Abstract
The molecular mechanisms that govern differential T cell development from CD4+CD25-conventional T (Tconv) into CD4+CD25+ forkhead-box-P3+ (FoxP3+) inducible regulatory T (iTreg) cells remain unclear. Herein, we investigated the relative contribution of protein kinase A (PKA) in this process. Mechanistically, we found that PKA controlled the efficiency of human iTreg cell generation through the expression of different FoxP3 splicing variants containing or not the exon 2. We found that transient PKA inhibition reduced the recruitment of cAMP-responsive element-binding protein (CREB) on regulatory regions of the FoxP3 gene, a condition that is associated with an impaired acquisition of their suppressive capacity in vitro. To corroborate our findings in a human model of autoimmunity, we measured CREB phosphorylation and FoxP3 levels in iTreg cells from treatment-naïve relapsing–remitting (RR)-multiple sclerosis (MS) subjects. Interestingly, both phospho-CREB and FoxP3 induction directly correlated and were significantly reduced in RR-MS patients, suggesting a previously unknown mechanism involved in the induction and function of human iTreg cells.
Similar content being viewed by others
Introduction
Regulatory T (Treg) cells are a specialized subset of T lymphocytes that arise from thymocytes during intrathymic maturation, with a key role in maintaining immunological self-tolerance, due to their ability to restrain inflammatory responses1,2,3. To date, several distinct lineages of T cells with regulatory properties have been identified4,5,6. Among these, the most extensively studied population of Treg cells comprises CD4+ T cells characterized by their high expression of the α-subunit of the interleukin 2 (IL-2) receptor and the presence of the master transcription factor forkhead box P3 (FoxP3)7,8. Although the majority of Treg cells derive from the thymus (thymic Treg, tTreg cells), they can differentiate also in the periphery (pTreg) from conventional T (Tconv) cells, upon sub-immunogenic antigen presentation9. Furthermore, induced Treg (iTreg) cells can be obtained in vitro by T cell receptor (TCR) stimulation of Tconv cells in the presence or absence of exogenous cytokines10,11,12,13. In the process of Treg cell induction, IL-2 and TGF-β signalling have been shown to support FoxP3 expression also promoting the proliferative capability of Treg cells14,15,16,17,18,19. Nonetheless, recent evidence has unveiled the crucial role of TCR engagement strength in the process of iTreg cell generation; indeed it has been shown that FoxP3 expression can be stably induced in Tconv cells through suboptimal in vitro TCR stimulation without the addition of exogenous cytokines10,11,12, consequently leading to the generation of fully suppressive iTreg cells10,11,12,20.
Compelling evidence suggests that the transcription factor FoxP3 not only serves as a Treg lineage-specific molecule but is also indispensable for their proper differentiation, stability and the acquisition of suppressive functions21,22,23. The human FoxP3 gene consists of 11 coding exons and includes a promoter region along with three conserved non-coding sequences (CNS) within the locus: CNS1, CNS2, and CNS324. These CNS elements are crucial functional enhancers for both the induction and stabilization of FoxP3 expression. CNS1 encompasses the TGF-β response element, which plays a role in the development of extrathymic Treg cells and the promotion of mucosal immune tolerance. CNS2 is responsible for maintaining FoxP3 stability in response to TCR stimulation, while CNS3 appears to influence the threshold of TCR stimuli required for FoxP3 expression25.
To date, several FoxP3 splicing variants have been described in human Treg cells26 and among them, those containing the sequence corresponding to exon 2 (FoxP3-E2) are pivotal to confer their suppressive capability and stability10,12,27. Indeed, the absence of FoxP3-E2 is associated with impaired Treg cell activity, uncontrolled effector T cell activation leading to the development of autoimmune diseases in both mice and humans10,12,27,28.
In this context, it has amply documented that the number and function of circulating Treg cells, the capacity to generate iTreg cells, as well as the FoxP3-E2-related suppressive activity of iTreg cells are all impaired in human autoimmune disorders, including relapsing–remitting multiple sclerosis (RR-MS)10,21,29,30, an immune-mediated disease of the central nervous system31.
However, the molecular pathways involved in the control of human FoxP3 gene expression remain incompletely understood.
FoxP3 transcription is known to be controlled by the recruitment of several transcription factors into FoxP3 promoter or to the CNS regions10. Among all these factors, the cyclic adenosine 3′,5′-monophosphate (cAMP) responsive element-binding protein (CREB) has attracted increasing interest for its possible role in Treg cell differentiation and function32,33,34. Indeed, while it has been shown that CREB is crucial to driving FoxP3 transcription32,33, another research group revealed that it is dispensable for FoxP3 expression and negatively regulates the survival of iTreg cells in mouse models of autoimmunity34. However, the role of the cAMP/PKA/CREB pathway in the control of FoxP3 expression to generate fully functional iTreg cells has not been investigated in human settings.
Since the strength of TCR signalling during T cell activation promoted the engagement of several signalling pathways including those associated with cAMP/PKA/CREB35,36, herein, we explored the involvement of PKA in the regulation of FoxP3 splicing variants, in the immunometabolic asset and in the suppressive function of human iTreg cells. Further, we investigated in depth the relative contribution of CREB in FoxP3 expression during iTreg cell differentiation from Tconv cells obtained from human RR-MS subjects.
Results
Low TCR engagement activates PKA favouring CREB binding on FoxP3 gene in human Tconv cells
To assess whether PKA is involved in the early events induced by low TCR stimulation, freshly isolated human CD4+CD25- Tconv cells were pre-treated for 1 h with the myristoylated PKA specific inhibitor fragment 14–22 (mPKAI)37,38 or not (mPKAI-Tconv and CTR-Tconv, respectively) and then stimulated or not with anti-CD3/CD28 microbeads (see the experimental procedure in Fig. S1). We found that transient inhibition of PKA reduced both catalytic activity of PKA and CREB phosphorylation in TCR-stimulated mPKAI-Tconv cells (Fig. 1a,b and Fig. S2). Further, we assessed the involvement of PKA in the regulation of FoxP3 gene expression during Tconv cell differentiation towards iTreg cells. To this end, we performed a chromatin immunoprecipitation (ChIP) assay to evaluate CREB binding on FoxP3 regulatory elements: promoter, non-coding conserved sequence (CNS)2 and CNS3 regions, respectively. ChIP assay revealed that upon Tconv cell stimulation, CREB was rapidly recruited on the regulatory elements of FoxP3, particularly on CNS2 region (Fig. 1c). Of note, transient PKA inhibition was significantly associated with reduced CREB binding on the regulatory elements of FoxP3 (Fig. 1c). Together, these findings suggest that during the early steps of iTreg cell induction, TCR-mediated stimulation of Tconv cells results in PKA activation, which in turn promotes CREB recruitment on FoxP3 regulatory elements39.
Low TCR stimulation activates PKA in human Tconv cells. (a) Scatter plots showing PKA activity in human mPKAI-Tconv and CTR-Tconv cells TCR-stimulated for 5 and 15 min. Data are shown from four independent experiments in duplicates (n = 8). (b) Left, immunoblot showing phosphorylated (p) and total CREB in human mPKAI-Tconv and CTR-Tconv cells TCR-stimulated or not for 15 min. Right, relative densitometric quantitation of p-CREB in the aforementioned experimental conditions. Data are shown from nine independent experiments (n = 9); uncropped blots are presented in Supplementary Fig. S2. (c) ChIP assay for CREB on FoxP3 promoter, CNS2 and CNS3 regions of mPKAI-Tconv and CTR-Tconv cells TCR-stimulated for 10 min. The horizontal line (bracket ± SEM) indicates the percent of input from a control ChIP (Ab:non-immune serum). Data are shown from three independent experiments in duplicates (n = 6). Independet experiments refer to different individuals.
PKA regulates FoxP3 expression during Tconv differentiation towards iTreg cells
To evaluate whether the cAMP/PKA/CREB axis participated in the regulation of FoxP3 gene expression, we measured, by quantitative real time PCR, total FoxP3 splicing forms (FoxP3-All) and its variants containing the Exon2 (FoxP3-E2) during iTreg cell induction, upon suboptimal TCR stimulation, to sustain a more effective induction of FoxP3, as previously demonstrated11,40,41. To this aim, mPKAI-Tconv and CTR-Tconv cells were stimulated via TCR for 24 and 36 h, respectively. While no significant difference was observed in terms of FoxP3-All mRNA levels in both conditions, a significant reduction of FoxP3-E2 mRNA levels at 24 h upon TCR stimulation in mPKAI-Tconv cells was found (Fig. 2a), suggesting the requirement of PKA for the selective FoxP3-E2 generation. We also evaluated at protein level the expression of FoxP3-All and FoxP3-E2 by Western Blot. Specifically, mPKAI-Tconv and CTR-Tconv cells were stimulated via TCR for 36 h, to measure the splicing forms of FoxP3 protein, using specific monoclonal antibodies (mAbs). Our results revealed that the induction of both FoxP3-All and FoxP3-E2 proteins was affected in mPKAI-Tconv cells (Fig. 2b and Fig. S3), differently from what has been observed at mRNA level.
Inhibition of PKA impairs FoxP3 induction. (a) Scatter plots showing FoxP3-All (left) and FoxP3-E2 (right) mRNA levels in mPKAI-Tconv and CTR-Tconv cells TCR-stimulated or not for 24 or 36 h, respectively. Data are shown from four independent experiments in duplicates (n = 8). (b) Left, immunoblot analysis of FoxP3-All, FoxP3-E2 and ERK1/2 in mPKAI-Tconv and CTR-Tconv cells TCR-stimulated for 36 h. Right, relative densitometric quantitation of FoxP3-All or FoxP3-E2 normalized on ERK 1/2 in the aforementioned experimental conditions. Data are shown from thirteen independent experiments (n = 13). (c) Left, immunoblot analysis of p-STAT5 and p-S6 in human Tconv cells, as described in B. Right, relative densitometric analysis of p-STAT5 and p-S6 normalized on their total proteins, respectively. Data are shown from five independent experiments in duplicates (n = 10). Independet experiments refer to different individuals. All the uncropped blots are presented in Supplementary Fig. S3.
Since IL-2R/STAT-5 and mechanistic target of rapamycin (mTOR) signalling pathways are also involved in iTreg cell differentiation and function18,42, we analysed phosphorylation of STAT-5 and S6 proteins (a mTOR downstream target) in the above mentioned experimental conditions. Biochemical analysis of these proteins revealed that PKA inhibition did not influence their phosphorylation (Fig. 2c and Fig. S3), suggesting that PKA selectively controls the expression of FoxP3 both at mRNA and protein level during iTreg cell differentiation.
Impact of PKA on the immunometabolic asset and phenotype of Tconv cells during their differentiation towards iTreg cells and on their suppressive activity
Next, we explored whether cAMP/PKA/CREB pathway influences glycolytic immunometabolism, phenotype and suppressive function of iTreg cells. It has been previously shown that glycolytic pathway was necessary for the induction of human iTreg cells in vitro43, and in particular for the expression of FoxP3-E2 splicing variants10,12. To this aim, we measured extracellular acidification rate (ECAR), an indicator of glycolysis, in mPKAI-Tconv and CTR-Tconv, during iTreg cell generation. Seahorse analysis revealed that transient inhibition of PKA significantly affected glucose metabolism, as testified by reduced glycolysis, maximal glycolysis and glycolytic capacity (Fig. 3a, left and right panels), suggesting that PKA acts as a positive determinant for glycolytic metabolism during Tconv differentiation towards iTreg cells.
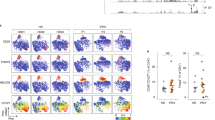
PKA inhibition affects the immunometabolic asset and phenotype of Tconv cells during their diffentiation towards iTreg cells and their suppressive function (a) Left, kinetic profile of ECAR in human mPKAI-Tconv and CTR-Tconv cells TCR-stimulated or not for 12 h. ECAR was measured in real time, under basal conditions and in response to glucose, oligomycin, and 2-DG. Data are shown from three independent experiments at least in technical duplicates (n = 10). Right, parameters of the glycolytic pathway were calculated from the ECAR profile of Tconv cells in the above-mentioned conditions. Data are expressed as mean ± SEM of three different measurements, each of them in ten replicates (n = 30). (b) Representative dot plots (left) and cumulative data (right) of CD25, FoxP3-All and FoxP3-E2 in mPKAI-Tconv and CTR-Tconv cells. Data are shown from eleven independent experiments (n = 11). (c) Scatter plots showing the expression of CTLA-4, PD-1 and GITR gated on CD4+FoxP3-All+ (top) or CD4+FoxP3-E2+ (bottom) in mPKAI-Tconv and CTR-Tconv cells. Data are shown as mean ± SEM from four independent experiments in duplicates (n = 8). (d) Left, flow cytometry histograms showing proliferation of CFSE+CD4+ T cells TCR-stimulated for 96 h in vitro and cultured alone (empty curves) or in the presence of various numbers of flow-sorted iTreg from mPKAI-Tconv and CTR-Tconv cells. Numbers in plots indicate the percent of CSFE dilution in CD4+ T cells cultured alone (top left) and co-cultured with iTreg cells (above bracketed lines), as indicated. Right, cumulative data of CD4+ T cell proliferation in the above conditions. Data are shown from four independent experiments in duplicates (n = 8). Independet experiments refer to different individuals.
Furthermore, we assessed whether impaired glycolysis associated with altered expression of the main Treg cell-markers. To this aim, mPKAI-Tconv and CTR-Tconv cells were stimulated for 36 h via low TCR engagement to generate iTreg cells. Flow cytometric analysis revealed that CD25 and FoxP3-All molecules were not affected by PKA inhibition, while a selective reduction of the FoxP3-E2 splicing forms was observed (Fig. 3b). Moreover, we found that mPKAI pre-treatment significantly reduced the expression of the suppressive markers CTLA-4, PD1 and GITR only in FoxP3-E2+ cells (Fig. 3c). Further, transient inhibition of PKA did not affect the expression level of the activation markers such as CD62L, CD69, CD71 and ICOS as well as the proliferation marker Ki-67 (Fig. S4).
Finally, we examined whether reduced FoxP3-E2 expression in iTreg cells, generated from mPKAI-Tconv, associated with an impaired suppressive function in vitro. Thus, iTreg cells were flow sorted on the basis of CD25 surface expression (see gating strategy for iTreg cell isolation in Fig. S5) and tested for their ability to suppress proliferation of TCR-activated CD4+ T cells. We found that flow sorted iTreg derived from mPKAI-Tconv cells displayed a significantly lower capability to suppress the proliferation of TCR-stimulated CD4+ T cells at different cell-to-cell ratios (Fig. 3d). In all, these findings indicate that PKA represents a molecular determinant important for a proper in vitro generation of fully active human iTreg from Tconv cells.
CREB phosphorylation influences FoxP3 induction in Tconv cells from autoimmune RR-MS patients
To investigate the role of cAMP/PKA/CREB axis in an autoimmune disorder such as RR-MS in which number and function of Treg cells are reduced, we measured CREB phosphorylation in Tconv cells during their differentiation towards iTreg cells, in healthy (n = 5) and naïve-to-treatment RR-MS patients (n = 5). Western blot analysis revealed that freshly isolated (basal) and shortly TCR-stimulated Tconv cells from RR-MS subjects displayed significantly lower levels of intracellular p-CREB compared to healthy subjects (Fig. 4a and Fig. S6). In parallel, in the same RR-MS subjects, we found a significant impairment of FoxP3-E2 induction in Tconv cells stimulated for 36 h (Fig. 4b and Fig. S6). Also, we correlated basal p-CREB in ex-vivo Tconv cells with FoxP3 induction upon TCR stimulation in RR-MS and healthy subjects. A positive correlation between basal p-CREB and FoxP3-E2 induction levels at 24 and 36 h post TCR stimulation was observed in the healthy controls; however, in RR-MS patients this correlation was present only at 36 h (Fig. 4c). No correlations between basal p-CREB and FoxP3-All induction were observed in both RR-MS and healthy individuals (data not shown). In all, these findings indicate that CREB phosphorylation is impaired in Tconv cells from RR-MS patients and associated with reduced induction of FoxP3-E2 protein.
Reduced CREB phosphorylation associates with low FoxP3 expression in Tconv cells during their differentiation towards iTreg cells from autoimmune RR-MS subjects. (a) Left, immunoblot analysis of total and p-CREB in human Tconv cells from healthy and RR-MS subjects TCR-stimulated at different time points. Right, relative densitometric quantitation of p-CREB normalized on total CREB in the aforementioned experimental conditions. (b) Left, immunoblot analysis of FoxP3-All, FoxP3-E2 and ERK 1/2, in TCR-stimulated Tconv cells from healthy (n = 5) and RR-MS (n = 5) subjects. Right, relative densitometric quantitation of FoxP3-All and FoxP3-E2 normalized on ERK 1/2 in above conditions. Data are shown as mean ± SEM from five independent experiments (5 healthy and 5 RR-MS subjects) in triplicates (n = 15). All the uncropped blots are presented in Supplementary Fig. S6. (c) Statistical correlation between expression levels of basal p-CREB from ex-vivo Tconv cells with FoxP3-E2 in healthy (n = 5) and RR-MS (n = 5) subjects, in the above mentioned conditions. All the uncropped blots are presented in Supplementary Fig. S6.
Discussion
Herein we investigated the role of cAMP/PKA/CREB pathway in the induction of FoxP3 during the generation of fully functional human iTreg cells. We found a positive role of PKA/CREB axis in regulating iTreg cell fate, as transient inhibition of PKA during iTreg cell differentiation affected glycolysis, expression of FoxP3 splicing forms, Treg-cell markers, and their suppressive capability in vitro.
Compelling evidence has revealed that PKA/CREB pathway is important in T cell immunobiology39, specifically in the differentiation and function of Treg cells. In particular, it has been reported a Treg-specific region in the FoxP3 locus, which contains a binding site for several transcriptional factors, including CREB44. Nevertheless, the role exerted by CREB on FoxP3 expression has been deeply investigated only in mouse models with contrasting results33,34. Specifically, published findings by Kim and Leonard revealed that CREB positively controls FoxP3 transcription through direct interaction with CNS2 region in a demethylated state33. On the other hand, Wang et al.34 have shown that CREB negatively regulates Treg cell survival and by using a CREB-knockout mouse model, have demonstrated that its deficiency increased Treg cell frequency protecting the mice from colitis.
These conflicting results need further investigation to clearly understand whether CREB may play a different role in the regulation and expression of FoxP3 and whether these findings can be also translated to human setting, which still lacks specific experimental evidence.
In this context, according to the results previously published by Kim and co-worker, our findings revealed that CREB was rapidly recruited on regulatory regions (CNS2 and CNS3) of the FoxP3 gene in human Tconv cells upon short TCR stimulation. In particular, we found that transient PKA inhibition, through mPKAI pre-treatment, reduced the recruitment of CREB on regulatory regions of the FoxP3 gene. Although other experimental studies have used H89 as PKA inhibitor, in our experimental setting we choose to utilize mPKAI, as it selectively impairs CREB phosphorylation, unlike H89, which has been shown to affect also additional pathways involved in Treg cell generation and functions, such as S6 phosphorylation37,38.
Together, these findings suggest that during the first phases of human iTreg cell induction, CREB rapidly binds FoxP3 locus into the nucleus, where it cooperates to the formation of a Treg-specific multifactorial transcriptional complex45. In particular, in this study, we characterized the molecular mechanism controlling FoxP3 induction and found that FoxP3-E2 expression is also supported by cAMP/PKA/CREB pathway.
It is well known that transcription of FoxP3 gene leads to the generation of several isoforms and our published study revealed that those containing the exon 2 variants are important in conferring suppressive capability to human iTreg cells10,12. Accordingly, a recent paper by Du et al.28, showed that subjects expressing only the shorter isoform produced by transcripts lacking exon 2 (FoxP3 ΔE2) failed to maintain self-tolerance and developed immunodeficiency, polyendocrinopathy, and enteropathy X-linked (IPEX) syndrome. In this context, it is now clear that the expression of different FoxP3 isoforms and Treg cell functions are closely associated with immunometabolic programs46,47.
Here we found a differential regulation of FoxP3-All and FoxP3-E2 mRNA expression triggered by PKA inhibition; indeed, the reduction in PKA activity may induce an increase in the transcription of some FoxP3 splicing variants in spite of others, whose expression is inhibited (as in the case of FoxP3-E2), thus unbalancing the relative ratios between the different isoforms.
Moreover, despite PKA inhibition in Tconv cells affected both FoxP3-All and FoxP3-E2 protein levels, we found that this treatment significantly reduced only FoxP3-E2 mRNA expression at 24 h, with no significant effects on FoxP3-All mRNA, suggesting that PKA may differently regulate transcriptional and translational mechanisms controlling FoxP3 expression.
In this context, it has been previously described that PKA can differentially modulate both the translation of different mRNAs and their stability48,49,50; therefore, PKA inhibition could induce the accumulation of mRNA encoding for FoxP3, which, however, is not associated with an increase in its protein level, as this treatment could possibly act by reducing the half-life of FoxP351,52.
Cellular metabolism has been shown to be necessary to control the fate of immune cells. During differentiation and activation, immune cells can change their metabolic programs to meet the increased energy demand53. In this context, environmental metabolites and intracellular metabolic intermediates have been recently shown to regulate FoxP3 expression in Treg cells. For instance, glycolysis plays a crucial role in modulating the generation of iTreg cells and expression of FoxP3-E2 splicing forms10,12. Here, we found that inhibition of the PKA was able to impair glycolysis and this was associated with an altered induction of FoxP3-E2 and iTreg-cell function. Thus, based on these findings it is possible to hypothesize that PKA represents a molecular determinant linking glycolytic pathway to FoxP3-E2 expression, necessary for functional activity of iTreg cells.
In our previous work, we observed that T lymphocytes from autoimmune RR-MS patients displayed a reduced engagement of glycolysis associated with an impairment of FoxP3-E2 induction10,54. In line with these results, here we showed that Tconv cells from RR-MS subjects displayed reduced levels of p-CREB, which correlated with low induction of FoxP3 during iTreg cell differentiation. Thus, we revealed that impaired CREB phosphorylation in RR-MS subjects influences a proper iTreg cell generation, through modulation of FoxP3 expression.
In summary, our data unveil that transient perturbation of cAMP/PKA/CREB pathway impinges on human iTreg cell fate and differentiation, as testified by their reduced FoxP3-E2 expression and suppressive capacity. These data suggest that exploring PKA/CREB axis in Tconv cells from subjects with immune disorders may open the way to the identification of a novel deranged molecular pathway leading to the loss of immunological self-tolerance in autoimmunity.
Methods
Study participants
The Institutional Review Board of the Università degli Studi di Napoli “Federico II” approved the study, and peripheral blood was obtained from healthy control and RR-MS subjects after they signed a written informed consent. Healthy donors were matched for age, body mass index and sex with the RR-MS subjects and had no history of inflammatory, endocrine or autoimmune diseases. All the experiments were performed in accordance with Helsinki Declaration.
Tconv cell purification and in vitro iTreg cell generation
Human Tconv cells were purified from peripheral blood mononuclear cells (PBMCs) by negative selection with a human CD4+CD25+ Regulatory T Cell Isolation kit (MiltenyiBiotec) (cell purity ≥ 98%). Cells (1 × 106 cells/well) were pre-incubated (37 °C, 5% CO2 for 1 h) in RPMI 1640 (Gibco) medium supplemented with 100 U/ml penicillin and 100 μg/ml streptomycin (all from Life Technologies), with or without 10 μM of mPKAI (Sigma Aldrich; P9115). After 1 h, cells were washed and activated in the presence or not of anti-CD3/CD28 Dynabeads (Invitrogen), at a density of 0.2 beads/cell for early stimulation (5, 15, 30, 60 min), and at a density of 0.1 bead/cell for 36 h to generate iTreg cells.
PKA activity assay
PKA activity in Tconv cells was detected by a standard sandwich ELISA, following the manufacturer’s instructions (PKA Kinase Activity Assay Kit; Abcam). Results were normalized on the total protein levels.
Western blotting analyses
Cells were lysed in RIPA buffer (Sigma-Aldrich) plus SIGMAFAST Protease Inhibitor (Sigma-Aldrich), and Sigma Phosphatase Inhibitor (Sigma-Aldrich), as previously described10. Nitrocellulose filter membranes were incubated with the following antibodies: pCREB (Ser133) (87G3; Cat: 9198), CREB (48H2; Cat: 9197), pSTAT5 (Tyr694; Cat: 9351), STAT5 (D2O6Y Cat: 94205), pS6 (Ser240/244; Cat: 2215), S6 (5G10; Cat: 2217) (all from Cell Signaling Technology), FoxP3-All (PCH101; Cat: 14-4776-82), FoxP3-E2 (150D/E4; Cat: 14-4774-82) (all from eBioscience) and ERK1/2 (C-9: Cat: sc-514302) (Santa Cruz Biotechnology). Densitometric analysis was performed using ImageJ Software (NIH).
RNA extraction, q-PCR and qRT-PCR
Total RNA was extracted as previously described55,56. Quantitative PCR (qPCR) and qRT-PCR were performed three times in triplicate on a StepOnePlus™ Real-Time PCR System (Applied Biosystems) with TaqMan gene expression assay (Taqman® Universal Master Mix II, Applied Biosystems), using primers for FoxP3 (Hs01092118_g1, Applied Biosystems) and, as internal standard control the 18S gene (Hs03928990_g1, Applied Biosystems).
ChIP assay
Cells were treated as indicated in the legends of the figures and were processed as described56. Pre-cleared chromatin was divided in aliquots and incubated at 4 °C for 16 h with 5 µl of anti-CREB (clone 48H2, Cell Signaling Technology) or non-immune IgG (normal rabbit IgG, Santa Cruz Biotechnology Inc.). Immunoprecipitated DNA was recovered and subjected to qPCR using the specific primers (Table S1), at 250 nM final concentrations.
Seahorse immunometabolic analyses
Metabolic profile (ECAR) processed as previously described10, was evaluated in TCR-activated or not Tconv cells, using an XFe-96 Extracellular Flux Analyzer (Seahorse Bioscience).
Flow cytometry analyses
For the simultaneous evaluation of surface and intracellular molecules, human iTreg cells were stained with the following antibodies: FITC anti-CD4 (Clone RPA-T4, Cat: 561842) (BD Pharmingen), PE-Cy7 anti-CD25 (Clone M-A251, Cat: 560920) (cBD Pharmingen), APC anti-CD152/CTLA-4 (Clone BNI3, Cat: 555855) (BD Pharmingen), PE-Cy5 anti-GITR (Clone REA1007, Cat: 130-116-842) (Miltenyi Biotec), BV421 anti-CD279/PD-1 (Clone EH12.1, Cat: 562516) (BD Horizon), PE anti-FoxP3-All (Clone PCH101, Cat: 560046) (eBioscience) and PE anti-FoxP3-E2 (Clone 150D/E4, Cat: 12-4774-42) (BD Pharmingen). Staining for intracellular factors was performed using fixation and a permeabilization FoxP3 buffer kit (BD Pharmingen), according to the manufacturer's instructions. Samples were acquired using a FACSCanto II (BD Bioscience) and cytofluorimetric analyses were performed using FlowJo Software (FlowJo, LLC).
T cell proliferation assay
For the assessment of iTreg cell suppression function, 36 h TCR-stimulated Tconv cells were flow-sorted with a BD FACSJazz (Becton–Dickinson) on the basis of CD25 expression (see the gating strategy in the Fig. S3), as previously described10.
Statistical analyses
Statistical analysis was carried out by GraphPad Prism Software (GraphPad, California). Comparisons were performed by Kruskal–Wallis test (corrected for Dunn’s multiple comparisons test) in Figs. 1a (for each time point), c (for each region), 2a and 3a. Two-tailed paired Wilcoxon test was performed in Figs. 1b, 2b,c, 3b–d. Two-tailed unpaired Mann–Whitney test was performed in Fig. 4a,b. Pearson’s correlation test was performed in Fig. 4c.
All data are shown as mean ± SEM; P < 0.05 denoting statistical significance.
Data availability
All data generated or analysed during this study are included in this published article [and its supplementary information files].
References
Sakaguchi, S., Yamaguchi, T., Nomura, T. & Ono, M. Regulatory T cells and immune tolerance. Cell 133, 775–787. https://doi.org/10.1016/j.cell.2008.05.009 (2008).
Kawai, K., Uchiyama, M., Hester, J., Wood, K. & Issa, F. Regulatory T cells for tolerance. Hum. Immunol. 79, 294–303. https://doi.org/10.1016/j.humimm.2017.12.013 (2018).
Dikiy, S. & Rudensky, A. Y. Principles of regulatory T cell function. Immunity 56, 240–255. https://doi.org/10.1016/j.immuni.2023.01.004 (2023).
Joosten, S. A. & Ottenhoff, T. H. Human CD4 and CD8 regulatory T cells in infectious diseases and vaccination. Hum. Immunol. 69, 760–770. https://doi.org/10.1016/j.humimm.2008.07.017 (2008).
Santamaria, J. C., Borelli, A. & Irla, M. Regulatory T cell heterogeneity in the thymus: impact on their functional activities. Front. Immunol. 12, 643153. https://doi.org/10.3389/fimmu.2021.643153 (2021).
Bolivar-Wagers, S., Larson, J. H., Jin, S. & Blazar, B. R. Cytolytic CD4(+) and CD8(+) regulatory T-cells and implications for developing immunotherapies to combat graft-versus-host disease. Front. Immunol. 13, 864748. https://doi.org/10.3389/fimmu.2022.864748 (2022).
Grover, P., Goel, P. N. & Greene, M. I. Regulatory T cells: Regulation of identity and function. Front. Immunol. 12, 750542. https://doi.org/10.3389/fimmu.2021.750542 (2021).
Sakaguchi, S. et al. Regulatory T cells and human disease. Annu. Rev. Immunol. 38, 541–566. https://doi.org/10.1146/annurev-immunol-042718-041717 (2020).
Curotto de Lafaille, M. A. & Lafaille, J. J. Natural and adaptive foxp3+ regulatory T cells: More of the same or a division of labor?. Immunity 30, 626–635. https://doi.org/10.1016/j.immuni.2009.05.002 (2009).
De Rosa, V. et al. Glycolysis controls the induction of human regulatory T cells by modulating the expression of FOXP3 exon 2 splicing variants. Nat. Immunol. 16, 1174–1184. https://doi.org/10.1038/ni.3269 (2015).
Sauer, S. et al. T cell receptor signaling controls Foxp3 expression via PI3K, Akt, and mTOR. Proc. Natl. Acad. Sci. U. S. A. 105, 7797–7802. https://doi.org/10.1073/pnas.0800928105 (2008).
Bruzzaniti, S. et al. An immunometabolic pathomechanism for chronic obstructive pulmonary disease. Proc. Natl. Acad. Sci. U. S. A. 116, 15625–15634. https://doi.org/10.1073/pnas.1906303116 (2019).
Shevach, E. M. & Thornton, A. M. tTregs, pTregs, and iTregs: Similarities and differences. Immunol. Rev. 259, 88–102. https://doi.org/10.1111/imr.12160 (2014).
Chen, W. et al. Conversion of peripheral CD4+CD25- naive T cells to CD4+CD25+ regulatory T cells by TGF-beta induction of transcription factor Foxp3. J. Exp. Med. 198, 1875–1886. https://doi.org/10.1084/jem.20030152 (2003).
Hernandez, R., Põder, J., LaPorte, K. M. & Malek, T. R. Engineering IL-2 for immunotherapy of autoimmunity and cancer. Nat. Rev. Immunol. 22, 614–628. https://doi.org/10.1038/s41577-022-00680-w (2022).
Giri, P. S. & Dwivedi, M. Altered expression of nuclear factor of activated T cells, forkhead box P3, and immune-suppressive genes in regulatory T cells of generalized vitiligo patients. Pigment Cell Melanoma Res. 33, 566–578. https://doi.org/10.1111/pcmr.12862 (2020).
Chen, W. TGF-β regulation of T cells. Annu. Rev. Immunol. 41, 483–512. https://doi.org/10.1146/annurev-immunol-101921-045939 (2023).
Chinen, T. et al. An essential role for the IL-2 receptor in Treg cell function. Nat. Immunol. 17, 1322–1333. https://doi.org/10.1038/ni.3540 (2016).
Chen, W. & Konkel, J. E. Development of thymic Foxp3(+) regulatory T cells: TGF-beta matters. Eur. J. Immunol. 45, 958–965. https://doi.org/10.1002/eji.201444999 (2015).
Kretschmer, K. et al. Inducing and expanding regulatory T cell populations by foreign antigen. Nat. Immunol. 6, 1219–1227. https://doi.org/10.1038/ni1265 (2005).
Fontenot, J. D., Gavin, M. A. & Rudensky, A. Y. Foxp3 programs the development and function of CD4+CD25+ regulatory T cells. Nat. Immunol. 4, 330–336. https://doi.org/10.1038/ni904 (2003).
Abbas, A. K. et al. Regulatory T cells: Recommendations to simplify the nomenclature. Nat. Immunol. 14, 307–308. https://doi.org/10.1038/ni.2554 (2013).
Ohkura, N., Kitagawa, Y. & Sakaguchi, S. Development and maintenance of regulatory T cells. Immunity 38, 414–423. https://doi.org/10.1016/j.immuni.2013.03.002 (2013).
Colamatteo, A. et al. Metabolism and autoimmune responses: The microRNA connection. Front. Immunol. 10, 1969. https://doi.org/10.3389/fimmu.2019.01969 (2019).
Kawakami, R. et al. Distinct Foxp3 enhancer elements coordinate development, maintenance, and function of regulatory T cells. Immunity 54, 947-961.e948. https://doi.org/10.1016/j.immuni.2021.04.005 (2021).
Mailer, R. K. W. Alternative splicing of FOXP3-virtue and vice. Front. Immunol. 9, 530. https://doi.org/10.3389/fimmu.2018.00530 (2018).
Melis, D. et al. Cutting edge: Increased autoimmunity risk in glycogen storage disease type 1b Is Associated with a reduced engagement of glycolysis in T cells and an impaired regulatory T cell function. J. Immunol. 198, 3803–3808. https://doi.org/10.4049/jimmunol.1601946 (2017).
Du, J., Wang, Q. & Yang, S. FOXP3 exon 2 controls T(reg) stability and autoimmunity. Sci. Immunol. 7, eabo5407. https://doi.org/10.1126/sciimmunol.abo5407 (2022).
Pesenacker, A. M., Cook, L. & Levings, M. K. The role of FOXP3 in autoimmunity. Curr. Opin. Immunol. 43, 16–23. https://doi.org/10.1016/j.coi.2016.07.004 (2016).
Wing, J. B., Tanaka, A. & Sakaguchi, S. Human FOXP3(+) regulatory T cell heterogeneity and function in autoimmunity and cancer. Immunity 50, 302–316. https://doi.org/10.1016/j.immuni.2019.01.020 (2019).
Kuhlmann, T. et al. Multiple sclerosis progression: Time for a new mechanism-driven framework. The Lancet Neurol. 22, 78–88. https://doi.org/10.1016/s1474-4422(22)00289-7 (2023).
Ogawa, C. et al. TGF-beta-mediated Foxp3 gene expression is cooperatively regulated by Stat5, Creb, and AP-1 through CNS2. J. Immunol. 192, 475–483. https://doi.org/10.4049/jimmunol.1301892 (2014).
Kim, H. P. & Leonard, W. J. CREB/ATF-dependent T cell receptor-induced FoxP3 gene expression: a role for DNA methylation. J. Exp. Med. 204, 1543–1551. https://doi.org/10.1084/jem.20070109 (2007).
Wang, X. et al. Cyclic AMP-Responsive element-binding protein (CREB) is critical in autoimmunity by promoting Th17 but inhibiting treg cell differentiation. EBioMedicine 25, 165–174. https://doi.org/10.1016/j.ebiom.2017.10.010 (2017).
Mosenden, R. & Tasken, K. Cyclic AMP-mediated immune regulation—overview of mechanisms of action in T cells. Cell. Signal. 23, 1009–1016. https://doi.org/10.1016/j.cellsig.2010.11.018 (2011).
Arumugham, V. B. & Baldari, C. T. cAMP: A multifaceted modulator of immune synapse assembly and T cell activation. J. Leukoc. Biol. 101, 1301–1316. https://doi.org/10.1189/jlb.2RU1116-474R (2017).
Murray, A. J. Pharmacological PKA inhibition: All may not be what it seems. Sci. Signal. 1, re4. https://doi.org/10.1126/scisignal.122re4 (2008).
Lochner, A. & Moolman, J. A. The many faces of H89: A review. Cardiovasc. Drug Rev. 24, 261–274. https://doi.org/10.1111/j.1527-3466.2006.00261.x (2006).
Wen, A. Y., Sakamoto, K. M. & Miller, L. S. The role of the transcription factor CREB in immune function. J. Immunol. 185, 6413–6419. https://doi.org/10.4049/jimmunol.1001829 (2010).
Haxhinasto, S., Mathis, D. & Benoist, C. The AKT-mTOR axis regulates de novo differentiation of CD4+Foxp3+ cells. J. Exp. Med. 205, 565–574. https://doi.org/10.1084/jem.20071477 (2008).
Turner, M. S., Kane, L. P. & Morel, P. A. Dominant role of antigen dose in CD4+Foxp3+ regulatory T cell induction and expansion. J. Immunol. 183, 4895–4903. https://doi.org/10.4049/jimmunol.0901459 (2009).
Huang, H., Long, L., Zhou, P., Chapman, N. M. & Chi, H. mTOR signaling at the crossroads of environmental signals and T-cell fate decisions. Immunol. Rev. 295, 15–38. https://doi.org/10.1111/imr.12845 (2020).
Pearce, E. L., Poffenberger, M. C., Chang, C. H. & Jones, R. G. Fueling immunity: Insights into metabolism and lymphocyte function. Science 342, 1242454. https://doi.org/10.1126/science.1242454 (2013).
Zheng, Y. et al. Role of conserved non-coding DNA elements in the Foxp3 gene in regulatory T-cell fate. Nature 463, 808–812. https://doi.org/10.1038/nature08750 (2010).
Oda, J. M., Hirata, B. K., Guembarovski, R. L. & Watanabe, M. A. Genetic polymorphism in FOXP3 gene: Imbalance in regulatory T-cell role and development of human diseases. J. Genet. 92, 163–171. https://doi.org/10.1007/s12041-013-0213-7 (2013).
Shi, H. & Chi, H. Metabolic control of treg cell stability, plasticity, and tissue-specific heterogeneity. Front. Immunol. 10, 2716. https://doi.org/10.3389/fimmu.2019.02716 (2019).
Galgani, M., Bruzzaniti, S. & Matarese, G. Immunometabolism and autoimmunity. Curr. Opin. Immunol. 67, 10–17. https://doi.org/10.1016/j.coi.2020.07.002 (2020).
Jungmann, R. A. & Kiryukhina, O. Cyclic AMP and AKAP-mediated targeting of protein kinase A regulates lactate dehydrogenase subunit A mRNA stability. J. Biol. Chem. 280, 25170–25177. https://doi.org/10.1074/jbc.M502514200 (2005).
Tian, D., Huang, D., Brown, R. C. & Jungmann, R. A. Protein kinase A stimulates binding of multiple proteins to a U-rich domain in the 3’-untranslated region of lactate dehydrogenase A mRNA that is required for the regulation of mRNA stability. J. Biol. Chem. 273, 28454–28460. https://doi.org/10.1074/jbc.273.43.28454 (1998).
Zhang, T. et al. Adenosine influences Foxp3 expression of T regs via the A2ar/Creb pathway in a mouse model of sepsis. Shock 61, 924–933. https://doi.org/10.1097/SHK.0000000000002281 (2024).
Clark, B. J. & Hudson, E. A. StAR protein stability in Y1 and Kin-8 mouse adrenocortical cells. Biology (Basel) 4, 200–215. https://doi.org/10.3390/biology4010200 (2015).
Liu, Q., Nguyen, E., Doskeland, S. & Segal-Bendirdjian, E. cAMP-dependent protein kinase A (PKA)-mediated c-myc degradation is dependent on the relative proportion of PKA-I and PKA-II isozymes. Mol. Pharmacol. 88, 469–476. https://doi.org/10.1124/mol.115.097915 (2015).
Kempkes, R. W. M., Joosten, I., Koenen, H. & He, X. Metabolic pathways involved in regulatory T cell functionality. Front. Immunol. 10, 2839. https://doi.org/10.3389/fimmu.2019.02839 (2019).
La Rocca, C. et al. Immunometabolic profiling of T cells from patients with relapsing-remitting multiple sclerosis reveals an impairment in glycolysis and mitochondrial respiration. Metabolism 77, 39–46. https://doi.org/10.1016/j.metabol.2017.08.011 (2017).
Galgani, M. et al. Regulatory T cells, inflammation, and endoplasmic reticulum stress in women with defective endometrial receptivity. Fertil. Steril. 103, 1579–1586. https://doi.org/10.1016/j.fertnstert.2015.03.014 (2015).
De Rosa, V. et al. A key role of leptin in the control of regulatory T cell proliferation. Immunity 26, 241–255. https://doi.org/10.1016/j.immuni.2007.01.011 (2007).
Acknowledgements
We thank M. Montagna for technical support. This paper was supported by Grants from Fondazione Italiana Sclerosi Multipla (FISM) no. 2022-PR-Single/013, Italian Ministry of Health, Next Generation EU, M6/C2_CALL 2022 cod. PNRR-MAD-2022-12376126, the National Recovery and Resilience Plan (NRRP), Mission 4, Component 2 (M4C2), Investment 1.1, Call for tender No. 1409 published on 14.9.2022 by the Italian Ministry of University and Research (MUR), funded by the European Union – NextGenerationEU– Project Title: Defining the role of the mTORC1-TFEB axis in the control of immune tolerance in Multiple Sclerosis (P2022T4PKT)-CUP B53D23024820001- Grant Assignment Decree No. 1365 published on 01.09.2023 to C.P.; Fondazione Italiana Sclerosi Multipla (FISM) n°2018/R/4, MAI Award Research Grant Prot. 2254 to V.D.R.; Fondazione Italiana Sclerosi Multipla (FISM) n° 2018/S/5, National Recovery and Resilience Plan (NRRP), Mission 4, Component 2 (M4C2), Investment 1.1, Call for tender No. 104 published on 02/02/2022 by the Italian Ministry of University and Research (MUR), funded by the European Union –NextGenerationEU– Project Title: Role of Overweight and EBV Infection in the Pathogenesis of Multiple Sclerosis: Metabolic Overwork as Key Mechanism Leading to Loss of Immune-Tolerance (Grant n. 2022LNHZAP) – CUP E53D23013240006 - Grant Assignment Decree No. 0001111 adopted on 20/07/2023; Italian Ministry of Health, Next Generation EU, M6/C2_CALL 2022 cod. PNRR-MAD-2022-12375634 and the MUR/PNRR Extended Partnership (MNESYS no. PE00000006) to G.M.; “Inflammation in human early life: targeting impacts on life-course health” (INITIALISE) consortium funded by the Horizon Europe Program of the European Union under Grant Agreement 101094099, European Foundation for the Study of Diabetes (EFSD)/Novo Nordisk Programme for Diabetes Research in Europe 2020, Fondazione Italiana Sclerosi Multipla (number 2020/R/13), National Recovery and Resilience Plan (NRRP), Mission 4, Component 2, Investment 1.1, Call for tender No. 1409 published on 14.9.2022 by the Italian Ministry of University and Research (MUR), funded by the European Union – NextGenerationEU– Project Title: Exploring PROteoMIc profile and homeostasis of regulatory TR3-56 cells in Type 1 Diabetes at onset – CUP: E53D23015370001 - Grant Assignment Decree No. 1409 adopted on 14-09-2022 by the Italian Ministry of Ministry of University and Research (MUR) and ECP Immunomodulation Award (Therakos, UK) to M.G; F.C. is funded by Ministry of Health, Bando Ricerca Finalizzata 2021 (Grant n. GR-2021-12373337) and by National Recovery and Resilience Plan (NRRP), Mission 4, Component 2 (M4C2), Investment 1.1, Call for tender No. 104 published on 2.2.2022 by the Italian Ministry of University and Research (MUR), funded by the European Union – NextGenerationEU– Project Title: Role of metabolic pressure and viral infections in the loss of immune tolerance and tissue damage in celiac disease (Grant n. 2022YMJXYT) – CUP B53D23003710006 - Grant Assignment Decree No. 972 adopted on 30/06/23 and by NRRP, M4C2, Investment 1.1, Call for tender No. 1409 published on 14.9.2022 by the Italian Ministry of University and Research (MUR), funded by the European Union – NextGenerationEU– Project Title: The Longevity Associated Variant- (LAV) of BPIFB4 protein as a potential therapeutic tool in senescence program associated to multiple sclerosis (P2022CMK43)-CUP B53D23031570001- Grant Assignment Decree No. 1369 published on 01.09.2023; C.L.R. is funded by National Recovery and Resilience Plan (NRRP), M4C2, Investment 1.1, Call for tender No. 104 published on 02.02.2022 by the Italian Ministry of University and Research (MUR), funded by the European Union – NextGenerationEU– Project Title: Regulatory T cell-mediated immunological nursing for direct cardiac reprogramming (IMMUNOCARD) (Grant n. 20225KH7BZ) – CUP B53D23012500006 - Grant Assignment Decree No. 971 adopted on 30/06/23. Schematic model showed in the Fig. S1 was prepared using the Motifolio Scientific Illustration Toolkit (Motifolio Inc).
Author information
Authors and Affiliations
Contributions
All authors have made substantial contributions to this work and have approved the final manuscript. M.T.L. and S.C. performed most of the experiments and data analyses; S.B., C.L.R, C.F., F.C. and S.D.S., performed experiments and data analyses; M.T.L., S.C., S.B., C.P., A.P. and V.D.R. analysed the data and interpreted the results; M.M., B.Z., R.L., V.B.M. and G.T.M. provided human samples; M.T.L., S.C., M.G. and G.M. designed the study and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lepore, M.T., Bruzzaniti, S., La Rocca, C. et al. Deciphering the role of protein kinase A in the control of FoxP3 expression in regulatory T cells in health and autoimmunity. Sci Rep 14, 17571 (2024). https://doi.org/10.1038/s41598-024-68098-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-68098-z