Abstract
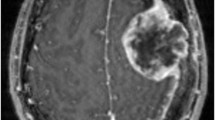
Meningiomas are the most common primary intracranial tumors, frequently requiring radiotherapy as a part of management. Effective radiotherapy planning for meningiomas necessitates accurate and consistent segmentation of target volumes on MRI, a process that is complex, labor-intensive, and dependent on expert expertise. The 2024 Brain Tumor Segmentation Challenge Meningioma Radiotherapy (BraTS-MEN-RT) Dataset addresses this problem by providing the largest multi-institutional collection of systematically annotated radiotherapy planning MRIs for meningiomas. Publicly accessible, this dataset comprises 570 radiotherapy planning 3D T1-weighted post-contrast MRIs at native resolutions, with 500 cases featuring expert-annotated gross tumor volumes (GTV). Annotations follow standardized radiotherapy planning protocols and include both intact and postoperative meningioma cases, ensuring wide clinical relevance. Contributions from seven diverse medical centers across the United States and the United Kingdom enhance the dataset’s generalizability. The dataset aims to accelerate the development of automated segmentation methods for radiotherapy planning, improving workflow efficiency, reducing interobserver variability, and ultimately enhancing patient outcomes.
Similar content being viewed by others
Background & Summary
Meningiomas are the most common intracranial tumors, encompassing approximately 37% of all primary central nervous system (CNS) neoplasms and up to 55% of non-malignant CNS tumors1,2,3. While the majority of these tumors (commonly WHO grade 1) are benign, higher-grade meningiomas (WHO grades 2 and 3) carry a significantly higher risk of recurrence and morbidity4,5. Management strategies for symptomatic meningioma generally involve maximal surgical resection when anatomically feasible, often followed by radiotherapy (RT) for higher-grade or incompletely resected disease6,7,8.
Despite wide adoption of RT in both the intact (unresected) and postoperative contexts, the process of defining the radiation target volume—namely the gross tumor volume (GTV) and surrounding tissues at risk—remains labor-intensive and highly reliant on specialized expertise. Established guidelines, such as those from EORTC 22042–026042 and RTOG 0539, detail the delineation of meningioma tumor beds on contrast-enhanced T1-weighted magnetic resonance tomography (MRI) for primary or adjuvant therapy9,10,11. However, segmenting meningiomas in real-world RT workflows is challenging: postoperative cavities may exhibit complex morphologies, postoperative changes, and artifact from adjacent hardware. In addition, some patients receive stereotactic radiosurgery (SRS) with dedicated headframes that can result in additional image artifacts12,13.
Although deep learning-based segmentation methods have seen considerable success for preoperative brain tumor imaging, relatively few solutions have targeted the unique requirements of RT planning14,15,16,17,18,19. Prior efforts—such as earlier Brain Tumor Segmentation (BraTS) challenges—commonly emphasize skull-stripped, isotropically resampled images to a standard anatomical space, along with multiple MRI sequences19,20. While valuable for diagnostic and predictive modeling purposes, these pre-processing steps can limit the direct clinical utility for RT, where native spatial resolution and inclusion of extracranial anatomy (e.g., stereotactic localizer fiducials, fixation devices) are essential21,22,23,24.
Herein, we introduce the 2024 Brain Tumor Segmentation Challenge Meningioma Radiotherapy (BraTS-MEN-RT) Dataset, a multi-institutional, expert-annotated resource encompassing 570 post-contrast T1-weighted (T1c) RT planning MRI scans for intact or postoperative meningiomas. Each study remains in its original resolution and orientation, with only defacing performed to preserve patient anonymity while retaining important extracranial structures25,26,27,28. A subset of 500 cases include corresponding GTV segmentations encompassing all meningioma lesions or at-risk resection cavities, harmonized according to established meningioma RT protocols9,10,11. This dataset is designed to facilitate the creation of clinically informed models that can effectively navigate the challenges posed by variability in image acquisition, SRS-specific artifacts, and postoperative imaging complexity.
In the sections that follow, we detail the data collection and annotation pipelines, describe robust quality assurance measures, and summarize the results from the BraTS-MEN-RT Dataset. By making this dataset freely available, we aim to accelerate the integration of automated segmentation algorithms into actual radiotherapy workflows, thereby improving the speed, consistency, and objectivity of meningioma RT planning.
Methods
Study population
The BraTS-MEN-RT Dataset comprises 3D T1c brain MRI scans obtained for clinical RT planning of meningiomas, including both intact and postoperative cases. Seven academic medical centers across the United States and the United Kingdom contributed. Participating institutions included Duke University, University of California San Francisco, State University of New York Upstate Medical University, University of Washington, University of Missouri, King’s College London, and University of California San Diego. Each institution retrospectively identified meningioma patients who underwent any modality of RT including Cobalt-60, external beam radiation therapy with photons, SRS, or proton therapy. Case inclusion criteria, whether diagnosis was made based on pathological, clinical, or radiological evidence, as well as case collection methods (e.g., random or consecutive sampling), were determined independently by each site, often leveraging pre-existing curated datasets developed for other purposes. For patients that underwent multiple courses of RT, each of their respective radiotherapy planning 3D T1c brain MRI scans could be included as a separate BraTS-MEN-RT case.
All centers contributing publicly accessible data adhered to their respective institutional review board (IRB) protocols ensuring compliance with ethical standards for research involving human subjects (DUKE IRB approval number: Pro00110695; MISS IRB approval number: 2096253 MU; SUNY Upstate IRB approval number: 2183481-1; UCSD IRB approval number: 809620; UCSF IRB approval number: 18-24633; UW Human Subjects Division IRB approval number: STUDY00020442). A waiver for informed consent was provided by each institution’s respective IRB. To protect patient confidentiality, all metadata within Digital Imaging and Communications in Medicine (DICOM) files were de-identified by each participating institution and reviewed to ensure that all patient identifiers were removed or anonymized before data release.
Imaging data
The imaging dataset includes exclusively 3D T1c brain MRI scans in native acquisition resolution in either the intact (Fig. 1) or postoperative setting (Fig. 2, 3), which mimics the data available for most radiotherapy planning. While additional sequences such as T1-weighted, T2-weighted, T2-FLAIR, and computed tomography are sometimes used in clinical workflows, these were not consistently available nor required for RT planning at all centers and were therefore not included in the dataset.
Panels A, B, and C depict two distinct intact meningiomas (red) on axial, sagittal, and coronal images respectively.
Panels A, B, and C depict a left anterior falcine meningioma (red) on axial, sagittal, and coronal images respectively. Panel B demonstrates an area of hypointense encephalomalacia between two separate anterior falcine meningioma components. Panels B and C highlight the defacing process, which removes pixels in and around the face to eliminate potentially identifying facial features.
Panels A, B, and C showing a postoperative left parietal meningioma target volume (red) in the axial, sagittal, and coronal planes, respectively, as delineated by the treating institution.
Participating sites exported physician-delineated gross tumor volumes (GTVs) from their available DICOM-RT structure sets when present. Imaging parameters including vendor, scanner model, magnetic field strength, series description, sequence identifiers (sequence name, scanning sequence, sequence variant), repetition and echo times (TR, TE), inversion time (TI), flip angle, in-plane resolution (pixel spacing), slice thickness, acquisition and reconstruction matrices, pixel bandwidth, and coil name; varied substantially both within and across contributing institutions. Complete parameter metadata were available for a subset of cases, reflecting the diversity of acquisition protocols in clinical practice. To minimize barriers to data sharing and encourage participation, documentation of these imaging parameters was not mandated. This variability and partial availability of detailed imaging parameters underscore the dataset’s realism and enhance its value for developing and benchmarking automated segmentation models that must generalize across heterogeneous imaging conditions.
Data splits
A total of 750 radiotherapy planning MRI exams from 747 unique patients were included in the BraTS-MEN-RT Dataset. These were divided into a training set (500/750, 66.7%), a validation set (70/750, 9.3%), and a private testing set (180/750, 24%). As feasible, the data splits were stratified by site to ensure balanced representation and consistency. In cases where there were multiple exams from a single patient, all such exams were included in only a single split. The University of Missouri data were included in the validation and testing set only, and the King’s College London data were included in the testing set only, due to the timing of data availability relative to the BraTS-MEN-RT Dataset release dates29. The 570 radiotherapy planning 3D T1c MRIs from the training and validation datasets, and expert-annotated GTV from the 500 training dataset cases are publicly available as part of this manuscript. The 180 cases from the testing dataset remain private to facilitate unbiased evaluation of new automated segmentation methods.
Clinical data
Clinical-pathologic information, including patient age at the time of imaging, sex, and WHO tumor grade, was obtained from the respective electronic medical records at each contributing institution when possible. The age range was 11–90 years (including private testing set data) and 13–90 years in the publicly available data. The male to female ratio was 229:482 (including private testing set data) and 181:360 in the publicly available data. WHO grade was available in 342 of the 750 total cases (including private testing set data) and in 271 of the 570 publicly available cases. For training and validation cases, image x-resolutions and y-resolutions ranged from 0.338 to 1.055 mm, and slice thickness ranged from 0.488 to 2.000 mm, as shown in Fig. 4. Aggregate clinical and demographic data for the dataset are summarized in Table 1, including training, validation, and private testing data splits. Individual case-level data for the publicly available training and validation cases are publicly available on the Synapse data repository29.
Violin plot depicting the distribution of T1c image resolution parameters (x-resolution, y-resolution, and slice thickness) from the BraTS-MEN-RT Dataset. Each violin shape represents the density distribution of the resolution measurements, with wider sections indicating higher frequency of measurements at specific resolution values.
Image data pre-processing
Following anonymized DICOM-RT data collection from each respective institution, the raw DICOM-RT data were converted to Neuroimaging Informatics Technology Initiative (NIfTI) format using open-source software30. This process ensured that each MRI scan was accurately captured in its native spatial orientation and voxel resolution without any intensity resampling. To preserve extracranial anatomy and anatomical structures relevant to RT (e.g., stereotactic headframes, fixation devices), no skull-stripping was applied22,23,24. Instead, patient facial features were removed using the Analysis of Functional NeuroImages (AFNI) automated defacing algorithm with AFNI’s default parameters26,27,28. The AFNI defacing process eliminates patient-identifiable facial structures while retaining the rest of the cranial and extracranial volume as shown in Figs. 2C, 5.
Example of a sagittal image of a brain MRI from the BraTS-MEN-RT training set, both before (A) and after (B) automated defacing.
After defacing was performed, a robust quality-control procedure was employed, whereby each converted and defaced MRI volume underwent manual slice-by-slice review using ITK-SNAP to confirm the completeness of the intracranial volume and ascertain that no meningioma tissue was inadvertently removed31. Cases with meningiomas extending beyond the region of the face, such as anterior skull base tumors, were closely inspected to ensure no tumor signal was truncated. If partial tumor removal was detected, manual post-processing was performed to add back intensity to clinically appropriate brain MRI voxels corresponding to tumor (when partially removed) or exclude the case (if fully removed). Figure 6 demonstrates the partial inclusion of an institutional GTV on the defaced MRI, and Fig. 7 demonstrates a case of a patient that underwent SRS to an intact meningioma in Meckel’s cave that would have been at least partially excluded by a skull-stripping approach.
Example of an axial MRI image-label pair slice where the treating institution’s meningioma GTV extended outside of the defaced image. Note that this left sphenoid meningioma involves the skull base and extents extracranially into the left masticator space. This case was ultimately excluded from the challenge dataset due to the excessive volume within the anonymized region that would have needed to be reintroduced.
Image panels depicting a case that utilizes a SRS planning Gamma Knife headframe. Panels A, B, and C depict an intact meningioma (red) in the right Meckel’s cave on T1c axial, sagittal, and coronal images, respectively. Note that this challenge’s defacing technique preserves this meningioma as compared to a skull-stripping pre-processing technique which would have excluded at least part of this lesion. Panel D shows the SRS localizer box fiducials attached to a standard Gamma Knife headframe.
Target volume definition
For each case, the binary segmentation consisted of a GTV segmentation corresponding to intact and/or postoperative meningioma. GTVs included areas of high-risk disease composed of any enhancing component of tumor on T1c MRI, including nodular dural tails when relevant, and the resection cavity for post-surgical cases9,10,11. Residual enhancing tissue in postoperative scans was considered part of the GTV if clinically determined to represent tumor.
In some patients with multiple intracranial meningiomas, the GTV segmentation included all visible meningiomas, regardless of whether every lesion had been clinically targeted in the respective case’s actual RT plan. This approach provides a comprehensive representation of meningioma burden and ensures that automated segmentation algorithms trained on these data are robust to multi-lesion scenarios.
These target volume definitions are clinically useful in radiotherapy planning and were agreed upon by a coalition of BraTS organizers consisting of board-certified radiation oncologists and board-certified fellowship trained neuroradiologists. The target volume segmentation definitions were agreed upon after review of the EORTC 22042-026042 and RTOG 0539 annotation protocols9,10,11.
Annotation protocol
Each participating center had the option to provide their own clinical workflow GTV segmentations from the DICOM-RT structure sets. These institution-submitted segmentations served as an initial reference if they followed standard meningioma RT guidelines9,10,11. When institutional GTVs were either unavailable or did not conform to the challenge-defined annotation protocol, a deep convolutional neural network-based automated segmentation model was applied to generate a preliminary GTV mask. This model, built using the nnU-Net framework, was initially trained on the BraTS Preoperative Meningioma (BraTS-MEN) challenge data, which is the largest multi-institutional expert annotated multilabel preoperative meningioma multi-sequence MR image dataset to date15,18,32,33,34. Only T1c images and the tumor core segmentation (including enhancing and non-enhancing tumor) from the 1000 publicly available BraTS-MEN Dataset cases were utilized for nnU-Net training to most closely resemble the image and label data included in the BraTS-MEN-RT challenge. Training was performed using the default nnU-Net hyperparameters and data augmentation for a total of 1000 epochs33. Despite fundamental differences in processing (i.e., skull-stripping and atlas registration in the 2023 BraTS-MEN dataset), iterative retraining on accumulating BraTS-MEN-RT samples steadily improved the model’s ability to identify postoperative and residual/recurrent tumors. No formal quantitative analysis of nnU-Net training performance was conducted throughout the iterative re-training process, as the primary purpose was to accelerate data labeling.
After initial institution GTV collection or automated presegmentation of GTV, a senior radiation oncology resident (D.L.) conducted an initial review and potential revision through a slice-by-slice evaluation (ITK-SNAP) of each GTV and revised as needed to adhere to the BraTS-MEN-RT target volume definition as shown in Fig. 831. During the initial revision phase of nnU-Net automated segmentation GTV, common edits included the following:
-
1.
Adding or removing dural tail areas.
-
2.
Modifying segmentation edges to conform accurately with enhancing tissue boundaries in postoperative meningioma.
-
3.
Adding additional meningioma tumors outside of the clinical RT treatment volumes.
Axial, sagittal, and coronal brain MRI of a patient with multiple meningioma demonstrating the difference between the provided institution’s GTV as seen in panels A1, B1, and C1 compared to the manually revised target label as seen in panels A2, B2, and C2. Note that this case’s corrections involved inclusion of additional meningioma, correction of label edges, and inter-axial slice label smoothening.
After initial revision, a board-certified attending neuroradiologist (E.C.) reviewed the corrected GTV for final approval.
This protocol, guided by clinical best practices in meningioma delineation, aimed to ensure segmentation consistency across institutions and tumor subtypes7,9,10,11,35,36.
Data Records
The BraTS-MEN-RT Dataset training data (500/750, 66.7%), validation data (70/750, 9.33%), and clinical metadata are publicly available on Synapse at https://doi.org/10.7303/syn5905977929. The testing data (180/750, 24%) will be kept private for the foreseeable future to allow for unbiased evaluation of algorithms developed in future challenges including the MICCAI 2025 Lighthouse Challenge: “BraTS-Meningioma-RT: Meningioma Radiotherapy Segmentation”30. The “Meningioma radiotherapy supplementary clinical data and imaging parameters for training and validation sets.xlsx” file on the Synapse data repository describes the case level clinical patient data and the available image parameters for the training and validation cases29. The supplementary file “BraTS-MEN-RT Dataset Access Steps” provides step by step instructions on how to access the training image and segmentation data, validation image data, and the clinical patient data. All publicly available images, labels, and clinical metadata are of the CC-BY-NC license per Synapse policy.
Technical Validation
Patient clinical and demographic data
The clinical characteristics of individuals included in the BraTS-MEN-RT Dataset were sourced from medical records at each participating academic institution. Specific details regarding the methods used for data collection at each site were not disclosed, an approach intended to promote broader data contribution. The raw clinical data included in the BraTS-MEN-RT Dataset were not subjected to further independent validation.
Image pre-processing and defacing
All images that underwent defacing using AFNI were manually reviewed by a fellowship-trained attending neuroradiologist (E.C.) and senior radiation oncology resident (D.L.) to ensure adequate defacing, presence of at least one intracranial meningioma or postoperative target, and absence of a non-meningioma intracranial tumor. Any pre-processing defacing errors were manually corrected before inclusion in the dataset. It is important to note that the AFNI defacing we used may have limitations. Its face coverage is similar to older programs like mri_deface and PyDeface, and studies have shown that replacing more of the face, and especially the eyebrow ridge, can potentially result in greater privacy protection albeit at the cost of greater patient anatomy elimination21,22,37. Future challenges should compare additional face anonymization methods, including algorithms that either replace facial features with synthetic faces or further expand the defacing mask while preserving tumor boundaries to reduce potential re-identification risk while preserving clinically relevant anatomy.
Meningioma segmentations
All GTV segmentations were manually reviewed by a board-certified attending neuroradiologist “approver” (E.C.). In cases where the approver identified an inaccurate or incomplete segmentation, the case was further refined (E.C.) until satisfactory.
Data availability
The BraTS-MEN-RT Dataset image data, label data, and clinical metadata are publicly available on Synapse at https://doi.org/10.7303/syn5905977929.
Code availability
In line with the scientific data principles of findability, accessibility, Interoperability, and reusability38, the tools used throughout the generation of these data are publicly available. Specifically, we used the dcmrtstruct2nii repository for conversion of DICOM-RT to NIfTI, which is publically available at (https://github.com/Sikerdebaard/dcmrtstruct2nii)30. We used the Analysis of Functional NeuroImages (AFNI) automated defacing algorithm for defacing, which is publically available at (https://github.com/afni/afni)26,27. The nnU-Net model as used for initial pre-automated segmentation and iterative re-pre-automated segmentation is publicly available at https://github.com/ecalabr/nnUNet_models39. Additional GTV segmentation modification code used for revision of some institution provided GTVs is publicly available as CC-BY 4.0 at https://github.com/dlabella29/BraTS-MEN-RT_Data_Resource40.
References
Ogasawara, C., Philbrick, B. D. & Adamson, D. C. Meningioma: A Review of Epidemiology, Pathology, Diagnosis, Treatment, and Future Directions. Biomedicines 9, 319 (2021).
Huntoon, K., Toland, A. M. S. & Dahiya, S. Meningioma: A Review of Clinicopathological and Molecular Aspects. Front Oncol 10 (2020).
Ostrom, Q. T. et al. CBTRUS Statistical Report: Primary Brain and Other Central Nervous System Tumors Diagnosed in the United States in 2016—2020. Neuro Oncol 25, iv1–iv99 (2023).
Zhao, L. et al. An Overview of Managements in Meningiomas. Front Oncol 10 (2020).
Louis, D. N. et al. The 2021 WHO Classification of Tumors of the Central Nervous System: a summary. Neuro Oncol 23, 1231–1251 (2021).
Saraf, S., McCarthy, B. J. & Villano, J. L. Update on Meningiomas. Oncologist 16, 1604–1613 (2011).
Kent, C. L. et al. Long-Term Outcomes for Patients With Atypical or Malignant Meningiomas Treated With or Without Radiation Therapy: A 25-Year Retrospective Analysis of a Single-Institution Experience. Adv Radiat Oncol 7, 100878 (2022).
Pinzi, V. et al. Hypofractionated Radiosurgery for Large or in Critical-Site Intracranial Meningioma: Results of a Phase 2 Prospective Study. International Journal of Radiation Oncology*Biology*Physics 115, 153–163 (2023).
Weber, D. C. et al. Adjuvant postoperative high-dose radiotherapy for atypical and malignant meningioma: A phase-II parallel non-randomized and observation study (EORTC 22042–26042). Radiotherapy and Oncology 128, 260–265 (2018).
Rogers, L. et al. Intermediate-risk meningioma: initial outcomes from NRG Oncology RTOG 0539. J Neurosurg 129, 35–47 (2018).
Rogers, C. L. et al. High-risk Meningioma: Initial Outcomes From NRG Oncology/RTOG 0539. International Journal of Radiation Oncology*Biology*Physics 106, 790–799 (2020).
Wiant, D. & Bourland, J. D. Simulated Gamma KnifeTM Head Frame Placement for Radiosurgical Pre-Planning. Technol Cancer Res Treat 8, 265–270 (2009).
Gallagher, T. A., Nemeth, A. J. & Hacein-Bey, L. An Introduction to the Fourier Transform: Relationship to MRI. American Journal of Roentgenology 190, 1396–1405 (2008).
Adewole, M. et al. The Brain Tumor Segmentation (BraTS) Challenge 2023: Glioma Segmentation in Sub-Saharan Africa Patient Population (BraTS-Africa). ArXiv (2023).
LaBella, D. et al. The ASNR-MICCAI Brain Tumor Segmentation (BraTS) Challenge 2023: Intracranial Meningioma. ArXiv (2023).
Kazerooni, A. F. et al. The Brain Tumor Segmentation (BraTS) Challenge 2023: Focus on Pediatrics (CBTN-CONNECT-DIPGR-ASNR-MICCAI BraTS-PEDs). ArXiv (2024).
Moawad, A. W. et al. The Brain Tumor Segmentation (BraTS-METS) Challenge 2023: Brain Metastasis Segmentation on Pre-treatment MRI. ArXiv (2023).
LaBella, D. et al. Analysis of the BraTS 2023 Intracranial Meningioma Segmentation Challenge. (2024).
Bakas, S. et al. Advancing The Cancer Genome Atlas glioma MRI collections with expert segmentation labels and radiomic features. Sci Data 4, 170117 (2017).
Menze, B. H. et al. The Multimodal Brain Tumor Image Segmentation Benchmark (BRATS). IEEE Trans Med Imaging 34, 1993–2024 (2015).
Schwarz, C. G. et al. Identification of Anonymous MRI Research Participants with Face-Recognition Software. New England Journal of Medicine 381, 1684–1686 (2019).
Schwarz, C. G. et al. Changing the face of neuroimaging research: Comparing a new MRI de-facing technique with popular alternatives. Neuroimage 231, 117845 (2021).
Thakur, S. P. et al. Skull-Stripping of Glioblastoma MRI Scans Using 3D Deep Learning. in 57–68, https://doi.org/10.1007/978-3-030-46640-4_6 (2020).
Thakur, S. et al. Brain extraction on MRI scans in presence of diffuse glioma: Multi-institutional performance evaluation of deep learning methods and robust modality-agnostic training. Neuroimage 220, 117081 (2020).
Bischoff‐Grethe, A. et al. A technique for the deidentification of structural brain MR images. Hum Brain Mapp 28, 892–903 (2007).
Cox, R. W. AFNI: Software for Analysis and Visualization of Functional Magnetic Resonance Neuroimages. Computers and Biomedical Research 29, 162–173 (1996).
Cox, R. W. & Hyde, J. S. Software tools for analysis and visualization of fMRI data. NMR Biomed 10, 171–8 (1997).
Theyers, A. E. et al. Multisite Comparison of MRI Defacing Software Across Multiple Cohorts. Front Psychiatry 12 (2021).
Calabrese, E. & LaBella, D. 2024 BraTS Meningioma Radiotherapy Dataset. https://doi.org/10.7303/syn59059779 (2024).
Phil, T., Albrecht, T., Gay, S. & Rasmussen, M. E. Sikerdebaard/dcmrtstruct2nii: dcmrtstruct2nii v5 (Version v5). Zenodo https://doi.org/10.5281/zenodo.4037864 (2023).
Yushkevich, P. A. et al. User-guided 3D active contour segmentation of anatomical structures: Significantly improved efficiency and reliability. Neuroimage 31, 1116–1128 (2006).
LaBella, D. et al. A multi-institutional meningioma MRI dataset for automated multi-sequence image segmentation. Sci Data 11, 496 (2024).
Isensee, F., Jaeger, P. F., Kohl, S. A. A., Petersen, J. & Maier-Hein, K. H. nnU-Net: a self-configuring method for deep learning-based biomedical image segmentation. Nat Methods 18, 203–211 (2021).
Calabrese, E. & LaBella, D. 2023 BraTS Meningioma Dataset. https://doi.org/10.7303/syn51514106.
Aghi, M. K. et al. Long-term recurrence rates of atypical meningiomas after gross total resection with or without postoperative adjuvant radiation. Neurosurgery 64, 56–60 (2009).
Martz, N. et al. ANOCEF Consensus Guideline on Target Volume Delineation for Meningiomas Radiotherapy. International Journal of Radiation Oncology*Biology*Physics 114, e46 (2022).
Schwarz, C. G. et al. A face-off of MRI research sequences by their need for de-facing. Neuroimage 276, 120199 (2023).
Wilkinson, M. D. et al. The FAIR Guiding Principles for scientific data management and stewardship. Sci Data 3, 160018 (2016).
Calabrese, E. https://github.com/ecalabr/nnUNet_models.
LaBella, D. BraTS-MEN-RT_Data_Resource. https://doi.org/10.5281/zenodo.17582447 (2025).
Acknowledgements
We are grateful to everyone who contributed data and to the development and review of the tumor volume segmentations including Radiation Oncology and Neuroradiology residents and attendings from Duke University, University of California San Francisco, State University of New York Upstate Medical University, University of Washington, University of Missouri, King’s College London, and University of California San Diego. Research reported in this publication was partly supported by the National Institutes of Health (NIH), under award numbers U24CA279629, U01CA242871, NCI K08CA256045, and NCI/ITCR U01CA242871. The content of this publication is solely the responsibility of the authors and does not represent the official views of the NIH.
Author information
Authors and Affiliations
Contributions
Conceptualization (D.L., M.P., S.A., S.F., P.W., J.A., A.J., M.M., V.C., R.C., N.M., R.S., F.K., C.G.S., P.L., P.V., L.G., M.A., H.L., A.F., N.T., U.A., A.M., B.M., M.L., M.A., B.W., U.B., G.C., A.R., A.N., A.A., R.H., M.C., J.R., S.B., E.C.), Methodology (D.L., T.M., J.K., E.V., Z.R., Sc.Fl., M.P., S.A., Sh.Fa., P.W., J.A., A.J., M.M., V.C., R.C., N.M., R.S., F.K., P.L., P.V., L.G., M.A., H.L., A.F., N.T., U.A., A.M., B.M., M.L., M.A., B.W., U.B., G.C., A.R., A.N., A.A., R.H., M.C., J.R., S.B., E.C.), Software (D.L., S.P., M.S., A.A., A.K., H.K., V.C., R.C., J.A., U.B., S.B., E.C.), Validation (E.C.), Formal Analysis (D.L., S.B., R.S., R.C., V.C., J.R., U.B., N.T., U.A., A.M., B.M., M.L., M.A., B.W., G.C., A.R., A.N., A.A., R.H., M.C., E.C.), Resources (D.L., J.K., S.P., M.S., A.A., A.K., H.K., V.C., R.C., J.A., U.B., S.B., E.C.), Data Curation (D.L., K.S., M.M., P.T., K.L., S.M., P.N., J.V., D.R., J.S., T.V., K.C., M.I., T.B., O.A., J.L., L.H., Y.V., C.W., J.K., S.F., Z.R., T.M., E.V., U.B., S.S., J.H., T.S., N.F., C.P., M.P., K.S., N.T., A.N., A.R., J.R., M.C., E.C.), Data Annotation (D.L., E.C.), Writing - Original Draft (D.L., E.C.), Writing - Review & Editing (D.L., C.S., M.C., E.C.), Supervision (S.B., E.C.), Project Administration (D.L., E.C.), Funding Acquisition (E.C., S.B., U.B.).
Corresponding author
Ethics declarations
Competing interests
T.S. reports honoraria from Varian Medical Systems, WebMD, GE Healthcare, and Janssen; he has an equity interest in CorTechs Labs, Inc. and serves on its Scientific Advisory Board; he receives research funding from GE Healthcare through the University of California San Diego. J.S. and T.V. are co-founders and shareholders of Hypervision Surgical whose interests are unrelated to the present work. P.L. reports honoraria for lectures from Blue Earth Diagnostics and for advisory board participation from Servier Pharmaceuticals.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
LaBella, D., Schumacher, K., Mix, M. et al. The 2024 Brain Tumor Segmentation Challenge Meningioma Radiotherapy (BraTS-MEN-RT) dataset. Sci Data 13, 306 (2026). https://doi.org/10.1038/s41597-026-06649-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-026-06649-x