Abstract
Glioblastomas (GBM) are the most prevalent primary brain tumors, affecting 5 in every 100,000 people. GBMs optimize proliferation through adaptive cellular metabolism, frequently exploiting the Warburg effect by increasing aerobic glycolysis and glucose utilization to facilitate rapid cell growth. This disproportionate reliance on glucose has driven interest in using the ketogenic diet (KD) as a treatment for GBM. In this study, we explored metabolic flux in three primary human GBM cell samples using a media simulating a KD. Flux analysis using a detailed metabolic modeling approach revealed three unique metabolic phenotypes in the patient GBMs that correlated with cell viability. Notably, these phenotypes are apparent in the flux modeling, but were not evidenced by changes in the metabolite pool sizes. This variability in metabolic flux may underlie the inconsistent results observed in preclinical and clinical studies using the KD as a treatment paradigm.
Similar content being viewed by others
Introduction
Glioblastoma multiforme (GBM) is an exceptionally dangerous grade IV glioma, with the 5-year survival rate being less than 5%1. Aside from the extremely aggressive nature of glioblastoma, contributions to the high mortality rate include late diagnosis, a shortage of effective treatment options, and a limited ability to measure treatment response2,3. The high mortality rate is also attributed to limited treatment methods that can effectively target divergent cellular profiles characterizing GBM. GBM is typically diagnosed and monitored through clinical imaging methods, primarily magnetic resonance imaging (MRI) and computed tomography (CT). Like other forms of cancer, GBM is known to display differential metabolism compared to standard brain tissue4. It is well recognized that glioblastomas demonstrate the Warburg effect and thus are typically more glycolytic than surrounding tissue5. The Warburg effect describes a metabolic phenotype defined by increased glucose utilization to support cell growth and proliferation6. However, glucose metabolism can vary widely among individual GBMs7.
The ketogenic diet (KD) focuses on limiting carbohydrate intake while supplementing fat. It has been suggested as an intervention in GBM management, as the KD should restrict availability of glycolytically derived ATP, as well as flux through multiple pathways used for cell growth. This includes the pentose phosphate pathway and one-carbon metabolism through the serine biosynthesis pathway8. While normal brain cells also use glucose for energy metabolism, carbohydrate derived acetyl-CoA can be supplemented by ketone oxidation, namely β-hydroxybutyrate and acetoacetate. Many tumors including gliomas, however, have reduced activity of enzymes needed to metabolize ketones, such as succinyl-CoA: 3-ketoacid CoA transferase (OXCT1)9. Thus, there is a possibility that tumors may not be able to effectively utilize ketones for energy and that tumor growth may be subsequently restricted in ketogenic conditions.
There have been multiple preliminary animal studies on ketogenic diet and glioblastoma that highlight differences in the glioblastoma response to ketogenic conditions. One study showed a decrease in the rate of tumor growth and an increase in the survival time of mice when using a highly glycolytic U87M glioblastoma model10. This study combined a ketogenic diet with a standard chemotherapeutic agent bevacizumab to further reduce the supply of glucose to the tumor. Although the ketogenic diet al.one did not have a survival benefit, the combination of this and bevacizumab demonstrated significantly improved survival. This study examined global metabolic changes using standard metabolomics approaches. Metabolite pool sizes for most amino acids were similar between tumor and brain tissues. However, without any isotopic tracing to gain insight into metabolic flux, these results have limited interpretation. Another study examined in vivo metabolism of 13C-β-hydroxybutyrate (BHB) oxidation in two rat glioma models, RG2 and 9 L11. The authors observed low uptake and metabolism of BHB in vitro in the RG2 cells, indicated by C-labeling of glutamate C4 in cell extractsC fractional enrichment: 7.9%±0.5%). In the 9 L model, uptake of BHB was not detectable and no substantialC fractional enrichment was detected above the 1.1% natural abundance level (1.9%±0.9%). The divergent responses between RG2 and 9 L models suggest phenotypic differences between the cell lines.
Additionally, there have been over 13 clinical trials that examined the application of the ketogenic diet to high-grade glioma treatment12. One study analyzed 53 patients with six receiving the ketogenic diet13. They concluded that the diet was well tolerated by the patients but the study did not have enough patients enrolled to conclude any survival benefit. Another clinical trial noted a longer trend in progression-free survival in patients in ketosis but concluded that this likely does not hold a significant therapeutic advantage as a monotherapy14. The authors suggest the diet is safe and a clinical trial assessing the ketogenic diet in conjunction with standard chemotherapy or radiotherapy is necessary. These studies demonstrate the clinical interest in the ketogenic diet but are limited in scope, report inconsistent results with respect to a survival benefit for glioblastoma patients, and often failed to measure consistent metabolic profiles in the tumor. A recent review on ketogenic metabolic therapy highlighted these limitations and introduced a clinical research framework to guide future translational work, emphasizing the need to understand metabolic adaptations to meet ATP synthesis requirements in the tumor12.
We used [2H7]glucose to evaluate changes in glycolytic and central carbon metabolism in primary glioblastoma cells from three patients with and without ketogenic conditions. We performed a detailed metabolic flux analysis after examining the fractional enrichment in downstream metabolites by mass spectrometry. This approach permits a deep assessment of metabolic phenotypes across cell lines. Furthermore, we assessed OXCT1 expression in all cell lines under standard and ketogenic conditions. The three patient-derived cell lines display nearly orthogonal metabolic flux phenotypes, which likely explains the varying success of the ketogenic diet in preclinical and clinical studies. These results suggest establishing the specific metabolic phenotype of each patient tumor might be highly advantageous for predicting the efficacy of a ketogenic therapy.
Results
Cell viability and HDO production indicated differences in the metabolic response to the ketogenic media
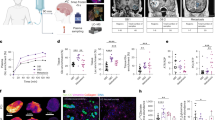
Three patient-derived primary glioblastoma cell lines (CA7, CA3, L2) displayed divergent effects of the ketogenic condition on central carbon metabolism (Fig. 1). Cell viability was analyzed to establish whether exposure to the ketogenic diet would impact cell survival. In both CA7 and CA3 cells, the cell viability was significantly lower in cells grown in the ketogenic media compared to the standard media (Fig. 1b). Noticeably, CA7 cells exposed to the ketogenic media had the lowest viability of all cell lines. L2 cells, on the other hand, displayed no difference in cell viability based on substrate availability. Oxygen consumption measurements were taken in the cell culture to determine whether media contents impacted overall metabolic health. In all three cell lines, there was no difference in oxygen consumption (µmol/(min×million cells)) between cells placed in the ketogenic and standard medias (Fig. 1c). This suggests that the baseline respiratory chain activity is similar across the tested cells and can be used for further comparative analysis. During glycolysis, water is generated from the protons attached to glucose at several points. If these protons are deuterated as they are in [2H7]glucose, partially deuterated water (HO2H or HDO) is produced. Intracellular HDO production in the cell pellet was obtained by 2H NMR analysis to quantify the incorporation of the deuterium into water during cellular metabolism (Fig. 1d and e). The analysis revealed that exposure to the ketogenic diet significantly reduced intracellular levels of HDO production from the [2H7]glucose in the CA7 cell line, indicating lower levels of glucose utilization15,16 (Fig. 1f). There was a 45% decrease of intracellular HDO in the ketogenic condition compared to the standard media in the CA7 cells, compared to 27% in CA3 and 2% in L2 (Fig. 1g).
Primary glioblastoma samples display differences in cell viability and HDO production after exposure to ketogenic media conditions. (a) Experimental design of isotopic tracing and analysis of GBM neurospheres created with Biorender. (b) Cell viability (n = 4) and (c) oxygen consumption (n = 3) measurements of three GBM samples after incubation in standard and ketogenic media. (d) Representative 2H nuclear magnetic resonance spectra of the lysed cell pellet with clear peaks corresponding to HDO and the internal standard [2H4]pyrazine. (e) Overlay of representative HDO peaks from each cell line after standard (S) and ketogenic (K) media. (f) HDO production by GBM neurospheres (n = 3–4/group). (g) Comparison of average HDO production between standard and ketogenic media samples within each GBM cell line. Data are represented as mean ± SD. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001 by two-way ANOVA (b and f) or Student’s t test (c).
Ketogenic media induced minimal changes in metabolite pool sizes
To examine changes in overall metabolite abundance, total pool sizes of metabolites (ng metabolite/million cells) were quantified in CA3 and CA7 cells. Figure 2 displays a heatmap (2a), a principal components analysis (2b), and a Venn diagram of important features identified by a two-way ANOVA analysis (2c). The PCA scores plot displays strong separation between the CA3 and CA7 cell lines, but minimal differences between the standard and ketogenic experimental groups within each cell line. Furthermore, the Venn diagram demonstrates significant differences in pool sizes can be primarily attributed to the cell line rather than the treatment group. Pool sizes of each metabolite are presented in Fig. 2d-r. Only CA3 cells demonstrated sensitivity to media conditions in terms of pool sizes, with decreased abundances of lactate, fumarate, malate, and glutamine after exposure to the ketogenic condition compared to the standard media. Pool sizes were universally unchanged by the ketogenic media in CA7 cells. The PCA scores plot emphasizes the cell line as the primary source of variability across conditions.
Metabolite pool sizes are driven by individual GBM rather than standard or ketogenic media. (a) Heatmap displaying pool sizes in CA3 and CA7 in both media conditions. (b) Principal components analysis (PCA) plot with minimal separation based on media, but strong separation by cell line. (c) Interaction effect examining Cell Line and Media Treatment based on two-way ANOVA that shows differences in pool sizes are based entirely on GBM cell line rather than media group. (a-c) were generated using Metaboanalyst software. (d–f) Pool size quantification of individual metabolites in CA3 and CA7 GBM cells after exposure to standard or ketogenic media. A significant decrease was found in CA3 cells in (d) lactate, (h) fumarate, (i) malate, and (l) glutamine after the ketogenic media compared to the standard media. Data are represented as mean ± SD. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001 by two-way ANOVA.
Despite minimal differences in metabolite pool sizes, the intracellular lactate:alanine ratio was significantly decreased in CA3 cells in the ketogenic condition compared to the standard media, with no difference in CA7 (Supplementary Fig. 1). Lactate:pyruvate is a common readout of the redox state of the cytosol due to its relation to the [NADH]/[NAD+] pool17. Alanine was used instead of pyruvate as pyruvate concentrations were too low to be accurately measured, and the pyruvate/alanine equilibrium is not redox dependent. The observed decrease in this ratio indicates the redox balance is negatively affected in the CA3 cells in the ketogenic media18.
Deuterium incorporation From [2H7]glucose varied between cell lines and conditions
Isotopologue distributions for a panel of central carbon metabolites were assessed to evaluate 2H incorporation from [2H7]glucose in both standard and ketogenic media conditions. In CA3 cells, exposure to the keto media before [2H7] glucose exposure increased 2H enrichment in several metabolites, such as lactate, alanine, and malate, compared to cells exposed to the standard media. Other metabolites displayed decreased 2H enrichment based on exposure to ketogenic conditions, such as in glutamate, succinate, GABA, and 2-OH-glutarate (Fig. 3, Supplementary Fig. 2). In CA7 cells, the keto media condition increased 2H enrichment in lactate, alanine, and malate. However, unlike CA3, there were similar fractional enrichment patterns across treatment groups in glutamate, succinate, and 2-OH-glutarate (Supplementary Fig. 3). L2 displayed very little isotopic enrichment in either treatment group (Supplementary Fig. 4). Furthermore, the only statistically significant difference found between the media conditions in the L2 cells was increased H incorporation in glutamate and glutamine in the ketogenic conditions. Ultimately, analysis of fractional enrichment on downstream metabolites revealed differential glucose utilization based on both the choice of standard or ketogenic media and the individual GBM cell lines.
Isotopologue distributions in central carbon metabolites in CA3 cells after standard and ketogenic media conditions. Statistically significant differences in fractional enrichment between standard and ketogenic conditions were found in (a) lactate, (b) alanine, (d) succinate, (f) malate, (h) N-acetylaspartate (NAA), (j) glutamate, (k) 2-OH-glutarate, and (l) GABA. Other measured metabolites that did not display differences in enrichment between media conditions are (c) citrate, (e) fumarate, (g) aspartate, (i) glutamine, and (m) β-hydroxybutryate (BHB). Data are represented as mean ± SD. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001 by Student’s t test.
Metabolic flux analysis reveals distinct metabolic phenotypes
To further investigate metabolic differences between the glioblastoma cell lines, a model of central carbon metabolism was developed using the software Isotopomer Network Compartment Analysis (INCA)19 to evaluate metabolic flux. This software generates estimates of absolute enzyme flux based on isotopic fractional enrichment. A map of metabolic pathways of interest, including glycolysis, the TCA cycle, the GABA shunt, and BHB metabolism, representing the INCA model is displayed in Fig. 4a with the exact equations reported in Supplementary Table 3. The metabolic flux analysis using fractional enrichment data provided further insight into the different metabolic phenotypes (Fig. 4b-m). Notably, the metabolic model included reactions relevant to pyruvate cycling20,21 and the malic enzyme22,23 as these pathways have been previously implicated in glioblastoma metabolism and tumorigenesis. Based on the metabolic flux analysis, there was a decrease in pyruvate dehydrogenase (PDH) flux in CA3 cells in response to the ketogenic media. After exposure to the ketogenic media, CA3 switched the direction of glutamate dehydrogenase flux such that α-ketoglutarate was used as a source for glutamate rather than glutamine anaplerosis fueling the TCA cycle (Fig. 4k)24. GABA was produced from this glutamate and did not reenter the TCA cycle through succinate (Fig. 4l). Utilization of GABA through the GABA shunt appeared to occur solely in L2 cells (Fig. 4m).
The CA7 cells displayed a pattern of increased flux through glycolytic enzymes under ketogenic conditions, with statistically significant increases in enolase and pyruvate kinase. However, PDH flux was significantly decreased with a corresponding increase in labeled lactate production. Flux throughout the TCA cycle, namely succinate dehydrogenase and malate dehydrogenase, were significantly decreased in ketogenic media compared to the standard media, and anaplerosis/cataplerosis (malic enzyme) was also significantly decreased. Flux through glutamate dehydrogenase and glutamate decarboxylase were quite low in both media conditions. The L2 cells displayed no significant changes in metabolic flux in the ketogenic condition compared to the standard media. However, the modeling data indicates increased flux through GABA transaminase and succinate semialdehyde dehydrogenase in this cell line compared to CA3 and CA7, suggesting imported GABA may be an important fuel source for L2.
Metabolic flux analysis reveals differences in enzyme activity among GBMs in response to ketogenic media. (a) Reactions present in the metabolic model, with a complete list in Supplementary Table 3. Created with Biorender. Specific enzymatic steps were subjected to statistical analysis including those involved in (b-d) glycolysis, (e-f) pyruvate utilization, (h-j) the Krebs cycle, and (k-m) the GABA shunt. Data are represented as mean ± SD. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001 by two-way ANOVA.
Patterns in metabolic flux were also analyzed using the multivariate statistical approach previously applied to the metabolite pool sizes (Fig. 5). Significant separation of all groups (p = 0.001) occurred in the principal components analysis (Fig. 5a), which indicates the metabolic flux profiles differed between experimental groups. The Venn diagram from the Two-way ANOVA indicates significant variations in metabolic flux can be attributed from the cell line and the interaction between the cell line and media treatment group. The correlation matrix displayed strong correlations between related enzymes in some metabolic pathways (Fig. 5c), For example, metabolic flux through individual enzymes involved in glycolysis was highly correlated with each other, namely hexokinase, aldolase, glyceraldehyde 3-phosphate dehydrogenase, enolase, and pyruvate kinase. This also occurred in the TCA cycle, where flux through pyruvate dehydrogenase was positively correlated with α-ketoglutarate dehydrogenase, fumarase, and succinate dehydrogenase flux.
Metabolic flux is altered by ketogenic media as well as cell line. (a) PCA plot of metabolic fluxes in CA3, CA7, and L2 cells in both standard and keto media conditions displays clear separation of all groups. (b) Venn diagram displaying the interaction effect from two-way ANOVA shows differences in flux arise from the cell line and the interaction between the cell line and media treatment group. (c) Correlation matrix of metabolic flux through enzymes display strong correlations among enzymes in the same metabolic pathway. Figures were generated using Metaboanalyst software.
OXCT1 expression was decreased in CA7
The KD diet logically should cause changes in OXCT1 expression, which prompted a study of the relative enzyme levels. Immunofluorescence studies were performed on the neurospheres after the isotopic tracing study to identify differences in expression level of OXCT1 (Fig. 6a). L2 displayed significantly less OXCT1 than both CA3 and CA7 cell lines in the standard media conditions (Fig. 6b). OXCT1 expression in the CA7 cells was downregulated in the ketogenic condition but was unchanged by KD in CA3 or L2 (Fig. 6c). PDH expression was also examined in CA3 and CA7 cells due to the decreased flux found in the metabolic modeling (Supplementary Fig. 5). However, there were no significant differences in PDH expression in either cell line between the standard and ketogenic conditions.
OXCT1 expression is decreased by ketogenic conditions in CA7 GBM neurospheres. (a) Representative images of OXCT1 and DAPI staining in neurospheres from each GBM. (b) Statistical analysis of corrected OXCT1 expression between different GBM reveal decreased OXCT1 expression in L2 relative to other GBMs. (c) A comparison of OXCT1 expression within each GBM in response to ketogenic conditions indicates only CA7 neurospheres alter OXCT1 expression based on media condition. Data are represented as mean ± SD. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001 by two-way ANOVA.
Discussion
The heterogeneity of the metabolic adaptation of the three patient cells was further established in the analysis of 2H incorporation in downstream metabolites. Each cell line displayed a unique flux phenotype that appears to correspond to the ability to survive in ketogenic conditions. Both CA3 and CA7 displayed increased enrichment in the labeled lactate and alanine isotopologues under KD. However, isotopic labeling in TCA cycle and glutamate-derived metabolites differed between CA3 and CA7. CA3 cells displayed a switch in glutamate metabolism in the KD condition, favoring net glutamate synthesis from α-ketoglutarate. This increase in glutamate synthesis was coupled with an increase in glutamate decarboxylase activity. In both media conditions, glutamine synthase and glutamate dehydrogenase flux in CA3 was much more substantial than in CA7 and L2. Based on previous work on CA3 (also known as BT53),26 it has been established that these cells display mutations in the epidermal growth factor receptor (EGFR). However, the specific EGFR variant III (vIII) mutation was not detected in this cell line. EGFR mutations have been shown to preserve cell proliferation and promote glutamine metabolism in a glutamate dehydrogenase-dependent manner in glioblastoma27. This is supported by a statistically significant decrease in glutamine concentrations in CA3 after placement in ketogenic conditions compared to standard media (Fig. 2l). Additionally, these metabolic effects are independent of glycolytic metabolism. Coupled with the apparent decrease in flux through glycolytic enzymes, CA3 does not show significant utilization of the Warburg effect under ketogenic conditions. Ultimately, the metabolic adaptations to the ketogenic media suggest measuring flux in future studies will be essential in understanding the effects of the ketogenic diet.
The ketogenic condition had a devastating effect on the CA7 cell line, which did not appear to implement any effective metabolic adaptations. Flux through the glycolytic enzymes enolase and pyruvate kinase were significantly upregulated. However, PDH flux was downregulated, so the pyruvate generated did not enter the TCA cycle. Instead of being oxidized to acetyl-CoA, the excess labeled pyruvate generated labeled lactate. Of note, the CA7 cell line (previously known as BT206) does not have any mutations in EGFR. These results indicate that CA7 is a highly glycolytic glioblastoma that is unable to survive when deprived of glucose. The increase in flux through glycolytic enzymes and increase in labeled lactate in the CA7 cell line suggests pronounced Warburg metabolism that is enhanced under ketogenic conditions. The commonly used U87M GBM cell line is also highly glycolytic, which may explain why the ketogenic diet limits growth for this tumor as well.
In contrast, the L2 cell line, which exhibits EGFR overexpression along with a single non-synonymous point mutation that substitutes arginine with lysine—a mutation reported in lung cancer but not commonly found in GBM—appears to be unaffected by the ketogenic diet. 2H enrichment in the downstream metabolites analyzed is low in both the standard and ketogenic media conditions, and there are no significant differences in metabolic flux between the two conditions. Thus, the L2 cell line does not appear to display a highly glycolytic (Warburg) phenotype. It is likely that glucose is used by these cells for glycogen synthesis or the pentose phosphate pathway, since downstream 2H labeling was preserved. Metabolites pertaining to these pathways were not observed in this analysis. Given this limitation, L2 was excluded in the assessment of pool size differences.
A multivariate statistical analysis of metabolite abundances that considered both the cell line and media condition revealed that differences in pool sizes were primarily driven by cell line rather than the ketogenic condition (Fig. 2a-c). Given the stark changes in fractional enrichment and metabolic fluxes, a lack of changes in pool sizes is surprising. However, both the PCA plot and Venn diagram of interaction effects from the Two-way ANOVA (Fig. 5a, b) demonstrate the presence of unique metabolic flux phenotypes arising from the ketogenic media treatment in all cell lines. These phenotypes were not evident using only the pool sizes. Furthermore, the metabolic flux analysis performed here was internally consistent. The correlation matrix of enzyme fluxes (Fig. 5c) demonstrates expected correlations based on known metabolic pathways. These correlations emphasize the relevance and accuracy of the metabolic flux analysis.
Due to the drastically different responses in the cell lines to the ketogenic media, OXCT1 expression was examined across all cells and media conditions. OXCT1, in conjunction with β-hydroxybutyrate dehydrogenase 1 (BDH1), is essential in the conversion of ketones into acetyl-CoA. BDH1 converts BHB to acetoacetate, which is then converted to acetoacetyl-CoA by OXCT1, releasing a free molecule of succinate. Generally, OXCT1 and BDH1 expression show similar trends and are lower in glioblastoma compared to normal tissue10. CA7 downregulates OXCT1 after exposure to ketogenic conditions, while OXCT1 expression was unchanged in L2 and CA3 between the two media conditions. Reduced expression of OXCT1 in the CA7 cells after ketogenic media contradicts our expectations. We would expect increased ketone availability to drive OXCT1 expression. It is likely that ketone availability for CA7 induces a signaling program that defies our current understanding. Previous work has reported changes as well as no differences in expression of both OXCT1 and BDH1 in various types of tumors in response to the ketogenic diet28. Interestingly, L2 displayed significantly lower expression in OXCT1 compared to both CA3 and CA7 when cultured in the standard media. Several studies, including one specific to glioma, have hypothesized that tumors with lower expression of OXCT1 and BDH1 are more sensitive to the ketogenic diet28,29. However, the findings were not uniform and confirm that not all glioblastomas or gliomas with low expression of these enzymes are responsive to the ketogenic diet. One limitation of the study presented here is that OXCT1 expression was examined in CA3, CA7, and L2 only at the endpoint of the study, after all experimental groups had been reintroduced to the higher glucose concentration. Changes in OXCT1 expression may occur earlier in this study prior to the timepoint analyzed. More research is needed to fully elucidate the mechanism behind changes in OXCT1 expression and its relationship to the ketogenic diet in glioblastoma.
Conclusion
Using three distinct human GBM derived cell lines, we have demonstrated divergent metabolic responses to media conditions that simulate the ketogenic diet. We note that concentrations of metabolites were largely unchanged by the KD, but metabolic flux was significantly altered in the ketogenic groups compared to controls. Some individual glioblastomas, such as CA3 and L2, are able to metabolically adapt to conditions of low glucose. However, ketogenic conditions resulted in massive cell death for the CA7 line, indicating that certain phenotypes may be highly sensitive to a KD intervention in humans. The strong increase in flux through glycolytic enzymes displayed in CA7 suggests this cell line relies heavily on the Warburg effect. Differences in utilization of Warburg metabolism in GBM may underlie the varying sensitivities to KD in patients. Ultimately, the results presented here highlight the metabolic differences in individual glioblastomas and emphasize the importance of measuring metabolic flux rather than only metabolite pool sizes. Whole genome sequencing for cancer patients is now practiced at some institutions. Further work may establish stronger links between genetics and the flux phenotypes of some tumors, enhancing decision-making for treatment avenues.
Experimental procedures
Cell culture
Three primary patient-derived glioblastoma cell lines L2 (Supplementary Table 1)30,31, along with CA3 and CA7 (previously named BT53 and BT206, respectively)26, were used in this study. The primary cell lines were isolated and cultured as previously described and approved by Institutional review committee (IRB)31,32,33,34,35,36. Informed consent was obtained from all subjects, and the experiments conformed to the principles set out in the WMA Declaration of Helsinki and Department of Health and Human Services Belmont Report. The cells were maintained as neurospheres using the human NeuroCult NS-A Proliferation Kit (Stem Cell Technologies, #05751) with the addition of EGF (20 ng/mL, Stem Cell Technologies, #78006), FGF (10 ng/mL, Stem Cell Technologies, #78003), and heparin (2 µg/mL, Stem Cell Technologies, #07980). The cells were incubated at 37ºC in an atmosphere of 5% CO2. To maintain optimal density and stimulate further proliferation, the neurospheres were split once they were approximately 150 μm in diameter. As such, this splitting was performed roughly every 7 days. To split, the cells were isolated by centrifugation at 110xg for 8 min, followed by supernatant aspiration. The cells were resuspended in 3 mL accutase (Stem Cell Technologies, #07920) for 8 min. The resulting single cell suspension was then centrifuged at 110xg for 8 additional minutes. The supernatant was aspirated and the cell pellet was resuspended in fresh media. This splitting process was repeated twice per cell line and all experiments were performed using cells at passage 3. These culture conditions were selected based on previous literature and have shown optimal conditions for proliferation and maintenance.
Isotopic tracing using [2H7]glucose
We examined metabolic changes in primary patient-derived human GBM cells using the stable isotope tracer [2H7]glucose ([1,2,3,4,5,6,6-2H7]-D-glucose, Sigma Aldrich, #552003). One day after the third passage, the neurospheres were switched to a standard media replicating the NeuroCult NS-A Proliferation media or a media representative of a ketogenic diet in cells (n = 3–4/group). The standard media is the same formulation as the NeuroCult NS-A Proliferation media for the culture of neural stem cells. The exact composition of the standard media used here was reported previously37. and was made in house to ensure the standard and ketogenic medias were identical except for the glucose and β-hydroxybutyrate concentrations. The standard stem cell media contained 33 mM glucose to replicate the glucose concentration in the NeuroCult NS-A Proliferation media, while the ketogenic stem cell media contained 4 mM glucose and 4mM β-hydroxybutyrate. These concentrations were selected based on previous reports of mimicking the ketogenic diet in cell culture11,38,39. The cells were maintained in their respective medias for 72 h. After this period, both treatment groups were incubated in the stem cell media with 33 mM [2H7]glucose for six hours.
Following the 6-hour incubation, a 1 mL aliquot of media was collected for O2 consumption measurements with the Oxygraph+ (Hansatech Instruments, UK). The remaining sample was centrifuged for 10 min at 200xg. Following centrifugation, the supernatant was collected for 2H Nuclear Magnetic Resonance (NMR) analysis of the media. The cells were also collected after creation of a single cell suspension as previously described with accutase. The remaining cell pellet was resuspended in 4 mL saline. 20 µL of sample was collected and mixed with 20 µL of trypan blue for automated cell counting. The remaining sample was centrifuged for 10 min at 200xg, followed by supernatant aspiration. The cells were then resuspended in 3 mL saline, centrifuged for 10 min at 200xg, and then followed by a final supernatant aspiration. The remaining rinsed cell pellet was rapidly immersed in liquid nitrogen to preserve metabolic profiles for gas chromatography-mass spectrometry analysis. Separate identical experiments were conducted for immunofluorescence of complete neurospheres andH NMR analysis of the dissociated cell pellet. All statistical analysis for this research was conducted in GraphPad Prism (La Jolla, CA, USA, v10.1.2). Oxygen consumption measurements were analyzed by unpaired Students’ t test for each cell line. Cell viability across cell lines and conditions was assessed by a two-way ANOVA with multiple comparisons between the standard and ketogenic conditions. Šídák statistical hypothesis testing was used to correct for multiple comparisons.
Nuclear magnetic resonance spectroscopy
2H NMR was used to measure HDO production in the cell pellet. Cells were lysed with 100 µL 100% acetonitrile which contained 2.5 mM each of unlabeled (Sigma Aldrich, #P56003) and [2H4]pyrazine (Sigma Aldrich, #340456) for chemical shift calibration and quantification. After vortexing and centrifugation at 4 °C at 10xg for 30 min, 80 µL was transferred into a 1.7 mm NMR tubes (Norell #C2384259). Spectra were acquired in a 600 MHz spectrometer equipped with an Avance III NMR console (Bruker Biospin) running TopSpin 4.1.1 and a 1.7 mm Micro CryoProbe2H-NMR spectra were acquired with 2172 complex data points for each of four FIDs for each cell pellet sample. The four FIDs were summed to generate a single spectrum for each sample. A spectral width of 11 ppm was used along with an acquisition time of 1 s. Processing was performed using MestReNova (v14.2.1) where 2 Hz exponential line broadening and zero filling to 4096 points were applied to each summed spectrum. HDO and pyrazine peak areas were obtained by fitting mixed Lorentzian–Gaussian line shapes. HDO concentrations were estimated using the 2.5 mM [2H4]pyrazine peak area, correcting for the number of deuterons responsible for each peak. A two-way ANOVA with multiple comparisons and Šídák statistical hypothesis testing between the standard and ketogenic conditions was used to assess statistical significance of HDO production.
Gas chromatography – mass spectrometry
Metabolites from the cell pellets were extracted using the acetonitrile isopropanol water (AIW) method. Each frozen cell sample was treated with a precooled mixture of 1 mL 3:3:2 AIW (v/v/v). 700 ng of the internal standards norleucine and 3-methyl-2-OH-butyrate as well as a scoop of zirconium oxide beads were also added. The cell samples were homogenized (FastPrep-24, M.P. Biomedicals) to facilitate cell lysis. Each sample was homogenized 3 times for 20 s each and placed on ice for 5 min between rounds. Following homogenization, the samples were resuspended and transferred to 2.0 mL Eppendorf tubes. The remaining beads in the original sample were then rinsed with 500 µL 3:3:2 AIW and added to the Eppendorf tube to optimize sample retention. The samples were centrifuged at 10xg for 45 min at 4ºC to separate the insoluble fraction, yielding a supernatant with extracted metabolites. This supernatant was transferred to a new Eppendorf tube and dried down overnight. A precooled mixture of 200 µL of 1:1 acetonitrile was added to each dried sample the following day. The sample was then vortexed to facilitate suspension, placed in the − 20ºC freezer for 2 h, and vortexed every 30 min. Following the 2 h, the samples were centrifuged at 10xg for 45 min at 4ºC. The supernatant was collected in new Eppendorf tubes and dried down overnight.
Fifty microliters of methoxyamine HCL in pyridine (Thermo Fisher Scientific, Waltham, MA, USA) was added and allowed to react for 90 min at 30 °C. Fifty microliters of N-tert-butyl-dimethylsilyl-N-methyltrifluoroacetamide (MTBSTFA; ProteoSpec MTBSTFA w/ 1% TBDCMS, Ricca Chemical Company, Arlington, TX, USA) was then combined with each sample and reacted at 70 °C for 30 min. Eighty microliters of derivatized sample was transferred to a glass insert (CAS #13-622-207, Thermo Fisher Scientific, Waltham, MA, USA) inside of a 2-mL autosampler vial with a 9-mm PFTE screw cap (CAS #60180-595, Thermo Fisher Scientific, Waltham, MA, USA). Eighty microliters of each sample was automatically injected into the gas chromatograph (Trace 1310) and single quadrupole mass spectrometer using the AI/AS 1300 autosampler (Thermo Fisher Scientific, USA). This system utilized a 30-m RTX-5MS Integra column (Crossbond 5% diphenyl/95% dimethylpolysiloxane CAT# 12623-127, Restek, Bellefonte, PA, USA) and 10-m guard column. Each sample was held at 60 °C for 3 min after injection. The temperature was increased by 10 °C/min up to 325 °C, followed by a 5-min bakeout period. The ion source was held at 230 °C, and the transfer line to the MS was maintained at 290 °C. Xcaliber software (version 4.1) was used to identify and integrate peaks. ICIS peak fitting with an S/N cutoff threshold of 3 and height cutoff at 5.0% of the peak was applied during batch processing. Metabolite concentrations and pool sizes were determined using an 8-point standard curve with a range of 50–2000 ng/sample for each metabolite. Previous databases and literature were used in metabolite identification (Supplementary Table 2). Statistical significance of individual metabolite pool sizes was assessed between the standard and ketogenic conditions in each cell line using a two-way ANOVA with multiple comparisons with Fisher’s LSD correction in GraphPad Prism (p < 0.05). Additionally, pool sizes of individual metabolites were analyzed with a PCA plot, a Venn diagram from two-way ANOVA, and a heatmap using Metaboanalyst40. Data were normalized in Metaboanalyst by autoscaling to maintain the highest level of rigor when comparing experimental groups.
Metabolic modeling
Mass isotopomer distributions across the glycolysis and the TCA cycle were visualized using the Escher-Trace software41. Furthermore, the mass isotopomer distributions were corrected for natural abundance using Isotopomer Network Compartment Analysis (INCA)19 for statistical analysis of isotopic enrichment. Fractional enrichment was analyzed in each isotopologue by Student’s t tests (p < 0.05). INCA software was also used to generate a model of glycolysis and central carbon metabolism based on the mass isotopomer distributions (Supplementary Table 3). To generate quantitative estimates of enzymatic flux through the model, average oxygen consumption measurements for each experimental group were included. Each replicate was modeled individually to ensure vigorous data collection and permit statistical analysis. Estimates of metabolic flux were analyzed using a two-way ANOVA with multiple comparisons between the standard and ketogenic conditions. Šídák statistical hypothesis testing was used to correct for multiple comparisons. Metabolic flux values were also analyzed by Metaboanalyst using a PCA plot, a Venn diagram from two-way ANOVA, and a correlation matrix40. Flux data were normalized in Metaboanalyst by autoscaling.
Immunofluorescence
Following exposure to standard or ketogenic conditions, CA3, CA7, and L2 neurospheres were transferred into a 24-well plate (Corning Costar #3527). The wells had been incubated with Laminin/Poly-D-Lysine Coating Solution for one hour to promote cell adherence (Sigma # LPDL001). The plate was centrifuged for 5 min at 40xg with an acceleration and deceleration set at 5. The plate was rotated 180 degrees, then again centrifuged for 5 min at 300xg with an acceleration and deceleration set at 5 to form an even monolayer. Cells were fixed with 4% paraformaldehyde for 30 min prior to storage at 4 °C. At a separate time, cells were permealibilized with 0.3% TX-100 in 1x PBS for 15 min prior to blocking with ReadyProbes™ 2.5% Normal Goat Serum (1X) (ThermoFisher Scientific, #R37624) for one hour at room temperature. The cells were then incubated with primary antibodies overnight at 4 °C. Primary antibodies used were 3-oxoacid CoA-transferase 1 (OXCT1) (Invitrogen, #PA5-52899, 1:250) and Pyruvate Dehydrogenase (PDH) (Santa Cruz Biotechnology, #sc-377092, 1:500).
The wells were washed three times with 400 µL PBS before application of secondary antibodies for one hour at room temperature. Goat Anti-Mouse (Invitrogen, #A32742 (Alexa Fluor Plus 594), 1:2000) and Goat Anti-Rabbit (Invitrogen, #A32733 (Alexa Fluor Plus 647), 1:2000) secondary antibodies were used. After rinsing an additional three times with PBS, NucBlue Fixed Cell Stain ReadyProbes Reagent (ThermoFisher Scientific, #R37606) was added to the well, covered, and incubated for 15 min at room temperature. After rinsing in PBS, cells were imaged on the Revolve microscope (Echo, San Diego, CA) with the 10x objective. Fluorescent images were taken with identical brightness, digital haze reduction, and other settings across wells. Multiple images were taken per well. Total fluorescence in each channel was quantified in ImageJ and corrected to the nuclear signal. Statistical significance was assessed with a two-way ANOVA with Šídák statistical hypothesis testing to correct for multiple comparisons.
Data availability
All data with the exception of the RNA sequencing data are contained within the manuscript. The RNA sequencing data has been deposited to the European Genome Archive with accession number EGAS00001003251 as part of the following reference: Hoang‐Minh, L. B., Siebzehnrubl, F. A., Yang, C., Suzuki‐Hatano, S., Dajac, K., Loche, T., Andrews, N., Schmoll Massari, M., Patel, J., Amin, K., Vuong, A., Jimenez‐Pascual, A., Kubilis, P., Garrett, T. J., Moneypenny, C., Pacak, C. A., Huang, J., Sayour, E. J., Mitchell, D. A., Sarkisian, M. R., Reynolds, B. A., and Deleyrolle, L. P. (2018) Infiltrative and drug‐resistant slow‐cycling cells support metabolic heterogeneity in glioblastoma. EMBO J. 10.15252/EMBJ.201798772/SUPPL_FILE/EMBJ201798772-SUP-0013-TABLEEV11.XLSX.
References
Oizel, K. et al. Glutamine uptake and utilization of human mesenchymal glioblastoma in orthotopic mouse model. Cancer Metab. 8, 9 (2020).
Czarnywojtek, A. et al. Glioblastoma multiforme: The latest diagnostics and treatment techniques. Pharmacology 108, 423–431 (2023).
Davis, M. E. Glioblastoma Overview of disease and treatment. Clin. J. Oncol. Nurs. 20, S2 (2016).
Trejo-Solis, C. et al. Role of glycolytic and glutamine metabolism reprogramming on the proliferation, invasion, and apoptosis resistance through modulation of signaling pathways in glioblastoma. Int. J. Mol. Sci. 24, (2023).
Agnihotri, S. & Zadeh, G. Metabolic reprogramming in glioblastoma: the influence of cancer metabolism on epigenetics and unanswered questions. Neuro Oncol. 18, 160–172 (2016).
Iranmanesh, Y. et al. Mitochondria’s role in the maintenance of Cancer stem cells in glioblastoma. Front. Oncol. 11, (2021).
Duraj, T. et al. Beyond the Warburg effect: Oxidative and glycolytic phenotypes coexist within the metabolic heterogeneity of glioblastoma. Cells 10, 1–23 (2021).
Mukherjee, P. et al. Therapeutic benefit of combining calorie-restricted ketogenic diet and glutamine targeting in late-stage experimental glioblastoma. Commun. Biol. 2, (2019).
Seyfried, T. N., Flores, R., Poff, A. M., D’Agostino, D. P. & Mukherjee, P. Metabolic therapy: A new paradigm for managing malignant brain cancer. Cancer Letters vol. 356 289–300 Preprint at (2015). https://doi.org/10.1016/j.canlet.2014.07.015
Maeyama, M. et al. Metabolic changes and anti-tumor effects of a ketogenic diet combined with anti-angiogenic therapy in a glioblastoma mouse model. Sci. Rep. 11, (2021).
De Feyter, H. M. et al. A ketogenic diet increases transport and oxidation of ketone bodies in RG2 and 9L gliomas without affecting tumor growth. Neuro Oncol. 18, 1079–1087 (2016).
Duraj, T. et al. Clinical research framework proposal for ketogenic metabolic therapy in glioblastoma. BMC Medicine 2024 22:1 22, 1–49 (2024).
Champ, C. E. et al. Targeting metabolism with a ketogenic diet during the treatment of glioblastoma multiforme. J. Neurooncol. 117, 125–131 (2014).
Rieger, J. et al. A pilot study of ketogenic diet in recurrent glioblastoma. Int. J. Oncol. 45. ERGO, 1843–1852 (2014).
Mahar, R., Donabedian, P. L. & Merritt, M. E. HDO production from [2H7]glucose quantitatively identifies Warburg metabolism. Sci. Rep. 10, (2020).
Chang, M. C. et al. Assessing cancer therapeutic efficacy in vivo using [2H7] glucose deuterium metabolic imaging. Sci. Adv. 11, 568 (2025).
Williamson, D. H., Lund, P. & Krebs, H. A. The redox state of free nicotinamide-adenine dinucleotide in the cytoplasm and mitochondria of rat liver. Biochem. J. 103, 514–527 (1967).
Go, S., Kramer, T. T., Verhoeven, A. J., Elferink, O., Chang, J. C. & R. P. J. & The extracellular lactate-to-pyruvate ratio modulates the sensitivity to oxidative stress-induced apoptosis via the cytosolic NADH/NAD + redox state. Apoptosis 26, 38–51 (2021).
Young, J. D. INCA: a computational platform for isotopically non-stationary metabolic flux analysis. Bioinformatics 30, 1333–1335 (2014).
Xue, S. et al. Dysregulation of phosphoenolpyruvate Carboxykinase in cancers: A comprehensive analysis. Cell. Signal. 120, (2024).
Oppermann, H. et al. Metabolic response of glioblastoma cells associated with glucose withdrawal and pyruvate substitution as revealed by GC-MS. Nutr. Metab. (Lond) 13, (2016).
Cheng, C. P. et al. The mechanisms of malic enzyme 2 in the tumorigenesis of human gliomas. Oncotarget 7, 41460–41472 (2016).
Yang, M. et al. ME2 promotes Proneural–Mesenchymal transition and lipogenesis in glioblastoma. Front. Oncol. 11, (2021).
Yang, C. et al. Glutamine oxidation maintains the TCA cycle and cell survival during impaired mitochondrial pyruvate transport. Mol. Cell. 56, 414–424 (2014).
Jue, T. R. & McDonald, K. L. The challenges associated with molecular targeted therapies for glioblastoma. Journal of Neuro-Oncology vol. 127 427–434 Preprint at (2016). https://doi.org/10.1007/s11060-016-2080-6
Cusulin, C. et al. Precursor States of brain tumor initiating cell lines are predictive of survival in xenografts and associated with glioblastoma subtypes. Stem Cell. Rep. 5, 1–9 (2015).
Yang, R. et al. EGFR activates GDH1 transcription to promote glutamine metabolism through MEK/ERK/ELK1 pathway in glioblastoma. Oncogene 2020 39:14 39, 2975–2986 (2020).
Zhang, J. et al. Low ketolytic enzyme levels in tumors predict ketogenic diet responses in cancer cell lines in vitro and in vivo. J. Lipid Res. 59, 625–634 (2018).
Chang, H. T., Olson, L. K. & Schwartz, K. A. Ketolytic and glycolytic enzymatic expression profiles in malignant gliomas: implication for ketogenic diet therapy. Nutr. Metab. (Lond) 10, (2013).
Hoang-Minh, L. B. et al. Infiltrative and drug‐resistant slow‐cycling cells support metabolic heterogeneity in glioblastoma. EMBO J. 37, (2018).
Deleyrolle, L. P. et al. Evidence for label-retaining tumour-initiating cells in human glioblastoma. Brain 134, 1331–1343 (2011).
Kelly, J. J. P. et al. Proliferation of human glioblastoma stem cells occurs independently of exogenous mitogens. Stem Cells. 27, 1722–1733 (2009).
Hoang-Minh, L. B. et al. Disruption of KIF3A in patient-derived glioblastoma cells: effects on ciliogenesis, Hedgehog sensitivity, and tumorigenesis. Oncotarget 7, 7029–7043 (2016).
Sarkisian, M. R. et al. Detection of primary cilia in human glioblastoma. J. Neurooncol. 117, 15 (2014).
Siebzehnrubl, F. A., Vedam-Mai, V., Azari, H., Reynolds, B. A. & Deleyrolle, L. P. Isolation and characterization of adult neural stem cells. Methods Mol. Biol. 750, 61–77 (2011).
Siebzehnrubl, F. A. et al. The ZEB1 pathway links glioblastoma initiation, invasion and chemoresistance. EMBO Mol. Med. 5, 1196–1212 (2013).
Louis, S. A. & Reynolds, B. A. Generation and differentiation of neurospheres from murine embryonic day 14 central nervous system tissue. Methods Mol. Biol. 290, 265–280 (2005).
Ji, C. C. et al. A ketogenic diet attenuates proliferation and stemness of glioma stemlike cells by altering metabolism resulting in increased ROS production. Int. J. Oncol. 56, 606–617 (2020).
Vallejo, F. A. et al. The contribution of ketone bodies to glycolytic Inhibition for the treatment of adult and pediatric glioblastoma. J. Neurooncol. 147, 317–326 (2020).
Pang, Z. et al. MetaboAnalyst 5.0: narrowing the gap between Raw spectra and functional insights. Nucleic Acids Res. 49, W388–W396 (2021).
Kumar, A., Mitchener, J., King, Z. A. & Metallo, C. M. Escher-Trace: A web application for pathway-based visualization of stable isotope tracing data. BMC Bioinform. 21, (2020).
Acknowledgements
This work was supported by T32DK108736, F31DK138741, and R01s EB032376, DK132254. A portion of this work was performed in the McKnight Brain Institute at the National High Magnetic Field Laboratory’s Advanced Magnetic Resonance Imaging and Spectroscopy (AMRIS) Facility, which is supported by National Science Foundation Cooperative Agreement DMR-2128556 and the State of Florida.
Author information
Authors and Affiliations
Contributions
A.R.: Conceptualization, Methodology, Writing - Original Draft. A.R., A.S., C.H.: Investigation A.R., A.S., C.H., L.D., M.M.: Formal Analysis, Writing - Review & Editing, M.M.: Supervision, Project Administration.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Rushin, A., Shaikh, A., Hardin, C. et al. Metabolic flux analysis of glioblastoma neural stem cells reveals distinctive metabolic phenotypes in ketogenic conditions. Sci Rep 15, 18736 (2025). https://doi.org/10.1038/s41598-025-02124-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-02124-6