Abstract
Indigenous goats exhibit strong adaptability to remote environments and provide a vital source of protein for residents in impoverished regions. Whole-genome sequencing (WGS) data can elucidate the economic traits of these goats. However, the limited genomic resources have constrained the functional dissection of advantageous traits and hampered the breeding process in goats. Here, we present a WGS dataset of 304 goat samples, from the Guizhou black (n = 104), Hezhang black (n = 100), and Tashi (n = 100) goat breeds. The dataset consists of 6.0 TB of paired-end sequences generated through the BGI-T7 sequencing platform. The data has an average sequencing depth of 7.5X, a mapping ratio of 97.0%, and genome coverage of 98.4%. Following the variant calling and hard filtration, a total of 27.13 million single nucleotide polymorphisms (SNPs) and 2.76 million insertions-deletions (InDels) were retained. To our knowledge, this is the largest goat WGS dataset from Southwest China, significantly enriching the global public genomic resources for the study of genetic diversity, environmental adaptations, and functional genes in goats.
Similar content being viewed by others
Background & Summary
The domestic goat is an important economic livestock in developing countries, especially in Asia and Africa, as it provides essential products such as milk, meat, and mohair, cashmere, and furs produced from cashmere goats1. China ranks first in the world both in the number of breeds and the total population of goats raised2. In 2024, 69 national goat breeds were recognized by the National Catalogue for Livestock and Poultry Genetic Resources (http://www.zys.moa.gov.cn/gsgg/), with numbers concentrated in mainly South China. Beyond these national-level goat breeds, numerous local breeds exist in smaller populations. Guizhou province, featuring a subtropical mountain climate, is the center of the Karst ecosystems in Southwest China3. The province is home to several goat breeds, including but not limited to the Guizhou black goat (GBG), the Hezhang black goat (HBG), and the Tashi goat (TG). Through long-term natural and artificial selection, these breeds have developed small body size and strong mountain foraging ability, well-suited to the local Karst ecosystem and providing a basic income for farmers.
As the GBG is the leading goat breed in Guizhou Province, it has gained extensive research attention, particularly regarding its growth performance. For example, Yuan et al.4 investigated the effect of allicin on the growth performance of GBG, while Long et al.5 focused on the effect of Chinese herbal medicine residues. Based on variant-trait associations, the Insertions/Deletions (InDels) in GATA46, and the copy number variants in CADM27, Opn48, SNX299, and MYLK410 were identified to be significantly associated with growth traits; the Single Nucleotide Polymorphisms (SNP) in ACADM11 was determined to be significantly associated with slaughter and meat quality traits. Additionally, this breed’s reproductive performance and population structure have generated some attention. Before this study, the largest genome resequencing dataset of GBG, comprising 30 individuals, was conducted by Prof. Wang’s team12,13 and by integrating this dataset and other downloaded data, they performed a population structure and selective signals analyses to explore genomic characteristics12 and causal structural variants linked to mutton flavor13. Compared to GBG, research on HBG and TG remains scarce, and for HBG, no relevant publications were obtained from the Web of Science or the PubMed database. For TG, there is only one study from this lab that identified genomic variants associated with body conformation traits14. Overall, the absence of sufficient genomic information for these local breeds severely hampers breed genetic evaluation, functional gene dissection, genetic improvement, and conservation efforts. Hence, additional genome sequence data is desperately needed.
Whole-genome sequencing (WGS) is a powerful technique that enables researchers to analyze an organism’s entire genetic makeup, and a considerable number of studies used WGS data for different reasons. For example, Cai et al.15 investigated the evolutionary history of cashmere-producing goats in China by integrating ancient and current goat genome sequencing data, while Liu et al.16 constructed a goat pan-genome using the WGS dataset that comprises 813 individuals and revealed the patterns of gene loss during domestication. Based on a genome-wide association study, WGS was used to identify genes associated with traits such as milk yield17, cashmere yield18, hair diameter19, and hair density20. Based on a selective sweep analysis, some breed-specific traits have been elucidated, such as the high-altitude adaptation of Tibetan cashmere goats21 and the rapid muscle growth of Boer goat22. Additionally, the WGS technique has also been used to evaluate breed genetic characteristics23,24 and provide core variants to develop SNP chips25,26. Altogether, WGS offers comprehensive genetic insights essential for evolutionary study, trait analysis, breeding, and conservation. Consequently, this makes the generation of genomic data for underrepresented breeds increasingly essential.
Here, we present a new WGS dataset from three indigenous goat breeds, including GBG (n = 104), HBG (n = 100), and TG (n = 100). The dataset, encompassing 6.0 TB of raw sequence data, constitutes the largest WGS dataset generated from the Karst region of Southwest China to date. Sequencing was performed at an average depth of 7.5X, ensuring the necessary power and resolution for genomic analyses. By aligning the sequencing data to the Capra hircus reference genome (ARS1.227), https://www.ncbi.nlm.nih.gov/datasets/genome/GCF_001704415.2/) and performing variant calling and variant filtration, a total of 27.13 million SNPs and 2.76 million InDels were identified. The reliability of this WGS dataset is evident from its sequencing base quality, variant quality, sample independence, and breed independence.
This dataset will fill gaps in the genomic resources of these goat breeds, allowing for the (1) Calculation of genetic metrics to evaluate their current status, and infer their genetic relationships; (2) Identification of genomic variants associated with biological traits; (3) Integration of other genomic resources and tracing species evolution and domestication; (4) Comparison of genomic data of different breeds to identify regions under positive selection; and (5) Use of core variants to develop SNP chips for future breeding purposes. Altogether, this large-scale WGS dataset from the Karst region of Southwest China significantly enriches global goat breed genome resources and is crucial for studying population genetics and elucidating economic traits.
Methods
Animals and sample collection
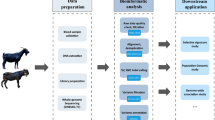
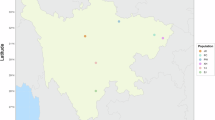
This experiment was approved by the Animal Care and Use Committee of Guizhou University (No.EAE-GZU-2023-E047). A total of 304 adult goats, including Guizhou black goats (GBG, male n = 84, female n = 20), Hezhang black goats (HBG, male n = 80, female n = 20), and Tashi goats (TG, male n = 19, female n = 81), were collected from Guizhou province, China (Fig. 1a). To thoroughly sample the genetic diversity of goat breeds, individuals were selected from dozens of smallholder farms in Anshun, Bijie, and Rongjiang City, respectively (Fig. 1b). These breeds exhibit distinct phenotypes, such as differences in coat color and horn type (Fig. 1c). The goats were allowed to graze naturally and were supplemented with corn and dry hay. Then, 3–5 mL jugular venous blood was sampled from each animal, anticoagulated with EDTA, and stored at −20 °C until DNA extraction.
Sampling locations and study workflow. (a) The three goat breeds included the Guizhou black goat (GBG, n = 104), the Hezhang black goat (HBG, n = 100), and the Tashi goat (TG, n = 100). (b) Sampling area and geographic distribution of the study populations. (c) Breed characteristics in terms of phenotype and production, and the purpose of the breed. (d) Workflow of this study.
Whole genome sequencing
The DNA was extracted from blood samples of 304 goats using the standard phenol-chloroform protocol (Fig. 1d). The concentration, integrity, and purity of the genomic DNA were assessed using agarose gel electrophoresis and a NanoDrop spectrophotometer (Thermo Scientific, Waltham, MA, USA). During library construction, the genomic DNA was amplified and randomly broken into fragments of about 150 bp, followed by the addition of sequencing adapters. Then, the DNA library was sequenced by Compson Biotechnology Company (Beijing, China) using the BGI-T7 platform.
Genomic alignment and variant calling
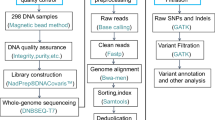
The genomic analysis pipeline primarily comprised quality control, read mapping, variant calling, variant filtering, and variant annotation (Fig. 1d). The WGS raw data in fastq files for all 304 goats, were quality-controlled using the fastp software (v0.23.428) to obtain clean data, while sequence alignment and variant detection were processed using the Sentieon Genomics software (v20230829). Briefly, clean reads were aligned to the goat reference genome (ARS1.227), whereafter the BAM files were sorted and duplicates marked. Variant calling was conducted using the Sentieon haplotyper module to independently generate a genomic Variant Call Format (gVCF) file for each individual. Finally, variant joint calling was performed in the Sentieon GVCFtyper module to create a common VCF file from all the gVCF files. It is important to note that the module tools in Sentieon software are fully faithful to the classic BWA-GATK pipeline30,31.
Variant types and variant annotation
Statistical metrics generated in the above variant calling step, include Mapping Quality (MQ), Quality by Depth (QD), Fisher Strand (FS), and Strand Odds Ratio (SOR), evaluating coverage depth and alignment quality at variant positions, which help filter out potential false positives and ensure accurate variant calling. Both SNP and InDel variants were filtered using the SelectVariants module in the GATK software (v4.1.8.131). Further filtration was performed using the Vcftools software (v0.1.1632) to filter out variants when the average depth was less than 5 and the missing genotype rate exceeded 10% in all samples. The variant depth and cumulative proportions were performed to assess variant depth, which in turn serves as an indicator of variant quality, while the filtered SNPs and InDels were functionally annotated using the snpEff software (v.5.133). Additionally, the variant locations in intronic, untranslated, upstream, downstream, and intergenic regions were calculated.
Principal component analysis, genetic kinship, and population structure
To evaluate sample and breed independence, the principal component analysis (PCA), kinship analysis, phylogeny analysis, and population structure analysis were conducted based on genome-wide SNPs. The PCA was performed in Plink (v0.7634), and each principal component was tested based on the twstat method using the EIGENSTRAT software (v6.1.435). The kinship matrix was calculated in the GEMMA software (v0.98.536) and visualized in the R package heatmap (v1.0.12). Phylogenetic distance was estimated using the VCF2Dis software (v1.54, https://github.com/BGI-shenzhen/VCF2Dis) and visualized using Figtree software (v1.4.4, http://tree.bio.ed.ac.uk/software/figtree/). Under the different hypothetical subpopulations (from K = 2 to K = 10), the population structure for these samples was calculated using the Admixture software (v1.3.037) and visualized in the Python package PONG (v1.538).
Data Records
All original genome sequencing data in FASTQ format have been deposited in the Genome Sequence Archive39 on the China National Center for Bioinformation (CNCB) platform under accession number CRA025744 (https://ngdc.cncb.ac.cn/gsa/browse/CRA025744)40, and Sequence Read Archive on National Center for Biotechnology Information (NCBI) under accession number PRJNA1281799 (https://www.ncbi.nlm.nih.gov/sra/SRP594418)41. The final SNP.vcf (27,134,128 SNPs) and InDel.vcf (2,762,642 InDels) files were deposited in the Genome Variation Map42 on the CNCB platform under accession number GVM00105143, and the European Variation Archive (EVA)44 under accession number PRJEB9083145.
Technical Validation
Quality control for sequencing data
The whole genome sequencing of 304 goats using the BGI-T7 platform yielded 5968.2 Gb of raw sequence data (Fig. 2a, Table 1). For each sample, the unfiltered reads ranged from 97 to 386 million reads, and sequencing yields ranging between 11.3 Gb and 57.9 Gb were obtained. The average sequencing depth was 7.5X for all the samples, which varied from 5.0X to 22.0X (Fig. 2a, Table 1). For all samples, 97.4–99.4% and 93.2–98.0% of the bases achieved the high Phred quality score of Q20 (sequencing error rate < 0.01) and Q30 (sequencing error rate < 0.001), indicating the high base calling accuracy, respectively. Besides, the average GC content varied from 39.7% to 44.4%. Each position in the 150 base pair read obtained high-quality scores of 35 (Fig. 2b), and almost all the reads had quality scores that varied from 35 to 40, confirming the overall high-quality scores of all the sequencing reads (Fig. 2c). For all samples, the genome coverage of the sequencing reads on the goat reference genome exceeded 95% (Fig. 2d), the mapped depth exceeded 5X (Fig. 2e), and the properly mapped rates varied from 73.54% to 99.2% (average of 97.0%). These indicators demonstrate the high quality of the sequencing data based on sequencing data yield, base quality, and genome coverage.
Statistics of sequencing data and read alignment. (a) The sequencing yield, reads, depth, phred quality scores (Q20 and Q30), GC content, and mapping rate of high-throughput sequencing data of 304 goats. (b) Average quality score per base in a 150-base-pair read. (c) The sequence quality score plot revealed that nearly all reads have quality scores ranging from 35 to 40. (d) The genome coverage of sequencing reads on the 29 autosomes of the goat reference genome exceeded 95%. (e) The mapped depth of sequencing reads on the 29 autosomes of the goat reference genome. The mapped depth of all samples exceeded 5X. Each colorful circle or line represents one sample of the 304 goats.
Quality control of SNP and InDel data
After performing joint calling for all samples, a total of 41,559,429 SNPs and 4,891,077 InDels were obtained. To ensure variant quality and minimize false positives, we performed hard filtration using the GATK software31 and assessed variant quality using statistical metrics including MQ, QD, FS, and SOR, obtaining 31,405,939 SNPs and 4,603,254 InDels. The Vcftools software (v0.1.1632) filtered out variants whose average depth was less than 5 and whose missing genotype rate exceeded 10%. Finally, a total of 27,134,128 SNPs and 2,762,642 InDels were retained (Fig. 3a,b).
Variant locations and their cumulative distribution using WGS data of 304 goats. Proportions of all filtered SNPs (a) and InDels (b) at specific chromosomal location categories. The cumulative depth distribution of SNPs (c) and InDels (d) in each of the 304 samples. The average depths for SNPs and InDels were 7.2 and 7.5, respectively. Each colorful line represents one of the 304 goats.
Summary statistics of SNPs and InDels
High-quality variants were evenly dispersed throughout the 29 autosomes of the goat genome, with an average frequency of one SNP per 85 bases and one InDel per 838 bases (Table 2). As shown in Fig. 3a,b, more than half of the SNPs (54.3%) and InDels (56.1%) were located in intronic regions, while only a small percentage (0.8% of SNPs and 0.2% of InDels) were located in Exons. Approximately 1% of variants were located in UTR regions, although the rate of total numbers of SNPs and InDels was approximately 10:1, their distribution and variant classes were similar (Fig. 3a,b). The cumulative depth distribution plots (Fig. 3c,d) illustrated the cumulative proportional curves of variant depth from 1 to 40, and the arithmetic average of variant depth for all SNPs and InDels was 7.2 and 7.5, respectively. These results point to the uniform distribution and high quality of the filtered variants.
Sample and breed independence
We conducted population structure analysis using PCA, kinship analysis, and phylogenetic tree analysis to assess both sample-level and breed-level genetics independence. The first two principal components explained 15.3% of the total variation and showed a clear distinction between the three goat breeds (Fig. 4a). Additionally, the kinship matrix heatmap showed almost no kinship between breeds, with low kinship coefficients within each breed (Fig. 4b). The histogram frequency distribution showed that 90% and 95% of the relatedness coefficients were lower than 0.037 and 0.093, respectively, indicating low kinship among samples and suggesting good sample independence (Fig. 4c). The phylogenetic tree (Fig. 4d) showed the evolutionary relationship of the three goat breeds, and while the three breeds were relatively independent, TG and HBG showed some influence from GBG. As shown in Fig. 4e, the lowest cross-validation error value was 0.268 at K = 3. Population structure analysis revealed that the ancestral composition of all samples can be easily distinguished by breed at K = 3 (Fig. 4f), and although a few samples showed ancestry from other breeds, this is considered normal given their geographical proximity of 200 km. These results effectively demonstrated sample independence, breed independence, and the high quality of the whole-genome resequencing dataset.
Population genetic analyses using WGS data from 304 goats indicated satisfactory sample independence and breed independence. (a) PCA reveals population stratification among the three goat breeds. (b) The kinship matrix calculated based on genome-wide SNPs can also differentiate the three goat breeds. (c) The histogram frequency distribution of the pairwise kinship coefficients, where low kinship coefficients among most samples indicate good sample independence. (d) Maximum-likelihood phylogeny was constructed using genome-wide SNPs of 304 goats. (e) The line chart of cross-validation (CV) error under different numbers of hypothetical subpopulations. (f) ADMIXTURE analysis of 304 samples based on genome-wide SNPs under a model with three ancestral components (K = 3). These results show breed independence and low intra-breed kinship, confirming the high quality of the WGS dataset. GBG, Guizhou black goat; HBG, Hezhang black goat; TG, Tashi goat.
Usage Notes
This study presents both original raw reads and processed variant files. Notably, these variants were obtained based on the most common reference genome of the San Clemente breed (ARS1.2, https://www.ncbi.nlm.nih.gov/datasets/genome/GCF_001704415.2/). Our dataset includes both bucks and does, but due to the lack of effective Y-chromosome information in the reference genome, variants on this chromosome were missing in the VCF files. The other two telomere-to-telomere genome assemblies of the goat genome are available in the NCBI database: ASM4082201v1 (https://www.ncbi.nlm.nih.gov/datasets/genome/GCA_040822015.1/), assembled by Prof. Su’s team46, and T2T-goat2.0 (https://www.ncbi.nlm.nih.gov/datasets/genome/GCA041735815.1/), assembled by Prof. Li’s team47. These genome versions offer greater flexibility for utilizing our WGS dataset in future applications.
Data availability
The genome sequencing data have been deposited in the CNCB platform under accession number CRA025744 and NCBI under accession number PRJNA1281799 (https://www.ncbi.nlm.nih.gov/sra/SRP594418). The identified variants data were deposited in the CNCB platform under accession number GVM001051 and the ENA under accession number PRJEB90831.
Code availability
All genomic analyses were conducted using standard bioinformatic tools on a Linux operating system. The main steps, software, and code/parameters used to process the data from raw sequencing reads to the VCF file are available on GitHub (https://github.com/jpanzhang/WGS.pipeline).
References
Naderi, S. et al. The goat domestication process inferred from large-scale mitochondrial DNA analysis of wild and domestic individuals. Proc Natl Acad Sci USA 105, 17659–17664, https://doi.org/10.1073/pnas.0804782105 (2008).
Li, M. H., Li, K. & Zhao, S. H. Diversity of Chinese indigenous goat breeds: A conservation perspective - A review. Asian-Australasian Journal of Animal Sciences 17, 726–732, https://doi.org/10.5713/ajas.2004.726 (2004).
Hu, C., Wu, W., Zhou, X. & Wang, Z. Spatiotemporal changes in landscape patterns in karst mountainous regions based on the optimal landscape scale: A case study of Guiyang City in Guizhou Province, China. Ecol. Indicators 150, https://doi.org/10.1016/j.ecolind.2023.110211 (2023).
Yuan, C. et al. Effects of allicin on growth performance, slaughter performance, antioxidant, immune parameters and economic benefits of black goats. Anim. Feed Sci. Technol. 324, https://doi.org/10.1016/j.anifeedsci.2025.116310 (2025).
Long, Y. et al. Partially substituting roughage with traditional Chinese herbal medicine residues in the diet of goats improved feed quality, growth performance, hematology, and rumen microbial profiles. Bmc Veterinary Research 20, https://doi.org/10.1186/s12917-024-04412-1 (2024).
Li, X. et al. Relationships between the mutations of the goat GATA binding protein 4 gene and growth traits. Gene 898, https://doi.org/10.1016/j.gene.2023.148095 (2024).
Xu, Z. et al. Copy number variation of CADM2 gene revealed its association with growth traits across Chinese Capra hircus (goat) populations. Gene 741, https://doi.org/10.1016/j.gene.2020.144519 (2020).
Li, L. et al. Association Analysis to Copy Number Variation (CNV) of Opn4 Gene with Growth Traits of Goats. Animals 10, https://doi.org/10.3390/ani10030441 (2020).
Wang, Q. et al. Detection distribution of CNVs of SNX29 in three goat breeds and their associations with growth traits. Frontiers in Veterinary Science 10, https://doi.org/10.3389/fvets.2023.1132833 (2023).
Shi, S.-Y. et al. Copy number variation of MYLK4 gene and its growth traits of Capra hircus (goat). Anim. Biotechnol. 31, 532–537, https://doi.org/10.1080/10495398.2019.1635137 (2020).
Li, Z. et al. Effect of genetic variation in ACADM on slaughter and meat quality traits in Guizhou black goat. Small Ruminant Research 240, https://doi.org/10.1016/j.smallrumres.2024.107376 (2024).
Chang, L. et al. Identification of genomic characteristics and selective signals in Guizhou black goat. BMC Genomics 25, https://doi.org/10.1186/s12864-023-09954-6 (2024).
Chang, L. et al. Detection of structural variants linked to mutton flavor and odor in two closely related black goat breeds. BMC Genomics 25, https://doi.org/10.1186/s12864-024-10874-2 (2024).
Yang, R. et al. Genome-Wide Association Study of Body Conformation Traits in Tashi Goats (Capra hircus). Animals 14, 1145, https://doi.org/10.3390/ani14081145 (2024).
Cai, Y. et al. Ancient Genomes Reveal the Evolutionary History and Origin of Cashmere-Producing Goats in China. Mol. Biol. Evol. 37, 2099–2109, https://doi.org/10.1093/molbev/msaa103 (2020).
Liu, J. X. et al. The goat pan-genome reveals patterns of gene loss during domestication. Journal of Animal Science and Biotechnology 15, https://doi.org/10.1186/s40104-024-01092-7 (2024).
Scholtens, M. et al. Genome-wide association studies of lactation yields of milk, fat, protein and somatic cell score in New Zealand dairy goats. Journal of Animal Science and Biotechnology 11, https://doi.org/10.1186/s40104-020-00453-2 (2020).
Rong, Y. et al. Genome-wide association study for cashmere traits in Inner Mongolia cashmere goat population reveals new candidate genes and haplotypes. BMC Genomics 25, https://doi.org/10.1186/s12864-024-10543-4 (2024).
Zhang, J., Fang, J., Zhang, S., Xu, J. & Zhao, Y. Several variants on chromosome 10 are associated with coarse hair diameter in Dazu black goats (Capra hircus). Anim. Genet. 56, https://doi.org/10.1111/age.13509 (2025).
Zhang, J., Xiao, M., Fang, J., Huang, D. & Zhao, Y. Phenotypic, transcriptomic, and genomic analyses reveal the spatial temporal patterns and associated genes of coarse hair density in goats. Zool. Res., https://doi.org/10.24272/j.issn.2095-8137.2025.034 (2025).
Li, C. et al. Markhor-derived Introgression of a Genomic Region Encompassing PAPSS2 Confers High-altitude Adaptability in Tibetan Goats. Mol. Biol. Evol. 39, https://doi.org/10.1093/molbev/msac253 (2022).
Yuan, Y. et al. A 1.1 Mb duplication CNV on chromosome 17 contributes to skeletal muscle development in Boer goats. Zool. Res. 44, 303-+, https://doi.org/10.24272/j.issn.2095-8137.2022.384 (2023).
An, Z. X., Shi, L. G., Hou, G. Y., Zhou, H. L. & Xun, W. J. Genetic diversity and selection signatures in Hainan black goats revealed by whole-genome sequencing data. Animal 18, https://doi.org/10.1016/j.animal.2024.101147 (2024).
Zhang, T. et al. Genetic diversity and population structure in five Inner Mongolia cashmere goat populations using whole-genome genotyping. Animal Bioscience 37, 1168–1176, https://doi.org/10.5713/ab.23.0424 (2024).
Guan, S. Y., Li, W. N., Jin, H., Zhang, L. & Liu, G. S. Development and Validation of a 54K Genome-Wide Liquid SNP Chip Panel by Target Sequencing for Dairy Goat. Genes 14, https://doi.org/10.3390/genes14051122 (2023).
Zhao, J. Q. et al. Design and verification of a 25 K multiple-SNP liquid-capture chip by target sequencing for dairy goat. BMC Genomics 26, https://doi.org/10.1186/s12864-025-11576-z (2025).
Bickhart, D. M. et al. Single-molecule sequencing and chromatin conformation capture enable de novo reference assembly of the domestic goat genome. Nat. Genet. 49, 643–650, https://doi.org/10.1038/ng.3802 (2017).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890, https://doi.org/10.1093/bioinformatics/bty560 (2018).
Kendig, K. I. et al. Sentieon DNASeq Variant Calling Workflow Demonstrates Strong Computational Performance and Accuracy. Frontiers in Genetics 10, https://doi.org/10.3389/fgene.2019.00736 (2019).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760, https://doi.org/10.1093/bioinformatics/btp324 (2009).
McKenna, A. et al. The Genome Analysis Toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303, https://doi.org/10.1101/gr.107524.110 (2010).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158, https://doi.org/10.1093/bioinformatics/btr330 (2011).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly 6, 80–92, https://doi.org/10.4161/fly.19695 (2012).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. GigaScience 4, 1–16, https://doi.org/10.1186/s13742-015-0047-8 (2015).
Patterson, N., Price, A. L. & Reich, D. Population structure and eigenanalysis. PLoS Genet. 2, 2074–2093, https://doi.org/10.1371/journal.pgen.0020190 (2006).
Zhou, X. & Stephens, M. Genome-wide efficient mixed-model analysis for association studies. Nat. Genet. 44, 821–824, https://doi.org/10.1038/ng.2310 (2012).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664, https://doi.org/10.1101/gr.094052.109 (2009).
Behr, A. A., Liu, K. Z., Liu-Fang, G., Nakka, P. & Ramachandran, S. pong: fast analysis and visualization of latent clusters in population genetic data. Bioinformatics 32, 2817–2823, https://doi.org/10.1093/bioinformatics/btw327 (2016).
Chen, T. et al. The Genome Sequence Archive Family: Toward Explosive Data Growth and Diverse Data Types. Genomics Proteomics & Bioinformatics 19, 578–583, https://doi.org/10.1016/j.gpb.2021.08.001 (2021).
CNCB Genome Sequence Archive https://ngdc.cncb.ac.cn/gsa/browse/CRA025744 (2025).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRP594418 (2025).
Li, C. et al. Genome Variation Map: a worldwide collection of genome variations across multiple species. Nucleic Acids Res. 49, D1186–D1191, https://doi.org/10.1093/nar/gkaa1005 (2021).
CNCB Genome Variation Map https://ngdc.cncb.ac.cn/gvm/getProjectDetail?project=GVM001051 (2025).
Cezard, T. et al. The European Variation Archive: a FAIR resource of genomic variation for all species. Nucleic Acids Res. 50, D1216–D1220, https://doi.org/10.1093/nar/gkab960 (2022).
ENA European Variation Archive https://identifiers.org/ena.embl:PRJEB90831 (2025).
Wang, Z. et al. Chromosome-level genome assembly of the cashmere goat. Scientific Data 11, https://doi.org/10.1038/s41597-024-03932-7 (2024).
Wu, H. et al. Telomere-to-telomere genome assembly of a male goat reveals variants associated with cashmere traits. Nature Communications 15, https://doi.org/10.1038/s41467-024-54188-z (2024).
Acknowledgements
This work was supported by the National Key Research and Development Program of China (No.2022YFD1300202) and the Gene Mining and Validation of Advantageous Traits in Local Goat Breeds (Guizhou Provincial Department of Agriculture and Rural Affairs).
Author information
Authors and Affiliations
Contributions
Jipan Zhang: Funding Acquisition, Conceptualization, Data Curation, Formal Analysis, Visualization, Writing-Original Draft; Di Zhou: Sample collection, Supervision; Rong Yang: Sample collection; Zhengang Guo: Sample collection; Xingzhou Tian: Sample collection; Yongju Zhao: Funding Acquisition, Supervision. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, J., Zhou, D., Yang, R. et al. Whole-genome sequencing and variants data of 304 indigenous goats from Southwest China. Sci Data 12, 1595 (2025). https://doi.org/10.1038/s41597-025-05893-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-025-05893-x