Abstract
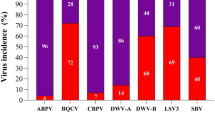
Honey bee viruses are serious pathogens that can cause poor colony health and productivity. We analyzed a multi-year longitudinal dataset of abundances of nine honey bee viruses (deformed wing virus A, deformed wing virus B, black queen cell virus, sacbrood virus, Lake Sinai virus, Kashmir bee virus, acute bee paralysis virus, chronic bee paralysis virus, and Israeli acute paralysis virus) in colonies located across Canada to describe broad trends in virus intensity and occurrence among regions and years. We also tested climatic variables (temperature, wind speed, and precipitation) as predictors in an effort to understand possible drivers underlying seasonal patterns in viral prevalence. Temperature was a significant positive predictor of the total number of viruses per sample, which was highest in British Columbia (mean = 5.0). Lake Sinai virus (LSV) was the most prevalent overall (at 89%) and had the highest infection intensity, at an average of 3.9 × 108 copies per bee. Acute bee paralysis virus was the least prevalent virus (at 4.7%) and had the lowest infection intensity (1.9 × 105 copies per bee). Surprisingly, including Varroa abundance as a covariate did not significantly improve model fit for any virus. All viruses, except Kashmir bee virus, varied by region, and one or more climatic variables were significant predictors for six of the nine viruses. Although climatic effects were often inconsistent among individual viruses, we show that climatic variables can be better predictors of virus intensity and occurrence than Varroa mite abundance, at least when infestation rates are low.
Similar content being viewed by others
Introduction
Since the discovery of the first viral disease of honey bees (sacbrood, documented by Gershom Franklin White in 19131), viruses have become widely recognized as serious honey bee pathogens2,3,4,5,6,7,8. Recent advances in high-throughput sequencing have now led to the discovery of at least 74 different viruses in honey bees9. While many of these appear to be asymptomatic, several—including Lake Sinai virus (LSV), black queen cell virus (BQCV), sacbrood virus (SBV), deformed-wing virus (DWV) A, B, C, and D,10 acute bee paralysis virus (ABPV), Israeli acute paralysis virus (IAPV), Kashmir bee virus (KBV), chronic bee paralysis virus (CBPV), and slow bee paralysis virus (SBPV), among others—do cause disease or are linked to colony weakening. Symptoms may include failure to pupate, deformed wings at emergence, trembling or paralysis, and shortened lifespan11,12, and some viruses are associated with colony dwindling and winter losses6,7,8,11,13,14,15,16. Despite being linked to these adverse outcomes, we still have large gaps in our knowledge of the factors governing honey bee virus epidemiology17.
In the absence of an additional vector, viruses spread through the fecal–oral route, sexually (from drones to queens during mating), vertically (from queens to her progeny), and, to a lesser extent, through direct physical contact mediated by worn setae18 (reviewed in Yañez et al.18). Through these transmission routes, viruses were historically not problematic (although significant outbreaks did occasionally occur).4 However, the threat of viruses intensified after the host-jumping and subsequent global spread of the Varroa destructor mite (hereon Varroa), which can act as a mechanical and biological vector (reviewed elsewhere17,18,19,20,21). Greater virulence is observed when viruses are inoculated via injection compared to orally, which is attributed to the fact that injection circumvents the bee’s normal immune defenses encountered in the digestive tract18,22,23. As a result, viruses transmitted via Varroa feeding wounds tend to lead to more intense infections than those transmitted by ingestion, even when the mite is not acting as a biological vector24,25. While the relative importance of viruses to other factors influencing colony health is difficult to ascertain,3 it is likely to be cumulatively substantial especially when Varroa abundance is high19.
Varroa is not the only parasite to have been implicated in virus pathogenicity. Based on observations made in the early 1980s, Vairimorpha apis (formerly Nosema apis26, but see Bartolomé et al.27) is thought to exacerbate BQCV infections by causing damage to the bee’s gut epithelium, thereby facilitating viral infection28,29. This mechanism has not been substantiated further, and it is not clear why this association would exist for BQCV and not other viruses, but the explanation is consistent with the observation that V. apis and BQCV tend to covary30,31.
Several surveys broadly tracking virus prevalence over time have documented annual and seasonal trends, with highly regional distributions. In a survey conducted in the U.S., Traynor et al.32 identified a worrying trend of annual doubling of CBPV prevalence, increasing from 0 to 25% between 2009 and 2014, as well as seasonal prevalence patterns of an LSV variant (LSV-2; highest in spring and early summer), and ABPV (highest in winter). In southwest Germany, D’Alvise et al.33 found that BQCV and LSV peaked in the spring and summer, but contrary to Traynor et al. 32, ABPV peaked in July. BQCV trends in China34, France31, and Lithuania35 agree with those in Germany33, and prevalence of SBV in Lithuania and Canada tends to be higher in spring or summer compared to fall35,36. BQCV and DWV were the most prevalent viruses measured in the US32, China34, and France31, but SBV was similarly prevalent in Lithuania and France (where it tied with BQCV for second place)31,35. BQCV, LSV, and SBV were the most prevalent viruses in the 2017 Canadian National Honey Bee Health Survey37, whereas BQCV, DWV, and IAPV were most prevalent across provinces in an independent Canadian dataset (in which LSV was not measured)38. In contrast to all other countries discussed here, ABPV was the second most prevalent virus in Austria30.
These seasonal, annual, and regional patterns reported above and in additional surveys39,40,41,42,43 suggest that climatic variables or other factors that covary with climate (such as foraging activity, colony population size, or relationships with pathogens or parasites which themselves are affected by climate) could conceivably influence viral occurrence or intensity. While high temperatures negatively affect viral replication in honey bees kept in the laboratory44,45, few studies have analyzed the effects of climate17. Piot et al.46 found that in Europe, precipitation and temperature significantly predicted prevalence of viruses in the AKI-complex (ABPV, KBV, and IAPV) as well as DWV and SBPV in bumble bees and solitary bees, but not in honey bees. Prado et al.47 found that warmer winters were associated with lower virus abundance, perhaps mediated by elevated immune effectors. To our knowledge, these are the only studies rigorously testing linkages between climatic data and honey bee viruses.
Here, we used a multi-year, longitudinal dataset of abundances of nine viruses in honey bee colonies located in five Canadian provinces to describe regional and temporal patterns in intensity and occurrence, as well as test the hypothesis that climatic variables (temperature, precipitation, and wind speed) are meaningful predictors. Because viruses can be transmitted by the fecal–oral route, we hypothesized that high wind speed and precipitation would limit defecation flights and therefore be associated with higher viral intensity within colonies, but lower overall occurrence due to reduced transmission opportunities between colonies. We also hypothesized that temperature would positively influence viral intensity and occurrence indirectly via positive effects on colony population size and foraging activity (since larger, crowded populations have more opportunities for transmission within and between colonies via drifting and robbing). We also used data for two other parasites (Varroa and Vairimorpha spp.) to test if their abundance was also positively linked to viral intensity or occurrence as has been previously documented19,29. Finally, we discuss potential mechanisms through which relationships with climatic variables could arise.
Results
Primary dataset
Overview of virus intensities, number of viruses per sample, and climatic variables
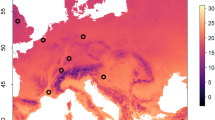
By conducting a new analysis of a dataset that was in part previously described48,49, we found clear regional trends in viral occurrence and intensities, with the most notable trends being (1) few viruses detected in Quebec (QC), (2) many viruses detected in British Columbia (BC), and (3) low virus intensities in samples from Ontario (ON) (Fig. 1a). We detected an overall average of 4.2 ± 1.2 (standard deviation) viruses per sample (of nine that were tested), with the largest mean number in BC (5.0), followed by southern Alberta (AB) (4.5), northern AB (4.3), QC near Lac St. Jean (QC (LSJ); 3.7), Manitoba (MB; 3.5), QC near Quebec City (QC (QC); 3.4), ON (3.3) and QC near Montreal (QC (Mon); 3.2) (Fig. 1b, c). We found significantly fewer viruses per sample in 2021 compared to 2020, but the magnitude of the difference is lower than the regional variation (coefficient = − 0.4). As reported previously48, in this dataset, overall mean temperature was 15.1 °C (2.9–22.7 °C), mean precipitation was 2.0 mm (0.1–19.6 mm), and mean wind speed was 9.5 km/h (3.9–17.5 km/h). Temperature predicted the number of viruses per sample better than sampling date (Fig. 1d), but region and the region-by-date interaction were the most influential terms in the model. Varroa abundance did not consistently correlate with viral abundance (Fig. 1e) and including mite abundance as a covariate did not improve model fit (p = 0.36, χ2 = 0.83, ΔAIC = 1.17). See Table 1 for complete statistical reporting.
Viruses in six Canadian beekeeping operations across two years of sampling. (A) Viral abundances (log10) in different pollination contexts and regions over time. Samples (n = 120 pooled samples representing 480 colonies) from colony cohorts near and far from focal crops were taken immediately before the pollination period (beginning), when the crop reached > 50% bloom (middle), and at the end of the pollination period (end), except for corn, for which middle samples coincided with sowing and end samples coincided with the end of the corn production season. (B,C) The total number of viruses per sample varied by year (χ2 = 6.6, df = 1, p = 0.010) and according to a region-by-date interaction (χ2 = 23.6, df = 7, p = 0.001) as well as (D) temperature (χ2 = 8.0, df = 1, p = 0.005). See Table 1 for complete statistical reporting. (E) Virus infection intensities did not consistently correlate with Varroa mite abundance using the full dataset (asterisks indicate significant correlations after Bonferroni corrections).
Lake Sinai virus (LSV)
Of the nine viruses analyzed, LSV had the highest prevalence and intensity, with a positivity rate of 89% overall and an average of 3.9 × 108 virus copies per bee (or 8.6 log10 LSV units). LSV intensity was higher in 2021 than 2020 (coefficient = 0.3 log10 LSV units) (Fig. 2a, b), but region and the region-by-date interaction were again the most influential factors. In some regions, LSV intensities increased over time and in others, intensities decreased over time. Interestingly, the temperature-by-precipitation interaction was highly influential, with LSV copies being positively linked to temperature when precipitation is low, but negatively linked to temperature when precipitation is high (Fig. 2c). A less influential three-way interaction between temperature, precipitation, and wind was also observed, whereby the effect of precipitation was exacerbated by high wind speed (Figure S1). See Table 2 for complete statistical reporting of LSV intensity analysis. Despite its overall high prevalence, LSV occurrence was still regionally distributed, with significantly fewer detections in ON and MB compared to BC (Fig. 2d). Including mite abundance as a covariate did not improve model fit (p = 0.54, χ2 = 0.38, ΔAIC = 1.62). See Table 3 for complete statistical reporting of LSV occurrence analysis.
LSV and BQCV are linked to region, date, and climatic variables. (A,B) LSV intensities differed between years (χ2 = 6.3, df = 1, p = 0.012), with mean log10 loads of 7.6 and 8.1 in 2020 and 2021, respectively, and according to a region-by-date interaction (χ2 = 38.4, df = 6, p < 0.001). (C) LSV intensities varied according to a significant temperature-by-precipitation interaction (χ2 = 12.5, df = 1, p < 0.001), and a less influential temperature-by-precipitation-by-wind interaction (Supplementary Figure S1; χ2 = 5.2, df = 1, p = 0.023). Data are shown in three equi-density precipitation bins to illustrate the nature of the interaction, but precipitation was analyzed as a continuous variable. See Table 2 for statistical parameters. (D) LSV occurrence (1 = detected, 0 = not detected) varied by region (χ2 = 29.1, df = 7, p < 0.001) and a temperature-by-precipitation interaction similar to (C) (Supplementary Figure S2; χ2 = 5.9, df = 1, p = 0.015; see Table 3 for statistical parameters), but not sample date (χ2 = 1.1, df = 1, p = 0.30). (E) BQCV intensities varied by a region-by-date interaction (χ2 = 37.5, df = 7, p < 0.001) and (F) temperature (χ2 = 5.8, df = 1, p = 0.016), as well as (G) a Vairimorpha spp.-by-precipitation interaction (χ2 = 10.3, df = 1, p = 0.001; again, with precipitation illustrated in three equi-density bins). See Table 4 for statistical parameters.
Black queen cell virus (BQCV)
BQCV was the second most prevalent virus (84% positivity rate). Broadly, we observed relatively low intensities in ON and year-dependent occurrence in BC (Fig. 1a). Formal analysis of BQCV intensities shows that temperature is again a significant positive predictor, in addition to region, date, and a region-by-date interaction (Fig. 2e, f). The region effect is driven by high intensities in southern AB and low intensities in QC (QC), whereas the region-by-date effect is driven by intensities increasing dramatically over time in QC (LSJ) (coefficient = 4.3 log10 BQCV units). We found that Vairimorpha spp. intensity significantly interacted with precipitation, with higher BQCV intensity associated with higher Vairimorpha spp. intensity, but not when precipitation was low (Fig. 2g). Including V. apis occurrence as a cofactor did not significantly improve model fit, nor did mite abundance. See Table 4 for statistical reporting.
Sacbrood virus (SBV)
SBV was the third most prevalent virus, with a 73.6% positivity rate, and was the only virus to be significantly influenced by proximity of colonies to focal crops, in this case in a region-by-date-by-proximity interaction (Fig. 3a). The interaction is driven by SBV intensities changing differently over time in near vs. far sites within southern AB and QC (LSJ): intensities increased faster at sites near lowbush blueberries (in QC (LSJ)) compared to sites far from lowbush blueberries, but intensities increase faster at sites far from seed canola and commodity canola in southern AB. SBV intensities were also linked to temperature and wind speed as an interactive effect, with high winds exacerbating the positive effect of temperature (Fig. 3b). SBV occurrence was highest in BC and southern AB, where nearly 100% of samples tested positive for the virus by midsummer, and occurrence was lower in 2021 compared to 2020 (Fig. 3c, d). Occurrence generally increased over time in most locations, except in MB, where it decreased over time, and in QC (QC), where it did not change. Including Varroa abundance did not significantly improve model fit for intensity or occurrence analysis for SBV. See Tables 5, 6 for statistical reporting.
Sacbrood virus intensities are linked to regional and climatic factors and occurrence probabilities follow region-dependent seasonal patterns. (A) SBV was the only pathogen significantly linked to site type (in this case, as part of a site type-by-region-by-date interaction; χ2 = 17.1, df = 4, p = 0.002). SBV intensities were more likely to increase over time in AB (N) and QC (LSJ) near sites compared to far, but the opposite was true in MB. See Table 5 for statistical parameters. (B) SBV intensities varied according to an interaction between wind and temperature (χ2 = 4.5, df = 1, p = 0.035), with the positive effect of temperature exacerbated at higher average wind speeds (wind is illustrated in three equi-density bins, but was analyzed as a continuous variable). (C,D) SBV occurrence probability tended to increase during the beekeeping season (χ2 = 9.0, df = 1, p = 0.003), except in MB (where occurrence probability decreased over time) and AB (S) (where occurrence probability was extremely high throughout the sampling periods). See Table 6 for statistical parameters.
Deformed wing viruses (DWV-A & DWV-B)
DWV-B was more prevalent than DWV-A, occurring in 65.3% and 45.3% of samples, respectively. Region and date were significant predictors of DWV-A occurrence, with the probability of occurrence being highest in BC and generally increasing over time (Fig. 4a). Region, date, and year were significant predictors of DWV-B occurrence (Fig. 4b, c). Occurrence was higher in 2021 than 2020, was consistently high in BC, and, similar to DWV-A, also generally increased over time (except in QC (QC)). The increase in prevalence over time was fastest in QC (LSJ), where probability of occurrence changed from zero in mid-May to 0.9 in early June. No climate variables were significant predictors of DWV-A or B occurrence. Surprisingly, mite abundance did not significantly improve model fit for either strain. See Tables 7, 8 for statistical reporting.
DWV-A and DWV-B occurrence varied by region and date. (A) DWV-A occurrence increased significantly over time (χ2 = 4.7, df = 1, p = 0.030) at similar rates between regions. 2020 and 2021 data are shown together, as there was no significant effect of year (χ2 = 3.6, df = 1, p = 0.058). See Table 7 for statistical parameters. (B,C) DWV-B occurrence was higher in 2021 than 2020 (χ2 = 29.4, df = 1, p < 0.001) and increased over time (χ2 = 5.5, df = 1, p = 0.019) in a pattern driven by Quebec sites (the only regions where DWV-B occurrence was initially low). See Table 8 for statistical parameters.
Paralysis viruses (ABPV, KBV, IAPV, and CBPV)
The paralysis viruses in the AKI-complex (ABPV, KBV, and IAPV) did not appear to covary, despite being closely related. ABPV was the least prevalent virus, occurring in just 4.7% of samples. Date, temperature, and a precipitation-by-wind speed interaction were significant predictors (Fig. 5a, b), but these associations are likely not reliable due to very few positive detections. See Table 9 for statistical reporting for ABPV. KBV was not significantly linked to any predictor (Fig. 5c). IAPV was the most prevalent paralysis virus, present in 36.7% of samples, and region, wind speed, and a region-by-year interaction (Fig. 5d–f). Occurrence was highest in BC (especially in 2021, when occurrence probability was 0.9–1.0 for all sample dates), and the region-by-year interaction was driven by differences between years in southern AB (where occurrence decreased dramatically with date in 2020 but not 2021). Occurrence did not change substantially with date in other regions. Curiously, IAPV occurrence appeared to be positively influenced by windspeed. See Table 10 for statistical reporting for IAPV.
Occurrence probabilities of viruses in the AKI-complex. (A) ABPV occurrence varied by date (χ2 = 3.9, df = 1, p = 0.047), temperature (Supplementary Figure S3; χ2 = 4.3, df = 1, p = 0.038), and (B) a precipitation-by-wind interaction (χ2 = 6.5, df = 1, p = 0.011), but not region (χ2 = 6.2, df = 7, p = 0.52). Precipitation is illustrated in three equi-density bins but was analyzed as a continuous variable. See Table 9 for statistical parameters. (C) KBV occurrence did not vary significantly by any factor. (D,E) IAPV occurrence varied according to a year-by-region interaction (χ2 = 15.2, df = 3, p = 0.002), and (F) was positively influenced by wind (χ2 = 4.4, df = 1, p = 0.036). See Table 10 for statistical parameters.
CBPV was present in 13.6% of samples and is the only virus to occur least often in BC and AB. Probability of occurrence was highest in ON and QC (LSJ) (Fig. 6a). Region, date, and temperature are significant predictors, but unlike the other viruses, temperature is a negative predictor of occurrence (Fig. 6b). Including mite abundance as a predictor did not improve model fit of viruses in the AKI-complex or CBPV. See Table 11 for statistical reporting for CBPV.
CBPV occurrence varies by region, date, and temperature. (A) CBPV occurrence probability tended to decrease over time (χ2 = 4.7, df = 1, p = 0.030). (B) Temperature predicts CBPV occurrence more strongly than date, with probabilities decreasing as temperatures increase (χ2 = 8.8, df = 1, p = 0.003). See Table 11 for statistical reporting.
Validation data
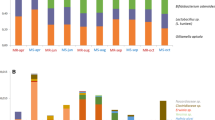
To determine if the climatic and temporal trends we observed were consistent in an independent dataset, we analyzed data collected by the Alberta Technology Transfer Program as part of the Alberta colony health monitoring project, for which qPCR data was obtained for BQCV, SBV, DWV-A, DWV-B, and CBPV. We found that BQCV and SBV varied according to climatic variables, but not in the same way as the primary dataset. BQCV intensities were significantly predicted by a date-by-year interaction (χ2 = 53.4, df = 4, p < 0.001), whereby intensities tended to peak mid-summer but to varying degrees in different years (Fig. 7a and Table S1). Whereas we previously identified a main effect of temperature in the primary dataset, here we found that BQCV intensities were significantly predicted by a temperature-by-wind interaction (χ2 = 9.0, df = 1, p = 0.0027), where windy conditions tended to enhance the positive relationship with temperature, but not in all years (Figure S4a). BQCV intensities also were positively linked to precipitation (χ2 = 29, df = 1, p < 0.001), which we had not previously observed (Figure S4b). SBV occurrence did not generally increase over time, as we had previously found; instead, occurrence remained steady throughout 2021, decreased over time in 2022, and slightly increased over time in 2023 (Fig. 7b and Table S2). SBV intensities were linked to precipitation (χ2 = 4.1, df = 1, p = 0.042), where a negative relationship was observed except in 2022, but not a temperature-by-wind interaction (χ2 = 0.25, df = 1, p = 0.61) (Fig. 7c). DWV-A and DWV-B occurrence tended to increase over time (Fig. 7d, e), similar to the primary dataset. CBPV occurrence was overall very low in Alberta (Fig. 7f), which is consistent with the primary dataset.
Analysis of validation data. The validation dataset consisted of qPCR data for BQCV, SBV, DWV-A, DWV-B, and CBPV derived from samples taken over time throughout Alberta in 2021, 2022, and 2023. See Tables S1-S5 for complete statistical summaries. (A) Unlike in the primary dataset, BQCV intensities varied according to a significant interaction between date (as a quadratic term) and year (χ2 = 53.4, df = 4, p < 0.001) and a significant interaction between temperature and wind (χ2 = 9.0, df = 1, p = 0.0027) (Supplementary Figure S4). (B) SBV occurrence varied according to a significant interaction between date and year (χ2 = 8.2, df = 1, p = 0.015), but no climatic variables. (C) SBV intensities (censored if log10(intensity) < 4.5 due to bimodality of the distribution) were significantly predicted by precipitation (χ2 = 4.1, df = 1, p = 0.042). Unlike the primary dataset, SBV intensity was not predicted by a temperature-by-wind interaction (χ2 = 0.25, df = 1, p = 0.61). (D) DWV-A and (E) DWV-B occurrence varied according to a significant interaction between date and year (χ2 = 11.4, df = 2, p = 0.0034 for DWV-A and χ2 = 10.3, df = 2, p = 0.0058 for DWV-B). (F) CBPV occurrence was low overall and insufficient for modelling.
Discussion
We used longitudinal datasets of viral abundances in Canadian honey bee colonies to describe temporal, regional, and climatic trends in virus intensity and occurrence. We hypothesized that, controlling for temporal and regional differences, we would find generally positive associations between temperature, wind speed, precipitation and viral intensity or occurrence, as warmer temperatures promote bee population growth and more crowded conditions, while inclement flight weather prevents defecation flights and may lead to more fecal–oral transmission within the hive. We also hypothesized that Varroa abundance would be a positive predictor of virus intensity and occurrence, particularly for DWV-B, for which it is a biological vector50,51,52. Finally, we hypothesized that Vairimorpha spp. intensity or V. apis occurrence would predict BQCV intensity, as suggested by Bailey29. Overall, we did find that climatic variables were significant predictors of viral intensity or occurrence for six of the nine viruses we tested, but the nature of these relationships were not always as expected nor consistent with our validation analysis. We also identified a positive association between BQCV and Vairimorpha (Fig. 2g), although the magnitude of influence is small (coefficient = 0.1).
Varroa abundance was not a significant predictor for any of the viruses tested here, despite strong existing evidence that Varroa acts as vector for viruses in a way that increases virulence and even changes infection dynamics18,19,20,50,51,53,54,55,56,57,58. Conversely, we found that although Varroa abundance did correlate with DWV-B abundance in 2021, it did not correlate in 2020 (Fig. 1E) and it was not a statistically significant covariate in any of our mixed models (Tables 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11). Our data do not refute the well-established role of Varroa as a viral vector (which has been supported by other surveys of honey bee colonies59,60,61); rather, we suggest that the abundance observed in our data (averaging 0.30 and 0.32 mites/100 bees in 2020 and 2021, respectively) was sufficiently low to not be influential. This provided an excellent opportunity for us to investigate climate variables as influential factors when the impact of Varroa is not the dominant factor.
In the primary dataset, while temperature was a positive predictor of the total number of viruses per sample (Fig. 1d), BQCV intensity (Fig. 2f), and SBV intensity (Fig. 3b), temperature negatively predicted CBPV occurrence (Fig. 6b), which was opposite to our prediction. The proposed mechanism underlying our hypothesis was that in the spring and summer (when these data were collected), rapid colony population growth during periods of warmer temperatures with abundant forage should result in denser colonies and more transmission, both through physical contact within the colony and contact at shared forage sources62,63. Because Bailey et al.64 described CBPV transmission occurring in crowded conditions, perhaps mediated by physical contact with broken setae, we especially expected a positive relationship with temperature to manifest for CBPV. We are unsure why this was not the case. Like many viruses, CBPV can also spread via fecal–oral transmission, and if this is the dominant infection route, it is conceivable that a relationship with temperature might be moderated by wind or precipitation, whereby a relationship with temperature is less apparent when wind and precipitation is low (favoring defecation flights) but more apparent when wind or precipitation are high (limiting defecation flights and promoting even more crowded conditions). However, we found no interactive effects between temperature, wind speed, or precipitation to support this scenario. There is likely some other influential biotic or abiotic variable explaining CBPV occurrence that we have not considered, such as relationships with the gut microbiome65 or transmission via robbing or drifting behaviors66,67. We were also unable to re-evaluate the factors affecting CBPV occurrence in the validation dataset, as this virus was too rare in Alberta to yield sufficient data (Fig. 7f).
We expected high precipitation and greater wind speeds to promote virus accumulation in the bees’ digestive tract, leading to more intense infections within bees confined to their hive and more opportunities for intra-colony transmission, either as fixed effects or as interactors. LSV14, BQCV53, ABPV68, CBPV12, DWV53, KBV69, and IAPV70 are all known to accumulate in the gut, so such an interaction is a reasonable prediction for all of them. In our primary analysis, precipitation or wind were significant predictors for SBV (precipitation interacted with temperature), IAPV (wind was a fixed effect), LSV (wind and precipitation interacted with temperature), and ABPV (precipitation interacted with temperature), but again the direction of change was not always as anticipated. High winds exacerbated positive effects of temperature for SBV (Fig. 3b), which is consistent with our proposed underlying mechanism, but SBV is one of the few viruses which is not known to be transmitted by the fecal–oral route among adults18 (though this is, admittedly, in part due to a lack of updated studies investigating SBV transmission routes). Wind positively predicted IAPV occurrence (Fig. 5f), which is consistent with our prediction, but there was no corresponding positive influence of temperature. For LSV, the interaction between precipitation and temperature had the opposite trajectory of our prediction, with high precipitation flipping the association with temperature from positive to negative (Fig. 2c). The link between precipitation and ABPV occurrence was also not as expected, with an overall negative effect of increasing precipitation (Fig. 5b). In our validation analysis, temperature and wind interacted to predict BQCV intensities, where high winds tended to intensify the positive effect of temperature, but not in all years, and precipitation also positively predicted BQCV intensities, which is the general scenario we predicted. However, our validation analysis also showed that for SBV, while a relationship with precipitation was identified, it tended to be a negative predictor (but again, not in all years).
These results are highly inconsistent and cannot be explained solely by inclement weather limiting dispersal opportunities alone; furthermore, they contrast with the largely intuitive results found when drawing similar associations between climatic variables and etiological agents with more well-defined epidemiology (Varroa, Vairimorpha spp., and M. plutonius)71. We refrain from speculating on possible explanations for these inconsistences, as there are many; rather, we will focus here on what these data collectively do allow us to conclude.
The association between Vairimorpha spp. spore loads and BQCV is intriguing. Bailey previously proposed that V. apis facilitates BQCV infection29, but the lack of evidence associating V. ceranae with BQCV may simply be a consequence of V. ceranae having not yet been recognized in A. mellifera at the time those observations were made (the early 1980s)72. Our spore load measurements were conducted by microscopy and do not discriminate between V. apis and V. ceranae, but our endpoint PCR analysis shows that V. apis was detected in only 20.3% of samples; therefore, V. ceranae is inferred as the main spore contributor. An association between both V. ceranae and V. apis and BQCV is conceivable, as both parasites infect honey bees via the midgut epithelium72. Some researchers have found that BQCV interacts with both V. apis and V. ceranae, but a reversed interaction was tested (BQCV intensifying Vairimorpha infections, rather than Vairimorpha intensifying BQCV)73. Nonetheless, the data collectively indicate that synergism between both Vairimorpha species and BQCV can occur. This may explain why occurrence of V. apis alone was a poor predictor of BQCV intensity, as every sample testing positive for V. apis also tested positive for V. ceranae, which had much higher prevalence (detected in 82.8% of samples).
Of all the regions tested, the colonies located in BC (where sampling was concentrated in the Lower Mainland) had consistently high virus occurrence, averaging at five viruses detected per sample. This was higher than previously reported virus occurrence data in pooled samples from Canada, where most samples contained two viruses (although only seven viruses were tested, compared to nine here)38. We also previously observed higher Varroa abundance and M. plutonius occurrence in BC compared to colonies in other regions71. While the data also vary by year, the pattern for colonies in BC to experience relatively high parasite and pathogen abundance is consistent with prior survey data published by Borba et al.74, where BC colonies again had the highest Varroa abundance and MB and BC had the highest pathogen abundances—traits associated with low fall colony weights and small spring and fall colony cluster sizes. BC also had the highest Varroa abundance among all 10 provinces in the 2017 Canadian National Honey Bee Health Survey37, and the highest DWV abundance in an independent survey of Canadian provinces38. The reasons for this pattern of BC colony affliction are not strictly known and are not consistent for all pathogens (BC had the fourth highest incidence of M. plutonius in the afore-referenced survey, for example), but we speculate that high densities of small-scale apiaries, particularly in the Lower Mainland, combined with short winter brood breaks and a longer beekeeping season may promote the development and spread of infections.
SBV was the only virus we identified as being significantly influenced by proximity to focal crops. We previously reported an association between SBV and highbush blueberry pollination—an analysis which utilized some of the data presented here, as well as an additional time point taken a further two weeks after the end of the pollination period (when the difference in SBV occurrence became apparent)48. This observation, combined with the interaction we identified between crop proximity, date, and region here (Fig. 3a), suggests that SBV may be influenced by landscape parameters more than the other viruses. Bailey and Fernando previously found that SBV-infected workers become poor pollen foragers, that when infected bees do collect pollen it is highly contaminated with the virus, and that when infected workers were pollen-deprived, they had longer lifespans than when they were not pollen-deprived75. These findings provide plausible mechanisms for how interactions between SBV and landscape could arise, which can be tested in future work. Concerningly, in a previous survey of viruses in Canada, SBV occurrence was associated with unhealthy colonies38, and in the present dataset we found that SBV was detected in a large fraction of samples (73.6% of samples), underscoring the need to better understand this understudied virus.
Conclusion
We present an analysis of viral abundance data which highlights regional, temporal, and climatic trends among viruses. The notable exception was KBV, which did not vary by any factor tested. Although the nature of the relationships between climatic variables were inconsistent, we make the following conclusions: (1) Climatic factors do not influence viral occurrence or intensity in the way we would expect if the fecal–oral route is a major mode of transmission, (2) warmer temperatures may promote BQCV as well as the total number of infections, possibly by leading to more crowded conditions, (3) Varroa abundance was a surprisingly weak predictor in all cases, possibly because mite abundances were too low to be influential in our data, (4) Vairimorpha spp. spore loads, but not V. apis occurrence, was linked to BQCV intensity, (5) virus intensity and occurrence was highly regional (which in our case is not distinguishable between differences between operations; see Methods), even when major climatic variables are accounted for, and (6) landscape effects may be more influential for SBV than other viruses. In addition to contributing to a larger collection of data on honey bee pathogens in Canada that can be used to determine how prevalence is changing over time, this study also improves our understanding of the potential effects of environmental variables on important viral pathogens.
Methods
This work involves the analysis of two datasets: a primary dataset derived from virus testing conducted on colonies across Canada, and a validation dataset derived from separate virus testing conducted on colonies within Alberta. The specific methods related to each dataset are described further below.
Ethics declaration
Honey bees, as non-cephalopod invertebrates, are not subject to Animal Ethics approval at the institutions involved in this study.
Primary dataset
Honey bee colonies and sampling
The honey bee colonies, sampling methods, and pathogen quantification methods have been described in detail elsewhere48,49. Briefly, a total of 480 colonies participated in this study, all of which were sampled at three time points. All were headed by locally overwintered queens and were maintained in standard deep Langstroth hives, consisting of one or two boxes as required by pollination standards for specific crops. Colonies were located in five provinces (BC, AB, MB, ON, and QC) with two regions within AB (southern AB, or AB (S), and northern AB, or AB (N)), three within QC (near Quebec City, or QC (QC), near Lac St. Jean, or QC (LSJ), and near Montreal, or QC (Mon)), and one each for BC, MB, and ON. The sampling was conducted in two years, 2020 and 2021, using different colonies in each year. In each region, colonies were derived from a single operation, and, although management practices were standardized across regions, other operational differences are thus indistinguishable from regional differences.
Colonies were assigned to one of eight focal crops (highbush blueberries, lowbush blueberries, commodity canola, seed canola, soybean, corn, apple, and cranberry), such that a total of 40 colonies were assigned to a focal crop in a given year. Highbush blueberry, cranberry, commodity canola, and seed canola were assessed in both years (totalling 80 colonies each), while lowbush blueberries, soybean, corn, and apple were assessed in one year (40 colonies each). For each focal crop, half the colonies were randomly assigned to pollination yards placed beside the crop fields (“near” group) at the beginning of the pollination period, while the other half were placed farther away from the crop (> 1.5 km away for all crops except highbush blueberries, which were > 1.3 km away; “far” group). Adult bee samples were collected from the brood nest at the beginning, middle, and end of the pollination period for each crop, except corn, which was sampled before sowing, during sowing, and at the end of the corn growing season. As such, the length of time between the first and last sampling events varied for each crop, from a minimum of 3 weeks for apple and maximum of 19 weeks for corn.
Virus, Varroa, and Vairimorpha spp. testing
As previously described48,49, adult bee samples were pooled for molecular analysis such that bees from four colonies were co-extracted from pooled samples (n = 15 bees from each colony; 60 bees total per pooled sample). The four colonies constituting a pooled sample were always located in the same apiary as one another when they travelled from holding yards to near or far yards. Two pooled samples for each group of four colonies were analyzed. One sample was used for determining Vairimorpha spp. spore counts by microscopy, and DNA was extracted from the same sample to determine occurrence of V. apis and V. ceranae by endpoint PCR. The other pooled sample was used for determining virus (LSV, BQCV, SBV, DWV-A, DWV-B, IAPV, ABPV, CBPV, and KBV) copy numbers by RT-qPCR. Detailed methods for PCR analysis, including primers, are described elsewhere48,49. Samples for determining Varroa infestation rates (mites per 100 bees, determined from samples of ~ 260 bees per colony) were taken at the same time as bees for molecular analysis. Mite counts for each colony were determined by the alcohol wash method74, and infestation rates were averaged among colonies belonging to the same pooled sample unit. All pathogen, parasite, and climate data (see below) are available in Supplementary Data 1 and 2.
Validation dataset
Honey bee colonies and sampling
The validation dataset includes a total of 4,910 colonies, sampled across 491 apiaries in Alberta over a three-year period from 2020 to 2023. Sampling took place in the spring, summer (from a subset of colonies), and fall each year, with different colonies sampled each year. The total number of colonies sampled each year in the spring, summer, and fall, respectively, was: 2021—690, 550, and 690; 2022—700, 470, and 680; and 2023—440, 290, and 400. Sampling was conducted within 34 distinct beekeeping operations distributed across Alberta’s five beekeeping regions. The number of operations in 2021, 2022, and 2023, respectively, was 26, 24, and 15.
Each apiary contributed 10 colonies to the dataset. Similar to the primary dataset, the sampling process involved collecting adult bee samples from the brood nest. In each apiary, bee samples from all 10 colonies were pooled into one live bee shipping box and shipped to the National Bee Diagnostic Center in Beaverlodge, Alberta, for viral analysis. The final sample size of the dataset after pooling was 491 samples across years and seasons. Collection times were adjusted to align with the seasonal activity of the colonies: early post-winter recovery in the spring, peak foraging and brood-rearing in the summer, and pre-winter preparation in the fall. All colonies were queenright at the time of sampling.
Virus testing
Virus copy numbers for the validation dataset were determined by RT-qPCR. Detailed methods to isolate total RNA, cDNA synthesis, qPCR quantification and analysis, including primers, are described elsewhere74. All pathogen and climate data (see below) are available in Supplementary Data 3.
Climate data extraction
Climate data extraction was performed exactly as previously described48. Briefly, data from Environment and Climate Change Canada climate stations76 were used to extract hourly and daily mean temperatures (°C), average daytime wind speeds (km/h), and total precipitation (mm). All variables were averaged over the three-week time period prior to the sampling date, as long-term averaging has been shown to improve climatic modelling of honey bee diseases77, and this is the longest time period for which we had knowledge of exact colony locations prior to every sampling event.
A subset of the colonies contributing to the validation dataset summer time point (21% of samples) were taken shortly after (1–2 weeks) the colonies were moved into pollination yards, and the exact GPS coordinates of the colonies prior to moving is unknown. Most of these colonies, however, were located in the general area of the pollination zone and likely experienced similar climatic conditions to the sampling location. We tested the impact of this discrepancy on our results by analyzing both the censored and uncensored data, and found that models fit similarly, with no change in effect sign nor significance for any factor. We therefore have reported the uncensored analysis.
Statistical analysis
We used the lme478 package in R (version 4.3.0), using R Studio (version 2023.09.1 + 494), to perform statistical analysis (see model descriptions for each virus below). In all cases, appropriateness of model fit was confirmed using tools within the DHARMa package79.
Primary dataset
Preliminary inspection of the data showed that only LSV, BQCV, and SBV had sufficient prevalence and unimodality to model viral intensities. For these viruses, intensity (non-zero viral copies per bee, log10-transformed) was analysed using linear mixed models with pooled sample unit included as a random intercept term. In all cases except BQCV, we fit an initial model with region, proximity, and calendar date as a three-way interaction, year as a fixed effect, and temperature, precipitation, and wind speed as a three-way interaction term. In a reductive modelling approach, we then simplified the model stepwise to remove non-significant interaction terms until the simplest model was achieved while still retaining appropriate residual distributions. BQCV intensities were modelled using the same strategy, but we also included log10 transformed Vairimorpha spp. spore loads as an interactor with the climate variables. LSV intensities followed a left-tailed distribution, and it was necessary to censor samples below 3.16 × 106 copies per bee to yield a well-fitting linear model. BQCV and SBV were censored at ≤ 35 copies per bee (35 copies per bee is below the limit of quantitation, but not the limit of detection). We modelled the number of viruses per sample using the same approach, but with no censoring.
In addition to modelling the intensities of the most prevalent viruses, we also modelled occurrence (1 = present, 0 = absent) of all viruses using a generalized linear mixed model. We followed the same approach as described above, again with pooled sample unit as a random intercept term, but with a binomial distribution specified.
The final models fit for each virus, as well as comparisons to more complex models describe below, are indicated in the corresponding tables describing statistical parameters for each model (Tables 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11). For all viruses, once the final model was determined, we compared that model to the same model with the added covariate of mite abundance using a χ2 test and the Akaike information criterion (AIC). Because some literature suggests that BQCV and V. apis interact synergistically (whereby V. apis infection enables greater BQCV infection by damaging the gut epithelium)28,29, we also tested including V. apis occurrence as a covariate in the model (instead of Vairimorpha spp. spore loads) using the same strategy.
Validation dataset
These data were derived from sampling that occurred over a longer time period than the primary dataset; therefore, we first filtered the data to cover a comparable duration (April–September; final N 181 apiaries sampled up to three times, for a total of 401 observations). We then modelled the data according to the same principles as described above, except that (1) we found that date was best modelled as a quadratic term (likely owing to sampling time points spanning a longer duration than the primary dataset), and (2) the distribution of SBV intensities was bimodal and as such we were unable to generate a well-fitting model using the complete dataset; therefore, we censored the data if log10(SBV copies per bee) < 4.5, which was both the local minimum between modes and the minimum filtering required to achieve adequate residual distributions.
Data availability
All data underlying the analyses presented in this manuscript are available in Supplementary Data files 1, 2. Supplementary Data 1 contains virus, Vairimorpha, and Varroa data and climate variables for 2020 (primary dataset). Supplementary Data 2 contains virus, Vairimorpha, and Varroa data and climate variables for 2021 (primary dataset). Supplementary Data 3 contains Alberta colony health monitoring project virus data and climate variables for 2021–2023 (validation dataset).
References
White GF. Sacbrood, a disease of bees. 1-5 (US Government Printing Office, Washington, D.C, U.S.A. 1913).
Brutscher, L. M., McMenamin, A. J. & Flenniken, M. L. The buzz about honey bee viruses. PLoS Pathog. 12, e1005757 (2016).
McMenamin, A. J. & Flenniken, M. L. Recently identified bee viruses and their impact on bee pollinators. Curr. Opin. Insect Sci. 26, 120–129 (2018).
McMenamin, A. J. & Genersch, E. Honey bee colony losses and associated viruses. Curr. Opin. Insect Sci. 8, 121–129. https://doi.org/10.1016/j.cois.2015.01.015 (2015).
Johnson, R. M., Evans, J. D., Robinson, G. E. & Berenbaum, M. R. Changes in transcript abundance relating to colony collapse disorder in honey bees (Apis mellifera). Proc. Natl. Acad. Sci. U. S. A. 106, 14790–14795. https://doi.org/10.1073/pnas.0906970106 (2009).
Nazzi, F. et al. Synergistic parasite-pathogen interactions mediated by host immunity can drive the collapse of honeybee colonies. PLoS Pathog. 8, e1002735 (2012).
Cox-Foster, D. L. et al. A metagenomic survey of microbes in honey bee colony collapse disorder. Science 318, 283–287 (2007).
Cornman RS et al. Pathogen webs in collapsing honey bee colonies. PLoS One 7, e43562 (2012).
Beaurepaire, A. et al. Diversity and global distribution of viruses of the western honey bee Apis mellifera. Insects 11, 239 (2020).
de Miranda, J. R. et al. Cold case: The disappearance of Egypt bee virus, a fourth distinct master strain of deformed wing virus linked to honeybee mortality in 1970’s Egypt. Virol. J. 19, 12 (2022).
Chen, Y. P. & Siede, R. Honey bee viruses. Adv. Virus Res. 70, 33–80 (2007).
Bailey, L. & Ball, B. Honey Bee Pathology 2nd edn. (UK. Academic Press Ltd, 1991).
Genersch, E. et al. The German bee monitoring project: a long term study to understand periodically high winter losses of honey bee colonies. Apidologie 41, 332–352 (2010).
Daughenbaugh, K. F. et al. Honey bee infecting Lake Sinai viruses. Viruses 7, 3285–3309 (2015).
Berthoud, H., Imdorf, A., Haueter, M., Radloff, S. & Neumann, P. Virus infections and winter losses of honey bee colonies (Apis mellifera). J. of Apic. Res. 49, 60–65 (2010).
Johnson, R. M., Evans, J. D., Robinson, G. E. & Berenbaum, M. R. Changes in transcript abundance relating to colony collapse disorder in honey bees (Apis mellifera). Proc. Natl. Acad. Sci. U. S. A. 106, 14790–14795 (2009).
McMenamin, A. J., Brutscher, L. M., Glenny, W. & Flenniken, M. L. Abiotic and biotic factors affecting the replication and pathogenicity of bee viruses. Curr. Opin. Insect Sci. 16, 14–21 (2016).
Yañez, O. et al. Bee viruses: Routes of infection in Hymenoptera. Front. Microbiol. 11, 537174 (2020).
Rosenkranz, P., Aumeier, P. & Ziegelmann, B. Biology and control of Varroa destructor. J. Invertebr. Pathol. 103, S96–S119 (2010).
Traynor, K. S. et al. Varroa destructor: A complex parasite, crippling honey bees worldwide. Trends Parasitol. 36, 592–606. https://doi.org/10.1016/j.pt.2020.04.004 (2020).
Genersch E, Aubert M. Emerging and re-emerging viruses of the honey bee (Apis mellifera L). Veterinary Research 41, 54 (2010).
De Miranda, J. R. et al. Standard methods for virus research in Apis mellifera. J. Apic. Res. 52, 1–56 (2013).
Al Naggar, Y. & Paxton, R. J. Mode of transmission determines the virulence of black queen cell virus in adult honey bees, posing a future threat to bees and apiculture. Viruses https://doi.org/10.3390/v12050535 (2020).
Bailey, L., Gibbs, A. & Woods, R. Two viruses from adult honey bees (Apis mellifera Linnaeus). Virology 21, 390–395 (1963).
Maori, E. et al. Isolation and characterization of Israeli acute paralysis virus, a dicistrovirus affecting honeybees in Israel: evidence for diversity due to intra-and inter-species recombination. J. Gen. Virol. 88, 3428–3438 (2007).
Tokarev, Y. S. et al. A formal redefinition of the genera Nosema and Vairimorpha (Microsporidia: Nosematidae) and reassignment of species based on molecular phylogenetics. J. Invertebr. Pathol. 169, 107279 (2020).
Bartolomé, C. et al. The recent revision of the genera Nosema and Vairimorpha (Microsporidia: Nosematidae) was flawed and misleads bee scientific community. J. Invertebr. Pathol. 206, 108146 (2024).
Bailey, L., Ball, B. V. & Perry, J. The prevalence of viruses of honey bees in Britain. Ann. Appl. Biol. 97, 109–118 (1981).
Bailey, L., Ball, B. V. & Perry, J. Association of viruses with two protozoal pathogens of the honey bee. Ann. Appl. Biol. 103, 13–20 (1983).
Berényi, O., Bakonyi, T., Derakhshifar, I., Köglberger, H. & Nowotny, N. Occurrence of six honeybee viruses in diseased Austrian apiaries. Appl. Environ. Microbiol. 72, 2414–2420 (2006).
Tentcheva, D. et al. Prevalence and seasonal variations of six bee viruses in Apis mellifera L. and Varroa destructor mite populations in France. Appl. Environ. Microbiol. 70, 7185–7191 (2004).
Traynor, K. S. et al. Multiyear survey targeting disease incidence in US honey bees. Apidologie 47, 325–347 (2016).
D’Alvise, P., Seeburger, V., Gihring, K., Kieboom, M. & Hasselmann, M. Seasonal dynamics and co-occurrence patterns of honey bee pathogens revealed by high-throughput RT-qPCR analysis. Ecol. Evol. 9, 10241–10252 (2019).
Chen, G. et al. Seasonal variation of viral infections between the eastern honey bee (Apis cerana) and the western honey bee (Apis mellifera). MicrobiologyOpen 10, e1162 (2021).
Blažytė-Čereškienė, L. et al. A three-year survey of honey bee viruses in Lithuania. J. Apic. Res. 55, 176–184 (2016).
Desai, S. D. & Currie, R. W. Effects of wintering environment and parasite–pathogen interactions on honey bee colony loss in north temperate regions. PLoS One 11, e0159615 (2016).
National Bee Diagnostic Center. Canadian National Honey Bee Health Survey: 2017 Report. Available at https://www.nwpolytech.ca/doc.php?d=2017NHBHS. (2017).
Desai, S. D., Kumar, S. & Currie, R. W. Occurrence, detection, and quantification of economically important viruses in healthy and unhealthy honey bee (Hymenoptera: Apidae) colonies in Canada. Can. Entomol. 148, 22–35 (2016).
Runckel, C. et al. Temporal analysis of the honey bee microbiome reveals four novel viruses and seasonal prevalence of known viruses, Nosema, and Crithidia. PLoS One 6, e20656 (2011).
Kandel, M., Paxton, R. J. & Al Naggar, Y. Nationwide screening for bee viruses in Apis Mellifera colonies in Egypt. Insects 14, 172 (2023).
Kukielka, D., Perez, A. M., Higes, M., del Carmen Bulboa, M. & Sánchez-Vizcaíno, J. M. Analytical sensitivity and specificity of a RT-PCR for the diagnosis and characterization of the spatial distribution of three Apis mellifera viral diseases in Spain. Apidologie 39, 607–617 (2008).
Kojima, Y. et al. Infestation of Japanese native honey bees by tracheal mite and virus from non-native European honey bees in Japan. Microb. Ecol. 62, 895–906 (2011).
Correa-Benítez, A. et al. Prevalence of adult honey bee (Apis mellifera L.) pests and pathogens in the five beekeeping regions of Mexico. Animals 13, 1734 (2023).
McMenamin, A. J., Daughenbaugh, K. F. & Flenniken, M. L. The heat shock response in the western honey bee (Apis mellifera) is antiviral. Viruses https://doi.org/10.3390/v12020245 (2020).
Dalmon, A., Peruzzi, M., Le Conte, Y., Alaux, C. & Pioz, M. Temperature-driven changes in viral loads in the honey bee Apis mellifera. J. Invertebr. Pathol. 160, 87–94 (2019).
Piot, N. et al. Honey bees and climate explain viral prevalence in wild bee communities on a continental scale. Sci. Rep. 12, 1904 (2022).
Prado, A. et al. Warmer winters are associated with lower levels of the cryoprotectant glycerol, a slower decrease in vitellogenin expression and reduced virus infections in winter honeybees. J. Insect Physiol. 136, 104348 (2022).
McAfee, A. et al. Higher prevalence of sacbrood virus in Apis mellifera (Hymenoptera: Apidae) colonies after pollinating highbush blueberries. J. Eco. Entomol. https://doi.org/10.1093/jee/toae119 (2024).
French, S. K. et al. Honey bee stressor networks are complex and dependent on crop and region. Curr. Biol. 34, 1893–1903. https://doi.org/10.1016/j.cub.2024.03.039 (2024).
Ryabov, E. V. et al. Dynamic evolution in the key honey bee pathogen deformed wing virus: Novel insights into virulence and competition using reverse genetics. PLoS Biol. 17, e3000502. https://doi.org/10.1371/journal.pbio.3000502 (2019).
Ryabov, E. V. et al. The vectoring competence of the mite Varroa destructor for deformed wing virus of honey bees is dynamic and affects survival of the mite. Front. Insect Sci. 2, 931352 (2022).
Gisder, S. & Genersch, E. Direct evidence for infection of Varroa destructor mites with the bee-pathogenic deformed wing virus variant B, but not variant A, via fluorescence in situ hybridization analysis. J. Virol. 95, 10–1128 (2021).
Chen, Y., Pettis, J. S., Collins, A. & Feldlaufer, M. F. Prevalence and transmission of honeybee viruses. Appl. Environ. Microbiol. 72, 606–611 (2006).
Bernardi, S. & Venturino, E. Viral epidemiology of the adult Apis mellifera infested by the Varroa destructor mite. Heliyon 2, e00101. https://doi.org/10.1016/j.heliyon.2016.e00101 (2016).
Francis, R. M., Nielsen, S. L. & Kryger, P. Varroa-virus interaction in collapsing honey bee colonies. PLoS One 8, e57540. https://doi.org/10.1371/journal.pone.0057540 (2013).
Mondet, F., de Miranda, J. R., Kretzschmar, A., Le Conte, Y. & Mercer, A. R. On the front line: quantitative virus dynamics in honeybee (Apis mellifera L.) colonies along a new expansion front of the parasite Varroa destructor. PLoS Pathog. 10, e1004323. https://doi.org/10.1371/journal.ppat.1004323 (2014).
Hasegawa, N. et al. Evolutionarily diverse origins of deformed wing viruses in western honey bees. Proc. Natl. Acad. Sci. U. S. A. 120, e2301258120 (2023).
Ryabov, E. V. et al. A virulent strain of deformed wing virus (DWV) of honeybees (Apis mellifera) prevails after Varroa destructor-mediated, or in vitro, transmission. PLoS Pathog. 10, e1004230. https://doi.org/10.1371/journal.ppat.1004230 (2014).
Nguyen, B. K. et al. Effects of honey bee virus prevalence, Varroa destructor load and queen condition on honey bee colony survival over the winter in Belgium. J. Apic. Res. 50, 195–202 (2011).
Lopes, A. R., Low, M., Martín-Hernández, R., de Miranda, J. R. & Pinto, M. A. Varroa destructor shapes the unique viral landscapes of the honey bee populations of the Azores archipelago. PloS Pathog. 20, e1012337 (2024).
Doublet, V. et al. Shift in virus composition in honeybees (Apis mellifera) following worldwide invasion by the parasitic mite and virus vector Varroa destructor. R. Soc. Open Sci. 11, 231529 (2024).
Alger, S. A., Burnham, P. A., Boncristiani, H. F. & Brody, A. K. RNA virus spillover from managed honeybees (Apis mellifera) to wild bumblebees (Bombus spp.). PLoS One 14, e0217822. https://doi.org/10.1371/journal.pone.0217822 (2019).
Burnham, P. A. et al. Flowers as dirty doorknobs: Deformed wing virus transmitted between Apis mellifera and Bombus impatiens through shared flowers. J. Appl. Ecol. 58, 2065–2074 (2021).
Bailey, L., Ball, B. & Perry, J. Honeybee paralysis: its natural spread and its diminished incidence in England and Wales. J. Apic. Res. 22, 191–195 (1983).
Deng, Y. et al. Chronic bee paralysis virus exploits host antimicrobial peptides and alters gut microbiota composition to facilitate viral infection. ISME J. 18, wrae051 (2024).
Fries, I. & Camazine, S. Implications of horizontal and vertical pathogen transmission for honey bee epidemiology. Apidologie 32, 199–214 (2001).
Betti, M. & Shaw, K. A multi-scale model of disease transfer in honey bee colonies. Insects 12, 700 (2021).
Bailey, L. & Milne, R. The multiplication regions and interaction of acute and chronic bee-paralysis viruses in adult honey bees. J. Gen. Virol. 4, 9–14 (1969).
Hung, A. PCR detection of Kashmir bee virus in honey bee excreta. J. Apic. Res. 39, 103–106 (2000).
Chen, Y. P. et al. Israeli acute paralysis virus: epidemiology, pathogenesis and implications for honey bee health. PLoS Pathog. 10, e1004261 (2014).
McAfee, A. et al. Climatic predictors of prominent honey bee (Apis mellifera) disease agents: Varroa destructor, Melissococcus plutonius, and Vairimorpha spp. PLOS Clim. 3, e0000485 (2024).
Goblirsch, M. Nosema ceranae disease of the honey bee (Apis mellifera). Apidologie 49, 131–150 (2018).
Gajda, A. M., Mazur, E. D., Bober, A. M. & Czopowicz, M. Nosema ceranae interactions with Nosema apis and black queen cell virus. Agriculture 11, 963 (2021).
Borba, R. S. et al. Phenomic analysis of the honey bee pathogen-web and its dynamics on colony productivity, health and social immunity behaviors. PLoS One 17, e0263273 (2022).
Bailey, L. & Fernando, E. Effects of sacbrood virus on adult honey-bees. Ann. Appl. Biol. 72, 27–35 (1972).
Government of Canada Environment and Natural Resources. Past weather and climate: Historical data. https://climate.weather.gc.ca/historical_data/search_historic_data_e.html. Accessed January 2024. (2024).
Rowland, B. W., Rushton, S. P., Shirley, M. D. F., Brown, M. A. & Budge, G. E. Identifying the climatic drivers of honey bee disease in England and Wales. Sci. Rep. 11, 21953. https://doi.org/10.1038/s41598-021-01495-w (2021).
Bates, D., Maechler, M., Bolker, B. & Walker, S. Fitting linear mixed-effects models using lme4. J. Stat. Softw. 67, 1–48. https://doi.org/10.18637/jss.v067.i01 (2015).
Hartig, F., Lohse, L., and de Souza leite, M. DHARMa: Residual diagnostics for hierarchical (multi-level/mixed) regression models. R package version 0.4.6. Available at https://CRAN.R-project.org/package=DHARMa. (2022).
Acknowledgements
We would like to acknowledge A. Achkanian, R. Bahreini, S. Bailey, V. Beger, C. Bryant, A. Chapman, M. Chihata, C. Currie, K. Dave, G.L. Ganesan, P. Grubiak, D. Holder, A. Ibrahim, M.A. Imrit, J. Janisse, J. Kearns, K. Lynch, D. Micholson, M. Munro, L. Ovinge, M. Peirson, K. Peters, Z. Rempel, R. Thygesen, D. Tran, A. Travas, B. Vinson, E. Olson, and N. McCormick for their supportive efforts. We are also grateful to all members of the BeeCSI Consortium (https://beecsi.ca/beecsi-consortium) who participated this project as well as all growers and beekeepers who let us use their field sites. This work was funded by Genome BC through the Genomic Innovation for Regenerative Agriculture, Food and Fisheries (GIRAFF) program to LJF, LT, MMG, CAB, and JD. Data analyzed in this work was derived from the BeeCSI project, which was funded and supported by the Ontario Genomics Institute (OGI-185) to AZ, Genome Canada (LSARP #16420) to AZ, LJF, PG, SEH, EG, RWC, MMG and SFP, the Ontario Research Fund (LSARP #16420) to AZ, Genome Quebec, and the Government of Canada through Agriculture and Agri-Food Canada (AAFC) Genomics Research and Development Initiative (GRDI) funding (AAFC J-002368) to MMG and SFP. LT and EMW received salaries in part paid from AAFC GDRI funds, and MC received a stipend as Research Affiliate from AAFC as well as funds derived from the GIRAFF program. Salaries for AM, JC, HH, SKF, IMC, and DB were derived in part from funds awarded by Genome Canada. Salary for AM ere also derived in part from funds derived from the GIRAFF program. This work was also supported by the Canadian Honey Council, the Ontario Technology Transfer Program (of which DB is a member) of the Ontario Beekeepers’ Association, and the Alberta Technology Transfer Program (led by RL) of the Alberta Beekeepers Commission.
Author information
Authors and Affiliations
Contributions
AM conducted the statistical analysis, generated the figures, and wrote the first draft of the manuscript. NA extracted and prepared the climate data. RL led the Alberta TTP honey bee health monitoring project, from which the validation dataset was derived, and curated the data. Funding awarded to LT, SFP, PG, SEH, EG, RWC, CAB, AZ, JD, MMG, and LJF, all of whom contributed to conceptualizing this project, supported this research. JC, HH, EMW, and DB made significant contributions to honey bee colony management for this study. PWV conducted the virus testing. LT, SKF, IMC, and EMW oversaw project management operations, sample organization, and transfer of materials and data between institutions. All authors reviewed and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
McAfee, A., Alavi-Shoushtari, N., Labuschagne, R. et al. Regional patterns and climatic predictors of viruses in honey bee (Apis mellifera) colonies over time. Sci Rep 15, 286 (2025). https://doi.org/10.1038/s41598-024-79675-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-79675-7
Keywords
This article is cited by
-
Spatial and environmental drivers of Varroa destructor detection in New South Wales, Australia
Scientific Reports (2025)
-
Neonicotinoid-induced signature dysbiosis identified via metagenomic sequencing of the honey bee gut microbiome
Scientific Reports (2025)