise identification and extraction of the corresponding segments from the raw data. This structure ensures a direct and consistent correspondence between the annotations and the dataset entries throughout the six rehearsals and three performances.
Technical Validation
Data Synchronization
The EEG, Empatica and OPALs data were synchronized with the rehearsal and performance video files. A four second subset of the time-synchronized data was chosen for further analysis, as shown in Fig. 8. During the experiment, the participants were asked to clap and clench their teeth at the same time. The research assistant added an event into the EEG data stream at the same time so that the claps could act as a marker for the start and end of the experiment.
(a) A short sample of time-synchronized subset of raw EEG, EOG, ACC, and BVP data for Dyad B and one member of the audience during Performance. The timeseries EOGv (vertical) and EOGh (horizontal) are computed as bipolar signals for the vertical and horizontal EOG channels. (b) Dyad B during the performance in front of an audience.
Usage Notes
All the datasets provided in this study consist of raw data. For data preprocessing and analysis, the authors recommend using MATLAB R2021a (The Mathworks Inc., Natick, MA) and/or EEGLAB9 (https://sccn.ucsd.edu/eeglab/index.php). De-noising algorithms have been suggested for identifying and removing both physiological and non-physiological artifacts from the EEG data, see Fig. 5.
A key limitation of the dataset is the absence of audio in the video recordings. While audio was originally captured during data collection, it is not included in the shared dataset due to copyright restrictions7. Researchers intending to pursue audio-dependent analyses should be aware of this constraint. To maintain transparency, we have retained a description of the audio recording methodology in the Methods section, despite the unavailability of the audio files.
Code availability
No custom code is needed to access the data. However, due to the large size and complexity of the datasets, opening the CSV files can be computationally intensive. We recommend using programming environments such as Python or MATLAB for accessing and analyzing the data, as Microsoft Excel may not efficiently handle the volume of information and could limit visualization capabilities. Based on internal testing, the time required to open individual CSV files ranged from approximately 12 minutes on a high-performance workstation (270 GB RAM, AMD®Ryzen Threadripper PRO 5975WX) to 50 minutes on a laptop (32 GB RAM and an Intel Core Ultra 9 processor).
References
Greaves, D. A. et al. Exploring Theater Neuroscience: Using Wearable Functional Near-infrared Spectroscopy to Measure the Sense of Self and Interpersonal Coordination in Professional Actors. Journal of Cognitive Neuroscience 34, 2215–2236, https://doi.org/10.1162/jocn_a_01912 (2022).
McDonald, B., Goldstein, T. & Kanske, P. Could Acting Training Improve Social Cognition and Emotional Control. Frontiers in Human Neuroscience 14, 1–5, https://doi.org/10.3389/fnhum.2020.00348 (2020).
Brown, s., Cockett, P. & Yuan, Y. The neuroscience of Romeo and Juliet: an fMRI study of acting. Royal Society Open Science 6, 3, https://doi.org/10.1098/rsos.181908 (2019).
Garbarino, M., Lai, M., Tognetti, S., Picard, R. & Bender, D. Empatica E3 - A wearable wireless multi-sensor device for real-time computerized biofeedback and data acquisition. Proceedings of the 4th International Conference on Wireless Mobile Communication and Healthcare - Transforming healthcare through innovations in mobile and wireless technologies. 3–6, 2014. https://doi.org/10.4108/icst.mobihealth.2014.257418
Cruz-Garza, J. G. et al. A novel experimental and analytical approach to the multimodal neural decoding of intent during social interaction in freely-behaving human infants. Journal of Visualized Experiments: JoVE 104, https://doi.org/10.3791/53406 (2015).
Marber, P. Closer. Grove Press, 1999.
Marber, P. Scene 11. In Closer. Grove Press, 1999.
Meisner, S., Longwell, D. & Pollack, S. Sanford Meisner on Acting. Vintage Books, 1987.
Delorme, A. & Makeig, S. EEGLAB: An open source toolbox for analysis of single-trial EEG dynamics including independent component analysis. Journal of Neuroscience Methods 134, 9–21, https://doi.org/10.1016/j.jneumeth.2003.10.009 (2004).
Kilicarslan, A., Grossman, R. & Contreras-Vidal, J. L. A robust adaptive denoising framework for real-time artifact removal in scalp EEG measurements. Journal of Neural Engineering 13, 2, https://doi.org/10.1088/1741-2560/13/2/026013 (2016).
Bigdely-Shamlo, N., Mullen, T., Kothe, C., Su, K. & Robbins, K. The PREP pipeline: Standardized preprocessing for large-scale EEG analysis. Frontiers in Neuroinformatics 9, 1–19, https://doi.org/10.3389/fninf.2015.00016 (2015).
Kilicarslan, A. & Contreras-Vidal, J. L. Characterization and real-time removal of motion artifacts from EEG signals. Journal of Neural Engineering 16, 5, https://doi.org/10.1088/1741-2552/ab2b61 (2019).
Kothe, C. A. E. & Jung, T. P. Artifact removal techniques with signal reconstruction. Google Patents 11, 417–441 (2016).
Mullen, T. R. et al. Real-time neuroimaging and cognitive monitoring using wearable dry EEG. IEEE Transactions on Biomedical Engineering 62, 2553–2567, https://doi.org/10.1109/TBME.2015.2481482 (2015).
Lee, T. W., Girolami, M. & Sejnowski, T. J. Independent component analysis using an extended infomax algorithm for mixed subgaussian and supergaussian sources. Neural computation 11, 417–441, https://doi.org/10.1162/089976699300016719 (1999).
Rejer, I. & Gorski, P.Benefits of ICA in the Case of a Few Channel EEG. Proceedings of the Annual International Conference of the IEEE Engineering in Medicine and Biology Society, EMBS, 7434–7437, https://doi.org/10.1109/EMBC.2015.7320110 (2015)
Baarsch, J. & Celebi, M. Investigation of internal validity measures for K-means clustering. Proceedings of the International MultiConference of Engineers and Computer Scientists (2012).
Hendry, M. F. et al. Mobile Brain-Body Imaging and Visual Data of Theatrical Actors During Rehearsal and Performance. FigShare Collection, https://doi.org/10.6084/m9.figshare.c.7271338.v4 (2024)
Acknowledgements
This research is partly supported by National Science Foundation (NSF) REU Award #1757949, NSF IUCRC BRAIN Site award #2137255, NSF AccelNet #2412731, and a research grant by the Zurich University of the Arts (ZHdK), Institute for the Performing Arts and Film, Zurich University of the Arts, Zurich, Switzerland. The authors would like to thank the University of Houston School of Theatre and Dance: Rachel Cendrick, Josh Rahman, Clare Keating, Sophie Mobbs, Tony Shortt, Leo Rojas, Olivia Swasey, Michael Duran, and Shelby Kesler for their insights in follow-up transdisciplinary discussions of the experiment. Students of the Jack J. Valenti School of Communications recorded the video data: Alexander Brovig, Guillermo Alán Cantú, Richard Davis, Devin Pruitt and Liana Rawley. At ZHdK, Prof. Anton Rey and Dr. Miriam Loertscher provided input for the experiment protocol, and Norbert Kottmann synchronized the audio/video data. Students at the Laboratory for Noninvasive Brain-Machine Interface Systems at the University of Houston assisted with logistics and data collection: Devon Bellman, Ryan Thankson, Kevin C. Nathan, Justin A. Brantley, Alexander Craik. Fangshi Zhu assisted with video data collection.
Author information
Authors and Affiliations
Contributions
Manuel Flurin Hendry: Conceptualization, Methodology, Validation, Investigation, Data Curation, Writing - Review & Editing, Visualization, Supervision, Project administration. Jesus G. Cruz-Garza: Methodology, Software, Validation, Formal Analysis, Investigation, Data Curation, Visualization, Writing - Review & Editing. Esther A. Delgado-Jimenez: Software, Validation, Formal Analysis, Investigation, Data Curation, Visualization, Writing - Original Draft. Yoshua E. Lima Carmona: Software, Validation, Formal Analysis, Investigation, Data Curation, Visualization, Writing - Original Draft. Aime J. Aguilar-Herrera: Software, Validation, Formal Analysis, Investigation, Data Curation, Visualization, Writing - Review & Editing. Mauricio A. Ramirez-Moreno: Software, Validation, Formal Analysis, Investigation, Data Curation, Writing - Review & Editing. Akshay Sujatha Ravindran: Investigation, Writing - Review & Editing. Andrew Y. Paek: Investigation, Writing - Review & Editing. Michelle Smith: Investigation, Writing - Review & Editing. Julia Kan: Formal Analysis, Data Curation, Visualization, Writing - Review & Editing. Mikayla Fors: Formal Analysis, Data Curation, Writing - Review & Editing. Amirah Alam: Formal Analysis, Data Curation, Writing - Review & Editing. Ruofan Liu: Formal Analysis, Data Curation, Writing - Review & Editing. Adam Noble: Methodology, Writing - Review & Editing. Jose L. Contreras-Vidal: Conceptualization, Methodology, Software, Validation, Formal Analysis, Investigation, Resources, Data Curation, Writing - Review & Editing, Visualization, Supervision, Project administration, and Funding acquisition.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hendry, M.F., Cruz-Garza, J.G., Delgado-Jiménez, E.A. et al. Mobile Brain-Body Imaging and Visual Data of Theatrical Actors During Rehearsal and Performance. Sci Data 12, 1421 (2025). https://doi.org/10.1038/s41597-025-05713-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-025-05713-2