Abstract
Even though head and neck squamous cell carcinoma (HNSCC) is the seventh most common cancer worldwide, there are only two PD-1 targeted immunotherapies (pembrolizumab and nivolumab) and one tumor intrinsic EGFR targeted therapy (cetuximab) that are FDA approved for treatment of HNSCC. Taking advantage of a high throughput inhibitor assay and computational tools originally showing success in leukemia, we designed and employed HNSCC-specific inhibitor panels that capture the diversity of aberrational pathways in HNSCC to test viable cells derived from patients’ HNSCC tumors. This provides a functional context to the multi-omic readouts conducted on these samples (mutations, protein expression and copy number alterations). In addition to generating these deeply characterized functional genomics datasets, we also developed additional visual analytics that have the potential to provide greater insight into HNSCC drug response patterns and potentially aid precision oncology tumor boards in evaluation and assessment of effective targeted therapeutic agents.
Similar content being viewed by others
Introduction
Head and neck squamous cell carcinoma (HNSCC) is the seventh most common cancer worldwide1, with approximately 58,450 new cases and 12,230 deaths estimated in the United States for 20242. The gradual increase in survival has been outpaced by the rising incidence of HNSCC3,4. Tumor heterogeneity continues to impede molecular characterization and subsequent improvement in outcomes5,6. Surgical treatment often results in comorbidities such as nerve pain, impairment in eating and vocalization and overall diminished quality of life7,8; whereas the efficacy of nonsurgical options such as radiotherapy and cytotoxic chemotherapy is limited by inherent and acquired resistance5,6. Precision therapies for HNSCC could spare tissue destruction, improve quality of life, and extend survival. However, in the United States this is currently limited to monoclonal antibodies against EGFR (cetuximab) and PD-1 (pembrolizumab and nivolumab)9. While an improvement over classic chemotherapy, the efficacy of EGFR targeting is not reliably predictable10 and patients eventually develop resistance11,12. Furthermore, immunotherapies targeting PD-1 are only effective for a minority of patients13,14,15 and biomarkers for predicting response are lacking16.
While large databases exist for HNSCC patient samples17 – including gene expression and mutation data – the majority of mutations and differentially expressed genes are of unknown functional significance. The genomic complexity of HNSCC has made it difficult to tie 'omics data to effective treatments. To address this, we established a deeply characterized HNSCC cohort with the goal of elucidating the drivers of drug response in HNSCC. Characterization of these patient samples included multiple genomics assays including whole exome sequencing, RNA sequencing, copy number arrays and reverse-phase protein arrays as wells as evaluation with a panel of small-molecule inhibitors and natural products. Our previous work focused on the development of an HNSCC-specific inhibitor assay through the identification of HNSCC-specific biological pathways that could potentially be targeted by current FDA approved cancer drugs18. The integration of high-throughput tumor drug response data with genomic characterization has proven useful in leukemia19, but similar study of HNSCC has been restricted to cell lines as part of multi-cancer20,21 and HNSCC specific studies22,23,24. This study highlights the feasibility of this approach in patient-specific cells tumor cell models in HNSCC. Here we provide a clinically-relevant dataset and highlight patient-specific analyses that potentially could be used for tumor-boards based on the integration of genomic and drug response data that can be used to help guide development of larger scale HNSCC studies.
Results
Clinical and genomics characteristics
The cohort consists of twenty patients enrolled at the time of surgical treatment for primary (n = 14) or first recurrence (n = 6) of HNSCC (Supplementary Table S1). The anatomical sites represented include the oral cavity (n = 12), larynx (n = 4), and oropharynx (n = 3) as well as a single rare (biologically and epidemiologically distinct) tumor of the maxillary sinus. No patient is known to have had more than one diagnosis of recurrent (n = 1) or metastatic (n = 4) HNSCC following the diagnosis of the cancer collected. Multiple patients (n = 5) had histories of other cancers prior to their first diagnosis of HNSCC, but only one (10139) was diagnosed with a non-HNSCC cancer (cutaneous SCC-MOHS) following their first diagnosis of HNSCC.
In total, we obtained results for at least one genomics-based assay for all 20 patients as summarized in Supplementary Table S2.
Comparative analyses with TCGA-HNSC
We obtained tumor tissue from 16 of the patients for whole exome sequencing, with whole blood to serve as a control for somatic genotyping. In The Cancer Genome Atlas HNSCC cohort (TCGA-HNSC)17, they reported 11 significantly mutated genes. Of these 11 genes we found mutations in 8 genes in our HNSCC cohort. When we compared frequencies between TCGA-HNSC and our cohort we found that 5 genes had < 10% difference. The most similar frequencies were for TP53 and NOTCH1, NSD1 and PIK3CA which differed by 1%, 1%, 3% and 6% respectively (Supplementary Fig. S1a). While TP53 and PIK3CA are also significantly mutated in other TCGA cohorts from Pan-Cancer analyses, NOTCH1 and NSD1 appear to be significantly mutated only in TCGA-HNSC25. We further assessed array-based copy number alterations (CNAs) on 8 / 16 patient samples, observing overall higher frequencies of the putative driver recurrent CNA driver genes than found in TCGA-HNSC (Supplementary Fig. S1b). The closest match was CDK2NA which was observed in 25% of our HNSCC cohort and 28% of TCGA.
Expression subtyping of the cohort
Additionally, we performed RNASeq on 15 patient samples, most of which were also characterized with DNA-based analyses (14 / 15; 93%). We obtained and ultimately included 499 samples from the TCGA-HNSC project from the Genomic Data Commons (GDC)26 and harmonized our samples with them (see “Methods”). We first evaluated our ability to assign these patient samples to four previously characterized expression subtypes (Atypical, Basal, Classical and Mesenchymal)27,28 that were called in a subset of the currently available TCGA-HNSC cohort17. We were able to re-call these subtypes with 96% accuracy in TCGA, allowing us to further annotate all available RNASeq samples in TCGA-HNSC and our HNSCC cohort. We determined that our HNSCC cohort represented all four subtypes (Fig. 1a) with both of the HPV+ patients being part of the Atypical subtype consistent with previously observations27.
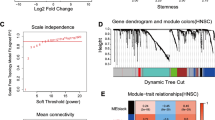
Weighted Gene Co-expression Analysis provides biological context to known HNSCC subtypes and correlates with clinical covariates in the HNSCC cohort. RNASeq from the HNSCC cohort tumor tissue samples were combined with TCGA and the four main subtypes were called for both cohorts. (a) A UMAP of expression data from the 643 genes used for determining subtypes is shown for TCGA with black dots indicating the HNSCC cohort samples. Samples that could not be confidently called (see “Methods”) are shown as transparent circles. (b) A heatmap of the median PC1 score (termed eigengene) in TCGA–HNSC is shown for each module (X-axis) and subtype (Y-axis) indicating the overall expression trend for genes in the module. (c) T-Statistics from specified comparisons (Y-axis) are shown for each module (X-axis). Stars indicate significance of the corresponding T-test where the unadjusted P-value < 0.001 is ***, < 0.01 **, < 0.05 * otherwise no significance. For the directionality of the tests not specified on the plot: One Year RFS–Yes vs. No, Smoker Pack Years–[0,15] vs. (59,100], Alcohol Use—Heavy vs. Minimal, Gender—Female vs. Male, Age–(40,62] vs. (62,90].
De-novo network analyses
To further characterize our expression data with regard to network-based signatures, we first separated the most variable 2,500 genes from the TCGA-HNSC patient samples into 12 co-expression modules using WGCNA29. These modules are denoted by a number and color (e.g. ‘Mod1 (turquoise)’). Note that the ‘Mod0 (grey) module’, which customarily contains genes with low co-expression, is not considered in any analyses. These modules were overrepresented for a variety of hallmark gene sets30 (Supplementary Fig. S2a). For each module, patients were scored by their first two principal components (PCs) and after harmonization, our HNSCC cohort clustered with the TCGA samples (Supplementary Fig. S2b). The first PC is termed the module eigengene31 and can represent the trend of the expression of the genes within the module. Using the median of the module eigengene values in TCGA, we found that at least one module was highly expressed in each HNSCC subtype (Fig. 1b). For instance, patient samples with the Mesenchymal subtype tended to have increased expression of genes in the Mod3 (brown) module which was significantly (FDR < 0.05) associated with the epithelial-mesenchymal transition hallmark. When we compared the module eigengene values between clinical groups we found that two modules, Mod5 (green) and Mod7 (black), were marginally significant (T-test; unadjusted P-value < 0.01) and suggestive (T-test; unadjusted P-value < 0.05) respectively with one-year relapse free survival (RFS) (Fig. 1c). The modules were seen to be associated in opposite directions, with Mod5 (green) decreased in patient samples who achieve one-year RFS while Mod7 (black) was increased. The correlation with the module eigengene (kME)31 is a measure of membership to a given module that can be used to prioritize biomarkers and can provide additional biological relevance. A clear instance of this is the INHBA gene which is the top ranked gene by kME in the Mod5 (green) module. INHBA is involved in EMT processes32, with high expression previously characterized to be associated with poor prognosis in HNSCC33.
Formation of tumor cell models
We established tumor cell cultures from patient solid tumor samples to facilitate inhibitor screening. Of approximately 90 cases attempted, 20 yielded epithelial tumor cultures sufficient for the inhibitor assays. The appearance of hematoxylin and eosin (H&E)-stained sections and pavement-like morphology of the corresponding patient-derived tumor cells in culture (Supplementary Fig. S3; top and bottom) were consistent with epithelial origin. Samples 10336, 10021 and 10356 had varying degrees of mesenchymal morphological characteristics, i.e. subsets of cells tending toward fusiform fibroblast cell morphology and swirls/whorls of colony morphology. To further characterize epithelial composition and ensure minimal fibroblast composition, cell lines were stained with epithelial (keratin 5 and K1/18) and mesenchymal vimentin markers as described in “Methods”. The majority of cell lines expressed keratinocyte markers and were found to contain minimal fibroblast cells (Supplementary Table S3). We further characterized keratin expressing epithelial cells as having an increased mesenchymal character if those cells were found to express the mesenchymal/fibroblast marker vimentin more strongly than the epithelial keratins (Examples of representative fibroblasts and epithelial cell lines expressing varying degrees of mesenchymal character are presented in Supplementary Fig. S4). Epithelial cells going through epithelial-mesenchymal transition (EMT) can progress to undifferentiated cells indistinguishable from fibroblasts; however, none in our cohort appeared to lose epithelial characteristics, consistent with the squamous cell carcinoma pathological assessment of the original tumor. Finally, to facilitate evaluation of drug-associated biomarkers we also generated RNASeq and reverse-phase protein array (RPPA) data for these tumor cell culture samples (Supplementary Table S2).
Description of inhibitor assays
Tumor-derived cell lines at low passage (P1–P2) were exposed to a panel of 89 single-agent targeted inhibitors and natural products. Using our high confidence set of Targetome interactions (see “Methods”) we found that most of the covered drugs (drugs that have known drug target interactions) were concentrated within the PanCancer RTK-RAS pathway34 (Fig. 2a, See Supplemental Table S4 for full target annotation). This included 9 of the 18 genes considered to be candidate therapeutic targets in TCGA-HNSC17 (Fig. 2b).
Summary of inhibitor pathway targeting. (a). A barplot indicating the number of targeted genes for each single-agent inhibitor used with the HNSCC cohort (X-axis) for each PanCancer pathway (Y-axis). (b) A barplot indicating the number of single-agent inhibitors used with the HNSCC cohort (Y-axis) for each candidate therapeutic targets for TCGA-HNSC. Colors correspond to the PanCancer pathways from (a). (c) Shown is a heatmap indicating the annotated co-targeting of the inhibitor combinations used in the HNSCC cohort. Text indicates the number of inhibitors with matching color intensity. (d). Paired t-test summary results for each drug combination (X-axis) grouped by single-agent PanCancer pathway (top). For each combination, the estimated difference (termed the log combination ratio; Y-axis) between the combination log AUC and the lowest single-agent log AUC is shown as a dot centered on a line representing the 95% confidence interval. Negative values indicate the combination is more sensitive than the corresponding single-agents. Stars indicate significance with P-value < 0.001 ***, < 0.01 **, < 0.05 * and N.S. indicating non-significance.
Tumor-derived cell lines were also treated with 49 inhibitor combinations with targets similarly concentrated on combinations of RTK-RAS pathway genes (Fig. 2c). Sensitivity to each agent and combination was determined by calculating the area under the dose-response curve (AUC). Comparing the combination log AUC to the lowest single-agent log AUC for each inhibitor over all samples resulted in 31 combinations being significantly (Paired T-Test; unadjusted P-value < 0.001) more sensitive than either single-agent suggesting non-redundancy of the underlying pathways (Fig. 2d).
Consistency of inhibitor responses with external data
Large scale efforts such as the Cancer Cell Line Encyclopedia20 and the Genomics of Drug Sensitivity in Cancer (GDSC)35 project have evaluated drug response in many commonly used cell line models across cancer types including HNSCC. In order to evaluate consistency of our drug response data to these cell line models, we retrieved mutation, CNA and drug response data (AUC) for 42 cell lines from the GDSC project with data for at least one common drug. Although several approaches such as CELLector36 can find relevant cell lines to TCGA or another large cohort based on their alteration status, we wanted to match our patient samples with cell lines in a manner consistent with drug response. We first found the significant pharmacogenomic interaction gene alterations in the PanCancer and HNSC-specific analysis from Iorio et al. 201637. These genes could be more relevant to drug response than those most frequently mutated. Using these pharmacogenomic interaction genes, we then evaluated the Jaccard distance from each patient sample’s mutation and or CNA profile to each cell line similar to the approach of Sinha et al. 202138 (Fig. 3a). A total of 9 patient samples could be matched to one or more cell lines using this approach. Of those, 7 had significantly correlated AUC values (Pearson’s correlation; unadjusted P-value < 0.001), the remaining 2 also had strong correlations (unadjusted P-value < 0.01) (Fig. 3b). We did observe a negative trend between correlation and alteration distance (i.e. higher correlation was seen on average for lower distances; Supplementary Fig. S5) suggesting the utility of this approach.
Drug response in the HNSCC cohort is consistent with large-scale cell line screens. Patient samples and GDSC cells lines were clustered according to alterations associated with drug response to one of the common inhibitors between the HNSCC cohort and GDSC. (a) Shown is a matrix plot where columns (samples or cell lines) and alterations (rows) are clustered based on their Jaccard distance. Purple indicates that the patient sample or cell line has the corresponding alteration. Only those patient samples (bold) or cell lines with at least one alteration are shown. Note that patient sample 10250 and 10356 have no matching cell lines based on their mutation profile. Pearson’s correlation was computed between the normalized AUC values for each patient sample and the AUCs of the matching inhibitors in GDSC. (b) Scatterplots of the AUC values for the matching drugs between the HNSCC cohort and GDSC are provided. Each scatterplot is faceted by patient sample id and the corresponding best GDSC cell line name (boxes on the top). The text indicates the Pearson’s correlation. Unadjusted significance is indicated by P-value: < 0.001 ***, < 0.01 ** < 0.05 *.
Exploration of inhibitor response in patients
We first devised a gene scoring approach for our AUC-based drug response as previous methods have used IC50 values39. Our statistic was based on the sum of centered/scaled drug AUC values (i.e. Zscores) for each gene and significance was determined using a permutation test per gene. Negative values of these gene scores indicate the degree of sensitivity to a given drug based on the inhibitor distributions. If a mutation or copy number alteration occurred in a gene with a highly negative score it suggested etiological relevance for that patient with respect to their drug response profile. We further refined our set of copy number alterations by comparing the concordance/discordance of the values with the cell culture-based RNASeq and RPPA directionality. For instance, if RNASeq and/or RPPA was available and both had a positive Zscore, then an amplification was considered a concordant amplification (cA). Likewise, a discordant amplification (dA) would have a negative Zscore for the available cell-based data. When looking at the TCGA candidate therapeutic targets we found that only in the case of EGFR amplifications did a mutation or CNA co-occur with a significant gene score (Permutation Test; unadjusted P-value < 0.001; Fig. 4a). Patient 10058 was of particular interest due to EGFR being both significant (Permutation Test; unadjusted P-value < 0.001) but not having a corresponding alteration. To further explore the direct relationships between drug response, alterations and genes, we created an approach to visualize these data called ‘Response Cards’. The Response Card for patient 10058 showed no corresponding direct alteration for any of the significant or suggestive genes (Permutation Test; unadjusted P-value < 0.001 or 0.05 respectively; Fig. 4b), indicating a potential indirect association with an altered gene. Even though this patient did not have CNA data, we would still expect a putative CNA to have a high or low (> abs(2)) RNASeq or RPPA Zscore; this was not observed to be the case. To prioritize the remaining mutations based on gene interaction network proximity, we used a random walk with restarts based approach40 in conjunction with the Reactome FIViz gene interaction network41. We found 7 mutations that occurred in genes considered to be neighbors of EGFR, ERBB2 or ERBB4, the three most significant genes by score (Fig. 4c, d). This combination of studying both direct and indirect alterations allows for a more effective interpretation of the integrated genomics and functional data via the response card and adds considerable value to this dataset.
Prioritization of genes based on drug sensitivity. (a). Shown is a summary of the gene score significance for the HNSCC cohort patients relative to the TCGA candidate therapeutic target genes. The EGFR gene scored highly in three patients, two of whom had amplifications of the gene via copy number. The other patient, 10058, didn’t have copy number data. Patient 10058 had EGFR as the most significant gene target and was unlikely to have had an underlying amplification as it showed mild down-regulation of EGFR from both RPPA and expression. (b). Shown is the Response Card for patient 10058. Inhibitor data is shown on the top in the form of a Zscore with negative values indicating increased sensitivity. From left to right, the scores of the top genes by significance are shown along with SYK which was a target of the most significant drug. In addition, a heatmap displays the drug target data from Targetome. Finally, any mutations (black rectangle) in these genes are shown along with RPPA and expression from the tumor cell culture models after being centered/scaled to Zscore values. If available, copy number data is also displayed. (c) Using a network propagation approach we prioritized the patient’s somatic mutations and the results are displayed as a barplot where the resulting prioritization score is on the Y-axis and corresponding gene is on the X-axis. (d) The mutations that scored highly in the prioritization (blue ovals) were seen to interact with EGFR and the other significant gene targets (green rectangles) either directly or indirectly in a network context.
Discussion
HNSCC represents a genomically heterogeneous malignancy in which mutation and copy number alteration data are not adequate for guiding the development of personalized targeted therapy. Indeed, histopathologically indistinguishable HNSCC samples are likely to have distinct somatic mutational abnormalities42 leading to clinical uncertainty regarding the best treatment options. This study highlights the feasibility and potential utility of functional screening in solid tumors like HNSCC with the assays and methods originally designed for hematological malignancies. By integrating genomic alterations with drug response data from patient-derived tumor cell cultures, our dataset provides the potential to achieve a more thorough understanding of the oncogenesis of individual HNSCCs.
When comparing our cohort to TCGA-HNSC we observed similar mutation distributions, with differences likely due to much smaller sample size in our HNSCC cohort. Additionally, the discordance in CNA frequency can also be explained by cohort size as well as differences in array platform (i.e. Affymetrix CytoScan HD vs. SNP 6.0 chips used in TCGA), as well as the resulting methodological differences. For instance, the Affymetrix CytoScan HD array has fewer probes than the SNP 6.0 and it has been seen to have poor performance in validating known CNVs ≤25 kb, suggesting that it is better suited for the detection of larger CNVs43. The same study also found that there was often high discordance depending on the software and parameterization used. Additionally, we found that our cohort also represented all four expression subtypes. The expression data could be further explored by using co-expression-based methods, which divided the most variable genes into modules. Two of these modules were seen to be associated with one year recurrence-free survival providing the opportunity to evaluate them as prognostic biomarkers in the future with larger datasets. Our inhibitor data were consistent with previously generated cell line data from GDSC. This further indicated the value of our dataset and approach since the data were derived from actual patient samples and are therefore more clinically relevant. When prioritizing genes with respect to drug response, we note that the genomic alterations were rarely in the targeted genes and therefore network-based approaches are necessary to provide a more complete picture of the genes that could underlie observed differences in drug response. Complicating these analyses is the potential for the heterogeneity of the cell lines to become masked by the competitive nature of cell growth in culture. Over time, evolution of the cell population can alter the functional (drug response) and genetic heterogeneity of the cultures relative to the original tumor. This is due to the fact that the fastest growing epithelial cell populations are arbitrarily selected as passage numbers increase. External heterogeneity can also develop from new mutations that can accumulate in culture over time44,45. We sought to mitigate this effect by collection of aliquots for genomic and functional studies from primary cultures directly or early passages (1–2). Future application of integrated analyses could also utilize drug response data from implantable microdevices in human tumors prior to surgery46, replacing tumor cell culture all together.
This dataset provides the first functional context for HNSCC in a representative cohort with in-depth characterization and epithelial tumor cell morphology in culture. More importantly, it provides a framework for the experimental and computational approaches to scale this work in larger cohorts and future studies. The patient-specific “Response Card” and corresponding network-based prioritization not only helps maximize the available information from this cohort but also provides a foundation for precision oncology trials in HNSCC. These tools in the context of SMMART trials being carried out at our institute47 will allow for the integration of intrinsic and extrinsic tumor targeting as we move closer to the goal of personalized targeted therapy for HNSCC.
Methods
Ethics
This study had the approval of the Institutional Review Board at Oregon Health and Science University. All patients gave informed consent to participate in this study and all samples were de-identified. All experiments and methods were performed in accordance with relevant guidelines and regulations.
Patient sample collection and clinical annotation
Tumor, uninvolved oral mucosa, and blood samples were collected from HNSCC patients undergoing surgical resection. Tissues were collected from patients undergoing surgery as the initial treatment for either a primary or first recurrence of HNSCC. Relapse-free survival was measured from the end of the primary (surgical) treatment to the date of next diagnosis by biopsy. Following resection, uninvolved and tumor tissues were each immediately divided into four pieces treated as follows: 20 mg of each tissue were placed in RNAlater (Invitrogen) for 24 h at 4 C prior to storage at -80 C. 20 mg were flash frozen in liquid nitrogen and stored at -140 C. The remaining tissues were divided between culture and 24-hour formalin fixation for paraffin embedding. Whole blood was frozen at -80 C in EDTA treated blood collection vacutainers.
Cell culture and immunostaining
Patient-derived cell cultures were utilized at low passage P1 – P2 (with first replating of primary cultures defined as passage number 1) so as to more closely reflect the epithelial tumor heterogeneity present in the patient’s tumor44,45. Tumor tissue was transported to the lab in transport media consisting of DMEM/F12 Media (Gibco, 11320082) supplemented with 2x antibiotic-antimycotic (Gibco 15240112) and 5% iron-supplemented bovine calf serum (Cytiva, SH3007203). Tissues were minced to approximately 1 mm pieces and grown in tissue-culture treated dishes (Corning, 430293) and roller bottles (Corning, CLS 430195). Roller steady state culture techniques were used to more closely replicate in-vivo conditions. Tissues were cultured in DMEM/F12 Media supplemented with 5% BCS, 1x penicillin-streptomycin (Gibco, 15070063), 5 µg/mL insulin (Sigma, 19278), 10 µg/mL epidermal growth factor (Gibco, PHG0311), 0.18 mM adenine (Sigma, A2786), 0.1 nM cholera toxin (Sigma, C8052), 0.02nM Triiodo-L-thyronine (Sigma, T6397), and 0.4 µg/mL hydrocortisone (Sigma, H0888). Fibroblasts were periodically removed from epithelial culture via cell scrapers and differential trypsinization (Trypsin-EDTA, Gibco, 25200114). All cells were passaged between 60 and 80% confluence. Cell culture methods for adherent epithelium were used48. For examination of epithelial mesenchymal transition (EMT), cells were formalin fixed and subjected to indirect immunofluorescence using primary antibodies to epithelial-specific intermediate filament markers Keratin 5 (K5; abcam-ab52635, rabbit), Keratins 8 and 18 (K8/18; Fitzgerald 20R-Cp004, guinea pig), and mesenchymal intermediate filament marker vimentin (abcam ab8978, mouse). Nuclei were stained with DAPI (405 nm). Secondary antibodies for K8/18 were Goat anti-Guinea Pig IgG with Alexa Fluor™ 488 (A-11073), for vimentin were Goat anti-Mouse IgG with Alexa Fluor™ Plus 647 (A32728), and for Keratin 5 were Goat anti-Rabbit IgG with Alexa Fluor™ 594 (A-11037). No primary antibody replicate cultures served as primary antibody controls.
Ten sets of four superimposable sub-images (CY5, DAPI, GFP, and TxRed channels) were captured for each cell line; five sets of the cells treated with primary antibodies and five of the no-primary controls. Images were captured with an EVOS FL Color Imaging System (AMEFC4300). ImageJ was used to quantify the relative intensity of each channel associated with each cell within each image. Fibroblastic cells were distinguished from epithelial cells based on their sole expression of vimentin. Epithelial cells (cells expressing keratin) with a vimentin signal more intense than the combined signal of K8/18 and K5 were described as having increased mesenchymal character.
Inhibitor assays
Drugs were selected for our inhibitor panel based upon initial responses of HNSCC and cutaneous SCC to a panel designed for leukemia19,49. The functional screening platform, which requires only the tumor cells, was developed over 10 years ago and has been validated for FDA clinical trials (under an IDE or as a correlative readout). Tumor cell lines were exposed to seven 3-fold serial dilutions of 89 single agents and 49 agent combinations within their first two passages. Adherent epithelial cell lines were trypsinized and strained at 70 μm (Corning, 431751) to achieve a near single cell suspension. Cells suspended in DMEM/F12 Media, 5% BCS, and 0.5x antibiotic were plated in 384 well plates (Greiner, 781182) containing pre-aliquoted inhibitors dissolved in 46nL DMSO per well, to a final volume of 48uL per well. Cells were plated at 4000 cells per well using a Multidrop Combi Reagent Dispenser. Following a 72-hour incubation at 37 deg C, 5uL of MTS (a colorimetric assay of metabolic activity used to infer cell viability) (Promega, G3581) was added to each well and allowed to incubate for one hour. For metabolically slow cell lines, incubation time was extended to ensure absorbances fell within the established linear range. Following incubation, absorption at 490nM was measured with a Biotek Synergy H1 Microplate Reader. Values on each plate were normalized to the average of 46 negative control wells containing only DMSO, media, and cells. Wells containing a lethal mixture of flavopiridol, staurosporine, and velcade provided blank (no cell) data.
Formation of response AUC values
Data were pre-processed following the workflow from Tyner et al. 201819 with the following modifications. Data were first harmonized by calculating an ordinary least squares (OLS) regression for those runs with within-panel replicates after applying a ceiling of 1 and a floor of 0 for the normalized viability. If the AUC values showed an increase (i.e., greater cell viability with higher inhibitor concentration), those curves were removed. Any negative cell viability values that had a floor of 0 applied were flagged. The maximum change in AUC amongst the replicates was noted and those runs with differences > 1 were flagged. Remaining within-plate replicates had their normalized viability averaged and subject to a ceiling of 1 and a floor of 0. An additional set of OLS AUCs was computed for sample-inhibitor pairs run on multiple panels. The maximum change in AUC amongst the across-panel replicates was noted and those runs with differences > 0.75 were flagged. The within and across plate replicates were then averaged together with a ceiling of 1 and a floor of 0. A probit regression was then fit to all possible run groups. The AUC values were rescaled relative to the range of log10 concentration values when compared with the GDSC AUC values and when comparing single-agents with drug combinations. These data are publicly available at https://biodev.github.io/HNSCC/.
Annotation of drug targets
In order to establish a high-quality list of targets we utilized the Cancer Targetome50. We required all targets to have assay values (termed Tier III) and further required these values be less than 100 nM. To maximize coverage while minimizing inclusion of lesser characterized targets, drugs-target interactions were categorized based on the number of supporting publications (> 1, 2 or 3). For each drug, only gene targets that were part of the highest category were kept. Note that since our drug screen also includes natural products, we interrogated the natural product database based on Targetome51, only finding one interaction meeting the 100 nM threshold (Coumarin -> CA14). However, since there was a large discrepancy in assay values between the two available references (48 vs. > 200000 nM), we decided to not include it.
Whole-exome sequencing
DNA was isolated from either previously snap-frozen, FFPE preserved or RNAlater treated tumor tissue. Patient matched controls were derived from EDTA treated whole blood or snap frozen uninvolved tissue. Preparation for sequencing was performed using either the DNeasy blood and tissue kit, QiaAmp DNA Mini Kit, AllPrep DNA/RNA/Protein Mini Kit (Qiagen) or NucleoSpin Tissue kit. Whole-exome sequencing was performed using Illumina’s Nextera Rapid Capture Enrichment kit on an Illumina HiSeq 2500 by the Oregon Health and Science University Massively Parallel Sequencing Shared Resource. Preprocessing was performed using an implementation of the GenomeAnalysisToolkits Best Practices52. Finally, Mutect253,54 was used to identify candidate somatic mutations in the tumor samples relative to their matched normal samples55. We annotated the candidate somatic mutations using the Ensembl Variant Effect Predictor (v99.1)56. We only kept mutations that passed the Mutect2 filter, had a predicted non-synonymous effect or were indels, had a gnomAD57 allele frequency < 0.01 and were not tolerated in SIFT58, benign in PolyPhen59 or were considered clinically pathogenic in ClinVar60. Finally we removed any potential false positive mutations/genes using a previously curated list61.
RNASeq
For tumor tissue and uninvolved (normal) samples, RNA was isolated from either Snap Frozen Tumor Tissue samples or from samples preserved in RNAlater using either the RNeasy mini Kit (Qiagen) or the AllPrep DNA/RNA/Protein Mini Kit (Qiagen). For tumor cell cultures, RNA isolation was performed using the RNeasy mini Kit (Qiagen).
Sequencing was performed at two locations, either at OHSU or OSU:
OHSU sequencing
At the Oregon Health and Science University Massively Parallel Sequencing Shared Resource, RNA quality was first assessed using the Agilent 2100 Bioanalyzer. Libraries were then prepared using the TruSeq Stranded mRNA kit (Illumina). Briefly, 100ng of total RNA was converted to cDNA using random hexamers. Synthesis of the second strand was done with the addition of dUTP, which enforced the stranded orientation of the libraries by blocking amplification off the second strand during the first round of PCR. Amplified libraries were profiled on the 4200 TapeStation (Agilent). Libraries were quantified for sequencing using an NGS Library Quantification Kit (Roche/Kapa Biosystems) on a StepOnePlus Real Time PCR Workstation (Thermo/ABI). Sequencing was performed on an Illumina HiSeq 2500 Instrument.
OSU sequencing
Libraries were prepped using Wafergen PrepX PolyA mRNA Isolation Kit followed by Wafergen PrepX RNA-Seq for Illumina. The preps were done on the Wafergen Apollo 324 liquid handler. Final RNAseq libraries were quantified by Qubit fluorometer using the Qubit dsDNA HS Assay. Then the libraries were run on the Agilent BioAnalyzer 2100 using the high sensitivity DNA chip to check sizing. Finally, all libraries were quantified by qPCR using the ABI 7500 fast instrument using the KAPA Biosystems Library Quantification Kit. Samples were normalized and pooled, then run on 6 lanes of the Illumina HiSeq 2000, 100 bp paired end.
Preprocessing
Alignment and counting of the reads was performed using the protocol published by the GDC26. Equivalent gene count data for TCGA-HNSC were also downloaded from the GDC for 520 patient samples derived from the primary tumor. Based on an initial expression filtering using ` filterByExpr` function in edgeR62 and transformation into counts per million, we removed 21 samples whose median expression was less than 2 standard deviations below the mean. Finally, for each collection of samples (tumor, normal, cell culture and TCGA-HNSC) we first filtered counts, normalized using the Trimmed mean of M-values methodology63 and finally computed the log2 RPKM using the functionality in edgeR.
WGCNA
Using the filtered and normalized log2 RPKM data, we first limited genes to the common set between the TCGA-HNSC and HSNCC cohort samples. We used ComBat64 to adjust for the batch effects between the different sequencing runs in the HNSCC cohort and the TCGA-HNSC cohort using TCGA-HNSC as the reference. The effectiveness of this approach was verified using BatchQC65. Limited to only the 499 TCGA samples, we evaluated a range of parameters for WGCNA including the number of variable genes, power and four variations of key module detection parameters using a combination of subsampling as well as assessment of module quality66. Based on these results, we chose to use the 2,500 most variable genes and create a signed-hybrid network using ‘bicor’ correlation67 with maxPOutliers = 0.1, power = 3, deepSplit = 2, detectCutHeight = 0.995, minModuleSize = 30, pamStage = T. For the HNSCC cohort, the principal component values used for the eigengenes were generated using ‘prcomp’ in R after centering and scaling based off the per-gene mean and standard deviation of the TCGA data. Enrichment of the module genes was computed using clusterProfiler68, for each module, limiting the universe to the 2,500 most variable genes.
Subtype calling
We used a variation of a prior method implemented in breast cancer69. We first downloaded the centroids used in the TCGA-HNSC publication as well as their published prior calls70. Next, we median centered the expression data by gene and computed the Pearson’s correlation between each centroid and the expression of the common genes. The subtype with the best correlation was assigned. This was the value used for assessing the performance in TCGA-HNSC. However, as seen in Fig. 1a if the correlation was negative or the second-best correlation was too close (< 0.1) we flagged these calls as indeterminate.
Copy number alterations
Copy number was assessed for eight cases with sufficient DNA after WES using the same samples. Copy number analysis was performed by the Oregon Health and Science University Gene Profiling Shared Resource, using a CytoScan HD Array Kit on a GeneChip™ Scanner 3000 7G System. First RawCopy71 was used to generate log ratios and B-allele frequencies for the tumor and matched normal samples relative to GRCh37. ASCAT72 was then used produce segmented allele-specific copy number for the tumor samples. The segments were then overlapped with Gencode v1973 protein coding genes. The total copy number was determined by taking the sum of the major and minor alleles. We median centered the total copy number for each sample and assigned each gene the largest (based on absolute value) centered copy number value. Amplifications and deletions were called based on genes having a centered value > 1 and < − 1 respectively.
Reverse phase protein array
Cell cultures were grown to 70–90% confluence in 6-well plates. Following protein extraction with lysis buffer (purchased; MD Anderson), lysate was further diluted in lysis buffer and 4x SDS buffer (purchased; MD Anderson) to 1.5ug/ul. Samples were stored at -20 C before delivery to MD Anderson. Core protocols for RPPA data generation, processing and normalization were utilized for the data74. For all analyses, we used the 278 antibodies considered to be validated75. Analyses were performed using log2 transformed normalized linear data.
Integrative analyses of inhibitor data
Drug target score
The database of drug-target interactions was structured as a g x d matrix \(\:T=\left[{t}_{ij}\right]\) where \(\:{t}_{ij}\) = 1 if gene i was a target for drug j and 0 otherwise. Next, we defined a d x n matrix X consisting of up to d AUC values for each of the n patients. For each drug j we first centered and scaled each sample k as \(\:{(X}_{jk}-mean\left({X}_{j}\right))/sd\left({X}_{j}\right)\) forming a d x n matrix \(\:Z\:=\:\left[{z}_{jk}\right]\). Finally, we define our g x n gene score matrix G \(\:=\left[{g}_{ik}\right]\) as:
P-values for each gene were generated by permuting the rows (drug labels) for \(\:Z\) 1,000 times, and recomputing G, counting the number of times the permuted version of the score for each gene was less than the original.
Network propagation
We propagated our drug target scores to the larger set of genes comprising of the Reactome Functional (FI) gene interaction database41. We first limited the Reactome interactions to only those with a score > 0.95. Next, we converted it into an adjacency matrix \(\:A=\left[{a}_{ij}\right]\:\)where \(\:{a}_{ij}=1\) if gene i is connected to gene j and 0 otherwise. We only considered genes that were part of the largest connected component. Note we are treating the graph as undirected whereas the Reactome FI framework also reports directionality for some interactions. We then normalized \(\:A\) by the diagonal matrix \(\:D\) containing the column degree:
For each patient we categorize the genes by the significance level of the gene scores. Top tier genes are P-value < 0.001, second tier < 0.01 and third is < 0.05. We define \(\:{P}_{0}\) as a vector containing the most significant genes by gene score for a given patient, normalized to sum to 1.0. Fixing \(\:\text{c}=0.7\), we solve:
Until convergence76. In this approach, high scores of \(\:{P}_{k}\) indicate a higher proximity to the selected genes and therefore more likely to be relevant to the drug response40. As the degree of genes in the interaction database can amplify the scores in \(\:{P}_{k}\), we implemented an approach known as random degree preserving networks (RDPN) to generate a P-value adjusting for degree77. Specifically, the above procedure in (3) is computed for 1,000 re-wired networks where the degree is preserved. Empirical P-values are computed for each gene comparing how many times the original score (i.e. the corresponding element of \(\:{P}_{k})\) is larger than those generated from re-wired graphs. Genes with a copy number alteration or mutation were prioritized for those genes with RDPN unadjusted P-values < 0.05. The subnetwork containing the top genes by score and alterations was formed using an adaptation of the STM Steiner tree approximation approach78. The resulting network plot was produced using Cytoscape79,80.
Data availability
The raw datasets generated and analyzed during the current study are available in the dbGaP repository (Accession number: phs003456.v1.p1), https://www.ncbi.nlm.nih.gov/projects/gap/cgi-bin/study.cgi?study_id=phs003456.v1.p1. Processed data as used in the analyses are publicly available at https://biodev.github.io/HNSCC/. Unedited microscopy images supporting Supplementary Figures S3 and S4 are available in FigShare: https://doi.org/10.6084/m9.figshare.28988237.v1.
References
Barsouk, A. et al. Risk factors, and prevention of head and neck squamous cell carcinoma. Med. Sci. 11, 42 (2023).
Cancer of the Oral Cavity and Pharynx - Cancer Stat Facts.
Wierzbicka, M., Pietruszewska, W., Maciejczyk, A. & Markowski, J. Trends in incidence and mortality of head and neck Cancer subsites among elderly patients: A Population-Based analysis. Cancers. 17. https://doi.org/10.3390/cancers17030548 (2025).
Miquel Dominguez, A. et al. Incidence, prevalence, and survival of head and neck cancers in the united Kingdom from 2000–2021. MedRxiv. https://doi.org/10.1101/2024.12.05.24318538 (2024).
Alsahafi, E. et al. Clinical update on head and neck cancer: molecular biology and ongoing challenges. Cell. Death Dis. 10, 540. https://doi.org/10.1038/s41419-019-1769-9 (2019).
Johnson, D. E. et al. Head and neck squamous cell carcinoma. Nat. Rev. Dis. Primers. 6, 92. https://doi.org/10.1038/s41572-020-00224-3 (2020).
Aarstad, H. J. et al. The amount of treatment versus quality of life in patients formerly treated for head and neck squamous cell carcinomas. Eur. Arch. Otorhinolaryngol. 263, 9–15. https://doi.org/10.1007/s00405-005-0961-y (2006).
Bose, P., Brockton, N. T. & Dort, J. C. Head and neck cancer: from anatomy to biology. Int. J. Cancer. 133, 2013–2023. https://doi.org/10.1002/ijc.28112 (2013).
Institute, N. C. Drugs approved for head and neck cancer. https://www.cancer.gov/about-cancer/treatment/drugs/head-neck (2021).
Licitra, L. et al. Evaluation of EGFR gene copy number as a predictive biomarker for the efficacy of cetuximab in combination with chemotherapy in the first-line treatment of recurrent and/or metastatic squamous cell carcinoma of the head and neck: EXTREME study. Ann. Oncol. 22, 1078–1087. https://doi.org/10.1093/annonc/mdq588 (2011).
Chong, C. R. & Janne, P. A. The quest to overcome resistance to EGFR-targeted therapies in cancer. Nat. Med. 19, 1389–1400. https://doi.org/10.1038/nm.3388 (2013).
Vermorken, J. B. et al. Platinum-based chemotherapy plus cetuximab in head and neck cancer. N Engl. J. Med. 359, 1116–1127. https://doi.org/10.1056/NEJMoa0802656 (2008).
Borel, C., Jung, A. C. & Burgy, M. Immunotherapy breakthroughs in the treatment of recurrent or metastatic head and neck squamous cell carcinoma. Cancers. 12. https://doi.org/10.3390/cancers12092691 (2020).
Tahara, M. et al. Real-world safety and effectiveness of nivolumab for recurrent or metastatic head and neck cancer in Japan: a post-marketing surveillance. Int. J. Clin. Oncol. 26, 1619–1627. https://doi.org/10.1007/s10147-021-01949-1 (2021).
Ferris, R. L. et al. Nivolumab for recurrent Squamous-Cell carcinoma of the head and neck. N Engl. J. Med. 375, 1856–1867. https://doi.org/10.1056/NEJMoa1602252 (2016).
Gavrielatou, N., Doumas, S., Economopoulou, P., Foukas, P. G. & Psyrri, A. Biomarkers for immunotherapy response in head and neck cancer. Cancer Treat. Rev. 84, 101977. https://doi.org/10.1016/j.ctrv.2020.101977 (2020).
Lawrence, M. S. et al. Comprehensive genomic characterization of head and neck squamous cell carcinomas. Nature 517, 576–582. https://doi.org/10.1038/nature14129 (2015).
Choonoo, G. et al. Illuminating biological pathways for drug targeting in head and neck squamous cell carcinoma. PLoS One. 14, e0223639. https://doi.org/10.1371/journal.pone.0223639 (2019).
Tyner, J. W. et al. Functional genomic landscape of acute myeloid leukaemia. Nature 562, 526–531. https://doi.org/10.1038/s41586-018-0623-z (2018).
Barretina, J. et al. The Cancer cell line encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483, 603–607. https://doi.org/10.1038/nature11003 (2012).
Garnett, M. J. et al. Systematic identification of genomic markers of drug sensitivity in cancer cells. Nature 483, 570–575. https://doi.org/10.1038/nature11005 (2012).
Ghasemi, F. et al. High-throughput testing in head and neck squamous cell carcinoma identifies agents with Preferential activity in human papillomavirus-positive or negative cell lines. Oncotarget 9, 26064–26071. https://doi.org/10.18632/oncotarget.25436 (2018).
Lepikhova, T. et al. Drug-Sensitivity screening and genomic characterization of 45 HPV-Negative head and neck carcinoma cell lines for novel biomarkers of drug efficacy. Mol. Cancer Ther. 17, 2060–2071. https://doi.org/10.1158/1535-7163.MCT-17-0733 (2018).
Tuomainen, K. et al. High-throughput compound screening identifies navitoclax combined with irradiation as a candidate therapy for HPV-negative head and neck squamous cell carcinoma. Sci. Rep. 11, 14755. https://doi.org/10.1038/s41598-021-94259-5 (2021).
Kandoth, C. et al. Mutational landscape and significance across 12 major cancer types. Nature 502, 333–339. https://doi.org/10.1038/nature12634 (2013).
Grossman, R. L. et al. Toward a shared vision for cancer genomic data. N. Engl. J. Med. 375, 1109–1112 (2016).
Walter, V. et al. Molecular subtypes in head and neck Cancer exhibit distinct patterns of chromosomal gain and loss of canonical Cancer genes. PLOS ONE. 8, e56823. https://doi.org/10.1371/journal.pone.0056823 (2013).
Chung, C. H. et al. Molecular classification of head and neck squamous cell carcinomas using patterns of gene expression. Cancer Cell. 5, 489–500 (2004).
Zhang, B. & Horvath, S. A general framework for weighted gene co-expression network analysis. Stat. Appl. Genet. Mol. Biol. 4 (2005).
Liberzon, A. et al. The molecular signatures database hallmark gene set collection. Cell. Syst. 1, 417–425 (2015).
Horvath, S. & Dong, J. Geometric interpretation of gene coexpression network analysis. PLoS Comput. Biol. 4, e1000117 (2008).
Yu, Y., Wang, W., Lu, W., Chen, W. & Shang, A. Inhibin β-A (INHBA) induces epithelial–mesenchymal transition and accelerates the motility of breast cancer cells by activating the TGF-β signaling pathway. Bioengineered 12, 4681–4696. https://doi.org/10.1080/21655979.2021.1957754 (2021).
Khera, N. et al. Identification of multidrug chemoresistant genes in head and neck squamous cell carcinoma cells. Mol. Cancer. 22, 146. https://doi.org/10.1186/s12943-023-01846-3 (2023).
Sanchez-Vega, F. et al. Oncogenic signaling pathways in the Cancer genome atlas. Cell 173, 321–337e310. https://doi.org/10.1016/j.cell.2018.03.035 (2018).
Yang, W. et al. Genomics of drug sensitivity in Cancer (GDSC): a resource for therapeutic biomarker discovery in cancer cells. Nucleic Acids Res. 41, D955–D961. https://doi.org/10.1093/nar/gks1111 (2012).
Najgebauer, H. et al. CELLector: genomics-guided selection of cancer in vitro models. Cell. Syst. 10, 424–432 (2020).
Iorio, F. et al. A landscape of Pharmacogenomic interactions in cancer. Cell 166, 740–754 (2016).
Sinha, R., Luna, A., Schultz, N. & Sander, C. A pan-cancer survey of cell line tumor similarity by feature-weighted molecular profiles. Cell. Rep. Methods. 1. https://doi.org/10.1016/j.crmeth.2021.100039 (2021).
Tyner, J. W. et al. Kinase pathway dependence in primary human leukemias determined by rapid inhibitor screening. Cancer Res. 73, 285–296. https://doi.org/10.1158/0008-5472.CAN-12-1906 (2013).
Bottomly, D. et al. HitWalker: variant prioritization for personalized functional cancer genomics. Bioinformatics 29, 509–510. https://doi.org/10.1093/bioinformatics/btt003 (2013).
Wu, G., Dawson, E., Duong, A., Haw, R. & Stein, L. ReactomeFIViz: a cytoscape app for pathway and network-based data analysis. F1000Res 3, 146. https://doi.org/10.12688/f1000research.4431.2 (2014).
Hunter, K. D. et al. Divergent routes to oral cancer. Cancer Res. 66, 7405–7413. https://doi.org/10.1158/0008-5472.CAN-06-0186 (2006).
Haraksingh, R. R., Abyzov, A. & Urban, A. E. Comprehensive performance comparison of high-resolution array platforms for genome-wide copy number variation (CNV) analysis in humans. BMC Genom. 18, 1–14 (2017).
Ben-David, U. et al. Genetic and transcriptional evolution alters cancer cell line drug response. Nature 560, 325–330. https://doi.org/10.1038/s41586-018-0409-3 (2018).
Liu, K. et al. Evaluating cell lines as models for metastatic breast cancer through integrative analysis of genomic data. Nat. Commun. 10, 2138. https://doi.org/10.1038/s41467-019-10148-6 (2019).
Jonas, O. et al. An implantable microdevice to perform high-throughput in vivo drug sensitivity testing in tumors. Sci. Transl Med. 7, 284ra257. https://doi.org/10.1126/scitranslmed.3010564 (2015).
Labrie, M. Multi-omics analysis of serial samples from metastatic TNBC patients on PARP inhibitor monotherapy provide insight into rational PARP inhibitor therapy combinations. https://doi.org/10.1101/2020.07.25.20146431v1 (2020).
Rasmussen, C., Thomas-Virnig, C. & Allen-Hoffmann, B. L. Classical human epidermal keratinocyte cell culture. Methods Mol. Biol. 945, 161–175. https://doi.org/10.1007/978-1-62703-125-7_11 (2013).
Anderson, A. N. et al. Functional genomic analysis identifies drug targetable pathways in invasive and metastatic cutaneous squamous cell carcinoma. Cold Spring Harb Mol. Case Stud. 6. https://doi.org/10.1101/mcs.a005439 (2020).
Blucher, A. S., Choonoo, G., Kulesz-Martin, M., Wu, G. & McWeeney, S. K. Evidence-Based precision oncology with the Cancer targetome. Trends Pharmacol. Sci. 38, 1085–1099. https://doi.org/10.1016/j.tips.2017.08.006 (2017).
Chamberlin, S. R. et al. Natural product target network reveals potential for cancer combination therapies. Front. Pharmacol. 10, 557 (2019).
Institute, B. gatk4-data-processing. https://github.com/gatk-workflows/gatk4-data-processing (2021).
MuTect2. Call somatic SNPs and indels via local re-assembly of haplotypes. https://software.broadinstitute.org/gatk/documentation/tooldocs/current/org_broadinstitute_gatk_tools_walkers_cancer_m2_MuTect2.php (2016).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31, 213–219. https://doi.org/10.1038/nbt.2514 (2013).
Institute, B. somatic-snvs-indels. https://github.com/gatk-workflows/gatk4-somatic-snvs-indels/tree/master (2020).
McLaren, W. et al. The ensembl variant effect predictor. Genome Biol. 17, 1–14 (2016).
Karczewski, K. J. et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581, 434–443. https://doi.org/10.1038/s41586-020-2308-7 (2020).
Ng, P. C. & Henikoff, S. S. I. F. T. Predicting amino acid changes that affect protein function. Nucleic Acids Res. 31, 3812–3814 (2003).
Adzhubei, I., Jordan, D. M. & Sunyaev, S. R. Predicting functional effect of human missense mutations using PolyPhen-2. Curr. Protoc. Hum. Genet. 76, 21–41 (2013).
Landrum, M. J. et al. ClinVar: public archive of interpretations of clinically relevant variants. Nucleic Acids Res. 44, D862–D868 (2016).
Bottomly, D. et al. Integrative analysis of drug response and clinical outcome in acute myeloid leukemia. Cancer Cell. 40, 850–864 (2022).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. EdgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Robinson, M. D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 11, 1–9 (2010).
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007).
Manimaran, S. et al. BatchQC: interactive software for evaluating sample and batch effects in genomic data. Bioinformatics 32, 3836–3838 (2016).
Langfelder, P., Luo, R., Oldham, M. C. & Horvath, S. Is my network module preserved and reproducible?? PLoS Comput. Biol. 7, e1001057. https://doi.org/10.1371/journal.pcbi.1001057 (2011).
Langfelder, P. & Horvath, S. Fast R functions for robust correlations and hierarchical clustering. J. Stat. Softw. 46 (2012).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. ClusterProfiler: an R package for comparing biological themes among gene clusters. Omics: J. Integr. Biology. 16, 284–287 (2012).
Ding, Y. C. et al. Molecular subtypes of triple-negative breast cancer in women of different race and ethnicity. Oncotarget 10, 198 (2019).
Network, T. C. G. A. Comprehensive genomic characterization of head and neck squamous cell carcinomas supplemental data. https://gdc.cancer.gov/about-data/publications/hnsc_2014 (2015).
Mayrhofer, M., Viklund, B., Isaksson, A. & Rawcopy Improved copy number analysis with affymetrix arrays. Sci. Rep. 6, 36158. https://doi.org/10.1038/srep36158 (2016).
Van Loo, P. et al. Allele-specific copy number analysis of tumors. Proc. Natl. Acad. Sci. 107, 16910–16915 (2010).
Harrow, J. et al. GENCODE: producing a reference annotation for ENCODE. Genome Biol. 7, 1–9 (2006).
Anderson, M. Technical Information. https://www.mdanderson.org/research/research-resources/core-facilities/functional-proteomics-rppa-core/technical-information.html (2020).
Anderson, M. Antibody Information and Protocols. https://www.mdanderson.org/research/research-resources/core-facilities/functional-proteomics-rppa-core/antibody-information-and-protocols.html (2020).
Cowen, L., Ideker, T., Raphael, B. J. & Sharan, R. Network propagation: a universal amplifier of genetic associations. Nat. Rev. Genet. 18, 551–562. https://doi.org/10.1038/nrg.2017.38 (2017).
Biran, H., Kupiec, M. & Sharan, R. Comparative analysis of normalization methods for network propagation. Front. Genet. 10. https://doi.org/10.3389/fgene.2019.00004 (2019).
Sadeghi, A. & Fröhlich, H. Steiner tree methods for optimal sub-network identification: an empirical study. BMC Bioinform. 14, 144. https://doi.org/10.1186/1471-2105-14-144 (2013).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504. https://doi.org/10.1101/gr.1239303 (2003).
Otasek, D., Morris, J. H., Bouças, J., Pico, A. R. & Demchak, B. Cytoscape automation: empowering workflow-based network analysis. Genome Biol. 20, 185. https://doi.org/10.1186/s13059-019-1758-4 (2019).
Acknowledgements
First and foremost, we would like to thank our patients, who are our partners in this work. We would also like to acknowledge Chris Harrington, Robert Searles, Jeffrey Tyner, Xiaoming Ouyang, Daniel Clayburgh, Gabrielle Choonoo, Ryan Li, Peter Andersen, Daniel Petrisor and Jennifer Jennings for their input on this project. We would also like to thank Clara Stemwedel for her administrative support. This work was supported by the National Institutes of Health, National Cancer Institute R01 CA192405 (M.K-M and S.K.M), T32 CA106195 (M.K-M. and N.E.). Short read sequencing assays were performed by the OHSU Massively Parallel Sequencing Shared Resource and SNP arrays were performed by the OHSU Integrated Genomics Laboratory, both of which receives support from the OHSU Knight Cancer Institute NCI Cancer Center Support Grant P30CA069533. The Functional Proteomics RPPA Core facility is supported by MD Anderson Cancer Center Support Grant # 5 P30 CA016672-40. The research reported in this publication used computational infrastructure supported by the Office of Research Infrastructure Programs, Office of The Director, of the National Institutes of Health under Award Number S10OD034224. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
DB: Designed and implemented final workflows and analyses, generated final computational figures and performed manuscript preparation; CM performed experimental design and manuscript preparation. MV: patient consent, cell culture, RPPA, manuscript preparation. SJ: data management, processing and assisted AB with Targetome updates. NE: Preprocessing and QC of inhibitor data. AA, AL: Sample prep, data curation. AB: Targetome updates and data integration, CZ: initial genomics workflow development. TL: Data Visualization. JJ, MK, SM: project conception, mentorship, experimental design, integration and interpretation of results, manuscript preparation.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bottomly, D., Mathieson, C., Vigoda, M. et al. Assessing individual head and neck squamous cell carcinoma patient response to therapy through integration of functional and genomic data. Sci Rep 15, 19742 (2025). https://doi.org/10.1038/s41598-025-03111-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-03111-7