Abstract
New HIV-1 infections are genotyped as part of standard of care testing to ensure that antiretroviral treatment will be efficacious against the virus. Historically this has been performed by sequencing the pol region of the HIV-1 genome only. The popularity of next-generation sequencing (NGS) methods during the SARS-CoV-2 pandemic has resulted in a shift towards using NGS in diagnostic sequencing, but there remain limited methodologies utilising the strengths of NGS for robust diagnostic sequencing of longer regions of the HIV-1 genome. Given the acceptance and success of tiling PCR methodologies during the SARS-CoV-2 pandemic, we aimed to design and verify a novel tiling PCR method for routine HIV-1 sequencing. A set of tiling PCR primers was designed to amplify the 5’ half of HIV-1 in six overlapping segments of 1,000 bp in only two PCR reactions. The assay can move from sample to sequencer in under a day. The tiling PCR was able to generate HIV-1 sequences from 90 (100%) samples in a comparison panel, and complete protease-reverse transcriptase and integrase regions were amplified in > 90% of samples with a viral load > 5000 copies/mL. Seven additional drug resistance mutations were identified when using this novel method. As such, this novel designer tiling PCR is a promising method for the routine NGS-based diagnostic sequencing of HIV-1.
Similar content being viewed by others
Introduction
Current antiretroviral therapies (ART), when taken daily, suppress viral loads of human immunodeficiency virus 1 infections (HIV-1) such that ongoing viral transmission is halted1,2,3. A variety of different ART regimes are available that target different parts of the HIV-1 lifecycle. As part of standard of care, viral sequencing is therefore performed to assess for the presence of drug resistance mutations (DRMs) for different therapies in the viral swarm, and tailor the therapy to the individual.
Commercial assays are available to perform this routine HIV-1 sequencing, while due to cost or support issues some laboratories use a specialised in-house assay to amplify and sequence the virus4,5,6.. Regardless of method, in most cases only subgenomic regions, approximately 1–2 kb, are amplified and sequenced, often using Sanger sequencing. More specifically, sequencing occurs primarily in the HIV-1 pol gene region, and results may include sequence derived from protease (PR), reverse transcriptase (RT), and/or integrase (IN).
In recent years, next-generation sequencing (NGS) technologies have improved in performance7 and ongoing innovation has decreased running costs8. These technological improvements, combined with the availability of robust and sensitive sequencing protocols9,10,11,12, led to the rapid adoption of NGS-based viral diagnostic sequencing during the SARS-CoV-2 pandemic. As such, there is now increased capacity for using NGS within diagnostic laboratories.
One key advantage for using NGS for HIV-1 sequencing is that longer regions of the genome can be more easily sequenced using these methods. This would ensure that sequencing can encompass novel DRM sites; curative, therapeutic and vaccine strategies that target regions of the genome outside of pol are in advanced stages of development13,14, and as such sequencing longer genomic regions ensures an assay will not have to be changed to capture additional mutations as these strategies mature. An increase in the data available per genome also increases the capacity to perform cross-genome analysis such as for the purpose of molecular epidemiology15, or for the identification of novel recombinant forms of HIV which may not be identified when sequencing smaller genome regions only16. Sequencing deeper, made possible using NGS, can also identify rare populations that may significantly impact the response to therapy, such as minor variant DRMs or cases of superinfection17,18,19,20. If a method can be derived that can sequence increased genome lengths using a time and cost-effective approach and non-specialised equipment, HIV-1 surveillance also becomes more accessible in lower resourced settings.
Key guidelines have already been developed for the implementation of NGS for HIV-1 diagnostics at the First and Second Winnipeg Consensus Symposia21,22. One of the issues identified is the requirement for consistent, verified protocols for NGS-based HIV-1 sequencing22. During the SARS-CoV-2 pandemic, many laboratories utilised a tiling PCR methodology for routine whole genome sequencing using NGS9,10,12. Tiling PCR is advantageous as it is faster than amplifying the genome in a single full-length amplicon, and thanks to the multiplexed amplification strategy only two PCR reactions are required for long-range sequencing. Moreover, tiling PCR is less susceptible to the impacts of sample degradation than full-genome PCR methods, and more effective in amplifying low viral load samples. Given tiling PCRs were widely adopted for SARS-CoV-2 sequencing in diagnostic laboratories, and bioinformatic pipelines for tiling PCR analysis have become increasingly accessible23,24, we chose to develop a similar method that could be used for routine diagnostic HIV-1 sequencing.
Here we report a tiling PCR method for bulk HIV-1 sequencing from plasma for use in an Australian epidemic. We determined the overall utility of this method in amplifying the 5’ HIV-1 genome in an efficient and cost-effective manner. We also verified this method following procedures from the WHO HIVResNet HIV Drug Resistance Laboratory Operational Framework25, and compared our method to previous sequencing performed using Sanger methods.
Methods
Study approval
Ethics approval to use archived HIV-1 samples from the ENCORE26 and SECOND-LINE27 cohorts was obtained from the St Vincent’s Hospital Human Research Ethics Committee, reference 2022/ETH01204. Informed consent for archived samples to be used in future studies was given as part of the original cohort consent process. All experiments were performed in accordance with the relevant guidelines and regulations.
Design of tiling PCR
We aimed to design a tiling PCR assay that could be used for routine HIV-1 screening in Australia. Given this regional usage, primers were designed to bind to the most common Australian HIV-1 subtypes: B, C and CRF01_AE28,29. All full-length (> 8500 bp) subtype B (n = 6757), C (n = 2588) and CRF01_AE (n = 1505) HIV-1 genome sequences were therefore downloaded from GenBank (19 January 2022). Sequences with a high proportion of non-coding sites, duplicate sequences, and those where the incorrect subtype had been identified were removed to create three subtype-specific reference alignments. To minimise computing time, a smaller alignment for each subtype, the representative alignment, was also generated by subsampling 16 sequences at random from the reference alignments. All downloaded sequences with metadata identifying them as of Australian origin (n = 111) were used to generate an Australian HIV-1 reference alignment.
To identify putative primers, PrimalScheme analysis (regular QC settings)23 was performed on the three subtype representative alignments. A design based on the amplification of 1 kb segments was chosen to minimise the primers needing to be designed while also minimising the amplicon length, and therefore the extension time of the PCR. To identify primers that could bind to all three subtypes, the collated primer list was remapped to the subtype representative alignments in Geneious Prime 2 (2022.1.1). Primers were then iteratively redesigned based on the following characteristics: < 3 mismatches to the representative alignments, overlap of generated segments > 100 bp, putative amplicon length 0.6–1.5 kb, Tm between 55 and 60 °C, presence of GC clamp, no self-dimer/hairpin formation with Tm > 40, no significant interaction found between primers, putative binding to the reference (subtype and Australian) alignments. If no pan-subtype specific primer could be designed that fit these characteristics, three subtype-specific primers were designed instead (Fig. 1). Once a prototypic primer set had been generated, primer quality control and assay optimisation (Supplementary Methods) was performed to determine the final primer set and assay conditions for the HIV-1 Tiling PCR. Reliable primers targeting env were unable to be designed that fit these requirements, therefore only the 5’ genome (gag-vpu) was targeted in the final methodology to ensure coverage of clinically important regions.
Primer design for use in a tiling PCR assay for HIV-1 sequencing. A combination of published30,31,32 and newly designed primers were utilised to amplify the 5’ end of HIV-1 in six 1 kb overlapping amplicons. Primers are split into two pools to ensure overlapping fragments are generated in separate reactions. Where a pan-subtype primer could not be designed, multiple subtype-specific primers were designed, targeting subtypes CRF01_AE, B and C as indicated. Blue highlighting in the genome indicates regions typically sequence for drug resistance screening.
Sample selection and preparation
The HIV-1 tiling PCR was developed and verified using plasma samples from archived ENCORE and SECOND-LINE cohorts. Detailed demographic information for all samples can be found in Supplementary Table 1.
Specific samples for assay verification were selected and processed based on the WHO HIVResNet HIV Drug Resistance Laboratory Operational Framework (Fig. 2). The testing panel consisted of 90 HIV-infected samples from 54 individuals, and six HIV-negative samples. Sample viral loads (VL) ranged from 1295 to 1,301,198 copies/mL, and encompassed subtypes common (CRF01_AE, B, C) and rare (CRF02_AG, D, F, G) in Australia (Supplementary Table 1). Sanger sequences were available from the protease and reverse transcriptase (PR-RT) region for samples from the ENCORE cohort, and PR-RT and integrase (IN) from the SECOND-LINE cohort. An uninfected plasma sample was sourced from the VIMM reference set (UNSW HREC HC200777) to act as a negative sample.
HIV-1 tiling PCR assay verification experimental and analytical flow.
Plasma samples were randomly shuffled and split into five batches of approx. 20 samples by an independent technician to ensure laboratory operators were blind to input, and that the assay could be run in five separate batches to assess reliability. Samples included as repeats for precision and reliability testing underwent 1:4 dilution with phosphate-buffered saline during the aliquoting stage, due to the increased volume requirements for these samples.
RNA extraction
For high-throughput extraction of RNA from samples prior to tiling PCR verification and benchmarking, the Roche MagNA Pure 96 Instrument was used. The DNA and Viral NA small volume kit (Roche #06543588001) and the pathogen universal 200 4.0 protocol was used, with input volume of 200 µl and output volume 50 µl. Extracted RNA was stored at − 80 °C until use.
Reverse transcription
Extracted RNA was reverse transcribed using SuperScript VILO IV (ThermoFisher #11756500). Briefly, 4 µl VILO enzyme was added to 8 µl sample in a final volume of 20 µl. The mixture was then incubated for 10 min at 25 °C, 20 min at 50 °C and 5 min at 85 °C, before being held at 4 °C until use.
Tiling PCR
To create the two tiling primer pools (A and B), the primers used to amplify non-overlapping segments of the genome were combined equivalently (Fig. 1), except for the segment six primers for which half volumes were added (Supplementary Table 2). Two PCR mastermixes were then made per run, containing 5 µl of cDNA, 4 µl of primer pool (1 × A, 1 × B) (10 mM) and 1X (10 µl) SuperFi II Green mastermix (ThermoFisher #12369050) in a final volume of 20 µl. Cycling was performed as follows: 98 °C for 30 secs followed by 40 cycles of 98 °C for 10 secs, 60 °C for 10 secs, and 72 °C for 90 secs, with a final extension at 72 °C for 5 min. The resulting amplicons were visualised on a 1% agarose gel to determine if PCR had been completed successfully. For each sample, the tiling PCR amplicons generated by the two mastermixes were then recombined in a 1:1 ratio prior to sequencing.
Library preparation and analysis: illumina sequencing
For high throughput sequencing of panel samples, Illumina based sequencing was performed. Equivalency of ONT and Illumina sequencing methods for tiling PCRs was demonstrated previously throughout the SARS-Cov-2 pandemic12 and was similarly found to be equivalent for the HIV tiling PCR (Supplementary Methods, Supplementary Fig. 1).
All panel samples, regardless of whether amplicons were visualised, were prepared for sequencing by cleaning using AMPure XP beads (Beckman Coulter #A63881) at a 1:1 ratio. Samples were then quantified using the Quant-iT PicoGreen dsDNA Assay Kit (ThermoFisher #P11496) as per the manufacturer’s protocol. Samples were then diluted to 0.2 ng/µl for Nextera XT library preparation (Illumina #FC-131–1096), which was performed as per standard manufacturer’s protocol except all volumes were halved. The resulting libraries were pooled equimolarly based on PicoGreen quantitation prior to sequencing. If the sample concentration was too low and did not allow for equimolar pooling, a maximum volume of 8 µl of prepared library was added to the pool. Samples were sequenced using a MiSeq device by the Ramaciotti Centre for Genomics (UNSW Sydney, Australia).
The resulting reads were assembled using a previously published virus assembly pipeline (https://github.com/laulambr/virus_assembly) on the UNSW computational cluster, Katana. The pipeline was modified to include trimming of the tiling PCR primers through the addition of the primers (Supplementary Table 2) as a database. The resulting de novo assemblies were then visualised using Geneious Prime 2 (2023.2.1). If the pipeline failed to produce an adequate assembly (e.g. if more than one assembly was generated per sample), a map to reference process was used as a secondary assembly method. Here, the trimmed reads generated by the pipeline were first mapped to the HIV-1 reference sequence HXB2 (accession: K03455) using the Geneious assembler (medium sensitivity). A secondary mapping of all reads to the consensus of that first assembly was then performed to account for the variability found in the HIV-1 genome. In all cases, the final consensus was taken as the major (> 50%) nucleotide at each position. For quality control, minimum acceptable coverage at any nucleotide position was 20 reads: to improve visualisation and identification of inadequate sequencing, a gap (−) was used in the consensus sequence if < 20 reads were present at any site.
Results
Advantage of a tiling PCR for HIV-1 long range sequencing
One advantage of moving to an NGS method for HIV-1 sequencing is the improvement in coverage over a larger genome section compared to what is possible using Sanger sequencing (< 1.5 kb). Using our novel tiling PCR assay, we found that all 90 samples from the HIV-1 testing panel were able to be successfully amplified, with 95% of samples generating a consensus length of > 4.5 kb (Fig. 3). This included amplification of samples of rarer subtypes such as F and G. The HIV-1 tiling PCR was found to generate over 80% coverage of gag and pol in 99% and 86% of samples respectively, increasing the genome length available for analysis for these samples. Env was not targeted as part of this assay.
Sequence coverage of panel samples using the HIV-1 tiling assay. Any site where under 20 reads were generated, that is where gaps or an N was present in the consensus sequence, is represented in black. The location of notable drug resistance mutations (DRMs) is highlighted in navy on the representative genome. Samples are clustered first by viral load, then by subtype.
Tiling PCRs are also advantageous due to the lower time and cost required to generate genome sequence. As such, the cost of running the tiling method was estimated at approximately $62 AUD/sample (October 2023) if running 48 samples on an ONT flow cell, or $120 AUD/sample (October 2023) if running 96 samples on a MiSeq device as per the manufacturer’s protocol. The time taken to generate a sequence was approximately one day if using ONT sequencing; approximately eight hours are required from starting the RNA extraction of the plasma sample through to library loading, including approximately five hours of direct hands-on time. Illumina sequencing adds approximately one extra day to the protocol due to the increased number of steps required for library preparation.
Precision and reliability of the HIV-1 tiling PCR over the 5’ HIV-1 genome
Precision of the HIV-1 tiling PCR assay over the entire 5’ genome region was assessed by including five repeats of one sample in a single batched assay run. Three such samples, each repeated five times in a batch, were included in the HIV-1 testing panel. For each sample, 10 total pairwise comparisons were performed, by comparing each sequence to every other sequence generated from the one sample in the one batched assay run. All possible comparisons for all three samples shared more than 99% genetic identity across the entire sequenced region (Table 1).
Reliability was assessed by again comparing sequences generated from the same sample, with each sample included once in five independent runs. The HIV-1 tiling assay was highly reliable across the entire sequenced region (Table 1), with all three samples having > 99% genetic identity in all pairwise comparisons across the five independent samples.
Additionally, 12 samples were included in duplicate as part of the HIV-1 testing panel. Of these 12 samples, when pairwise analysis of the entire sequenced region was performed, 11 were greater than 98% identical across the two repeats (Supplementary Table 3).
Utility of tiling PCR for diagnostic HIV-1 sequencing: assay sensitivity
We then assessed the results of the tiling PCR assay for use as a diagnostic assay for HIV-1 drug resistance sequencing, following the WHO HIVResNet HIV Drug Resistance Laboratory Operational Framework (Fig. 2)25. Consensus sequences generated by tiling PCR were broken into subgenomic regions, encompassing protease-reverse transcriptase (PR-RT) and integrase (IN), and these sequences were compared with historical Sanger sequencing as per the WHO HIVRestNet Framework25. Based on these guidelines, samples were also split into two categories for analysis, > 5000 copies/mL or < 5000 copies/mL.
Using a highly stringent cut-off of complete gene coverage, we found over 95% of samples with both low (< 5000 copies/mL) and high (> 5000 copies/mL) viral loads amplified PR. However, sensitivity was slightly diminished within RT, with 92.3% of samples with a VL > 5000 copies/mL and 76.0% samples with a VL < 5000 copies/mL having a complete RT. 95.4% of high VL samples and 84% of low VL samples had a complete IN region amplified using the tiling PCR.
Utility of tiling PCR for diagnostic HIV-1 sequencing: consensus sequence accuracy
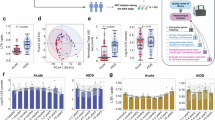
For the subgenomic regions sequenced using tiling PCR, 79.5% of PR-RT and 93.3% of IN sequences shared > 98% identity with their Sanger equivalents (Fig. 4). For the higher viral load samples (VL > 5000 copies/mL), 91% and 100% of PR-RT and IN sequences were > 98% identical to their Sanger equivalents respectively. However, concordance was 74% and 80% for PR-RT and IN when the viral load fell below 5000 copies/mL. Drops in sequence accuracy were due to sequence drop out within the subgenomic region: for the 10 samples with VL > 5000 copies/mL where the initial shared identity was < 98% in PR-RT, the average shared identity rose to 99.0% when gap sites were stripped. The tiling PCR did not generate any HIV-1 sequence from any of the HIV-negative control samples.
Shared genetic identity between the HIV-1 tiling PCR method and diagnostic Sanger sequencing of HIV-1 infected samples. Consensus sequences generated by a novel tiling PCR and next-generation sequencing were compared to prior Sanger sequences of PR-RT (ENCORE and SECOND-LINE samples) and IN (SECOND-LINE samples only). Matched data is available for 88 samples in PR-RT, and 15 samples in IN. Shared genetic identity between matched sequences > 98% is highlighted in purple. Note that shared identity between samples at different locations in the panel is due to samples from the same participant being run multiple times for precision and reliability testing.
Utility of tiling PCR for diagnostic HIV-1 sequencing: assessment of drug resistance mutations
We also assessed assay accuracy through the proportions of DRMs present or absent when comparing the new sequences to the original sequencing results, using the most updated version of the Stanford Drug Resistance algorithm (HIVDB version 9.5.1; February 2024)33. A total of 107 DRM sites were identified within the Sanger sequences, and 99 of these sites were also found with the tiling assay. Eight DRMs in RT were missed by the tiling PCR due to sequencing failure over those codon positions. Of the 99 DRM sites identified by tiling PCR, 20 had mismatching outputs to the Sanger results. All of these were due to an ambiguous character being called in the Sanger consensus sequence, but a nucleotide being resolved in the tiling consensus sequence.
An additional seven DRMs were identified in the tiling PCR consensus sequences that were not present in the Sanger data. Five samples were found to encode resistance to NNRTIs, (N348I), not previously identified as the site was outside the genomic region amplified for Sanger sequencing. An additional NRTI-associated DRM, A62V, was observed in one sample with a low viral load (1977 copies/mL), which had failed initial diagnostic sequencing (Supplementary Table 1). Finally, one DRM (V106I) was identified in a sample with a low viral load (4946 copies/mL), however this was not seen in the original Sanger generated consensus. The sample was sequenced twice as part of the panel, and V106I was identified in both repeats. To confirm the presence of this mutation, sequencing was also performed using an alternative NGS based method4 and the V106I mutation was observed at the subconsensus level in this assay. It is therefore likely this mutation was naturally occurring in this sample; however the true proportion of the variant is difficult to detect accurately between methods due to the low viral load of the sample.
Utility of tiling PCR for diagnostic HIV-1 sequencing: subgenomic precision and reliability
Precision of the HIV-1 tiling PCR assay over the subgenomic areas was assessed by including five repeats of one sample in a single batched assay run. The HIV-1 tiling assay was highly precise across subgenomic regions, with all possible comparative sequences (10 total pairwise comparisons per sample) sharing more than 99% genetic identity in PR-RT and IN (Table 2). Reliability was assessed by comparing subgenomic sequences generated from three samples included once each in five independent runs. The assay was also highly reliable within the subgenomic regions, with all three reliability samples sharing more than 99% genetic identity in all pairwise comparisons (Table 2).
Discussion
We have successfully developed a tiling PCR protocol that is able to quickly and efficiently generate HIV-1 consensus sequence from varying subtypes and viral loads. We have demonstrated the diagnostic utility of this method for DRM detection by detecting complete and comparable PR-RT and IN regions in > 90% of samples with a viral load > 5000 copies/mL. This method was also able to identify seven additional DRMs not originally found due to the sensitivity and length limitations of Sanger sequencing. The tiling PCR for HIV-1 is therefore a promising solution for generating HIV-1 sequence using NGS for patient care.
The assay success of tiling PCR approaches is particularly reliant on the successful binding of multiple primers to allow for the amplification of the genome in multiple segments9,10,23,34. Previous tiling PCRs have been designed in outbreak scenarios for use on specific viruses where the overall genetic diversity is low9,10,23,34. Attempts to develop tiling PCRs for other highly genetically diverse viruses have resulted in lower overall success rates: as an example, a tiling PCR designed for HCV was only able to sequence more than > 50% of the genome in 57/90 cases35. As such, the reliability of our novel tiling PCR approach for HIV-1 is particularly noteworthy. The success of our assay may be in part due to the use of the SuperFi II enzyme, which has a universal annealing temperature. This enzyme may have allowed for more efficient binding of the primer pool, given the range of melting temperatures present in the primer sets.
The primer set in our approach may not yet be optimal, however, as some losses of sensitivity were observed when the viral load dropped below 5000 copies/mL, with segment drop out particularly observed within the RT gene. Future work optimising the balance of primers in the pool to minimise competition for binding in these lower VL samples may help to overcome this issue. Extraction of larger volumes of sample in cases where the viral load is known to be low, increasing the concentration of HIV-1 RNA present in the eluate, may also overcome this issue. However, despite the primer set being designed to predominately bind subtype B, C and CRF01_AE viruses, the method was still able to generate sequence from samples with other subtypes, such as F and G28,29. While some balancing may therefore still optimise sequencing from lower VL samples, the primer set may have a greater utility than initially expected based on primer design when amplifying from non-B subtypes.
While the tiling PCR was able to amplify the 5’ half of the HIV-1 genome (~ 6 kb), we found during optimisation (Supplementary Data) that targeting of the entire genome hampered robust sequence generation of the 5’ end. This was likely in part due to the high genetic variability and longer length of env36, and the relative difficulty of designing a comprehensive primer set over this region. However, given env remains an important target for novel therapies and/or curative strategies13, if this method is to be used for DRM sequencing future adaptations may have to incorporate strategies for env sequencing to ensure that standard of care can continue.
Some differences were noted in the DRMs sequenced using tiling PCR compared to Sanger sequencing, as 20 originally noted DRMs were not called at the consensus level due to an ambiguous nucleotide being resolved at that site. Given Sanger sequencing is only capable of distinguishing viral population diversity down to the 20%, while NGS has been found to be able to clearly distinguish genetic diversity of at least 5% frequency, this result is not surprising37. Over 90% of previously observed DRMs in our data set were recovered using the tiling PCR; only eight previously identified DRMs were missed due to coverage dropout in low viral load (< 5000 copies/mL) samples. Moreover, an equivalent number of newly identified mutations were found using tiling PCR that were missed due to the technical limitations of the original Sanger-based assay. Notably the same pattern has also been observed in other studies looking at NGS, though with a focus on sequencing deeper, not wider38,39,40.
The tiling PCR methodology is designed not only as an alternative method for HIV-1 NGS-based sequencing for DRM surveillance, but also for public health use as part of a near-real-time molecular epidemiological surveillance system. Such surveillance is reliant on a cohesive data set between jurisdictions, and sequencing turn-around time must be minimised to ensure a timely response can occur. The tiling PCR, which amplified the 5’ half of HIV-1 with a turnaround time of under 24 h, makes molecular epidemiological surveillance of HIV-1 one step closer in an Australian setting. Moreover, given the success of generating sequence of rarer subtypes, as well as the considerably low cost and time to run the assay, the tiling PCR may allow for surveillance of HIV-1 in surrounding regions where commercial assays are prohibitively expensive, or technological assistance for these methods is not routinely available. Given the increasing awareness of the role played by cases of HIV-1 acquired outside Australia in the local epidemic41,42, the development of a cost-effective viral sequencing assay that is usable in both a local and regional setting is key for timely surveillance of HIV-1 transmission in the Indo-Pacific region.
In conclusion, we have successfully developed a tiling PCR protocol that is able to robustly generate HIV-1 sequence from archived patient samples. This method has been able to fulfill the requirements of the WHO HIVResNet HIV Drug Resistance Laboratory Operational Framework for samples > 5000 copies/mL. As such, this novel designer tiling PCR is a promising method for routine long-range sequencing of HIV-1 in Australia for the purpose of both patient care and routine molecular epidemiology.
Data availability
Sequence data is available on GenBank, accessions PQ893199–PQ893288. Primers for the tiling PCR are available in Supplementary Table 2. Requests for additional data can be made to the corresponding author.
References
Bavinton, B. R. et al. Viral suppression and HIV transmission in serodiscordant male couples: An international, prospective, observational, cohort study. Lancet HIV 5, e438–e447 (2018).
Rodger, A. J. et al. Risk of HIV transmission through condomless sex in serodifferent gay couples with the HIV-positive partner taking suppressive antiretroviral therapy (PARTNER): Final results of a multicentre, prospective, observational study. The Lancet 393, 2428–2438 (2019).
Cohen, M. S. et al. Antiretroviral therapy for the prevention of HIV-1 transmission. N. Engl. J. Med. 375, 830–839 (2016).
Jenkins, F. et al. Validation of an HIV whole genome sequencing method for HIV drug resistance testing in an Australian clinical microbiology laboratory. J. Med. Virol. 95, e29273 (2023).
Ayitewala, A., Ssewanyana, I. & Kiyaga, C. Next generation sequencing based in-house HIV genotyping method: Validation report. AIDS Res. Ther. 18, 64 (2021).
Saravanan, S. et al. Evaluation of two human immunodeficiency virus-1 genotyping systems: ViroSeq™ 2.0 and an in-house method. J. Virol. Methods 159, 211–216 (2009).
Ni, Y., Liu, X., Simeneh, Z. M., Yang, M. & Li, R. Benchmarking of Nanopore R10.4 and R9.4.1 flow cells in single-cell whole-genome amplification and whole-genome shotgun sequencing. Comput. Struct. Biotechnol. J. 21, 2352–2364 (2023).
Arslan, S. et al. Sequencing by avidity enables high accuracy with low reagent consumption. Nat. Biotechnol. 42, 132–138 (2024).
Freed, N. E., Vlková, M., Faisal, M. B. & Silander, O. K. Rapid and inexpensive whole-genome sequencing of SARS-CoV-2 using 1200 bp tiled amplicons and Oxford Nanopore Rapid Barcoding. Biol. Methods Protoc. 5, bpaa014 (2020).
Tyson, J. R. et al. Improvements to the ARTIC multiplex PCR method for SARS-CoV-2 genome sequencing using nanopore. bioRxiv 2020.09.04.283077 (2020) https://doi.org/10.1101/2020.09.04.283077.
Plitnick, J. et al. Whole-genome sequencing of SARS-CoV-2: Assessment of the ion torrent ampliseq panel and comparison with the Illumina MiSeq ARTIC protocol. J. Clin. Microbiol. 59, e00649-e721 (2021).
Bull, R. A. et al. Analytical validity of nanopore sequencing for rapid SARS-CoV-2 genome analysis. Nat. Commun. 11, 6272 (2020).
Temereanca, A. & Ruta, S. Strategies to overcome HIV drug resistance-current and future perspectives. Front. Microbiol. 14, 1133407 (2023).
Jin, H., Li, D., Lin, M. H., Li, L. & Harrich, D. Tat-based therapies as an adjuvant for an HIV-1 functional cure. Viruses 12, 415 (2020).
Horsburgh, B. A. et al. Next-generation sequencing methods for near-real-time molecular epidemiology of HIV and HCV. Rev. Med. Virol. 34, e70001 (2024).
Obasa, A. E., Engelbrecht, S. & Jacobs, G. B. Near full-length HIV-1 subtype B sequences from the early South African epidemic, detecting a BD unique recombinant form (URF) from a sample in 1985. Sci. Rep. 9, 6227 (2019).
Obasa, A. E., Ambikan, A. T., Gupta, S., Neogi, U. & Jacobs, G. B. Increased acquired protease inhibitor drug resistance mutations in minor HIV-1 quasispecies from infected patients suspected of failing on national second-line therapy in South Africa. BMC Infect. Dis. 21, 214 (2021).
Parikh, U. M., McCormick, K., van Zyl, G. & Mellors, J. W. Future technologies for monitoring HIV drug resistance and cure. Curr. Opin. HIV AIDS 12, 182–189 (2017).
Redd, A. D., Quinn, T. C. & Tobian, A. A. R. Frequency and implications of HIV superinfection. Lancet Infect. Dis. 13, 622–628 (2013).
Casadellà, M. & Paredes, R. Deep sequencing for HIV-1 clinical management. Virus Res. 239, 69–81 (2017).
Ji, H. et al. Bioinformatic data processing pipelines in support of next-generation sequencing-based HIV drug resistance testing: The Winnipeg Consensus. J. Int. AIDS Soc. 21, e25193 (2018).
Ji, H. et al. Are we ready for NGS HIV drug resistance testing? The second ‘Winnipeg Consensus’ symposium. Viruses 12, 586 (2020).
Quick, J. et al. Multiplex PCR method for MinION and Illumina sequencing of Zika and other virus genomes directly from clinical samples. Nat. Protoc. 12, 1261–1276 (2017).
Metrichor & Oxford Nanopore Technologies. EPI2ME. https://labs.epi2me.io/ (2024).
HIVREsnet. WHO HIVResNet HIV Drug Resistance Laboratory Operational Framework. (2020).
ENCORE1 Study Group. Efficacy of 400 mg efavirenz versus standard 600 mg dose in HIV-infected, antiretroviral-naive adults (ENCORE1): A randomised, double-blind, placebo-controlled, non-inferiority trial. The Lancet 383, 1474–1482 (2014).
SECOND-LINE Study Group. Ritonavir-boosted lopinavir plus nucleoside or nucleotide reverse transcriptase inhibitors versus ritonavir-boosted lopinavir plus raltegravir for treatment of HIV-1 infection in adults with virological failure of a standard first-line ART regimen (SECOND-LINE): A randomised, open-label, non-inferiority study. The Lancet 381, 2091–2099 (2013).
Di Giallonardo, F. et al. Increased HIV subtype diversity reflecting demographic changes in the HIV epidemic in new south wales, Australia. Viruses 12, 1402 (2020).
Castley, A. et al. A national study of the molecular epidemiology of HIV-1 in Australia 2005–2012. PLoS ONE 12, e0170601 (2017).
Hiener, B. et al. Identification of genetically intact HIV-1 proviruses in specific CD4+ T cells from effectively treated participants. Cell Rep. 21, 813–822 (2017).
Sanders-Buell, E., Salminen, M. & McCutchan, F. Sequencing primers for HIV-1. Human retroviruses and AIDS (1995).
Chrysostomou, A. C., Topcu, C., Stylianou, D. C., Hezka, J. & Kostrikis, L. G. Development of a new comprehensive HIV-1 genotypic drug resistance assay for all commercially available reverse transcriptase, protease and integrase inhibitors in patients infected with group M HIV-1 strains. Infect. Genet. Evol. 81, 104243 (2020).
Liu, T. F. & Shafer, R. W. Web resources for HIV Type 1 genotypic-resistance test interpretation. Clin. Infect. Dis. 42, 1608–1618 (2006).
Quick, J. et al. Real-time, portable genome sequencing for Ebola surveillance. Nature 530, 228–232 (2016).
Yakovleva, A. et al. Hepatitis C Virus in people with experience of injection drug use following their displacement to Southern Ukraine before 2020. BMC Infect. Dis. 23, 446 (2023).
Li, G. et al. An integrated map of HIV genome-wide variation from a population perspective. Retrovirology 12, 18 (2015).
Houldcroft, C. J., Beale, M. A. & Breuer, J. Clinical and biological insights from viral genome sequencing. Nat. Rev. Microbiol. 15, 183–192 (2017).
Alidjinou, E. K. et al. RNA and DNA Sanger sequencing versus next-generation sequencing for HIV-1 drug resistance testing in treatment-naive patients. J. Antimicrob. Chemother. 72, 2823–2830 (2017).
Tzou, P. L. et al. Comparison of an in vitro diagnostic next-generation sequencing assay with Sanger sequencing for HIV-1 genotypic resistance testing. J. Clin. Microbiol. 56, e00105-e118 (2018).
Mohamed, S. et al. Clinical impact of ultra deep versus Sanger sequencing detection of minority mutations on HIV-1 drug resistance genotype interpretation after virological failure. BMC Infect. Dis. 14, O1 (2014).
Taiaroa, G. et al. Characterising HIV-1 transmission in Victoria, Australia: a molecular epidemiological study. Lancet Reg Health West Pac 47, 101103 (2024).
Di Giallonardo, F. et al. Subtype-specific differences in transmission cluster dynamics of HIV-1 B and CRF01_AE in New South Wales, Australia. J. Int. AIDS Soc. 24, e25655 (2021).
Acknowledgements
This research includes analysis performed using the computational cluster Katana, supported by Research Technology Services at UNSW Sydney. Illumina sequencing of the tiling PCR samples was performed by the Ramaciotti Centre for Genomics at UNSW Sydney. Samples of HIV-1 BaL, used as a positive control for PCR assays, were kindly donated by Dr Scott Ledger from the Kirby Institute, UNSW. We would like to acknowledge all participants who donated samples as part of the ENCORE and SECOND-LINE cohorts. We would also like to acknowledge all those involved in the ENCORE and SECOND-LINE studies, as well as the AMR biobank team, for their help in collecting and accessing these samples.
Funding
This work was supported by an Australian Government Department of Health and Aged Care Medical Research Future Fund (MRFF) grant (FSPGN000047).
Author information
Authors and Affiliations
Consortia
Contributions
BAH, RAB and FDG conceived the experiments. BAH designed the method. BAH, AA, FJ, HL and CSPF carried out the experiments, WR, ADK, SVH, RAB and FDG supervised. BAH performed the analysis and wrote the original draft manuscript. AL, LC, WR, ADK, RAB and FDG acquired funding for the H2Seq project. All authors provided feedback and helped shape the research, analysis and manuscript.
Corresponding author
Ethics declarations
Competing interest
The authors declare no competing interests.
Ethical approval
This study was approved by the St Vincent’s Hospital Human Research Ethics Committee, reference 2022/ETH01204.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Horsburgh, B.A., Abayasingam, A., Jenkins, F. et al. Development and verification of a novel tiling PCR method for long-range HIV-1 sequencing in a diagnostic setting. Sci Rep 15, 22057 (2025). https://doi.org/10.1038/s41598-025-03190-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-03190-6