Abstract
Contaminated lake water and fish can be sources of bacterial pathogens of public health concern, including pathogenic E. coli. Within Ethiopia, specifically, Central Oromia, raw fish consumption is a common practice. Although there are few reports on occurrence of E. coli O157 in fish destined for human consumption and children under five years, information on the transmission pathways of E. coli O157 and other sorbitol non-fermenting (SN-F) E. coli from water-to-fish-to-human, and their virulence factors and antimicrobial resistant determinants along the fish supply chain is lacking. The study aimed to investigate the occurrence, molecular characteristics, and antimicrobial susceptibility of E. coli O157 and other SN-F E. coli strains in fish, lake water and humans in central Oromia, Ethiopia. A total of 750 samples (450 fish samples, 150 water samples, 150 human stool samples) were collected from five lakes and three health facilities. The samples were processed following the standard protocol recommended by European Food Safety Authority and Kirby–Bauer disc diffusion method for detection of the bacteria, and antimicrobial susceptibility tests, respectively. Molecular characterization of presumptive isolates was performed using Whole-Genome Sequencing (WGS) for serotyping, determination of virulence factors, antimicrobial resistance traits, and genetic linkage of the isolates. Overall, 3.9% (29/750) of the samples had SN-F E. coli; of which 6.7% (n = 10), 1.8% (n = 8) and 7.3% (n = 11) were retrieved from water, fish, and diarrheic human patients, respectively. The WGS confirmed that all the isolates were SN-F non-O157: H7 E. coli strains. We reported two new E. coli strains with unknown O-antigen from fish and human samples. All the strains have multiple virulence factors and one or more genes encoding for them. Genetic relatedness was observed among strains from the same sources (water, fish, and humans). Most isolates were resistant to ampicillin (100%), tetracycline (100%), cefotaxime (100%), ceftazidime (100%), meropenem (100%), nalidixic acid (93.1%) and sulfamethoxazole/trimethoprim (79.3%). Majority of the strains were resistant to chloramphenicol (58.6%) and ciprofloxacin (48.3%), while small fraction showed resistance to azithromycin (3.45%). Isolates had an overall MDR profile of 87.5%. Majority, (62.1%; n = 18) of the strains had acquired MDR traits. Genes encoding for mutational resistance and Extended-spectrum beta-lactamases (ESBL) were also detected. In conclusion, our study revealed the occurrence of virulent and MDR SN-F E. coli strains in water, fish, and humans. Although no genetic relatedness was observed among strains from various sources, the genomic clustering among strains from the same sources strongly suggests the potential risk of transmission along the supply chain at the human–fish-environment interface if strict hygienic fish production is not in place. Further robust genetic study of the new strains with unknown O-antigens, and the epidemiology of SN-F E. coli is required to elucidate the molecular profile and public health implications of the pathogens.
Similar content being viewed by others
Introduction
Foodborne diarrheal diseases, including those acquired through fish consumption, are among the leading cause of morbidity and mortality globally with a mortality rate of 22·4 deaths per 100, 000 person-years, and a substantial impairment to socioeconomic development worldwide1,2. The mortality rate is higher among children under 5 years, elderly people over 70 years, and in low-income countries3.
Perishable food items like ground beef, raw milk, meat, fish, vegetables, unpasteurized fruit juices originated from contaminated sources or contaminated during production or processing may harbor potential foodborne pathogens, including E. coli4,5,6. For instance, cross-contamination, inadequate cooking and storage are the dominant food safety impediments responsible for 60–78% of the foodborne disease burden (FBD); while raw foods accounted for an estimated 23–41% of the disease burden in France7.
Both wild and cultured fish are sources of a wide variety of bacterial pathogens of public health concern8,9. Human infections due to potential pathogens acquired from fish or the aquatic environment are also quite common depending on season, poor hygienic fish handling practices, raw fish consumption habits and the immune status of the exposed individuals10; implying that detection of the microbial quality of fish intended for human consumption is crucial11.
Although most strains of E. coli are normal inhabitants of the intestinal tract of humans, animals and fish12, some strains have acquired genetic determinants encoding for various virulence factors giving them the capacity to exert intestinal and extra-intestinal illness in humans with a characteristic watery or bloody diarrhea13. Based on their clinical, epidemiological, and pathogenic characteristics, there are seven pathotypes of diarrheagenic E. coli strains; including enteropathogenic E. coli (EPEC), enterotoxigenic E. coli (ETEC), enteroinvasive E. coli (EIEC), enterohemorrhagic E. coli (EHEC) or Shiga toxin producing E. coli (STEC), enteroaggregative E. coli (EAEC), diffusely adherent E. coli (DAEC)13, and a new pathotype, adherent invasive E. coli (AIEC)14. Each pathotype has a distinctive virulence factor responsible for colonization and subsequent pathogenic effects of the pathogen15,16, among which EPEC, ETEC, and EAEC are the dominant causes of infantile diarrhea in developing countries with a relatively low standard of living17.
E. coli may contaminate fish products destined for human consumption and become a potential fish safety concern12. For instance, from 2009–2015 in the United States and Puerto Rico alone, the FBD Outbreak Surveillance System (FDOSS) section of CDC received reports of 344 outbreaks and 2,288 illnesses associated with consumption of contaminated food of aquatic origin including fish18. A report from the Zhejiang Province of China has also shown, from 2010–2020 aquatic products including fish were responsible for 109 outbreaks, 1073 cases, and 77 hospitalizations, though no death was recorded19. Previous reports have also shown that zoonotic pathogens including E. coli can be transmitted to humans via consumption of contaminated and improperly processed fish or fish products20.
However, despite increasing evidence of detecting E. coli strains harboring many virulence genes from human and animal feces, only a few studies have investigated the occurrence of pathogenic E. coli strains in environmental water including lake water21, and fish12. Although Ethiopia has a huge potential of fish production and its consumption is common in the country, the fish quality and safety aspect is overlooked by regulatory bodies due to limited implementation of food safety regulations22. The report of Bedane et al. 2022 indicated lack of infrastructure for fish production and processing, unavailability of cold chain facility for transportation, unhygienic handling practices, and the habit of consuming raw fish in central Oromia, Ethiopia23. Besides, some studies reported the occurrence of Shiga toxin producing E. coli O157 in fish and humans from various parts of the country. A 1.6% and 1.3% occurrence of E. coli O157 in fish in Lake Hayq and Tekeze dam, respectively, was reported from northern Ethiopia24. Similarly, Tilahun and Engdawork25, have reported 2.3% prevalence of the pathogen from fish in Lake Hawassa, southern Ethiopia. In humans, 15.3% prevalence and 28.9% isolation rate of E. coli O157 related diarrhea in children under five years was reported from Eastern Ethiopia26, and Bahir Dar town of northern Ethiopia27, respectively. In a similar study, Gutema et al.28, have detected E. coli O157 in 2.8% of 216 diarrheic patients' stool samples investigated in Bishoftu town, central Ethiopia. Besides, E. coli is the dominant donor and recipient of resistance genes through horizontal gene transfer, and directly or indirectly, it is the prime cause of treatment failures in both animals and humans. Thus, at a global scale, antimicrobial resistant E. coli is a public health concern29.
In Ethiopia, Central Oromia in particular, although raw fish consumption is a common practice, and there are few reports of E. coli O157 from fish destined for human consumption and children under five years, information on the transmission pathways of E. coli O157 and other sorbitol non-fermenting E. coli strains along the fish supply chain and, the virulence factors and antimicrobial resistant determinants of the pathogens is lacking. Therefore, this study was aimed to investigate the occurrence, molecular characteristics, and antimicrobial susceptibility of E. coli O157: H7 and other sorbitol non-fermenting E. coli strains in lake water, fish, and humans to examine potential transmission pathways along fish supply chain in central Oromia, Ethiopia.
Materials and methods
Study sites and settings
The study was conducted from December 2020 to June 2022 in East Shewa zone (Bishoftu, Koka and Batu). East Shewa zone is located in the upper rift-valley region and endowed with, a number of crater lakes. The zone has a total population of 1,356,342 within an area of 8,370.90 square kilometers with a population density of 162.03. Three-fourths (74.9%) of the population are mixed crop-livestock farmers dwelling in rural settings, while 25.1% are urban inhabitants30. The rift-valley lakes including lakes Dambel, Koka, Babogaya, Hora-Arsedi and Koftu are important source of fish for consumers in the adjacent towns (Bishoftu, Koka and Batu towns) and beyond31.
Ethics statement
The study was reviewed and approved by Addis Ababa University Ethical Review Committee (Ref No.VM/ERC/14/05/13/2021) and Oromia Health Bureau Health Research Ethical Review Committee (Ref No. BEFO/4BTW/1-16/10393); and all methods were performed in accordance with ARRIVE guidelines (https://arriveguidelines.org). Moreover, after explaining the purpose of the study, informed consent was obtained from all subjects and/or their legal guardians.
Study design and sampling
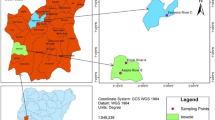
A cross-sectional study design was employed to collect data from five lakes that were conveniently selected based on their accessibility and fishing potential. These include Babogaya, Hora-Arsedi and Koftu Lakes in Bishoftu town; Koka Lake nearby Koka town, and Dambel Lake at Batu town. Stool samples were collected from diarrheic out-patients at Bishoftu hospital, Koka health center and Batu health center (Fig. 1). On each sampling day, live fish were purchased at the lake shore from the fishermen who harvested them for commercial purposes, and placed in a bucket of water so that they feel as if they are in their natural environment. Then, the fish were stunned by mixing a clove oil (having an anesthetic nature) into the bucket of water in which they were placed few minutes before severing with a sharp knife to reduce pain during slaughter. Thirty fish from each Lake, comprising of six fish species (Oreochromis niloticus, Clarias gariepinus, Tilapia Zilli, Cyprinus carpio, Labeobarbus intermedius, Barbus ethiopicus), which are commonly used for human consumption were included in the study (n = 150), from which 150 fecal, 150 meat, 150 skin swabs were sampled. A total of 750 samples, comprising of 150 water samples, 450 fish samples, and 150 stool samples were collected and processed. A maximum of ten fish were identified and sampled per sampling day from which 30 samples consisting of each ten fecal, meat and swab samples were collected on a visiting day.
Map of the study areas and the Oromia reginal state within the federal democratic republic of Ethiopia.
Sample collection
Skin swabs were collected using 2 × 3 cm sterile cotton tipped swabs soaked in 10 ml of buffered peptone water (BPW) (Oxoid Ltd., Hampshire, England), as described by McEvoy et al.32. The skin was swabbed several times first horizontally and then, vertically both in the left and right lateral commissars of the fish from the gill area towards the caudo-ventral region covering the entire dissection area of ~ 10 × 10 cm (an area with substantial risk of contact with a fillet during scaling). Then, the swabs were placed in a sterile universal bottle containing 10 ml of BPW by removing extra shafts and the bottles were screw capped.
After dissection, scaling, and gutting, ~ 10 g of meat (fillet) samples were taken from different parts of the fillet nearby the dissection area (the most suspicious part for external and fecal contamination) and pooled as a single sample; placed in a sterile universal bottle containing 10 ml of BPW.
After gutting, ~ 10 g of fecal samples were collected as described by Elder et al.33. Briefly, the whole intestinal organs were separated from the remaining parts of the dissected fish and placed in a sterile plastic bag. Then, the intestinal lumen was opened using a sterile surgical blade and the fecal samples were put into a sterile universal bottle containing 10 ml of BPW.
Water samples (~ 20 ml each) were collected by immersing sterile universal bottles containing 10 ml of BPW into a grossly visible dirty area of the water bodies near the Lake shore with an interval of about 2 m between each sampling points. Similarly, about 10 g of stool samples were collected from the stools submitted by the diarrheic patients for laboratory analysis, into a sterile universal bottle containing 10 ml of BPW in collaboration with the laboratory technicians using a sterile toothpick for each sampling. The procedures were done as aseptic as possible using a sterile disposable glove to avoid the risk of cross contamination during each sampling. Finally, all samples were carefully labeled using a permanent marker, kept in an ice box containing ice packs and transported to Addis Ababa University College of Veterinary Medicine and Agriculture Veterinary Public Health Laboratory for processing.
Detection of sorbitol non-fermenting E. coli and E. coli O157
The samples were primarily processed targeting sorbitol non-fermenting E. coli, and then Shiga toxin producing E. coli O157 strain. Briefly, all the samples in the universal bottle containing 10 ml BPW were incubated at 41.5°c for 6 h to revive and increase the recovery rate of stressed bacterial cells34. Then, the meat samples were blended using stomacher (Seward Stomacher 400, Seward, London, UK) at a speed of 150 rpm for 60 s and, the aliquots from the meat, water, fecal, stool and swab cultures were selectively enriched by measuring 10 ml of samples into 90 ml modified tryptone soya broth (mTSB) (Oxoid, Hampshire, England) and agitating using the same blending machine at 150 rpm for 60 s. After homogenization, the selectively enriched samples were incubated overnight at 41.5°c.
A loop-full of aliquots from selective enrichment was streaked on cefixime-potassium tellurite Sorbitol MacConkey agar (CT-SMAC) and incubated at 37°c for 24 h. After incubation, sorbitol non-fermenting colonies (colorless colonies) were sub-cultured on CT-SMAC and incubated at 37°c for 24 h to obtain adequate number of pure colonies. To rule-out the growth of other Gram negative organisms with colorless colonies on CT-SMAC, including Burkholderia, Vibrio, Proteus, Klebsiella, Aeromonads and Pseudomonas35, the pure colonies on CT-SMAC were further sub-cultured onto eosin methylene blue (EMB) agar and incubated at 37°c for 24 h. Isolates with typical characteristic of green metallic sheen color of the generic E. coli on EMB agar were further tested for indole production. All indole positive presumptive colonies were preserved on tryptone soya agar after incubating at 37°c for 24 h and adding 80% glycerol for further analysis. The presumptive isolates were shipped to Belgian National Reference Centre, Brussels, Belgium for immunological testing and whole genomic sequencing.
E. coli O157 latex agglutination test
At the reference laboratory in Brussels, Belgium, the isolates were further sub-cultured on CT-SMAC to perform latex agglutination test. Then, each presumptive isolate was emulsified with normal saline on disposable reaction cards provided in the kit and mixed with a drop of test latex (a latex particle sensitized with specific rabbit antibody reactive with the O157 somatic antigen) and agitated for one minute. After one minute, the result was interpreted based on the standard protocol described by DeBoer and Heuvelink36. Due to lack of reference strains, known positive controls were not used. However, after the agglutination test, a suspension of inactivated E coli O157 and E coli O116 cells in buffer were used as a positive and negative controls, respectively.
Whole-genome sequencing (WGS)
Genomic DNA was extracted from pure cultures of SN-F E. coli isolates grown overnight on SMAC Agar; and its purity and quantity were measured with a Qubit double stranded DNA (dsDNA) BR assay kit. Then, fragmentation of 500 ng of genomic DNA was carried out using the NEBNext® Ultra™ II FS module. Sequencing libraries, with an insert size of on average 550 bp, were prepared using a KAPA Hyper Plus kit (Kapa Biosystems, Wilmington, USA) and a Pippin Prep (Sage Science, Beverly, MA, USA) size selection with a CDF1510 1.5% agarose dye-free cassette. Every sample was assigned an in-house truseq style adapter with a unique dual-indexed 8-bp barcode. PCR amplification (6 cycles) of the library was performed using the KAPA HiFi HotStart Libray Amplification kit (Kapa Biosystems, Wilmington, USA) according to the manufacturer’s instructions. After equimolar pooling, libraries were sequenced on a Novaseq 6000 instrument (Illumina, San Diego, CA, USA) using a NovaSeq 6000 SP Reagent kit (500 cycles) generating 2 × 250 bp reads. For this, the library was denaturated and diluted according to the manufacturer’s instructions. A 1% PhiX control library was included in each sequencing run. Lastly, the raw reads were uploaded, and de novo assembled, using SPAdes v.3.15.3, in BioNumerics v.8.1., and sequence quality was assessed using the quality metrics incorporated in BioNumerics v.8.1.
Comparison of SN-F E. coli strains retrieved from water, fish and humans in EnteroBase
The raw reads of 29 SN-F E. coli genomes were uploaded and automatically assembled in the public genome database, EnteroBase. All genome assemblies were subsequently compared to the available E. coli genomes using hierarchical clustering of cgMLST (HierCC) at different levels of resolution, ranging from HC0 (hierarchical clusters consisting of identical genomes with no AD) to HC200 (hierarchical clusters consisting of genomes with up to 200 ADs)37. However, up on quality check, the genomes of six isolates were poor and only 23 isolates were included in the cluster analysis.
In silico identification of genes encoding for serotype, virulence factors, and antimicrobial resistance traits
The E. coli genotyping tool v.2.1, available in BioNumerics v.8.1 was used to predict E. coli serotypes, virulence gene profiles, acquired resistance genes, and point mutations starting from the genome assemblies. The presence of virulence and resistance genes was determined with a minimum % identity (ID) threshold of 85% and a minimum length for coverage of 60%.
Antimicrobial susceptibility test
Antimicrobial susceptibility test was conducted on a total of 29 SN-F E. coli strains retrieved in the present study following the standard protocol described by CLISI 202238, using ten essential antimicrobial agents obtained from commercial market (Thermo Scientific Fisher); namely, (ampicillin 10 μg, tetracycline 30 μg, ciprofloxacin 5 μg, azithromycin 15 μg, chloramphenicol 30 μg, ceftazidime 30 μg, cefotaxime 30 μg, meropenem 10 μg, sulfamethoxazole /trimethoprim 25 μg, and nalidixic acid 30 μg) grouped under eight antimicrobial classes using Kirby-Bauer disc diffusion method38. The antibiotics were selected based on their common usage in humans and animals and AMR reports. The discs were funded by Michigan State University through the 2019 faculty exchange program for one of our co-authors. To estimate the concentration of the isolates in the culture broth, 0.5 McFarland Standard (≈ 1.5 × 108 CFU/ml) was prepared by measuring 0.05 ml of 1% BaCl2 and 9.95 ml of 1% H2SO4; and the turbidity of the culture broth was adjusted towards this standard using sterile saline solution. A sterile cotton swab was immersed into the broth culture and uniformly swabbed to Mueller–Hinton agar plates. The plates were allowed to dry for few minutes, and the antimicrobial discs were randomly placed on the surface of the agar plates with gentle pressure using sterile forceps. The plates were incubated at 37 °C for 18 h; and after incubation, the inhibition zones of each antimicrobial agent were carefully measured using a digital caliper. The test results were qualitatively interpreted as susceptible, intermediate, or resistant based on zone diameter interpretative standards established for E. coli and other enteric Gram negative rods38. Controls were accomplished with an in-silico identification of genes encoding for antimicrobial resistance traits using E. coli genotyping tool v.2.1, available in BioNumerics v.8.1 to predict acquired resistance genes, and point mutations following the standard procedure described by Feldgarden et al.39.
Data analysis
The data were entered into an excel spreadsheet (Microsoft office® excel 2013, Cengage Learning, version 15) and analyzed using Stata version 15.0 software (Stata Corp, College Station, TX). Descriptive statistics such as frequency and percentages were used to express the sociodemographic characteristics of the diarrheic patients and the antimicrobial susceptibility profiles of the isolates. The proportions of occurrence of sorbitol non-fermenting E. coli in diarrheic patients and, water and fish samples were calculated by dividing the number of culture positive samples by the total number of samples tested from each sample source. Fisher’s exact test was used to assess the difference in the proportion of sorbitol non-fermenting E. coli among the different sample sources.
The assembled sequencing data was analyzed using the Escherichia /Shigella cgMLST typing scheme in BioNumerics v.8.1 (core Enterobase). Both assembly algorithms, namely, the assembly-free k-mer-based approach using the raw reads and the assembly-based BLAST approach were used for allele calling. The default settings were used for both the assembly-free and assembly-based algorithms. The quality of the assembly-free and the assembly-based allele calls were verified using the quality statistics window in BioNumerics. The MLST profile of each isolate was determined using the three basic allele mapping experiments (Pub (Achtman) MLST, Pasteur MLST, Whittam MLST) incorporated in BioNumerics. Minimum spanning tree (MSTs) diagrams of the cgMLST data were generated using the MSTree V2 algorithm and visualized by GrapeTree in EnteroBase.
Results
Demographic characteristics and proportion of SN-F E. coli strains per sample sources
Among the total of 150 diarrheic patients participated in the study, 52.67% of them were males. The mean age of the patients was 30.7 years (range: 9 months to 70 years). From the patients investigated, 8.7% (13), 17.3% (26), 17.3% (26), and 56.7% (85) were observed with the clinical history of bloody, mucoid, mixed and watery diarrhea, respectively (Table 1).
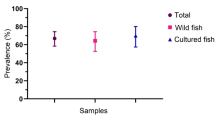
The overall culture-based proportion of sorbitol non-fermenting E. coli strains in all samples was 3.9% (29/750). Specific sample level detection rate was 6.7% (10/150), 1.8% (8/450), and 7.3% (11/150) in water, fish, and human samples, respectively (Table 1).
Among the total of 10 SN-F E. coli strains detected in water samples, higher proportion (40%, n = 4) was detected in samples collected from lake Koftu, followed by lake Hora-Arsedi (30%; n = 3). Similarly, from a total of 450 fish samples examined, only eight SN-F E. coli strains were detected (n = 8; 1.8%, 95%CI: 0.7, 3.5%). The majority, 87.5% (n = 7), of the isolates were recovered from fish collected from lake Koka. Fifty percent (n = 4) of the fish isolates were detected in meat samples, as compared to the skin swab and fecal samples. Nevertheless, none of the fish samples collected from Babogaya, Koftu and Dambel Lakes were tested positive (Table 2).
In diarrheic patients, the overall detection rate of sorbitol non-fermenting E. coli strains was 7.3% (95% CI: 3.7, 12.7%) with higher prevalence in younger age groups (5–17 years) and children under 5 years; at a detection rate of 10.5% and 8.3%, respectively (Table 2).
Core genome multi locus sequence typing (cgMLST) confirmed that all the strains detected from water, fish and humans were SN-F E. coli strains. None of the E. coli strains were tested positive for E. coli O157 using E. coli O157 latex agglutination test, and whole genome sequencing (WGS). Core genome MLST also detected a new strain with unknown O-antigen from fecal sample of fish obtained from Lake Koka and stool sample of diarrheic patient presented to Bishoftu Hospital (Table 3).
Comparison of SN-F E. coli Genomes in EnteroBase
Comparison of the genetic linkage among the SN-F E. coli strains retrieved from the three sample sources (water, fish and humans) in EnteroBase showed lack of genetic relationships among the isolates. However, genetic relatedness was observed among strains from the same sample sources: 2 clusters of E. coli strains from humans (5 and 2 strains), water (2 strains) and Fish (2 strains) (Fig. 2).
Grape-Tree diagram generated in EnteroBase based on core genomic MLST profiles of 23 E. coli strains isolated from fish, lake water and humans showing lack of clustering among strains from the three sample sources. Nodes are color-coded based on the sample types and sample ID.
In silico identification of genes encoding for virulence factors and antimicrobial resistance traits
An in silico MLST showed that all the strains have multiple virulence factors and one or more genes encoding for them. The most repetitive virulence factors and respective encoding genes detected were survival (iss), adherence proteins (papC, lpfA, irp2, k88ab), invasion (ompT, cia), toxin production (hlyE, hlyF, cvaC, mchF), and metabolic regulation (terC, eilA). It also revealed genes encoding for acquired resistance of ampicillin, amoxicillin, cephalothin, piperacillin, and ticarcillin (blaTEM-1B), doxycycline and tetracycline (tet(A)), trimethoprim (dfrA1and dfrA5), sulfamethoxazole (sul1), fosfomycin (fos7), chloramphenicol (catA1), ciprofloxacin (qnrS1), and nalidixic acid (gyrA). Moreover, genes encoding for mutational resistance against ciprofloxacin and nalidixic acid ((parC, gyrA), and extended spectrum beta-lactamase (ESBL) against aztreonam, cefepime, ceftaxime, ceftazidime, and ceftriaxone (blaCTX-M15) were detected (See the supplementary file).
Overall, 18 (62.1%) of the isolates were virulent strains; of which, 7 (38.9%), 6 (33.3%), and 5 ((27.8%) were retrieved from water, fish, and human, respectively. The majority (55.6%; n = 10) of the virulent isolates have genes encoding for antimicrobial resistance. Among this, 5 (50%), 4 (40%) and 1 (10%) isolates were from humans, fish and water respectively (See the supplementary file).
Antimicrobial susceptibility
Of the 29 sorbitol non-fermenting E. coli strains tested, 96.6% (n = 28) of them were susceptible to azithromycin followed by ciprofloxacin (20.7%, n = 6), chloramphenicol (17.3%, n = 5) and sulfamethoxazole /trimethoprim (13.8%, n = 4). All of the isolates were resistant to ampicillin, cefotaxime, ceftazidime, meropenem and tetracycline (Table 4).
A multi-drug resistance (MDR) of 87.5% was observed; where the strains were resistant to seven antimicrobial classes from the eight antimicrobial classes tested. Five isolates and one isolate were resistant to 90% (n = 9) and 100% (n = 10) of the antimicrobial agents used, respectively. An in silico MLST analysis also confirmed that the majority (62.1%; 18/29) of the strains have multiple acquired resistance against ampicillin, amoxicillin, cephalothin, piperacillin, ticarcillin, doxycycline, tetracycline, trimethoprim, sulfamethoxazole, and one or more genes encoding for them. Genes encoding for mutational resistance against ciprofloxacin and nalidixic acid, and extended spectrum beta-lactamase (ESBL) against aztreonam, cefepime, ceftaxime, ceftazidime, and ceftriaxone were also detected (See the supplementary file).
Discussion
In this study, we investigated the occurrence, molecular characteristics, and antimicrobial susceptibility of sorbitol non-fermenting E. coli with particular emphasis to E. coli O157 in Lake Water, fish and humans in central Oromia, Ethiopia. The study showed that none of the identified isolates are E. coli O157. Interestingly, the study revealed antimicrobial resistant SN-F E. coli and new SN-F E. coli strains with unknown O-antigen, and genetic relatedness among strains from the same sources with no linkage among strains from different sample sources (water, fish and humans). The majority of the strains have multiple genes encoding for a single virulence factor and antimicrobial resistance traits; implying that the isolates are pathogenic and antimicrobial resistant E. coli strains.
The genetic relationships among the isolates from the same sources suggests the circulation and the potential risk of dissemination of virulent and antimicrobial resistant strains along the fish supply chain in the study area. Similar report from Southeastern Nigeria revealed the occurrence of MDR SN-F E. coli strains in other species of animals (broiler chicken, cattle and pig), and suggested that the pathogens may spread from animals to humans and the environment making it a public health threat40. Likewise, the recent report of Bedane et al.23, on hygienic fish handling practices in the study area indicated that the Lakes are used as watering points for cattle, which are the principal carriers of E. coli O157:H7, and fully accessible to run-off water, which may carry many other pathogens of public health concern, including SN-F E. coli.
Reports on SN-F non- O157 E. coli strains are limited and as a result, our findings are discussed in relation to previous reports on other E. coli strains and E. coli O157 as appropriate to provide an overall insight. The observed 6.7% SN-F E. coli strains in lake water samples in the present study is lower than the finding of Mekonnen et al.41, who reported 93.3% E. coli in water samples collected from Lake Dambel, and nearly comparable with the report of Dissasa et al.42, who retrieved 5.9% of generic E. coli from water samples of Lakes Dambel, Langano and Hawasa of Ethiopia. Similarly, Akoachere et al.43, in Cameron and Ribeiro et al.44, in Brazil found that bacteria of the family Enterobacteriaceae, including E. coli, are widely distributed in the aquatic ecosystem. Once the aquatic environment is contaminated with bacterial pathogens of public health concern, fish can easily acquire the pathogens naturally by direct physical contact44. This substantiates the higher odds of exposure of fish to pathogenic organisms and the subsequent higher odds of human exposure to the pathogens. The observed variations could be due to differences in the method of detection and targeted E. coli strains in this study. We primarily investigated the occurrence of sorbitol-non fermenting E. coli as compared to other studies that reported the prevalence of the generic E. coli which yield higher detection rate. Moreover, the variations could be attributed to the temporo-spatial disparity of the studies, and related anthropogenic and natural elements which affect the magnitude of contamination of lake water.
Sorbitol non-fermenting E. coli strains were detected in 1.8% of fish samples analyzed during the present study. An important notable finding is that 50% of the fish isolates were detected in meat, showing a remarkable contamination of the filet either from the external environment including skin, or intestinal contents due to poor handling and processing practices. Similar study conducted in India reported 19.2% occurrence of SN-F E. coli in cattle feces45, implicating the potential risk of contamination of Lake water and fish with SN-F E. coli strains of cattle fecal origin. The poor hygienic conditions along the fish supply chain in the study area might have contributed to the contamination of the lakes with animal feces during watering and through water run-off23.
Compared to the previous studies by Haile and Getahun46, Yohans et al.47, and Dissasa et al.41, who reported the generic E. coli in 12%, 20%, and 5.7% of fish samples collected from Lakes Dambel, Tana, and three rift-valley Lakes (Dambel, Langano and Hawasa), respectively, our report is exceedingly lower. It is also extremely lower than the finding of Marijani48, who recovered E. coli from 39% of fish sampled from marine and freshwater fish in Tanzania. Our detection of sorbitol non-fermenting E. coli on the edible part of fish agrees with the findings of Akoachere et al.43, from Cameron, and Ribeiro et al.44, from Brazil who previously recovered E. coli from fish destined for human consumption. The report of Anyanwu et al.40, from Nigeria have also shown 13.4% prevalence of SN-F E. coli in other food animals (broiler chicken, cattle and pigs), among which 0.51% of them were E. coli O157.
The 7.3% occurrence of SN-F E. coli strains in diarrheic patients is lower than the systematic review and meta-analysis findings of Zenebe et al.49, who reported 25% pooled prevalence of a generic E. coli in diarrheic children under five years in Ethiopia, and 15.3% prevalence and 28.9% isolation rate of E. coli O157 related diarrhea in children under five years from Eastern Ethiopia26, and Bahir Dar town of northern Ethiopia27, respectively.
The occurrence of highly virulent MDR E. coli strains in humans may be related to unhygienic handling of animal products including fish or consumption of raw or undercooked products. Likewise, the reports of Haenen et al.50, Santos and Vieira51, and Mekonnen et al.52, showed unhygienic handling and consumption of raw or undercooked infected fish may pose the risk of infection in susceptible individuals. Similarly, Wiriyaprom et al.53, have reported 8.82% prevalence of SN-F Shiga toxin-producing E. coli (STEC) isolated from goats in Thailand; among which only 0.77% was E. coli O157.
The majority, (62.1%) of the strains have virulence determinants encoding for adhesion, host invasion, toxin production, and promoting survival of the pathogen. In addition, genes encoding for deterioration of metabolic regulation and iron uptake of the susceptible hosts were detected in majority of the isolates retrieved from the three sample sources, implying that the strains are sharing many virulence characteristics. A similar study in Mozambique also showed the coexistence of different combinations of two or more virulence genes encoding for a single virulence factor in E. coli strains retrieved from ready to eat food items54.
Sorbitol non-fermenting E. coli strains carrying virulence determinant traits were detected in all the study lakes, all health facilities and fish sampled from lakes Koka and Koftu. Among them, a higher proportion, (38.9%; n = 7) was detected from water samples; suggesting that the pathogen is exceedingly disseminated across the aquatic ecosystem in the study areas.
The higher prevalence of virulent strains in lake water implies that fish can be easily infected with such virulent strains from the aquatic environment, and the pathogens can be disseminated among the fish population in that ecosystem and become a potential public health threat, especially among raw fish consumers. Previous reports also showed that if the water bodies are contaminated with pathogenic organisms, fish can be easily infected from its immediate environment55.
The new SN-F E. coli strains with unknown O-antigens, O-unknown: H28 and O-unknown: H40, from fish and human, respectively might be mutant progenies of known E. coli strain or newly emerging strains not yet characterized. Moreover, in silico MLST have shown that both the new strains with unknown O-antigen have multiple virulence factors, and O-unknown: H40 is MDR strain. Thus, further robust molecular study is required to determine the lineages of these strains relative to E. coli strains in the sequence database to elucidate fundamental information on the epidemiology and public health implications of the strains. Similar previous studies conducted on bacterial genomics have also shown that comparison of bacterial 16S rRNA gene (the most conserved genetic marker of a bacteria) with a known sequence of related bacteria in the database has emerged as a preferred genetic technique to identify new strains56,57.
Among the ten antimicrobial agents tested for their level of efficacy, azithromycin was effective for 96.6% (n = 28) of the isolates and recognized as the most promising drug for the treatment of infections related to SN-F non-O157 E. coli strains. Studies indicated that azithromycin is the most promising alternative and excellent drug for the treatment of diarrhoeagenic enteric infections caused by E. coli, Shigella, Salmonella, and Campylobacter species58,59,60. Reduced efficacy of ciprofloxacin, chloramphenicol and sulfamethoxazole /trimethoprim to SN-F non-O157 E. coli strains was observed as compared to the efficacy of azithromycin.
The lower efficacy of ciprofloxacin and sulfamethoxazole /trimethoprim is in agreement with the current report of Yasmin et al.61, from Pakistan, who reported 93% and 92% resistance of E. coli strains to the two drugs, respectively. Other study reported a better and moderate efficacy of ciprofloxacin and sulfonamides to E. coli strains recovered from neonates in China62. On the other hand, the low efficacy of chloramphenicol to SN-F non-O157 E. coli strains is lower than the finding of Ashenafi et al.63, who reported 27.3% efficacy of the drug to E. coli O157 retrieved from raw cow milk in central Ethiopia.
Conversely, in the present study, it was noted that all isolates are resistant to five antimicrobial agents, namely: ampicillin, cefotaxime, ceftazidime, meropenem and tetracycline; and besides all human isolates were resistant to nalidixic acid. These findings are comparable with the recent findings of Dejene et al.64, from Ethiopia, who have reported an absolute resistance of E. coli O157 to ampicillin, and Yasmin et al.61, from Pakistan, who have shown 91% resistance of other E. coli strains to the same drug. Thus, based on the result of the present study and previous findings, avoiding the use of ampicillin for the treatment of E. coli O157 and other E. coli infections is recommended. The absolute resistance to cefotaxime, ceftazidime and meropenem observed in the present study is also comparable with the finding of Yasmin et al.61, who reported a 98%, 86% and 71% resistance of E. coli strains to the same drugs, respectively.
The study revealed the widespread occurrence of MDR SN-F E. coli strains along the fish supply chain in central Oromia, Ethiopia, where the strains were resistant to 87.5% (seven-out of eight) antimicrobial classes. Based on in silico MLST, the majority (55.6%) of the virulent strains, have multiple acquired antimicrobial resistant traits and one or more genes encoding for them. This implies the higher AMR profile of virulent strains retrieved from humans as compared to fish and water isolates. Similar report from the city of Maputo, Mozambique have also shown that MDR E. coli strains was detected in drinking water54. Another study reported MDR Gram-negative rods, including E. coli as a public health threat globally65. Thus, as MDR Gram-negative bacteria, including E. coli, represent a global public health challenge66, it is crucial to note that these resistant organisms may be disseminated along the fish supply chain, and become a potential public health threat. Moreover, genes encoding for mutational resistance against ciprofloxacin and nalidixic acid, and extended spectrum beta-lactamase (ESBL) against aztreonam, cefepime, ceftaxime, ceftazidime, and ceftriaxone were detected. Besides, previous reports have shown that E. coli strains may serve as reservoir of antimicrobial resistant genes which can be transferred to other pathogenic strains primarily through horizontal gene transfer29, and become a serious public health concern67. Therefore, since they are highly prone to exchanging genetic material68, the risk of horizontal gene transfer to other pathogenic or non-pathogenic E. coli strains and dissemination of mutant strains along the fish supply chain may be inevitable.
In general, although resistance of E. coli to older antibiotics such as tetracycline and ampicillin and those that acquire resistance through plasmid transfers may not be surprising, resistance to the last-resort drugs like meropenem which are used for the treatment of human infections caused by MDR enteric pathogens and classified by WHO as high priority critically important antimicrobials69, is a potentially discerning public health threat; and rational use of drugs are critically important.
Conclusion
The study revealed the occurrence of considerable proportion of virulent and multidrug resistant sorbitol non-fermenting E. coli strains in Lake Water, fish and humans in Central Oromia, Ethiopia. We also reported new SN-F E. coli strains with unknown O-antigen from fish and stool samples. However, the strains detected from the three sample sources are not genetically linked; and none of them were E. coli O157. The observed occurrence of virulent and multidrug resistant E. coli strains and the genetic linkage among strains originated from the same samples compounded with the poor fish hygienic production and raw fish consumption habit of the community suggest the potential risk of dissemination of the organisms along the supply chain and the transmission pathways from contaminated water-to-fish-to-humans. Further robust molecular study is required to establish the epidemiology of sorbitol non-fermenting E. coli strains along fish supply chain and characterize the new strains with unknown O-antigen to guide public health intervention programs to ensure fish safety in Ethiopia in general and in central Oromia in particular.
Data availability
The DNA Sequence datasets generated during the current study are available in the National Library of Medicine repository, with accession numbers: SUB13951691, SUB13951614, SUB13951611, SUB13951598, SUB13951583, SUB13951579, SUB13951565, SUB13951559, SUB13951552, SUB13951544, SUB13951396, SUB13951541, SUB13951294, SUB13951282, SUB13951298, SUB13951274, SUB13951308, SUB13951320, SUB13951382, SUB13951387, SUB13951393, SUB13942239, SUB13951735, SUB13951753, SUB13951569, SUB13949288, SUB13949315, SUB13949345, and SUB13949358. All other data were incorporated to the manuscript.
References
World Health Organization (WHO). Who Estimates of The Global Burden of Foodborne Diseases. Foodborne Disease Burden Epidemiology Reference Group 2007–2015 (2015).
Hoffmann, S. et al. Attribution of global foodborne disease to specific foods: Findings from a World Health Organization structured expert elicitation. PLoS ONE 12(9), e0183641 (2017).
Troeger, C. et al. Estimates of the global, regional, and national morbidity, mortality, and etiologies of diarrhoea in 195 countries: A systematic analysis for the Global Burden of Disease Study 2016. Lancet Infect. Dis. 18, 1211–1228 (2018).
Linscott, A. J. Food-borne illnesses. Clin. Microbiol. Newsl. 33(6), 41–45 (2011).
Puig, P. Y., Leyva, C. V., Maceo, R. B. & Muñoz, A. P. Y. Bacterial agents associated with outbreaks of food-borne diseases in Havana, 2006–2010. Rev. Cubana Hig. Epidemiol. 51, 74–83 (2013).
Amezquita-Lopez, B. A., Quinones, B., Soto-Beltran, M., Lee, B. G., Yambao, J. C. & Lugo-Melchor, O. Y. et al. Antimicrobial resistance profiles of Shiga toxin-producing Escherichia coli O157 and non O157 recovered from domestic farm animals in rural communities in Northwestern Mexico. Antimicrob. Resist. Infect. Control. 5, 1 (2018).
Augustin, J. C. et al. Contribution of foods and poor food-handling practices to the burden of foodborne infectious diseases in France. Foods. 9, 1644 (2020).
Petronillah, R., Robert, K., John, V. & Nyoni, S. Isolation and identification of pathogenic bacteria in edible fish: A case study of Fletcher Dam in Gweru, Zimbabwe. Int. J. Sci. Res. 2, 269–273 (2013).
Abowei, J. & Briyai, O. A review of some bacterial diseases in Africa culture fisheries. Asian J. Med. Sci. 3, 206–217 (2011).
Acha, P. & Szyfres, B. Zoonoses and communicable diseases common to man and animals: Bacterioses and mycoses 3rd ed. Scientific and Technical Publication No 580, Pan American Health Organization, Regional Office of the WHO, USA. 384 (2003).
Costa, R. A. Escherichia coli in seafood: A brief overview. Adv. Biosci. 4(03), 450 (2013).
Soliman, M., Khalil, R., Saad, T., El-Gamal, M. & Gebril, A. Isolation and identification of E. coli from cultured freshwater fish. J. Arab. Aquat. Sci. 5, 8 (2010).
Nataro, J. P. & Kaper, J. B. Diarrheagenic Escherichia coli. Clin. Microbiol. Rev. l 11, 142–201 (1998).
Croxen, M. A. et al. Recent advances in understanding enteric pathogenic Escherichia coli. Clin. Micro. Rev. 26(4), 822–880 (2013).
Merchant, L. E. et al. Characterization of antibiotic-resistant and potentially pathogenic Escherichia coli from soil fertilized with litter of broiler chickens fed antimicrobial-supplemented diets. Can. J. Microbiol. 58(9), 1084–1098 (2012).
Cui, Y., Li, D. F. & Yang, R. F. Shiga toxin-producing Escherichia coli O104:H4: an emerging important pathogen in food safety. Chinese Sci. Bull. 58(14), 1625–1631 (2013).
Bii, C. C., Taguchi, T. T., Ouko, L. W., Muita, N. & Kamiya, S. Detection of virulence related genes by multiplex PCR in multidrug-resistance diarrheagenic E. coli isolates from Kenya and Japan. Epidemiol. Infect. 133, 627–633 (2005).
Dewey-Mattia, D., Manikonda, K., Hall, A. J., Wise, M. E. & Crowe, S. J. Surveillance for foodborne disease outbreaks-United States, 2009–2015. MMWR Surveill. Summ. 67, 1–11 (2018).
Chen, L. et al. An 11-year analysis of bacterial foodborne disease outbreaks in Zhejiang Province, China. Foods 11, 2382 (2022).
Mina, Z. et al. Zoonotic diseases of fish and their prevention and control. Vet. Q. 42(1), 95–118 (2022).
Titilawo, Y., Obi, L. & Okoh, A. Occurrence of virulence gene signatures associated with diarrhoeagenic and non-diarrhoeagenic pathovars of Escherichia coli isolates from some selected rivers in South-Western Nigeria. BMC Microbiol. 15(204), 1–14 (2015).
Gazu, L. et al. Foodborne disease hazards and burden in Ethiopia: A systematic literature review, 1990–2019. Front. Sustain Food Syst. 7, 1058977 (2023).
Bedane, T. D., Agga, G. E. & Gutema, F. D. Hygienic assessment of fish handling practices along production and supply chain and its public health implications in Central Oromia, Ethiopia. Sci. Rep. 12, 13910 (2022).
Ayalew, A. Prevalence of Escherichia coli O157:H7 in foods of animal origin in Ethiopia: A meta-analysis. Cog. Food Agric. 5(1) (2019).
Tilahun, A. & Engdawork, A. Isolation, Identification and antimicrobial susceptibility profile of E. coli O157:H7 from Fish in Lake Hawassa, Southern Ethiopia. Int. J. Vet. Sci. Technol. 3(1), 013–019 (2019).
Getaneh, D. K., Hordofa, L. O., Ayana, D. A., Tessema, T. S. & Regassa, L. D. Prevalence of Escherichia coli O157:H7 and associated factors in under-five children in Eastern Ethiopia. PLoS ONE. 16(1), e0246024 (2021).
Adugna, A., Kibret, M., Abera, B., Nibret, E. & Adal, M. Antibiogram of E. coli serotypes isolated from children aged under five with acute diarrhea in Bahir Dar town. Afr. Health Sci. 15(2), 656–664 (2015).
Gutema, F. D. et al. Occurrence, molecular characteristics, and antimicrobial resistance of Escherichia coli O157 in cattle, beef, and humans in Bishoftu Town, Central Ethiopia. Foodborne Pathol. Dis. 18(1), 1–7 (2021).
Poirel, L., Madec, J. Y., Lupo, A., Schink, A. K., Kieffer, N., Nordmann, P. & Schwarz, S. Antimicrobial resistance in Escherichia coli. Microbiol. Spectr. 6(4) (2018).
Central Statistical Authority (CSA). The 2007 Population and Housing Census of Ethiopia; Results for Oromia Region, 1, Archived November 13, 2011, at the Wayback Machine (2012).
ESZOLF, East Shewa zone Office of Livestock and Fisheries. Major lakes and towns of east Shewa zone where fishes are potentially harvested and consumed. Unpublished information obtained through official communication (2021).
Mc Evoy, J. et al. The prevalence and spread of E. coli O157:H7 at a commercial beef abattoir. J. Appl. Microbiol. 95, 256–266 (2003).
Elder, R. et al. Correlation of Enterohemorrhagic E. coli O157 prevalence in feces, hides and carcasses of beef cattle during processing. Proc. Natl. Acad. Sci. 97, 2999–3003 (2000).
European food safety authority. Pathogenicity assessment of Shiga-toxin producing Escherichia coli (STEC) and the public health risk posed by contamination of food with STEC. (2020).
Müller, E. E. & Ehlers, M. M. Biolog identification of non-sorbitol fermenting bacteria isolated on E. coli O157 selective CT-SMAC agar. Water SA. 31(2), 247 (2005).
DeBoer, E. & Heuvelink, A. Methods for the detection and isolation of shiga-toxin producing Escherichia coli. Symp. Ser. Soc. Appl. Microbiol. 29, 133–143 (2000).
Zhou, Z., Alikhan, N. F., Mohamed, K., Fan, Y., Agama, Study, G., Achtman, M. The EnteroBase user’s guide, with case studies on Salmonella transmissions, Yersinia pestis phylogeny, and Escherichia core genomic diversity. Genome Res. 30, 138–152 (2020).
CLSI. Clinical and laboratory standards institute, antimicrobial susceptibility testing updates. Ochsner Medical Center, New Orleans. Ochsner Health. 8 (2022).
Feldgarden, M. et al. Validating the AMRFinder tool and resistance gene database by using antimicrobial resistance genotype-phenotype correlations in a collection of isolates. Antimicrob. Agents Chemother. 63, 483 (2019).
Anyanwu, U. M. et al. Sorbitol non-fermenting Escherichia coli and E. coli O157: Prevalence and antimicrobial resistance profile of strains in slaughtered food animals in Southeast Nigeria. Access Microbiol. 4, 00043 (2022).
Mekonnen, M., Assefa, F., Lemma, B., Haaren, V. & C., Casper, P.,. Assessing the occurrence of waterborne pathogens in Lake Ziway and drinking water system of Batu (Ziway) Town, Ethiopia. Ethiop. J. Health Dev. 28(2), 120 (2014).
Dissasa, G., Lemma, B. & Mamo, H. Isolation and identification of major bacteria from three Ethiopian rift valley lakes live and processed fish, and water samples: implications in sanitary system of fish products. BMC Vet. Res. 18, 439 (2022).
Akoachere, J. F., Bughe, R. N., Oben, B. O., Ndip, L. M. & Ndip, R. N. Phenotypic characterization of human pathogenic bacteria in fish from the coastal waters of South West Cameron: Public health implications. Rev. Environ. Health 24, 147–156 (2009).
Ribeiro, L. F. et al. Shiga toxigenic and enteropathogenic Escherichia coli in water and fish from pay-to-fish ponds. Lett. Appl. Microbiol. 62, 216–220 (2016).
Manna, S. K. et al. Serogroup distribution and virulence characteristics of sorbitol-negative Escherichia coli from food and cattle stool. J. Appl. Microbiol. 108, 658–665 (2010).
Haile, A. B. & Getahun, T. K. Isolation and identification of Escherichia coli and Edwardsiella tarda from fish harvested for human consumption from Zeway Lake, Ethiopia. Afr. J. Microbiol. Res. 12(20), 476–480 (2018).
Yohans, H., Mitiku, B. A. & Tassew, H. Levels of Escherichia coli as bio-indicator of contamination of fish food and antibiotic resistance pattern along the value chain in Northwest Ethiopia. Vet. Med. Res. Rep. 13, 299–311 (2022).
Marijani, E. Prevalence and Antimicrobial resistance of bacteria isolated from marine and freshwater fish in Tanzania. Int. J. Microbiol. (2022).
Zenebe, T., Mitiku, M. & Alem, Y. Prevalence of Escherichia coli in under-five children with diarrhea in Ethiopia: A systematic review and meta-analysis. Int. J. Microbiol. (2020).
Haenen, O. L., Evans, J. J. & Berthe, F. Bacterial infections from aquatic species: Potential for and prevention of contact zoonoses. Rev Sci. Tech. Off. Int. Epiz. 32(2), 497–507 (2013).
Santos, C. A. M. L. & Vieira, R. H. S. F. Bacteriological hazards and risks associated with seafood consumption in Brazil. Rev. Inst. Med. Trop. Sao. Paulo. 55, 219–228 (2013).
Mekonnen, S., Gezahegne, M. & Lemma, A. Major fish-borne bacterial and parasitic zoonoses in Ethiopia. A review. Int. J. Fauna Biol. Stud. 6(4), 50–58 (2019).
Wiriyaprom, R., Ngasaman, R., Kaewnoi, D. & Prachantasena, S. Prevalence and virulent gene profiles of sorbitol non-fermenting Shiga toxin-producing Escherichia coli isolated from goats in Southern Thailand. Trop. Med. Infect. Dis. 7, 357 (2022).
Salamandane, A., Alves, S., Chambel, L., Malfeito-Ferreira, M. & Brito, L. Characterization of Escherichia coli from water and food sold on the streets of Maputo: Molecular typing, virulence genes, and antibiotic resistance. Appl. Microbiol. 2, 133–147 (2022).
Fouz, B., Toranzo, A. E., Milan, M. & Amaro, C. Evidence that water transmits the disease caused by the fish pathogen, Photobacterium damselae subsp. damselae. J. Appl. Microbiol. 88, 531–535 (2000).
Magray, M. S., Kumar, A., Rawat, A. K. & Srivastava, S. Identification of Escherichia coli through analysis of 16S rRNA and 16S–23S rRNA internal transcribed spacer region sequences. Bioinformation. 6(10), 370–371 (2011).
Olowe, B. M., Oluyege, J. O., Famurewa, O., Ogunniran, A. O. & Adelegan, O. Molecular identification of Escherichia coli and new emerging enteropathogen, Escherichia fergusonii, from drinking water sources in Ado-Ekiti, Ekiti State, Nigeria. J. Microbiol. Res. 7(3), 45–54 (2017).
Andrea, T. One Health and development (2012).
Lubbert, C. Antimicrobial therapy of acute diarrhea: A clinical review. Expert Rev. Anti. Infect. Ther. 14, 193–206 (2016).
Gomes, C., Martinze-Puchol, S. & Palma, N. Macrolide resistance mechanisms in enterobacteriaceae: focus on azithromycin. Crit. Rev. Microbial. 43, 1–30 (2017).
Yasmin, S., Karim, A. M., Lee, S. H. & Zahra, R. Temporal variation of Meropenem resistance in E. coli isolated from sewage water in Islamabad, Pakistan. Antibiotics. 11, 635 (2022).
Wu, D., Ding, Y., Yao, K., Gao, W. & Wang, Y. Antimicrobial resistance analysis of clinical Escherichia coli isolates in neonatal ward. Front. Pediatr. 9, 1 (2021).
Ashenafi, A., Dereje, E. & Haben, F. Isolation and antimicrobial susceptibility profile of Escherichia coli O157: H7 from raw milk of dairy cattle in Holeta District, Central Ethiopia. Hindawi Int. J. Microbiol. 8 (2020).
Dejene, H., Abunna, F., Tuffa, C. A. & Gebresenbet, G. Epidemiology and antimicrobial susceptibility pattern of E. coli O157:H7 along dairy milk supply chain in central Ethiopia. Vet. Med. Res. Rep. 13, 131–142 (2022).
Ejaz, H. et al. Molecular epidemiology of extensively drug-resistant mcr encoded colistin-resistant bacterial strains co-expressing multifarious-lactamases. Antibiotics. 10, 467 (2021).
Mdegela, R. H. et al. Antimicrobial use, residues, resistance and governance in the food and agriculture sectors, Tanzania. Antibiotics. 10, 454 (2021).
Aworth, M. K., Kwaga, J., Okolocha, E., Mba, N. & Thakur, S. Prevalence and risk factors MDR E. coli among poultry workers in the federal capital territory, Abuja, Nigeria. PLoS ONE. 14(11), 25379 (2019).
Amador, P., Fernandes, R., Duarte, I., Brito, L. & Prudêncio, C. In vitro transference and molecular characterization of blaTEM genes in bacteria isolated from Portuguese ready-to-eat foods. World J. Microbiol. Biotechnol. 27, 1775–1785 (2011).
Loest, D. et al. Carbapenem-resistant Escherichia coli from shrimp and salmon available for purchase by consumers in Canada: A risk profile using the Codex framework. Epidemiol. Infect. 150(e148), 1–20 (2022).
Acknowledgements
The authors would like to acknowledge Ministry of Education of Ethiopia, Addis Ababa University postgraduate research funding, and Salale University for the financial support; Bishoftu town, Lome Adama and Batu district office of livestock development and fisheries, Bishoftu Hospital, Koka and Batu health Centers, the fisher men, and diarrheic patients for their generous support, sample provision, and substantial information provided during data collection; Michigan State University for funding the antimicrobial discs; BRIGHTcore (Belgian National Reference Centre for STEC/VTEC) for their collaboration in performing E. coli O157 Latex agglutination test, and whole genome sequencing.
Funding
The study was funded by Ministry of Education, Addis Ababa University thematic and postgraduate research funding, and Salale University.
Author information
Authors and Affiliations
Contributions
All the authors made significant contributions to the manuscript and agreed to its publication. The contributions were performed in the following order: Conceptualization, T.D.B. and F.D.G.; Study design, T.D.B. and F.D.G.; data collection, T.D.B., M.T., and E.S., Conventional laboratory analysis, T.D.B., M.T. and E.S.; data analysis and interpretation, T.D.B., F.D.G. and B.M.; original draft preparation, T.D.B.; writing-critical review and editing, F.D.G., B.M., F.A., H.W., F.T.; supervision, F.D.G. and B.M.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bedane, T.D., Megersa, B., Abunna, F. et al. Occurrence, molecular characterization, and antimicrobial susceptibility of sorbitol non-fermenting Escherichia coli in lake water, fish and humans in central Oromia, Ethiopia. Sci Rep 14, 12461 (2024). https://doi.org/10.1038/s41598-024-61810-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-61810-z