Abstract
Plant diseases pose a significant threat to global food security, with severe implications for agricultural productivity. Early and accurate detection of these diseases is crucial, yet it remains a challenging task, significantly impacting crop yields and food supply chains. Despite the progress in artificial intelligence, particularly deep learning, challenges persist in real-world applications due to environmental noise, varying light conditions, and other complicating factors that hinder detection accuracy. This study introduces the AttCM-Alex model, a novel deep-learning framework designed to boost the detection and classification of plant diseases under challenging environmental conditions. By integrating convolutional operations with self-attention mechanisms, AttCM-Alex effectively addresses the variability in light intensity and image noise, ensuring robust performance. To simulate practical agricultural scenarios, the study employs bilinear interpolation for image dimension adjustment and introduces Salt-and-Pepper noise. Additionally, the model’s robustness was evaluated by varying image brightness levels by ±10%, ±20%, and ±30%. Experimental results demonstrate that AttCM-Alex significantly outperforms traditional models, particularly in scenarios involving fluctuating light conditions and noise interference. The model achieved a peak detection accuracy of 0.97 with a 30% increase in image brightness and maintained an accuracy of 0.93 even with a 30% decrease in brightness, highlighting its robustness and reliability. The findings affirm the AttCM-Alex model as a powerful tool for real-world agricultural applications, capable of enhancing disease detection systems’ accuracy and efficiency. This advancement not only supports better crop management practices but also contributes to sustainable agriculture and global food security.
Similar content being viewed by others
Introduction
Plant diseases are a major threat to crop yields and food safety, significantly impacting agricultural economies and contributing to global food shortages1. Historical events, such as the Irish Potato Famine and the Bengal Famine, underscore the devastating effects of plant diseases2. To prevent such historical tragedies from recurring, maintaining crop yields will be of paramount importance. To minimize losses caused by plant diseases, it is necessary to continuously monitor crop diseases and take effective and timely measures, which will significantly reduce potential damages3. Traditional manual detection methods are costly and inefficient for large-scale disease monitoring, often failing to capture disease progression in a timely manner4. Consequently, the accurate detection of plant diseases and the timely implementation of appropriate countermeasures are of utmost importance.
While existing methods in the field focus primarily on using image-based classification algorithms, they often overlook the unique challenges presented by real-world agricultural environments. These environments typically involve complex backgrounds, such as soil and weeds, making accurate disease detection particularly difficult5. Furthermore, the scarcity of high-quality annotated datasets for rare diseases and varying crop conditions continues to limit the effectiveness of current AI models. In contrast to common image classification problems, plant diseases exhibit highly similar characteristics, with both biological and abiotic factors contributing to the characteristics of similar diseases6. As plants become afflicted by diseases, pathological features typically manifest in leaves, stems, flowers, or fruits7. Among these, leaf features are the most pronounced and readily detectable, making them the primary source of information for assessing plant disease severity8. However, while the progress made in using deep learning-based image algorithms for plant disease detection is notable, many AI models struggle to adapt to the dynamic conditions of agricultural production environments. Specifically, lighting variations, image noise, and model computational demands are significant barriers to achieving reliable and scalable disease detection9.
Although considerable progress has been made in using deep learning-based image algorithms for detecting plant diseases under the support of AI technologies, many issues remain unresolved when considering the application of academic achievements in real-world agricultural production10. For example, many scholarly studies perform detection tasks in single backgrounds, making it difficult to achieve ideal detection results in actual production environments that include complex backgrounds such as soil and weeds. Furthermore, in real agricultural production, factors such as lighting conditions during image capture and noise introduced by imaging equipment can affect the accuracy of plant disease detection. Given these challenges, AI-driven approaches, particularly computer vision models, offer a promising solution for scalable, efficient, and timely plant disease detection.
This research tackles these challenges by introducing a hybrid deep learning model, Attention Convolutional Mixed AlexNet (AttCM-Alex), designed to enhance detection accuracy in complex, noisy, and resource-limited environments. By combining the strengths of convolutional neural networks (CNNs) and Vision Transformers (ViT), the model leverages the self-attention mechanism to capture global information while maintaining computational efficiency. In recent years, the rapid development of artificial intelligence (AI) has effectively addressed significant issues in various fields11. The scalability of AI models in resource-constrained agricultural environments is a significant challenge, as sophisticated models require powerful hardware and computational resources, which may not be available in small-scale or low-resource settings. Designing lightweight, efficient models is crucial for these environments. Data availability is another significant challenge, as high-quality, annotated datasets are scarce, especially for plant disease detection. Acquiring labeled data for rare diseases or diverse crops and environmental conditions limits the generalizability of AI models. Furthermore, collecting data under varying real-world conditions adds complexity. Ethical considerations, such as data privacy, model transparency, and fair decision-making processes, are also crucial for AI applications in agriculture.
Computer vision has provided powerful methods for solving problems in other domains, promoting advances in multiple areas, and achieving significant progress in object recognition and classification. Convolutional neural networks (CNNs) have demonstrated robust capabilities in image classification. Numerous excellent models based on CNN have been designed, including EfficientNet12, AlexNet13, ResNet14, DenseNet15, and MobileNet16. Despite these successes, these methods have certain limitations, such as the constrained receptive field of CNNs due to the fixed size of convolutional kernels, which leads to significant loss of global information in images.
However, despite these advancements, existing AI models, especially CNNs, face significant limitations when applied to real-world agricultural environments. These limitations include high computational demands, threatening macroeconomic stability, lack of adaptability to varying environmental conditions, sustainable development, and reduced performance in dynamic field settings. To overcome the drawback of CNNs in losing global information, researchers have drawn inspiration from the Transformer algorithm applied to computer vision17. Vision Transformer (ViT) has achieved remarkable results in various typical computer vision tasks, such as image recognition, object detection, and image segmentation18. Benefiting from the self-attention mechanism’s ability to capture global information, the output data of the model processing include comprehensive global details, allowing the computer’s perceptual field to encompass the entire image from the start of training. Furthermore, the dynamically calculated weight matrix of the self-attention mechanism enables it to handle irregular inputs, allowing it to distinguish it from CNNs easily.
This research addresses the practical challenges in plant disease detection by developing a deep learning-based model tailored for complex agricultural environments. We introduce the Attention Convolutional Mixed AlexNet (AttCM-Alex), designed to enhance robustness under varying light intensities. The proposed model’s key mechanisms, such as the self-attention mechanism, capture global dependencies within the data, enabling the model to weigh different parts of the input image more effectively. Additionally, the channel attention mechanism enhances the model’s ability to focus on crucial features by assigning varying importance to each channel. These mechanisms are central to improving the robustness and accuracy of our plant disease detection approach. This hybrid approach improves accuracy and computational efficiency, especially in resource-constrained environments. It is designed to operate effectively in real-world agricultural settings, where environmental variability and device limitations pose significant challenges. Furthermore, this study pioneers a methodology for simulating real-world agricultural conditions, such as brightness variation and image noise, which are critical to testing model robustness. Our methodology systematically adjusts the brightness of images in a plant disease dataset by ±10%, ±20%, and ±30% to rigorously evaluate the model’s performance across diverse lighting conditions. This approach significantly improves the detection accuracy despite environmental variability, thus contributing to a more reliable disease detection in agricultural practices. In addition, this work provides a theoretical foundation for future research on plant disease detection models in complex environments. The proposed plant disease detection model for complex environments overcomes issues such as lighting intensity and image noise. It also implements the functionality on mobile devices through the application, providing a theoretical foundation for future research on plant disease detection models in these environments.
Deficiencies in conventional approaches
Farmers currently rely on distinct detection methods such as chemical detection, spectral technology, and image processing technology to identify pests and diseases in crops. However, these conventional methods are labor intensive, expensive, complex, and have long detection times, making them inefficient for continuous monitoring. In addition, traditional machine learning methods employed to identify plant diseases face challenges when deploying in real-world applications due to high computing resource requirements, lack of resilience in varied environments, and unsuitability for devices with limited resources. Deep learning models like the Convolutional Swin Transformer also face challenges despite their extensive parameter sets and significant storage demands, which make them unsuitable for mobile or edge devices. They also suffer performance declines under different environmental conditions, reducing their reliability in practical agricultural settings.
Furthermore, while these traditional models often achieve high precision in specific datasets, they lack the resilience and adaptability required for deployment in real agricultural environments. Models usually experience performance degradation under varied environmental conditions, such as changing lighting, background noise, or weather effects, significantly affecting their reliability. In addition, these models are prone to performance decline after optimization and cannot adapt effectively to the dynamic nature of agricultural fields. There is a pressing need for robust, adaptive, and computationally efficient models capable of maintaining high performance in unpredictable and resource-constrained environments. Such models should be able to operate effectively on mobile and edge devices, addressing real-world challenges like lighting variability, environmental noise, and low-resource conditions while ensuring scalability and practical application in agricultural contexts.
Research motivation and contribution
This study leverages deep learning and computer vision technologies to achieve high-precision plant disease detection, effectively improving detection efficiency and reducing the high costs associated with traditional detection methods. To facilitate the practical application of the research outcomes in agricultural production, this study primarily focuses on plant disease detection in complex environments, considering the impact of lighting conditions and noise on detection accuracy. Corresponding solutions are proposed to address these challenges; this research develops and validates the AttCM-Alex, integrating advanced components to enhance feature extraction and classification accuracy.
-
The study proposed a hybrid AttCM-Alex model that ViT with CNNs, enhancing its ability to capture both local and global features, addressing the limitations of traditional CNN-based models in agricultural environments.
-
The AttCM-Alex model adopts AlexNet as the backbone, utilizing the AttCM module to combine convolution operations and self-attention mechanisms.
-
The proposed AttCM-Alex architecture evaluates their performance under noisy conditions to assess their robustness against noise interference. It adds different levels of salt-and-pepper noise to the images, simulating real-world scenarios.
-
The model’s performance is validated under different brightness levels through extensive testing, demonstrating superior accuracy compared to baseline models, with an accuracy of 0.95 on the cucumber dataset and 0.97 on the banana dataset.
-
The proposed model maintains high performance even with increased image brightness up to 30%, making it suitable for deployment in real-world agricultural applications where lighting conditions can change drastically.
Study organization
The research study is organized as follows: “Literature Review” reviews the challenges for the detection of plant disease, highlighting current limitations and the need for robust models. “Proposed methodology” describes the methodology, describing the AttCM-Alex model, data management, and methods. “Proposed hybrid deep learning architectures” presents results, comparing the AttCM-Alex’s performance across varying brightness levels. “Experiments and result analysis” concludes with a summary of research results, potential future enhancements, and implications for real-world agricultural applications, sustainable practices, and food security.
Literature review
With rapid development, many researchers have introduced AI techniques into the field of plant disease detection. Among these, computer vision models such as AlexNet, ResNet, Faster R-CNN19, and VGG20 are typical examples. CNNs have demonstrated excellent performance in various computer vision tasks, including image classification, object detection, and instance segmentation.
Many studies use the Plant Village dataset as experimental material to detect problems in stable environments, achieving high detection accuracy. Bajpai et al.21 introduces a new perception-based framework for detecting potato leaf diseases, aiming to reduce agricultural losses and increase crop productivity. The model uses MDSCIRNet and SEResNet101V2 strengths for feature extraction and classification, with advanced data augmentation techniques improving training and model generalization. The model achieves training and test accuracy of 99.89% and 99.67%, offering a promising solution for early disease diagnosis. Huang et al.15 proposed a neural architecture search model for detecting diseases in different plant leaves, which learns features from the provided dataset and applies these features to detect plant diseases, achieving a recognition accuracy of 99.01% on the Plant Village dataset. Khandelwal et al.22 designed a plant disease detection model combining transfer learning and deep residual learning for multi-plant and multi-disease detection, achieving a detection accuracy of 99.374% on a test set that included 86,198 images of 57 classes (healthy and specific diseases) of 25 crops from the Plant Village dataset. Singh et al.23, addressing the issue of anthracnose-infected mango leaf recognition and diagnosis, proposed a multilayer CNN for the early detection of fungal diseases, achieving an accuracy of 97.13% on a real-time dataset captured at Shri Mata Vaishno Devi University in India. Zeng et al.24 designed a new detection model by introducing the self-attention mechanism into CNN, achieving a detection accuracy of 0.98 in the MalayaKew Plant Leaf subset, which contains 988 images of 44 disease categories.
In the field of plant disease detection, several studies have used CNN to address the challenges encountered when detecting plant diseases in complex environments, such as varying light intensity and leaf occlusion. Zhu et al.25 presents CBF-YOLO network for detecting common soybean pests in complex environments. The network includes CSE-ELAN, Bi-PAN, and FFE modules, which enhance feature extraction and fuse features for more accurate pest detection. The FFE module refines multi-scale fused features, improving their expression ability. Experimental results show an mAP of 86.9% for common soybean pests, with average precisions for detecting Caterpillar and Diabrotica speciosa pest-damaged leaves reaching 86.5% and 87.3%, respectively. Nawaz et al.26 develop CoffeeNet, a novel deep-learning model that addresses challenges in cultivation and quality due to environmental changes and plant diseases. Using a spatial-channel attention strategy-based ResNet-50 model, the model achieved a classification accuracy of 98.54% and an mAP of 0.97, demonstrating its usefulness in localizing and categorizing various types of coffee plant leaf disorders. Mingyue et al.27 research on plant leaf disease detection suggests deep learning techniques can enhance accuracy by overcoming traditional methods’ drawbacks like misjudgment and high labor costs. They emphasize the need to address light intensity, leaf occlusion, complex backgrounds, and high disease similarity. Juncheng et al.28 developed a CNN-based system to detect cucumber mildew in greenhouse environments. Despite uneven light distribution and noise, they used composite color features to achieve a precise segmentation and detection accuracy of 97.29% and 95.7%, respectively.
The Vision Transformer architecture, introduced in 2020, has shown promising performance in computer vision, but its application in plant disease detection is less prevalent than CNN. The model typically incorporates an attention mechanism, which is usually introduced into CNN. Thai et al.29 introduce MobileH-Transformer, a hybrid model that uses CNN and Transformer architectures for accurate detection of leaf disease. It uses a dual convolutional block for feature extraction and reduces input size for the transformer component. The model achieves competitive F1-score values of 97.20% on corn leaf disease and 96.80% on PlantVillage datasets, surpassing previous studies with 0.4 Giga Floating Point Operations. Sharma et al.30 proposed a novel model SoyaTrans network combining random shifting and CNN architecture, which is being developed to enhance plant leaf disease detection for crop safety. Tested on four datasets, it achieved high accuracy rates of 98%, 97%, 76%, and 92% with minimal computational complexity of 5.2 million parameters, making it a promising solution for detecting plant leaf diseases. Monisha et al.31 suggest that hybrid CNNs and Transformers (ViTs) can overcome challenges such as spatial hierarchy, computational complexity, object location, and data prerequisites in natural language processing, thus creating novel opportunities for CV tasks, plant disease detection, sustainability, and agricultural productivity. Fu et al.32 uses VIT methods for pattern recognition and deep learning to classify and predict crop pest images. Its self-attention mechanism helps identify unique solutions and improves accuracy. This method serves as a reference for research on agricultural diseases and pests and optimizes crop disease and pest control work for agrarian workers in need. Barman et al.33 present a smartphone-based solution using a Vision Transformer model to identify healthy and unhealthy tomato plants with diseases. The model detected 10 different tomato disease classes from 10,010 images. An Android app using a ViT-based model performed better than Inception V3 in 90.99% of testing, promising large-scale implementation in smart agriculture and inspiring future research. Borhani et al.34 combined CNN and Vision Transformer architectures to design a new network structure for plant disease detection, achieving a detection accuracy of 0.96 on the WRCD dataset, which contains three disease categories and 3,679 images taken in real environments. Zhang et al.35 designed a new Vision Transformer network structure called RCAA-Net, achieving a detection accuracy of 0.89 in the 2018 AI Challenge dataset, which contains 36,258 images of 61 disease categories. Qian et al.36 designed a new detection model by introducing the attention mechanism into CNN, achieving a detection accuracy of 0.99 on a custom dataset with 7701 images of 4 disease categories.
Combining the findings of earlier discussions, it is evident that CNN, as a typical model structure in the field of computer vision, has achieved excellent detection performance when plant disease detection is viewed as an image classification problem. However, these achievements are often obtained in stable environments and their performance tends to decline when applied to detect plant diseases in complex environments. Therefore, to improve the practical value of research, this study focuses on the detection of plant diseases in complex environments and further explores the impact of variations in noise and light intensity on model performance in the detection of plant diseases. In addition, Table 1 provides critical analysis, synthesis, and identification of research gaps by comparing plant disease detection with other traditional approaches. This comparative approach highlights the unique challenges associated with environmental variability in the detection of plant diseases. It underscores the need for robust, adaptable models capable of maintaining high performance under diverse real-world conditions.
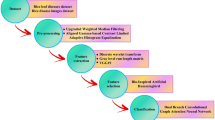
Proposed methodology
This research improves plant disease detection by addressing the limitations of traditional approaches by proposing a deep learning-based model as the backbone and efficiently combining attention mechanisms with convolution through the AttCM module. The core of the AttCM-Alex model lies in its integration of CNNs and ViTs, leveraging the benefits of both approaches for plant disease detection. The ViT architecture is particularly suited for tasks where capturing global information is crucial. Contrary to CNNs, which have a limited receptive field, ViTs use self-attention mechanisms to weigh different parts of the image based on their relevance, allowing the model to learn complex patterns and relationships in the data more effectively. The self-attention mechanism works by capturing relationships between distant parts of the input data, allowing the model to understand the global context better. The model focuses on spatially distant areas of an image that may still have important correlations. The channel attention mechanism, on the other hand, allows the model to adjust the weight of each channel in the feature map, thereby enhancing its ability to focus on the most informative channels for disease classification. These two mechanisms work synergistically to improve the overall performance of the AttCM-Alex model and show superior robustness in addressing image brightness’s impact on detection accuracy. Experimental results show that AttCM-Alex accurately identifies and classifies diseases in plants, achieving the highest accuracy in both experimental tests. The proposed model codes are publicly available at: https://github.com/tmsherazi-psu/Smart_Agriculture_AttCMAlex with DOI: https://doi.org/10.5281/zenodo.15762466.
Proposed datasets
The study evaluated the performance of a proposed model using three public datasets, which include leaves, weeds, and soil, making the images more complex than those from controlled backgrounds. Instances of the datasets are illustrated in Fig. 1.
This figure shows some images from cucumber, banana, and tomato datasets and image preprocessing like resizing and data augmentation.
This study uses a cucumber disease dataset39 to assess the model’s ability to detect plant leaf diseases. The dataset comprises 679 images captured in real-world environments, including leaves, soil, and weeds, as shown in Table 2. 80% images are classified into healthy and diseased cucumber leaves for training, while the remaining 20% images are used for testing. The dataset presents a classic binary classification problem and is referred to as the cucumber dataset throughout the paper.
The study utilized the banana leaf disease dataset to further evaluate the architecture’s disease detection capabilities40. The dataset comprises healthy, Sigatoka-infected leaves and bacterial wilt-affected leaves classifications, as depicted in Table 3. All images in this dataset have dimensions of \(150 \times 113 \times 3\). The dataset contains 1284 images, with 20% images from each category used for testing and the rest for training.
Finally, this study uses the Plant Village tomato dataset41, a well-known dataset for plant disease detection, to compare the performance of a proposed model. The dataset consists of 4021 images, each with dimensions of \(256 \times 256 \times 3\), across ten categories, with 50 images used for testing and the remaining images for training, as shown in Table 4. The tomato dataset is used for a multi-class classification problem focusing on tomatoes and is used in this study to compare its performance with other studies.
Image preprocessing
The study utilized bilinear interpolation to adjust the image dimensions in the dataset due to the original dimensions being unsuitable. The bilinear interpolation calculation is as follows: To determine the function f at the point (x, y), assume we know four points \(Q_{11}, Q_{12}, Q_{21}, Q_{22}\), where the coordinates of \(Q_{11}\) are \((x_1, y_1)\), \(Q_{12}\) are \((x_1, y_2)\), \(Q_{21}\) are \((x_2, y_1)\), and \(Q_{22}\) are \((x_2, y_2)\). First, linear interpolation is performed in the x-direction, as shown in Eqs. (1) and (2).
After the calculations, we obtain two temporary points R1(x, y1) and R2(x, y2) . Then, linear interpolation is performed in the y-direction to obtain the desired value f(x, y) , as shown in Eq. (3).
Additionally, the results of Eq. (2) can be directly substituted into Eq. (3) to derive Eq. (4).
This study used bilinear interpolation to convert all images from the original dimensions to \(224 \times 224 \times 3\). The transformed images have a height and width of 224, with the number of channels remaining the same at 3, representing red, green, and blue.
Noise augmentation
To further test the model’s robustness, we considered the impact of substantial noise introduced by imaging equipment in real-world applications, which can affect detection accuracy. In this study, we introduced varying levels of salt-and-pepper noise to the images to simulate practical conditions. The trained model was then tested on this noise-augmented data to evaluate its detection performance.
This figure shows the image from the banana dataset in its original form after adding 10, 20, and 30% noise.
As depicted in Fig. 2, we artificially added three different proportions of salt-and-pepper noise to the original images: 10%, 20%, and 30%.
Simulating light intensity variations
In practical agricultural applications, light intensity is a major interfering factor, affecting detection accuracy. To simulate the varying light intensities encountered in real-world scenarios, we manually adjusted the image brightness in this study. The model was tested under these six different conditions to evaluate its detection accuracy under varying brightness levels, as illustrated in Fig. 3.
Depicts tomato dataset image after adjusting brightness to \(\pm 20, \pm 40, \pm 60 \%\).
Plant disease recognition and the impact of image brightness under complex backgrounds
The progressive emergence of exceptional performance in computer vision can be attributed to Visual Transformers, largely due to the effectiveness of the attention mechanism within this architecture42. Currently, the field of computer vision leverages Visual Transformer architectures in two primary directions: the development of pure Transformer models devoid of convolutional layers and the incorporation of attention mechanisms into CNNs42. The latter approach, predominantly utilized in this study, has demonstrated promising outcomes. In the specific context of plant disease detection within complex backgrounds, Albahli et al.43 introduced EANet by integrating attention mechanisms into CNNs, achieving an impressive detection accuracy of 99.89% on a custom dataset. Similarly, Alirezazadeh et al.44 employed a comparable strategy by embedding attention mechanisms into the CNN architecture, incorporating channel attention into EfficientNet, and attaining a detection accuracy of 86.89%. In agricultural production scenarios, variations in image brightness due to factors such as time of image acquisition, weather conditions, and light intensity can significantly impact detection accuracy. Addressing this challenge, this study proposes the AttCM-Alex for detecting plant diseases under varying image brightness conditions.
Proposed hybrid deep learning architectures
This study explores various deep learning models, including AlexNet, MobileNet, ResNet, SqueezeNet, and ConvNeXt, which are tailored for tasks like plant disease detection and classification. These models were chosen for their ability to handle challenges such as computational efficiency and depth-related accuracy issues. Transformer-based models, including the Vision Transformer, Swin Transformer, and Deep ViT, use attention mechanisms to enhance image feature extraction. The study introduces hybrid models such as the convolutional Swin Transformer and the proposed AttCM-Alex architecture, which combine convolutional and attention mechanisms for robust feature learning while maintaining computational efficiency. These models demonstrate the evolution of deep learning architectures, combining the strengths of convolutional neural networks with the flexibility of transformers.
AlexNet
This paper proposes a plant disease detection model based on an improved AlexNet convolutional neural network. AlexNet45, proposed by Alex Krizhevsky in 2012, is a classic convolutional neural network initially used for ImageNet classification. AlexNet is composed of stacked basic CNN modules, using stacked convolutional layers to extract image features. The AlexNet model adopted in this paper consists of eight layers: the first five are convolutional layers, and the last three are fully connected layers. Different features are added to each layer to train the network and evaluate its performance. The Local Response Normalization (LRN) and max-pooling layers are located after the first two convolutional layers, providing necessary support for subsequent processing. The model begins with an initial layer featuring 96 convolutional kernels, followed by subsequent layers incorporating 256 kernels, two additional sets of convolutional layers with 384 kernels each, and a final layer with 256 kernels. The resulting feature maps from these convolutional layers are then processed through two fully connected networks, each comprising 2048 neurons. The output layer produces tensors of varying dimensions—specifically configured to \(1 \times 1 \times 2\), \(1 \times 1 \times 3\), and \(1 \times 1 \times 10\) depending on the number of categories present in the dataset. Due to the large number of parameters, the network model is prone to overfitting, which reduces recognition efficiency. The enormous size of the network leads to significant computational demands, making it difficult to use in practical applications. This is because the hardware available during deployment often cannot handle such extensive computational tasks. Due to the depth of the network structure, it is susceptible to gradient loss problems, which also increase the difficulty of model optimization.
MobileNet
In this study, MobileNetV3 was selected for the classification of plant diseases. The MobileNet network46 is mainly based on separable depthwise convolutions to reduce computational complexity and the number of model parameters. The separable depthwise convolution consists of the depthwise convolution and the pointwise convolution. Unlike traditional convolution layers, depth-wise convolution does not compute the convolution kernel across all channels. Instead, a single convolution kernel is applied only to one channel. On the other hand, pointwise convolution is essentially a \(1 \times 1\) convolution layer. The computation process of depthwise separable convolution is input is passed through the depthwise convolution, which performs convolution operations separately on each channel. The resulting feature maps are then passed through the pointwise convolution. This computation is similar to that of standard convolutions, but the design significantly reduces the model’s complexity and parameter count, resulting in better performance on devices with limited hardware capabilities.
ResNet
In this study, ResNet101V2, a smaller version of the original ResNet model47, was selected to classify plant diseases. This network achieves a good balance between model size and recognition accuracy. MobileNet’s success is largely due to its internal depthwise separable convolution structure. As the number of layers in a network increases, the model’s performance improves. However, with increasing depth, the accuracy tends to saturate and then rapidly decrease. Kaiming He’s proposed ResNet effectively solves this problem. The most notable difference between deep residual networks and traditional network architectures is that deep residual networks include several bypasses that allow the input to skip several convolutional layers and directly connect to the subsequent convolution layers. ResNet has since become one of the most popular convolutional neural network architectures. The final output layer was modified according to the number of plant disease categories in the dataset.
SqueezeNet
This study proposes the SqueezeNet model, proposed by Forrest N. Iandola48 for plant disease detection and classification, which achieves similar accuracy to AlexNet. SqueezeNet consists of the squeeze stage for dimensionality reduction and the expand stage for dimensionality expansion. In the Squeeze stage, the model mainly uses \(1 \times 1\) convolutions to reduce the dimensionality of the input data. The reduced feature information is then passed to the Expand stage, where a combination of \(1 \times 1\) and \(3 \times 3\) convolutions is used to increase the dimensionality of the features. In the design of SqueezeNet, the pooling operations are placed after each module, helping provide larger activated feature maps to the convolutional layers. This allows for the retention of richer feature information, effectively improving classification accuracy.
ConvNeXt
This study used the ConvNeXt model49 to detect plant diseases, which have surpassed convolutional neural network models in computer vision due to their local attention mechanism. These models, often adopting a pyramid structure, reduce model computation and focus more on relationships between features. ConvNeXt, a new convolutional neural network model, integrates the advantages of ResNet and Swin Transformer, resulting in ConvNeXt. This network structure has been tested and shown to exceed the accuracy and segmentation performance of the Swin Transformer in image classification and instance segmentation tasks. Furthermore, as the dataset and model size increase, ConvNeXt’s performance continues to improve.
Deep ViT
This study uses DeepViT to detect and classify plant diseases, leveraging its improved attention mechanism to improve the extraction of characteristics and the precision of classification in complex agricultural environments. This algorithm suggests that stacking multiple transformer layers does not necessarily enhance model performance. Instead, as the network depth increases, the performance of the vision transformers quickly saturates. To address this issue, Daquan Zhou50 proposed the Re-Attention mechanism and developed DeepViT. The re-attention mechanism regenerates attention maps at a minimal computational cost, enhancing the diversity of feature representations across layers. In the Vision transformer model design, simply replacing the self-attention mechanism with the Re-Attention mechanism significantly improves performance.
Vision transformer (ViT)
This study employs VIT51 relatively smaller models for plant disease detection and classification. Although vision transformer models outperform convolutional neural network models in accuracy metrics, Transformer-based models are becoming more like convolutional neural networks due to their powerful local feature processing capabilities. It transforms images into sequences for computational handling in the visual domain, dividing them into smaller patches and arranging them as a sequence. Specifically, the image is first divided into several smaller patches of the same size, which are then placed in a sequence. This entire sequence is fed into the Transformer encoder for feature extraction. The resulting feature map is then passed into a fully connected neural network for image classification.
Swin transformer
This study employs the Swin transformer model52 for plant disease detection and classification and is implemented using a hierarchical transformer structure. This hierarchical design is based on a sliding window operation, which includes non-overlapping local windows and overlapping cross-windows. By limiting attention computation within a single window, the Swin Transformer introduces the locality of CNN convolutions while also saving computational resources. The Swin transformer has shown excellent performance across various image tasks.
Convolutional swin transformer (CST)
This research proposed a CST network based on the Swin Transformer architecture. It utilizes image data to build a network that can automatically diagnose plant diseases and assess their severity. The model is designed to be highly robust, ensuring reliable detection accuracy for agricultural production. Additionally, Swin Transformer’s window attention design greatly reduces model complexity, making it more suitable for deployment. The CST design integrates many features of the Swin transformer, but the attention mechanisms differ. The model consists of the following stages: image cutting, convolution layer, layer normalization, window attention, multilayer perceptron, sliding window attention, and image fusion using two Swin Transformer modules. The first module employs a fixed window attention mechanism, while the second uses a sliding window attention mechanism. The number of channels in the hidden layer and the model layers represent the number of Swin Transformer modules in each stage. The proposed CST uses a single \(7 \times 7\) convolutional layer to produce a \(224 \times 224 \times 3\) feature map, matching the original Swin transformer’s input image size. Image fusion and classification using a fully connected neural network.
Proposed AttCM-Alex algorithm architecture
Vision Transformers have proven successful in computer vision due to their attention mechanisms. Two main directions in the field are designing pure Transformer models without convolutional layers or integrating attention mechanisms into convolutional neural networks. CNNs excel at learning pixel-level features, making them ideal for detecting changes in image brightness. This study uses a CNN framework to explore the impact of light intensity on image analysis. The AttCM module integrates attention mechanisms into \(3 \times 3\) convolutional layers, enhancing feature learning but increasing model complexity. To maintain efficiency for mobile devices, we use the simpler AlexNet architecture with the AttCM module.
Figure depicts the model architecture of AttCM-Alex that integrates the AttCM module into AlexNet, enhancing feature learning with attention mechanisms in \(3 \times 3\) convolutional layers. The architecture includes convolutional layers, ReLU activations, pooling layers, and fully connected layers, ensuring robust plant disease detection under varying light conditions while maintaining efficiency for mobile devices.
Figure 4 illustrates the AlexNet architecture, which is divided into two stages: the first stage consists of convolutional and pooling layers, while the second stage comprises fully connected layers. For a clear comparison between the proposed model and the original network structure, Figure 4 presents the AlexNet structure above and the AttCM-Alex structure below. The modifications introduced in this study are as follows: First, all \(3 \times 3\) convolutional layers are replaced with AttCM modules to obtain more comprehensive image feature information. Specifically, three \(3 \times 3\) convolutional layers are replaced with \(3 \times 3\) AttCM modules. Second, the number of filters in certain convolutional layers is adjusted. SpeciA simple convolutional layer was added to the Swin Transformer to introduce inductive bias into the model convolutional layer, from 256 to 192. Third, considering the richer information represented by the feature maps processed by AttCM, three additional fully connected layers are added to mAlexNet. In contrast, the proposed mAlexNet includes five fully connected layers with parameters of 2048, 1024, and 512, respectively, in addition to the original design. These modifications enhance the feature learning capabilities of the network, making it more robust for plant disease detection under varying light conditions while maintaining computational efficiency.
Attention convolution module (AttCM)
The AttCM block is calculated by splitting feature maps into convolution and attention channels after three convolutional layers. Convolution involves fully connected layers and fusion, while attention uses self-attention. Outputs are combined with learnable weights to produce the final feature map.
The computation process of the AttCM module is divided into two distinct parts, as illustrated in Fig. 5. First, the original feature maps are passed through three \(1 \times 1\) convolutional layers, reshaping the feature maps into four parts, resulting in \(3 \times 4\) feature maps enriched with feature information. Second, these feature maps are processed through two separate channels: the convolution channel and the attention channel. In the convolution channel, the feature maps first pass through a fully connected network to generate nine feature maps. These maps undergo shifting and aggregation operations ?, resulting in convolution-processed feature maps. The shifting operation is computed as per Eq. (5).
In Eq. (5), (i, j) represents the pixel position of the feature tensor; f is the input pixel before processing; \(\Delta x\) and \(\Delta y\) correspond to horizontal and vertical displacements, respectively. The shifting operation is \(f_{i+\Delta x, j+\Delta y}\), which outputs the pixel with added horizontal and vertical displacements based on \(f_{i, j}\). Assuming K represents the size of the convolution kernel, g is the output feature map, and (p, q) represents the position coordinates of the convolution kernel, the convolution calculation formula for the convolution channel can be derived as Eq. (6).
The aggregation operation is defined in Eq. (7). Summing the results of Eq. (6) across the entire image yields the output of the convolution channel, denoted as \(F_{\text {convolution}}\).
Subsequently, the feature maps are processed in the attention channel, comprising three parts derived from different \(1 \times 1\) convolutions. These parts function as Queries (Q), Keys (K), and Values (V). Self-attention scores are then computed, and the results are fused into a single feature map, \(F_{\text {attention}}\). Finally, the outputs from both the convolution and attention channels are combined, with learnable scalar weights \(\alpha\) and \(\beta\) controlling their relative importance, as illustrated in Eq. (8).
Optimized attention mechanisms and multi-head self-attention
The Vision Transformer excels in computer vision due to its use of the attention mechanism, distinct from CNNs. It employs the scaled dot-product attention mechanism17, as shown in Eq. (9).
In Eq. (9), Q, K, and V are Queries, Keys, and Values. Attention represents the computed attention scores, and \(\sqrt{d_q}\) is the dimension of Q. Q, K, and V are calculated using Eq. (10) with trainable weight matrices \(W_q\), \(W_k\), and \(W_v\).
The calculation process for scaled dot-product attention involves the following steps:
-
1.
Firstly, the input image \(X = [X_1, X_2,..., X_n]\) is transformed using trainable weight matrices to obtain three initial vector representations.
-
2.
The attention scores for each part’s contribution weight are determined by performing the dot product operation on the Q and K matrices, with an initial score of 112, as shown in Eq. (10)
-
3.
The model’s gradients are stabilized by scaling down the dot product result by \(\sqrt{d_q}\) and applying softmax normalization, resulting in a scaled-down attention score of 14, normalized to 0.88.
-
4.
Finally, multiply the result by V to obtain a weighted V value, which is the output tensor z.All outputs are sequentially arranged and combined to obtain the output tensor Z.
Multi-head attention is a technique that enhances the attention mechanism’s ability to learn data features by learning the same set of information multiple times and concatenating multiple outputs into a fully connected neural network, thereby capturing richer feature information.
Label smoothing in cross-entropy loss for improved model regularization
The study emphasizes deep learning and regularization methods to enhance model performance on the training set and generalization on the test set. It employs label smoothing to convert hard labels into soft ones, thereby improving generalization and smoothing the optimization process. This regularization method prevents overfitting during training and enhances classification accuracy by utilizing the cross-entropy function as a loss function. The label smoothing formula is illustrated in Eq. (11).
In Eq. (11), the process of label smoothing is described where K is the number of label categories, \(\alpha\) is a very small number (specifically \(\alpha = 0.1\) in this study), \(y_k\) denotes the original label typically in a one-hot encoded format, and \(y_{LS_k}\) signifies the smoothed label, where LS stands for Label Smoothing. The Eq. (11) indicates that label smoothing involves multiplying the original label by \((1 - \alpha )\) and then adding \(\frac{\alpha }{K}\), resulting in the final label for the sample.
This study applied the label smoothing cross-entropy loss function to enhance network accuracy in plant disease classification. The formulation of the cross-entropy loss function is depicted in Eq. (12).
In binary classification, \(y_k\) represents the original label, with 1 for positive class samples and zero otherwise, and p represents the probability of predicting the positive class for each class prediction. The computational illustration of the label smoothing cross-entropy loss function is illustrated in Eq. (13). Specifically, it replaces the label \(y_k\) in Eq. (12) with the smoothed label \(y_{LS_k}\) as shown in Eq. (11).
Experiments and result analysis
Experimental environment and hyperparameters fine tuning
This study utilized an Ubuntu 20.04 system equipped with an Intel Core(TM) i7-10875H CPU. Deep learning tasks were performed using the PyTorch framework and accelerated on an NVIDIA GeForce RTX 3050 Ti GPU. The GPU features 3584 CUDA cores and 20GB of GDDR6X memory, operating at a core frequency of 1500 MHz, providing a floating-point operation capability of 16.2 TFLOPs. The detailed training hyperparameters for all the models utilized in this investigation are outlined in Table 5.
Performance analysis proposed models
Based on the Plant Village dataset, the study examines the effectiveness of CNN and Transformer models for plant disease detection in complex and stable environments. Table 6 presents performance metrics of transformer and CNN-based models on Cucumber, Banana, and Tomato datasets. It details their complexity, parameters, memory usage, inference time, and performance metrics (accuracy and F-score). The models compared include AlexNet, EfficientNetV2, MobileNetV3, SqueezeNet, ResNet, ConvNeXt, Swin Transformer, DeepViT, and MaxViTsmall. The average and standard deviation of accuracy and F-score are also provided. CNN-based models showed higher average accuracy than Transformer models in challenging conditions, with AlexNet achieving a detection accuracy of 0.909, as depicted in Table 6. At the same time, MaxViTSmall was the leading Transformer model with an accuracy of 0.891. In the banana dataset, CNN models outperformed Transformers, with MobileNetV3 achieving an accuracy of 0.878. In the Tomato dataset, MobileNetV3 recorded an accuracy of 0.964, while MaxViTSmall achieved 0.901. The consistency in accuracy between CNN and Transformer models across these datasets demonstrates their robust performance in various plant disease detection scenarios. In stable environments, CNN models achieved an average accuracy of 0.9359, significantly higher than the Transformer models’ 0.799 average accuracy. MobileNetV3 achieved a detection accuracy of 0.964, while LeViT was the best-performing Transformer model with an accuracy of 0.966. CNN models exhibit superior performance and consistency in stable environments, outperforming Transformer models without specific optimizations for plant disease detection.
We compared the AttCM-Alex model with EfficientNet models to evaluate performance differences. Our results show that the AttCM-Alex model outperforms EfficientNet in terms of accuracy and computational efficiency, particularly in agricultural applications where robustness to environmental factors such as lighting variations and noise is crucial.
Detection performance analysis proposed AttCM-Alex model
(a) Confusion matrix under the cucumber dataset (b) Confusion matrix under the banana dataset (c) Confusion matrix under the tomato dataset.
The study evaluates AttCM-Alex’s performance in detecting plant diseases in complex backgrounds, specifically in a cucumber dataset. The model achieved a detection accuracy of 0.953, a 5.3% improvement from the original AlexNet. Despite increased parameters, the complexity decreased from 0.61 to 0.5GFLOPS, indicating a significant improvement in the model’s ability to diagnose plant diseases. The proportion of correctly classified and misclassified samples was similar, with 102 out of 109 healthy samples correctly classified and 103 out of 109 diseased samples correctly classified, as shown in Fig. 6a.
Subsequently, we analyzed the performance of AttCM-Alex in detecting various plant diseases in complex backgrounds. The banana dataset, comprising images of two diseases and healthy leaves, presents a typical multi-class classification problem. The detection results on this dataset provide insights into the model’s capability to identify different types of plant diseases. As shown in Table 6, AttCM-Alex achieved the highest classification accuracy of 0.971, representing a 14% improvement over the original AlexNet. It was the only model to surpass an accuracy of 0.91 on the dataset. The number of correctly classified images was consistent across categories, with AttCM-Alex correctly classifying 0.99, 0.98, and 0.97 images in healthy, black stripe leaf spot, and yellow bacterial blight categories, indicating minimal classification bias, as depicted in Fig. 6b. These results demonstrate AttCM-Alex’s effectiveness in complex environments and outperform other models in detecting plant diseases in plant leaves. The study evaluates the performance of the AttCM-Alex model in detecting plant diseases in stable environments. The tomato dataset includes ten disease categories and is a multiclass classification problem. AttCM-Alex achieved a detection accuracy of 0.979, while LeViT-128s achieved 0.996. Out of 50 test images for each category, only one had fewer than 46 correctly classified images. The remaining categories had more than 45 correctly classified images, as shown in Fig. 6c. Despite this, AttCM-Alex demonstrated excellent performance in detecting different plant diseases in stable environments without significant classification bias.
Robustness analysis in noisy image detection
The study reveals that increasing image brightness doesn’t significantly impact a model’s detection accuracy, with some models even showing higher accuracy under increased brightness conditions, as depicted in Table 7. However, when image brightness decreases, the model’s accuracy is affected to varying degrees. Only three out of 14 models show a decrease in accuracy when brightness increases by 10%, while the remaining models either increase or remain unchanged. The proposed model, AttCM-Alex, maintains stable recognition accuracy under six different brightness levels, demonstrating excellent robustness in reducing the impact of light intensity on accuracy. CNN-based models outperformed Transformer models in this evaluation, achieving an average accuracy of 0.847 without changes to image brightness. AlexNet and LeViT accuracy does not exceed 0.832 when image brightness increases. However, the proposed AttCM-Alex maintains an accuracy of no less than 0.94, starting with an accuracy of 0.94 without increasing image brightness. As image brightness decreases, the accuracy of the three models decreases, posing challenges for plant disease detection tasks.
Robustness analysis based on grayscale image
To assess the AttCM-Alex model’s robustness further, we performed an additional experiment using grayscale images generated from the original multispectral dataset. These grayscale images remove color information and retain only luminance, allowing us to evaluate the model’s ability to detect plant diseases using texture and shape features. The results, as shown in Table 8, demonstrate that while the model’s accuracy decreases slightly with grayscale images, it still performs robustly, particularly for diseases relying more on texture. These findings highlight the AttCM-Alex model’s adaptability to different types of data, confirming its potential utility in real-world agricultural scenarios where color information may not always be available.
Ablation analysis
(a) Detection accuracy when the screen brightness increases. (b) Detection accuracy when the screen brightness decreases.
The study demonstrates that the proposed mAlexNet model increases the model’s parameter count by 54.16 million, occupies an additional 674.2M of memory, and increases the detection time by 1.21 s. However, when the model only incorporates the AttCM module, the model’s complexity decreases by 0.22G FLOPs, and the model parameters decrease by 1.35 million. The model’s memory is reduced by 16M, but the detection time increases by 1.63 seconds, as illustrated in Table 6. The proposed model simultaneously uses the AttCM module and the mAlexNet, reducing complexity by 0.5GFLOPs, parameters by 66.24 million, and memory by 813.21 m. However, the model requires larger memory space for storage and significantly increases the time needed for model inference under unchanged hardware conditions. Despite increasing hardware requirements, the model significantly improves the detection performance. When using the newly designed mAlexNet and introducing the AttCM module separately, the model’s detection accuracy improves to varying degrees. Under the cucumber dataset, the model’s detection accuracy is maximally improved from 0.953 to 0.979. However, under the tomato dataset, the detection accuracy decreases by 3.2% when only the mAlexNet is used.
The proposed model, AttCM-Alex, achieves a recognition accuracy of 0.97 even when image brightness increases by 30%, surpassing the accuracy of other models, as plotted in Fig. 7a. This is significantly higher than models using only the mAlexNet or the AttCM module. The model shows an increasing trend in accuracy as the brightness increases, while the other models decrease with increasing brightness, as shown in Fig. 7b. When image brightness decreases by 30% all four models’ accuracy decreases to varying degrees. AttCM-Alex still achieves a detection accuracy of 0.978, while the original network model only achieves a detection accuracy of 0.799. It is noteworthy that AttCM-Alex has the highest detection accuracy at all brightness levels. The proposed model significantly improves the model’s accuracy and demonstrates superior robustness when image brightness changes.
(a) ROC comparison curve showing AUC values for the proposed models. (b) Performance metrics of the proposed models demonstrate the benefits of attention mechanisms. (c) Training and validation accuracy curves over 50 epochs, indicating steady progress and convergence. (d) Training and validation loss curves over 50 iterations, illustrating consistent error rate reduction and model resilience.
Figure 8 depicts the thorough performance study of the AttCM-Alex model developed for the identification of plant diseases. The AUC curve assesses classification model performance by plotting the True Positive Rate against the False Positive Rate, with higher values indicating better performance. Figure 8a shows the AUC comparison of four models such as AlexNet, mAlexNet, AttCM, and AttCM-Alex. The AUC values for each model were 0.73, 0.82, 0.87, and 0.98, respectively. The results demonstrated that AttCM-Alex had the highest AUC of 0.98, indicating the best performance among the evaluated models. AttCM also showed strong performance with an AUC of 0.87, benefiting from including attention mechanisms. mAlexNet improved upon the standard AlexNet with an AUC of 0.82, while AlexNet had the lowest AUC of 0.73, indicating the least discriminatory power. Figure 8b compares AlexNet, AttCM, mAlexNet, and AttCM-Alex in accuracy, specificity, sensitivity, and AUC. AlexNet has an accuracy of 82%, specificity of 65%, sensitivity of 75%, and an AUC of 0.73. AttCM performs better with 90% accuracy, 80% specificity, 85% sensitivity, and an AUC of 0.87. mAlexNet shows improvements with 85% accuracy, 75% specificity, 80% sensitivity, and an AUC of 0.82. AttCM-Alex excels with 98% accuracy, 97% specificity, 97% sensitivity, and an AUC of 0.98. This highlights the performance boost from incorporating attention mechanisms in AttCM and AttCM-Alex. Figure 8c displays the accuracy curves for training and validation over 50 epochs. Initially (0–10 epochs), both accuracies rise rapidly from 0.84 to 0.94. Between epochs 10 and 30, the accuracy improves steadily, with the training accuracy slightly higher. From epochs 30 to 50, the curves plateau, with training accuracy reaching 0.96 and validation accuracy at 0.98. This steady progress and convergence indicate that the model has learned effectively and can generalize well across various datasets, demonstrating strong learning performance and good generalization with minimal overfitting. Figure 8d exhibits the training and validation loss curves for AttCM-Alex, with training loss up to 0.08 and validation loss up to 0.04. The graph shows that from iterations 0–10, losses decrease rapidly from 0.45 to 0.1, steadily decreasing between iterations 10 and 20, and finally stabilize at around 0.05 and 0.08 between iterations 20 to 50. This graph demonstrates strong learning performance, effective training, and sound generalization to unseen data, highlighting the model’s resilience and effectiveness.
The AttCM-Alex model ranks at the top of tested models and has significant complexity and execution time. However, it has higher hardware requirements, with a parameter count and memory usage. The model’s average rank across these metrics is 15. The model ranks first in accuracy metric tests in complex and stable environments. However, when considering all detection performance, it drops to second place. The SqueezeNet model, which requires the least memory among the proposed models, achieves an average ranking of lower in accuracy tests. Despite these improvements, the model’s detection accuracy and robustness are not as high as those of other models. Therefore, the AttCM-Alex model improves its performance due to increased hardware requirements.
Discussion
This study presents a novel AttCM-Alex model that combines CNNs and ViTs to improve plant disease detection. The empirical findings depict that the suggested model outperforms several state-of-the-art models based on the same datasets, including traditional CNN-based architectures and other deep learning methods. Specifically, the AttCM-Alex model achieves higher accuracy and better robustness in handling environmental factors such as lighting variations and noise, which are common in agricultural settings. Our research first evaluates the performance of Transformer-based and CNN-based models in stable environments using the Plant Village dataset, a widely adopted benchmark for plant disease detection. The results confirm that most high-performance models excel in controlled settings. Several Transformer and CNN models are trained and tested on Plant Village subsets, revealing their respective advantages and limitations in detecting plant diseases. However, performance in stable environments does not guarantee effectiveness in real-world agricultural applications, where factors such as illumination variations, background noise, and crop type diversity significantly impact detection accuracy.
To assess practical applicability, we further evaluate model performance in complex environments using three publicly available datasets (cucumber, banana, and tomato datasets). The results indicate that AttCM-Alex achieves the highest accuracy of 0.953 on the cucumber dataset, while it reaches 0.971 on the banana dataset. Compared to previous studies, our model demonstrates superior performance in classifying Cucumber, Banana, and Tomato diseases, with notable improvements in precision and FScore. Prior CNN-based models, such as AlexNet and ResNet, showed high accuracy under ideal conditions but struggled with noise interference and fluctuating lighting conditions, leading to significant performance degradation in real-world applications.
Noise interference is a common issue in real-world agricultural image acquisition, as cameras may introduce various noise levels during data collection. This study evaluates the robustness of AttCM-Alex by introducing salt-and-pepper noise at different intensities to simulate real-world conditions. Experimental results show that detection accuracy decreases as noise increases, but AttCM-Alex maintains superior robustness. Specifically, AttCM-Alex achieves 0.90 accuracy with 30% noise on grayscale images, while other models show a significant drop in performance. These results confirm that AttCM-Alex effectively mitigates noise-related performance degradation. Beyond noise resistance, this study also examines the impact of varying illumination levels on disease detection accuracy, a crucial factor in agricultural applications. Illumination changes directly affect image brightness, which in turn alters the leaf features essential for disease identification. Although grayscale images slightly reduced the model’s overall accuracy, particularly in cases where color is a key distinguishing factor, the model still exhibited robust generalization capabilities, achieving relatively high performance. This suggests that while color information is beneficial, AttCM-Alex is sufficiently flexible to handle variations in input data, making it suitable for scenarios where multispectral or color information is limited.
Our methodology overcomes the limitations of previous approaches by integrating a self-attention mechanism, which allows the model to capture long-range dependencies within image data, and a channel attention mechanism, which dynamically emphasizes important features during classification. These innovations enhance the AttCM-Alex model’s ability to handle real-world agricultural data, making it a more reliable tool for disease detection in diverse and uncontrolled environments. Despite its advancements, AttCM-Alex still has limitations. While it performs well in noisy and varying lighting conditions, it may require further fine-tuning when applied to datasets with significantly different characteristics. Future research should explore transfer learning techniques to improve adaptability to different crop types and disease categories, further enhancing generalizability and robustness. Additionally, the large parameter size of AttCM-Alex poses challenges for mobile and edge deployment, making model compression and optimization an essential area for future development.
Conclusion
In this study, we introduce AttCM-Alex, an advanced methodology for plant disease detection under complex environmental conditions. The AttCM module combines convolutional operations with self-attention mechanisms, significantly enhancing the model’s robustness and accuracy. Our experimental results reveal that AttCM-Alex outperforms traditional models like AlexNet in various challenging scenarios, including different image brightness and noise levels. AttCM-Alex demonstrated significant improvements in detection accuracy, maintaining high performance even with considerable levels of salt-and-pepper noise. For instance, with 30% noise added, the model achieved a detection accuracy of 0.97 on the banana dataset, underscoring its robustness against noisy inputs. Furthermore, the model excelled under varying image brightness conditions, achieving a detection accuracy of 0.956 with a 30% increase in brightness and maintaining an accuracy of 0.91 with a 30% decrease in brightness. These results indicate a substantial improvement over other models tested. Therefore, AttCM-Alex represents a significant advancement in plant disease detection, offering high accuracy and robustness under various challenging conditions. Future work will focus on optimizing the model to reduce its parameter size and computational requirements, enhancing its applicability to resource-constrained environments such as mobile devices and edge computing platforms.
Data availability
This study used the publicly available dataset from the “Cucumber Disease Recognition Dataset” from Kaggle Dataset1 and “Banana Leaf Spot Diseases” from Kaggle Dataset2, as key supporting evidence.
Change history
30 September 2025
The original online version of this Article was revised: In the original version of this Article, Muhammad Akram Mujahid was incorrectly affiliated with ‘Shandong Research Institute of Industrial Technology, Jinan, 250000, China’. The correct affiliation is ‘Department of Information Sciences, University of Education, Township Campus, Lahore, 54000, Pakistan’.
References
Yu, S., Xie, L. & Huang, Q. Inception convolutional vision transformers for plant disease identification. Internet Things 21, 100650 (2023).
He, D. C. et al. Triple bottom-line consideration of sustainable plant disease management: From economic, sociological and ecological perspectives. J. Integr. Agric. 20, 2581–2591 (2021).
Pfordt, A. & Paulus, S. A review on detection and differentiation of maize diseases and pests by imaging sensors. J. Plant Dis. Prot. 132, 1–21 (2025).
Altieri, M. A. Agroecology: The Science of Sustainable Agriculture (CRC Press, 2018).
Aboelenin, S., Elbasheer, F. A., Eltoukhy, M. M., El-Hady, W. M. & Hosny, K. M. A hybrid framework for plant leaf disease detection and classification using convolutional neural networks and vision transformer. Complex Intell. Syst. 11, 142 (2025).
Abade, A., Ferreira, P. A. & de Barros Vidal, F. Plant diseases recognition on images using convolutional neural networks: A systematic review. Comput. Electron. Agric. 185, 106125 (2021).
Li, L., Zhang, S. & Wang, B. Plant disease detection and classification by deep learning-a review. IEEE Access 9, 56683–56698 (2021).
Upadhyay, A. et al. Deep learning and computer vision in plant disease detection: a comprehensive review of techniques, models, and trends in precision agriculture. Artif. Intell. Rev. 58, 1–64 (2025).
Ulukaya, S. & Deari, S. A robust vision transformer-based approach for classification of labeled rices in the wild. Comput. Electron. Agric. 231, 109950 (2025).
Sharma, P., Berwal, Y. P. S. & Ghai, W. Performance analysis of deep learning cnn models for disease detection in plants using image segmentation. Inf. Process. Agric. 7, 566–574 (2020).
Moen, E. et al. Deep learning for cellular image analysis. Nat. Methods 16, 1233–1246 (2019).
Tan, M. & Le, Q. Efficientnet: Rethinking model scaling for convolutional neural networks. In International Conference on Machine Learning, 6105–6114 (2019).
Krizhevsky, A., Sutskever, I. & Hinton, G. E. Imagenet classification with deep convolutional neural networks. Adv. Neural. Inf. Process. Syst. 25, 1097–1105 (2012).
He, K., Zhang, X., Ren, S. et al. Deep residual learning for image recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 770–778 (2016).
Huang, G., Liu, Z., Van Der Maaten, L. et al. Densely connected convolutional networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 4700–4708 (2017).
Howard, A. G., Zhu, M., Chen, B. et al. Mobilenets: Efficient convolutional neural networks for mobile vision applications. ArXiv preprint arXiv:1704.04861 (2017).
Dosovitskiy, A., Beyer, L., Kolesnikov, A. et al. An image is worth 16x16 words: Transformers for image recognition at scale. In International Conference on Learning Representations (2020).
Zhu, X., Su, W., Lu, L. et al. Deformable detr: Deformable transformers for end-to-end object detection. ArXiv preprint arXiv:2010.04159 (2020).
Ren, S., He, K., Girshick, R. et al. Faster r-cnn: Towards real-time object detection with region proposal networks. Adv. Neural Inf. Process. Syst. 28 (2015).
Simonyan, K. & Zisserman, A. Very deep convolutional networks for large-scale image recognition. ArXiv preprint arXiv:1409.1556 (2014).
Bajpai, A., Sahu, S. & Tiwari, N. K. Integrating attention mechanisms and squeeze-and-excitation blocks for accurate potato leaf disease detection. Potato Res. 1–21 (2025).
Khandelwal, I. & Raman, S. Analysis of transfer and residual learning for detecting plant diseases using images of leaves. In Computational Intelligence: Theories, Applications and Future Directions-Volume II: ICCI-2017, 295–306 (2019).
Singh, U. P. et al. Multilayer convolution neural network for the classification of mango leaves infected by anthracnose disease. IEEE Access 7, 43721–43729 (2019).
Zeng, T., Li, C., Zhang, B. et al. Rubber leaf disease recognition based on improved deep convolutional neural networks with a cross-scale attention mechanism. Front. Plant Sci.13 (2022).
Zhu, L., Li, X., Sun, H. & Han, Y. Research on cbf-yolo detection model for common soybean pests in complex environment. Comput. Electron. Agric. 216, 108515 (2024).
Nawaz, M. et al. Coffeenet: A deep learning approach for coffee plant leaves diseases recognition. Expert Syst. Appl. 237, 121481 (2024).
Mingyue, S. et al. Research progress of deep learning in plant leaf disease detection and recognition [j]. Smart Agric. 4, 29–46 (2022).
Ma JunCheng, M. J., Du KeMing, D. K., Zheng FeiXiang, Z. F., Zhang LingXian, Z. L. & Sun ZhongFu, S. Z. Disease recognition system for greenhouse cucumbers based on deep convolutional neural network. (2018).
Thai, H.-T. & Le, K.-H. Mobileh-transformer: Enabling real-time leaf disease detection using hybrid deep learning approach for smart agriculture. Crop Prot. 189, 107002 (2025).
Sharma, V., Tripathi, A. K., Mittal, H. & Nkenyereye, L. Soyatrans: A novel transformer model for fine-grained visual classification of soybean leaf disease diagnosis. Expert Syst. Appl. 260, 125385 (2025).
Monisha, R., Tamilselvan, K. & Sharmila, A. Advancing plant disease detection with hybrid models: Vision transformer and cnn-based approaches. In Computational Intelligence in Internet of Agricultural Things, 275–307 (Springer, 2024).
Fu, X. et al. Crop pest image recognition based on the improved vit method. Inf. Process. Agric. 11, 249–259 (2024).
Barman, U. et al. Vit-smartagri: vision transformer and smartphone-based plant disease detection for smart agriculture. Agronomy 14, 327 (2024).
Borhani, Y., Khoramdel, J. & Najafi, E. A deep learning based approach for automated plant disease classification using vision transformer. Sci. Rep. 12, 11554 (2022).
Zhang, Q., Sun, B., Cheng, Y. et al. Residual self-calibration and self-attention aggregation network for crop disease recognition. Int. J. Environ. Res. Public Health. 18 (2021).
Qian, X., Zhang, C., Chen, L. et al. Deep learning-based identification of maize leaf diseases is improved by an attention mechanism: Self-attention. Front. Plant Sci. 13 (2022).
Wang, P., Niu, T., Mao, Y. et al. Identification of apple leaf diseases by improved deep convolutional neural networks with an attention mechanism. Front. Plant Sci. 12 (2021).
Tang, Z. et al. Grape disease image classification based on lightweight convolution neural networks and channelwise attention. Comput. Electron. Agric. 178, 105735 (2020).
Sultana, N., Shorif, S. B., Akter, M. & Uddin, M. S. Cucumber disease recognition dataset. Mendeley Data. 10, y6d3z6f8z9 (2022).
Arman, S. E. et al. Bananalsd: A banana leaf images dataset for classification of banana leaf diseases using machine learning. Data Brief. 109608 (2023).
Hughes, D., Salathé, M. et al. An open access repository of images on plant health to enable the development of mobile disease diagnostics. arXiv preprint arXiv:1511.08060 (2015).
Khan, S. et al. Transformers in vision: A survey. ACM Comput. Surv. CSUR 54, 1–41 (2022).
Albahli, S. & Masood, M. Efficient attention-based CNN network (eanet) for multi-class maize crop disease classification. Front. Plant Sci. 13 (2022).
Alirezazadeh, P., Schirrmann, M. & Stolzenburg, F. Improving deep learning-based plant disease classification with attention mechanism. Gesunde Pflanzen 75, 49–59 (2023).
Chen, H.-C. et al. Alexnet convolutional neural network for disease detection and classification of tomato leaf. Electronics 11, 951 (2022).
Howard, A. et al. Searching for mobilenetv3. In Proceedings of the IEEE/CVF International Conference on Computer Vision, 1314–1324 (2019).
Kumar, V., Arora, H., Sisodia, J. et al. Resnet-based approach for detection and classification of plant leaf diseases. In 2020 International Conference on Electronics and Sustainable Communication Systems (ICESC), 495–502 (IEEE, 2020).
Iandola, F. N. et al. Squeezenet: Alexnet-level accuracy with 50x fewer parameters and< 0.5 mb model size. arXiv preprint arXiv:1602.07360 (2016).
Brock, A., De, S., Smith, S. L. & Simonyan, K. High-performance large-scale image recognition without normalization. In International Conference on Machine Learning, 1059–1071 (PMLR, 2021).
Zhou, D. et al. Deepvit: Towards deeper vision transformer. arXiv preprint arXiv:2103.11886 (2021).
Thakur, P. S., Khanna, P., Sheorey, T. & Ojha, A. Vision transformer for plant disease detection: Plantvit. In International Conference on Computer Vision and Image Processing, 501–511 (Springer, 2021).
Liu, Z. et al. Swin transformer: Hierarchical vision transformer using shifted windows. In Proceedings of the IEEE/CVF International Conference on Computer Vision, 10012–10022 (2021).
Acknowledgements
This research is supported by the Artificial Intelligence & Data Analytics Lab (AIDA) CCIS Prince Sultan University, Riyadh, Saudi Arabia. The authors are thankful for the support.
Funding
This research received no specific funding from any funding agency.
Author information
Authors and Affiliations
Contributions
Conceptualization, Z.Z., T.M., A.R., Y.W., and M.A.M. Data curation, Z.Z., T.M., A.R., and Y.W. Formal analysis, Z.Z., T.M., Y.W., and M.A.M. Methodology, Z.Z., T.M., A.R., and M.A.M. Software, Z.Z., A.R., Y.W., and M.A.M. Supervision, T.M.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Institutional Review Board Statement
All methods were carried out in accordance with relevant guidelines and regulations.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zeng, Z., Mahmood, T., Wang, Y. et al. AI-driven smart agriculture using hybrid transformer-CNN for real time disease detection in sustainable farming. Sci Rep 15, 25408 (2025). https://doi.org/10.1038/s41598-025-10537-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-10537-6