Abstract
Esophageal adenocarcinoma (EAC) is an aggressive cancer characterized by a high risk of relapse post-surgery. Current follow-up methods (serum carcinoembryonic antigen detection and PET-CT) lack sensitivity and reliability, necessitating a novel approach. Analyzing cell-free DNA (cfDNA) from blood plasma emerges as a promising avenue. This study aims to evaluate the cost-effective and genome-wide cell-free reduced representation bisulfite sequencing (cfRRBS) method combined with computational deconvolution for effective disease monitoring in EAC patients. cfDNA methylation profiling with cfRRBS was performed on 162 blood plasma samples from 33 EAC cancer patients and 28 blood plasma samples from 20 healthy donors. The estimated tumor fraction for EAC patients at the time of diagnosis was significantly different from the healthy donor plasma samples (one-sided Wilcoxon rank-sum test: p-value = 0.032). Tumor fractions above 15% and focal gains/amplifications in MYC (chr8), KRAS (chr12), EGFR (chr7) and NOTCH2 (chr1) were observed in four samples of distinct patients at the time metastatic disease was detected. This study showed feasibility to estimate tumor fractions in blood plasma of EAC patients based on cfDNA methylation using cfRRBS and computational deconvolution. Nevertheless, in this study only cancer patients with evidence of metastatic disease show high tumor fractions and copy number alterations.
Similar content being viewed by others
Introduction
Esophageal adenocarcinoma (EAC) is a histological subtype of esophageal cancer that occurs in the lower part of the esophagus. It is an aggressive cancer with a poor five-year survival of only 25%, despite improved multimodal treatment regimens. The current gold standard for the diagnosis of EAC is based on imaging techniques and requires an upper endoscopy with a tissue biopsy for confirmation. Treatment consists of neoadjuvant therapy (depending on tumor and lymph node stage) and surgical resection of curable (localized) disease. Despite these interventions, the risk of relapse and metastasis remains notably high post-surgery: 35% of the patients that received neoadjuvant therapy and surgery (CROSS trial) eventually relapse (i.e. local recurrence or metastasis), despite apparently curative and complete tumor resection1. Routine follow-up methods used in most centers include gastroscopy (with biopsy), computed tomography (CT) and positron emission tomography (PET) if a relapse is suspected based on symptoms (ESMO guidelines). The combination of the last two imaging modalities (PET-CT) is more sensitive and specific for the detection of local recurrence and distant metastasis compared to PET alone2,3. However, PET-CT is only done in follow-up when metastasis or local recurrence is suspected from the CT scan or increasing serum carcinoembryonic antigen (CEA) values (in some centers). Unfortunately, distinguishing small tumor lesions from normal esophageal metabolic activity or post-radiation esophagitis using PET-CT within 2–3 months post-operatively is unreliable4. Moreover, CEA values have shown not to be reliable for EAC, with sensitivities between 19 and 39% for detecting recurrence and specificity of 89%5. Given high chance of relapse post-surgery, there is an urgent need for improved disease monitoring of EAC patients to identify patients at risk for relapse.
Analysis of cell-free DNA (cfDNA) from blood plasma offers a minimally-invasive approach to follow-up the tumor during therapy as plasma sampling can be performed longitudinally. Through active secretion via vesicles and apoptosis/necrosis of cancer cells, cfDNA originating from tumor cells (ctDNA) is released in the bloodstream. Genomic analyses of circulating tumor DNA (ctDNA) in blood plasma have shown great potential in the context of diagnosis, treatment response monitoring and minimal residual disease detection in various cancer types6. Another advantage of cfDNA monitoring is that cfDNA is mostly shed from the various tumor (sub)clones at both primary and metastatic sites and therefore better represents the tumor heterogeneity. An important challenge with cfDNA isolated from a cancer patient’s blood plasma is that it also contains cfDNA originating from healthy cells7. For that reason, mutation detection for disease monitoring requires prior knowledge about the mutations that are present in the tumor when using a targeted deep sequencing approach, or need to sequence cfDNA from white blood cells to enable correction for clonal hematopoiesis of indeterminate potential (CHIP) variants. Given the tissue-specificity of DNA-methylation profiles8, this genome-wide approach can be used to discriminate tumor-derived cfDNA fragments from those originating from healthy cells. Subsequently, the fraction of tumoral fragments can be calculated, which might be indicative for the tumoral burden.
For various cancer entities cfDNA methylation profiling of longitudinally collected plasma samples has shown to be promising for disease monitoring9,10,11. For esophageal cancer methylome-wide studies have solely focused on esophageal squamous cell carcinoma12. For EAC, cfDNA methylation has mostly been studied for diagnostic and screening purposes13,14. Regarding disease monitoring using cfDNA, so far only one relevant study was done, where authors showed that hypomethylation of LINE-1 was associated with EAC development15.
We put forward that monitoring the tumoral cfDNA fraction in blood plasma through DNA methylation profiling may aid in the follow-up of patients with esophageal adenocarcinoma and pinpoint the patients at risk for relapse. To enable cost-effective genome-wide methylation profiling on highly fragmented and low abundant cfDNA, cell-free reduced representation bisulfite sequencing (cfRRBS) is used as described16,17,18. From these DNA methylation profiles, the tumoral fraction can be estimated using computational deconvolution17. In this study, we aim to evaluate the potential of genome-wide cfRRBS in combination with deconvolution for disease monitoring in 33 EAC patients (predominantly T3 tumor stage) that were eligible for surgery and received neo-adjuvant therapy.
Results
Quality control of the cfRRBS sequencing data
cfRRBS sequencing was successfully performed on 200 cfDNA samples isolated from blood plasma of 33 EAC patients (162 samples, nine repeated samples) and of 20 healthy donors (28 samples) as well as 7 FFPE EAC tissue samples. All samples included in this study showed high sequencing quality and were included in downstream analyses. The median number of raw sequencing reads per sample was 23.75 M (range 10.67–165.84 M) (Supplementary Fig. 1A). All samples had an acceptable conversion rate (greater than 95%17), with a median bisulfite conversion percentage of 99.6% (range 97.7–99.8) (Supplementary Fig. 1B). Enzymatic digestion of the DNA by MspI results in fragments with CpG rich ends, thus mapped reads were expected to mostly map to these regions. The median percentage of bases mapping to these bait regions (MspI regions between 20 and 200 bp) was 89.48% (80.79–95.05%) (Supplementary Fig. 1C). The median mapping efficiency (aligned reads/(aligned + ambiguous + discarded reads) * 100) was 65.48% (60.93–72.26%) (Supplementary Fig. 1D). The median sequencing read length was 41.01 bp (36.95–42.54 bp). Overall, the read length in sequencing run 128 was slightly shorter compared to other runs (Supplementary Fig. 1E), but can be explained by the hard trimming settings used for these samples due to complete adapter sequence removal in the demultiplexing step. Per patient at least three collected blood plasma samples were included in this study, divided over six sequencing runs. Hierarchical clustering (Euclidean distances with Ward.D clustering) of the blood plasma samples based on the beta values (output Bismark pipeline) showed clustering by patient rather than by sequencing run, thus indicating no batch effect (Supplementary Fig. 2).
Estimated tumor fractions in EAC patients at diagnosis and healthy controls
In order to estimate the EAC tumor fraction in the cfDNA samples, beta values were used as input for computational deconvolution with MethAtlas (NNLS-based method). The estimated tumor fractions of the 162 EAC samples (from citrate blood collection tubes) and 14 healthy donor samples (from EDTA blood collection tubes) show tumor fractions between 0 and 31% (Fig. 1A). For EAC patients the median estimated tumor fraction at the time of diagnosis (t0) is 2.2% (0.0–22.6%), for healthy donors the median is 1.6% (0.0–2.6%) which is a significant difference (one-sided Wilcoxon rank-sum test: p-value = 0.032). Based on the highest estimated tumor fraction in the healthy donor samples, a cut-off value of 2.6% was set as lower limit for detection of EAC derived cfDNA.
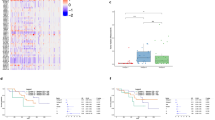
Dynamic change of tumor fractions and cfDNA concentration in plasma from EAC patients. (A) Boxplots of estimated EAC tumor fractions in the patient cfDNA sampled at different timepoints and in the 14 healthy donor samples. Based on the highest estimated tumor fraction of the healthy donor samples, a cut-off was set at 2.6%. (B) cfDNA concentration (ng/µL) in blood plasma of healthy donors and EAC patients at the different timepoints.
The comparison of tumor fractions in citrate tube EAC samples with EDTA tube healthy controls could be justified as the estimated tumor fractions in EDTA versus citrate plasma originating from the same healthy donor (six healthy donors) did not show a statistically significant difference (Wilcoxon signed-rank test: p-value = 0.46) (Fig. 2, Supplementary Table S1). The reproducibility of our approach was demonstrated by generating cfRRBS libraries from cfDNA isolated from two different blood plasma aliquots from nine patients at one specific timepoint. The median difference in estimated tumor fraction between the replicates of the nine cases that were re-prepped and re-sequenced was 0.6% (0.1–1.8%), and showed high reproducibility (Wilcoxon signed-rank test, p-value = 0.89) (Supplementary Fig. 3).
Tumor fractions in matched plasma samples collected in two blood collection tubes. No significant difference (paired Wilcoxon rank-sum test, p-value = 0.46) in EAC tumor fraction between six matched healthy donor plasma samples collected in citrate and EDTA blood collection tubes.
Increased tumor fractions after surgery for patients with metastatic disease
An important advantage of liquid biopsy sampling is the possibility to perform longitudinal analysis. Longitudinal analysis of the estimated tumor fractions showed an increase within one week after surgery (t2, Fig. 1A), which is a significant increase compared to samples at the time of diagnosis (t0) (Wilcoxon signed-rank test: p-value = 2.0e−3). While this tumor fraction increase from t0 to t2 coincided with an increase in cfDNA concentration (Fig. 1B) (Wilcoxon signed-rank test: p-value < 0.0001), there was no correlation between tumor fraction and cfDNA concentration (R2 < 0.01; p-value = 0.57) (Supplementary Fig. 4).
Next, we performed a more detailed longitudinal analysis of the estimated tumor fractions using a heatmap visualization, centering all patients’ timelines around the time of surgery (timepoint t1) (Fig. 3A). The time of metastasis (black squares) and time of death (red crosses) was indicated in the heatmap. From 6/19 patients with metastatic disease (no adjuvant therapy), a blood plasma sample was collected within one month of the detected metastasis. Three of these patients (ID25, ID48 and ID55) had estimated tumor fractions above 15% (dark blue color) between two and five months after surgery, and all eventually died due to disease progression (between 4.5 and 9.5 months after detection of metastasis) (Fig. 3B–D). Patient ID4 has a bone metastasis that was detected six months after surgery, but the blood sample collected at that time showed no increase in tumor fraction. Patient UZL11 had a liver metastasis detected shortly after surgery. The estimated tumor fraction in the blood plasma sample collected few weeks earlier showed an estimated tumor fraction of 8.5%. The last patient with a blood sample collection around the time of the detected metastasis was patient ID58. This was the only patient with a tumor fraction above 15% at the time of diagnosis and was also the only patient with metastatic disease (lymph nodes) at the time of diagnosis.
Dynamic change in estimated tumor fraction in longitudinally collected blood plasma samples from 33 EAC patients. (A) Heatmap showing estimated tumor fractions per patient and sampled timepoint. The x-axis indicates the time in months relative to the time of surgery (timepoint t1). Grey boxes indicate tumor fractions below the cut-off of 2.6%, blue between 2.6% and 15% and dark blue more than 15%. (B–D) Timelines showing increasing tumor fractions of three patients that eventually presented with a tumor fraction that is higher than 15% (after surgery).
Copy number variation analysis points at amplified oncogenes
In addition to methylation profiles, copy number variation (CNV) profiles can be retrieved from cfRRBS data (200 kb bin sizes). Focal gains/amplifications were observed in all four samples that had estimated tumor fractions above 15% (Fig. 4A–D). These focally gained/amplified regions encompass genes that are associated with tumorigenesis in EAC: MYC (chr8), KRAS (chr12), EGFR (chr7) in three samples collected post-surgery (ID25, ID48 and ID55, respectively) and NOTCH2 (chr1) in a diagnostic sample (ID58). For patient ID48, this focal gain/amplification was also observed earlier, in the blood plasma sample that was collected three weeks after surgery (sample ID48_EAC_3; tumor fraction 10.1%) (Supplementary Fig. 5). For two cases (ID25 and ID48) we also analyzed the resected tumor tissue and confirmed the presence of the focal amplification using shallow whole genome sequencing (sWGS) (Supplementary Fig. 6A,B). Comparison of the cfRRBS-based CNV profile of the cfDNA with the sWGS-based CNV profile of the tumor tissue showed that the less abundant aberrations (heterozygotic losses or gains) are (almost) not detected in the cfRRBS data.
CNV profiles based on cfRRBS data show aberrations in plasma samples with tumor fractions above 15%. (A) CNV profile of ID25_EAC_4, showing a focal gain/amplification in chromosome 8; (B) CNV profile of ID48_EAC_4, showing a focal gain/amplification in chromosome 12; (C) CNV profile of ID55_EAC_3, showing a focal gain/amplification in chromosome 7; (D) complete CNV profile of ID58_EAC_0 and a zoomed in profile of chromosome 1. The profile of this patient blood sample collected at the time of diagnosis showed a focal gain/amplification in chromosome 1.
Discussion
The current methods for follow-up of patients with EAC are imaging-based techniques such as (PET-)CT, and endoscopy with biopsies. As blood-based biopsies offer many advantages, we aimed to evaluate the potential of genome-wide cfDNA methylation profiling using cfRRBS in combination with deconvolution for disease monitoring of EAC cancer patients. The cfRRBS method is cost-effective and allows methylation profiling of fragmented cfDNA in plasma. Unlike panel/targeted sequencing for mutation detection, cfRRBS does not require prior knowledge about the (epi)genetic profile of the tumor. Nevertheless, mutation detection is known to be a sensitive and specific method for the detection of low allele frequencies in cfDNA with low tumoral fractions. For example, for longitudinally collected blood samples from EAC patients, allele frequencies as low as 0.1% were reported based on TP53 mutations (the most common mutation detected in EAC)19. The exact sensitivity and specificity of cfRRBS for disease monitoring of EAC patients has to be determined in a separate follow-up study, in which the cfRRBS method is compared to other methods.
In this study we showed the feasibility of using this method in combination with computational deconvolution to estimate tumoral cfDNA fractions allowing disease monitoring of EAC patients. We used MethAtlas as deconvolution tool given our successful application in pediatric and adult solid tumors17,18,20. Importantly, we could show that the pipeline is robust, generating reproducible results in replicate experiments and when using different blood collection tubes (such as citrate and EDTA tubes).
The overall estimated tumor fractions of the (predominantly stage T3) cancer patients are low (0–22.6%) and were significantly different from healthy individuals (0.0–2.6%). Based on the estimated tumor fractions in healthy donors, a cut-off value 2.6% was applied (highest estimated tumor fraction in the healthy donors). This is a conservative approach, and further statistical analyses based on a larger dataset are needed to more accurately establish a cut-off value that optimizes either sensitivity or specificity.
In our data we observed an increase in tumor fraction only in blood samples (4/6) that were collected around the time (within a month) metastatic disease was clinically detected. This may explain the low tumor fraction at the time of diagnosis (when no metastasis was present). Moreover, the mean tumor fraction at the time of diagnosis for EAC patients was 3.7%, which is slightly lower than reported for EAC in literature (7%, based on variant allele fractions)21. Overall, the low tumor fractions in patients with lower tumor stages (T1–T3) and non-metastatic disease patients (at the time of diagnosis), as is the case in our study, is consistent with reports from literature19,21,22.
Intriguingly, both cfDNA concentration and estimated tumor fraction peak within one week after surgery (timepoint t2). An increase in cfDNA concentration after surgery is expected, as the surgical resection causes release of healthy cfDNA23. Similarly, an overall increase in cfDNA up to four weeks after surgery has been reported for colorectal and bladder cancer23. An increased tumor fraction after surgery however is rather surprising as no additional ctDNA is expected in circulation when the tumor is removed. The lack of correlation between cfDNA concentration and tumor fraction suggests that the increase in tumor fraction is not a result of the increase in cfDNA concentration. For deconvolution, we have used a reference dataset based on cfDNA methylation profiles generated with cfRRBS of healthy donor cfDNA and FFPE tumor tissue DNA from EAC patients. Moreover, DNA methylation profiles based on publicly available whole genome bisulfite sequencing data from healthy esophageal tissue and healthy stomach tissue was included in the reference dataset to rule out that the increased tumor fraction was a result of an increase in healthy esophageal/stomach cfDNA. However, this was not the case as healthy tissue fractions did not increase (data not shown). Further refinement of the reference dataset and use of other deconvolution tools could be investigated to improve specificity.
By analyzing longitudinal samples, we identified four patients with increased estimated tumor fractions (> 15%) that matched with the clinical diagnosis of metastasis, suggesting a clinically relevant signal. Whether the increase in tumor fraction with this cfRRBS workflow is detected prior to current diagnostic tools (PET-CT) remains to be further investigated and requires additional sampling points along the timeline. However, for the 2/6 patients for which blood plasma samples were collected at the time of metastatic disease, we could not observe a similar increase in estimated tumor fraction, demonstrating that this biomarker is only marginally relevant within this patient group. Future follow-up studies should focus on determining the sensitivity of the tumor fraction estimation with the cfRRBS method with computational deconvolution, and compare the outcomes directly to clinical data (e.g. PET-CT data) obtained at the same time.
Lastly, our results demonstrate that DNA copy number variation profiles (CNV) can be retrieved from cfRRBS data. Even though the generated CNV profiles are noisier compared to more conventional methods like sWGS, abundant aberrations (focal gains/amplifications) could be identified in blood plasma samples with estimated tumor fractions greater than 15%. These focal gains/amplifications involved regions that included MYC, KRAS, EGFR and NOTCH2. Three out of four of these gene gains/amplifications have been previously reported in literature in the context of EAC: MYC, KRAS and EGFR amplification have been reported in approximately 30%, 14% and 15% of EAC cases, respectively24. While NOTCH2 amplification has not yet been reported in EAC, the Notch signaling pathway is known to play an important role in cell proliferation, differentiation and apoptosis, and dysregulation has been shown to contribute to tumorigenesis25.
In this proof-of-concept study, cfRRBS followed by computational deconvolution with MethAtlas was used to estimate tumor fractions in the blood plasma of EAC patients. This approach has demonstrated its value in three studies17,18,20, but is also characterized with some limitations that warrant consideration for future improvements. A limitation is that cfRRBS uses bisulfite conversion of unmethylated cytosines which is a chemical treatment that may cause further DNA degradation. Future developments might be directed towards enzymatic-based conversion or long-read sequencing approaches. A further limitation is that cfRRBS covers on average 3M CpG sites, which is higher compared to microarray platforms (27K or 450K CpGs), but lower compared to, for example, whole genome bisulfite sequencing (28M CpGs) or CapSeq Epi26 (5.5M CpGs). However, the computational deconvolution approach uses median beta values of CpG clusters and therefore incomplete coverage of CpGs will minimally impact the results. A final limitation and potential future improvement is the choice of deconvolution algorithm. In this study and previous studies, we have demonstrated a good performance with the MethAtlas tool. However, other more recently developed tools8,27 might further improve the sensitivity of EAC detection. Finally, as EAC is a highly heterogenous cancer type, expanding the FFPE tissue sample set in the reference data may improve the accuracy.
This study shows the feasibility of using cfRRBS and computational deconvolution for tumor fraction estimation in cfDNA isolated from blood plasma of EAC patients. Moreover, for some cases, copy number variations, including gains/amplifications, could be identified from the cfRRBS sequencing data. However, given the rather low tumor fractions in patients with stage T1-T3 EAC disease, the clinical utility of this approach for tumor monitoring is likely to be limited to patients that develop metastatic disease. More sensitive methylation profiling and deconvolution approaches that can also accurately detect lower tumor fractions are warranted for further clinical application in this disease type.
Methods
Sample collection
Blood was drawn into Vacuette Tube 9 mL 9NC Coagulation sodium citrate 3.2% (Greiner Bio-One) tubes from 33 EAC patients at different timepoints during treatment (162 collections in total) (Table 1, Supplementary File 1). Patient plasma samples were divided into seven time categories (t0–t6). In case more than one plasma sample per patient was categorized in one timepoint, only the sample that was first collected was included for statistical analyses between timepoints. Blood samples from 20 age-matched healthy donors were collected in BD Vacutainer Plastic K2EDTA (Becton Dickinson and Company, n = 18) and PAXgene Blood ccfDNA Tube (Qiagen, n = 9) blood collection tubes. Sample collection was approved by the ethics committee of Ghent University Hospital (registration numbers B670201628317, B670201628319 and B670201733701). Plasma samples were collected at Ghent University Hospital (UZ Ghent) or University Hospital Leuven (UZ Leuven) and all patients signed an informed consent prior to blood collection. Blood samples were processed at room temperature within 10 h after collection, except for ID37_EAC_0, UZL2_EAC_4, UZL14_EAC_1, UZL16_EAC_3, UZL16_EAC_5 and ID22_EAC_7 (processed within 22 h). Blood samples from EAC patients were prepared by double-spun centrifugation (two times 10 min at 2500g). Plasma was aliquoted in 2 mL DNA LoBind PCR clean tubes (Eppendorf) and stored at − 80 °C until further processing. Blood samples from healthy donors were prepared by double-spin centrifugation (EDTA tubes, two times 10 min at 2500g) or single-spun centrifugation (PAXgene tubes, 15 min at 1900g). Tumor tissue samples (FFPE: formalin-fixed, paraffin embedded) from seven EAC patients were obtained through the Department of Pathology of the Ghent University Hospital. For each patient, tumor tissue was collected from slides with hematoxylin–eosin stained FFPE sections. Per patient, tumoral regions were macro-dissected and collected in a 1.5 mL DNA LoBind PCR clean tube (Eppendorf).
DNA isolation
cfDNA from plasma from 33 EAC patients and 20 healthy donors was purified from 0.9–2.5 mL (mean: 2.0 mL) using the Maxwell RSC LV ccfDNA kit (Promega), according to the manufacturer’s instructions. Plasma was thawed at 37 °C and processed immediately. For samples with plasma volumes below 2 mL, 1× PBS (Gibco, Thermo Fisher Scientific) was added up to 2 mL before adding an equal amount of binding buffer and 140 µL beads. Samples were incubated for 45 min on the Multi Bio RS-24 Rotator (Biosan) to allow binding of cfDNA to the magnetic beads. cfDNA was recovered with the fully automated protocol on the Maxwell RSC device in 75 µL elution buffer (Promega). All cfDNA samples were stored at − 20 °C.
DNA was isolated from FFPE tumor tissue samples using the QIAamp DNA FFPE Tissue kit (Qiagen) following the manufacturer’s recommendation, except using deparaffinization solution (Qiagen) instead of xylene. DNA was eluted in 50 µL ATE buffer (Qiagen) and stored at − 20 °C.
DNA quantification and quality assessment
cfDNA was isolated from 162 plasma samples from 33 EAC patients (3–11 longitudinal samples per patient) and 28 plasma samples (EDTA and/or PAXgene tubes) from 20 healthy donors. To determine the robustness of the workflow, 9 EAC plasma samples were re-prepped and re-sequenced once (n = 8) or twice (n = 1) (Supplementary Table S2). The median cfDNA concentration measured by TapeStation (Agilent) was 0.22 ng/µL (0.01–5.72 ng/µL) (Supplementary Table S3). Typically, cfDNA has an average fragment length of around 170 bp. High molecular weight (HMW) DNA, originating from lysed cells, consists of larger fragments (greater than 700 bp). Based on fragment analysis, all samples had a cfDNA/HMW ratio greater than one (median: 5.98; range 1.15–54.56). The DNA concentration from the seven tumor tissue samples was measured with a Varioskan LUX Microplate Fluorometer (Thermo Scientific) using the Quant-iT™ 1X dsDNA kit (broad range) according to the manufacturer’s protocol. The median concentration was 32.27 ng/µL (22.49–139.50 ng/µL).
cfRRBS library construction
Library construction for cfRRBS was performed as described16. The input amount for cfDNA samples was aimed at 10 ng in 11.1 µL. Due to low cfDNA concentration in blood plasma, it was not possible to use 10 ng as input for some samples. The median cfDNA input amount for library preparation was 4.38 ng (0.66–48.50 ng) for the patient and healthy donor plasma samples (n = 200), and the input for the tissue samples was 200 ng (n = 7). If necessary, eluate volumes were concentrated by means of vacuum centrifugation (Eppendorf, Concentrator Plus, program V-AQ at 30 °C) and nuclease-free water was added to a volume of 11.1 μL. Afterwards, 0.5 µL of 0.01 ng/µL unmethylated lambda phage spike-in DNA was added to the eluate. After library construction, libraries were cleaned by magnetic bead selection (AMPure XT beads, NEB) and eluted in 0.1× TE buffer. The libraries were visualized with the Fragment Analyzer (Agilent) and quantified with a Varioskan LUX/Fluoroskan Microplate Fluorometer (Thermo Scientific) using the Quant-iT™ 1× dsDNA kit (high sensitivity). The median library concentration was 15.0 ng/µL (0.6–68.69 ng/µL). Libraries were equimolarly pooled and quantified using the Kapa library quantification kit (Kapa Biosystems). Sequencing was done in six separate runs, on the NovaSeq 6000 (Illumina) instrument (for details see Supplementary Table S4).
Pre-processing of the sequencing data
Pre-processing of the sequenced libraries was done using an in-house developed Bismark pipeline as described by Van Paemel et al.17. First, demultiplexing of the data (bcl files) was done using bcl2fastq2 v2.20.0. Adapter trimming was done with TrimGalore v0.6.6 (flags --three_prime_clip_R1 1 -- three_prime_clip_R2 1 --clip_R1 3 -- clip_R2 4 to remove methylation bias at the three or five prime end due to the MspI restriction digest). For sequencing run 128 hard trimming was needed as Illumina adapter sequences were incorrectly removed and raw sequencing data (bcl files) were not available anymore (flags --three_prime_clip_R1 3 --three_prime_clip_R2 3 --clip_R1 3 --clip_R2 4).
Bismark v0.23.1 (default parameters) was used for mapping to GRCh37. Mapped reads were sorted using SAMtools v1.14. For paired end sequencing reads, clustering duplicates were removed using Picard v2.21 with the following parameters: OPTICAL_DUPLICATE_PIXEL_DISTANCE = 12,000 (for NovaSeq 6000), REMOVE_SEQUENCING_DUPLICATES = true, TAGGING_POLICY = OpticalOnly, READ_NAME_REGEX = [a-zA-Z0-9] + :[0–9] + :[a-zA-Z0-9] + :[0–9] + :([0–9] +):([0–9] +):([0–9] +)_[0–9] + :[a-zA-Z0-9] + :[0–9] + :[a-zA-Z0-9] +\+ [a-zA-Z0-9] + (for dual index).
Bait regions are defined as MspI regions between 20–200 bp in GRCh37. CollectHsMetrics (Picard v2.21.6) was used to determine the number of reads covering these regions. The output of the in-house developed Bismark pipeline are coverage files, that includes the coverage of each methylated and unmethylated cytosine. Based on these coverage files, beta values are determined (per cytosine, reads methylated C/total reads of that C).
Establishing tumor fractions through data deconvolution
cfDNA tumor fractions were estimated using computational deconvolution of the cfDNA methylation profile. To select the most informative regions, differentially methylated region (DMR) selection was performed between the healthy donor samples (EDTA tubes only) and the seven EAC FFPE tissue samples using the default settings in DMRfinder v0.3. The resulting 9195 clusters were all used for the deconvolution. As reference dataset for deconvolution we used cfRRBS data from 14 healthy donor plasma samples (EDTA), cfRRBS data from seven EAC FFPE tissue samples, as well as publicly available whole genome bisulfite sequencing data from healthy esophagus tissue samples (n = 2) and stomach tissue samples (fundus, n = 3; body, n = 3). Data deconvolution to estimate tumor fractions was done using MethAtlas (default settings), based on non-negative least squared (NNLS) matrix decomposition28. Data visualization was done using R v4.2.329. Statistical tests were applied to compare estimated tumor fractions between healthy donors and cancer patients (at time of diagnosis) using the Wilcoxon rank-sum test (unpaired data) and Wilcoxon signed-rank test (paired data) of the R package coin (v1.4-3).
Data availability
Raw DNA sequencing data (cfRRBS data) that was generated in this study is deposited in the European Genome-phenotype Archive (EGA) under accession number EGA50000000482. Processed data (beta values) of all sequenced samples are available from ArrayExpress under accession number E-MTAB-14370. R scripts to generate the figures in this manuscript are available on GitHub (https://github.com/VIBTOBIlab, repository name: cfRRBS_EAC).
References
Oppedijk, V. et al. Patterns of recurrence after surgery alone versus preoperative chemoradiotherapy and surgery in the CROSS trials. J. Clin. Oncol. 32, 385–391. https://doi.org/10.1200/jco.2013.51.2186 (2014).
Bruzzi, J. F. et al. PET/CT of esophageal cancer: its role in clinical management. Radiographics 27, 1635–1652. https://doi.org/10.1148/rg.276065742 (2007).
Roedl, J. B. et al. Assessment of treatment response and recurrence in esophageal carcinoma based on tumor length and standardized uptake value on positron emission tomography-computed tomography. Ann. Thorac. Surg. 86, 1131–1138. https://doi.org/10.1016/j.athoracsur.2008.05.019 (2008).
Valkema, M. J. et al. Accuracy of (18)F-FDG PET/CT in predicting residual disease after neoadjuvant chemoradiotherapy for esophageal cancer. J. Nucl. Med. 60, 1553–1559. https://doi.org/10.2967/jnumed.118.224196 (2019).
Moyes, L. H., Anderson, J. E. & Forshaw, M. J. Proposed follow up programme after curative resection for lower third oesophageal cancer. World J. Surg. Oncol. 8, 75. https://doi.org/10.1186/1477-7819-8-75 (2010).
Cescon, D. W., Bratman, S. V., Chan, S. M. & Siu, L. L. Circulating tumor DNA and liquid biopsy in oncology. Nat. Cancer 1, 276–290. https://doi.org/10.1038/s43018-020-0043-5 (2020).
Siravegna, G., Marsoni, S., Siena, S. & Bardelli, A. Integrating liquid biopsies into the management of cancer. Nat. Rev. Clin. Oncol. 14, 531–548. https://doi.org/10.1038/nrclinonc.2017.14 (2017).
Loyfer, N. et al. A DNA methylation atlas of normal human cell types. Nature 613, 355–364. https://doi.org/10.1038/s41586-022-05580-6 (2023).
Silva, R. et al. Longitudinal analysis of individual cfDNA methylome patterns in metastatic prostate cancer. Clin. Epigenet. 13, 168 (2021).
Chen, K. et al. Individualized dynamic methylation-based analysis of cell-free DNA in postoperative monitoring of lung cancer. BMC Med. 21, 255 (2023).
Barault, L. et al. Discovery of methylated circulating DNA biomarkers for comprehensive non-invasive monitoring of treatment response in metastatic colorectal cancer. Gut 67, 1995–2005 (2018).
Liu, J. et al. Multimodal analysis of cfDNA methylomes for early detecting esophageal squamous cell carcinoma and precancerous lesions. Nat. Commun. 15, 3700 (2024).
Macedo-Silva, C. et al. DNA methylation biomarkers accurately detect esophageal cancer prior and post neoadjuvant chemoradiation. Cancer Med. 12, 8777–8788. https://doi.org/10.1002/cam4.5623 (2023).
Zhai, R. et al. Genome-wide DNA methylation profiling of cell-free serum DNA in esophageal adenocarcinoma and Barrett esophagus. Neoplasia 14, 29–33. https://doi.org/10.1593/neo.111626 (2012).
Boldrin, E. et al. Detection of LINE-1 hypomethylation in cfDNA of esophageal adenocarcinoma patients. Int. J. Mol. Sci. 21. https://doi.org/10.3390/ijms21041547 (2020).
De Koker, A., Van Paemel, R., De Wilde, B., De Preter, K. & Callewaert, N. A versatile method for circulating cell-free DNA methylome profiling by reduced representation bisulfite sequencing. bioRxiv, 663195. https://doi.org/10.1101/663195 (2019).
Van Paemel, R. et al. Minimally invasive classification of paediatric solid tumours using reduced representation bisulphite sequencing of cell-free DNA: a proof-of-principle study. Epigenetics 16, 196–208. https://doi.org/10.1080/15592294.2020.1790950 (2021).
De Wilde, J. et al. A fast, affordable and minimally-invasive diagnostic test for Cancer of Unknown Primary (CUP) using DNA methylation profiling. Lab. Investig., 102091 (2024).
Ococks, E. et al. Longitudinal tracking of 97 esophageal adenocarcinomas using liquid biopsy sampling. Ann. Oncol. 32, 522–532. https://doi.org/10.1016/j.annonc.2020.12.010 (2021).
Cornelli, L. et al. Diagnosis of pediatric central nervous system tumors using methylation profiling of cfDNA from cerebrospinal fluid. Clin. Epigenet. 16, 87. https://doi.org/10.1186/s13148-024-01696-w (2024).
Azad, T. D. et al. Circulating tumor DNA analysis for detection of minimal residual disease after chemoradiotherapy for localized esophageal cancer. Gastroenterology 158, 494–505 e496. https://doi.org/10.1053/j.gastro.2019.10.039 (2020).
Maron, S. B. et al. Circulating tumor DNA sequencing analysis of gastroesophageal adenocarcinoma. Clin. Cancer Res. 25, 7098–7112. https://doi.org/10.1158/1078-0432.CCR-19-1704 (2019).
Henriksen, T. V. et al. The effect of surgical trauma on circulating free DNA levels in cancer patients-implications for studies of circulating tumor DNA. Mol. Oncol. 14, 1670–1679. https://doi.org/10.1002/1878-0261.12729 (2020).
The Cancer Genome Atlas Research Network. Integrated genomic characterization of oesophageal carcinoma. Nature 541, 169–175. https://doi.org/10.1038/nature20805 (2017).
Anusewicz, D., Orzechowska, M. & Bednarek, A. K. Notch signaling pathway in cancer-review with bioinformatic analysis. Cancers (Basel) 13. https://doi.org/10.3390/cancers13040768 (2021).
Wendt, J., Rosenbaum, H., Richmond, T. A., Jeddeloh, J. A. & Burgess, D. L. Targeted bisulfite sequencing using the SeqCap epi enrichment system. Methods Mol. Biol. 1708, 383–405. https://doi.org/10.1007/978-1-4939-7481-8_20 (2018).
Passemiers, A. et al. MetDecode: methylation-based deconvolution of cell-free DNA for noninvasive multi-cancer typing. Bioinformatics 40. https://doi.org/10.1093/bioinformatics/btae522 (2024).
Moss, J. et al. Comprehensive human cell-type methylation atlas reveals origins of circulating cell-free DNA in health and disease. Nat. Commun. 9, 5068. https://doi.org/10.1038/s41467-018-07466-6 (2018).
R: A language and environment for statistical computing (R Foundation for Statistical Computing, 2023).
Acknowledgements
The authors would like to thank the study nurses P. Horckmans and I. Vandenbroucke from the UZ Ghent Department of Gastro-Intestinal Surgery for helping with the patient sample collection.
Funding
This work was supported by Kom op tegen Kanker [Stand up to Cancer, the Flemish cancer society; 2022/13037 to KS]; Ghent University Special Research Fund [BOF; 2–4 year research project and starting grant awarded to KDP]; the Fund for Scientific Research Flanders [FWO; 11H7523N to PD, 1S90621N to JDW].
Author information
Authors and Affiliations
Contributions
JV, KDP and KS contributed to the conceptualization; AP, HV, JDW, KS and PD contributed to data curation; formal analysis was done by JDW, KS, MFDS, SVdV, LM and OT; JV, KDP and KS performed funding acquisition; investigation was done by JDW, KS, MFDS, SR and PD; methodology was performed by JV, KDP and KS; project administration was done by KS; EVD, HV, JV, KDP, LD, PN and PP contributed to resources; JDW, KS, MFDS and SVdV contributed to software; supervision was performed by JV, KDP and PP; visualization was performed by KS; KS performed the writing—original draft; all authors contributed to reviewing and editing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics
Written informed consent was obtained from all cancer patients and healthy donors. Sample collection was approved by the ethics committee of Ghent University Hospital (registration numbers B670201628317, B670201628319 and B670201733701). The research was conducted according to the local legislation and institutional requirements.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Schoofs, K., Ferro Dos Santos, M., De Wilde, J. et al. Therapy response monitoring in blood plasma from esophageal adenocarcinoma patients using cell-free DNA methylation profiling. Sci Rep 14, 31112 (2024). https://doi.org/10.1038/s41598-024-82325-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-82325-7