Abstract
To explore the effect of PAX1 and JAM3 gene methylation on pathological upgrading before conization. A total of 549 patients who underwent colposcopy at our hospital were enrolled for analysis from December 2020 to April 2022. PAX1 and JAM3 gene methylation results in preoperative cervical exfoliated cells were collected. Univariate analysis and multivariate logistic regression analysis were conducted to identify the independent risk factors influencing the pathological upgrading of conization, aiming to establish a prediction model. A total of 88 patients were finally included for statistical analysis according to the inclusion and exclusion criteria. Based on univariate analysis and multivariate logistic regression analysis, ∆Ct PAX1 (P = 0.016, OR: 0.784, 95%CI 0.644–0.956) and cervical canal lesions (P = 0.048, OR: 3.469, 95%CI 1.014–11.870) were identified as independent risk factors for pathological upgrading for conization. Using the above results, we established a prediction model for pathological upgrading and plotted the receiver operator characteristic curve (ROC). The area under the curve (AUC) was calculated when the Youden index was maximized with an AUC value of 0.818 (95%CI 0.720–0.916), specificity of 94.4%, sensitivity of 60%. The cut-off value for ∆Ct PAX1 was determined as 4.34 when maximizing the Youden index. PAX1 could be a promising triage marker in predicting the pathological upgrading of CIN before conization. We found that if the ∆Ct PAX1 cut-off value is lower than 4.34, it is highly suggestive of pathological upgrading.
Similar content being viewed by others
Introduction
Cervical cancer is a common malignant tumor that seriously threatens women’s health worldwide. Its incidence is second only to breast cancer, and it ranks the fourth among cancer-related death factors in women1,2. In 2020, there were about 604 000 new cases and 342 000 deaths of cervical cancer worldwide, of which 88.1% of new cases and 91.4% of deaths occurred in low and middle-income countries1. In order to further reduce the global disease burden of cervical cancer, the World Health Organization (WHO) launched the global action to eliminate cervical cancer in 2018 and released the Global Strategy to accelerate the elimination of Cervical Cancer in 2020, which specified the phased goals of human papillomavirus (HPV) vaccination, cervical cancer screening and treatment by 2030. So far, it has received support from more than 194 countries and regions2.
In 2020, WHO classification of female genital tumours was updated from the original three-level classification of cervical intraepithelial neoplasia (CIN1, CIN2, CIN3) to a two-level classification that included low-grade squamous intraepithelial lesions (LSIL/CIN1) and high-grade squamous intraepithelial lesions (HSIL/CIN2 and CIN3)2. Persistent high-risk HPV(hr-HPV) infection is the cause of cervical cancer. It takes about 15–20 years for people with normal immune function to progress from HPV infection to cervical cancer. The natural progression of CIN leads to three directions: regression, persistence and progression. Among them, CIN3 has a higher risk of progressing to cancer. The model estimates that 15-23% of untreated CIN3 will develop into invasive cancer in an average of 13 years3, and the lifetime risk is 40%4. The risk of CIN2 was intermediate between CIN1 and CIN3. In CIN2, 50% of patients had spontaneous resolution within 2 years, 32% had persistence, and only 18% had progression. In young women under 30 years old, 60% regressed, 23% persisted and 11% progressed5. At present, the main treatment for CIN is cervical conization. Although cervical conization is a relatively safe and simple operation, it may still cause risks such as bleeding and infection. In addition, it may also change the original morphology of the cervix, causing cervical insufficiency, thus increasing the incidence of adverse events of pregnancy and delivery.
In patients with CIN confirmed by colposcopy directed biopsy(CDB), the pathological results of cervical conization are not completely consistent, which may upgrade to early invasive carcinoma or downgrade to LSIL, or even no lesions in the specimen. Literature reported that the probability of pathological upgrading after conization was 23.1%, and the probability of pathological downgrade was as high as 33.6%6. Underdiagnosis leads to multiple follow-up visits, and either delayed or progressed the lesion, exacerbating the potential harm to patients. Alternatively, overdiagnosis can result in unnecessary or premature treatment, especially in younger women, as inappropriate treatment significantly increases the risk of adverse outcomes in subsequent pregnancies7.
Therefore, how to screen patients with possible pathological upgrading by non-invasive examination is a clinical problem that needs to be solved.The PAX1 gene is located on chromosome 20p11.2, and it’s an essential tumor suppressor gene which plays a role in regulating cell maturation and differentiation. The expression of PAX1 is associated with embryogenesis, especially the development of the skeleton, thymus, and the parathyroid glands8,9.As an important methylation silencing gene in the occurrence and development of cervical cancer, the hypermethylation of PAX1 gene can be observed both in the stage of cervical cancer and precancerous lesions10. It has been reported that the disruption between kinases and phosphatases caused by PAX1 methylation is involved in cervical carcinogenesis11.An increasing number of studies have confirmed PAX1 methylation as a promising biomarker for cervical cancer based on its ability to discriminate between HSIL and normal tissues, resulting in a reduced necessity for colposcopy referral and biopsy12,13.
The JAM3 gene is located on chromosome 11q25. JAM-C, the production of JAM3 expression, is a protein at the tight junction between endothelial cells and polar epithelial cells, belonging to the immunoglobulin superfamily. JAM-C is expressed in epithelial cells, endothelial cells, natural killer cells, lymphocytes, etc. It is involved in many complex physiological and pathological processes, such as blood vessel formation, sperm formation, leukocyte migration and tumor metastasis. Recent studies have shown that the methylation level of JAM3 gene promoter is associated with the occurrence and development of cervical cancer and its precancerous lesions14,15,16.
The aim of this study is to investigate the correlation between various factors and pathological upgrading in patients with CIN by CDB who underwent cervical conization, and to provide a basis for clinical diagnosis and treatment.
Materials and methods
Participants and sample collection
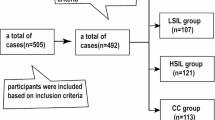
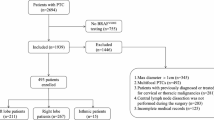
A total of 549 patients with PAX1 and JAM3 gene methylation results who underwent colposcopy in our hospital from December 2020 to April 2022 were collected for analysis. The exclusion criteria were as follows: (1) CDB revealed the presence of squamous cell cancer, adenocarcinoma in situ (AIS), or adenocarcinoma, (2) inability to undergo conization or undergo conization in other hospitals, (3) lost to follow-up, (4) patients that had a history of cervical disease treatment, hysterectomy, or chemoradiotherapy. Finally, 88 patients were enrolled for statistical analysis. The inclusion of patients was shown in Fig. 1. We informed patients of the research programs and obtained written consent before conization. Informed consent was obtained from all the patients. This study was approved by the Zhejiang Provincial People’s Hospital IRB on June 25th, 2024. Approval number: 2024(158). I confirm that all methods were performed in accordance with the relevant guidelines and regulations.
Flow chart showing selection of the study population.
∆Ct JAM3 and ∆Ct PAX1 levels of the upgrade and not-upgrade groups.The ∆Ct PAX1 and ∆Ct JAM3 level of not-upgrade group is significantly higher than that of upgrade group, with statistically significant differences(P < 0.01).
Clinical prediction models to distinguish pathological escalation of conization.The results of logistic regression analysis were included to establish a prediction model to distinguish the upgrading and downgrading of conization surgery.
ROC curve of the clinical prediction model to distinguish pathological escalation of conization. Based on the Cut-off value of ∆Ct PAX1, the area under the curve (AUC) of ∆Ct PAX1 high expression group and low expression group was 0.818 (95%CI 0.720–0.916), sensitivity: 94.4%, specificity: 60%.
Methylation detection methods
Methylation detection was using the “Human PAX1 and JAM3 gene methylation detection kit (Real-time PCR)” for Cervical Cancer [Class: III medical devices: approved by the National Medical Products Administration (No.20233400253)] with glyceraldehyde-3-phosphatedehydrogenase (GAPDH) as an internal control (OriginPoly Bio-Tec Co., Ltd., Beijing, China) by SLAN-96 S automatic medical- PCR- analysis system (Shanghai Hongshi Med- Tech Co., Ltd, Shanghai, China) per the manufacturer’s instructions. This kit uses multiple fluorescent PCR technology to detect the methylation status of the target PAX1 and JAM3 genes. The technology is based on the bisulfite conversion. DNA samples are treated with bisulfite, leading to the conversion of unmethylated cytosines into uracil, while methylated cytosines remain unchanged. Treated DNA following bisulfite conversion is called bisDNA. The hypermethylation level of the PAX1 gene and JAM3 gene was determined by the difference between the two Ct values (ACtPAX1 = CtPAX1-CtGAPDH and ACtJAM3-CtJAM3-CtGAPDH): According to the manufacturer’s instructions, the result of the PAX1/JAM3(+) is defined as ΔCtPAX1 < 6.6 orΔCtJAM3 < 10.0. Colposcopic impressions were made according to the American Society for Colposcopy and Cervical Pathology(ASCCP) standard, multiple biopsies targeting all areas with acetowhitening, metaplasia, or higher abnormalities are recommended. The knife is angled toward the endocervical canal and cuts deeper into the stroma, the depth of excision depends on the type of TZ according to 2011 Colposcopic Terminology of International Federation for Cervical Pathology and Colposcopy (IFCPC)17.
Pathological diagnosis of upgraded disease
Cervical biopsy and conization were histologically examined and classified according to the 2020 WHO classification of female genital tumours2.The highest pathological grade was taken as the final pathological diagnosis. Pathological upgrading of disease, which is shown as upgrade group, is defined as colposcopy confirmed CIN1 to conization confirmed CIN2\CIN3 or cervical cancer, CIN2 to conization confirmed CIN3 or cervical cancer, and colposcopy confirmed CIN3 to conization confirmed cervical cancer. The cervical lesions were diagnosed by two professional pathologists.
Statistical methods
SPSS (version 26) statistical software was used to analyze the data. For the measurement data, normality test was performed by using the mean ± standard deviation (X ̅±S). The count data were expressed by frequency and percentage, and the comparison between groups was performed by X^2 test. t test and X^2 test were used to screen out the risk factors related to the upgrading of conization. Logistic regression analysis was used to obtain the odds ratio (OR) and 95%Confidence Interval (95%CI) of the screened risk factors, and to determine the independent risk factors affecting the upgrading of conization.According to the results of logistic regression analysis, R studio (version 4.1.0) was used to construct the prediction model, and the receiver operator characteristic curve (ROC) was used to analyze the data. area under the curve (AUC) value, 95% confidence interval (CI), Youden index, sensitivity and specificity were evaluated. The Bootstrap method was used for verification, and the calibration chart was drawn. The performance of the model was evaluated by Hosmer-Lemeshow test, AUC and goodness of fit of the prediction model. Decision Curve Analysis (DCA) was used to verify the net clinical benefit rate of the prediction model. P < 0.05 was considered statistically significant.
Data availability
The data that support the findings of this study are available from the scientific research system of our institute but restrictions apply to the availability of these data, which were used under license for the current study, and so are not publicly available. Data are however available from the authors upon reasonable request and with permission of our institute. If someone wants to request the data from this study, the first author Xiaoyan Chen, whose E-mail is3090103449@zju.edu.cn, can be contacted.
Results
The ∆Ct PAX1 and ∆Ct JAM3 level of not-upgrade group is significantly higher than that of upgrade group, with statistically significant differences(P < 0.01) (Fig. 2).
Using t test, non-parametric test and X^2 test analysis, Age, height, weight, BMI, conization method, menopause, number of vaginal delivery, postcoital bleeding, mycoplasma infection, type of transformation zone, adequate colposcopy, biopsy conization interval, ∆Ct PAX1, ∆Ct JMA3, lesion/biopsy ratio, HPV infection and cervical canal lesions were included for analysis. Univariate results showed that ∆Ct PAX1, ∆Ct JMA3, cervical canal lesions and conization surgery were significantly improved (P < 0.05), while age, height, weight, BMI, conization method, menopause, number of vaginal delivery, postcosexual bleeding, mycoplasma infection, type of transformation zone, adequate colposcopy, interval between biopsy conization, lesion/biopsy ratio, and HPV infection were not associated with upgrading or downgrading of conization surgery (Table 1).
Logistic regression analysis showed that ∆Ct PAX1 (P = 0.016, OR: 0.784, 95%CI 0.644–0.956) and cervical canal lesions (P = 0.048, OR: 3.469, 95%CI 1.014–11.870) were independent risk factors for upgrading after conization. (Table 2)The results of logistic regression analysis were included to establish a prediction model to distinguish the upgrading and downgrading of conization surgery (Fig. 3).
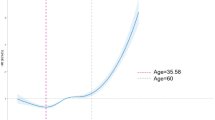
The ROC curve was established according to the results of logistic regression, and the results showed that the area under the curve (AUC) of ∆CtPAX1 and cervical canal lesions for predicting the upgrading and downstaging of conization was 0.818 (95%CI 0.720–0.916).When the Youden index was the maximum, the specificity was 94.4%, the sensitivity was 60.0%, and the discrimination was good. The Receiver Operating Characteristic curve (ROC curve) is one of the commonly used tools to evaluate the performance of binary classification models. It shows the performance of the model under different classification conditions through different threshold Settings, and the AUC value can help judge the merits of the model quickly. In this study, the AUC reached 0.818, indicating that the positive prediction model based on ΔCt PAX1 and ECC has a good ability to distinguish the escalation of conical pathology, and can play a good prediction role in clinical practice. When the Youden index was maximum, the cut-off value of ∆Ct PAX1 was 4.34 (P = 0.02, 95%CI 0.04–0.277, OR: 0.035). Based on the Cut-off value of ∆Ct PAX1, the area under the curve (AUC) of ∆Ct PAX1 high expression group and low expression group was 0.818, 95%CI 0.685–0.889, sensitivity: 94.4%, specificity: 60% (Fig. 4). And the curve of calibration is shown as Fig. 5. Patients with a ΔCt PAX1 value greater than 4.34 have a greater risk of pathological upgrading than those with a ΔCt PAX1 value less than 4.34, which requires special attention in clinical practice. The definition of ΔCt PAX1 cut-off value is more conducive to the clinical application of risk stratification.
Bootstrap was used to resample the original data set 1000 times to create the simulated data set. The calibration curve showed good agreement between the predicted discrimination rate and the actual discrimination rate of the prediction model, with an average absolute error of 0.031 (Fig. 5). DCA showed that the benefit of intervention was significantly higher than that of no intervention or full intervention across thresholds of 0-0.3, suggesting that the model has some clinical utility. (Fig. 6).
Curve of calibration.The calibration curve showed good agreement between the predicted discrimination rate and the actual discrimination rate of the prediction model, with an average absolute error of 0.031.
Clinical Model Prediction efficacy. DCA showed that the benefit of intervention was significantly higher than that of no intervention or full intervention across thresholds of 0-0.3, suggesting that the model has some clinical utility.
Discussion
Our study showed that 20.5% (18/88) of conization patients had pathological upgrading, and colposcopy evaluation alone had a certain risk of underdiagnosis of more advanced lesions. Using accurate tests to determine whether CIN have a tendency to progress is crucial for subsequent disease management.
DNA methylation, histone modification and non-coding RNA are the main mechanisms of epigenetic regulation18. During tumorigenesis, aberrant hypermethylation is usually observed on the promoters of genes involved in the prevention of tumor changes regulating key pathways such as cell cycle, DNA repair and apoptosis18. Hypermethylation of promoter CpG island sites can inhibit transcription by preventing the binding of transcription factors or changing the local binding position of chromatin, leading to gene silencing19. Therefore, changes in DNA methylation are considered as biomarkers for tumor or disease diagnosis20. Current studies mainly focus on the hypermethylation changes of tumor suppressor gene promoter CpG, and believe that the detection of specific DNA methylation sites can be used for early diagnosis, prognosis prediction and treatment evaluation of tumors21,22.
DNA methylation detection improves the specificity and sensitivity of cytology and HPV detection in cervical cancer screening, and has a good clinical application prospect in early diagnosis or prognosis prediction of cervical cancer. Clinical research experience has shown that DNA methylation detection technology has good clinical significance in the stratification management of hr-HPV positive women, the stratification management of women with abnormal cytology, and the screening of cervical cancer in women23,24. After infection of epithelial cells with hr- HPV, its DNA can be integrated into the cell genome. Because of the disorder of E2 gene expression, it cannot effectively inhibit the transcription of E6 and E7 genes, resulting in the continuous high expression of two oncoproteins, E6 and E7. In the process of transcription, E6 and E7 interact with various cellular proteins in differentiated epithelial cells, and genetic remodeling of viral chromatin including histone modification and DNA methylation level changes25, results in instability of chromosomal DNA synthesis, affecting cell differentiation cycle and inducing epithelial cell carcinogenesis26. Further intensified local hypermethylation of CpG-rich promoter regions also exacerbates tumor suppressor gene silencing and promotes cancer development27,28.
At present, DNA methylation of many specific genes has been found to be associated with the pathogenesis and progression of cervical cancer, such as PAX1、 JAM3、 EPB41L3、 ZNF582、 SOX1、FAM19A4 and miR124-2 and so on15,29,30,31. In 2008, Lai et al. first reported that abnormal methylation of PAX1(PAX1m) was associated with cervical cancer, and that the PAX1 gene was found to be silenced by hyper-methylation and under-expressed in cervical cancer biopsies32. In particular, the efficacy of PAX1m as a biomarker for the detection of CIN3 or worse (CIN3+) has been demonstrated in various studies12,32,33,34. It has been reported that the detection performance of CIN3 + in hrHPV positive women using PAX1 gene alone was 71.15-76.9%, and the specificity was 80.17-91.6%33,34. In a follow-up study of 452 hrHPV-positive women without a diagnosis of cervical cancer at baseline, the incidence of CIN3 + was significantly higher with PAX1m-positive or cytologic HSIL + than with PAX1m-negative or no cytologic HSIL+. This suggests that PAX1m detection is an effective molecular marker that can be used to predict disease progression33. PAX1m has a comparable clinical performance to cytology and better accuracy and specificity than HPV16/18 as a triage tool for detecting CIN3 + in women with hr-HPV33. PAX1m has also been reported to predict the efficacy of concurrent chemo-radiotherapy in cervical cancer35, and is a potential biomarker for monitoring the prognosis of cervical adenocarcinoma36. At the same time, PAX1 combined with other genes can increase the sensitivity and specificity of methylation in hrHPV positive triage, which is also the focus of current clinical research37,38,39. Liou et al. indicated that significant association was observed between CIN3 + and methylated PAX1 or ZNF582 in combination with HPV16/18 (OR:15.52, 95% CI7.73–31.18). The sensitivities and specificities of methylated PAX1 or ZNF582 combined with HPV16/18 for CIN3 + women were 89.2 and 76.0%, or 85.4 and 80.1%, respectively37.These studies suggest that detection of specific cervical cancer gene methylation can be used as a triage technique for hrHPV.
Shang Xiao et al. Indicates that the sensitivity and specificity of PAX1m/JAM3m testing for detecting CIN2 or more severe lesions (CIN2+) were 74.1% and 95.9%, respectively. The sensitivity and specificity of PAX1m/JAM3m testing for detecting CIN3 + were 87.6% and 86.8%, respectively. Receiver operating characteristic curve analysis showed that, for detecting CIN3+, the area under curve of PAX1m/JAM3m testing (0.872, 95%CI 0.847–0.897) was significantly superior to TCT testing (0.580, 95%CI 0.551–0.610) or hrHPV testing (0.503, 95%CI 0.479–0.515) (all P values < 0.05)40. Jing Fei et al. proposed that the PAX1/JAM3 test demonstrated demonstrated superior triage performance over cytology, even when used in combination with cytology, showing a high sensitivity of 89.0% (95% CI 82.9–95.1%) and specificity of 95.3% (95% CI 92.6–98.0%)41. We previously found that PAX1m/JAM3m testing shows promise as a reliable supplemental method to HPV-DNA testing for the triage of women with cytologic ASC-US. The methylation assay demonstrated a sensitivity of 83.8% and a specificity of 95.8%, outperforming HPV-DNA testing in differentiating high-grade cervical lesions among women with ASC-US. Moreover, PAX1m/JAM3m testing significantly reduced the colposcopy referral rate for further diagnostic procedures in high-risk HPV-positive women by 79.5%42.
However, few studies have evaluated PAX1m before conization, it has been previously reported that PAX1m would be a suitable alternative method to conventional options and it has the ability to predict the outcome of conization in CIN3 cases43. However, the role of PAX1m in predicting the pathological upgrading of CIN2 is unclear.
In the HPV-related study, the combination of PAX1 /ZNF582 and HPV16/18 testing reduced the number of colposcopy visits by 31.3%, thus providing a feasible follow-up solution in areas where colposcopy is not easily available35,36. In addition, JAM3 could significantly improve specificity and positive predictive value by regulating tumor cell growth through cell-cell adhesion and interaction. EPB41L3 /JAM3 methylation was validated clinically to have similar positive predictive value with hrHPV testing (0. 930 and 0. 954, respectively, P = 0.395). It can be used in combination with HPV screening or triage HPV positive patients15. The combined detection of PAX1 /ZNF582 and PAX1 /JAM3 methylation and HPV16/18 can be used as a reference for further colposcopy screening to improve the accuracy of cervical lesions screening and avoid repeated colposcopy and cancer panic in women.
Our study showed that PAX1 and cervical canal lesions were independent risk factors for pathological escalation after conization. We established a prediction model for pathological upgrading and plotted the ROC. When the Youden index was the largest, the AUC was calculated, which was 0.818 (95%CI 0.720–0.916), the specificity was 94.4%, the sensitivity was 60.0%. The ∆Ct PAX1 cut-off value was taken to be 4.34 when the Youden index was maximum. PAX1 could be a promising triage marker in predicting the pathological upgrading of CIN before conization. If biopsy pathology indicates CIN with a ∆Ct PAX1 less than 4.34, then cervical conization is inevitable because of the increased risk of pathological upgrading. On the other hand, women with ∆Ct PAX1 more than 4.34 do not need immediate conization because of there being a relatively low short-term progression risk. For young women who have fertility requirements, this approach seems to be particularly important, since only hypermethylated lesions require treatment and the risk of preterm abortion due to treatment could be reduced.
This study has several limitations. First, the sample size was not large enough and a larger longitudinal study is necessary to validate the natural history of CIN progression in relation to DNA methylation. Only 88 patients were eligible for enrollment in this study, so the sample size was small. Although the sample size was small, in our statistical analysis, we still found the value of methylation detection for the pathological escalation of cervical precancerous lesions, so we believe it is necessary to present our research results to provide necessary references for clinical diagnosis and treatment. Secondly, it’s a retrospective study and our research has not reached the follow-up endpoint, residual and recurrence of lesions have not been discussed here, and continued follow-up is needed in the future. We have already performed methylation tests in the real world and look forward to further validation of our findings with additional cases. In addition, further studies are needed to explore PAX1m levels after treatment and to compare PAX1m changes before and after conization. At last, further studies are also needed to determine the ideal interval of monitoring using PAX1m to avoid underdiagnosis and overdiagnosis. While the biological Mechanisms are still need to be elaborated on how PAX1 methylation may drive pathological upstaging.
Based on current clinical studies, methylation detection of many genes may have clinical value for cervical cancer screening or pathological progression and degradation of cervical conectomy. At present, we have only preliminarily investigated the PAX1 and JAM3 genes, and more clinical studies can be conducted to confirm the clinical value of the genes reported in the literature. As of October 2024, China has authorized more than 50 patents related to cervical cancer DNA methylation detection, and more than 30 kits using methylation detection technology have been approved, of which 4 have been applied to cervical cancer DNA methylation detection. Methylation gene markers include ASTN1, DLX1, ITGA4, RXFP3, SOX17, ZNF671, SOX1 and other genes21.
In this exploratory study, we found that PAX1 could be a promising triage marker in predicting the pathological upgrading of CIN before conization. We found that if the ∆Ct PAX1 cut-off value is lower than 4.34, it is highly suggestive of pathological upgrading. Using PAX1m as a monitoring tool could help prevent inappropriate conservative observation or ablation therapy.Further validation and prospective clinical trials are needed to confirm these findings in the future.
Data availability
The data that support the findings of this study are available from the scientific research system of our institute but restrictions apply to the availability of these data, which were used under license for the current study, and so are not publicly available. Data are however available from the authors upon reasonable request and with permission of our institute. If someone wants to request the data from this study, the first author Xiaoyan Chen, whose E-mail is 3090103449@zju.edu.cn, can be contacted.
References
Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71(3), 209–249 (2021).
WHO Guideline for Screening and Treatment of Cervical pre-cancer Lesions for Cervical cancer Prevention (Geneva, 2021).
Gustafsson, L. & Adami, H. O. Natural history of cervical neoplasia: Consistent results obtained by an identification technique. Br. J. Cancer. 60(1), 132–141 (1989).
Peto, J. et al. The cervical cancer epidemic that screening has prevented in the UK. Lancet 364(9430), 249–256 (2004).
Tainio, K. et al. Clinical course of untreated cervical intraepithelial neoplasia grade 2 under active surveillance: systematic review and meta-analysis. BMJ 360, k499 (2018).
Jung, Y. et al. Clinical factors that affect diagnostic discrepancy between colposcopically directed biopsies and loop electrosurgical excision procedure conization of the uterine cervix. Obstet. Gynecol. Sci. 61(4), 477–488 (2018).
Kyrgiou, M. et al. Obstetric outcomes after conservative treatment for cervical intraepithelial lesions and early invasive disease. Cochrane Database Syst. Rev. 11(11), CD012847 (2017).
Paixao-Cortes, V. R., Salzano, F. M. & Bortolini, M. C. Origins and evolvability of the PAX family. Semin Cell. Dev. Biol. 44, 64–74 (2015).
Yamazaki, Y. et al. PAX1 is essential for development and function of the human thymus. Sci. Immunol. 5(44) (2020).
Da Silva, M. L. R. et al. The role of HPV-induced epigenetic changes in cervical carcinogenesis (review). Biomed. Rep. 15(1), 60 (2021).
Su, P. H. et al. Paired box-1 (PAX1) activates multiple phosphatases and inhibits kinase cascades in cervical cancer. Sci. Rep. 9(1), 9195 (2019).
Luan, T. et al. PAX1 methylation as a potential biomarker to predict the progression of cervical intraepithelial neoplasia: A meta-analysis of related studies. Int. J. Gynecol. Cancer. 27(7), 1480–1488 (2017).
Fang, C. et al. The promising role of PAX1 (aliases: HUP48, OFC2) gene methylation in cancer screening. Mol. Genet. Genomic Med. 7(3), e506 (2019).
Boers, A. et al. Discovery of new methylation markers to improve screening for cervical intraepithelial neoplasia grade 2/3. Clin. Epigenet. 8, 29 (2016).
Kong, L. et al. DNA methylation for cervical cancer screening: A training set in China. Clin. Epigenetics 12(1), 91 (2020).
Boers, A. et al. DNA methylation analysis in self-sampled brush material as a triage test in hrHPV-positive women. Br. J. Cancer. 111(6), 1095–1101 (2014).
Bornstein, J. et al. 2011 Colposcopic terminology of the international federation for cervical pathology and colposcopy. Obstet. Gynecol. 120(1), 166–172 (2012).
Urbano, A. et al. Gene-Specific targeting of DNA methylation in the mammalian genome. Cancers (Basel) 11(10) (2019).
Smith, Z. D. & Meissner, A. DNA methylation: Roles in mammalian development. Nat. Rev. Genet. 14 (3), 204–220 (2013).
Koch, A. et al. Analysis of DNA methylation in cancer: Location revisited. Nat. Rev. Clin. Oncol. 15(7), 459–466 (2018).
Lu, Q., Ma, D. & Zhao, S. DNA methylation changes in cervical cancers. Methods Mol. Biol. 863, 155–176 (2012).
Kremer, W. W. et al. Clinical regression of High-Grade cervical intraepithelial neoplasia is associated with absence of FAM19A4/miR124-2 DNA methylation (CONCERVE Study). J. Clin. Oncol. 40(26), 3037–3046 (2022).
Zhu, H. et al. DNA methylation and hydroxymethylation in cervical cancer: Diagnosis, prognosis and treatment. Front. Genet. 11, 347 (2020).
Burley, M., Roberts, S. & Parish, J. L. Epigenetic regulation of human papillomavirus transcription in the productive virus life cycle. Semin Immunopathol. 42(2), 159–171 (2020).
Ehrlich, M. DNA hypomethylation in cancer cells. Epigenomics 1(2), 239–259 (2009).
Durzynska, J., Lesniewicz, K. & Poreba, E. Human papillomaviruses in epigenetic regulations. Mutat. Res. Rev. Mutat. Res. 772, 36–50 (2017).
Steenbergen, R. D. et al. Clinical implications of (epi)genetic changes in HPV-induced cervical precancerous lesions. Nat. Rev. Cancer 14(6), 395–405 (2014).
Hsu, Y. W. et al. Genotype-specific methylation of HPV in cervical intraepithelial neoplasia. J. Gynecol. Oncol. 28 (4), e56 (2017).
Szalmas, A. & Konya, J. Epigenetic alterations in cervical carcinogenesis. Semin Cancer Biol. 19(3), 144–152 (2009).
Zhao, J. et al. SOX14 hypermethylation as a tumour biomarker in cervical cancer. BMC Cancer. 21(1), 675 (2021).
Lai, H. C. et al. Identification of novel DNA methylation markers in cervical cancer. Int. J. Cancer. 123(1), 161–167 (2008).
Chang, C. L. et al. DNA methylation marker for the triage of HrHPV positive women in cervical cancer screening: Real-world evidence in Taiwan. Gynecol. Oncol. 161(2), 429–435 (2021).
Tian, Y. et al. Utility of gene methylation analysis, cytological examination, and HPV-16/18 genotyping in triage of high-risk human papilloma virus-positive women. Oncotarget 8 (37), 62274–62285 (2017).
Li, X. et al. Methylation of PAX1 gene promoter in the prediction of concurrent chemo-radiotherapy efficacy in cervical cancer. J. Cancer. 12(17), 5136–5143 (2021).
Zhao, Z. et al. SOX1 and PAX1 are hypermethylated in cervical adenocarcinoma and associated with better prognosis. Biomed. Res. Int. 2020, 3981529 (2020).
Liou, Y. L. et al. Combined clinical and genetic testing algorithm for cervical cancer diagnosis. Clin. Epigenetics. 8, 66 (2016).
Kremer, W. W. et al. The use of host cell DNA methylation analysis in the detection and management of women with advanced cervical intraepithelial neoplasia: A review. BJOG 128(3), 504–514 (2021).
Zhang, L. et al. Detection of host cell Gene/HPV DNA methylation markers: A promising triage approach for cervical cancer. Front. Oncol. 12, 831949 (2022).
Fu, K. et al. Triage by PAX1 and ZNF582 methylation in women with cervical intraepithelial neoplasia grade 3: A multicenter case-control study. Open. Forum Infect. Dis. 9(5), ofac013 (2022).
Xiao, S. et al. Multicenter study on PAX1/JAM3 dual gene methylation detection in cervical cytology for cervical cancer screening. Chin. Med. J. 104(20), 1852–1859 (2024)
Fei, J. et al. Evaluating PAX1/JAM3 methylation for triage in HPV 16/18-infected women. Clin. Epigenet. 16, 190 (2024).
Chen, X. et al. Cervical cancer screening: Efficacy of PAX1 and JAM3 methylation assay in the triage of atypical squamous cell of undetermined significance (ASC-US). BMC Cancer 24, 1385 (2024).
Acknowledgements
This work was partly supported by Medical Science and Technology Foundation of Zhejiang Province (2020KY020, 2023KY013).
Author information
Authors and Affiliations
Contributions
Data collection, study conception, and study design was performed by X.C. Personnel organization, data collection, and the conization procedure was performed by H.X. and L.Z. Sample collection was performed by H.J. Supervision of the research program and manuscript review and guidance during the study was provided by H.S. The first draft of the manuscript was written by X.C. and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chen, X., Xu, H., Zhao, L. et al. The role of PAX1 and JAM3 methylation in predicting the pathological upgrading of cervical intraepithelial neoplasia before conization. Sci Rep 15, 17684 (2025). https://doi.org/10.1038/s41598-025-01422-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-01422-3