Abstract
Loss of function in the chromatin remodeler CHD7 causes CHARGE syndrome, characterized by variable penetrance and diverse abnormalities. However, establishing genotype-phenotype correlations has been challenging, as most CHD7 inactivating mutations are null alleles. Through CHD7 missense variant analysis at potential phosphorylation sites, we identified T730 (T720 in mice) as a critical residue associated with pathogenesis. Using a CHD7 T730 missense variant (Chd7T720A) and a frameshift null allele (Chd7fs) in a mouse model, we found that Chd7fs/fs mice were non-viable, while Chd7fs/+ mice exhibited haploinsufficiency-related circling behavior. Notably, Chd7fs/T720A mice died before postnatal day 2, indicating the Chd7T720A allele is hypomorphic. Micro-CT analysis at E18.5 revealed that heterozygous mice primarily exhibited hypertrophic cardiomyopathy (HCM), while homozygous mice developed both HCM and dilated cardiomyopathy (DCM). RNA-seq analysis of neonatal Chd7T720A/T720A hearts revealed a disrupted transcriptome, which in males and females was characterized by downregulation of mitochondrial energy metabolism genes and enrichment of ETS family transcription factor targets. We further identified GSK3β, GSK3α, HIPK1, and DYRK2 as candidate kinases for this site, suggesting a regulatory role in CHD7. This missense variant causing developmental heart abnormalities establishes the first genotype-phenotype correlation for CHD7, and offers new insights into CHARGE syndrome pathogenesis.
Similar content being viewed by others
Introduction
Modulation of chromatin structure by ATP-dependent remodeling factors is central to the regulation of transcription and, consequently, key biological processes. These enzymes alter chromatin organization, in part, by moving or evicting nucleosomal histones that considerably restrict DNA accessibility1. There are 4 major families of chromatin remodeling factors: the Switch/Sucrose Non-Fermenting (SWI/SNF), Imitation Switch (ISWI), Inositol 80 - Swi2/Snf2-Related 1 (INO80-SWR1) and Chromodomain Helicase DNA-binding (CHD) families. This classification is based on protein domains associated with their central catalytic domain, which is homologous to the yeast SNF2 enzyme2. For instance, SWI/SNF-family ATPases harbor bromodomains while CHD-family enzymes are characterized by their chromodomains. CHD proteins form the largest family of remodeling enzymes and all 9 members are involved in vertebrate development3. CHD7 (Chromodomain Helicase DNA-binding Protein 7) is a key chromatin remodeler involved in developmental gene expression, and its loss-of-function mutations cause CHARGE syndrome, a multisystem disorder characterized by Coloboma of the eye, Heart defects, Atresia of the choanae, Retardation of growth, Genital and Ear abnormalities4.
Despite extensive research on CHD7 mutations, most studies have focused on null or frameshift mutations that lead to complete CHD7 function loss5,6,7,8. While sequencing of CHD7 is used as a molecular diagnostic for CHARGE9, how different pathogenic variants affect CHD7 function is largely unknown. For instance, truncating/stop codon variants in CHD7 are typically found in individuals with CHARGE, while individuals with IHH predominantly have missense variants10. CHD7 variants that do not generate null alleles may even be classified as ‘benign’ or ‘variants of uncertain significance’ despite producing amino acid substitutions at conserved CHD7 residues and the presence of CHARGE phenotypes. The inability to ascertain whether CHD7 missense mutations are benign or pathogenic is further compounded by their distribution along the entire gene, making it difficult to determine whether they affect a specific or multiple CHD7 biochemical functions9.
To investigate genotype-phenotype relationships for CHD7, we focused on a highly conserved phosphorylatable residue, threonine 730 (T730; T720 in mice), and generated a missense variant allele (Chd7T720A) to examine its role in heart development. Notably, mice carrying the Chd7T720A allele had distinct cardiomyopathies, linking this missense variant to defects in heart development. Furthermore, in vitro kinase assays identified GSK3β, GSK3α, HIPK1, and DYRK2 as kinases for T730 suggesting this may be a phosphorylation site on CHD7, and RNA-seq revealed transcriptional changes in the heart of animals carrying the T720A missense variant linked to mitochondrial dysfunction and ETS family transcription factor dysregulation. These in vivo and in vitro findings establish the first genotype-phenotype correlation for a CHD7 missense variant, and lay the groundwork for exploring upstream regulatory pathways governing CHD7 function, with potential implications for CHARGE syndrome and related cardiomyopathies.
Results
CHD7 threonine 730 (T730): a conserved and clinically relevant site on CHD7
To explore genotype-phenotype correlations and dissect how loss of specific CHD7 functions contribute to disease pathogenesis, we focused on potential phosphorylation sites, reasoning that these could potentially regulate specific CHD7 functions. We conducted an extensive review of previously identified phosphorylation sites on CHD7 involving collation of all documented phosphorylation sites from the PhosphoSitePlus database (https://www.phosphosite.org/)11. These phosphorylation sites were then correlated with clinical data for missense variants listed in the single nucleotide polymorphism database (dbSNP) at NextProt (https://www.nextprot.org/entry/NX_Q9P2D1/sequence)12. A total of eighteen clinically significant missense variants were identified at phosphorylatable serine (S), threonine (T) or tyrosine (Y) residues: S734P, T730I, T2472A, S2229P, S2395P or Y, T2952S, T298A, Y313C, S316I, S559L or W, S612N or T, T689M, S782C, T786I, Y1217D, S2110G, S2231C or F, T2532M. Clinical relevance of these variants ranged from associations with CHARGE syndrome to manifestations in hypogonadotropic hypogonadism, neurodevelopmental disorders, and various neoplasms, as detailed in Supplementary Table 1. Among these sites, T730 emerged as a key site of interest, as a T730 missense variant (T730I c.2189 C > T), cataloged as rs552946889 in reference SNP reports with an allele frequency of 0.00476% (38 cases in 797927 samples), was associated with both Kallmann syndrome and CHARGE syndrome (ClinVar; [VCV000363451.13]). In gnomAD, CHD7-T730A and CHD7-T730I have been reported with allele counts of 1 and 57, respectively. While CHD7-T730I has been observed in individuals with hypospadias and cryptorchidism, its clinical significance remains uncertain, as it has been classified as ‘uncertain significance’ or ‘likely benign’ in ClinVar (Variation ID: 363451). Given the variable penetrance observed in CHARGE syndrome, changes in cell signaling, perhaps in response to environmental variation, could impact CHD7 activity during key periods of development and contribute to disease pathogenesis. Given that cardiac defects are a major feature of CHARGE syndrome, understanding how phosphorylation affects CHD7 activity provides valuable insights into the molecular mechanisms underlying CHD7-related congenital heart disease. T730 is evolutionarily conserved from Drosophila(Kismet) to humans, underscoring its biological significance. Phosphorylation at this site (pT730) has been observed in cell lines13,14,15 and mouse tissues (Guo A, CST Curation Set 2923 and 2717; Year 2007, https://www.phosphosite.org/curatedInfoAction.action?record=3366534). Consequently, we focused on T730 for further exploration into regulatory pathways that could link specific genetic alterations to the multifaceted phenotypes of CHARGE syndrome and, more broadly, CHD7-related disorders.
Pathogenic effects of a CHD7 T730 missense variant in a murine model
To establish the functional importance of T730, we generated mice carrying a murine mT720A (hT730A equivalent) Chd7 allele variant (Chd7T720A) using a Chd7 cDNA (NM_001277149.1) and CRISPR-Cas9 (Fig. 1a and Supplementary Fig. 1a). T730I has been identified as a clinically relevant CHD7 variant, but its functional consequences remain uncharacterized. Our goal is to investigate the broader functional effects of phosphorylation loss at this site, rather than solely replicating the human variant’s effects. T730 in CHD7 is a conserved threonine residue, and a predicted phosphorylation site, although a functional role for this residue is unknown. Indeed, T730 has been identified as clinically relevant, as a T730I mutation has been linked to CHD7 (ClinVar, Variation ID: 363451). To better understand the potential impact of loss of phosphorylation loss at this site, we generated variants substituting threonine with a non-phosphorylatable alanine (T730A) residue. Concurrently, we generated a null allele carrying a frameshift deletion (ΔC2718) that introduces a premature stop codon upstream of the CHD7 chromodomains, subsequently referred to as Chd7fs (Fig. 1b). Given that approximately 78% of individuals with CHARGE syndrome have either nonsense or frameshift mutations (44% and 34%, respectively)8, the Chd7fs allele was also characterized as a potential new murine CHARGE model.
Subsequent breeding experiments among the various genotypes carrying these alleles were conducted to evaluate offspring viability. While Chd7fs/+ mice with a wild-type (WT) allele were viable, no Chd7fs/fs homozygotes were obtained from crosses between heterozygous Chd7fs/+ x Chd7fs/+ mice, as expected for null alleles; four separate matings between heterozygous Chd7fs/+ mice, including one that was left undisturbed for four months, yielded no homozygous Chd7fs/fs offspring. Chd7fs/+ mice carrying one null allele exhibited characteristic circling behavior indicative of inner ear defects due to Chd7 haploinsufficiency16, as well as small eyes (microphthalmia) and, in one case, a round head consistent with hydrocephalus (Table 1). In contrast, crosses between heterozygous Chd7T720A/+ mice produced offspring in Mendelian ratios (Fig. 1c), including Chd7T720A/T720A homozygotes. These mice did not display circling behavior, nor grossly observable eye defects, suggesting phosphorylation at T720 is not linked to pathogenesis of these defects.
Strikingly, however, Chd7fs/T720A progeny from Chd7T720A/T720A females crossed with Chd7fs/+ males all displayed either prenatal or immediate postnatal lethality, with none surviving beyond postnatal day 2 (Fig. 1c; Table 1), indicating the Chd7T720A allele was not able to fully substitute for WT Chd7. Of the live-born Chd7fs/T720A pups that survived long enough to be genotyped (but later died), 5 were males and only 1 was female. Although this observation suggests females may be more adversely affected than males, Fisher’s exact test showed that the deviation from the expected 1:1 ratio was not statistically significant (p = 0.55). While both Chd7fs/+ and Chd7fs/T720A mice exhibited eye defects such as microphthalmia, Chd7 mice displayed more severe phenotypes, including anophthalmia (Table 1 and Supplementary Fig. 1b).
Pathogenicity of the Chd7 T720A allele and its impact on cardiac development
To further investigate these findings, we conducted a phenotypic analysis using micro-computed tomography (micro-CT) of E18.5 mouse embryos, allowing us to examine Chd7T720A/+, Chd7T720A/T720A, Chd7fs/+, and Chd7fs/T720A mice before viability loss. This analysis included assessments of gross morphological abnormalities, such as dysmorphisms, growth delays, edema, and craniofacial anomalies, as well as anatomical defects in major organs. Notably, the most pronounced and consistently observed abnormalities were found in the brain, eyes, and heart (Fig. 2a and b). We observed highly penetrant eye defects, including unfused eyelids and microphthalmia, in Chd7fs/+ and Chd7 embryos, suggesting a strong association with the Chd7fs allele. In contrast, all Chd7T720A/+ and Chd7T720A/T720A embryos had normal eyes. In the brain, reduced lateral ventricle size was observed across most mutant genotypes. These findings corroborate previous research on CHD7 haploinsufficiency, with the Chd7fs allele phenotypes closely aligning with the Chd7COA1/+mouse model carrying the K719X nonsense mutation17.
We also observed highly penetrant heart abnormalities in both male and female mice carrying Chd7fs and Chd7T720A alleles (Fig. 2b). Micro-CT section analysis revealed embryos carrying these alleles exhibited both enlarged and reduced ventricular chambers compared to WT embryos. Embryos with enlarged ventricular chambers displayed thinning of the ventricular walls, characteristic of DCM. In contrast, embryos with reduced ventricular chambers showed thickened ventricular walls and septa, consistent with HCM. Additionally, two of the Chd7fs/+embryos (one male and one female) exhibited a ventricular septal defect (VSD), a hallmark in CHARGE syndrome and its mouse models16,18, characterized by a discontinuous gap in the ventricular septum. Interestingly, these embryos also showed enlarged ventricular chambers, consistent with a DCM-like phenotype. Given that CHD7 functions as a chromatin remodeler involved in cardiac development, its haploinsufficiency may disrupt transcriptional programs required for myocardial homeostasis. This could contribute to chamber dilation through mechanisms involving cardiomyocyte survival, extracellular matrix remodeling, or altered contractility. In more severe cases, progressive myocardial thinning may further impair ventricular septation, ultimately leading to VSD. The presence of VSD in some, but not all, Chd7fs/+ embryos suggests that while these defects may arise through overlapping mechanisms, they are not necessarily linked in all cases. Further studies will be necessary to elucidate the precise relationship between CHD7 function, myocardial development, and ventricular septation. Notably, this ventricular septal defect was not observed in mice carrying the Chd7T720A allele.
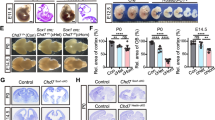
Murine T730 missense and frameshift-null alleles. (a) Alignment of mouse and human CHD7 proteins showing conservation between the human T730 and mouse T720 phosphorylation sites. (b) Schematic representation of the T720A point mutation and the frameshift mutation generated by CRISPR/Cas9-mediated technology. The frameshift null allele (Chd7fs) terminates at the N-terminal region due to the deletion of a cytosine at position 2718 in the Chd7 cDNA (NM_001277149.1). (c) Offspring produced by mating mice with Chd7T720A and/or Chd7fs alleles. The number of pups is shown along with the expected and observed percentages. Chd7T720A/fs mice died by postnatal day 2, and all pups from Chd7fs/+ mothers died after delivery.
Gross morphology and 2D digital section phenotyping with iodine-contrasted micro-CT. (a) Table summarizing the occurrence of organ abnormalities in Chd7 mutants across different genotypes: WT (Chd7+/+), T720A/+ (Chd7), T720A/T720A (Chd7T720A/T720A), frameshift null fs/+ (Chd7fs/+), and compound heterozygous fs/T720A (Chd7fs/T720A). The analysis was performed on mouse embryos at embryonic day 18.5 (E18.5) using micro-CT. Shaded areas in the table indicate the presence of abnormalities in various organs including the brain, eye, heart, and kidney. (b) Representative micro-CT images showing the phenotypic abnormalities observed in Chd7 mutants. The abnormalities are indicated by red arrows and are organized by organ: Brain (reduced lateral ventricular size in Chd7T720A/+ and Chd7fs/T720A mutants), eye (microphthalmia observed in two Chd7fs/+ mutants, heart (enlarged ventricular chamber, thinning of the ventricular wall, and ventricular septal defect in Chd7fs/+ mutant; or reduced ventricular chamber and thickened ventricular wall in Chd7fs/T720A mutant), and kidney (enlarged kidney in Chd7T720A/T720A mutant; or hydronephrosis in Chd7fs/T720A mutant). Scale bars for reference are included in the images (2.5 mm, 5 mm, and 2 mm).
Quantitative 3D analysis of cardiac morphology
To quantify the impact of the Chd7fs and Chd7T720A alleles on heart morphology, we developed a 3D segmentation method to measure the volumes of the left and right ventricular chambers and ventricular walls using micro-CT datasets (Fig. 3a). The volume of the ventricular chambers (left and right), normalized by the total ventricular volume (chambers + walls), was defined as the Volume of Chambers (VoC). A Z-score was computed for each sample, comparing it to WT average. A Z-score < −2 (2 standard deviations below the mean) with a Bonferroni-corrected p-value < 0.05 indicated a significantly smaller VoC than WT embryos, while a Z-score > 2 with p < 0.05 indicated a significantly larger VoC than WT embryos (Supplementary Table 2).
This 3D quantification method refined our cardiac phenotype characterization, differentiating between ventricular chamber volume changes and ventricular wall thickness variations. Four out of five Chd7fs/+ embryos exhibited altered VoC, with three showing enlarged VoC (DCM-like phenotype) and one showing reduced VoC (HCM-like phenotype) (Fig. 3, Supplementary Tables 2, and Supplementary Fig. 2). All Chd7T720A/+ embryos (n = 8) had significantly reduced VoC, consistent with an HCM-like phenotype. Among Chd7T720A/T720A embryos (n = 7), three exhibited enlarged VoC (DCM-like), three exhibited reduced VoC (HCM-like), and one was not significantly different from WT embryos. In Chd7fs/T720A embryos (n = 6), five showed reduced VoC (HCM-like phenotype), while one was not significantly different from WT embryos. These quantitative data in Fig. 3 directly support and validate the qualitative observations shown in Fig. 2, reinforcing the reliability of our phenotype assessment. Thus, 3D quantification confirmed the cardiomyopathy phenotypes identified in 2D sections.
Impact of the Chd7 T720A allele on the transcriptome in heart
To investigate the functional differences between WT and Chd7T720A alleles, we evaluated how the T730 missense variant affects CHD7’s activity as a transcriptional co-activator/co-repressor using RNA-seq at postnatal day 0 (P0). In Chd7T720A/T720A hearts, a total of 2012 differentially expressed genes (DEGs) (FDR < 0.01, ≥ 1.5-fold change) were identified in males (1264 up-regulated, 748 down-regulated) and 581 in females (489 up-regulated, 92 down-regulated) relative to age- and sex-matched controls. These changes were visulaized through heat maps, principal component analysis (PCA), and volcano plots (Fig. 4a and b, and Supplementary Fig. 3a). Interestingly, there was no overlap between DEGs that were down-regulated in male and female Chd7T720A/T720A hearts, while 179 DEGs were up-regulated in both sexes. This finding highlights an unexpected sex-specific difference, particularly given the young age of these animals and that both males and females presented with cardiomyopathy (Supplementary Fig. 3b).
To investigate the functional significance of these gene expression changes, we performed over-representation analysis (ORA) using the Gene Ontology: Biological Processes (GO: BP) compendium to identify pathways enriched in the disrupted transcriptomes of Chd7T720A/T720A mice. In males, the most significantly enriched pathways were related to cytoskeletal organization, cell adhesion, lipid metabolic process, and cell cycle (Fig. 4c and Supplementary Fig. 3c). In female Chd7T720A/T720A hearts, up-regulated DEGs were primarily associated with immune cell activation, while cell adhesion pathways were enriched in both sexes (Fig. 4c and Supplementary Fig. 3c). Figure 4c presents volcano plots highlighting the most significantly up- and down-regulated DEGs in males, as well as up-regulated DEGs in females within these pathways. Given that many cytoskeletal elements are involved in both cell division and cell adhesion, we examined the extent of overlap between these GO terms. As shown in Supplementary Fig. 3 d, only a quarter of the identified genes were shared across two or more of these pathways, suggesting that they reflect distinct gene sets involved in cell cycle, cytoskeletal organization, and cellular adhesion.
To identify common transcriptional changes in the neonatal hearts of both male and female Chd7A/T720A mice, we performed gene set enrichment analysis (GSEA) on the entire transcriptome of P0 hearts. As shown in the enrichment plots and bar charts in Fig. 4d and e, neonatal Chd7T720A/T720A hearts from both sexes exhibited negative enrichment for genes involved in mitochondrial respiration, indicating downregulation of this pathway. This finding suggests that CHD7 regulates the expression of oxidative phosphorylation genes which is impaired in Chd7T720A/T720A hearts of both sexes. While Chd7T720A/T720A homozygotes exhibited widespread transcriptional dysregulation affecting multiple biological processes, in the presence of a WT allele (Chd7T720A/+), heterozygote hearts showed more limited pathway-level disruption despite a considerable number of DEGs (Supplementary Fig. 4a and 4b). Unlike the strong down-regulation observed in homozygotes, Chd7T720A/+ hearts exhibited positive enrichment of genes involved in mitochondrial respiration, indicating a possible compensatory response of the WT allele to maintain cellular energy homeostasis (Supplementary Fig. 4c).
Comparative analysis and quantitation of cardiomyopathy in E18.5 mouse embryos. (a) Cardiac imaging and 3D reconstruction. Cross-sectional views of hearts are shown on the left, with 3D reconstructions highlighting the left ventricular wall (yellow) and right ventricular chamber (green) in the middle panel. The right panel shows the ventricular wall and septum. Three phenotypes are shown: normal, enlarged ventricular chamber (DCM), and reduced ventricular chamber (HCM). (b-c) Quantitative analysis of cardiac morphology. Ratios of the volumes of the left and right ventricular chamber were normalized to total ventricle volumes by measuring individual volumes in cubic millimeters (mm³). Comparisons were made among Chd7+/+ (WT), Chd7fs/+, Chd7T720A/+, Chd7T720A/T720A, and Chd7fs/T720A mice. Z scores were calculated using the average and standard deviation of WT, with significant thresholds marked at Z = ± 2. DCM and HCM phenotypes are color-coded in pink and blue, respectively.
CHD7 T730 missense variant alters ETS family transcription factor target gene expression
The role of CHD7 as a transcriptional co-regulator is well established19,20,21,22,23,24, and previous studies have shown that CHD7 co-localizes on chromatin with the ETS family transcription factor RUNX125. To further investigate this interaction, we performed an in silico comparison of CHD7 and ETS family transcription factor target genes. Using the Harmonizome ENCODE Transcription Factor Target gene set for CHD7 and the MSigDB Transcription Factor Target gene sets for the ETS family transcription factors26,27, we found a significant overlap of 57–73% (Fig. 5a). Next, we asked if DEGs in neonatal hearts of Chd7T720A/T720A homozygotes were significantly enriched for ETS recognition motifs. As shown in Fig. 5b, up-regulated DEGs in both male and female Chd7T720A/T720A hearts were significantly enriched for targets of ETS1, ETS2, RUNX1, PU1, PEA3, ELF1 and GABPβ, with -log p-values ranging from p ≤ 10−7 to 10−36, depending on sex and the specific ETS family transcription factor (notably, no down-regulated DEGs were shared between males and females, Supplementary Fig. 3b). Additionally, RUNX1 target gene enrichment was observed using GSEA against the ENRICHR RUNX1 LINCS L1000 CRISPR KO Sigs gene set28, as shown in the enrichment plots in Fig. 5c. These findings suggest that T720 palys a key role in CHD7 function as a co-regulator of ETS family transcription factors, potentially influencing RUNX1-mediated transcriptional programs. Given that RUNX1 activation has been linked to pathological myocardial remodeling and heart failure via TGF-β signaling, these findings suggest that CHD7 loss-of-function may contribute to DCM through transcriptional dysregulation. While the exact mechanism remains unclear, CHD7 may regulate RUNX1 directly or through broader chromatin remodeling effects that influence cardiac transcriptional networks. Further studies will be necessary to determine the precise role of CHD7 in myocardial remodeling and disease progression.
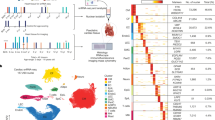
Transcriptional profiling of neonatal hearts from Chd7T720A/T720A mice. (a-b) Heat maps (a) and PCA plots (b) showing DEGs in Chd7T720A/T720A males (left) and females (right) neonatal hearts compared to Chd7+/+ (WT) controls (N = 3 per group for both WT and mutant). (c) Volcano plots highlighting the most significantly enriched pathways for DEGs determined by ORA of GO-BP gene sets in male and female Chd7T720A/T720A hearts. Key pathways such as cytoskeleton organization, lipid metabolic process, cell adhesion, and cell cycle are shown. (d) GSEA enrichment plots showing downregulation of the mitochondrial protein containing complex gene sets in homozygous Chd7T720A/T720A mouse neonatal hearts compared to those of wild type controls. The NES values are displayed for both males (NES = −3.72) and females (NES = −8.12). (e) Bar chart showing NES from GSEA of commonly disrupted pathways in male and female Chd7T720A/T720A neonatal hearts. Pathways include mitochondrial components, the inner mitochondrial membrane, and the tricarboxylic acid cycle enzyme complex.
Altered ETS family transcription factor target gene expression in the hearts of CHD7 T730 missense mutant mice. (a) Overlap between CHD7 and ETS family transcription factor targets as reported in the Harmonizome database. The table shows the number of target genes for each transcription factor, the overlap with CHD7 targets, and the percentage of overlap. (b) Bar chart depicting enrichment scores for ETS family transcription factors targets among DEGs from homozygous Chd7T720A/T720A male and female neonatal hearts. (c) GSEA enrichment plots for RUNX1 upregulated target genes in the Chd7T720A/T720A transcriptome of male and female neonatal hearts. The NES values are displayed for male (NES = 1.35) and female (NES = 1.18).
Multiple kinases phosphorylate CHD7 at T730 and the adjacent S734 residue
Because T730 can potentially be phosphorylated, with regulatory consequences for CHD7 activity, we analyzed data from Kinase Library29, accessible through PhosphoSitePlus to ascertain if T730 was a potential kinase target. The Kinase Library algorithm predicts kinase activity by integrating phosphorylated residues in proximal sequences (phospho-priming) with empirical data. This in silico analysis identified two isoforms of glycogen synthase kinase-3 (GSK3) as the primary candidates for T730 phosphorylation, with a confidence score of 100.0% for GSK3β and 99.996% for GSK3α (Supplementary Fig. 5a) when adjacent residues at S725 (pS725) or S734 (pS734) were phosphorylated (primed). Interestingly, a CHD7 serine-to-proline mutation (S734P) was identified in a patient with a neurodevelopmental disorder and classified as a variant of unknown significance (Supplementary Table 1). Notably, both pT730 and pS734 phosphorylation events have been observed in human cell lines, mouse heart and brain tissues13,15, and mouse NIH3T3 cells14. These findings suggest that phosphorylation of these CHD7 residues may be part of a yet-to-be-characterized but functionally important kinase signaling pathway.
A preliminary round of in vitro kinase assays was performed using a peptide containing T730 (Fig. 6a and b) and 20 kinases with aligned motifs to identify potential kinases that phosphorylate this site. The results showed that this peptide was phosphorylated by GSK3β, and even more robustly DYRK2 and HIPK1 (Supplementary Fig. 5b). However, this assay could not distinguish between the two potential phosphorylation sites, T730 and S734, nor determine the impact of S734 priming on GSK3β activity. To address this, we designed peptides in which either T730 or the potential priming site S734 was substituted with non-phosphorylatable alanine (T730 A or S734A, respectively), as well as a “pre-primed” peptide containing pS734 (Fig. 6b). These substitutions allowed us to determine whether specific kinases target T730 or S734 and how phosphorylation at S734 influences T730 phosphorylation efficiency. S734A was chosen as a non-phosphorylatable control to determine the role of priming phosphorylation at this site without introducing structural perturbations. While potentially clinically relevant, proline substitution (S734P) was not used in this context, as this amino acid induces local conformational rigidity, which could independently alter kinase recognition and activity. Instead, our kinase assays were specifically designed to assess phosphorylation site preference, ensuring that observed changes reflect phosphorylation loss rather than structural effects. Using these peptides, in vitro kinase assays revealed distinct phosphorylation preferences among kinases. DYRK2 preferentially phosphorylated S734, as indicated by the loss of activity on the S734A peptide (Fig. 6c). HIPK1 specifically targeted T730, with phosphorylation abolished only in the T730A variant, indicating that HIPK1 functions independently of S734 priming (Fig. 6c). GSK3α and GSK3β exhibited significantly enhanced phosphorylation activity on the pre-primed pS734 peptide, suggesting that their phosphorylation of T730 is facilitated by prior priming at S734 (Fig. 6d). GSK3α primarily targets T730, but its activity was enhanced when S734 is pre-phosphorylated. GSK3β, in contrast, strongly preferred S734 over T730, yet its phosphorylation of T730 was markedly increased when S734 was pre-phosphorylated. These findings suggest that T730 phosphorylation by GSK3α and GSK3β occurs in conjunction with pS734 priming, with both GSK3β and DYRK2 potentially acting as priming kinases. Meanwhile, HIPK1 phosphorylates T730 independently of S734 phosphorylation, indicating an alternative regulatory mechanism.
To confirm that full-length CHD7 serves as a bona fide substrate for phosphorylation, we performed in vitro kinase assays using purified GSK3β protein and full-length human CHD7 protein produced in Sf9 insect cells. As shown in Fig. 6e, recombinant GSK3β successfully phosphorylated full-length CHD7 protein, confirming CHD7 as a direct target for GSK3β-mediated phosphorylation. While the roles of these kinases in heart development remain incompletely understood, previous studies have reported cardiac defects, including hypertrophic cardiomyopathy, in GSK3β-deficient (Gsk3b−/−) mice30. Additionally, DYRK2 functions as a negative regulator of cardiomyocyte growth and suppresses the hypertrophic response to adrenergic stimulation, a critical pathway in cardiomyopathy progression31,32. These findings suggest that GSK3β, DYRK2, and HIPK1 are critically involved in heart development, suggesting that CHD7 phosphorylation is an integral component of these kinase-dependent regulatory pathways in the heart.
Pathogenic missense variants impact T730 phosphorylation
Guided by the Single Nucleotide Polymorphism database (dbSNP) at NextProt12, we explored how pathogenic missense variants near T730 impact its phosphorylation. Given that GSK3α and β are proline-directed kinases that phosphorylate T730 and S734, our attention was drawn to proline residues adjacent to these sites. Among the six prolines in this region, missense mutations in P732 and P737 have clinical significance. Notably, a male patient with probable CHARGE syndrome and hypopituitarism was identified with a heterozygous c.2194 C > G (p.Pro732Ala) missense variant, catalogued as rs200277422 (ClinVar; [VCV000095779.15])5,8,33. This individual exhibited a range of phenotypes, including Tetralogy of Fallot, sensorineural hearing loss, micropenis with bilateral undescended testes, abnormally shaped ears, strabismus and hypermetropia, trigonocephaly, a small occipital meningocele, and severe developmental delay with short stature33. Other missense variants identified at P737 were P737del (in-frame deletion, rs748363597, ClinVar; [VCV000421199.13]) and P737L (rs899004166). The P737del variant is associated with CHARGE syndrome, (RCV001392601.13)34 in an individual with neurodevelopmental disorder of uncertain significance (RCV000717371.1). Both the P732A and P737L variants have also been observed in lymphoma and skin cancer, respectively (https://www.cbioportal.org).
In vitro kinase assays using peptides containing these specific missense mutations (Fig. 6b) revealed that DYRK2 and GSK3β activities were significantly decreased in the presence of the P737L mutation compared to the WT peptide (Fig. 6c and d). In contrast, the P732A mutation did not noticeably affect those kinase activities. These findings suggest that mutations adjacent to T730 can modulate kinase activity (summarized in Fig. 6f) and potentially impact CHD7 function. Furthermore, these data support the model for phosphorylation of CHD7 by GSK3α/β, HIPK1, and DYRK2, as illustrated in Fig. 6f. Mutations near T730 may differentially alter CHD7 phosphorylation dynamics, thereby influencing its role as a transcriptional regulator. Given the clinical significance of these variants, further studies are warranted to elucidate how these phosphorylation changes contribute to CHD7-related developmental disorders and disease mechanisms.
Kinases phosphorylate human CHD7 at T730 and S734, pivotal sites in a regulatory kinase pathway for CHD7. (a) Schematic diagram of the T730 and S734 sites along with the key protein domains in CHD7. The consensus sequence of the GSK3β substrate, ‘Ser/Thr-X-X–X-Ser/Thr-P’, is highlighted in a red box and aligned with the amino acid sequence near T730. The WT T730 peptide used for the in vitro kinase assays is underlined. (b) List of synthesized peptides surrounding T730 used in the in vitro kinase assays, including a peptide with pre-priming phosphorylation at S734 (pS734) and several mutant peptides. (c) In vitro kinase assays using DYRK2 or HIPK1 kinases on the indicated peptides near CHD7 T730 and its mutant variants. Relative kinase activities were normalized to the luminescence signal of WT controls, measured using a GloMax plate reader. (d) In vitro kinase assay using GSK3α or GSK3β on the indicated peptides near CHD7 T730. Enzyme activity was measured by luminescence as shown in (c). (e) Autoradiography and Coomassie blue staining of an SDS-PAGE gel showing the results of in vitro kinase reactions using GSK3β on human full-length CHD7 protein in the presence of radioactive [γ−32P]ATP. (f) Model for the regulatory phosphorylation of CHD7 by kinases and the effects of nearby P737L mutation. Statistical significance is indicated as follows: *, **, ***; P < 0.05, P < 0.01, and P < 0.001, respectively (Student t-test).
Discussion
In this study, we identified a T720A variant in Chd7 that leads to developmental heart abnormalities, providing the first example of a genotype-phenotype correlation linking a specific CHD7 defect (loss of phosphorylation) to a pathogenic phenotype, namely cardiomyopathy, in a mouse model. While the loss of CHD7 function due to null alleles is well established as the underlying cause of CHARGE syndrome, no clear genotype-phenotype correlations have emerged to date that associate specific CHD7 functions with CHARGE-associated syndromic defects. This lack of correlation is not surprising, given that CHARGE syndrome results from haploinsufficiency caused by heterozygosity for a CHD7 null allele. Animal models have similarly replicated the human disease by employing null alleles to investigate the role of this chromatin remodeler in development and pathogenesis. Therefore, our finding that cardiomyopathy develops in E18.5 embryos was unexpected, as neither CHD7 null alleles in CHARGE syndrome patients nor previous studies using Chd7-deficient mouse models have reported an association with cardiomyopathy.
Notably, the T720A phosphorylation-deficient allele did not fully recapitulate the phenotypes observed in CHD7 null alleles. Heterozygous Chd7T720A/+ mice did not display the characteristic circling behavior associated with inner ear defects observed in Chd7fs/+ mice, and homozygous Chd7T720A/T720A mice were viable, in contrast to null alleles, which are embryonic lethal. Similarly, septal defects, which are frequent in mice carrying a null allele and also seen in mice carrying the Chd7fs allele, were not found in Chd7T720A/+ heterozygotes, Chd7T720A/T720A homozygotes, or Chd7fs/T720A compound heterozygotes. These findings suggest that Chd7fs null mutation specifically contributes to the development of septal defects, whereas the presence of the Chd7T720A allele mitigates this effect in compound heterozygotes.
Our quantitative analyses revealed that both a CHD7 missense mutation and gene dosage could significantly influence cardiomyopathy phenotypes. Chd7T720A/+ mice exhibited features characteristic of HCM, including reduced ventricular chamber volume and thickened ventricular walls, whereas Chd7T720A/T720A mice displayed a mixed phenotype, with some embryos showing mild HCM while others exhibited chamber dilation, ultimately leading to DCM. Most Chd7fs/+ mice developed DCM, characterized by significantly enlarged ventricular chambers and thinned ventricular walls, consistent with impaired myocardial function, whereas Chd7fs/T720A mice exhibited reduced ventricular chamber volume and ventricular wall thickening, displaying an HCM-like phenotype rather than DCM. These findings indicate that CHD7 haploinsufficiency (Chd7fs/+) is primarily associated with DCM, whereas the presence of the missense allele (Chd7T720A) partially compensates for CHD7 loss, triggering hypertrophic remodeling instead.
The distinct cardiomyopathy phenotypes observed in Chd7T720A, Chd7T720A/T720A, and Chd7fs/T720A mice indicate that CHD7 phosphorylation and dosage interact to regulate cardiac function. RNA-seq analysis of Chd7T720A/T720A hearts revealed disruptions in mitochondrial energy metabolism, suggesting a critical role for CHD7 phosphorylation in maintaining mitochondrial function. Consistent with this, Chd7T720A/T720A mice exhibited myocardial dysfunction and DCM, indicating that complete loss of CHD7 phosphorylation impairs mitochondrial energy homeostasis. In contrast, Chd7T720A/+ mice did not develop DCM but instead exhibited signs of compensatory hypertrophic remodeling, characteristic of HCM. RNA-seq analysis of Chd7T720A/+ hearts revealed compensatory up-regulation of mitochondrial metabolism pathways, suggesting the presence of a WT allele supports adaptive mechanisms to maintain cardiac function (Supplementary Fig. 4c). This suggests that while mitochondrial function is impaired in homozygotes, heterozygotes engage transcriptional compensation to mitigate metabolic deficiencies, possibly promoting hypertrophic remodeling rather than contractile dysfunction. Thus, while partial CHD7 phosphorylation loss activates compensatory hypertrophic pathways, complete loss disrupts mitochondrial homeostasis, leading to DCM. Similarly, in Chd7fs/T720A mice, the presence of a phosphorylation-deficient allele appears to provide partial functional compensation, possibly mitigating the deleterious effects of CHD7 haploinsufficiency and leading to HCM rather than DCM. On the other hand, Chd7fs/+ mice, which carry a null allele, developed DCM, suggesting that CHD7 haploinsufficiency contributes to impaired myocardial contractility, potentially through mechanisms involving mitochondrial dysfunction. The Chd7fs allele introduces a premature termination codon (PTC) upstream of the chromodomain, making it a likely target for nonsense-mediated mRNA decay (NMD)35. Although we did not directly assess whether Chd7fs transcripts undergo NMD, the absence of viable Chd7fs/fs homozygous mice is consistent with previous studies demonstrating that complete CHD7 loss is embryonically lethal36. Viability, and consistency with known haploinsufficient CHARGE-like phenotypes including microphthalmia, anophthalmia, and vestibular dysfunction leading to circling behavior, observed in Chd7fs/+ heterozygotes, further support Chd7fs as a null allele. While we did not directly assess transcriptomic effects in Chd7fs/+ hearts, it is expected that CHD7 haploinsufficiency results in broader transcriptional dysregulation compared to the T720A phosphorylation-deficient model. Future transcriptomic analysis of Chd7fs/+ hearts will be essential to directly compare the molecular consequences of haploinsufficiency versus phosphorylation loss. This will further clarify the distinct roles of CHD7 dosage and post-translational modification in developmental regulation.
RNA-seq analysis of neonatal Chd7T720A/T720A hearts revealed transcriptomic differences between males and females. While some subtle sex-based variations have been observed in human patients with CHARGE syndrome, their biological significance remains unclear. Our data indicate that female Chd7T720A/T720A mice exhibit greater changes in mitochondrial and immune-related pathways compared to males, though the functional consequences of these differences require further investigation. GSEA for the mitochondrial protein-containing complex showed a greater degree of down-regulation in females (NES = −8.12) compared to males (NES = −3.72). While this could suggest a stronger impact on mitochondrial function and oxidative stress in females, the physiological relevance of this difference remains to be determined. Additionally, female hearts exhibited increased expression of genes involved in immune response and lymphocyte activation, which could indicate differences in immune regulation. In contrast, transcriptomic changes in male hearts were more associated with lipid metabolism and cell cycle regulation. One possible explanation is that CHD7, as a chromatin remodeler, may regulate sex-specific transcriptional programs through interactions with epigenetic regulators or transcription factors that are differentially expressed or regulated in male and female embryonic hearts. Notably, the target genes of ETS family transcription factors, including RUNX1, were significantly enriched among differentially expressed genes in Chd7T720A/T720A hearts. This suggests that CHD7 may influence ETS factor-mediated transcription in a sex-dependent manner, potentially altering chromatin accessibility at key regulatory loci involved in cardiac development. Prior studies have reported that RUNX1 interacts with estrogen receptors (ERs) to modulate gene expression in a sex-dependent manner37,38. RUNX1 plays a crucial role in cardiac development and exhibits sex-dependent transcriptional regulation, which could contribute to the observed transcriptomic differences in Chd7T720A/T720A hearts. Additionally, the enrichment of immune-related pathways in female embryonic hearts suggests that sex-linked differences in inflammatory signaling may influence the phenotype. It has been reported that fetal immune responses can be shaped by sex chromosome-linked gene expression differences, particularly in Toll-like receptor (TLR) signaling and interferon-mediated immune responses39. Given that CHD7 has been implicated in immune regulation40, it is plausible that the female-specific immune pathway enrichment observed in Chd7T720A/T720A hearts reflects an interaction between CHD7-mediated chromatin remodeling and immune gene networks, potentially contributing to sex-biased immune regulation in cardiac development. Furthermore, the stronger downregulation of mitochondrial metabolism genes in female hearts may be related to known sex differences in oxidative stress response mechanisms. Sex differences in oxidative stress response have been reported, with female hearts often exhibiting greater resistance to oxidative damage compared to males41. Estrogen has been shown to protect cardiac cells from oxidative damage and mitochondrial dysfunction, potentially contributing to the observed differences in mitochondrial resilience between male and female hearts41. However, whether this hypothesis applies to embryonic heart development remains unclear and requires further investigation. Together, these findings highlight the importance of considering sex as a biological variable in CHD7-associated cardiomyopathies and suggest that sex-specific differences in transcriptional regulation, immune signaling, and oxidative stress responses contribute to the observed phenotypic differences.
Gene expression analysis in CHD7 phosphorylation-deficient (Chd7T720A/T720A ) hearts revealed significant enrichment of ETS family transcription factor targets, including RUNX1, suggesting that CHD7 normally functions as a co-repressor of RUNX1-mediated transcription. Given that RUNX1 has been implicated in cardiomyopathy and is upregulated in pathological cardiac conditions42,43,44,45,46, these findings suggest that CHD7 phosphorylation is necessary to suppress aberrant RUNX1 activity in the heart and maintain transcriptional homeostasis. However, it remains to be determined whether CHD7 directly suppresses RUNX1 activity or if its effects are mediated through chromatin remodeling. Interestingly, our study identified GSK3β as a kinase that phosphorylates T730 of CHD7, suggesting that CHD7 phosphorylation is integrated into GSK3β-mediated signaling. Given that GSK3β is essential for cardiac development, its loss (Gsk3b−/−) results in severe heart defects, some of which overlap with CHARGE30,47. These findings suggest the existence of a CHD7-GSK3β regulatory network in cardiac homeostasis, where loss of CHD7 phosphorylation alters transcriptional regulation and disrupts signaling pathways critical for heart development.
The genotype-phenotype correlation identified in this study for the Chd7T720A allele, which significantly influences cardiac development, enhances our understanding of CHD7 function across various organ systems and provides valuable insights into the pathophysiology of CHD7-related anomalies. These phenotypes, characterized by distinct cardiomyopathy features, underscore the pivotal role of CHD7 in normal heart development. Notably, a CHD7 variant of ‘uncertain significance’ linked to primary dilated cardiomyopathy was recently documented in the NIH ClinVar database (Accession: SCV003932388.1 and RCV003319232.1), suggesting that CHD7 variants in humans might also contribute to cardiomyopathy. This finding expands the spectrum of CHD7-related cardiac pathologies and highlights the need for further investigation into CHD7 function in cardiac development and disease. Our findings lay the groundwork for further research to unravel the mechanisms underlying specific pathophysiological features of CHD7-associated genetic disorders. In particular, assessing how structural changes, such as alterations in ventricular wall thickness and chamber size, impact functional cardiac parameters, including ejection fraction, stroke volume, and cardiac output, would provide valuable insights. Furthermore, investigating clinical manifestations of heart failure or reduced exercise capacity in these animals may further clarify the physiological impact of CHD7 deficiency. Such evaluations could strengthen the clinical relevance of our findings and facilitate the translation of these insights into therapeutic strategies for CHD7-related cardiac conditions.
Conclusions
This study identifies a novel genotype-phenotype correlation by linking a CHD7 missense variant (Chd7) to developmental heart abnormalities, specifically cardiomyopathy. It demonstrates that the T730A variant, and potentially loss of CHD7 phosphorylation at this site, disrupts transcriptional regulation, particularly of mitochondrial energy metabolism genes, and leads to altered cardiac morphology. These findings expand the understanding of CHD7’s role in CHARGE syndrome and related pathologies, providing valuable insights into the molecular mechanisms underlying these conditions and identifying potential avenues for therapeutic exploration.
Methods
Sex as a biological variable
Our study included both male and female animals, with similar findings observed across both sexes.
Generation of Chd7 T720A knock-in mouse & genotyping
The Chd7T720A knock-in mouse model (human T730 equivalent in mouse) was generated using CRISPR-Cas9 technology. Zygotes from C57BL/6 mice were electroporated with sgRNA at the Baylor College of Medicine’s Genetically Engineered Rodent Models (GERM) Core. As detailed in Supplementary Fig. 1, this sgRNA was specifically engineered to include two additional silent mutations to prevent the Cas9-sgRNA complex from re-targeting the site post-modification. The designed sgRNA induced a codon change from Thr (ACA) to Ala (GCG) in the Chd7 gene and introduced a novel HaeII restriction site to facilitate genotyping. Genotyping was performed by PCR using genomic DNA extracted from mouse tails with the following primers: Chd7-T720A-HaeII-F: 5’-GCC TTT GCC ACC ACT CTT TAT TA-3’ and Chd7-T720A-HaeII-R: 5’-TAA GCT GGG TTT CTG TGG GC-3’. This PCR amplification yielded a 417 bp product, which resulted in fragments of 226 bp and 191 bp upon digestion with HaeII in mutant mice.
In vitro kinase assay using human full-length CHD7 protein
Full-length CHD7 protein was purified from Sf9 insect cells, as previously described48. The kinase reaction included either 12.8 ml of BC100 buffer (10% glycerol, 20 mM HEPES pH 7.9, 0.4 mM EDTA, and 100 mM NaCl supplemented with 1 mM β-mercaptoethanol and cOmplete™ protease inhibitors from Roche/Millipore Sigma) as negative control, or 360 ng of CHD7 protein in the same buffer. Two hundred nanograms (2 µl) of GST-GSK3β (R&D SYSTEMS/bio-techne, catalog number 2506-KS-010) in a buffer containing 50 mM Tris-HCl, pH 7.5, 150 mM NaCl, 0.25 mM DTT, 10 mM glutathione, 0.1 mM EDTA, 0.1 mM PMSF, and 25% glycerol was then added. Reactions were initiated by adding 100 mM ATP and 300 mM MgCl2, supplemented with 0.5 ml of 32P-radiolabeled ATP (3000 Ci/mmol; catalog number BLU502 A250UC, PerkinElmer®) and incubated at 37 ℃ for 1 h. Reactions were terminated by adding SDS-PAGE loading buffer. Samples were then subjected to SDS-PAGE on 4–20% gradient gels, followed by fixation and Coomassie blue staining. For radiolabeled kinase activity detection, gels were dried and exposed to a Phosphorimager screen for 1 h or to X-ray film overnight.
Candidate kinase selection and in vitro kinase assays on T730 peptides
A panel of 20 protein kinases was selected for initial screening, based on predictive rankings from the PhosphoNET database (www.phosphonet.ca). The in vitro kinase assays were conducted at Kinexus Bioinformatics Corporation using the ADP-Glo™ Kinase Assay kit (Promega), following the manufacturer’s protocols. Kinase activity was quantified by measuring adenosine diphosphate (ADP) production, as indicated by an increase in luminescence signal using the GloMax® Discover System (Promega, Cat# E7031). Results were expressed in relative light units (RLU). Kinase activity was normalized against T730A-S734A, a non-phosphorylatable control peptide, to compensate for potential autophosphorylation of the kinases or nonspecific background signals. Assays with four candidate kinases were conducted in duplicate. Post-assay data were normalized against control peptide values and analyzed statistically using the Student’s t-test to compare kinase activity against that of the WT peptide. Peptide substrates used in the assays were synthesized by Kinexus Bioinformatics Corporation with 85% purity.
Iodine-contrast Micro-CT imaging
Iodine-contrast micro-CT imaging was performed as previously described49. Mouse embryos at embryonic day 18.5 were fixed in 10% (v/v) neutral buffered formalin (NBF) for at least three days at 4 ℃. Post-fixation, embryos were immersed in STABILITY buffer, consisting of 4% (w/v) paraformaldehyde, 4% (w/v) acrylamide, 0.05% bis-acrylamide, 0.25% (w/v) VA044 initiator, and 0.05% (w/v) saponin in phosphate buffered saline (PBS) at pH 7.2, and were stored at 4 ℃ for three days. The crosslinking reaction was facilitated using the X-CLARITYTM hydrogel polymerization system (Logos Biosystem, Cat# C200001) under conditions at −90 kPa and 37 ℃ for three hours. After crosslinking, the polymer was carefully removed, and embryos were washed twice in PBS for one hour each, followed by an overnight wash at 4 ℃. The embryos were then preserved in PBS supplemented with 0.1% (w/v) sodium azide at 4 ℃. For staining, the embryos were immersed in 0.1 N iodine solution (Sigma, Cat#38060) for at least three days at room temperature. Before imaging, embryos were embedded in 1% (w/v) agarose. The Skyscan 1272 micro-CT scanner (Bruker) was used to acquire raw data requisite for 3D imaging. Projection images were reconstructed with NRecon Reconstruction software (Version 1.6.9.8; Bruker). The reconstructed 3D data were visualized volumetrically using CTVox (Version 2.6.0; Bruker) or alternatively, Slicer freeware (Version 4.4.0; www.slicer.org), specifically in the coronal section to facilitate detailed morphological analysis of cardiac structures.
RNA extraction and sequencing
Mouse hearts from neonatal mice at postnatal day 0 (P0) were immediately processed for total RNA extraction. The study included three biological replicates for each experimental group, encompassing male and female hearts of Chd7+/+, Chd7T720A/+, and Chd7T720A/T720A mice. Total RNA was extracted using the RNeasy kit (Qiagen), following the manufacturer’s protocol. RNA libraries were prepared and sequenced at the University of Houston Sequencing Core (UH-SEQ). Quality control of the RNA samples was performed using the Tapestation 4200 (Agilent), and RNA quantification was carried out with the Qubit Fluorometer (Thermo Scientific). RNA libraries were prepared using the QIAseq Stranded Total RNA library Kit (Qiagen) with an input of 100 ng RNA. Ribosomal RNA depletion was performed using the QIAseq FastSelect HMR kit (Qiagen). RNA was then fragmented, reverse-transcribed into cDNA, and ligated with Illumina sequencing adaptors. Size selection for libraries was performed using SPRIselect beads (Beckman Coulter), and library purity was assessed on the Tapestation 4200 (Agilent) using the High Sensitivity DNA 1000 tape, targeting a size of approximately 300 bp. Libraries were pooled and sequenced on the NextSeq 500 system (Illumina), generating approximately 20 million 2 × 76 bp paired-end reads per sample. RNA-seq data were processed using trimGalore for trimming low-quality base pairs, and mapped to the mouse genome (UCSC build mm10) using star 2.7.10b50.
RNA-seq analysis
Gene expression was quantified using featureCounts51 and the GENCODE gene model. Normalization of gene expression data was conducted with RUVSeq52. Differential expression analysis between the groups was performed using the edgeR53 package in R, considering both sex and genotype as variables. Genes were considered differentially expressed if they had an FDR-adjusted p-value below 0.05 and a fold change greater than 1.5x. Pathway enrichment analysis was performed using the Hallmark, KEGG, GO.BP, GO.CC, Reactome, Enrichr, and Harmonizome database to identify biological processes and pathways significantly affected by the T720 A variant. Both GSEA54 and ORA as implemented by the Molecular Signatures Database (MSigDB) were used55. Volcano plots were generated using VolcanoseR, a web app for creating, exploring, labeling, and sharing volcano plots56. Bar charts were created with GraphPad Prism version 10.1.1 for macOS, GraphPad Software, Boston, Massachusetts USA (www.graphpad.com).
Statistical analysis for diagnosis of cardiomyopathy
To determine whether the ventricular volumes in different genotypes were significantly different from the WT control, we performed the following statistical analyses: The data included left and right ventricular chamber volumes normalized by the total volume for each sample across various genotypes, including WT, fs/+, T720A/+, T720A/T720A, and fs/T720A. The mean and standard deviation of the normalized ventricular volumes for the WT control group were calculated. For each sample in the different genotypes, the Z-score was calculated using the formula: Z = (X − µ)/σ, where X is the sample value, µ is the mean of the WT control group, and σ is the standard deviation of the WT control group. The p-value for each Z-score was then computed to assess the significance of the deviation from the WT control mean, using the formula: p = 2 × (1 − Φ(∣Z∣)), where Φ represents the cumulative distribution function of the standard normal distribution. To account for multiple comparisons, a Bonferroni correction was applied to adjust the significance threshold. The corrected significance level (αcorrected) was calculated as: αcorrected = α/N, where α is the initial significance level (0.05) and N is the number of comparisons. Each sample was classified based on the calculated Z-scores:
-
HCM: Z-score < −2 and Bonferroni-corrected p-value < 0.05, indicating significantly smaller ventricular volumes compared to WT.
-
DCM: Z-score > 2 and Bonferroni-corrected p-value < 0.05, indicating significantly larger ventricular volumes compared to WT.
-
Normal: Z-score between − 2 and 2, or Bonferroni-corrected p-value ≥ 0.05.
All statistical analyses were performed using Python 3.11.8 with the ‘numpy’ and ‘scipy’ libraries.
Statistics
All data are expressed as means with actual values. For comparison between two groups, an unpaired two-tailed Student’s t- test was performed with p values less than 0.05 considered statistically significant. For the diagnosis of cardiomyopathy, a Z-test was conducted, with a Z-score < −2 or > 2 followed by a Bonferroni-corrected p-value < 0.05 considered significant.
Study approval and ethical considerations
All mice were housed and maintained in a facility accredited by the Association for Assessment and Accreditation of Laboratory Animal Care (AAALAC). All animal sacrifice and experimental procedures were conducted in accordance with protocols approved by the Baylor College of Medicine Institutional Animal Care and Use Committee (IACUC) (protocol number AN-7189, approval period from 7/11/2022 to 7/10/2025). Furthermore, this study is reported in accordance with the ARRIVE guidelines.
Animal euthanasia
Adult mice were euthanized by CO₂ asphyxiation in a chamber. Neonates were euthanized by decapitation, and embryos were obtained following removal from the mother, which was euthanized by CO₂ asphyxiation.
Source of mice
The mice used in this study were generated at the Baylor College of Medicine as described above.
Data availability
The datasets generated and/or analyzed during the current study are available within the article and its supplementary materials. The RNA-seq data supporting the findings of this study have been deposited in the NCBI Gene Expression Omnibus (GEO) repository under accession number GSE292553.
References
Clapier, C. R. & Cairns, B. R. The biology of chromatin remodeling complexes. Annu. Rev. Biochem. 78, 273–304. https://doi.org/10.1146/annurev.biochem.77.062706.153223 (2009).
Eisen, J. A., Sweder, K. S. & Hanawalt, P. C. Evolution of the SNF2 family of proteins: subfamilies with distinct sequences and functions. Nucleic Acids Res. 23, 2715–2723. https://doi.org/10.1093/nar/23.14.2715 (1995).
Ho, L. & Crabtree, G. R. Chromatin remodelling during development. Nature 463, 474–484. https://doi.org/10.1038/nature08911 (2010).
van Ravenswaaij-Arts, C. M. et al. in GeneReviews((R)) (eds M. P. Adam (1993).
Jongmans, M. C. et al. CHARGE syndrome: the phenotypic spectrum of mutations in the CHD7 gene. J. Med. Genet. 43, 306–314. https://doi.org/10.1136/jmg.2005.036061 (2006).
Lalani, S. R. et al. Spectrum of CHD7 mutations in 110 individuals with CHARGE syndrome and genotype-phenotype correlation. Am. J. Hum. Genet. 78, 303–314. https://doi.org/10.1086/500273 (2006).
Bergman, J. E. et al. CHD7 mutations and CHARGE syndrome: the clinical implications of an expanding phenotype. J. Med. Genet. 48, 334–342. https://doi.org/10.1136/jmg.2010.087106 (2011).
Bergman, J. E. et al. A novel classification system to predict the pathogenic effects of CHD7 missense variants in CHARGE syndrome. Hum. Mutat. 33, 1251–1260. https://doi.org/10.1002/humu.22106 (2012).
Janssen, N. et al. Mutation update on the CHD7 gene involved in CHARGE syndrome. Hum. Mutat. 33, 1149–1160. https://doi.org/10.1002/humu.22086 (2012).
Balasubramanian, R. & Crowley, W. F. Jr. Reproductive endocrine phenotypes relating to CHD7 mutations in humans. Am. J. Med. Genet. C Semin Med. Genet. 175, 507–515. https://doi.org/10.1002/ajmg.c.31585 (2017).
Hornbeck, P. V. et al. PhosphoSitePlus, 2014: mutations, PTMs and recalibrations. Nucleic Acids Res. 43 (D512-520). https://doi.org/10.1093/nar/gku1267 (2015).
Lane, L. et al. NeXtProt: a knowledge platform for human proteins. Nucleic Acids Res. 40, D76–83. https://doi.org/10.1093/nar/gkr1179 (2012).
Sharma, K. et al. Ultradeep human phosphoproteome reveals a distinct regulatory nature of Tyr and Ser/Thr-based signaling. Cell. Rep. 8, 1583–1594. https://doi.org/10.1016/j.celrep.2014.07.036 (2014).
Gnad, F. et al. Systems-wide analysis of K-Ras, Cdc42, and PAK4 signaling by quantitative phosphoproteomics. Mol. Cell. Proteom. 12, 2070–2080. https://doi.org/10.1074/mcp.M112.027052 (2013).
Carrier, M., Joint, M., Lutzing, R., Page, A. & Rochette-Egly, C. Phosphoproteome and transcriptome of RA-Responsive and RA-Resistant breast Cancer cell lines. PLoS One. 11, e0157290. https://doi.org/10.1371/journal.pone.0157290 (2016).
Bosman, E. A. et al. Multiple mutations in mouse Chd7 provide models for CHARGE syndrome. Hum. Mol. Genet. 14, 3463–3476. https://doi.org/10.1093/hmg/ddi375 (2005).
Jiang, X. et al. The mutation in Chd7 causes misexpression of Bmp4 and developmental defects in telencephalic midline. Am. J. Pathol. 181, 626–641. https://doi.org/10.1016/j.ajpath.2012.05.006 (2012).
Polito, M. V. et al. CHARGE syndrome and congenital heart diseases: systematic review of literature. Monaldi Arch. Chest Dis. https://doi.org/10.4081/monaldi.2023.2661 (2023).
Bajpai, R. et al. CHD7 cooperates with PBAF to control multipotent neural crest formation. Nature 463, 958–962. https://doi.org/10.1038/nature08733 (2010).
Engelen, E. et al. Sox2 cooperates with Chd7 to regulate genes that are mutated in human syndromes. Nat. Genet. 43, 607–611. https://doi.org/10.1038/ng.825 (2011).
He, D. et al. Chd7 cooperates with Sox10 and regulates the onset of CNS myelination and remyelination. Nat. Neurosci. 19, 678–689. https://doi.org/10.1038/nn.4258 (2016).
Stathopoulou, A. et al. CHARGE syndrome-associated CHD7 acts at ISL1-regulated enhancers to modulate second heart field gene expression. Cardiovasc. Res. 119, 2089–2105. https://doi.org/10.1093/cvr/cvad059 (2023).
Liu, Y. et al. CHD7 interacts with BMP R-SMADs to epigenetically regulate cardiogenesis in mice. Hum. Mol. Genet. 23, 2145–2156. https://doi.org/10.1093/hmg/ddt610 (2014).
Yan, S. et al. CHD7 regulates cardiovascular development through ATP-dependent and -independent activities. Proc. Natl. Acad. Sci. U S A. 117, 28847–28858. https://doi.org/10.1073/pnas.2005222117 (2020).
Hsu, J. et al. CHD7 and Runx1 interaction provides a braking mechanism for hematopoietic differentiation. Proc. Natl. Acad. Sci. U S A. 117, 23626–23635. https://doi.org/10.1073/pnas.2003228117 (2020).
Rouillard, A. D. et al. The harmonizome: a collection of processed datasets gathered to serve and mine knowledge about genes and proteins. Database (Oxford) (2016). (2016) https://doi.org/10.1093/database/baw100
Liberzon, A. et al. The molecular signatures database (MSigDB) hallmark gene set collection. Cell. Syst. 1, 417–425. https://doi.org/10.1016/j.cels.2015.12.004 (2015).
Chen, E. Y. et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinform. 14, 128. https://doi.org/10.1186/1471-2105-14-128 (2013).
Johnson, J. L. et al. An atlas of substrate specificities for the human serine/threonine Kinome. Nature 613, 759–766. https://doi.org/10.1038/s41586-022-05575-3 (2023).
Kerkela, R. et al. Deletion of GSK-3beta in mice leads to hypertrophic cardiomyopathy secondary to cardiomyoblast hyperproliferation. J. Clin. Invest. 118, 3609–3618. https://doi.org/10.1172/JCI36245 (2008).
Weiss, C. S. et al. DYRK2 negatively regulates cardiomyocyte growth by mediating repressor function of GSK-3beta on eIF2Bepsilon. PLoS One. 8, e70848. https://doi.org/10.1371/journal.pone.0070848 (2013).
Cotecchia, S., Vescovo, D., Colella, C. D., Caso, M., Diviani, D. & S. & The alpha1-adrenergic receptors in cardiac hypertrophy: signaling mechanisms and functional implications. Cell. Signal. 27, 1984–1993. https://doi.org/10.1016/j.cellsig.2015.06.009 (2015).
Gregory, L. C. et al. Structural pituitary abnormalities associated with CHARGE syndrome. J. Clin. Endocrinol. Metab. 98, E737–743. https://doi.org/10.1210/jc.2012-3467 (2013).
Nykamp, K. et al. Sherloc: a comprehensive refinement of the ACMG-AMP variant classification criteria. Genet. Med. 19, 1105–1117. https://doi.org/10.1038/gim.2017.37 (2017).
Karousis, E. D. & Muhlemann, O. Nonsense-Mediated mRNA decay begins where translation ends. Cold Spring Harb Perspect. Biol. 11 https://doi.org/10.1101/cshperspect.a032862 (2019).
Hurd, E. A. et al. Loss of Chd7 function in gene-trapped reporter mice is embryonic lethal and associated with severe defects in multiple developing tissues. Mamm. Genome. 18, 94–104. https://doi.org/10.1007/s00335-006-0107-6 (2007).
Stender, J. D. et al. Genome-wide analysis of Estrogen receptor alpha DNA binding and tethering mechanisms identifies Runx1 as a novel tethering factor in receptor-mediated transcriptional activation. Mol. Cell. Biol. 30, 3943–3955. https://doi.org/10.1128/MCB.00118-10 (2010).
Chimge, N. O. & Frenkel, B. The RUNX family in breast cancer: relationships with Estrogen signaling. Oncogene 32, 2121–2130. https://doi.org/10.1038/onc.2012.328 (2013).
Pujantell, M. & Altfeld, M. Consequences of sex differences in type I IFN responses for the regulation of antiviral immunity. Front. Immunol. 13, 986840. https://doi.org/10.3389/fimmu.2022.986840 (2022).
Wong, M. T., Scholvinck, E. H., Lambeck, A. J. & van Ravenswaaij-Arts, C. M. CHARGE syndrome: a review of the immunological aspects. Eur. J. Hum. Genet. 23, 1451–1459. https://doi.org/10.1038/ejhg.2015.7 (2015).
Cadeddu Dessalvi, C. et al. Sex differences in anthracycline-induced cardiotoxicity: the benefits of estrogens. Heart Fail. Rev. 24, 915–925. https://doi.org/10.1007/s10741-019-09820-2 (2019).
McCarroll, C. S. et al. Runx1 deficiency protects against adverse cardiac remodeling after myocardial infarction. Circulation 137, 57–70. https://doi.org/10.1161/CIRCULATIONAHA.117.028911 (2018).
Gattenlohner, S. et al. NCAM(CD56) and RUNX1(AML1) are up-regulated in human ischemic cardiomyopathy and a rat model of chronic cardiac ischemia. Am. J. Pathol. 163, 1081–1090. https://doi.org/10.1016/S0002-9440(10)63467-0 (2003).
Qi, P., Zhai, Q. & Zhang, X. RUNX1 facilitates heart failure progression through regulating TGF-beta-induced cardiac remodeling. PeerJ 11, e16202. https://doi.org/10.7717/peerj.16202 (2023).
Liu, X. et al. Lineage-specific regulatory changes in hypertrophic cardiomyopathy unraveled by single-nucleus RNA-seq and Spatial transcriptomics. Cell. Discov. 9, 6. https://doi.org/10.1038/s41421-022-00490-3 (2023).
Kattih, B. et al. Single-nuclear transcriptome profiling identifies persistent fibroblast activation in hypertrophic and failing human hearts of patients with longstanding disease. Cardiovasc. Res. 119, 2550–2562. https://doi.org/10.1093/cvr/cvad140 (2023).
Meisner, J. K. & Martin, D. M. Congenital heart defects in CHARGE: the molecular role of CHD7 and effects on cardiac phenotype and clinical outcomes. Am. J. Med. Genet. C Semin Med. Genet. 184, 81–89. https://doi.org/10.1002/ajmg.c.31761 (2020).
Bouazoune, K. & Kingston, R. E. Chromatin remodeling by the CHD7 protein is impaired by mutations that cause human developmental disorders. Proc. Natl. Acad. Sci. U S A. 109, 19238–19243. https://doi.org/10.1073/pnas.1213825109 (2012).
Hsu, C. W. et al. High resolution imaging of mouse embryos and neonates with X-Ray Micro-Computed tomography. Curr. Protoc. Mouse Biol. 9, e63. https://doi.org/10.1002/cpmo.63 (2019).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21. https://doi.org/10.1093/bioinformatics/bts635 (2013).
Liao, Y., Smyth, G. K. & Shi, W. FeatureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930. https://doi.org/10.1093/bioinformatics/btt656 (2014).
Risso, D., Ngai, J., Speed, T. P. & Dudoit, S. Normalization of RNA-seq data using factor analysis of control genes or samples. Nat. Biotechnol. 32, 896–902. https://doi.org/10.1038/nbt.2931 (2014).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. EdgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140. https://doi.org/10.1093/bioinformatics/btp616 (2010).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. U S A. 102, 15545–15550. https://doi.org/10.1073/pnas.0506580102 (2005).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740. https://doi.org/10.1093/bioinformatics/btr260 (2011).
Goedhart, J. & Luijsterburg, M. S. VolcaNoseR is a web app for creating, exploring, labeling and sharing volcano plots. Sci. Rep. 10, 20560. https://doi.org/10.1038/s41598-020-76603-3 (2020).
Funding
This research was supported by grants from the 2020 CHARGE Syndrome Foundation Scientific Research Grant Program to IYP; the Optical Imaging & Vital Microscopy Core at Baylor College of Medicine and NIH UM1 HG006348 to C-WH; NIH R01 DC018404 to DMM; R01 CA203012, R35 CA231993, and P30 ES030285 to CLW.
Author information
Authors and Affiliations
Contributions
IYP and CLW conceived the overall concept. IYP, C-WH, and KB designed and conducted the primary experiments. CEE performed mouse breeding and sample collection. C-WH analyzed the micro-CT imaging. MHMA, CC, SG, and CLW analyzed RNA-sequencing data. CLW, IYP, and KB wrote the manuscript. JFM and DMM reviewed the data and the manuscript and offered feedback and suggestions.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Park, I.Y., Hsu, CW., Bouazoune, K. et al. A novel cardiomyopathy phenotype linked to a CHD7 missense variant. Sci Rep 15, 19429 (2025). https://doi.org/10.1038/s41598-025-00606-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-00606-1