Abstract
To develop a deep learning (DL) model based on MRI to predict muscle-invasive bladder cancer (MIBC). A total of 559 patients, including 521 patients in our center and 38 patients in external centers were collected from 2012 to 2023 to construct the DL model. In this study, the DL model was utilized to differentiate between MIBC and NMIBC based on three-channel image inputs, including original T2WI images, segmented bladder, and regions of interest. Inception V3 was employed for model construction. The accuracy, sensitivity (SN), specificity (SP), positive predictive value (PPV) and negative predictive value (NPV) for predicting MIBC by DL model were 92.4%, 94.7%, 91.5%, 81.8% and 97.7% in the validation set and 92.1%, 86.8%, 94.6%, 88.5% and 93.8% in the internal test set. In the external test set, these values were 81.6%, 57.1%, 87.1%, 50.0% and 90.0%. Additionally, the accuracy, SN, SP, PPV, and NPV for predicting MIBC were 93.5%, 100%, 93.4%, 11.1%, and 100% in VI-RADS 2; 80.0%, 66.7%, 87.2%, 73.7% and 82.9% in VI-RADS 3; 90.3%, 91.7%, 85.7%, 95.7%, 75.0% in VI-RADS 4. The accuracy, SN, and PPV were 93.9%, 93.9%, and 100% in VI-RADS 5. The DL model based on T2WI can effectively predict MIBC and serve as a valuable complement to VI-RADS 3.
Similar content being viewed by others
Introduction
Bladder cancer (BCa) is the 10th most commonly diagnosed cancer worldwide, with approximately 573,000 new cases and 213,000 deaths in 20201. Treatment options primarily depend on the stage of the disease at diagnosis. In addition, approximately 25% of patients are initially diagnosed with muscle-invasive bladder cancer (MIBC), while the remaining cases are classified as non-muscle-invasive bladder cancer (NMIBC)2. Radical cystectomy (RC) is the standard treatment for MIBC and certain high-risk or very high-risk cases of NMIBC3. Research has shown that 9–49% of BCa cases were incorrectly staged, leading to some patients receiving inappropriate treatment4. Importantly, accurately identifying muscle-invasive states is essential for the effective treatment and management of patients.
Currently, cystoscopy, computed tomography, and magnetic resonance imaging (MRI) play a significant role in the preoperative diagnosis of bladder cancer (BCa). A recent study indicated that it is possible to refer potential MIBC patients from cystoscopy to multi-parametric MRI (mpMRI) for staging instead of performing transurethral resection of bladder tumor (TURBT)5. MRI offers superior anatomical visualization with high spatial and contrast resolution, demonstrating its vast potential for determining muscle-invasive states. Recently, various diagnostic methods based on mpMRI have emerged. In 2018, Panebianco et al. proposed a vesical imaging-reporting and data system (VI-RADS), which utilizes the size, location, number, and morphology of tumors on mpMRI to predict the risk of muscle invasion6.
Artificial intelligence (AI)-based research methods are becoming increasingly prevalent in the medical field. Among these methods, machine learning is being leveraged to enhance the diagnosis and treatment of various diseases. Commonly applied machine learning methods include deep learning (DL), support vector machines, random forest, kernel methods, etc7. DL offers significant advantages in diagnosing various diseases, such as liver fibrosis, pancreatic cancer, pulmonary nodules, prostate cancer, etc8,9,10. Moreover, extensive research has focused on predicting drug response and assisting in radiotherapy11,12,13. In urinary, the preoperative prediction of MIBC has attracted considerable interest. The potential of DL to enhance diagnosis using cystoscopic and CT images has been explored14,15. Li et al. established a DL model based on MRI and compared it to the VI-RADS score, however, it lacked validation from a large sample and multiple centers16. Thus, this study aims to develop the DL model based on MRI to predict MIBC and conduct multicenter validation.
Materials and methods
Patient population
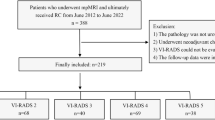
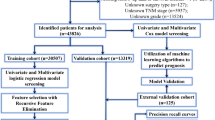
The study was approved by the Ethics Committees (2021-SR-409) and registered with ClinicalTrials.gov (NCT05096533). As shown in Fig. 1, patients who underwent preoperative MRI and surgery were enrolled between February 2012 and December 2023. Additionally, all written informed consent to participate in the study was obtained from the patient. The inclusion criteria and exclusion criteria were shown in the supplementary material.
The workflow of population screening process and model design process in multi-center research.
Lesion assessment and preprocessing
All pathological specimens were obtained from RC or TURBT. Two senior pathologists, who were unaware of the clinical data, reviewed all specimens. All specimens from NMIBC patients were confirmed to contain bladder muscle. MRI examinations of the bladder were conducted with patients in the supine position. Before the MRI, patients were instructed to void 2 h before the examination and to limit fluid intake or further urination until the completion of the MRI. Detailed parameters of MRI were provided in Table S1.
To construct the models, radiologists selected slides containing tumors for evaluation. For patients with NMIBC, slides containing tumors were all selected for the training set as a label for non-muscle-invasive. For patients with MIBC, all slides containing tumors were independently defined as muscle-infiltrating or non-muscle-infiltrating by two radiologists. A third senior radiologist with 20 years of experience, reviewed any disputed slides to make a final decision.
Besides, the radiologists evaluated the VI-RADS for all cases. All evaluation procedures were independently performed by two experienced radiologists. A third radiologist reviewed any controversial results before the final conclusions were made. All radiologists were blind to the patient’s pathology results. Furthermore, the diagnostic performance of the VI-RADS scores for MIBC was assessed using the AUC with cut-off scores of greater than 2 or 3. Then the images were resized and histograms equalized and normalized.
MIBC prediction model based on 2D- inception V3
As shown in Fig. 1b, automatic segmentation was performed by the Cascade Path Augmentation Unet (CPA-Unet) developed by our center17. The radiologists subsequently re-examined the segmented images. Based on the segmented T2WI images, the DL model was comprised of Inception V3, reconstruction blocks, and classification blocks for the classification of MIBC and NMIBC through an end-to-end approach. Inception V3 was utilized as the main model to extract image features from the T2WI slices18. Additionally, a reconstruction block was incorporated within a multitask framework to support the primary classification task (Fig. 1c). Detailed information regarding model construction is described in Supplementary Methods.
MRI images of each patient were input into the model slide by slide. The cut-off value was 0.5 (MIBC: probability value ≥ 0.5; NMIBC: probability value < 0.5). For a patient, if there were two or more slides’ probability values exceeding 0.5, it was MIBC. Otherwise, it was classified as NMIBC.
Statistical analysis
Statistical analysis was conducted using IBM SPSS statistics (v26.0). Evaluation metrics included sensitivity, specificity, accuracy, and the area under the curve (AUC) of the receiver operating characteristic (ROC). P < 0.05 was deemed statistically significant, with a 95% confidence level. The Chi-square and ANOVA tests were used to analyze the differences between groups. The interobserver agreement between the two radiologists for classifying the MRI slides and evaluating the VI-RADS was determined using kappa (k) scores. The sample size was calculated by PASS software (tests for one-sample sensitivity and specificity, version 15, NCSS, LLC). The experiments are performed on an NVIDIA 2080Ti with 12 GB of memory, and all networks are implemented in TensorFlow with Python version 3.7.
Results
The characteristics of the patients
The clinicopathologic characteristics across the groups are summarized in Table 1. A total of 559 BCa patients were included, with 521 patients from our center and 38 patients from external centers. There were no statistically significant differences among groups regarding clinical characteristics. Based on the sensitivity and specificity of the DL model in the validation set, the required sample size calculated by PASS software was at least 135. Finally, a total of 164 patients in the internal test set and 38 patients in the external test set were included for testing.
The inter-reader agreement for slides evaluation and VI-RADS
As shown in Table S2, an excellent diagnostic agreement on slides evaluation between the two readers was observed (k = 0.869, P < 0.001). Table S3 demonstrated robust diagnostic agreement between the two readers in VI-RADS scores in the validation test set (k = 0.927, P < 0.001), the internal test set (k = 0.922, P < 0.001), and the external test set (k = 0.831, P < 0.001).
The performance metrics of the DL model and VI-RADS
The performance of the DL model was shown in Table 2 and Fig. 2a. The accuracy, sensitivity (SN), specificity (SP), positive predictive value (PPV), and negative predictive value (NPV) for MIBC were 92.4% (61/66), 94.7% (18/19), 91.5% (43/47), 81.8% (18/22) and 97.7% (43/44) in the validation set. Meanwhile, the accuracy, SN, SP, PPV, and NPV were 92.1% (151/164), 86.8% (46/53), 94.6% (105/111), 88.5% (46/52) and 93.8% (105/112) in the internal test set. In the external test set, these values were 81.6% (31/38), 57.1% (4/7), 87.1% (27/31), 50.0% (4/8) and 90.0% (27/30).
The DL model results and typital cases. (a) The predictive performance of deep learning model across various sets (validation set, internal test set and external test set). (b) The predictive performance of deep learning model in different VI-RADS. (c–f) The predictive performance of deep learning model, VI-RADS and combined model across various sets. (g) The performance metrics of the DL model in different bladder anatomical locations. (h) The process of determining the muscle infiltration status of patients based on deep learning model.
To further explore the differences between the DL models and VI-RADS, we compared the performance metrics of the mode with VI-RADS (cut-off VI-RADS > 2 or 3). The performance metrics are summarized in Table 2 and Fig. 2c–f. There were no statistically significant differences in the AUCs between the DL model and VI-RADS in the validation set (0.931 vs. 0.976, P = 0.172) and the internal test set (0.907 vs. 0.930, P = 0.427). Besides, the combined model based on the DL model and VI-RADS score demonstrated better performance than a single model in validation and internal test sets (P < 0.05). However, in the external test set, the combined model did not show an improvement in prediction performance for VI-RADS (P = 0.224).
The performance metrics of the DL model among VI-RADS
As shown in Table 3 and Fig. 2b, the accuracy of the DL model was 100% (21/21) in VI-RADS 1. Meanwhile, the accuracy, SN, SP, PPV and NPV were 93.5% (115/123), 100% (1/1), 93.4% (114/122), 11.1% (1/9) and 100% (114/114) in VI-RADS 2; 80.0% (48/60), 66.7% (14/21), 87.2% (34/39), 73.7% (14/19) and 82.9% (34/41) in VI-RADS 3; 90.3% (28/31), 91.7% (22/24), 85.7% (6/7), 95.7% (22/23), 75.0% (6/8) in VI-RADS 4. The accuracy, SN, and PPV were 93.9% (31/33), 93.9% (31/33), and 100% (31/31) in VI-RADS 5.
The performance metrics of the DL model in different bladder anatomical locations
As shown in Table S4 and Fig. 2g, the accuracy, SN, SP were 63.2 (12/19), 71.4 (5/7), 58.3 (7/12) in the bladder neck and trigone; 92.0 (138/150), 85.4 (35/41), 94.5 (103/109) in left and right wall; 88.2 (15/17), 81.8 (9/11), 100.0 (6/6) in dome; 94.6 (35/37)80.0 (4/5), 96.9 (31/32) in posterior wall; 93.3 (14/15), 100.0 (6/6), 88.9 (8/9) in anterior wall and 95.2 (20/21), 100.0 (4/4), 94.1 (16/17) in multiple bladder walls. Additionally, in patients near the ureteral orifice, the accuracy, SN, SP, PPV, and NPV of the DL model in predicting MIBC were 71.4 (35/49), 75.0 (15/20), 69.0 (20/29), 62.5 (15/24), 80.0 (20/25).
Analysis of typical cases
The process of determining the muscle infiltration status of patients based on a DL model is illustrated in Fig. 2h. The six MRI slides of the patient containing tumors were input into the model. After classification recognition, the predicted probability values for each slide were obtained. All six slides of this patient were predicted as MIBC (probability values > 0.5), consistent with the final pathological results.
Additionally, misclassified cases by DL model were further analyzed, as shown in Table S5 and Fig. S1. The results indicated that the most common characteristics in misclassified cases were tumors located near the ureteral orifice (14/25) and tumors located on the dome or bladder neck (8/25).
Discussion
Currently, numerous studies focus on muscle-invasive states using imaging approaches, including VI-RADS score based on MRI, predictive models based on radiomic features, and DL models based on CT6,15,19. Considering the excellent anatomical visualization provided by MRI and the advanced artificial intelligence algorithms, we developed MRI-based DL models for MIBC diagnosis and conducted a multi-center clinical study.
Recently, the medical applications of radiomics based on machine learning have advanced significantly. By extracting radiomic features of the region of interest (ROI), a series of clinical prediction models with AUCs ranging from 0.798 to 0.986 have been developed19. However, most predictive models were created after manually outlining the ROI, extracting and analyzing the features, followed by the extraction and analysis of features. This process is time-consuming and considerably diminishes diagnostic utility11. The DL algorithm can construct models by automatically segmenting the ROI and extracting optimal features intelligently and objectively11. Several studies15,20 explored the value of DL models in BCa with the AUCs ranging from 0.861 to 0.998, but all were retrospective CT-based studies. Furthermore, many studies suggested that MRI has greater potential in diagnosing the muscle-invasive states of BCa6,21. Building upon T2WI of MRI, we developed a DL model and executed a multi-center clinical study, achieving impressive results with an accuracy rate exceeding 90% at our institution.
VI-RADS proposes standardized reporting criteria based on mp-MRI, which has been validated and improved at many centers22. However, the score only indicates the probability of MIBC, and 4.4–25.7% patients classified as VI-RADS 3 were unable to differentiate muscle-invasive states23,24,25,26. Additional analysis was conducted to compare the difference between the DL model and the VI-RADS. Remarkably, the performance of our DL model was found to be comparable to that of VI-RADS in both the validation and internal test sets (P > 0.05). Interestingly, the combined model of the DL model and VI-RADS score demonstrated better performance than a single model in the validation and internal test sets. The preprocessing of multicenter medical imaging data remains an open challenge27. In our research, the performance of the DL model in the external test set was not ideal, showing results similar to those of other studies15,28.
We further explored the performance metrics of the DL model under various VI-RADS categories. Promisingly, the models achieved satisfactory performance metrics (accuracy > 90%) in VI-RADS 2, 4, and 5. In VI-RADS 3, the muscle-invasive states of BCa are ambiguous, posing a challenge for each center. Research has suggested that the characteristics of tumor morphology can serve as an effective tool for the preoperative detection of muscle invasion in VI–RADS 327. However, the evaluation process was inherently subjective. For the first time, we elaborated on the performance metrics of the AI model in VI-RADS 3. Notably, our research achieved an accuracy of 80% with the DL model, underscoring the potential of AI as a complementary tool in the diagnostic workflow for patients with VI-RADS 3.
Several critical factors still affect the accuracy of the model. Firstly, non-muscle-invasive tumors near the ureteric orifice were mostly misclassified as muscle-invasive tumors due to their encroachment into the ureteral orifice. Secondly, due to the limitations of 2D transverse images, non-muscle-invasive tumors located at the bladder neck and parietal wall were more likely to be misclassified as muscle-invasive tumors. Ongoing research aims to develop a 3D deep learning model, which could reduce the impact of numerous variables, such as tumor location and human factors. Finally, image quality also affects performance metrics.
However, several limitations were identified in this research. The “black box” nature of DL dictated that its decision-making process is unexplainable. Therefore, illustrating DL principles through visualization techniques is a promising direction for future research. Secondly, the external centers incorporated a limited number of patients and exhibited disparities in the distribution of clinicopathologic characteristics compared to our center, introducing bias in the performance evaluation of the DL model at the external set. Additionally, the MRI imaging quality at external centers deviated from ours, and the DL model was trained exclusively on cases sourced from our center, consequently influencing the performance metrics in the external test set. To tackle these challenges, future research endeavors should prioritize enhancing the diversity of the training dataset, potentially through the application of data augmentation techniques and the adoption of multi-center training strategies. Moreover, establishing a dedicated data pre-processing pipeline to address the discrepancies in multicenter data is essential27.
While most previous imaging-based studies were retrospective and single-center, we developed DL models based on MRI to diagnose MIBC and carried out multi-center clinical research. Our findings indicate that the DL model can accurately determine MIBC and provide valuable information for clinical decision-making.
Conclusion
The DL model based on T2WI and Inception V3 was constructed for predicting MIBC. The research results indicated that the DL model had certain advantages in predicting MIBC and providing additional diagnostic value in the context of VI-RADS 3.
Data availability
The datasets used and analysed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- AUC:
-
The area under the curve of ROC
- BCa:
-
Bladder cancer
- CPA-Unet:
-
Cascade Path Augmentation Unet
- DL:
-
Deep learning
- MIBC:
-
Muscle-invasive bladder cancer
- MRI:
-
Magnetic resonance imaging
- Mp-MRI:
-
Multi-parametric MRI
- NMIBC:
-
Non-muscle-invasive bladder cancer
- NPV:
-
Negative predictive value
- PPV:
-
Positive predictive value
- RC:
-
Radical cystectomy
- ROC:
-
Receiver operating characteristic
- SN:
-
Sensitivity
- SP:
-
Specificity
- T2WI:
-
T2 weighted images
- TURBT:
-
Transurethral resection of bladder tumor
- VI-RADS:
-
Vesical imaging-reporting and data system
References
Sung, H. et al. Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249. https://doi.org/10.3322/caac.21660 (2021).
Witjes, J. A. et al. European association of urology guidelines on muscle-invasive and metastatic bladder cancer: Summary of the 2020 guidelines. Eur. Urol. 79, 82–104. https://doi.org/10.1016/j.eururo.2020.03.055 (2021).
D’Agostino, D. et al. Postoperative quality of life in patients with ileal neobladder at short-, intermediate- and long-term follow-up. Urol. Int. 97, 54–60. https://doi.org/10.1159/000443185 (2016).
Xu, X. et al. Study progress of noninvasive imaging and radiomics for decoding the phenotypes and recurrence risk of bladder cancer. Front. Oncol. 11, 704039. https://doi.org/10.3389/fonc.2021.704039 (2021).
Bryan, R. T. et al. Comparing an imaging-guided pathway with the standard pathway for staging muscle-invasive bladder cancer: Preliminary data from the bladderpath study. Eur. Urol. 80, 12–15. https://doi.org/10.1016/j.eururo.2021.02.021 (2021).
Panebianco, V. et al. Multiparametric magnetic resonance imaging for bladder cancer: Development of VI-RADS (vesical imaging-reporting and data system). Eur. Urol. 74, 294–306. https://doi.org/10.1016/j.eururo.2018.04.029 (2018).
Wang, P. P., Deng, C. L. & Wu, B. Magnetic resonance imaging-based artificial intelligence model in rectal cancer. World J. Gastroenterol. 27, 2122–2130. https://doi.org/10.3748/wjg.v27.i18.2122 (2021).
Vente, C., Vos, P., Hosseinzadeh, M., Pluim, J. & Veta, M. Deep learning regression for prostate cancer detection and grading in bi-parametric MRI. IEEE Trans. Biomed. Eng. 68, 374–383. https://doi.org/10.1109/TBME.2020.2993528 (2021).
Ardila, D. et al. Author correction: End-to-end lung cancer screening with three-dimensional deep learning on low-dose chest computed tomography. Nat. Med. 25, 1319. https://doi.org/10.1038/s41591-019-0536-x (2019).
Liu, K. L. et al. Deep learning to distinguish pancreatic cancer tissue from non-cancerous pancreatic tissue: a retrospective study with cross-racial external validation. Lancet Digit Health 2, e303–e313. https://doi.org/10.1016/S2589-7500(20)30078-9 (2020).
Skrede, O. J. et al. Deep learning for prediction of colorectal cancer outcome: a discovery and validation study. Lancet 395, 350–360. https://doi.org/10.1016/S0140-6736(19)32998-8 (2020).
Tran, K. A. et al. Deep learning in cancer diagnosis, prognosis and treatment selection. Genome Med. 13, 152. https://doi.org/10.1186/s13073-021-00968-x (2021).
Min, H. et al. Automatic radiotherapy delineation quality assurance on prostate MRI with deep learning in a multicentre clinical trial. Phys. Med. Biol. https://doi.org/10.1088/1361-6560/ac25d5 (2021).
Shkolyar, E. et al. Augmented bladder tumor detection using deep learning. Eur. Urol. 76, 714–718. https://doi.org/10.1016/j.eururo.2019.08.032 (2019).
Zhang, G. et al. Deep learning on enhanced ct images can predict the muscular invasiveness of bladder cancer. Front. Oncol. 11, 65468510 (2021).
Li, J. et al. Predicting muscle invasion in bladder cancer by deep learning analysis of MRI: comparison with vesical imaging-reporting and data system. Eur. Radiol. 33, 2699–2709. https://doi.org/10.1007/s00330-022-09272-7 (2023).
Yu, J. et al. Cascade path augmentation unet for bladder cancer segmentation in MRI. Med. Phys. 49, 4622–4631. https://doi.org/10.1002/mp.15646 (2022).
C Szegedy, VVS Ioffe, J Shlens, Z Wojna. Rethinking the Inception Architecture for Computer Vision. 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Las Vegas, NV, USA. 2818–2826. https://doi.org/10.1109/CVPR.2016.308. (2016).
Kozikowski, M. et al. Role of radiomics in the prediction of muscle-invasive bladder cancer: A systematic review and meta-analysis. Eur. Urol. Focus 8, 728–738. https://doi.org/10.1016/j.euf.2021.05.005 (2022).
Yang, Y., Zou, X., Wang, Y. & Ma, X. Application of deep learning as a noninvasive tool to differentiate muscle-invasive bladder cancer and non-muscle-invasive bladder cancer with CT. Eur. J. Radiol. 139, 109666. https://doi.org/10.1016/j.ejrad.2021.109666 (2021).
Wang, H. et al. Elaboration of a multisequence MRI-based radiomics signature for the preoperative prediction of the muscle-invasive status of bladder cancer: a double-center study. Eur. Radiol. 30, 4816–4827. https://doi.org/10.1007/s00330-020-06796-8 (2020).
Yuan, B. et al. Role of vesical imaging-reporting and data system in predicting muscle-invasive bladder cancer: A diagnostic meta-analysis. Int. J. Urol. 29, 186–195. https://doi.org/10.1111/iju.14748 (2022).
Marchioni, M. et al. Could bladder multiparametric MRI be introduced in routine clinical practice? Role of the new VI-RADS score: Results from a prospective study. Clin. Genitourin. Cancer 18, 409–415. https://doi.org/10.1016/j.clgc.2020.03.002 (2020).
Wang, Z. et al. Evaluation of the value of the VI-RADS scoring system in assessing muscle infiltration by bladder cancer. Cancer Imaging 20, 26. https://doi.org/10.1186/s40644-020-00304-3 (2020).
Wang, H. et al. Multiparametric MRI for bladder cancer: Validation of VI-RADS for the detection of detrusor muscle invasion. Radiology 291, 668–674. https://doi.org/10.1148/radiol.2019182506 (2019).
Liu, S. et al. Evaluation of vesical imaging-reporting and data system (VI-RADS) scoring system in predicting muscle invasion of bladder cancer. Transl. Androl. Urol. 9, 445–451. https://doi.org/10.21037/tau.2020.02.16 (2020).
Cackowski, S., Barbier, E. L., Dojat, M. & Christen, T. ImUnity: A generalizable VAE-GAN solution for multicenter MR image harmonization. Med. Image Anal. 88, 102799. https://doi.org/10.1016/j.media.2023.102799 (2023).
Wichtmann, B. D., Albert, S., Zhao, W., Maurer, A., Rödel, C., Hofheinz, R. D., Hesser, J., Zöllner, F. G., & Attenberger, U. I. Are We There Yet? The Value of Deep Learning in a Multicenter Setting for ResponsePrediction of Locally Advanced Rectal Cancer to Neoadjuvant Chemoradiotherapy. Diagnostics (Basel, Switzerland), 12(7), 1601. https://doi.org/10.3390/diagnostics12071601. (2022).
Acknowledgements
The authors thank the patients and their families for agreeing to provide personal information.
Funding
This work was funded by National Natural Science Foundation of China, 82072832, 82273152, Jiangsu Province Hospital (the First Affiliated Hospital of Nanjing Medical University) Clinical Capacity Enhancement Project, JSPH-MC-2021–15, JSPH-MA-2022–5, Jiangsu Province Capability Improvement Project through Science, Technology and Education, NO.ZDXK202219.
Author information
Authors and Affiliations
Contributions
Cai, L: conceptualization, formal analysis, writing—original draft, visualization. Yang, X: visualization, writing—original draft. Yu, J: methodology, validation, software. Shao, Q: resources. Wang, G: resources. Yuan, B: data curation, investigation. Zhuang, J: investigation, Visualization. Li, K: investigation, visualization. Wu, Q: data curation, investigation. Liu, P: data curation, investigation. Yu, R: data curation, investigation. Cao, Q: data curation. Li, P: data curation. Sun, X: formal analysis, resources. Zou, Y: software, visualization. Fu, X: software. Fang, X: supervision, writing—review & editing. Chen, C: supervision, project administration. Lu, Q: supervision, funding acquisition, writing—review & editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
All procedures performed in studies involving human participants were in accordance with the ethical standards of the Research Ethics Committee (2021-SR-409) and with the 1964 Helsinki Declaration and its later amendments. All written informed consent to participate in the study was obtained from the patient.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Cai, L., Yang, X., Yu, J. et al. Deep learning on T2WI to predict the muscle-invasive bladder cancer: a multi-center clinical study. Sci Rep 15, 9942 (2025). https://doi.org/10.1038/s41598-024-82909-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-82909-3