Abstract
Cerebral amyloid angiopathy (CAA) and insomnia are age-related neurological disorders increasingly recognized as being closely associated. However, research on the shared genes and their biological mechanisms remains limited. This study aims to identify common genes between CAA and insomnia and explore their potential molecular mechanisms, offering new insights for diagnosis and treatment. Blood samples were collected from 11 CAA patients and 11 healthy controls, followed by RNA sequencing (RNA-seq). Additionally, the microarray dataset GSE208668 for the insomnia cohort was downloaded from the Gene Expression Omnibus (GEO) database. Differential expression analysis was performed to identify common differentially expressed genes (DEGs). Protein-protein interaction (PPI) networks and machine learning methods Random Forest (RF) and Extreme Gradient Boosting (XGBoost) were used to narrow down key genes. We explored the biological functions of these genes through immune cell infiltration, metabolic and Hallmark pathway analyses, and clinical correlation analysis. Co-expression networks, drug-mRNA networks, transcription factor (TF)–mRNA–miRNA networks, and competing endogenous RNA (ceRNA) networks were also constructed. Finally, hub gene expression patterns were analyzed using the Human Protein Atlas (HPA) database, and validation was performed in clinical samples and animal models using quantitative Reverse Transcription Polymerase Chain Reaction (qRT-PCR) and Western blot. Differential expression analysis identified 185 DEGs. PPI network construction and machine learning methods identified CBX5 and POLR1B as common hub genes for both insomnia and CAA. Immune infiltration, metabolic, and Hallmark pathway analyses revealed these hub genes play distinct roles in each disease. Various network models were constructed to explore their regulatory mechanisms. The reliability of the hub genes was validated using bioinformatics analyses and experimental approaches. This study, combining bioinformatics and experimental validation, identifies CBX5 and POLR1B as shared hub genes for CAA and insomnia. These findings offer new molecular targets for the diagnosis and treatment of both diseases, providing a foundation for future research.
Similar content being viewed by others
Introduction
CAA is a type of cerebral small vessel disease predominantly affecting the elderly. Its characteristic pathological feature is the deposition of amyloid-β (Aβ) in the walls of small arteries, veins, and capillaries in the leptomeninges, cerebral cortex, and cerebellar cortex, leading to structural and functional vascular damage, ultimately impairing neurological function1. CAA is primarily sporadic and rarely manifests as a monogenic disease. The main clinical manifestations of CAA include spontaneous intracerebral hemorrhage, transient focal neurological episodes, cognitive impairment, and dementia2. Emerging evidence suggests a strong link between cerebrovascular dysfunction and sleep disturbances. Cerebral small vessel diseases (CSVDs) such as CAA are increasingly recognized for their role in disrupting sleep–wake regulation, possibly through shared mechanisms including neuroinflammation, glymphatic system dysfunction, and blood–brain barrier breakdown3,4,5,6,7,8,9. Insomnia is the most common sleep disorder, with 19–50% of adults reporting insomnia, which is closely related to aging10,11,12,13,14,15. The primary clinical feature of insomnia is chronic difficulty in initiating or maintaining sleep, which severely impacts daily life. Insomnia is not only associated with various neurodegenerative diseases such as Alzheimer’s disease (AD)16 and Parkinson disease(PD)17, but also with CSVD. Studies show that 54% of CSVD patients meet the criteria for chronic insomnia, a rate significantly higher than in healthy elderly individuals18. These patients exhibit increased sleep fragmentation, reduced sleep efficiency, and a decreased percentage of slow-wave sleep. Moreover, in CSVD patients, the severity of insomnia is linked to cognitive decline19. Treatment of insomnia has also been shown to alleviate comorbid disorders20. While some research suggests that improving sleep can reduce early Aβ accumulation, and that acute sleep disruption leads to increased amyloid production, these studies are limited to the context of AD21. Our preliminary research found that sleep deprivation exacerbates Aβ vascular deposition in a CAA mouse model, leading to memory and cognitive deficits. However, KDM6B knockdown reduced memory impairment induced by sleep deprivation22.
Although previous studies have explored the comorbidity between CSVD and insomnia, most have focused on clinical or pathological perspectives and often treated CSVD as a unified entity. However, CAA is a distinct subtype of CSVD, characterized by the deposition of amyloid-β in the walls of cortical and leptomeningeal vessels, and its specific association with insomnia remains largely unexplored.
Notably, no prior studies have directly investigated shared molecular signatures between CAA and insomnia using patient-derived biological data. This represents a critical gap in the current understanding of the pathophysiological overlap between these two conditions. Our study addresses this gap by being the first to perform an integrative transcriptomic analysis aimed at identifying shared gene expression alterations in CAA and insomnia. Peripheral blood, as a minimally invasive and readily accessible tissue source, offers a practical window into systemic molecular dysregulation and serves as an ideal substrate for early-stage exploration of disease comorbidity at the transcriptomic level. Moreover, this approach enhances translational potential by allowing for biomarker discovery in clinically feasible samples.
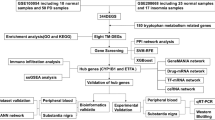
To this end, we analyzed RNA-Seq data from peripheral blood of clinically diagnosed CAA patients and microarray-based transcriptomic data from an insomnia cohort in the GEO database. By comparing each disease group with matched healthy controls, we identified significant DEGs, and further pinpointed commonly dysregulated genes and enriched biological pathways23,24. Through integrated bioinformatics analysis and experimental validation in both human and animal models, we identified CBX5 and POLR1B as key hub genes and explored their associations with immune responses, metabolic pathways, potential therapeutic compounds, and non-coding RNA networks.These findings provide the first molecular-level evidence linking CAA and insomnia, offering new insights into their shared pathophysiology. By uncovering common transcriptomic alterations, this study lays a foundation for future mechanistic research and the development of diagnostic and therapeutic strategies for CAA–insomnia comorbidity. The overall workflow is summarized in Fig. 1.
Overall workflow of this study.
Materials and methods
Study design
From January 1, 2023, to December 1, 2024, CAA patients (n = 11) were consecutively recruited from the Department of Neurology at the First Affiliated Hospital of Zhengzhou University. Diagnosis was independently performed by two experienced neurologists according to the Boston Criteria Version 2.025. The control group (n = 11) consisted of participants without any signs of neurological disorders. There were no significant differences in age and sex between the two groups. Peripheral venous blood samples from all 22 participants were collected for RNA-seq. The clinical characteristics of the groups are summarized in Supplementary Table S1. Additionally, 5 CAA patients and 5 healthy controls, along with 5 chronic insomnia patients and 5 healthy controls, were recruited for subsequent experimental validation. Chronic insomnia inclusion criteria were as follows: all insomnia patients reported at least three episodes of sleep difficulties per week, lasting for more than three months, with other sleep disorders excluded. All participants were free of significant physical diseases, such as malignancies or autoimmune disorders, and had no acute infections or recent use of medications that could significantly interfere with the experiment (e.g., anti-inflammatory or immunosuppressive drugs)26,27. This study was approved by the Ethics Committee of the First Affiliated Hospital of Zhengzhou University and was conducted in accordance with the principles of the Helsinki Declaration. Informed consent was obtained from all CAA patients, insomnia patients, and their corresponding healthy control participants (approval number: 2023-KY-0990).
RNA-seq and raw read processing
Total RNA from animal tissues was extracted using TRIzol Reagent (Life Technologies, California, USA) following the manufacturer’s instructions. The RNA-seq was performed by the Biomarker Technologies. RNA concentration and purity were assessed using the NanoDrop 2000 (Thermo Fisher Scientific, Wilmington, DE). RNA integrity was evaluated using the Agilent Bioanalyzer 2100 system (Agilent Technologies, CA, USA) with the RNA Nano 6000 kit. mRNA libraries were prepared using the Hieff NGS Ultima Dual-mode mRNA Library Prep Kit for Illumina (Yeasen Biotechnology, Shanghai, China). Sequencing was performed on the Illumina NovaSeq platform, generating 150 bp paired-end reads. Raw sequencing data in FASTQ format underwent quality control, read alignment, and transcript quantification to generate to generate log₂-CPM values. The RNA-seq data processing pipeline and quality control results including sequencing statistics and PCA-based sample distribution plots are provided in Supplementary Material 1.
Public data provenance and differential expression analysis
The microarray datasets GSE208668, corresponding to the insomnia cohort, and GSE39445, used as the validation cohort, were both obtained from the GEO database. Detailed information regarding these datasets is provided in Supplementary Table S2. The microarray datasets were processed using standard gene expression normalization followed by log₂ transformation, as per conventional microarray preprocessing workflows. DEGs were identified between 11 CAA samples and 11 controls, as well as between 17 insomnia samples and 25 control samples from the GSE208668 dataset, using the R package ‘limma‘(v3.54.2)28. Genes with a p-value < 0.05 and an absolute log2 fold change (|log2FC|) > 0.5 were considered significant DEGs. Volcano plots and heatmaps were generated to visualize DEG expression patterns.
Functional enrichment analysis
Gene Ontology (GO)29,30 and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses31,32,33 were performed on common DEGs using the R package ‘clusterProfiler’ (v4.7.1)34. Results with an adjusted p-value (FDR) < 0.05 were considered statistically. A complete list of enriched GO terms and KEGG pathways is provided in Supplementary Table S3.
PPI network construction
The STRING database was used to construct the PPI network35, which was visualized using Cytoscape v3.9.1 software36. The CytoHubba plugin v0.1(compatible with Cytoscape_v3.9.1) was employed to identify hub genes based on topological analysis37. Specifically, nine centrality algorithms were applied: Maximal Clique Centrality (MCC), Density of Maximum Neighborhood Component (DMNC), Maximum Neighborhood Component (MNC), Closeness Centrality, Degree Centrality, Edge Percolated Component (EPC), Radiality Centrality, Betweenness Centrality, and Stress Centrality. For each algorithm, the top 40 ranked genes were retained, and the final hub gene list was obtained by intersecting the results from all nine methods. The detailed gene selection process is provided in Supplementary Table S4.
Identifying hub genes using machine learning
Machine learning–based feature selection was performed using the randomForest (v4.7-1.1)38 and xgboost (v1.7.5.1)39 packages in R (v4.2.2). Data preprocessing and visualization used tidyverse (v2.0.0)40 and ggplot2 (v3.5.0)41, while sparse matrix conversion for XGBoost was handled with the Matrix package (v1.5-4)42. ROC and AUC calculations were performed using pROC (v1.18.1)43.
Co-expression network of hub genes
To identify genes associated with the function of hub genes, a co-expression network analysis was performed using GeneMANIA, an online tool for constructing gene co-expression networks44.
Immune infiltration analysis
For the CAA and insomnia datasets, we analyzed immune cell infiltration in both control and disease groups using Cell-type Identification By Estimating Relative Subsets Of RNA Transcripts (CIBERSORT)45. Differentially expressed immune infiltrating cells were identified using the t-test, and the correlation between characteristic genes and these immune infiltrating cells was explored through Spearman correlation analysis.
Hallmark pathways and metabolic pathway enrichment analysis
Hallmark gene sets were downloaded from the Molecular Signatures Database (MSigDB) database46, and metabolic pathway gene sets were obtained from the KEGG database31,32,33 as input files for single-sample gene set enrichment analysis (ssGSEA) enrichment analysis47. Significant differential pathways were calculated based on the enrichment scores for each sample and analyzed for correlation with hub genes48.
Construction of Drug-mRNA, cerna, and TF-mRNA-miRNA networks
Drugs predicted to target the hub genes were identified from the drug signatures database (DSigDB) database49, and drug-RNA interaction networks were constructed. To predict miRNAs targeting these biomarkers, we utilized the StarBase database, selecting the top 10 miRNAs for each hub gene based on clipExpNum (number of supporting CLIP-seq experiments ≥ 5) for subsequent lncRNA identification50. Ultimately, mRNA-miRNA-lncRNA regulatory networks were established. Additionally, transcription factors (TFs) regulating these biomarkers were predicted using the JASPAR database51, leading to the construction of TF-mRNA-miRNA interaction networks. All networks were visualized using Cytoscape software.
Animals
All animal experiments were approved by the Institutional Animal Care and Use Committee of the First Affiliated Hospital of Zhengzhou University (approval number: ZZU-LAC20240322[03]) and were conducted in accordance with the ARRIVE 2.0 guidelines.The 3R principles (Reduction, Refinement, and Replacement) were strictly followed to ensure statistical robustness while minimizing animal use and suffering. To assess gene expression differences in CAA, five 12-month-old homozygous male TgSWDI transgenic mice and five age-matched wild-type littermates were utilized. The TgSWDI transgenic mice were sourced from Jackson Laboratory. In addition, five 12-week-old male C57BL/6J wild-type mice were subjected to sleep deprivation in a transparent cylindrical device (KW-BD, Mouse Sleep Deprivation System, Calvin Biotech, China), where a rotating metal rod (10 rotations per minute) prevented the mice from sleeping for 20 h daily. Five control mice were housed in the same apparatus with unrestricted access to food and water, thereby maintaining a natural sleep cycle. After one week of continuous sleep deprivation, the mice were used for subsequent experimental procedures. All animals were maintained under standard laboratory conditions at 25 °C, with a 12-hour light/dark cycle, and had free access to food and water.
Experimental sample Preparation
Peripheral blood mononuclear cells (PBMCs) were extracted from the venous blood of human participants using Ficoll-Paque PLUS (Cytiva) density gradient centrifugation. Mice were deeply anesthetized with 3% isoflurane and subsequently euthanized by cervical dislocation, in accordance with institutional animal care guidelines. Brains were rapidly harvested on ice from both CAA and insomnia models, and hippocampi were dissected for downstream analyses.
qRT-PCR
Total RNA was extracted from the aforementioned PBMCs and hippocampus samples using TRIzol reagent (Invitrogen) according to the manufacturer’s instructions. Reverse transcription was performed using the HiScript IV RT SuperMix (Vazyme), and reactions were carried out using the qRT-PCR kit (Vazyme). Each group had triplicate controls. RNA concentration and purity were assessed using a Nanodrop-2000 spectrophotometer (ThermoFisher Scientific). Primers used in this study are listed in Supplementary Table S5 and were synthesized by Sangon Biotech (Shanghai, China).In addition, we performed 2% agarose gel electrophoresis to validate the specificity of primers used in each qRT-PCR assay. Detailed experimental procedures and electrophoresis results are provided in Supplementary Material 2.
Western blot
Tissue samples from the hippocampus and PBMCs were lysed in Radioimmunoprecipitation assay (RIPA) buffer (Solarbio, China) containing protease and phosphatase inhibitors. Protein concentrations were determined using the bicinchoninic acid (BCA) protein assay kit (Beyotime, Shanghai, China). Proteins were diluted in 5× SDS sample buffer (Beyotime, China) and boiled at 100 °C for 10 min. Protein samples were separated by 12% SDS-PAGE and transferred to polyvinylidene fluoride membranes. After blocking, membranes were incubated overnight at 4 °C with appropriate primary antibodies (CBX5 antibody, Proteintech, 1:1000; POLR1B antibody, ThermoFisher Scientific, 1:1000). After washing, membranes were incubated with secondary antibodies at room temperature (AffiniPure goat anti-rabbit, 1:5000; AffiniPure goat anti-mouse, 1:5000). Protein bands were visualized using enhanced chemiluminescence (ECL™) reagents Western Blotting Reagents (Millipore, USA), and band intensities were quantified using ImageJ software (NIH, USA).The corresponding uncropped gel electrophoresis images are provided in Supplementary Material 3.
Statistical analysis
Statistical analyses were conducted using R software version 4.2.2 and GraphPad Prism 8. For comparisons between two groups, normally distributed data with equal variances were analyzed using the Student’s t-test, while data with unequal variances were assessed using the Wilcoxon test. The correlations between continuous variables were calculated using the Spearman rank correlation coefficient. A p-value of less than 0.05 was considered statistically significant. The significance levels are indicated as *P < 0.05, **P < 0.01, and ***P < 0.001.
Ethics statement
All animal procedures were approved by the Institutional Animal Care and Use Committee of the First Affiliated Hospital of Zhengzhou University (approval number: ZZU-LAC20240322[03]) and complied with the Animals Research: Reporting of In Vivo Experiments (ARRIVE) guidelines 2.0. All methods adhered to relevant guidelines and regulations.
Results
Identification and functional enrichment analysis of DEGs
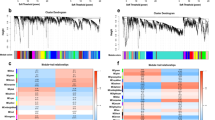
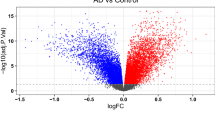
We collected blood samples from 11 patients with CAA and 11 age- and sex-matched healthy controls for RNA-seq. Principal component analysis demonstrated clear segregation of cases and controls in both cohorts (Supplementary Material 1). A total of 2,345 DEGs were identified, including 16 upregulated genes and 2,329 downregulated genes (Fig. 2A and B). Additionally, we obtained gene expression data from the peripheral blood of 17 insomnia patients and 25 age- and sex-matched controls from the GEO database. A total of 6,069 DEGs were identified between insomnia and control samples, with 3,438 upregulated genes and 2,631 downregulated genes (Fig. 2C and D). We then conducted an intersection analysis of the upregulated and downregulated genes in both the CAA and insomnia datasets, identifying 7 commonly upregulated genes and 178 commonly downregulated genes. (Fig. 2E). In total, 185 DEGs were identified as shared between insomnia and CAA. To further elucidate their biological functions, we performed GO enrichment analysis and KEGG pathway enrichment analysis. GO enrichment analysis revealed that these shared DEGs were predominantly involved in biological processes related to protein synthesis and modification, DNA metabolism, and cellular regulation, including protein demannosylation, protein alpha-1,2-demannosylation, and positive regulation of the DNA metabolic process. KEGG pathway enrichment analysis further highlighted significant associations with peripheral immune processes and key signal transduction pathways31,32,33, such as the T cell receptor signaling pathway, B cell receptor signaling pathway, Ras signaling pathway, and prolactin signaling pathway (Fig. 2F).
Identification and enrichment analysis of DEGs. (A) Volcano plot of differential expression analysis between the CAA and control groups. (B) Heatmap of differentially expressed gene clusters between the CAA and control groups. (C) Volcano plot of differential expression analysis between the insomnia and control groups. (D) Heatmap of differentially expressed gene clusters between the insomnia and control groups. (E) Venn diagram illustrating the intersection of upregulated and downregulated genes shared between the CAA and insomnia groups. (F) GO and KEGG enrichment analysis of DEGs.
PPI network and machine learning model selection
We constructed a PPI network of 185 DEGs using the STRING database and visualized it in Cytoscape v3.9.1. After filtering, the final network included 128 nodes and 225 edges (Fig. 3A). Next, we analyzed the network with 9 algorithms from the cytoHubba plugin, which led to the identification of 17 marker genes at their intersection (Fig. 3B). To refine the gene selection process, we applied RF and XGBoost models to both the CAA and insomnia datasets. This analysis pinpointed two shared hub genes: POLR1B and CBX5 (Fig. 3C and D). Following this, we validated the significant downregulation of these hub genes in the disease cohorts compared to control groups (Fig. 3E). ROC curve analysis demonstrated that, in the CAA cohort, POLR1B and CBX5 achieved AUCs of 0.793 (95% CI 0.554–0.975) and 0.826 (95% CI 0.628–0.975), respectively, reflecting strong diagnostic performance. In the insomnia cohort, their AUCs were 0.633 (95% CI 0.440–0.819) and 0.616 (95% CI 0.426–0.798), with lower confidence limits below 0.50, indicating only modest and uncertain discriminative utility (Fig. 3F). Furthermore, correlation analysis revealed a positive relationship between POLR1B and CBX5 across both datasets, suggesting their synergistic roles in the shared pathogenic mechanisms of CAA and insomnia (Fig. 3G).
Identification of hub genes using PPI network and machine learning. (A) Construction of the PPI network. (B) Identification of core genes using multiple PPI algorithms. (C) Selection of hub genes in the CAA dataset using XGBoost and RF models. (D) Selection of hub genes in the insomnia dataset using XGBoost and RF models. (E) Expression validation of hub genes in both the insomnia and CAA datasets. (F) ROC curve analysis of hub genes. (G) Correlation analysis between POLR1B and CBX5.
Immune infiltration analysis
There is substantial evidence suggesting a close relationship between CAA, insomnia, and peripheral immunity26,52. To further investigate this connection, we analyzed the immune landscape of the CAA and insomnia datasets using CIBERSORT (Fig. 4A). We excluded immune cell types with less than 75% cell abundance, retaining 14 immune cell types in the CAA dataset and 15 in the insomnia dataset. In CAA patients, monocyte levels were elevated, while the number of activated NK cells was significantly reduced. Conversely, in insomnia patients, the levels of monocytes, CD4 memory activated T cells, gamma-delta T cells, and resting mast cells were markedly decreased. Meanwhile, CD4 memory resting T cells, CD8 T cells, and memory B cells were notably increased. These findings reveal distinct immune responses between CAA and insomnia (Fig. 4B). Further correlation analysis showed that the expression of the hub genes was strongly correlated with peripheral immune cell infiltration, with consistent trends across both diseases. However, the correlations between hub genes and specific immune cells varied significantly between the two conditions. In the CAA dataset, CBX5 and POLR1B expression were positively correlated with CD4 memory T cells and negatively correlated with M0 macrophages. In the insomnia dataset, both genes showed positive correlations with memory B cells, while exhibiting negative correlations with gamma-delta T cells and monocytes (Fig. 4C). These results suggest a close interaction between the hub genes and the immune microenvironment, providing a theoretical foundation for further exploration of their specific mechanisms in these diseases.
Immune infiltration analysis in the CAA and insomnia datasets. (A) Distribution of 22 immune cell types in the disease and control groups. (B) Box plot illustrating differences in immune cell infiltration between disease and control groups. (C) Correlation analysis between hub genes and infiltrating immune cells.
Correlation of hub genes with metabolic and hallmark pathways
To explore the shared functional pathways of the hub genes in both diseases, we conducted ssGSEA analysis on metabolic and Hallmark pathways. Disruptions in PI metabolism, inositol phosphate metabolism, creatine metabolism, phospholipid metabolism, and the metabolism of ingested semet sec mesec into H2SE were observed in both diseases (Supplementary Material 4 A). Additionally, Hedgehog signaling, peroxisome signaling, UV response, and E2F target pathways were also imbalanced in both conditions (Supplementary Material 4B). Notably, insomnia patients exhibited more pronounced disruptions in these pathways. Correlation analysis of the hub genes with these key pathways revealed that CBX5 was significantly associated with PI metabolism and phospholipid metabolism, showing positive correlations in CAA and negative correlations in insomnia (Supplementary Material 4 C). POLR1B was positively correlated with the dysregulation of inositol phosphate metabolism in both diseases. Regarding Hallmark pathways, CBX5 showed differential correlations with the E2F target pathway and Hedgehog signaling across the two diseases. In CAA, POLR1B exhibited a positive correlation with the E2F target pathway, whereas in insomnia, a negative correlation was observed (Supplementary Material 4D). These results underscore the distinct metabolic and Hallmark functional pathways linked to the two hub genes, POLR1B and CBX5, in both insomnia and CAA, providing deeper insights into their potential roles in the pathophysiology of these disorders.
Correlation analysis of hub genes with clinical parameters
We conducted a correlation analysis between the expression of the two hub genes and clinical parameters. The results revealed that, in the CAA dataset, both CBX5 and POLR1B were negatively correlated with the percentage of monocytes and homocysteine levels, and positively correlated with total protein levels. Additionally, only CBX5 was positively correlated with free thyroxine levels.In the insomnia dataset, both CBX5 and POLR1B were negatively correlated with Beck Depression Inventory (BDI) and BDI-NS scores, while only CBX5 was positively correlated with education level (Fig. 5).
Correlation analysis between hub genes and multiple clinical indicators.
Co-expression network and functional analysis of hub genes
The co-expression network for the hub genes was constructed using GeneMANIA with an automatic weighting method. This gene-gene interaction network consisted of two central hub genes (CBX5 and POLR1B) and 20 peripheral predicted genes, including POLR1A, POLR1C, POLR3B, and others (Fig. 6A). We performed GO enrichment analysis for the two hub genes and the 20 predicted genes. The results revealed significant enrichments in aspects of gene transcription regulation, particularly RNA polymerase I-mediated transcription, epigenetic regulation, and chromatin structure regulation. Additionally, KEGG pathway enrichment highlighted associations with immune and inflammatory processes, such as the Cytosolic DNA-sensing pathway and Neutrophil extracellular trap formation (Fig. 6B and C).
Co-expression network and functional analysis. (A) Co-expression network model of hub genes. (B) GO enrichment analysis of the co-expression network model. (C) KEGG enrichment analysis of the co-expression network model.
Drug-mRNA, CeRNA network, and TF-mRNA-miRNA construction
Using the DSigDB53 database, we predicted small molecule drugs targeting the hub genes CBX5 and POLR1B. A total of 45 small molecules were identified as targeting CBX5, while 22 small molecules targeted POLR1B (Fig. 7A, Supplementary Table S6). Notably, 11 drugs were found to target both hub genes, including sanguinarine, vincristine, and hydrogen peroxide, among others. These drugs may simultaneously modulate the activity of these hub genes in the context of CAA comorbid with insomnia, potentially leading to improved therapeutic outcomes. We selected the top 10 miRNAs associated with CBX5 and POLR1B, based on clipExpNum, along with their corresponding predicted lncRNAs to construct a ceRNA network (Fig. 7B, Supplementary Table S7). Notably, a single lncRNA can regulate multiple miRNAs, such as lncRNA XIST, which concurrently regulates hsa-miR-3200-5p, hsa-miR-212-3p, hsa-miR-132-3p, and hsa-miR-214-3p.Based on the JASPAR database, we constructed a TF-mRNA-miRNA regulatory network. Our analysis revealed 20 transcription factors regulating CBX5 and 13 regulating POLR1B expression (Fig. 7C, Supplementary Table S8), with FOXL1, FOXC1, and PPARG identified as common transcriptional regulators of both hub genes.
Regulatory networks involving hub genes, including Drug-mRNA, TF-mRNA, and ceRNA interactions. (A) Drug-mRNA interaction network of hub genes. (B) ceRNA regulatory network of hub genes. (C) TF-mRNA-miRNA interaction network of hub genes.
Expression of CBX5 and POLR1B in peripheral blood single cells in the HPA database
We further investigated the expression patterns of CBX5 and POLR1B in PBMCs using the HPA database. The results revealed that CBX5 was primarily expressed in plasmacytoid dendritic cells and naive B cells, whereas POLR1B exhibited predominant expression in mucosal-associated invariant T (MAIT) cells and NK cells. These findings indicate that CBX5 and POLR1B exhibit preferential expression across distinct immune cell subsets. Such cell type-specific transcriptional patterns may underlie their specialized roles in immune regulation and offer mechanistic clues to the divergent immunopathological features observed in CAA and insomnia (Fig. 8).
Expression patterns of hub genes in PBMCs based on the HPA database.
Validating hub gene expression: bioinformatics and experimental validation
We first validated the expression of the hub genes using the sleep deprivation dataset (GSE39445). The results showed that, consistent with previous findings, the expression of the hub genes was significantly reduced in the sleep deprivation group (Fig. 9A). To further validate our bioinformatics findings, we recruited 5 CAA patients, 5 insomnia patients, and age- and sex-matched controls. qRT-PCR and Western blot analyses were conducted to assess the transcriptional and protein expression changes of CBX5 and POLR1B in PBMCs from both CAA and insomnia patients. Both qRT-PCR and Western blot results demonstrated significantly reduced expression of the hub genes in CAA and insomnia patients, in line with our bioinformatics analysis (Fig. 9B and C). Next, we examined the expression of these two hub genes in brain tissue. We assessed the protein expression levels of CBX5 and POLR1B in the hippocampus of CAA and insomnia model mice using qRT-PCR and Western blotting. The results revealed a decrease in CBX5 expression in the hippocampus of both CAA and insomnia model mice compared to the control group. Although POLR1B exhibited a downward trend, the change was not statistically significant (Fig. 9D and E).
Validation of hub genes. (A) Validation of hub gene expression in the GSE39445 dataset. (B) qRT-PCR validation of differential mRNA expression of hub genes in human peripheral blood samples. (C) Western blot validation of differential protein expression of hub genes in human peripheral blood samples. (D) qRT-PCR validation of mRNA expression of hub genes in mouse hippocampal samples. (E) Western blot validation of differential protein expression of hub genes in mouse hippocampal samples.
Discussion
CAA is a subtype of cerebral small vessel disease, characterized by the deposition of Aβ in the walls of small and medium-sized arteries, arterioles, and venules in the cerebral cortex and leptomeninges54. With the aging population, the clinical manifestations of CAA, primarily lobar hemorrhage and cognitive dysfunction, have gained increasing attention55. Sleep disturbances are common and challenging issues in aging, significantly affecting the overall health and well-being of the elderly, with potential negative impacts on cognition and mood56. Moreover, growing evidence supports a strong association between sleep disorders and CSVD. Previous cohort studies have shown that fragmented sleep is associated with cognitive dysfunction in CSVD patients, particularly affecting executive function and delayed recall57. A cross-sectional and longitudinal analysis involving 26,354 participants reported that unhealthy sleep behaviors are linked to increased white matter hyperintensities (WMH) and worse white matter microstructure in specific tracts58. Similarly, another study found that gray matter volume (GMV) of the right caudate and bilateral calcarine cortex was associated with sleep quality in CSVD patients59. In addition to clinical studies, animal models have further elucidated the pathological mechanisms underlying the relationship between CSVD and sleep disorders. Long-term fragmented sleep has been associated with pathological effects on the deep white matter in CSVD rat models, such as reduced myelin integrity, impaired oligodendrocyte precursor cell (OPC) maturation, and pro-inflammatory microglia polarization3.
Our previous research confirmed that sleep deprivation increases Aβ deposition in the microvasculature of CAA mouse models22. However, these findings were confined to animal studies, and to date, no clinical investigations have explored the molecular relationship between CAA and sleep disorders. In this study, we addressed this gap by utilizing clinical peripheral blood samples and employing an integrative multi-omics strategy to comprehensively identify shared hub genes and convergent molecular pathways linking CAA and insomnia.
Through differential expression analysis, in combination with PPI network construction and machine learning-based gene prioritization, we identified CBX5 and POLR1B as key hub genes commonly dysregulated in both conditions. Importantly, this dual-cohort, multi-modal study represents a novel paradigm in comorbidity research—shifting the focus from clinical associations to the molecular-level dissection of disease intersections. By directly linking neurovascular pathology and sleep dysfunction through shared transcriptomic alterations, our findings uncover a previously unrecognized molecular bridge between CAA and insomnia. Furthermore, CBX5 and POLR1B emerge not only as potential diagnostic biomarkers but also as promising cross-disease therapeutic targets. Their dual relevance suggests the possibility of developing targeted interventions that could simultaneously preserve vascular integrity and restore sleep homeostasis. Collectively, these findings provide a critical foundation for future mechanistic investigations and translational strategies aimed at addressing the complex interplay between neurovascular dysfunction and sleep disorders.
POLR1B, a key subunit of RNA polymerase I, plays an essential role in ribosomal RNA synthesis, protein synthesis, and fundamental cellular functions60. To date, research on POLR1B within neurological disorders remains limited, with the only reported association being cluster headache61, leaving its potential involvement in CAA and sleep disorders unexplored. Currently, studies on POLR1B primarily focus on cancers, including B-cell lymphoma, lung cancer, and prostate cancer62,63,64. These investigations suggest that POLR1B may influence tumorigenesis by regulating pathways linked to cell proliferation and survival—pathways similarly implicated in both CAA and insomnia. Further experimental studies are necessary to validate this mechanistic overlap. CBX5, which encodes heterochromatin protein 1α (HP1α), is a member of the HP1 family and a key reader of histone H3 lysine 9 di- and tri-methylation (H3K9me2/3). By mediating chromatin compaction and transcriptional repression, HP1α plays a central role in maintaining genome stability and regulating various cellular processes65. Functionally, HP1α has been implicated in several physiological systems. In vascular biology, it orchestrates endothelial progenitor cell (EPC) differentiation by forming distinct regulatory complexes that simultaneously repress stemness-associated genes and activate endothelial and angiogenic programs, thereby facilitating lineage transition and maintaining vascular integrity66. Similarly, in the innate immune system, CBX5 contributes to macrophage tolerance; its overexpression abolishes the trained immunity phenotype induced by MEK1/2 inhibition, suggesting a role as a negative regulator of macrophage priming via H3K9 methylation recognition67. Beyond these physiological contexts, most mechanistic insights into CBX5 have come from cancer models. In myelogenous leukemia, the RBMX/L1 complex regulates CBX5 transcription post-transcriptionally, thereby influencing chromatin dynamics and cell survival. Forced expression of CBX5 rescues proliferation and apoptosis defects caused by RBMX/L1 depletion, highlighting its therapeutic potential68. In intrahepatic cholangiocarcinoma, the HP1α–HDAC1 complex modulates STAT1 epigenetic activity and interferon signaling69. Additionally, in EGFR-mutant lung adenocarcinoma, CBX5 deficiency contributes to acquired resistance to EGFR tyrosine kinase inhibitors, identifying it as a possible target for overcoming treatment resistance70. Although these findings originate from oncologic studies and should not be directly extrapolated, they offer mechanistic insights into the broader regulatory functions of CBX5 in chromatin remodeling and transcriptional control, which may be relevant across disease contexts. Recent evidence also supports a role for CBX5 in the nervous system, though the literature remains relatively limited. In Alzheimer’s disease, Tau protein helps maintain pericentromeric heterochromatin (PCH) integrity by stabilizing H3K9me3 and HP1α distribution—a function disrupted in Tau-deficient or AD neurons36. In C9orf72-associated ALS/FTD, poly(PR) dipeptides interfere with HP1α phase separation and heterochromatin organization, resulting in aberrant histone methylation, nuclear lamina defects, and activation of repetitive elements71. In multiple sclerosis, decreased HP1α expression is associated with impaired Integrator complex function, global transcriptional dysregulation, and increased inflammatory vulnerability, underscoring its broader role in chromatin surveillance and gene expression homeostasis72.
Given the pivotal regulatory roles of CBX5 in angiogenesis, immune modulation, and neurological disorders, we hypothesize that CBX5 may also be involved in the pathological mechanisms underlying CAA and sleep disorders. Our previous research showed that sleep deprivation significantly increased KDM6B expression in the cerebral cortex of CAA mouse models. Knockdown of KDM6B inhibited PARP16 expression by enhancing H3K27me3 methylation, thereby mitigating memory deficits, neuronal damage, and vascular injury induced by sleep deprivation22. In the present study, we observed that CBX5, which encodes the HP1α protein and regulates H3K9me3 methylation, was commonly downregulated in both CAA and sleep disorder datasets. These findings suggest that CBX5, similar to KDM6B, may regulate immune and cellular function via epigenetic mechanisms, further highlighting the therapeutic potential of targeting epigenetic regulators in these conditions.
Consistent with this view, numerous studies have emphasized the role of the peripheral immune system in both CAA and insomnia52. Immune infiltration analysis revealed that monocyte activation was significantly elevated in the peripheral blood of CAA patients but reduced in those with insomnia. Interestingly, CBX5 and POLR1B expression levels were positively correlated with CD4 + memory T cells and negatively correlated with M0 macrophages; however, their associations with other immune cell types varied between the two conditions, suggesting disease-specific immune regulatory roles. In parallel, metabolic pathway analysis identified phosphatidylinositol metabolism, inositol phosphate metabolism, and phospholipid metabolism as commonly dysregulated pathways potentially influenced by the hub genes. POLR1B was particularly associated with inositol phosphate metabolism, and E2F targets and Hedgehog signaling emerged as hallmark pathways that may be co-regulated by CBX5 and POLR1B. Expression analysis showed that both CBX5 and POLR1B were significantly downregulated in the CAA group compared to normal controls, which is consistent with the observed increase in monocyte infiltration. In the insomnia cohort, hub gene expression was negatively correlated with BDI scores, implying a potential link between gene dysregulation and depressive symptoms. Together, these findings align with clinical observations and reinforce the potential involvement of CBX5 and POLR1B in the shared and divergent pathophysiological features of CAA and insomnia73.
Gene interaction networks constructed using GeneMANIA show that POLR1B and CBX5 interact with 22 related genes, with functional enrichment analysis indicating a strong association with RNA polymerase complex functions. Additionally, we further explored disease mechanisms and potential therapeutic targets by constructing drug-mRNA, TF-mRNA, and ceRNA networks. Regarding pharmacological research, cellular model experiments show that acetaminophen inhibits NF-κB activation and reduces inflammation induced by Aβ-derived peptides, suggesting its potential therapeutic effect in CAA74. Clinical studies have also found that acetaminophen can improve sleep75, supporting the hypothesis that CBX5 or POLR1B might play a key role in acetaminophen’s mechanism of action in treating CAA and insomnia. However, this hypothesis requires further experimental validation. Moreover, treatment of APP/PS1 mice, a classical CAA animal model, with retinoic acid reduced the activation of microglia and astrocytes, alleviated neuronal degeneration, and improved spatial learning and memory functions. These findings further support the potential of CBX5 and POLR1B as therapeutic targets for retinoic acid in CAA. Beyond acetaminophen and retinoic acid (RA), drugs like acacetin, vorinostat, apigenin, trichostatin A, and quercetin have shown improvement in AD and other neurodegenerative disease models76,77,78,79,80,81, but their therapeutic potential in CAA still needs further investigation. Due to the limited data on small RNA (miRNA) and drug studies in CAA, we referred to miRNAs reported in AD research, given the similarities in their pathological mechanisms36. Several miRNAs, such as hsa-miR-106a-5p, hsa-miR-125b-5p, hsa-miR-93-5p, and hsa-miR-17-5p, have been linked to AD pathogenesis82,83,84. This suggests that these miRNAs may also play a crucial role in the development of CAA. According to the ceRNA hypothesis, lncRNAs can regulate mRNA expression by competitively binding shared miRNAs, thereby forming post-transcriptional regulatory networks. Guided by this concept, we constructed a predicted ceRNA network centered on the hub genes CBX5 and POLR1B. While the proposed interactions suggest plausible regulatory relationships, the absence of matched lncRNA and miRNA expression data precluded correlation-based validation in this study. Nonetheless, this network serves as a hypothesis-generating framework that highlights potential regulatory axes worthy of further investigation. In future work, we plan to experimentally validate these predicted interactions using qRT-PCR, luciferase reporter assays, and other functional approaches.
To validate the reliability of our findings, we collected peripheral blood samples from both patients and controls and assessed the expression levels of hub genes using qRTPCR and Western blot methods. Consistent with our bioinformatics analysis, the hub genes were significantly downregulated in both diseases. Additionally, we investigated the relevance of these hub genes in the hippocampus of animal models. The results showed that CBX5 expression was reduced, while POLR1B exhibited no significant changes. The highly consistent expression trends of CBX5 in both peripheral blood and brain tissue suggest that CBX5 may play a critical bridging role in both insomnia and CAA, supporting its potential as a shared therapeutic target for both conditions.
Our study has several limitations. First, the insomnia dataset was drawn from public repositories, whereas the CAA cohort was prospectively assembled in-house, resulting in disparities in sample provenance and cohort size as well as in measurement platforms (microarray versus RNA-Seq), each employing unique collection protocols and preprocessing workflows that may introduce confounding biases. Second, the modest cohort size limits statistical power and may have constrained our ability to identify robust, shared biomarkers. Validation in larger, more diverse clinical populations will be essential to confirm and extend these findings. Third, although POLR1B and CBX5 demonstrated strong discriminative power in the CAA cohort, their diagnostic utility in the insomnia cohort was minimal—AUCs only marginally above 0.6, with lower confidence bounds falling below 0.50. This likely reflects our relatively small insomnia sample size, differences in sample origin and data processing workflows, and the distinct pathophysiological mechanisms of insomnia versus CAA as captured in peripheral blood. Consequently, although the point estimates suggest some ability to distinguish insomnia cases from controls, the wide, low-lying confidence intervals demand very cautious interpretation. Validation in larger, well‐characterized insomnia cohorts will therefore be essential. Fourth, direct functional validation experiments for POLR1B and CBX5 were not conducted in this study. To address this limitation, future investigations will include targeted in vitro and in vivo experiments, including gene knockout and overexpression approaches, to comprehensively delineate their mechanistic roles in the pathogenesis of both CAA and insomnia. Finally, our analysis primarily focused on peripheral blood transcriptomic data, despite the central nervous system being the predominant site of pathophysiology for both conditions. Although our findings were partially corroborated using mouse hippocampal tissue, human brain tissue samples were not evaluated. Consequently, future research will emphasize the collection and analysis of human brain tissues, alongside the inclusion of larger sample sizes, to substantiate and enhance the translational relevance and robustness of our conclusion.
Conclusion
This study identified CBX5 and POLR1B as key shared genes in both CAA and insomnia. Notably, CBX5 exhibited a more consistent expression pattern in human peripheral blood and mouse hippocampus across both diseases. However, the precise mechanisms underlying their involvement in CAA and insomnia remain unclear. Future research should focus on elucidating their regulatory pathways, which could pave the way for novel diagnostic and therapeutic strategies for these disorders.
Data availability
The RNA-seq data generated in this study for the CAA cohort have been deposited in the NCBI Sequence Read Archive (SRA) under accession number PRJNA1220702, and the corresponding expression matrix has also been submitted to the GEO database under accession number GSE289809. These datasets will be made publicly available upon publication. For the insomnia cohort, microarray datasets were retrieved from the GEO under accession numbers GSE208668 (training dataset) and GSE39445 (validation dataset). All datasets used in this study are publicly accessible through the NCBI GEO repository or are available from the corresponding author upon reasonable request. Supporting data are also provided in the main article and Supplementary Materials.
References
Charidimou, A. et al. Emerging concepts in sporadic cerebral amyloid angiopathy. Brain: J. Neurol. 140 (7), 1829–1850 (2017).
Lu, C. Q. et al. Quantitative comparison of CSVD imaging markers between patients with possible amyloid small vessel disease and with non-amyloid small vessel disease. NeuroImage. Clinical 44, 103681 (2024).
Fu, X. et al. Effects of sleep fragmentation on white matter pathology in a rat model of cerebral small vessel disease. Sleep, 47 (4), zsad225. (2024).
Bellesi, M. et al. Sleep loss promotes astrocytic phagocytosis and microglial activation in mouse cerebral cortex. J. Neuroscience: Official J. Soc. Neurosci. 37 (21), 5263–5273 (2017).
Kaneshwaran, K. et al. Sleep fragmentation, microglial aging, and cognitive impairment in adults with and without alzheimer’s dementia. Sci. Adv. 5 (12), eaax7331 (2019).
Parhizkar, S. et al. Sleep deprivation exacerbates microglial reactivity and Aβ deposition in a TREM2-dependent manner in mice. Sci. Transl. Med. 15 (693), eade6285 (2023).
Medina-Flores, F. et al. Sleep loss disrupts pericyte-brain endothelial cell interactions impairing blood-brain barrier function. Behav. Immun. 89, 118–132 (2020). Brain.
Carreras, A. et al. Chronic sleep fragmentation induces endothelial dysfunction and structural vascular changes in mice. Sleep 37 (11), 1817–1824 (2014).
Sauvet, F. et al. Total sleep deprivation alters endothelial function in rats: a nonsympathetic mechanism. Sleep 37 (3), 465–473 (2014).
Walsh, JK. et al. Nighttime insomnia symptoms and perceived health in the America insomnia survey (AIS). Sleep, 34 (8), 997–1011. (2011).
Abraham, O. et al. Factors contributing to poor satisfaction with sleep and healthcare seeking behavior in older adults. Sleep. Health. 3 (1), 43–48 (2017).
Morin, C. M. et al. Epidemiology of insomnia: prevalence, self-help treatments, consultations, and determinants of help-seeking behaviors. Sleep Med. 7 (2), 123–130 (2006).
Ryden, A. M. et al. Insomnia disorder among older veterans: results of a postal survey. J. Clin. Sleep. Medicine: JCSM : Official Publication Am. Acad. Sleep. Med. 15 (4), 543–551 (2019).
Koopman, A. D. M. et al. Prevalence of insomnia (Symptoms) in T2D and association with metabolic parameters and glycemic control: Meta-Analysis. J. Clin. Endocrinol. Metab. 105 (3), 614–643 (2020).
Sutton, E. L. Insomnia Annals Intern. Med. 174 (3), ITC33–ITC48 (2021).
Irwin, M. R. & Vitiello, M. V. Implications of sleep disturbance and inflammation for alzheimer’s disease dementia. Lancet Neurol. 18 (3), 296–306 (2019).
Li, X. Y. et al. Tryptophan metabolism-related gene CYP1B1 serves as a shared biomarker for both parkinson’s disease and insomnia. Sci. Rep. 15 (1), 1362 (2025).
Huang, B. H. et al. Sleep and physical activity in relation to all-cause, cardiovascular disease and cancer mortality risk. Br. J. Sports Med. 56 (13), 718–724 (2022).
Yan, W. et al. Reduced left hippocampal perfusion is associated with insomnia in patients with cerebral small vessel disease. CNS Spectr. 28 (6), 702–709 (2023).
Roth, T. Comorbid insomnia: current directions and future challenges. Am. J. Manag. Care. 15, S6–13 (2009).
Insel, P. S. et al. Association of sleep and β-Amyloid pathology among older cognitively unimpaired adults. JAMA Netw. Open. 4 (7), e2117573 (2021).
Yu, W. et al. KDM6B knockdown alleviates sleep deprivation-induced cerebrovascular lesions in APP/PS1 mice by inhibiting PARP16 expression. Biochem. Pharmacol. 231, 116650 (2025).
Han, Y. et al. Advanced applications of RNA sequencing and challenges. Bioinform. Biol. Insights. 9 (1), 29–46 (2015).
Rosati, D. et al. Differential gene expression analysis pipelines and bioinformatic tools for the identification of specific biomarkers: A review. Comput. Struct. Biotechnol. J. 23, 1154–1168 (2024).
Charidimou, A. et al. The Boston criteria version 2.0 for cerebral amyloid angiopathy: a multicentre, retrospective, MRI-neuropathology diagnostic accuracy study. Lancet Neurol. 21 (8), 714–725 (2022).
Piber, D. et al. Sleep disturbance and activation of cellular and transcriptional mechanisms of inflammation in older adults. Brain, Behavior, and Immunity, 106, 67–75 (2022).
Riemann, D. et al. The neurobiology, investigation, and treatment of chronic insomnia. Lancet Neurol. 14 (5), 547–558 (2015).
Ritchie, M. E. et al. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43 (7), e47 (2015).
Aleksander, S. A. et al. The gene ontology knowledgebase in 2023. Genetics, 224 (1), iyad031. (2023).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The gene ontology consortium. Nat. Genet. 25 (1), 25–29 (2000).
Kanehisa, M. et al. KEGG: biological systems database as a model of the real world. Nucleic Acids Res. 53 (D1), D672–D677 (2025).
Kanehisa, M. Toward Understanding the origin and evolution of cellular organisms. Protein Science: Publication Protein Soc. 28 (11), 1947–1951 (2019).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28 (1), 27–30 (2000).
Wu, T. et al. ClusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innov. (Cambridge (Mass)). 2 (3), 100141 (2021).
Szklarczyk, D. et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 47 (D1), D607–D613 (2019).
Mansuroglu, Z. et al. Loss of Tau protein affects the structure, transcription and repair of neuronal pericentromeric heterochromatin. Sci. Rep. 6, 33047 (2016).
Chin, C. H. et al. CytoHubba: identifying hub objects and sub-networks from complex interactome. BMC Syst. Biol. 8 (4), S11 (2014).
Altmann, A. et al. Permutation importance: a corrected feature importance measure. Bioinf. (Oxford England). 26 (10), 1340–1347 (2010).
Chen, T. & Guestrin, C. XGBoost: A Scalable Tree Boosting System, in Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining. Association for Computing Machinery: San Francisco, California, USA. 785–794. (2016).
and Max Kuhn et al. Welcome to the tidyverse. J. Open. Source Softw. 4, 1686 (2019).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis. 2 ed. Use R! New York, NY: Springer-Verlag New York. 260, (2016).
Bates, D. et al. Matrix: Sparse and Dense Matrix Classes and Methods. Comprehensive R Archive Network (CRAN). (2025).
Robin, X. et al. pROC: an open-source package for R and S + to analyze and compare ROC curves. BMC Bioinform. 12, 77 (2011).
Franz, M. et al. GeneMANIA update 2018. Nucleic Acids Res. 46 (W1), W60–W64 (2018).
Newman, A. M. et al. Determining cell type abundance and expression from bulk tissues with digital cytometry. Nat. Biotechnol. 37 (7), 773–782 (2019).
Liberzon, A. et al. The molecular signatures database (MSigDB) hallmark gene set collection. Cell. Syst. 1 (6), 417–425 (2015).
Wu, L. et al. IGSA: individual gene sets analysis, including enrichment and clustering. PloS One. 11 (10), e0164542 (2016).
Cui, H. K. et al. An integrative analysis of single-cell and bulk transcriptome and bidirectional Mendelian randomization analysis identified C1Q as a novel stimulated risk gene for atherosclerosis. Front. Immunol. 14, 1289223 (2023).
Yoo, M. et al. DSigDB: drug signatures database for gene set analysis. Bioinf. (Oxford England). 31 (18), 3069–3071 (2015).
Li, J. H. et al. StarBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res. 42 (Database issue), D92–D97 (2014).
Castro-Mondragon, J. A. et al. JASPAR 2022: the 9th release of the open-access database of transcription factor binding profiles. Nucleic Acids Res. 50 (D1), D165–D173 (2022).
Munsterman, D. et al. Cerebral amyloid angiopathy and the immune system. Alzheimer’s Dement. J. Alzheimer’s Assoc. 20 (7), 4999–5008 (2024).
García-Nieto, E. et al. Clinical and molecular characterization of steatotic liver disease in the setting of immune-mediated inflammatory diseases. JHEP Reports: Innov. Hepatol. 6 (10), 101167 (2024).
van Veluw, S. J. et al. Is CAA a perivascular brain clearance disease? A discussion of the evidence to date and outlook for future studies. Cell. Mol. Life Sci. 81 (1), 239 (2024).
Koemans, E. A. et al. Progression of cerebral amyloid angiopathy: a pathophysiological framework. Lancet Neurol. 22 (7), 632–642 (2023).
Kaur, S. et al. Sleep quality mediates the relationship between frailty and cognitive dysfunction in non-demented middle aged to older adults. Int. Psychogeriatr. 32 (5), 663 (2020).
Wang, J. et al. The influence of non-breathing-related sleep fragmentation on cognitive function in patients with cerebral small vessel disease. Neuropsychiatr. Dis. Treat. 15, 1009–1014 (2019).
Ning, J. et al. Association of sleep behaviors with white matter hyperintensities and microstructural injury: a cross-sectional and longitudinal analysis of 26 354 participants. Sleep, 46 (5), zsad020. (2023).
Zhao, J. et al. Differences in Gray matter volume in cerebral small vessel disease patients with and without sleep disturbance. Brain Sci., 13 (2), 294. (2023).
Andersen, J. S. et al. Nucleolar proteome dynamics. Nature 433 (7021), 77–83 (2005).
Harder, A. V. E. et al. Genetic susceptibility loci in genomewide association study of cluster headache. Ann. Neurol. 90 (2), 203–216 (2021).
Bywater, M. J. et al. Inhibition of RNA polymerase I as a therapeutic strategy to promote cancer-specific activation of p53. Cancer Cell. 22 (1), 51–65 (2012).
Yang, F. et al. POLR1B is upregulated and promotes cell proliferation in non-small cell lung cancer. Oncol. Lett. 19 (1), 671–680 (2020).
Craddock, J. et al. Alterations in the epigenetic machinery associated with prostate Cancer health disparities. Cancers, 15 (13), 3462 (2023).
Machida, S. et al. Structural basis of heterochromatin formation by human HP1. Mol. Cell, 69 (3), 385–397 (2018).
Maeng, Y. S. et al. Heterochromatin protein 1 alpha (HP1α: CBX5) is a key regulator in differentiation of endothelial progenitor cells to endothelial cells. Stem Cells (Dayton Ohio). 33 (5), 1512–1522 (2015).
Low, R. et al. Prolonged Inhibition of the MEK1/2-ERK Signaling Axis Primes Interleukin-1 Beta Expression through Histone 3 Lysine 9 Demethylation in Murine Macrophages. International Journal of Molecular Sciences, 24 (19), (2023).
Prieto, C. et al. Transcriptional control of CBX5 by the RNA binding proteins RBMX and RBMXL1 maintains chromatin state in myeloid leukemia. Nat. Cancer. 2, 741–757 (2021).
Xiong, F. et al. Correction: Unveiling the role of HP1α-HDAC1-STAT1 axis as a therapeutic target for HP1α-positive intrahepatic cholangiocarcinoma. J. Experimental Clin. Cancer Research: CR. 43 (1), 186 (2024).
Bugide, S. et al. CBX5 loss drives EGFR inhibitor resistance and results in therapeutically actionable vulnerabilities in lung cancer. Proc. Natl. Acad. Sci. U.S.A. 120 (4), e2218118120 (2023).
Zhang, Y. J. et al. Heterochromatin Anomalies and double-stranded RNA Accumulation Underlie C9orf72 poly(PR) Toxicity363, 6428. (Science, 2019).
Porozhan, Y. et al. Defective integrator activity shapes the transcriptome of patients with multiple sclerosis. Life Sci. Alliance, 7 (10), e202402586 (2024).
Staner, L. Comorbidity of insomnia and depression. Sleep Med. Rev. 14 (1), 35–46 (2010).
Bisaglia, M. et al. Acetaminophen protects hippocampal neurons and PC12 cultures from amyloid beta-peptides induced oxidative stress and reduces NF-kappaB activation. Neurochem. Int. 41 (1), 43–54 (2002).
Smith, G. M. & Smith, P. H. Effects of Doxylamine and acetaminophen on postoperative sleep. Clin. Pharmacol. Ther. 37 (5), 549–557 (1985).
Bu, J. et al. Acacetin improves cognitive function of APP/PS1 alzheimer’s disease model mice via the NLRP3 inflammasome signaling pathway. Translational Neurosci. 13 (1), 390–397 (2022).
Kilgore, M. et al. Inhibitors of class 1 histone deacetylases reverse contextual memory deficits in a mouse model of Alzheimer’s disease 35 (4), 870–880 (Official Publication of the American College of Neuropsychopharmacology, 2010).
Correction : Curcumin and Apigenin - novel and promising therapeutics against chronic neuroinflammation in Alzheimer’s disease. Neural Regeneration Research, 10 (12), 2017. (2015).
Venigalla, M., Gyengesi, E. & Münch, G. Curcumin and Apigenin - novel and promising therapeutics against chronic neuroinflammation in alzheimer’s disease. Neural Regeneration Res. 10 (8), 1181–1185 (2015).
Su, Q. et al. Trichostatin A ameliorates Alzheimer’s disease-related pathology and cognitive deficits by increasing albumin expression and Aβ clearance in APP/PS1 mice 13 (1), 7 (Alzheimer’s Research & Therapy, 2021).
Zhang, X. et al. Quercetin stabilizes Apolipoprotein E and reduces brain Aβ levels in amyloid model mice. Neuropharmacology 108, 179–192 (2016).
Yılmaz, Ş. G. et al. Can peripheral MicroRNA expression data serve as epigenomic (Upstream) biomarkers of alzheimer’s disease?? Omics: J. Integr. Biology. 20 (8), 456–461 (2016).
Quan, X. et al. Related network and differential expression analyses identify nuclear genes and pathways in the Hippocampus of alzheimer disease. Med. Sci. Monitor: Int. Med. J. Experimental Clin. Res. 26, e919311 (2020).
Nguyen, H. D., Kim, W. K., Huong, G. & Vu Molecular mechanisms implicated in protein changes in the alzheimer’s disease human hippocampus. Mech. Ageing Dev. 219, 111930 (2024).
Acknowledgements
Thanks for the fund support of the National Natural Science Foundation of China [Grant No.82171300] and major project co-constructed by the Ministry of Medical Science and Technology in Henan Province [Grant No. SBGJ202101016].
Funding
This study was supported by grants from a major project co-constructed by the Ministry of Medical Science and Technology in Henan Province (Grant No. SBGJ202101016) and National Natural Science Foundation of China (Grant No. 82171300).
Author information
Authors and Affiliations
Contributions
LXY and LYS conceived the study and were responsible for the study design and manuscript writing. LXY and LK analyzed the RNA-seq sequencing results. YWK and WYH performed the experiments. GK, HWJ, and WJH were responsible for data collection and study supervision. All co-authors have read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
All experimental procedures were approved by the Institutional Animal Care and Use Committee of the First Affiliated Hospital of Zhengzhou University (ZZU-LAC20240322[03]).
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, XY., Li, K., Wei, YH. et al. Exploration of common pathogenic genes between cerebral amyloid angiopathy and insomnia based on bioinformatics and experimental validation. Sci Rep 15, 26365 (2025). https://doi.org/10.1038/s41598-025-12553-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-12553-y
Keywords
This article is cited by
-
Target identification and mechanism elucidation of Chai Hu Shu Gan-San against the comorbidity of insomnia and depression based on integrative computational pharmacology
Network Modeling Analysis in Health Informatics and Bioinformatics (2025)