Abstract
Maize is a critical staple crop globally. Enhancing maize yield per unit area is essential to meet the rising food demand, and increasing planting density has emerged as a key strategy to achieve this goal. Optimizing plant architecture, a strategy central to the “Green Revolution”, is crucial for maize’s adaption to high-density planting. This study reports genome assemblies of two maize inbred lines, D132 and Yu82, characterized by significantly different plant architectures. By leveraging advanced sequencing technologies, we assembled the genomes of D132 and Yu82, achieving total lengths of 2,166.50 Mb and 2,193.33 Mb, respectively, and identifying 40,951 and 40,935 protein-coding genes. These genome data provide valuable resources for in-depth understanding of the genetic mechanisms underlying maize plant architecture and hold promise for contributing to maize breeding improvement.
Similar content being viewed by others
Background & Summary
Maize is a globally important staple crop, serving as a primary source of nutrition for humans and livestock, as well as a key ingredient in various industrial products1. Increasing maize yield per unit area is crucial for sustainably meeting the growing food demand driven by global population growth and worsening environmental conditions2. Enhancing planting density has emerged as a key strategy for boosting maize yield per unit area, as it maximizes land use efficiency and overall production3,4,5. In the United States and China, the world’s two leading maize-producing countries, the development of hybrids tolerant to increased planting density has significantly contributed to the continuous increase in total maize yield over the past few decades4,6,7,8.
Optimizing plant architecture is a well-established approach for enhancing crop tolerance to increased planting density9. This strategy was a cornerstone of the first “Green Revolution”, which led to remarkable yield increases in wheat and rice10. The success of this revolution was primarily attributed to improvements in lodging resistance and the ability to sustain higher planting densities, achieved through significant reductions in plant height—a key plant architecture trait10. These improvements were primarily driven by the rapid deployment of superior haplotypes of two key genes: SD1 in rice and Rht-B1/Rht-D1 in wheat11,12. The identification and utilization of key plant-architecture regulating genes and their favorable alleles thus hold great potential for advancing plant architecture optimization in crops13.
In maize, enhancing tolerance to high-density planting requires the simultaneous optimization of multiple plant architecture traits, including plant height, ear height, leaf angle, leaf length, leaf width, and root system architecture14,15,16,17. A comprehensive understanding of the genomic basis underlying these traits is essential for advancing maize breeding efforts. To contribute to this understanding, we report genome assemblies of two maize inbred lines, D132 and Yu82, which exhibit significantly different plant architectures (Fig. 1). D132, derived from the Lancaster heterotic group, features an expanded plant architecture and has been instrumental in identifying three genes and 17 QTLs associated with plant architecture in maize18,19,20,21. In contrast, Yu82, developed from the Reid heterotic group, exhibits a compact plant architecture characterized by significantly lower ear height, reduced plant height, and smaller leaf angles, widths, and lengths compared to D13222,23 (Fig. 1). By integrating multiple sequencing technologies, the genomes of D132 and Yu82 were assembled with a total length of 2,166.50 and 2,193.33 Mb, respectively, encompassing 40,951 and 40,935 protein-coding genes.
Plant architecture and genomic analyses of D132 and Yu82. (A,B) The whole plant of D132 and Yu82. (C,D) The genome assembly of D132 and Yu82. A window size of 500 Kb was used in the calculation of the density of different genome features.
Methods
Sample collection and DNA extraction
Young leaves of D132 and Yu82 were collected from plants grown in the field of Zhengzhou (E 113.82°, N 34.80°), Henan province, China. Two distinct DNA extraction protocols were employed based on the sequencing requirements. For high-molecular-weight (HMW) DNA isolation, required for PacBio long-read sequencing and BioNano optical mapping, a modified CTAB-based protocol was used. This involved tissue homogenization in liquid nitrogen, incubation in CTAB buffer (Solarbio, LS00066), purification using a DNA extraction reagent (Acmec, AP1012), and precipitation with cold isopropanol. The resulting HMW DNA was washed with 75% ethanol and absolute ethanol before resuspension. Critically, the integrity and high molecular weight of this DNA were confirmed by Pulsed-Field Gel Electrophoresis (PFGE) using a CHEF Mapper XA Chiller System (Bio-Rad). For the construction of Illumina short-read libraries, genomic DNA was extracted separately using the DNeasy Blood & Tissue Kit (QIAGEN).
Genome sequencing of D132
The genome of D132 was sequenced using a combination of Illumina HiSeq X Ten and Pacific Biosciences Sequel II platforms to ensure comprehensive coverage (Table 1).
For Illumina sequencing, three paired-end libraries (200 bp, 300 bp, and 500 bp) and four mate-pair libraries (2 kb, 5 kb, 10 kb, and 20 kb) were constructed, each requiring over 1 μg of genomic DNA. The mate-pair libraries were prepared using the Cre-loxP method24.
For Pacific Biosciences sequencing, 8 μg of genomic DNA was sheared using g-Tubes (Covaris) and concentrated with AMPure PB magnetic beads. Single-strand ends of the sheared DNA were digested with exonuclease VII. Single-strand breaks, nucleotide deletions, and oxidation were repaired using a DNA Damage Repair kit. SMRTbell libraries were constructed with the Pacific Biosciences SMRTbell Template Prep Kit 1.0 and size-selected using Sage ELF for fragments ranging from 14–17 kb. Primer annealing and binding of SMRTbell templates to polymerases were performed using the DNA Polymerase Binding Kit. Sequencing was carried out on the Sequel II platform at Annoroad Gene Technology Company (Beijing, China) with a runtime of 30 hours.
Illumina sequencing generated 645.4 Gb of clean data, providing over 308.4 × coverage of the genome (Table 1)25,26. PacBio single-molecule sequencing produced 47.9 Gb of data, achieving 21.8 × genome coverage27. Additionally, 1132.0 Gb of Hi-C sequencing data from a previous study was used to support chromosome-level genome assembly28.
Genome sequencing of Yu82
The genome of Yu82 was sequenced using a combination of Illumina HiSeq X Ten, Pacific Biosciences Sequel, and BioNano Genomics platforms, ensuring high-quality and comprehensive coverage (Table 2).
For Illumina sequencing, over 1 μg of genomic DNA from Yu82 was used to construct a short-fragment library with an insert size of 350 bp, which was sequenced on the Illumina HiSeq X Ten platform.
For Pacific Biosciences sequencing, at least 8 μg of genomic DNA was sheared using Covaris g-Tubes and concentrated with AMPure PB magnetic beads. Hairpin adapters were ligated to both ends of the DNA fragments, and unligated fragments were digested using exonucleases. The sequencing libraries were prepared using the Pacific Biosciences SMRTbell Template Prep Kit 2.0 and size-selected for molecules ≥ 15 kb using the BluePippin™ system. Primer annealing and polymerase binding were performed using the DNA/Polymerase Binding Kit, and sequencing was carried out on the Pacific Biosciences Sequel platform at Annoroad Gene Technology Company (Beijing, China) with a runtime of up to 10 hours.
Optical mapping of Yu82 was performed using the BioNano Genomics Irys technology. Purified DNA was embedded in a thin agarose layer and labeled and counterstained following the IrysPrep Reagent Kit protocol (BioNano Genomics). Samples were then loaded into IrysChips and imaged using the Irys imaging instrument. Single molecules under 150 kb in size or with fewer than 500 labels were removed. An optical map was produced in two instrument runs with labeled single molecules by IrysSolve (https://bionanogenomics.com/support/software-downloads/).
Illumina paired-end sequencing provided approximately 63.9 × genome coverage29,30, while PacBio single-molecule sequencing achieved ~62.6 × coverage with an N50 read length of 23,602 bp (Table 2)31,32. Optical mapping generated ~438.4 × genome coverage, with an N50 read length of 152,250 bp. Furthermore, 561.7 Gb of Hi-C sequencing data from a previous study was used to support chromosome-level genome assembly28,33,34. Collectively, the sequencing data for Yu82 provided over 820 × coverage of its genome.
Sample collection and transcriptome sequencing of D132 and Yu82
For RNA sequencing, D132 plants were grown in growth chambers (2.8 × 5.6 × 8.2 m) at 25 °C with alternating photoperiods of 16 hours light/8 hours darkness and 8 hours light/16 hours darkness, with a light intensity of 100 μmol m−2 s−1, during the spring of 2016. Then, leaves were collected from D132 plants at the 4th, 5th, 6th, and 7th fully expanded leaf stages. Additionally, for both D132 and Yu82, we collected their leaves at the 3–11 fully expanded leaf stages in the experimental field of Henan Agricultural University, Zhengzhou (E 113.82°, N 34.80°). Three biological replicates of each sample were used for RNA extraction with TRIzol reagent (Invitrogen), followed by paired-end sequencing on the Illumina HiSeq 3000 platform35,36.
In addition to RNA sequencing, we performed PacBio Iso-Seq library preparation and sequencing following the Isoform Sequencing protocol provided by Pacific Biosciences. For each inbred line, pooled cDNA from multiple tissues was used for library construction and sequencing37,38,39,40.
Genomic estimation
To perform an initial genome survey prior to assembly, the genome size and repeat content of D132 and Yu82 were estimated using k-mer depth analysis of sequencing reads. Jellyfish (version v2.1.3) was employed to count the number of 35-mers in the sequencing reads41. For this analysis, specific short-insert paired-end libraries were selected to ensure consistency and comparability. For Yu82, the only available Illumina library (350 bp insert size) was utilized. For D132, which had multiple short-insert libraries (200, 300, and 500 bp insert size), we specifically selected the 300 bp insert size library to maintain comparability with the Yu82 data. Based on this analysis, the genome sizes of D132 and Yu82 were estimated to be 2092.62 and 2162.99 Mb, respectively (Fig. 2, Tables 3, 4). These sizes are close to the genome size of the reference maize inbred line B7342. The proportions of repeat sequences were estimated to be 58.68% for D132 and 64.76% for Yu82.
Distribution of 35-mers analyzed from Illumina sequencing data of D132 and Yu82. The X-axis represents the depth of 35-mers counted using Jellyfish, and the Y-axis shows the frequency of 35-mers at varying depths.
Genome assembly of D132
A total of 11,546,831 polymerase reads were generated for D132 using the PacBio Sequel II platform27,43. These reads were processed into 57,491,036 subreads by adapter trimming and removal of low-quality sequences. Multiple subreads corresponding to the same SMRTbell molecule were merged, resulting in 47.9 Gb of high-fidelity (HiFi) PacBio sequencing data (3,030,776 reads; mean read length: 15,799 bp). The HiFi reads were assembled into 689 contigs with an N50 of 48.8 Mb using HiFiasm44. Redundant and heterozygous contigs were identified and removed based on read coverage and sequence similarity using Purge Haplotigs45.
Next, we used the Hi-C sequencing data of D132 to build the pseudomolecules46. Over 3,454.6 million pairs of clean Hi-C sequencing reads were generated, of which 360,723,879 pairs (10.44%) were uniquely aligned to the D132 contigs by Bowtie2 (version 2.2.3, end-to-end algorithm)47. Ligation sites in the uniquely mapped reads were identified using HiC-Pro (v2.7.8), resulting in 1.5 billion valid interaction read pairs48. These valid read pairs were analyzed with Lachesis to cluster, order, and orient the contigs49. In total, 587 contigs (covering 2,153,520,238 bp, 99.38% of the assembly) were grouped into 10 chromosome groups using an agglomerative hierarchical clustering algorithm (Fig. 3). Within each group, a minimum spanning tree was constructed based on contig interaction frequencies, and the longest path was extracted as the trunk to determine the contig order. Contig orientations were further resolved using a weighted directed acyclic graph (WDGA), with a scoring function to evaluate and select the most credible orientation. The final assembly comprised 10 pseudomolecules and 90 unanchored contigs, with an N50 of 227,258,638 bp (Fig. 1, Table 3)50. Interaction matrices were visualized as heatmaps using Juicebox (https://github.com/aidenlab/Juicebox, version 1.9.9)51. Over 99.4% of the assembled sequences were successfully anchored to the 10 maize chromosomes (Table 3).
Heatmap of Hi-C sequencing data showing genome-wide interactions in D132. Interactions are displayed between 500-kb non-overlapping windows, with interaction intensity represented by the color scale.
Genome assembly of Yu82
Over 135.4 Gb of PacBio Sequel II sequencing data (10 million reads; mean read length: 13,087 bp) were generated for Yu8231,32. Initially, error correction was performed by aligning all reads to a subset of the longest reads, with a minimum overlap of 2000 bp required. The error-corrected reads were assembled into contigs using Falcon (https://github.com/PacificBiosciences/falcon, version v0.3.0), resulting in 9802 contigs longer than 2000 bp, with an N50 of 426,513 bp and a total length of 2186.68 Mb. Meanwhile, the same set of error-corrected PacBio reads were assembled by SMARTdenovo (https://github.com/ruanjue/smartdenovo), producing 4003 contigs longer than 2000 bp, with a total length of 2145.12 Mb and an N50 of 999,298 bp. The assemblies from Falcon and SMARTdenovo were integrated using Quickmerge, yielding 3048 contigs with an N50 of 1,491,368 bp and a total length of 2,143.50 Mb52. Next, the PacBio sequencing reads were aligned back to the assembled contigs using BLASR (https://github.com/PacificBiosciences/blasr, version 5.3.3) with the follwing parameters: “–bam –bestn 5 –minMatch 18 –nproc 4 –minSubreadLength 1000 –minAlnLength 500 –minPctSimilarity 70 –minPctAccuracy 70 –hitPolicy randombest –randomSeed 1”. Contigs were then polished using the Arrow algorithm (https://github.com/PacificBiosciences/GenomicConsensus, version 2.3.3) with the parameter -j 30. Finally, Illumina sequencing data of Yu82 was used to further correct the contigs with Pilon (https://github.com/broadinstitute/pilon, version 1.2.3) using default parameters53. After this step, 3,048 contigs were obtained, with an N50 of 1,503,064 bp and a total length of 2,149.07 Mb.
Scaffolding of the contigs was subsequently performed using Irys optical mapping technology (BioNano Genomics). A total of 877 genome maps, with a mean length of 1.64 Mb, were obtained, covering 1440.08 Mb of the genome. The N50 of these genome maps was 1.65 Mb. The optical maps were aligned to PacBio contigs using IrysSolve (hybridScaffold.pl -B 2 -N 2) to build scaffolds based on overlaps between contigs and optimal maps. Conflicts between the optical maps and PacBio contigs were resolved according to the rules employed in a previous study54. This process yielded 2,401 scaffolds totaling 2,194,445,600 bp, with an N50 of 3,295,496 bp and a maximum scaffold length of 16,916,866 bp.
Chromosome-scale pseudomolecules for Yu82 were constructed using 326 Gb of clean Hi-C sequencing data33,34, following the same approach as described for D132 (Fig. 4). The final assembly consisted of 1,553 scaffolds with a total length of 2,193,328,715 bp and an N50 of 205,377,771 bp (Fig. 1, Table 3). Over 90.7% of the assembled sequences were successfully anchored to the 10 maize chromosomes.
Heatmap of Hi-C sequencing data showing genome-wide interactions in Yu82. Interactions are displayed between 500-kb non-overlapping windows, with interaction intensity represented by the color scale.
Annotation of transposable elements
Transposable elements (TEs) were annotated using RepeatMasker (http://www.repeatmasker.org/, version 1.323) with the recommended parameters: “-pa 30 -no_is -norna -nolow -div 40 -cutoff 225”. The annotation was based on the curated TE library from the Maize TE Consortium (MTEC), originally created on October 10, 2014, and last updated on November 12, 201955. Overall, a total of 1,135,580 and 1,154,679 transposable elements were identified in the genomes of D132 and Yu82, representing 81.46% and 82.59% of their genome sizes, respectively (Tables 3, 5, 6). The composition and abundance of TEs in D132 and Yu82 were comparable to those in other maize genomes, such as B73 and Mo1756,57.
Annotation of protein-coding genes
To annotate protein-coding genes, an integrated approach combining de novo prediction, protein-based homology searches, and transcript assembly using RNA-Seq and Iso-Seq data was applied. Four tools were used for ab initio prediction: SNAP (version 38926, default parameters), Augustus (version 3.3, parameter “–genemodel = partial”), GlimmerHmm (version 3.04, default parameters), and GeneMark (version 4.33, default parameters)58,59,60,61. For protein homology-based predictions, GeneWise (version 2.2.0, default parameters) was employed to align protein sequences from rice, sorghum, millet, Brachypodium distachyon, and maize B73 to the genome assemblies of D132 and Yu8242,62,63,64,65,66. For transcript-based evidence, we utilized Trinity67 (version v2.13.1, default parameters) to de novo assemble contigs from two sources: (1) our in-house RNA sequencing data from each cultivar, and (2) 172 additional public RNA-Seq datasets downloaded to enhance transcript coverage. These assembled contigs, along with full-length transcripts obtained from our Iso-Seq, were subsequently aligned to the genome assemblies using PASA68 (version 2.1, parameter “-m = 50”) to model transcript-based gene structures. Finally, gene predictions from these different approaches were integrated using EvidenceModeler69 (version 1.11, default parameters). The resulting gene models were further refined with PASA to update and finalize the annotation.
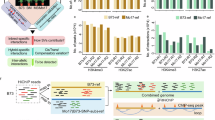
A total of 40,951 and 40,935 protein-coding genes were annotated in the genomes of D132 and Yu82, respectively (Table 3). The average coding sequence (CDS) lengths were 1226.48 bp for D132 and 1223.57 bp for Yu82. Overall, these key statistics (number of genes, average CDS length, average exon/intron length, and exon number) are consistent with those reported for other maize genomes, such as B73, SK, and W22 (Table 3, Fig. 5)42,54,70.
Comparative analysis of annotated gene models in D132, Yu82, and five previously reported maize genome assemblies. The comparison includes B73, Mo17, CML247, SK, and W22, highlighting differences in annotated gene model statistics.
Functional annotation of protein-coding genes
We employed Trinotate to functionally annotate the protein-coding genes predicted for D132 and Yu8271. Trinotate integrates multiple annotation strategies, including homology searches against well-established sequence databases (UniProt), identification of protein domains via PFAM, and functional mapping to resources such as eggNOG and Gene Ontology (GO). The coding sequences (CDS) of protein-coding genes were aligned to the UniProt database using BLASTX. Likewise, protein sequences were aligned to the UniProt database using BLASTP. Alignments with E-values higher than 1e-05 were excluded. In total, 39,418 (96.26%) and 39,513 (96.53%) protein-coding genes in D132 and Yu82, respectively, were successfully annotated by at least one method in Trinotate (Table 7). These results further highlight the high quality of genome annotations for both inbred lines.
Genome synteny analysis
We conducted a genome synteny analysis to investigate the collinearity between the assembled genomes of D132 and Yu82, and four previously published maize genomes: B73, Mo17, W22, and SK42,54,57,70. The analysis was performed by aligning the genome assemblies using MUMMER (https://github.com/mummer4/mummer, version 4.0)72. The nucmer program was used to align the reference genome (D132 or Yu82) against each query genome (B73, Mo17, W22, or SK), with the following parameter: “-L 2000 -l 80–mum -t 50 -g 200 -d 0.08 -b 100”. The resulting alignments were encoded in a delta file, which was then filtered by the delta-filter program with parameter: “-l 5000 -i 99.5 -u 100”. The filtered delta file was visualized as a two-dimensional dot plot using mummerplot.
Gene collinearity analysis
We next investigated gene collinearity between D132 and four previously reported maize genomes—B73, Mo17, SK, and W22—using MCScan (Materials and methods)73. A parallel analysis was also performed for Yu82. The Python version of MCScan (https://github.com/tanghaibao/jcvi/wiki/MCscan-(Python-version)) was used for all analyses. Taking B73 and D132 as an example, genome annotations were first converted into BED format using the command: “python -m jcvi.formats.gff bed --type = mRNA --key = Name”. Similarly, protein sequences from both genomes were formatted using the command: “python -m jcvi.formats.fasta format”. Orthologous gene pairs and synteny blocks were then identified with the following command: “python -m jcvi.compara.catalog ortholog --dbtype prot B73 D132 --no_strip_names”. To retain high-confidence blocks, synteny regions were filtered using: “python -m jcvi.compara.synteny screen --minspan = 30 --simple”. Finally, pairwise synteny between the reference and query genomes was visualized with the command: “python -m jcvi.graphics.karyotype seqids layout”. Here, the seqids file defines the chromosome names of the reference and query genomes, while the layout file specifies the configuration of the plot.
Data Records
The genome assemblies and annotations of D132 and Yu82 are available in NCBI GenBank and the Genome Warehouse of the National Genomics Data Center (NGDC, https://ngdc.cncb.ac.cn/) with the following accession codes: D132, GCA_044509805.174 (NCBI) or GWHETKL00000000.275 (NGDC); Yu82, GCA_042768475.176 (NCBI) or GWHETKM00000000.277 (NGDC). The genome assembly and annotation of D132 and Yu82 are also deposited at FigShare78 (https://doi.org/10.6084/m9.figshare.30595493.v1).
The RNA sequencing data, Illumina short read sequencing data, Hi-C data, PacBio long read sequencing data, and Iso-Seq data are deposited in NCBI Sequence Read Archive and NGDC Genome Sequence Archive with the following accession codes: RNA sequencing data of D132 and Yu82, SRP53762935 (NCBI) or CRA01716136 (NGDC); Illumina short read sequencing data of D132, SRP57787725 (NCBI) or CRA02714226 (NGDC); Illumina short read sequencing data of Yu82, SRP57806829 (NCBI) or CRA02720130 (NGDC); Hi-C data of D132, SRP14265579 (NCBI) or CRA02706446 (NGDC); Hi-C data of Yu82, SRP57918533 (NCBI) or CRA02713934 (NGDC); PacBio long read sequencing of D132, SRP59182527 (NCBI) or CRA02681343 (NGDC); PacBio long read sequencing of Yu82, SRP58017831 (NCBI) or CRA02717932 (NGDC); Iso-Seq data of D132, SRP59211337 (NCBI) or CRA02681538 (NGDC); and Iso-Seq data of Yu82, SRP59211139 (NCBI) or CRA02681740 (NGDC).
Technical Validation
Quality control of nucleic acids and libraries
To ensure the high quality of extracted DNA, we first evaluated its integrity using the Agilent 4200 Bioanalyzer (Agilent Technologies, CA, USA). DNA purity was then assessed with a Nanodrop spectrophotometer, ensuring OD260/280 ratios between 1.8 and 2.0, and OD260/230 ratios between 2.0 and 2.2. Finally, DNA quantity was measured using a Qubit fluorometer to confirm that sufficient amounts were available for downstream applications.
For RNA quality control, we initially used 1% agarose gel electrophoresis to check for degradation or contamination. The purity of RNA was subsequently measured using the K5500 spectrophotometer (Kaiao Technology, Beijing, China). RNA integrity and concentration were further evaluated using the Agilent 2100 system with the RNA Nano 6000 Assay Kit (Agilent Technologies, CA, USA).
Quality control of Illumina sequencing data
For the Illumina sequencing data of D132 and Yu82, adaptor sequences were first removed from the raw reads. Subsequently, reads containing more than 50% low-quality bases (Phred quality score ≤ 19) or more than 5% ambiguous “N” bases were discarded. If one end of a paired-end read was removed during this filtering process, the entire read pair was also discarded.
Assessment of genome assembly
To assess the quality of both genome assemblies, we aligned over 3,200 BUSCO genes (liliopsida_odb10) to the assembled genomes80. Approximately 92.99% and 94.08% BUSCO genes were completely recovered in the genome assemblies of D132 and Yu82, respectively (Table 8). Assembly quality was further evaluated using the LTR Assembly Index (LAI), which estimates assembly continuity based on LTR retrotransposons56,81. The LAI score of D132 was slightly lower than that of B73 (version 5) but higher than those of Yu82 and B73 (version 4) (Fig. 6). In addition, we assessed base-level accuracy and completeness using two methods. First, more than 99.3% and 99.7% of the Illumina sequencing reads could be aligned to the genome assemblies of D132 and Yu82, respectively, without mismatches. Second, k-mer-based consensus quality was evaluated using Merqury82. The D132 assembly showed an excellent consensus QV of 60.02 and 95.45% k-mer completeness, while the Yu82 assembly showed a QV of 35.62 and 97.01% k-mer completeness. These results collectively indicate that the genome assemblies of D132 and Yu82 have achieved chromosome-level contiguity and high base-level accuracy.
Assessment of genome assembly quality using the LTR Assembly Index (LAI). The LAI scores for B73 v4 (A), B73 v5 (B), D132 (C), and Yu82 (D) are shown. The mean and standard deviation of LAI scores across the genome are indicated at the top left of each plot.
Synteny and collinearity assessment
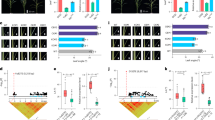
To further validate the large-scale structural accuracy and gene order of our assemblies, we compared them against four established maize reference genomes (B73, Mo17, W22, and SK). Whole-genome alignments revealed a high degree of synteny for both D132 and Yu82 against all references (Figs. 7, 8). This provides strong validation for the correct large-scale orientation and ordering of our assembled pseudomolecules.
Synteny analysis between the genome assembly of D132 and four previously reported maize genome assemblies. The analysis compares D132 with B73, Mo17, SK, and W22. Each dot represents an aligned genomic region longer than 5 kb. The Y-axis corresponds to the 10 chromosomes of D132, while the X-axes of the four plots correspond to the 10 chromosomes of B73, Mo17, SK, and W22, respectively.
Synteny analysis between the genome assembly of Yu82 and four previously reported maize genome assemblies. The analysis compares Yu82 with B73, Mo17, SK, and W22. Each dot represents an aligned genomic region longer than 5 kb. The X-axis corresponds to the 10 chromosomes of Yu82, while the Y-axes of the four plots correspond to the 10 chromosomes of B73, Mo17, SK, and W22, respectively.
Furthermore, we assessed gene-level collinearity. This analysis confirmed the high quality of the assemblies at the genic level, identifying more than 33,827 collinear gene pairs for D132 and 30,594 for Yu82 when compared to the reference genomes (Figs. 9, 10). The high degree of conserved synteny and gene collinearity thus serves as a critical technical validation, demonstrating that our assemblies accurately reflect the established structure of the maize genome.
Collinearity of protein-coding genes between D132 and four previously reported maize genome assemblies. The comparison involves B73, Mo17, SK, and W22 across 10 chromosomes. Each dot represents a collinear gene pair. The X-axis corresponds to the 10 chromosomes of D132, while the Y-axes of the four plots correspond to the 10 chromosomes of B73, Mo17, SK, and W22, respectively.
Collinearity of protein-coding genes between Yu82 and four previously reported maize genome assemblies. The comparison involves B73, Mo17, SK, and W22 across 10 chromosomes. Each dot represents a collinear gene pair. The X-axis corresponds to the 10 chromosomes of Yu82, while the Y-axes of the four plots correspond to the 10 chromosomes of B73, Mo17, SK, and W22, respectively.
Gene annotation validation
The completeness of the genome annotations was assessed using a BUSCO analysis against a set of 4,896 conserved genes. The results revealed that 95.12% and 93.57% of the BUSCO genes were complete in the genome annotations of D132 and Yu82, respectively (Table 9).
Code availability
In this study, no custom code was developed. All software and pipelines used for data processing and analysis are publicly available and were utilized according to the developers’ instructions, as described in the cited references. Default parameters were used as suggested in the references if no detailed parameters are mentioned for a software.
References
Yan, P. et al. Biofortification of iron content by regulating a NAC transcription factor in maize. Science 382, 1159–1165, https://doi.org/10.1126/science.adf3256 (2023).
Tilman, D., Balzer, C., Hill, J. & Befort, B. L. Global food demand and the sustainable intensification of agriculture. Proc Natl Acad Sci USA 108, 20260–20264, https://doi.org/10.1073/pnas.1116437108 (2011).
Tian, J. et al. Teosinte ligule allele narrows plant architecture and enhances high-density maize yields. Science 365, 658–664, https://doi.org/10.1126/science.aax5482 (2019).
Wang, B. et al. Genome-wide selection and genetic improvement during modern maize breeding. Nature Genetics 52, 565–571, https://doi.org/10.1038/s41588-020-0616-3 (2020).
Duvick, D. Genetic progress in yield of United States maize (Zea mays L.). Maydica 50, 193–202 (2005).
Lauer, S. et al. Morphological Changes in Parental Lines of Pioneer Brand Maize Hybrids in the U.S. Central Corn Belt. Crop Science 52, 1033–1043, https://doi.org/10.2135/cropsci2011.05.0274 (2012).
Luo, N. et al. China can be self-sufficient in maize production by 2030 with optimal crop management. Nature Communications 14, 2637, https://doi.org/10.1038/s41467-023-38355-2 (2023).
Mansfield, B. D. & Mumm, R. H. Survey of plant density tolerance in U.S. maize germplasm. Crop Science 54, 157–173, https://doi.org/10.2135/cropsci2013.04.0252 (2014).
Duan, H. et al. Genetic dissection of internode length confers improvement for ideal plant architecture in maize. The Plant Journal 121, e17245, https://doi.org/10.1111/tpj.17245 (2025).
Khush, G. S. Green revolution: the way forward. Nature Reviews Genetics 2, 815–822, https://doi.org/10.1038/35093585 (2001).
Peng, J. et al. Green revolution’ genes encode mutant gibberellin response modulators. Nature 400, 256–261, https://doi.org/10.1038/22307 (1999).
Sasaki, A. et al. A mutant gibberellin-synthesis gene in rice. Nature 416, 701–702, https://doi.org/10.1038/416701a (2002).
Zhao, S. et al. Genetic dissection of maize plant architecture using a novel nested association mapping population. Plant Genome 15, e20179, https://doi.org/10.1002/tpg2.20179 (2022).
Jafari, F., Wang, B., Wang, H. & Zou, J. Breeding maize of ideal plant architecture for high-density planting tolerance through modulating shade avoidance response and beyond. Journal of Integrative Plant Biology 66, 849–864, https://doi.org/10.1111/jipb.13603 (2024).
Mock, J. J. & Pearce, R. B. An ideotype of maize. Euphytica 24, 613–623, https://doi.org/10.1007/BF00132898 (1975).
Yan, Y. et al. Photosynthetic capacity and assimilate transport of the lower canopy influence maize yield under high planting density. Plant Physiology 195, 2652–2667, https://doi.org/10.1093/plphys/kiae204 (2024).
Ren, W. et al. Genome-wide dissection of changes in maize root system architecture during modern breeding. Nature Plants 8, 1408–1422, https://doi.org/10.1038/s41477-022-01274-z (2022).
Zhang, J. et al. The ZmCLA4 gene in the qLA4-1 QTL controls leaf angle in maize (Zea mays L. Journal of Experimental Botany 65, 5063–5076, https://doi.org/10.1093/jxb/eru271 (2014).
Ren, Z. et al. ZmILI1 regulates leaf angle by directly affecting liguleless1 expression in maize. Plant Biotechnology Journal 18, 881–883, https://doi.org/10.1111/pbi.13255 (2020).
Wei, X. et al. Epistatic and QTL × environment interaction effects on leaf area-associated traits in maize. Plant Breeding 135, 671–676, https://doi.org/10.1111/pbr.12422 (2016).
Dou, D. et al. CLA4 regulates leaf angle through multiple hormone signaling pathways in maize. Journal of Experimental Botany 72, 1782–1794, https://doi.org/10.1093/jxb/eraa565 (2021).
Cao, Y. et al. ZmDWF1 regulates leaf angle in maize. Plant Science 325, 111459, https://doi.org/10.1016/j.plantsci.2022.111459 (2022).
Cao, Y. et al. ZmIBH1-1 regulates plant architecture in maize. Journal of Experimental Botany 71, 2943–2955, https://doi.org/10.1093/jxb/eraa052 (2020).
Van Nieuwerburgh, F. et al. Illumina mate-paired DNA sequencing-library preparation using Cre-Lox recombination. Nucleic Acids Research 40, e24, https://doi.org/10.1093/nar/gkr1000 (2012).
NCBI Sequence Read Archive https://identifiers.org/insdc.sra:SRP577877 (2025).
NGDC Genome Sequence Archive https://ngdc.cncb.ac.cn/gsa/browse/CRA027142 (2025).
NCBI Sequence Read Archive https://identifiers.org/insdc.sra:SRP591825 (2025).
Tian, L. et al. Large-scale reconstruction of chromatin structures of maize temperate and tropical inbred lines. Journal of Experimental Botany 72, 3582–3596, https://doi.org/10.1093/jxb/erab087 (2021).
NCBI Sequence Read Archive https://identifiers.org/insdc.sra:SRP578068 (2025).
NGDC Genome Sequence Archive https://ngdc.cncb.ac.cn/gsa/browse/CRA027201 (2025).
NCBI Sequence Read Archive https://identifiers.org/insdc.sra:SRP580178 (2025).
NGDC Genome Sequence Archive https://ngdc.cncb.ac.cn/gsa/browse/CRA027179 (2025).
NCBI Sequence Read Archive https://identifiers.org/insdc.sra:SRP579185 (2025).
NGDC Genome Sequence Archive https://ngdc.cncb.ac.cn/gsa/browse/CRA027139 (2025).
NCBI Sequence Read Archive https://identifiers.org/insdc.sra:SRP537629 (2025).
NGDC Genome Sequence Archive https://ngdc.cncb.ac.cn/gsa/browse/CRA017161 (2025).
NCBI Sequence Read Archive https://identifiers.org/insdc.sra:SRP592113 (2025).
NGDC Genome Sequence Archive https://ngdc.cncb.ac.cn/gsa/browse/CRA026815 (2025).
NCBI Sequence Read Archive https://identifiers.org/insdc.sra:SRP592111 (2025).
NGDC Genome Sequence Archive https://ngdc.cncb.ac.cn/gsa/browse/CRA026817 (2025).
Marçais, G. & Kingsford, C. A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics 27, 764–770, https://doi.org/10.1093/bioinformatics/btr011 (2011).
Jiao, Y. et al. Improved maize reference genome with single-molecule technologies. Nature 546, 524, https://doi.org/10.1038/nature22971 (2017).
NGDC Genome Sequence Archive https://ngdc.cncb.ac.cn/gsa/browse/CRA026813 (2025).
Cheng, H., Concepcion, G. T., Feng, X., Zhang, H. & Li, H. Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nature Methods 18, 170–175, https://doi.org/10.1038/s41592-020-01056-5 (2021).
Roach, M. J., Schmidt, S. A. & Borneman, A. R. Purge Haplotigs: allelic contig reassignment for third-gen diploid genome assemblies. BMC Bioinformatics 19, 460, https://doi.org/10.1186/s12859-018-2485-7 (2018).
NGDC Genome Sequence Archive https://ngdc.cncb.ac.cn/gsa/browse/CRA027064 (2025).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nature Methods 9, 357–359, https://doi.org/10.1038/nmeth.1923 (2012).
Servant, N. et al. HiC-Pro: an optimized and flexible pipeline for Hi-C data processing. Genome Biology 16, 259, https://doi.org/10.1186/s13059-015-0831-x (2015).
Burton, J. N. et al. Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions. Nat Biotechnol 31, 1119–1125, https://doi.org/10.1038/nbt.2727 (2013).
Wang, Y. et al. shinyCircos-V2.0: Leveraging the creation of Circos plot with enhanced usability and advanced features. Imeta 2, e109, https://doi.org/10.1002/imt2.109 (2023).
Durand, N. C. et al. Juicebox Provides a Visualization System for Hi-C Contact Maps with Unlimited Zoom. Cell Systems 3, 99–101, https://doi.org/10.1016/j.cels.2015.07.012 (2016).
Chakraborty, M., Baldwin-Brown, J. G., Long, A. D. & Emerson, J. J. Contiguous and accurate de novo assembly of metazoan genomes with modest long read coverage. Nucleic Acids Research 44, e147 https://doi.org/10.1093/nar/gkw654 (2016).
Walker, B. J. et al. Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 9, e112963, https://doi.org/10.1371/journal.pone.0112963 (2014).
Yang, N. et al. Genome assembly of a tropical maize inbred line provides insights into structural variation and crop improvement. Nature Genetics 51, 1052–1059, https://doi.org/10.1038/s41588-019-0427-6 (2019).
Ou, S. et al. Benchmarking transposable element annotation methods for creation of a streamlined, comprehensive pipeline. Genome Biology 20, 275, https://doi.org/10.1186/s13059-019-1905-y (2019).
Hufford, M. B. et al. De novo assembly, annotation, and comparative analysis of 26 diverse maize genomes. Science 373, 655–662, https://doi.org/10.1126/science.abg5289 (2021).
Sun, S. et al. Extensive intraspecific gene order and gene structural variations between Mo17 and other maize genomes. Nature Genetics 50, 1289–1295, https://doi.org/10.1038/s41588-018-0182-0 (2018).
Korf, I. Gene finding in novel genomes. BMC Bioinformatics 5, 59, https://doi.org/10.1186/1471-2105-5-59 (2004).
Keller, O., Kollmar, M., Stanke, M. & Waack, S. A novel hybrid gene prediction method employing protein multiple sequence alignments. Bioinformatics 27, 757–763, https://doi.org/10.1093/bioinformatics/btr010 (2011).
Majoros, W. H., Pertea, M. & Salzberg, S. L. TigrScan and GlimmerHMM: two open source ab initio eukaryotic gene-finders. Bioinformatics 20, 2878–2879, https://doi.org/10.1093/bioinformatics/bth315 (2004).
Lomsadze, A., Ter-Hovhannisyan, V., Chernoff, Y. O. & Borodovsky, M. Gene identification in novel eukaryotic genomes by self-training algorithm. Nucleic Acids Research 33, 6494–6506, https://doi.org/10.1093/nar/gki937 (2005).
Birney, E., Clamp, M. & Durbin, R. GeneWise and Genomewise. Genome Research 14, 988–995, https://doi.org/10.1101/gr.1865504 (2004).
Kawahara, Y. et al. Improvement of the Oryza sativa Nipponbare reference genome using next generation sequence and optical map data. Rice 6, 4, https://doi.org/10.1186/1939-8433-6-4 (2013).
Vogel, J. P. et al. Genome sequencing and analysis of the model grass Brachypodium distachyon. Nature 463, 763–768, https://doi.org/10.1038/nature08747 (2010).
Paterson, A. H. et al. The Sorghum bicolor genome and the diversification of grasses. Nature 457, 551–556, https://doi.org/10.1038/nature07723 (2009).
Varshney, R. K. et al. Pearl millet genome sequence provides a resource to improve agronomic traits in arid environments. Nat Biotechnol 35, 969–976, https://doi.org/10.1038/nbt.3943 (2017).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol 29, 644–652, https://doi.org/10.1038/nbt.1883 (2011).
Haas, B. J. et al. Improving the Arabidopsis genome annotation using maximal transcript alignment assemblies. Nucleic Acids Research 31, 5654–5666, https://doi.org/10.1093/nar/gkg770 (2003).
Haas, B. J. et al. Automated eukaryotic gene structure annotation using EVidenceModeler and the Program to Assemble Spliced Alignments. Genome Biology 9, R7, https://doi.org/10.1186/gb-2008-9-1-r7 (2008).
Springer, N. M. et al. The maize W22 genome provides a foundation for functional genomics and transposon biology. Nature Genetics 50, 1282–1288, https://doi.org/10.1038/s41588-018-0158-0 (2018).
Bryant, D. M. et al. A Tissue-Mapped Axolotl De Novo Transcriptome Enables Identification of Limb Regeneration Factors. Cell Rep 18, 762–776, https://doi.org/10.1016/j.celrep.2016.12.063 (2017).
Marçais, G. et al. MUMmer4: A fast and versatile genome alignment system. PLOS Computational Biology 14, e1005944, https://doi.org/10.1371/journal.pcbi.1005944 (2018).
Tang, H. et al. Unraveling ancient hexaploidy through multiply-aligned angiosperm gene maps. Genome Research 18, 1944–1954, https://doi.org/10.1101/gr.080978.108 (2008).
Yao, W. et al. Chromosome-level genome assemblies of two maize inbred lines with contrasting plant architectures. NCBI GenBank https://identifiers.org/ncbi/insdc:JBITNO000000000 (2025).
NGDC Genome Warehouse https://ngdc.cncb.ac.cn/gwh/Assembly/102346/show (2025).
Yao, W. et al. Chromosome-level genome assemblies of two maize inbred lines with contrasting plant architectures. NCBI GenBank https://identifiers.org/ncbi/insdc:JBHUPN000000000 (2025).
NGDC Genome Warehouse https://ngdc.cncb.ac.cn/gwh/Assembly/102345/show (2025).
Yao, W. et al. Chromosome-level genome assemblies of two maize inbred lines with contrasting plant architectures. FigShare https://doi.org/10.6084/m9.figshare.30595493.v1 (2025).
NCBI Sequence Read Archive https://identifiers.org/insdc.sra:SRP142655 (2025).
Manni, M., Berkeley, M. R., Seppey, M. & Zdobnov, E. M. BUSCO: Assessing Genomic Data Quality and Beyond. Current Protocols 1, e323, https://doi.org/10.1002/cpz1.323 (2021).
Ou, S., Chen, J. & Jiang, N. Assessing genome assembly quality using the LTR Assembly Index (LAI). Nucleic Acids Research 46, e126–e126, https://doi.org/10.1093/nar/gky730 (2018).
Rhie, A., Walenz, B. P., Koren, S. & Phillippy, A. M. Merqury: reference-free quality, completeness, and phasing assessment for genome assemblies. Genome Biology 21, 245, https://doi.org/10.1186/s13059-020-02134-9 (2020).
Acknowledgements
This research was supported by the National Key Research and Development Program of China (No. 2021YFF1000301), the National Natural Science Foundation of China (No. 32201852), the Henan Agricultural (Maize) Improved Variety Joint Tackling Project (No. 2022010203), and the Natural Science Foundation of Henan Province (No. 242300420155 and 252300420196).
Author information
Authors and Affiliations
Contributions
L.K., Z.R. and W.Y. conceived and designed the overall study. W.Y., S.L. and Y.W. carried out genome assembly and annotation. W.Y., S.L., Z.R., L.K., J.R., J.S., F.W., S.Z., H.S. and Y.W. analyzed the data. J.R., J.S., F.W., S.Z., and H.S. performed the molecular experiments. Z.R. and Y.C. contributed to fieldwork. W.Y. and L.K. wrote the manuscript, with input from all authors. W.Y., L.K., Z.R. and Y.C. revised and finalized the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yao, W., Li, S., Ren, J. et al. Chromosome-level genome assemblies of two maize inbred lines with contrasting plant architectures. Sci Data 13, 276 (2026). https://doi.org/10.1038/s41597-026-06603-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-026-06603-x