Abstract
Species-specific efficacy necessitates accurate identification of medicinal leeches, but standard DNA barcoding often fails with degraded DNA from traditional medicines. This deficiency highlights the need for mini-barcoding. This study aimed to develop and validate mini-barcode markers for three Chinese Pharmacopoeia-listed leech species: Whitmania pigra, Whitmania acranulata and Hirudo nipponia. Four novel mini-barcode primer sets (ND1F1/R1, 12SF1/R1, 16SF1/R1 and COX1F1/R1) were developed and validated using seven morphologically identified specimens and subsequently tested on 16 commercial leech products. DNA extractions were performed using both single-tube and column purification kits, with the latter yielding superior DNA quality and meeting the requirements for following PCR amplification. The PCR results confirmed the validation of four candidate mini-barcodes targeting specific genetic regions, which produced results in 13 out of 16 commercial leech products. Mini-barcode sequences from morphologically identified W. pigra specimens exhibit > 95% identity to the complete ND1, 12S rDNA, 16S rDNA, and COX1 sequences (EU304459), whereas sequences from H. nipponia and W. acranulata show < 85% identity, and among leech-derived products only the proprietary Chinese medicine Maxuekang exhibits lower identity. Both the optimal partition of ASAP and phylogenetic tree identified three distinct groups correlating with the morphological species: W. pigra, W. acranulata, and H. nipponia. Mislabeled species have been uncovered in proprietary Chinese medicine, notably the claimed Hirudo nipponia, which was replaced by W. pigra. The results highlight the value of mini-barcodes in enhancing product quality control and offer a reliable method for accurate species identification in traditional and commercial leech-based medicines. This advance supports safer and more effective utilization of medicinal leeches and advocates for their integration into regulatory standards.
Similar content being viewed by others
Introduction
Leeches, members of the family Clitellata within the phylum Annelida, are characterized by the fact that their posterior suckers are used for attachment to surfaces, while their anterior suckers are specialized for feeding on small invertebrates or blood-feeding on hosts1,2,3. Beyond their ecological significance, leeches have attracted considerable attention due to their unique medicinal properties, which trace back over 2,000 years and remain relevant in contemporary medical sciences, owing to their therapeutic applications4,5,6. The recognition of certain blood-feeding leeches in traditional medicine and other therapeutic practices is largely attributed to their remarkable anticoagulant properties7,8. Leeches are valued for their ability to reduce blood stasis and promote fluid movement, making them effective in treating conditions such as edema and congestion9. Their anticoagulant is crucial in the treatment of cardiovascular diseases and in the control of hypercoagulable states in some chronic kidney diseases such as primary nephrotic syndrome and lupus nephritis10,11,12,13. In addition, for diseases such as diabetes, organ fibrosis, tumors, arthritis and diseases that require hypolipidemia and anti-inflammatory treatment, treatment with leeches can have a powerful effect on improving blood circulation and reducing inflammation14,15. Therefore, the wide spread use of leeches in medical practice is supported by their diverse biological functions that address various aspects of these diseases.
The therapeutic benefits of leeches are well documented, and it is crucial to identify leech species ensuring both safety and effectiveness in medical treatments. A systematic review of 277 cases found a significant success rate of approximately 78% with leech therapy for plastic and reconstructive surgery, and the selection of leech species was found to significantly influence the clinical outcomes and safety of medical leech therapy16. The evaluation of different leech species with regard to their treatment capability is also of crucial importance in traditional medicine. The 2020 edition of the Chinese Pharmacopoeia recognizes three medicinal leech species for use in traditional medicine: Whitmania pigra (commonly known as Mahuang), Whitmania acranulata, and the blood-feeding leech Hirudo nipponia (commonly known as Shuizhi). H. nipponia and Poecilobdella manillensis (another blood-feeding leech) have similar anticoagulant properties but possess 50–60% distinct different amino acid residues17, indicating that they have different immunosuppressive activities and anticoagulant mechanisms. This was further illustrated by Cheng18, who discovered distinct different stereoscopic structures and configurations in antithrombin peptides from H. nipponia, W. pigra, W. acranulata, and P. manillensis which exhibited different stereoscopic structures and configurations. Different leech species secrete a variety of bioactive substances, such as Antistasin from Haementeria officinalis and Helobdella austinensis19,20, Guamerin from Hirudo nipponia21, and Eglins, Hirudin, Saratin, as well as Bdellins from Hirudo medicinalis22,23,24. These substances exhibit diverse effects, including analgesic, anti-inflammatory, platelet inhibitory, anticoagulant and thrombin regulatory functions25. This diversity of bioactive substances among leech species highlights the need for precise species selection in therapy to exploit specific therapeutic benefits and minimize potential risks. Consequently, ignoring the different medicinal mechanisms of different leech species not only undermines the potential therapeutic benefits but also poses significant risks to the safety and effectiveness of medicinal leech clinical therapy and related herbal preparations, such as decoction pieces.
DNA barcoding is a standardized method utilizing a specific, sufficiently variable and short DNA fragment to identify species based on previously barcoded specimens26. It is helpful in identifying animal and plant species and has been proven to be effective27,28. The European medicinal leech Hirudo medicinalis is approved by the United States Food and Drug Administration (FDA) as a prescription medical device. It is a rare example of an animal used directly in modern medicine. However, recent DNA barcoding studies have revealed that what is commonly sold as the European medicinal leech, Hirudo medicinalis, actually consists of at least three different species8,29,30,31. This finding highlights the importance of accurate species identification in medical therapy and scientific investigations. DNA barcoding has also been applied to various medicinal animal raw materials, such as earthworms, deer antlers and tortoise shells32,33,34. Despite its widespread use, there is a notable lack of DNA barcoding studies on processed leech-derived medicinal ingredients. This disadvantage may be due to the complexity of these medicines, which often contain multiple ingredients and are processed in a way that make DNA identification difficult35. Traditional preparation methods such as stir-frying, stewing, boiling and steaming can damage DNA, resulting in quite short DNA sequences. This makes it difficult for universal COI barcode primers to amplify the entire target fragment. A study conducted by Xin et al.36 highlighted these challenges and found that the success rate of DNA amplification was particularly low due to DNA degradation. However, the DNA mini-barcoding approach effectively solves the amplification problems caused by DNA degradation in processed medicinal materials, because it requires only a shorter DNA template than the conventional DNA barcoding while still providing sufficient specificity37. Based on 20 empirical datasets covering approximately 30,000 specimens (5,500 species) and six clade-specific datasets from GenBank covering approximately 98,000 specimens (over 20,000 species), medium-length mini-barcodes (more than 200 bp) function similarly full-length barcodes for species-level and specimen-level identification38. Mini-barcoding can be considered as a complementary barcoding technique for identifying and authenticating traditional medicine ingredients, helping to ensure consistency and quality in the traditional medicine industry.
Given the significant findings that the FDA-approved medicinal leech, Hirudo medicinalis, includes several distinct species, the precision of species identification in commercial leech derived medicinal products is of paramount importance to ensure therapeutic efficacy and safety. These results highlight the necessity of using mini-barcoding techniques for species identification in traditional medicine, where the specificity of the active biological compounds is crucial for the desired therapeutic outcomes. This study aims to develop specific mini-barcoding primers based on mitochondrial genomes of three medicinal leech species listed in the Chinese Pharmacopoeia to validate the effectiveness of these primers on morphologically identified specimens of three leeches and to ensure that the leech species from leech derived products can be accurately identified, including traditional medicines.
Results
Leech DNA extraction and comparison of DNA quality
A total of 11 leech specimens representing Whitmania pigra, Whitmania acranulata, Hirudo nipponia and Poecilobdella manillensis, were morphologically identified (Table 1), and the former three species were also authenticated by mini-barcoding in the subsequent analysis to ensure accurate identification of these reference specimens. DNA extraction was initially performed on 11 leech specimens and leech derived products, including two traditional Chinese medicines (TCMs) and 14 proprietary Chinese medicines (PCMs), using the One-tube Universal Sample DNA Extraction Kit. The DNA extracted from 11 leech specimens were all successfully subjected to PCR amplification. However, DNA isolated from leech derived products using this kit did not produce successful PCR results in subsequent PCR amplification, with the exception of two TCMs samples, one hang-dried and one processed medicinal leeches (sample IDs were YQ016 and YQ015, respectively). Consequently, DNA extractions of 14 PCMs were performed again using the Ezup Column Animal Genomic DNA Purification Kits. DNA extracted using the column-based extraction method generally showed higher quality compared to that obtained using the one-tube extraction method repeated three times, as evidenced by the OD260/OD280 ratios (Fig. 1b). The concentrations of DNA extracted from PCMs using two different kits varied from 1.9 to 522.2 ng/μL.
General information and the quality of extracted DNA from 16 leech derived products. (a) The profiles of each leech derived product contain detailed information about its form (capsules, pills or tablets), pharmaceutical production sites and batch number. (b) A heatmap comparing two different extraction methods in leech derived products based on OD260/OD280 ratios. Each extraction method was repeated three times. The left part of the heatmap represents DNA quality obtained using Ezup Column Animal Genomic DNA Purification Kit, and the right part corresponds to DNA quality using the One-tube Universal Sample DNA Extraction Kit. The gradient bar represents a range of OD260/OD280 ratio values, with 1.0 indicating the lightest shade and 2.0 indicating the darkest shade, where higher values indicated better DNA quality. The abbreviations represent the names of leech products, detailed in Table 1.
Design and validation of specific mini-barcoding primers
Four candidate primer pairs for mini-barcoding were designed using Oligo 7 (Table 2) and refer to an alignment of the mitochondrial genome sequences of Whitmania pigra, Whitmania acranulata and Hirudo nipponia. Each of these primer pairs for ND1, 12S rDNA, 16S rDNA and COX1 sequences were designed to generate amplicons with the following sizes: 251 bp, 212 bp, 158 bp and 210 bp, respectively. Primer BLAST analysis was performed using these primers to confirm the accurate and specific identification of the corresponding leech species in the NCBI GenBank database. The specificity of these primers was also assessed in vitro by performing PCR on leech specimens (see Table 2 for PCR primer pairs and protocols) and the results were visualized through gel electrophoresis. The primer pair ND1F1/R1 achieved the best amplification results, followed by 12SF1/R1, 16SF1/R1 and COX1F1/R1 (Fig. 2). Consequently, these primer pairs were used in the subsequent PCR using DNA templates extracted from 16 leech derived products. Consistent with the PCR results obtained from leech specimens, the primer pair ND1F1/R1 showed the most effective amplification, while the primer pairs 12SF1/R1, 16SF1/R1 and COX1F1/R1 showed progressively decreasing amplification efficiency (Fig. 2). All amplicons were then sequenced and run in NCBI’s BLAST. The results showed that all but two PCMs matched the corresponding species very well. The PCR performed failed for sample CZTL and sequencing did not yielded any sequences for sample QZ. Furthermore, all primer pairs either failed to amplify DNA or produced nonspecific fragments when used with DNA templates from Poecilobdella manillensis specimens (a common adulterant for medicinal leeches).
Matrix layout of the PCR profiles of four primer pairs across 23 leech samples. The abbreviations represent the names of leech products, detailed in Table 1. The vertical bar represents the total number of successfully amplified sequences for each leech sample, while the horizontal bar represents the total number of samples that successfully amplified the sequences for each primer pair. The filled or empty dots indicate the success or failure of the corresponding PCR (results of leech derived products were heavily highlighted with gray background), and the number in each dot represents the ratio of the actual sequence obtained to the target amplicon through Sanger single-stranded chain termination sequencing, and the symbol " × " in the dots indicates the presence of PCR products but the absence of sequencing.
Comparison between mini-barcoding sequences and reference sequences
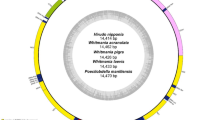
Target regions were separately amplified using Sanger sequencing with four primer pairs (ND1F1/R1, 12SF1/R1, 16SF1/R1, and COX1F1/R1). After removing primer sequences and low-quality regions, the final sequences from the four targets ranged from 101 to 252 base pairs. Specifically, the ND1, 12S rDNA, 16S rDNA and COX1 Sequences had average lengths of 118 bp, 153 bp, 208 bp, and 156 bp, respectively (Supplementary Fig. S1). The comparison of the positions and lengths of newly obtained amplicons with the complete sequences of ND1, 12S rDNA, 16S rDNA and COX1 from Whitmania pigra (EU304459) was shown in Fig. 3. The color of each arc indicated an identity of greater than 95% to the blast results of each newly amplified sequence with the reference sequences of W. pigra (the middle black circle in Fig. 3), while desaturated colors in arcs indicated sequences with identity less than 95% compared to reference sequences. Arcs within the circle represent the mini-barcode sequences amplified from morphologically identified leech specimens. Each amplified sequence from a morphologically identified W. pigra specimen (LS004, LS009 and LS018) is nearly identical (> 95%) to the corresponding W. pigra reference sequence (EU304459), confirming their classification as W. pigra and validating the accuracy of barcoding identification. The arches of Hirudo nipponia (LS002, LS077 and LS078) and Whitmania acranulata (LS013) were somewhat desaturated (less than 85%), suggesting that H. nipponia and W. acranulata were genetically distinct from W. pigra (Fig. 3). Arcs outside the circle represent mini-barcoding sequences obtained from leech derived products. The sequences for QZ and CTZL were not included due to unsuccessful PCR amplification or sequencing procedure. Compared to W. pigra, only the mini-barcode sequences of a proprietary Chinese medicine, MXK, showed lower identity, while all other sequences from leech derived products were almost identical.
Circular comparison between sequences of Whitmania pigra from GenBank (EU304459) and the mini-barcoding sequences obtained from leech samples visualized by BRIG. The black circle represents the complete ND1, 12S rDNA, 16S rDNA and COX1 sequences obtained from Whitmania pigra (EU304459). Colored arcs of different lengths within the circle represent the mini-barcoding sequences amplified from morphologically identified leech specimens, while arcs outside the circle represent mini-barcoding sequences obtained from leech derived products (including one processed medicinal leech sample, one hang-dried medicinal leech and other commercial traditional medicines). The four solid or hollow dots after each arc located on ND1 consolidate the results and represent the presence or absence of the obtained mini-barcoding sequences of ND1, 12S rDNA, 16S rDNA and COX1. The visualized area shows that the percent identity similar genes between the Whitmania pigra reference sequences and other mini-barcoding sequences in BLASTn was at least 85%. Color saturation in the arcs indicates identity: 100% saturation represents complete identity, 50% represents 95% identity, and grays represent identity below 85%. The abbreviations represent the names of leech products, detailed in Table 1.
Identify leech species from leech derived products
In order to identify original leech species from leech derived products, a matrix comprising partial ND1, 12S rDNA, 16S rDNA and COX1 genes was constructed. These sequences were obtained from seven specimens representing three medical leech species and 16 leech derived products. In addition, GenBank sequences of W. pigra (EU304459), W. acranulata (KC688271 and KM655838) and H. nipponia (NC_023776 and MZ507570) were also incorporated in this matrix. Using the data matrix of ND1, 12S rDNA, 16S rDNA and COX1 sequence data, the molecular species delimitation method ASAP identified eight distinct partitions (Fig. 4 and supplementary Fig. S2). The best ASAP partition, indicated by the lowest ASAP scores and marked with an asterisk, distinguished three different groups corresponding to the morphological species: W. pigra (common name Mahuang), W. acranulata and H. nipponia (common name Shuizhi). All but MXK leech derived products, including traditional Chinese medicines (TCMs) and proprietary Chinese medicines (PCMs), along with W. pigra specimens (LS004, LS009 and LS018) clustered with the GenBank sequence of W. pigra (EU304459) as one group. MXK formed a second group together with H. nipponia specimens (LS002, LS077 and LS078 from this study, NC_023776 and MZ507570 from GenBank). The three W. acranulata specimens (KC688271 and KM655838 from GenBank and LS013 from this study) are grouped together as a third group. The ASAP result showed that among the leech derived products, only MXK contained Shuizhi (H. nipponia), all others were based on W. pigra (Mahuang), and none used W. acranulata. Furthermore, the best partition result from the ASAP analyses based on data from ND1 sequences only consistently supported the current species delimitation results (Figs. 4 and supplementary Fig. S2).
Species delimitation of all leech samples through ASAP based on a concatenated dataset of ND1, 12S rDNA, 16S rDNA and COX1 sequences. The four solid or hollow dots after each sample ID indicate the presence or absence of these sequences for each sample. The best output partition is marked with an asterisk, indicating the existence of three different species. The bar chart displays the results of different species delimitation schemes, with each column representing a unique partitioning result from the ASAP analysis. The numbers above the columns show the ASAP scores and the proposed group number. Colorful segments in each column represent proposed groups, with numbers in each segment indicating the number of leech species assigned to that proposed group. The images in the upper right corner showed the dorsal view of three morphologically identified leech species. GenBank sequences of W. pigra (EU304459), W. acranulata (KC688271 and KM655838) and H. nipponia (NC_023776 and MZ507570) were also included in this matrix and highlighted in boldface. The abbreviations represent the names of leech products, detailed in Table 1.
Discussion
The quality and quantity of DNA extracted from leech derived products, including traditional Chinese medicines (TCMs) and proprietary Chinese medicines (PCMs), play a crucial role in ensuring the correct identification of medicinal animals through DNA barcoding. The special features of TCMs and PCMs are the specific processing methods used in the preparation of raw materials, which often lead to a change in their external morphological properties and to DNA degradation. The application of DNA barcodes in TCMs and PCMs can be challenging due to the low quality and quantity of DNA after the manufacturing process39. Given the critical role of DNA quality and quantity in identifying medicinal animals through DNA barcodes, purity ratios measured with microvolume spectrophotometers become relatively variable when DNA concentrations drop below 50 ng/µl40. This is consistent with the current results of low concentration of DNA extracted from PCMs and large fluctuations in A260/A280 ratios observed in the current study (Fig. 1b). To address these challenges, One-tube Universal Sample DNA Extraction Kit was initially used, completing the entire process in a single centrifuge tube and extending the sample cleavage time to minimize DNA losses. However, this method can increase DNA concentration but also introduces significant contamination, potentially affecting subsequent amplification experiments. In contrast, column-based extraction methods, although sacrificing some of the DNA concentration, could improve the purity of the DNA solution, as indicated by a better A260/A280 ratio (Fig. 1b), which in the current study ensured successful amplification. Since some TCMs consist predominantly of plant components, which can pose a challenge for the extraction of high-quality animal DNA, similar situations have been reported when using DNA isolation methods to extract high-quality DNA from complex mixtures41,42. These methods included an initial step using CTAB (typically used for plant DNA extraction), and PCR amplification was successful only with a DNA template extracted from TCMs using CTAB43. These results highlight the importance of selecting appropriate DNA extraction techniques that improve the reliability of species identification through mini-barcoding. DNA extraction from leech derived products is therefore a crucial step to ensure that only authenticated animals are used in the production of TCMs and PCMs, and to maintain their quality control and safety.
DNA barcoding has proven to be highly effective in numerous taxonomic studies, and a typical study recommends the utilization of a universal primer pair that targets relatively longer and standard DNA regions44. However, such primers can be challenging when attempting to amplify target DNA from TCMs and PCMs, which often consist of mixtures of multiple species with degraded DNA due to processing preparations37. This issue is of paramount importance as it involves the accurate identification of medical animal used in the preparation of TCMs and PCMs through DNA barcoding technology. Given this, the development of mini-barcoding technology has significantly expanded the scope of DNA barcoding. Mini-barcodes can improve the successful amplification and identification of shorter DNA fragments, even if they are partially degraded. Compared to full-length barcodes, mini-barcodes (> 200 bp) generally perform well in detecting and identifying species across a wide range of metazoan taxa in an in silico study38. In addition, according to Pharmacopoeias of China, Europe and America45,46,47, species identification in TCMs and PCMs that combine multiple barcodes instead of relying on a single barcode can improve identification accuracy. This result highlights the importance of using multiple barcodes, especially when dealing with complex mixtures such as TCMs, as they can compensate for the limitations of a single barcode. To accommodate approximately shorter DNA regions for extracting degraded DNA from TCMs and PCMs, four candidate mitochondrial primer pairs of mini-barcoding DNA for three medicinal leeches were designed in the current study. Primers designed for amplifying ND1, 12S rDNA, 16S rDNA and COX1 sequences showed progressively decreasing amplification success rates (Figs. 2 and 3). The mini-barcode ND1 was successfully amplified in almost all samples, with amplification success rates for the 12S rDNA (70.8%) and 16S rDNA (66.7%) primers exceeding those reported in the previous study (69.2% and 53.8%, respectively)48. The amplification efficiency of the COX1 primers was equivalent to that of primers for amplifying DNA from animal samples, but was slightly lower than the efficiency of primers for amplifying the sequence of W. pigra from the proprietary Chinese medicine (PCM), Naoxintong49. Despite the differences in detection efficiency of different mini-barcodes among raw materials, TCMs and PCMs, the successful validation of specific primers for ND1, 12S rDNA, 16S rDNA, and COX1 sequences highlights the potential of these markers together for the accurate identification of leech species from leech derived products.
The authentication and standardization of scientific names and pseudonyms of medicinal animals are crucial steps in ongoing efforts to correct the standardization of traditional Chinese Medicine (TCM) nomenclature in the Chinese Pharmacopoeia50. In the current study, the availability of mitochondrial genome sequences from three medicinal leeches provides essential reference sequences. These sequences enable the design of mini-barcode primers and increase the accuracy and credibility of species discrimination based on DNA barcodes. These efforts are essential to ensure the safety and efficacy of TCMs as well as the quality of raw materials and processed products, e.g., PCMs. To accurately identify leech species from TCMs and PCMs, the successful validation of specific primers for ND1, 12S rDNA, 16S rDNA and COX1 sequences from leech specimens has laid a solid foundation for subsequent analysis (Figs. 3 and 4). Shuizhi (H. nipponia) were claimed to be included in the ingredient lists of all PCMs used in the current study. However, surprisingly, the ASAP results based on mini-barcode data reveal that only one PCM, MXK, definitely contains Shuizhi (H. nipponia). In contrast, all other 11 PCMs contain only Mahuang (W. pigra) but not Shuizhi (H. nipponia). It may that not enough medicinal leeches were used in two PCMs, QZ and CTZL, however, it is important to note that this final conclusion must take into account various factors such as complex processing methods that can lead to serious DNA degradation and the possibility of contamination. This may affect the PCR results of that two PCMs, QZ and CTZL. However, it is crucial to close this gap by integrating DNA mini-barcoding technology with other analytical methods such as high throughput sequencing, HPTLC51,52, HPLC-MS53, HPLC-ELSD54 and high-resolution melting55,56 to provide a holistic quality approach to achieve control in PCMs. For example, to maximize the recovery of amplicons from processed products with complex ingredient mixtures, mini-barcodes can be combined with metabarcoding via high throughput sequencing to enable more accurate and detailed analysis of the species present in TCMs37. This integration will help ensure the accuracy and reliability of results when assessing the authenticity of medicinal ingredients.
Conclusion
The results of this study highlight the importance of using mini-barcoding technology to identify leech species from traditional Chinese medicines and proprietary Chinese medicines. The validation of ND1, 12S rDNA, 16S rDNA and COX1 primers on morphologically identified specimens demonstrated their effectiveness and specificity and allowed distinction between closely related medicinal leech species. The ASAP result based on mini-barcode showed discrepancies in the medicinal leech species used in traditional leech derived medical products (TCMs and PCMs), with most of these productions being based on Whitmania pigra (Mahuang), contrary to the claims of containg Hirudo nipponia (Shuizhi). This discovery is crucial for the authentication and standardization of scientific names and pseudonyms of medicinal leeches used in traditional medicine. Furthermore, DNA quality and appropriate extraction methods are crucial for the successful identification of medicinal animals using mini-barcoding.
Materials and methods
Sample collection and morphological identification of medicinal leeches
In the current study, various medicinal leech samples were collected, including seven specimens collected with hydrobiological nets from Xuzhou and two traditional Chinese medicine (TCM) leech samples purchased from traditional medicinal herb markets in Anhui Province. The two traditional Chinese medicinal leech samples included a sample of dried medicinal leeches and a sample of processed medicinal leeches. In addition, four specimens of Poecilobdella manillensis, commonly used as an adulterant for medicinal leeches, were also collected in China using hydrobiological nets (see Table 1). All leech specimens collected via hydrobiological nets were fixed in 95% ethanol and frozen at −20 °C and identified using Yang’s criteria57. The external morphology of all leech specimens was examined based on body size and shape, annulation, genital pore position and dorsal and ventral color patterns. The reproductive system of each specimen was examined under a dissecting microscope.
To further increase sample diversity, 14 types of proprietary Chinese medicines (PCMs), including powders, pills and tablets containing leech components, were purchased from several pharmacies (Fig. 1a). The aim of this selection was to ensure that diverse pharmaceutical production sites from a wide geographical range were represented across the country. This approach was chosen to avoid potential biases in the sampling process and to guarantee a comprehensive representation of available medicinal leech products.
DNA extraction from leech specimens and leech derived products
DNA was extracted from each leech sample using the One-tube Universal Sample DNA Extraction Kit (Sangon Biotech, Shanghai, China). The caudal sucker of the leech was first cleaned with 75% alcohol and approximately 1.5 mg of caudal sucker tissue was excised and placed in a 1.5 ml Eppendorf tube (EP tube). 100 μL of Qlysis-G reagent and 10 μL of Proteinase K were added to each EP tube and then shaken for 5 s to mix. The mixture was exposed to a 65 °C water bath for 60 min and then to a 95 °C water bath for 30 min to ensure complete tissue lysis. After the water bath, 100 μL of buffer NST was added and the tube was shaken well. The sample was then centrifuged at 12,000 rpm for 5 min at room temperature. The supernatant containing the DNA was collected and stored at −20 °C for subsequent PCR. For leech derived products, including traditional Chinese medicines and proprietary Chinese medicines, approximately 25 mg of the powder was added to a sterilized 1.5 mL centrifuge tube. DNA extraction for leech derived products was performed using the Ezup Column Animal Genomic DNA Purification Kits (Sangon Biotech, Shanghai, China), with the adjusted lysis time extended from 1 to 1 h and 30 min depending on the material consistency. The resultant supernatant was stored at −20 °C for further PCR. DNA quality was measured by calculating the absorbance ratio at 260/280 nm and DNA concentration using a Nanodrop 2000 Spectrophotometer.
Designing mini-barcoding primers for leech identification
The complete mitochondrial genomes of three medicinal species, namely Whitmania pigra (accession number: EU304459), Hirudo nipponia (accession number: KC667144, MZ50750) and Whitmania acranulata (accession number: KM655838, KC688271), were retrieved from the GenBank database in NCBI. The sequences of protein-coding genes (CDS) and ribosomal RNA genes (rDNA) from these genomes were aligned using Geneious Prime 2021.2.2. Regions that exhibited high alignment consistency across all three species were identified for primer development. Primer design was performed using Oligo 7.0 based on three principal criteria: targeting a fragment length of approximately 200 base pairs, which is considered optimal for the mini-barcoding primer length of approximately 20 ± 5 bp38,58, and ensuring high primer specificity. This high primer specificity was achieved by developing primer sequences that uniquely bind to the target region while exhibiting minimal cross-reactivity with non-target sequences.
DNA amplification and sequencing
The PCR reaction system with a total volume of 25 μL, consisted of 15 μL 2 × Taq Master Mix (Dye Plus), 6 μL diethy pyrocarbonate (DEPC) treated water, 2 μL DNA sample, and 1 μL of each primer (forward and reverse). The PCR conditions were categorized into two groups: The first group, including 12SF1/R1, 16SF1/R1, and ND1F1/R1, was first subjected to pre-denaturation at 95 °C for 5 min, followed by 35 cycles of denaturation at 95 °C for 45 s, annealing at 48 °C for 1 min and a final extension at 72 °C for 10 min. The samples were then stored at 4 °C. The second group, consisting of COX1F1/R1, followed a similar protocol but with an annealing temperature of 45.5 °C. The amplified fragments were examined by electrophoresis in a 1.5% agarose gel stained with 3 μL Vic Gel Red (Vicmed, Xuzhou China) at 120 V/200 mA for 25–30 min and visualized using the Gel imaging system (BIO-RAD, American). The PCR products, which showed a bright and single stripe, were sent to the company (GENEWIZ, Suzhou, China) for Sanger sequencing. To assess the specificity of the primers, amplifications were also performed on a common adulterant of the medicinal leech, Poecilobdella manillensis.
Data processing and species delimitation
The identity of the target sequences amplified by each primer pair was confirmed through the Basic Local Alignment Search Tool (BLAST) against the NCBI database, ensuring a high degree of sequence similarity. The newly obtained DNA sequences were manually corrected, which involving the removal of primer sequences and low-quality sequencing regions. The reference sequence including one Whitmania pigra (accession number: EU304459), two Hirudo nipponia (NC_023776 and MZ507570) and two Whitmania acranulata (KM655838 and KC688271) were downloaded and included in the current analysis. All ND1 and COX1 sequences were aligned separately using MAFFT version 7.490, while the sequences of 12S and 16S rDNAs were aligned via LocARNA59. These aligned sequences were then concatenated into a single FASTA file that was used for subsequent species delimitation. Molecular species identification was then performed using the distance-based Assemble Species by Automatic Partitioning (ASAP) method, with the best result selected based on the partition with the lowest ASAP score60. The Jukes-Cantor model (JC69) was used for the ASAP analysis. The prior intraspecific divergence (P) was set between 0.001 and 0.1, the minimum relative gap width (X) was set to 0.5, and all other parameters were kept as default settings. In addition, ASAP species delimitation was performed separately for all ND1, 12S rDNA, 16S rDNA and COXI sequence to reduce distance calculation bias associated with missing data.
Data availability
The data included in the current study are assigned detailed accession numbers in GenBank (Table 1).
References
Phillips, A. J., Govedich, F. R. & Moser, W. E. Leeches in the extreme: Morphological, physiological, and behavioral adaptations to inhospitable habitats. Int. J. Parasitol. Parasites Wildl. 12, 318–325. https://doi.org/10.1016/j.ijppaw.2020.09.003 (2020).
Sket, B. & Trontelj, P. Global diversity of leeches (Hirudinea) in freshwater. Hydrobiologia 595, 129–137. https://doi.org/10.1007/s10750-007-9010-8 (2007).
Kampowski, T., Eberhard, L., Gallenmuller, F., Speck, T. & Poppinga, S. Functional morphology of suction discs and attachment performance of the Mediterranean medicinal leech (Hirudo verbana Carena). J. R. Soc. Interface 13, 20160096. https://doi.org/10.1098/rsif.2016.0096 (2016).
Lui, C. & Barkley, T. W. J. Medicinal leech therapy: New life for an ancient treatment. Nursing 45, 25–30. https://doi.org/10.1097/01.NURSE.0000472561.41030.2d (2015).
Kumar, S., Dobos, G. J. & Rampp, T. Clinical significance of leech therapy in Indian medicine. J. Evid. Based Complement. Alternat. Med. 18, 152–158. https://doi.org/10.1177/2156587212466675 (2013).
Dong, H. et al. Chinese Medicinal Leech: Ethnopharmacology, Phytochemistry, and Pharmacological Activities. Evid. Based Complement Alternat. Med. 2016, 7895935. https://doi.org/10.1155/2016/7895935 (2016).
Qiu, J., Lingna, W., Jinghong, H. & Yongqing, Z. Oral administration of leeches (Shuizhi): A review of the mechanisms of action on antiplatelet aggregation. J. Ethnopharmacol. 232, 103–109. https://doi.org/10.1016/j.jep.2018.12.010 (2019).
Lemke, S. & Vilcinskas, A. European medicinal leeches-new roles in modern medicine. Biomedicines 8, 99. https://doi.org/10.3390/biomedicines8050099 (2020).
Koeppen, D., Aurich, M., Pasalar, M. & Rampp, T. Medicinal leech therapy in venous congestion and various ulcer forms: Perspectives of Western, Persian and Indian medicine. J. Tradit. Complement Med. 10, 104–109. https://doi.org/10.1016/j.jtcme.2019.08.003 (2020).
Tang, X. J., Feng, Y. J., Cui, Y. M. & Cui, J. B. Research progress on clinical application of hirudo in internal medicine. J. Pract. Tradit. Chin. Internal Med. 34, 86–89. https://doi.org/10.13729/j.issn.1671-7813.z20200300 (2020).
Kvist, S., Manzano-Marin, A., de Carle, D., Trontelj, P. & Siddall, M. E. Draft genome of the European medicinal leech Hirudo medicinalis (Annelida, Clitellata, Hirudiniformes) with emphasis on anticoagulants. Sci. Rep. 10, 9885. https://doi.org/10.1038/s41598-020-66749-5 (2020).
Marin, E. et al. Structural insights into thrombolytic activity of destabilase from medicinal leech. Sci. Rep. 13, 6641. https://doi.org/10.1038/s41598-023-32459-x (2023).
Liu, S.-J., Cao, Y.-L. & Zhang, C. Hirudin in the treatment of chronic kidney disease. Molecules 29, 1029. https://doi.org/10.3390/molecules29051029 (2024).
Tian, F. et al. Research progress on the treatment of diabetic nephropathy with leech and its active ingredients. Front. Endocrinol. 15, 1296843. https://doi.org/10.3389/fendo.2024.1296843 (2024).
Chen, J. et al. Pharmacological activities and mechanisms of Hirudin and its derivatives—A review. Front. Pharmacol. 12, 660757. https://doi.org/10.3389/fphar.2021.660757 (2021).
Sig, A. K., Guney, M., Uskudar Guclu, A. & Ozmen, E. Medicinal leech therapy-an overall perspective. Integrat. Medi. Res. 6, 337–343. https://doi.org/10.1016/j.imr.2017.08.001 (2017).
Xiao, L. & Chen, K. Review and analysis of processing technology and active ingredients of medicinal leech. China Pharm. 17, 1760–1762 (2014).
Cheng, S. Studies on the polypeptide compositions and anticoagulant mechanism of leeches Master thesis, Hubei University of Chinese Medicine, (2018).
Dunwiddie, C. et al. Antistasin, a leech-derived inhibitor of factor Xa. Kinetic analysis of enzyme inhibition and identification of the reactive site. J. Biol. Chem. 264, 16694–16699 (1989).
Kwak, H.-J., Park, J.-S., Medina Jiménez, B. I., Park, S. C. & Cho, S.-J. Spatiotemporal Expression of Anticoagulation Factor Antistasin in Freshwater Leeches. Int. J. Mol. Sci. 20, 3994. https://doi.org/10.3390/ijms20163994 (2019).
Jung, H. I., Kim, S. I., Ha, K. S., Joe, C. O. & Kang, K. W. Isolation and characterization of guamerin, a new human leukocyte elastase inhibitor from Hirudo nipponia. J. Biol. Chem. 270, 13879–13884. https://doi.org/10.1074/jbc.270.23.13879 (1995).
Al-Sayed, S. E., Abdel-Latif, M., Abdel-Haleem, H. M., El-Shahawy, G. & Abdel-Tawab, H. Protective efficacy of Eglin C from Hirudo medicinalis against Eimeria papillata-induced coccidiosis. Vet. Parasitol. 314, 109869. https://doi.org/10.1016/j.vetpar.2022.109869 (2023).
Baskova, I. P. & Zavalova, L. L. Proteinase inhibitors from the medicinal leech Hirudo medicinalis. Biochemistry 66, 703–714. https://doi.org/10.1023/a:1010223325313 (2001).
Gronwald, W. et al. Structure of the leech protein saratin and characterization of its binding to collagen. J. Mol. Biol. 381, 913–927. https://doi.org/10.1016/j.jmb.2008.06.034 (2008).
Cheng, X. M., Zhang, M., Wang, J. H., Lv, L. & Ning, S. X. Research process of hirudin. J. Jilin Med. Univ. 42, 135–137. https://doi.org/10.13845/j.cnki.issn1673-2995.2021.02.024 (2021).
Hebert, P. D. N., Cywinska, A., Ball, S. L. & de Waard, J. R. Biological identifications through DNA barcodes. Proc. R. Soc. Lond. B Biol. Sci. 270, 313–321. https://doi.org/10.1098/rspb.2002.2218 (2003).
Antil, S. et al. DNA barcoding, an effective tool for species identification: a review. Mol. Biol. Rep. 50, 761–775. https://doi.org/10.1007/s11033-022-08015-7 (2023).
Chac, L. D. & Thinh, B. B. Species identification through DNA barcoding and its applications: a review. Biol. Bull. 50, 1143–1156. https://doi.org/10.1134/s106235902360229x (2023).
Trontelj, P. & Utevsky, S. Y. Celebrity with a neglected taxonomy: molecular systematics of the medicinal leech (genus Hirudo). Mol. Phylogenet. Evol. 34, 616–624. https://doi.org/10.1016/j.ympev.2004.10.012 (2005).
Siddall, M. E., Trontelj, P., Utevsky, S. Y., Nkamany, M. & Macdonald, K. S. Diverse molecular data demonstrate that commercially available medicinal leeches are not Hirudo medicinalis. Proc. R. Soc. B: Biol. Sci. 274, 1481–1487. https://doi.org/10.1098/rspb.2007.0248 (2007).
Trontelj, P. & Utevsky, S. Y. Phylogeny and phylogeography of medicinal leeches (genus Hirudo): fast dispersal and shallow genetic structure. Mol. Phylogenet. Evol. 63, 475–485. https://doi.org/10.1016/j.ympev.2012.01.022 (2012).
Tian, N. et al. Identification of Pheretima by multiplex allele-specific PCR. Chin. J. Exp. Tradit. Med. Formulae 25, 124–129. https://doi.org/10.13422/j.cnki.syfjx.20191716 (2019).
Qian, R., Tian, N., Zhang, X. Y., Jiang, C. & Yuan, Y. Identification of Cervi Cornu Pantotrichum and Cervi Cornu by allele-specific PCR. Chin. J. Exp. Tradit. Med. Formulae 25, 118–123. https://doi.org/10.13422/j.cnki.syfjx.20191712 (2019).
Zhang, X. Y., Yuan, Y., Hu, Q. T., Jiang, C. & Fang, C. W. Identification of Testudinis Carapax et Plastrum by specific PCR method and high - resolution melting analysis. Mod. Chin. Med. 21, 1215–1220. https://doi.org/10.13313/j.issn.1673-4890.20190410002 (2019).
Parveen, I., Gafner, S., Techen, N., Murch, S. J. & Khan, I. A. DNA barcoding for the identification of botanicals in herbal medicine and dietary supplements: Strengths and limitations. Planta Med. 82, 1225–1235. https://doi.org/10.1055/s-0042-111208 (2016).
Xin, T. Y. et al. Application of DNA barcoding technology to national drug sampling inspection. Acta Pharm. Sin. 56, 1497–1508. https://doi.org/10.16438/j.0513-4870.2021-0089 (2021).
Chen, S. L. et al. DNA barcoding in herbal medicine: Retrospective and prospective. J. Pharm. Anal. 13, 431–441. https://doi.org/10.1016/j.jpha.2023.03.008 (2023).
Yeo, D., Srivathsan, A. & Meier, R. Longer is not always better: optimizing barcode length for large-scale species discovery and identification. Syst. Biol. 69, 999–1015. https://doi.org/10.1093/sysbio/syaa014 (2020).
Lu, Z. et al. Visualization of DNA in highly processed botanical materials. Food Chem. 245, 1042–1051. https://doi.org/10.1016/j.foodchem.2017.11.067 (2018).
Koetsier, G. & Cantor, E. J. A Practical Guide to Analyzing Nucleic Acid Concentration and Purity with Microvolume Spectrophotometers (New England Biolabs, Inc. 2019). Retrieved from https://www.neb.com/en/-/media/nebus/files/application-notes/technote_mvs_analysis_of_nucleic_acid_concentration_and_purity.pdf?rev=c24cea043416420d84fb6bf7b554dbbb.
Arulandhu, A. J. et al. Development and validation of a multi-locus DNA metabarcoding method to identify endangered species in complex samples. Gigascience 6, 1–18. https://doi.org/10.1093/gigascience/gix080 (2017).
Zhu, X., Wu, H. Y., Shaw, P. C., Peng, W. & Su, W. Specific DNA identification of Pheretima in the Naoxintong capsule. Chin. Med. 14, 41. https://doi.org/10.1186/s13020-019-0264-7 (2019).
Cheng, X. W. et al. DNA extraction protocol for biological ingredient analysis of Liuwei Dihuang Wan. Genom. Proteom. Bioinform. 12, 137–143 (2014).
Liu, Y. K. & Erséus, C. New specific primers for amplification of the Internal Transcribed Spacer region in Clitellata (Annelida). Ecol. Evol. 7, 10421–10439. https://doi.org/10.1002/ece3.3212 (2017).
Chen, S. et al. A renaissance in herbal medicine identification: From morphology to DNA. Biotechnol. Adv. 32, 1237–1244. https://doi.org/10.1016/j.biotechadv.2014.07.004 (2014).
Ha, W. Y., Wong, K. L., Ma, W. Y., Lau, Y. Y. & Chan, W. H. Enhancing testing laboratory engagement in plant DNA barcoding through a routine workflow-a case study on Chinese Materia Medica (CMM). Plants 11, 1317. https://doi.org/10.3390/plants11101317 (2022).
Sally Rahayu, Y. Y. Assessing adoption of DNA barcoding in herbal value chain: A multistakeholder analysis. Nat. Prod. J. 13, 54–62. https://doi.org/10.2174/2210315513666220928112238 (2023).
Xiao, L. Research on the DNA barcoding, activity peptide isolation and its anticoagulation mechanism of leeches. Doctor thesis, Hubei University of Chinese Medicine (2015).
Zhu, X. X., Wu, H., Shaw, P., Peng, W. & Su, W. Identification of specific DNA markers for Hirudo in the Naoxintong capsule. Act Sci. Nat. Univ. Sunyatseni 59, 114–124. https://doi.org/10.3471/j.cnki.acta.snus.2020.01.014 (2020).
Jiang, C., Huang, L. Q., Yuan, Y. & Li, J. D. Standard citations of zoological scientific names and Chinese names in Chinese Pharmacopoeia. Sci. Sin. Vitae 48, 772–782. https://doi.org/10.1360/N052017-00254 (2018).
Mück, F. et al. Complementary authentication of Chinese herbal products to treat endometriosis using DNA metabarcoding and HPTLC shows a high level of variability. Front. Pharmacol. 14, 1305410. https://doi.org/10.3389/fphar.2023.1305410 (2023).
Raclariu, A. C. et al. What’s in the box? Authentication of Echinacea herbal products using DNA metabarcoding and HPTLC. Phytomedicine 44, 32–38. https://doi.org/10.1016/j.phymed.2018.03.058 (2018).
Sánchez, M., GonzálezBurgos, E., Divakar, P. K. & Gómez-Serranillos, M. P. DNA-based authentication and metabolomics analysis of medicinal plants samples by DNA barcoding and ultra-high-performance liquid chromatography/triple quadrupole mass spectrometry (UHPLC-MS). Plants 9, 1601. https://doi.org/10.3390/plants9111601 (2020).
Wang, W. P. et al. Study on the molding process and quality standard of Qizhi Yifei granules. Global Tradit. Chin. Med. 11, 1525–1533 (2018).
Buddhachat, K. et al. Using mini-barcodes coupled with high resolution melting (minibar-HRM) method for species discrimination across Pangasianodon gigas, Pangasianodon hypophthalmus and Pangasius larnaudii.. Aquaculture 530, 735773. https://doi.org/10.1016/j.aquaculture.2020.735773 (2021).
Fadzil, N.A.-O., Wagiran, A.A.-O., Mohd Salleh, F.A.-O.X., Abdullah, S. & Mohd Izham, N. H. Authenticity testing and detection of Eurycoma longifolia in commercial herbal products using bar-high resolution melting analysis. Genes 9, 408. https://doi.org/10.3390/genes9080408 (2018).
Yang, T. Fauna Sinica.Annelida.Hirudinea (Science Press, 2016).
Meusnier, I. et al. A universal DNA mini-barcode for biodiversity analysis. BMC Genom. 9, 214. https://doi.org/10.1186/1471-2164-9-214 (2008).
Will, S., Joshi, T., Hofacker, I. L., Stadler, P. F. & Backofen, R. LocARNA-P: Accurate boundary prediction and improved detection of structural RNAs. RNA 18, 900–914. https://doi.org/10.1261/rna.029041.111 (2012).
Puillandre, N., Brouillet, S. & Achaz, G. ASAP: Assemble species by automatic partitioning. Mol. Ecol. Resour. 21, 609–620. https://doi.org/10.1111/1755-0998.13281 (2020).
Acknowledgements
This article was sponsored by the National Natural Science Foundation of China [32100334, 31801957], the Natural Science Foundation of Jiangsu, China [BK20210897, BK20181471] and the Scientific Research Foundation for the Talents by Xuzhou Medical University [D2020009]. The authors thank Siye Guo, Haodong Guan, Chi Gu and Jing-ai Shi for assistance with DNA extractions, Yan Zhang, Xingyu Xu, and Ying Zhu for purchasing medicines.
Author information
Authors and Affiliations
Contributions
Yingkui Liu initiated the study and designed the research framework, and the initial draft of the manuscript was prepared by Yingkui Liu and Youqun Xie; Youqun Xie and Yu Wang were responsible for data collection; Youqun Xie analyzed and visualized data; Yong Liu and YanYan Zong conducted a thorough literature review. Significant improvements and insightful feedback were provided by Yong Liu, Chong Li, and Jiajia Dong. This research was a collaborative effort among all authors. Each author has approved the final manuscript and agreed to its publication.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, Y., Xie, Y., Wang, Y. et al. The development and application of mini-barcodes from mitochondrial DNA for identifying medicinal leeches from traditional medicines. Sci Rep 15, 20444 (2025). https://doi.org/10.1038/s41598-025-04610-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-04610-3