Abstract
Bone marrow-derived mesenchymal stem cells (BMSCs) are capable of developing into osteoblastic cell lines in vitro and regenerating bone tissue in vivo, and they are considered to be a reliable source for bone regenerative medicine. In recent years, studies have shown that the immune microenvironment is important for osteogenesis, in which macrophages are an important component of innate immunity and coordinate with stem cells. Metformin (Met), a hypoglycemic drug that exerts a powerful effect on metabolic signaling, has been shown to modulate inflammatory responses and osteogenic activity. However, whether metformin modulates macrophage polarization and subsequently affects osteogenesis remains to be elucidated. In the present study, we investigated the potential immunomodulatory capacity of metformin on macrophage inflammatory responses and phenotypic switching, and the subsequent effects on osteogenic differentiation of BMSCs. Flow cytometry and qPCR were used to study the effects of metformin on macrophage phenotypic regulation. qPCR, ALP, ARS and calcium content measurement and ALP activity assay were used to determine the effects of macrophage-secreted activators on the osteogenic differentiation of BMSCs. Our study demonstrates that metformin can improve the immune microenvironment by modulating macrophage polarization towards an anti-inflammatory phenotype, promoting an increase in a range of anti-inflammatory factors and inhibiting pro-inflammatory factors. This was characterized by increased expression of IL-10 and CD206, Arg-1 and decreased expression of IL-1β, TNF-α, IL-6 and iNOS. In addition, metformin-modulated macrophage-conditioned medium promoted osteogenic differentiation of BMSCs, increased the expression levels of genes (ALP, Runx-2, OCN, and Col-1), enhanced ALP activity, and significantly formed mineralized nodules. In conclusion, our new study elucidates that metformin can promote osteogenic differentiation of BMSCs by modulating macrophage phenotype and thereby.
Similar content being viewed by others
Introduction
Fractures are a widespread public health issue and frequent surgical damage. The number of new fracture patients worldwide is estimated to be over 178 million in 2019; this represents an increase of 33.4% from 19901. 1.9–4.9% of patients still have issues from fracture treatment procedures, such as delayed bone healing or bone nonunion, which lengthens the healing process and raises patient healthcare expenses2,3. As a result, the treatment focus in bone repair has shifted to ways to encourage bone healing.
Bone marrow-derived mesenchymal stem cells are capable of developing into an osteoblastic cell line in vitro and regenerating bone tissue in vivo, and they are considered a reliable source for bone regenerative medicine4,5. In recent years, it has been shown that the immune microenvironment is important for osteogenesis6,7, in which macrophages are an important component of innate immunity and coordinate with stem cells8,9. Macrophages are key cells in the inflammatory cascade response and play an important role in the bone healing process. Macrophages are highly plastic and can differentiate into two types, the pro-inflammatory M1 phenotype and the anti-inflammatory, pro-tissue regeneration M2 phenotype10. Different phenotypes of macrophages play different roles in various stages of bone regeneration, with M1 macrophages enriched at early inflammatory sites, phagocytizing bacteria and apoptotic cell debris. In contrast, M2 macrophages play roles such as inhibiting inflammation and repairing tissues in the later stages of fracture, while promoting BMSCs-mediated bone formation and healing11. The ratio of the number of M1/M2 macrophage taxa changes over time during the development of inflammation, and the two phenotypes of macrophages can be interconverted, which makes them attractive targets for therapeutic intervention. Recent studies have shown that targeting and controlling macrophage phenotypes can be used to treat a number of diseases, including atherosclerosis, obesity, and bone disorders11,12,13. This milieu is then favourable to tissue repair and regeneration. Studies have determined that regulating the crosstalk between macrophages and stem cells is critical for bone healing14,15.
Metformin is a hypoglycemic agent that has a powerful effect on metabolism. Several studies have demonstrated the beneficial effects of metformin in bone disease and inflammatory disorders that can ameliorate inflammation and modulate bone homeostasis, including osteolysis, periodontitis, osteoarthritis, and osteoporosis16,17,18. In addition, previous studies have shown that metformin inhibits inflammation and avoids chronic inflammation at the site of injury by polarizing macrophages to an anti-inflammatory M2 phenotype through activation of the AMPK pathway19,20. When injury occurs, metformin modulates the surrounding microenvironment. Studies have shown that metformin reduces the transcription of TNF-α and IL-6 and decreases the expression of IL-1β, COX-2, and iNOS by inhibiting NF-κB and MAPKs. In addition, metformin increases the secretion of two anti-inflammatory cytokines, IL-4 and IL-10, in an inflammatory setting. It is believed that metformin induces these effects through activation of AMPK, inhibition of ROS formation, NF-kB activation, JNK and ERK activity21.
However, the mechanism of crosstalk between macrophages and bone marrow mesenchymal stem cells (BMSCs) and the role played by metformin remain to be further explored. Therefore, in the present study, we investigated the effects of metformin on macrophage polarization and inflammatory responses, and co-cultured BMSCs with different macrophage-conditioned media to explore the effects of macrophage polarization phenotype on the osteogenic differentiation ability of BMSCs. We aimed to explore whether metformin could be a bioactive stimulator of bone regeneration and enhance bone regeneration by modulating macrophage-BMSC intercellular communication to further promote osteogenic differentiation.
Results
Metformin has no pharmacotoxic effect on Raw264.7 cultured in vitro
In order to find a suitable concentration of metformin, the CCK-8 test was used to study the pharmacotoxic effects of different concentrations of metformin on Raw264.7 after 24 h of action. As shown in Fig. 1a, there was no statistically significant difference in Raw264.7 proliferation with 0.5 mM, 1 mM, and 2 mM metformin. Therefore, metformin was not cytotoxic to Raw264.7 and had no effect on Raw264.7 proliferation in our tested dose range. Finally, 0.5 mM and 2 mM concentrations of metformin were selected for subsequent experiments.
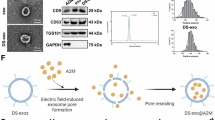
(a) Effect of different concentrations of metformin on the proliferative activity of Raw264.7. (b) Flow cytometry analysis of macrophage polarization (M2 type for CD206). (c) Mean fluorescence intensity of CD206. (d) Relative mRNA expression levels of M2-related genes Arg-1, CD206, and IL-10. (e) Relative mRNA expression levels of M1-related genes iNOS, IL-6, IL-1β. Data are presented as mean ± SD from triplicate experiments. *P < 0.05, **P < 0.01, ***P < 0.001, and****P < 0.0001.
Metformin modulates macrophage phenotype towards M2 polarization in both inflammatory and non-inflammatory settings
In view of the previous literature showing that metformin can modulate inflammatory responses, we investigated the macrophage phenotype modulation by metformin. First, flow cytometry results showed that 0.5 mM and 2 mM metformin induced high expression of CD206 (Fig. 1b), and there was a statistically significant difference in MFI in the experimental group compared with the control group (Fig. 1c). As shown in Fig. 1d, the expression of M2 macrophage-associated factors CD206, Arg-1, and IL-10 was detected to increase significantly with the increase of metformin concentration by qRT-PCR analysis. In addition, the expression levels of M1-related genes iNOS, IL-6, and IL-1β gradually decreased with the increase of drug concentration compared with the control group (Fig. 1e). All these results confirmed the potential of metformin to upregulate M2 polarization in macrophages.
To further validate the regulatory effect of metformin on macrophage phenotype under inflammatory state, we stimulated macrophages using 100 ng/ml LPS for 24 h with or without metformin for 24 h. The results of flow cytometry showed a significant increase in the expression of CD86 compared to the control group after 24 h of LPS stimulation (Fig. 2a), and the mean fluorescence intensity was statistically different from that of the control group (Fig. 2b). This indicated that LPS-stimulated macrophages had elevated levels of inflammation and were polarized towards the M1 type. Subsequent to the addition of different concentrations of metformin in the inflammatory state, flow cytometry showed a significant increase in the expression level of CD206 (Fig. 2c), and the mean fluorescence intensity was statistically different from that of the control group (Fig. 2d). In addition, by qRT-PCR analysis, we found that the expression levels of inflammatory factors and M1-related factors TNF-α and IL-1β in the LPS-stimulated group were significantly higher compared with the control group, while the expression levels of the related pro-inflammatory factors were significantly decreased 24 h after the addition of 0.5 mM and 2 mM of metformin (Fig. 2f); In contrast, the expression of CD206 and Arg-1 was suppressed under LPS stimulation, whereas the expression of both increased significantly and statistically differently compared to the control group after the addition of metformin(Fig. 2e). All of these data suggest that even in the inflammatory state metformin can modulate macrophage phenotype towards M2 polarization. In addition, 2 mM Met promoted the expression of Arg-1 in M0 and M1 type macrophages more than 0.5 mM Met with statistically significant difference, while in the expression of other genes, although there was no statistically significant difference between the two concentrations, 2 mM Met showed a stronger tendency to induce macrophage M2 polarization, so 2 mM Met was used in part of the subsequent osteogenesis experiments.
(a) Flow cytometry analysis of macrophage polarization after Lps stimulation (CD86 for M1 type). (b) Mean fluorescent intensities of CD86 by flow cytometry. (c) Flow cytometry analysis of macrophage polarization after metformin treatment in the inflammatory state (CD206 for M2 type). (d) Mean fluorescent intensities of CD206 by flow cytometry. (e) Relative mRNA expression levels of the M2-related genes Arg-1 and CD206 in the macrophage after metformin treatment in the inflammatory state. (f)Relative mRNA expression levels of the M1-related genes IL-1β and TNF-α in the macrophage after metformin treatment in the inflammatory state. *P < 0.05, **P < 0.01, ***P < 0.001, and****P < 0.0001.
Macrophage-conditioned medium promotes osteogenic differentiation of BMSCs after metformin action
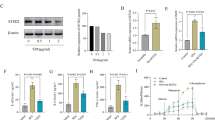
To further investigate the effects on osteogenic differentiation of BMSCs in different macrophage OCM,the experiment was divided into six groups :Control group: BMSCs cultured in normal osteogenic medium; Met group: BMSCs cultured in osteogenic medium containing 2 mM Met; M0 group: BMSCs cultured in untreated macrophage OCM; M0+ Met group: BMSCs cultured in macrophage OCM after 2 mM Met treatment for 24 h; LPS group: BMSCs cultured in macrophage OCM treated with LPS (100 ng/mL) for 24 h; and LPS + Met group: BMSCs cultured in macrophage OCM treated with Met and LPS. Figure 3a shows a schematic diagram of the cell processing method and Fig. 3b shows a timeline of the cell processing. CCK8 results showed significant proliferation of BMSCs cultured in the M0 + Met group after 7 days, while the LPS group exhibited suppressed cell proliferation vigor, and the rest of the macrophage-conditioned media had no significant effect on the proliferative vigor of BMSCs (Fig. 4a). We used qRT-PCR to detect the expression of osteogenesis-related genes (i.e., ALP, OCN, Col-1, Runx2) (Fig. 4b). On day 7, the Met group and M0-OCM showed a minor promotion of osteogenesis-related genes in BMSCs, whereas M0 + Met-OCM showed a statistically significant promotion of ALP, OCN, Col-1, and Runx2 expression in BMSCs compared to the control and Met groups as well as the M0 group. On the contrary, in LPS-OCM , the osteogenesis-related genes ALP, OCN, Col-1, and Runx2 were significantly reduced in BMSCs compared with the control group, and osteogenic gene expression was improved in LPS + Met-OCM treated with Met.
(a) Schematic diagram of different macrophage OCM treatments. (b) Timeline of different macrophage OCM treatments.
(a) The activity of BMSCs under different macrophage OCM was assessed after 7 days of culture. (b) Relative mRNA expression levels of osteogenic genes OCN, Runx2, ALPand COL-1 in BMSCs cultured with different macrophage OCMs.*P < 0.05, **P < 0.01, ***P < 0.001, and****P < 0.0001.
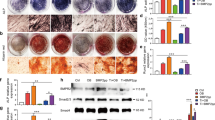
Two important markers, ALP activity and calcium deposition, were used to characterize the osteogenic differentiation capacity of BMSCs. As shown in Figs. 5 and 6, under non-inflammatory conditions, Met and RAW264.7 by itself exhibited a minor promoting effect on the osteogenic differentiation of BMSCs, whereas M0 + Met-OCM pretreated with Met for 24 h exerted a significant promoting effect on the osteogenic capacity of BMSCs, and the staining colors of ALP and calcium (alizarin red) became darker. In contrast, under inflammatory conditions, osteogenic differentiation capacity was significantly inhibited in LPS-OCM, with ALP and calcium (alizarin red) staining colors becoming lightest in comparison. In contrast, the osteogenic differentiation capacity of BMSCs improved in the inflammatory macrophage conditioned base after metformin induction. From the quantitative analyses of ALP and calcium content shown in Fig. 7, it is possible to distinguish more clearly the differences between the different groups, and the results are consistent with the staining. Based on these results, we concluded that macrophage-conditioned medium after metformin action promoted BMSCs osteogenic differentiation.
Representative images of alkaline phosphatase (ALP) staining of BMSCs cultured with the different macrophage OCM at day 7. Scar bar = 100 μm (×100).
Representative images of Alizarin Red staining (ARS) of BMSCs cultured with the different macrophage OCM at 21 days.Scar bar = 100 μm (×100).
(a) Quantitative analysis of ALP staining. (b) The relative quantitative analysis of ARS.*P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001.
Discussion
In this study, we investigated the effects of metformin on macrophage polarization and inflammatory responses and cultured BMSCs with different macrophage-conditioned media to explore the effects of exogenous substrates secreted by different macrophage phenotypes on the osteogenic differentiation capacity of BMSCs. The results indicated that metformin plays a regulatory role in converting M1 macrophages to M2 macrophages, which contributes to the transition from an inflammatory state to a tissue regenerative state. In addition, the extracellular matrix of macrophages after metformin stimulation improved the osteogenic differentiation capacity of BMSCs regardless of inflammatory or non-inflammatory conditions.
In recent years, studies have demonstrated the importance of the immune microenvironment for osteogenesis, and an increasing number of studies have focused on the association between macrophages and BMSCs and provided innovative strategies to enhance bone regeneration through targeted immunomodulation8,22. Functional dynamic transformation of macrophage phenotype plays a crucial role in inflammatory response and bone repair, and with this in mind, strategies to modulate macrophage activity to generate an ideal immune microenvironment and thereby promote osteogenic differentiation of BMSCs were the focus of this study.
Recent studies have reported that functional components in certain drugs may affect macrophage activity and convert the phenotype from M1 to M215,23. In the present study, metformin modulated macrophage phenotypic polarization toward the M2 phenotype. Met not only promoted the polarization of untreated Raw264.7 toward the M2, but also inhibited the polarization of LPS-stimulated macrophages toward the pro-inflammatory phenotype and increased the conversion to the anti-inflammatory phenotype. Thus, metformin has great potential to create favorable immune conditions for tissue regeneration. Previous studies have reached consistent conclusions regarding the immunomodulatory effects of metformin on macrophages. Wang et al. concluded that metformin inhibits LPS-induced macrophage inflammatory responses and modulates macrophage phenotypes to influence the immune environment thereby ameliorating allergic contact dermatitis in mice24. In addition, Jing et al. concluded that metformin ameliorated the low-grade inflammatory state in obese patients and regulated macrophage polarization to an anti-inflammatory M2 phenotype partly through AMPK activation25. Metformin also plays an immunomodulatory role in osteoarthritis. Li et al. further found that metformin was more effective in the treatment of arthritis in obese mice, mainly because it reduced synovial macrophage infiltration and regulated macrophage polarization18. Similarly Yan et al. found that metformin inhibited polyethylene particles-induced cranial osteolysis in mice by promoting macrophage polarization as an anti-inflammatory phenotype and improved the immune microenvironment for tissue regeneration17. Our study is consistent with the above studies that metformin has a promising immunomodulatory effect, creating an immune microenvironment that promotes tissue regeneration by modulating the inflammatory response and macrophage phenotypic polarization.
Regarding the direct promotion of stem cell osteogenesis by metformin, Lei et al. showed that low concentrations of metformin could promote the proliferation of BMSCs, with the best proliferative effect of metformin at 100 uM, and the proliferation of BMSCs was inhibited by too high a concentration of metformin26. Dong et al. also showed that, although low concentrations of metformin promoted the proliferation of BMSCs and the expression of the related osteogenic protein factors OPN, Runx-2 and Col-1, this promotion was substantially reduced at higher concentrations27. In this study, 2 mM metformin did not significantly affect the osteogenic effect of BMSCs, which may be due to the high concentration of metformin. However, the osteogenic effect of BMSCs in M0 + Met-OCM was significantly higher than that in all other groups, suggesting that the regulation of macrophages by metformin plays an important role in this.
The immune microenvironment generated by macrophages of different polarization types plays an important role in osteogenic differentiation. In the present study, metformin promoted the polarization of macrophages to M2 type, and the conditioned medium formed by such M2-polarized macrophages promoted the osteogenic differentiation ability and proliferative activity of BMSCs. Previous studies have shown that M2-type macrophages can secrete a range of anti-inflammatory factors and growth factors such as IL-10, TGF-β, IGF-1, and VEGF to promote osteogenic differentiation of BMSCs28,29, which is consistent with the results of the present study. Meanwhile, we found that Lps-stimulated macrophages had a significant inhibitory effect on the osteogenic differentiation ability of BMSCs and suppressed the cell proliferation viability of BMSCs, which is consistent with the findings of Gong et al.30. Although there are other studies suggesting that M1 enhances the osteogenic differentiation capacity of stem cells, these contrary findings may be due to the different induction of M1-type macrophages31,32. Our results showed that under inflammatory conditions, metformin was able to reduce the release of proinflammatory factors and reverse the switch in polarization of M1 macrophages under LPS conditions, thereby promoting osteogenic differentiation of BMSCs. This is consistent with the findings of Shen et al. who suggested that metformin could promote osteogenesis in human umbilical cord MSCs by promoting macrophage polarization and thus osteogenesis13. These results suggest that metformin has some immunomodulatory effects and that its stimulated macrophages can promote BMSCs osteogenesis under both non-inflammatory and inflammatory conditions.
In summary, metformin improves the immune microenvironment formed by macrophages by regulating the crosstalk between macrophages and BMSCs, and then promotes osteogenic differentiation of BMSCs, suggesting that metformin has great potential for bone regeneration. However, the specific mechanism behind this phenomenon needs to be further investigated and further validation in vivo is required.
Materials and methods
Cell culture and treatment
Mouse mononuclear RAW264.7 macrophages were provided by the Shanghai Cell Bank of the Chinese Academy of Sciences and cells were cultured in Dulbecco's modified Eagle's medium (DMEM) supplemented with 10% fetal bovine serum (FBS; Gibco, Waltham, MA, USA) and 1% penicillin–streptomycin in a humidified environment at 37 °C containing 5% CO2, and passaged every 1–2 days once without trypsin digestion. Mouse bone marrow mesenchymal stem cells (purchased from Zhongqiaoxinzhou Biotech, Shanghai, China) were cultured in DMEM/F12 medium (Gibco, Waltham, MA, USA)containing 10% FBS and 1% penicillin–streptomycin. The cell culture medium was changed every three days, and when the cells reached 80–90% confluence, the cells were passaged with 0.05% trypsin–EDTA (Gibco, CA, USA). BMSCs from generation 4 to 6 were used in this study.
To investigate the effect of metformin on macrophage polarization and inflammatory response in a non-inflammatory environment, RAW 264.7 cells were cultured in 6-well plates with Dulbecco's modified Eagle's medium containing 10% FBS at 37 °C overnight and treated with different doses (0.5, 2 mmol/L) of metformin (Sigma-Aldrich, Saint Louis, MO, USA) treatment for 24 h for further experiments. In addition, to investigate the regulatory role of metformin in the inflammatory milieu, macrophages were stimulated using 100 ng/mL E. coli lipopolysaccharide (Sigma-Aldrich, Saint Louis, MO, USA) for 24 h followed by the addition of different concentrations of metformin for 24 h.
To assess the effect of macrophages on BMSCs osteogenesis after treatment with different treatments, macrophage conditioned medium (MCM) was collected, centrifuged at 4◦ for a continuous period of 1 h, and filtered through a 0.22 μm pore filter (Millipore, Billerica, MA,USA). Then, MCM was mixed with osteogenic differentiation medium containing 10% FBS, 1% penicillin–streptomycin, 0.1 μM dexamethasone, 50 mM ascorbic acid, and 10 mM β-glycerophosphate in a 1:2 volume ratio as osteogenic conditioned medium (OCM) to induce osteogenic differentiation of BMSCs (Fig. 3) . BMSCs changed the OCM every other day during osteogenic induction.
CCK-8 assay
The cell viability of RAW264.7 at different concentrations of metformin and the viability of BMSCs at different OCM were detected using the Cell Counting Kit-8 (CCK-8, Dojindo, Japan). Briefly, cells were inoculated into 96-well plates at a concentration of 3 × 103 cells per well. Different concentrations of metformin were then added to the medium to culture Raw264.7 for 24 h, or BMSCs were cultured with different macrophage-conditioned medium for 7 days. The medium was then replaced with 100 μl of freshly prepared DMEM containing 10 μl of CCK-8 solution. The cells were incubated in the dark at 37 °C for 2 h. The absorbance at 450 nm was measured using a spectrophotometer (BioTek SynergyH1, USA). Experiments were performed in triplicate.
Flow cytometry
RAW264.7 cells were inoculated in 6-well plates (4 × 105 cells per well) and treated according to the section of Cell culture and treatment. Then, the plates were blocked with FcR receptor blocker (Miltenyi Biotec, Bergisch Gladbach, Germany) for 10 min. Subsequently, the plates were incubated in the dark with anti-mouse CD86 antibody (BioLegend, SanDiego, CA, USA) and anti-mouse CD206 antibody (BioLegend. SanDiego, CA, USA) at 4 °C for 30 min for cell surface staining. Cells were then washed and tested on a flow cytometer acquisition (BD FACSCalibur). Data were analyzed using FlowJo software (FlowJo, USA).
Real-time quantitative polymerase chain reaction (RT-qPCR)
RAW264.7 cells were inoculated in 6-well plates and treated according to the section of Cell culture and treatment; BMSCs were inoculated in 6-well plates (2 × 105 cells per well) overnight and incubated with different preparations of OCM for 5 days. Total RNA was extracted from the cells using the Total Cellular RNA Extraction Kit (Novizan, Nanjing, China) according to the manufacturer's instructions, and then the PrimeScriptTM II 1st Strand cDNA Synthesis Kit was used for real-time polymerase chain reaction) (Noviza, Nanjinng, China) for further reverse transcription to cDNA. qRT-PCR was performed using ChamQ Universal SYBR qPCR Master Mix (Novizan, Nanjing, China). The transcription levels of macrophage phenotype-related genes and osteogenesis-related genes were evaluated by qRT-PCR. The results were calculated by 2−∆∆CT method. qPCR primers are shown in Table 1.
Alkaline phosphatase (ALP) activity and staining
BMSCs were inoculated in 24-well plates (5 × 104 cells per well) overnight and incubated with different preparations of OCM. The medium was changed every other day. On day 7, cells were washed three times with PBS and lysed with lysis buffer without phosphatase inhibitors. Then, ALP activity of different samples was measured using an ALP assay kit (Beyotime, Shanghai, China) according to the manufacturer's instructions. ALP staining was also performed on day 7. Samples were fixed with 4% paraformaldehyde for 30 min, washed three times with PBS, and stained for 6 h using the BCIP/NBT Alkaline Phosphatase Staining Kit (Beyotime, Shanghai, China). Colorimetric changes were analyzed using a microscope and the stained cells were imaged.
Alizarin red staining
BMSCs were inoculated and treated as described in section 2.5. On day 14, fixed cells were stained with 2% alizarin red S staining solution (Cyagen, Suzhou, China) for 5 min, and then the excess dye was washed with distilled water. After observation and photography, mineralized nodules were dissolved with 10% (w/v) cetylpyridinium chloride to detect the absorbance at 520 nm.
Statistical analysis
All experiments were performed three times. All data are shown as mean ± standard deviation (SD). Statistical analysis was performed using GraphPad Prism 6.0 software. Differences between groups were analyzed using one-way analysis of variance (ANOVA) and Tukey post hoc tests. Differences were considered statistically significant when P < 0.05.
Data availability
The data used to support the findings of this study are available from the corresponding author upon request.
References
Collaborators, G. B. D. F. Global, regional, and national burden of bone fractures in 204 countries and territories, 1990–2019: A systematic analysis from the Global Burden of Disease Study 2019. Lancet Healthy Longev. 2, e580–e592. https://doi.org/10.1016/S2666-7568(21)00172-0 (2021).
Mills, L. A., Aitken, S. A. & Simpson, A. The risk of non-union per fracture: Current myths and revised figures from a population of over 4 million adults. Acta Orthop. 88, 434–439. https://doi.org/10.1080/17453674.2017.1321351 (2017).
Zura, R. et al. Epidemiology of fracture nonunion in 18 human bones. JAMA Surg. 151, e162775. https://doi.org/10.1001/jamasurg.2016.2775 (2016).
Zhang, J. & Chen, J. Bone tissue regeneration: Application of mesenchymal stem cells and cellular and molecular mechanisms. Curr. Stem Cell Res. Ther. 12, 357–364. https://doi.org/10.2174/1574888X11666160921121555 (2017).
Arthur, A. & Gronthos, S. Clinical application of bone marrow mesenchymal stem/stromal cells to repair skeletal tissue. Int. J. Mol. Sci. https://doi.org/10.3390/ijms21249759 (2020).
Walsh, M. C. et al. Osteoimmunology: Interplay between the immune system and bone metabolism. Annu. Rev. Immunol. 24, 33–63. https://doi.org/10.1146/annurev.immunol.24.021605.090646 (2006).
Tsukasaki, M. & Takayanagi, H. Osteoimmunology: Evolving concepts in bone–immune interactions in health and disease. Nat. Rev. Immunol. 19, 626–642. https://doi.org/10.1038/s41577-019-0178-8 (2019).
Pajarinen, J. et al. Mesenchymal stem cell-macrophage crosstalk and bone healing. Biomaterials 196, 80–89. https://doi.org/10.1016/j.biomaterials.2017.12.025 (2019).
Wang, Y. H. et al. The crosstalk between macrophages and bone marrow mesenchymal stem cells in bone healing. Stem Cell Res. Ther. 13, 511. https://doi.org/10.1186/s13287-022-03199-y (2022).
Liu, Y. C., Zou, X. B., Chai, Y. F. & Yao, Y. M. Macrophage polarization in inflammatory diseases. Int. J. Biol. Sci. 10, 520–529. https://doi.org/10.7150/ijbs.8879 (2014).
Goodman, S. B., Gibon, E., Gallo, J. & Takagi, M. Macrophage polarization and the osteoimmunology of periprosthetic osteolysis. Curr. Osteoporos. Rep. 20, 43–52. https://doi.org/10.1007/s11914-022-00720-3 (2022).
Eshghjoo, S., Kim, D. M., Jayaraman, A., Sun, Y. & Alaniz, R. C. Macrophage polarization in atherosclerosis. Genes https://doi.org/10.3390/genes13050756 (2022).
Shen, M. et al. Metformin facilitates osteoblastic differentiation and M2 macrophage polarization by PI3K/AKT/mTOR pathway in human umbilical cord mesenchymal stem cells. Stem Cells Int. 2022, 9498876. https://doi.org/10.1155/2022/9498876 (2022).
Qiao, D. et al. Bio-inspired glycosylated nano-hydroxyapatites enhance endogenous bone regeneration by modulating macrophage M2 polarization. Acta Biomater. 162, 135–148. https://doi.org/10.1016/j.actbio.2023.03.027 (2023).
Chen, S. et al. Curcumin modulates the crosstalk between macrophages and bone mesenchymal stem cells to ameliorate osteogenesis. Front. Cell Dev. Biol. 9, 634650. https://doi.org/10.3389/fcell.2021.634650 (2021).
Nicolini, A. C., Grisa, T. A., Muniz, F., Rosing, C. K. & Cavagni, J. Effect of adjuvant use of metformin on periodontal treatment: A systematic review and meta-analysis. Clin. Oral. Investig. 23, 2659–2666. https://doi.org/10.1007/s00784-018-2666-9 (2019).
Yan, Z. et al. Metformin suppresses UHMWPE particle-induced osteolysis in the mouse calvaria by promoting polarization of macrophages to an anti-inflammatory phenotype. Mol. Med. 24, 20. https://doi.org/10.1186/s10020-018-0013-x (2018).
Li, D. et al. Metformin attenuates osteoarthritis by targeting chondrocytes, synovial macrophages and adipocytes. Rheumatology (Oxford) 62, 1652–1661. https://doi.org/10.1093/rheumatology/keac467 (2023).
Qing, L. et al. Metformin induces the M2 macrophage polarization to accelerate the wound healing via regulating AMPK/mTOR/NLRP3 inflammasome singling pathway. Am. J. Transl. Res. 11, 655–668 (2019).
Yuan, D. et al. Metformin regulates alveolar macrophage polarization to protect against acute lung injury in rats caused by paraquat poisoning. Front. Pharmacol. 13, 811372. https://doi.org/10.3389/fphar.2022.811372 (2022).
Schuiveling, M., Vazirpanah, N., Radstake, T., Zimmermann, M. & Broen, J. C. A. Metformin, a new era for an old drug in the treatment of immune mediated disease?. Curr. Drug Targets 19, 945–959. https://doi.org/10.2174/1389450118666170613081730 (2018).
Schlundt, C. et al. The multifaceted roles of macrophages in bone regeneration: A story of polarization, activation and time. Acta Biomater. 133, 46–57. https://doi.org/10.1016/j.actbio.2021.04.052 (2021).
Zhang, Y. et al. Catalpol modulating the crosstalking between mesenchymal stromal cells and macrophages via paracrine to enhance angiogenesis and osteogenesis. Exp. Cell Res. 418, 113269. https://doi.org/10.1016/j.yexcr.2022.113269 (2022).
Wang, M., Qu, S., Ma, J., Wang, X. & Yang, Y. Metformin suppresses LPS-induced inflammatory responses in macrophage and ameliorates allergic contact dermatitis in mice via autophagy. Biol. Pharm. Bull. 43, 129–137. https://doi.org/10.1248/bpb.b19-00689 (2020).
Jing, Y. et al. Metformin improves obesity-associated inflammation by altering macrophages polarization. Mol. Cell. Endocrinol. 461, 256–264. https://doi.org/10.1016/j.mce.2017.09.025 (2018).
Lei, T. et al. Metformin enhances the osteogenesis and angiogenesis of human umbilical cord mesenchymal stem cells for tissue regeneration engineering. Int. J. Biochem. Cell Biol. 141, 106086. https://doi.org/10.1016/j.biocel.2021.106086 (2021).
Dong, K., Zhou, W. J. & Liu, Z. H. Metformin enhances the osteogenic activity of rat bone marrow mesenchymal stem cells by inhibiting oxidative stress induced by diabetes mellitus: An in vitro and in vivo study. J. Periodontal Implant Sci. 53, 54–68. https://doi.org/10.5051/jpis.2106240312 (2023).
Shi, M. et al. Deproteinized bovine bone matrix induces osteoblast differentiation via macrophage polarization. J. Biomed. Mater. Res. A 106, 1236–1246. https://doi.org/10.1002/jbm.a.36321 (2018).
Lin, T. et al. Preconditioned or IL4-secreting mesenchymal stem cells enhanced osteogenesis at different stages. Tissue Eng. Part A 25, 1096–1103. https://doi.org/10.1089/ten.TEA.2018.0292 (2019).
Gong, L., Zhao, Y., Zhang, Y. & Ruan, Z. The macrophage polarization regulates MSC osteoblast differentiation in vitro. Ann. Clin. Lab. Sci. 46, 65–71 (2016).
Tu, B. et al. Macrophages derived from THP-1 promote the osteogenic differentiation of mesenchymal stem cells through the IL-23/IL-23R/beta-catenin pathway. Exp. Cell Res. 339, 81–89. https://doi.org/10.1016/j.yexcr.2015.10.015 (2015).
Valles, G. et al. Influence of inflammatory conditions provided by macrophages on osteogenic ability of mesenchymal stem cells. Stem Cell Res. Ther. 11, 57. https://doi.org/10.1186/s13287-020-1578-1 (2020).
Funding
This study was supported by the Project of the Nanjing Medical Science and Technique Development Foundation (YKK21183).
Author information
Authors and Affiliations
Contributions
S.F., C.Z. and R.D. designed the research. S.F., X.S. and C.S. performed the experiments. S.F, Y.X. and X.S. acquired and analyzed the data. S.F., C.Z. conceived the study and wrote the manuscript. The integrity of this work is guaranteed by R.D. All authors contributed to the article and approved the submitted version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Fan, S., Zhang, C., Sun, X. et al. Metformin enhances osteogenic differentiation of BMSC by modulating macrophage M2 polarization. Sci Rep 14, 20267 (2024). https://doi.org/10.1038/s41598-024-71318-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-71318-1
This article is cited by
-
The Impact of Large-Scale Expansion on the Functional Properties of Mesenchymal Stem Cells
Stem Cell Reviews and Reports (2026)
-
Multi-omics analysis of synovial tissue and fluid reveals differentially expressed proteins and metabolites in osteoarthritis
Journal of Translational Medicine (2025)
-
Ionizing radiation-mediated dendritic cell maturation exacerbates inflammatory response of bone marrow mesenchymal stem cells and impairs osteogenesis in radiation-induced jaw injury
Stem Cell Research & Therapy (2025)
-
Sirtuin 3 promotes osteogenic differentiation of bone marrow mesenchymal stem cells by regulating macrophage polarization under high glucose
Scientific Reports (2025)
-
Trolox, r-irisin and resveratrol cocktail to counteract osteoblast metabolism alterations in osteoarthritis and osteoporosis
Journal of Bone and Mineral Metabolism (2025)