Abstract
Chilli cultivars are highly sensitive to diverse agroclimatic conditions and the present investigation provide a significant contribution by identifying high-yielding and stable hybrids for wider adaptation. The study was conducted in seven diverse environments by following conventional farming under field conditions in five locations of North-western Himalaya along with naturally ventilated polyhouse conditions using conventional and natural farming practices using 16 genotypes to identify the most stable ones for fruit yield and its attributes. Joint regression and AMMI analysis revealed significant Genotype × Environment interaction for all the traits. Eberhart and Russel (E-R) model revealed DPCHYB 10 (627.68 g/plant) and DPCHYB 5 (583.50 g/plant) as top ranked with stability for fruit yield. The AMMI1 biplot identified DPCHYB 2 as most resilient and productive across environments for fruit number and fruit yield which was also described by mean vs. stability biplot of GGE. Further, GGE suggested E5 (Berthin) as the most representative and discriminating environment for fruit yield. ‘Which won where’ biplot represented hybrid DPCHYB 10 as stable and high yielding with wider adaptability in most of the environments except Palampur (E1). DPCHYB 5 was the most responsive and adaptive in E1 that was also depicted as stable by E-R.
Similar content being viewed by others
Chilli (Capsicum annuum L. var. annuum) is known for its delicious and pungent taste and is a major spice crop worldwide particularly in tropical Asia and equatorial America1. It is consumed fresh or in dried forms and is the second most significant solanaceous vegetable after tomato. It is originated from South and Central America having chromosome number 2n = 2x = 24 and genome size of 3.48Gb2. It is a rich source of vitamin C, carotenoids, capsaicinoids, and capsinoids which have potential applications in cosmeceutical, food, spice, and pharmaceutical industries1,3. Capsaicin is the major cause of pungency in chilli which have varied preventive and beneficial uses in medicines (allopathic and ayurveda) and defence weapon industry3,4. Chilli is also known as hot pepper, paprika, red peppers, cayenne peppers, pimento, and capsicum in different parts of the world5, and is cultivated over an area of 2.06 million ha with a total production of 38.3 million tons and average productivity of 18.59 t ha−16.
It is a vital constituent of Indian cuisine and valuable cash crop which is cultivated over an area of 419 thousand hectares with production of 4436 thousand MT with productivity of 10.59 MT/ hectare7 during summer-rainy season as rainfed crop and during autumn/spring-summer season under irrigated conditions8. India is the largest exporter of chilli globally with a share of 30% of its total production and has exported 2,732 metric tonnes of green chillies worth 234.5 million USD in 2023-249. A considerable loss to chilli production is recorded due to its inherent sensitivity to environmental fluctuations10. The productivity of chilli can be enhanced in the shortest span of time by adopting hybrids which in general are uniform for different attributes along with resistance to various biotic and abiotic stresses. Male sterility is a crucial mechanism for controlled pollination and is widely exploited in commercial hybrid seed production of chilli. More than twelve monogenic recessive male sterile mutants have been discovered from natural sources and mutagenesis techniques11.
Genotype-environment interaction (GEI) is a major concern and can seriously destabilize efforts to select superior genotypes12. It is, therefore, essential to identify phenotypically stable genotypes with wider adaptability to varied climatic conditions across environments13. Such stable genotypes can be commercial exploited as they find favour by the farmers on account of high productivity vis-à-vis economic returns. It will further help in extenuating future worries viz., climate change, declining natural resources, and varied biotic and abiotic concerns etc. Multi-Environment Trials (MET) are conducted to evaluate advanced breeding lines or hybrids to regulate their stability and adaptability before recommendation for cultivation.
The assessment of GEI provides an idea of buffering potential of the population. The main result of GEI interaction would be to significantly reduce the influence of environment on the final appearance of plants14. A genotype with low GEI (stable) possesses a little or no change in the performance regardless of the weather vagaries. GEI reduces the predictability of genotypes performance in target environments. One of the major factors in plant breeding is the selection of appropriate test locations because it accounts for GE and maximizes gain from selection15. It is, therefore, necessary to evaluate varieties/ hybrids in multilocation to identify the most promising ones based on the parameters of stability and adaptability16. The identification of stable and high yielding cultivars/hybrids by the partitioning of genotype, environment, and GEI using various statistical tools forms the main objective of MET data analysis17.
A single stability model may not be representative to regulate the performance of hybrids over the environments and may lead to inconsistent conclusions about their stability. There are three groups of stability parameter namely, one associated only with yield and has least information on stability, second consider yield and stability simultaneously (AMMI, GGE biplot) while third group focus on stability and show little/no association with yield (joint regression analysis of Eberhart and Russel18. Earlier, the Eberhart and Russell18 regression model has been extensively used to assess linear (regression, bi) and nonlinear (deviation from regression, S−2di) components of GEI variation with independent response and considers both yield and stability with prediction of a particular genotypes for specific environments18. The stability of genotypes cannot reliably be predicted from this approach because it considers response of genotype to environment to be univariate issue which is otherwise a multivariate situation.
The AMMI biplot is a valuable tool for identifying stable and high-performing genotypes across varying environments. The AMMI offers various elucidations and information by dividing the main effect and interaction effects19, and is relatively effective for expecting G × E interaction20. Identification of site-specific genotypes following AMMI needs authentication of yield of the genotypes21 and mainly focusing G × E interaction and ignore main effects of genotypes. Yan et al.22 developed a biplot technique that was named as GGE biplot that graphically represents the main effect of genotype and, Genotype × Environment (G + G×E or GGE) effects and is an efficient method based on principal component analysis (PCA) to fully explore multi-environment trial data. The most interesting aspect of the GGE biplot is its ability to represent the ‘Which-won-where’ pattern of a genotype by environment data set. The genotypes in the GGE biplots that fall on the vertices of the polygons represent their degree of performance in a specific environment15. The earlier researchers have also employed these different models to predict stability and adaptability of chilli genotypes/hybrids in different growing conditions across globe23,24,25,26,27. Keeping these aspects in view, the present investigation was planned with objectives to compare the effectiveness of the Eberhart and Russell joint regression model, AMMI, and GGE biplot analysis to identify high yielding and stable GMS based chilli hybrids. This will simultaneously help to recommend hybrids for cultivation with least influence of climate fluctuations and better adaptation to biotic and abiotic stresses.
Materials and methods
Experimental material and layout plan
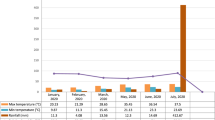
The present investigation was carried out involving 12 chilli hybrids and four checks (Table 1) which were evaluated at seven diverse environments during summer 2021 viz., field conditions at the Research Farm of the Department of Vegetable Science and Floriculture, College of Agriculture, Chaudhary Sarwan Kumar Himachal Pradesh Krishi Vishvavidyalaya, Palampur (E1), conventional farming under polyhouse conditions at Palampur (E2), natural farming under polyhouse conditions at Palampur (E3), field conditions at Bajaura (E4), Berthin (E5), Mandi (E6) and Dhaulakuan (E7). The characteristics of the different environments and soil properties are presented in Table 2 that clearly depicts the varied agro-climatic conditions of different locations, situated at an elevation of 468 m to 1290 m above mean sea level with humid temperate to sub-tropical climatic conditions. The soil texture varied from clay to sandy clay loam while soil reaction varied from slightly acidic (5.6) to neutral (7.1) with medium organic carbon, N, P and K contents in the soil of the respective environments. The weather data across the environments has been presented in Fig. 1 which clearly revealed diversity in temperature, rainfall, and humidity in the respective environments. E2 and E3 represent cultivation of chilli under naturally ventilated polyhouse conditions and hence the crop had not received direct rainfall.
Maximum temperature, minimum temperature, relative humidity and rainfall of all environments during growing season at different locations.
The experimental material was sown in the nursery bed of size 3 m × 1 m × 15 cm and seedlings were ready for transplanting in about eight weeks after seed sowing in the respective environments. The seedlings of the respective hybrids and check genotypes were laid out in Randomized Complete Block Design with three replications at each location by transplanting as detailed in Table 3. Each genotype was planted in 1.8 m × 1.8 m plots consisting of 16 plants in each replication with inter and intra row spacing of 45 cm × 45 cm, respectively.
Cultural practices
A three-disc tractor was used to plough the field under field conditions followed by a tractor-driven rotavator while soil in the polyhouse was prepared using power tiller under both conventional and natural farming systems. The appropriate amount of well-rotten farmyard manure @ 20 tonnes per hectare was mixed into the soil during field preparation. The fertilizers were administered at 75: 60: 60 kg N, P2O5 and K2O per hectare, with half of the required N, full P2O5 and K2O applied at transplanting time and the remaining N applied in two equal splits at one-month intervals. Under field conditions, irrigation was used one week before planting and shortly after transplanting to ensure optimal plant establishment in the soil and then every 7–10 days intervals until the monsoon arrived. On the other hand, drip irrigation system was used to apply water under polyhouse conditions at every alternate day. To keep the fields weed-free, three manual weedings were performed at 3–4 weeks intervals under field conditions.
Natural farming practice includes the application of Ghanjeevamrit (made from Indian cow dung, cow urine, gram flour, soil, and Jaggery in 10:5:1:1:1) @ 250 kg/ha by incorporating in the soil at the time of transplanting and subsequently during the flower initiation stage. Moreover, a 10% Jeevaamrit solution (dissolving 1 kg Ghanjeevamrit in 10 L of water) was sprayed after three weeks regularly until the last harvest.
Data recording and statistical analysis
The observations were recorded on five competitive plants selected randomly from each entry over the replications for nine traits viz., days to flowering, days to first harvest, primary branches per plant, plant height (cm), fruit length (cm), fruit diameter (cm), green fruits per plant (number), green fruit yield per plant (g), and harvest duration (days).
The pooled analysis of variance was undertaken after examining the homogeneity of experimental errors following Bartlett’s test using SAS v.9.1 software.The analysis of the stability performance of genotypes under different environments was done as per the model suggested by Eberhart and Russell18following OPSTAT by PAU Ludhiana and DOSBox 0.74-3 by TNAU. The AMMI and GGE biplots were constructed using the basic model of Gauch28 and Yan and Kang29 using Eigen vector distance matrix through software R-Package version 4.3.222,30.
Results
Analysis of variance and mean performance
The analysis of variance (Supplementary Table S1) for individual environments revealed that the mean sum of squares due to genotypes were significant for majority of the traits across environments except days to flowering at Bajaura (E4) indicating differences in the performance of different hybrids for yield and yield attributes irrespective of the impact of environments. The mean performance of different hybrids (Supplementary Table S2 to S10) showed a considerable variation for yield and yield related traits that construed enormous scope for improvement of chilli. Hybrid DPCHYB 10 consistently performed as top ranked for fruit yield in most of the environments except E1 with average fruit yield of 627.68 g per plant across environments with an advantage of 66.11% over the best yielding check variety Him Palam Mirch 2 (377.86 g). DPCHYB 5 recorded maximum fruit yield in E1 with mean fruit yield of 583.50 g over environments. Both these hybrids significantly superseded remaining hybrids and check varieties/hybrid (S-A8). The other hybrids viz., DPCHYB 12, DPCHYB 8 and DPCHYB 6 got ranking among top five hybrids for green fruit yield. The correlation between weather parameters and fruit yield revealed a positive linear relationship between mean yield and average maximum/minimum temperature (Table 4) while it was negative with relative humidity indicating the adverse effect of humidity on fruit yield.
Join regression analysis
The results of joint regression analysis proposed by Eberhart and Russell18 revealed that the mean sum of squares due to genotypes and environments were significant for most of the traits exceptnumber of primary branches per plant (Table 5). The mean sum of squares due to GEI were significant for all the traits. Combined environment and genotype × environment interaction [E + (G × E)] variance when tested against pooled error and pooled deviation construed the significance for majority of the traits except number of primary branches. G × E (linear) found to be significant for all the traits when tested against pooled error mean sum of squares directing to proceed for stability analysis.
Individual regression analysis and estimation of stability parameters of individual genotypes
The genotypes viz., DPCHYB 2, DPCHYB 4, and DPCHYB 5 were stable both for early flowering and first fruit picking across the environments along with DPCHYB 7 and DPCHYB 10 for flowering and DPCHYB 1 and DPCHYB 6 for first picking with average response (bi=1) and non-significant deviation from regression (S2di = 0) [Table 6]. In contrary, DPCHYB 6 for early flowering and DPCHYB 11 for first harvest had regression coefficient value more than one (1.10) indicating better response under favourable conditions while DPCHYB 12 for both these traits and DPCHYB 1 for flowering revealed regression coefficient less than one indicating their better response under unfavourable environments. The hybrids DPCHYB 8 and DPCHYB 9 for fruit length (Table 6) and that of DPCHYB 1, DPCHYB 2, DPCHYB 5, DPCHYB 6, DPCHYB 11 and DPCHYB 12 for harvest duration (Table 7) demonstrated stable and predictable performance (S2di = 0 and bi= 1). In contrast, hybrids namely, DPCHYB 1, DPCHYB 2, DPCHYB 3, DPCHYB 5, DPCHYB 6, DPCHYB 10 and DPCHYB 11 for fruit length, DPCHYB 8 for fruit diameter (Table 6), DPCHYB 1, DPCHYB 3, DPCHYB 4, DPCHYB 12, Him Palam Mirch 1 and Him Palam Mirch 2 for primary branches, DPCHYB 3, DPCHYB 5, DPCHYB 9 and DPCHYB 10 for plant height, and DPCHYB 7 and DPCHYB 8 for harvest duration (Table 7) had significant deviation from regression pointing towards their unpredictable performance. Besides, DPCHYB 4, DPCHYB 7 and DPCHYB 11 for plant height and DPCHYB 10 for fruit diameter revealed better response under favourable conditions (bi > 1) whereas, DPCHYB 1 and DPCHYB 8 for plant height, and DPCHYB 9 for harvest duration had better response under unfavourable environments (bi < 1).
All hybrids exhibited significant deviations from the regression (S2di = 0) for green fruits per plant and green fruit yield indicating unpredictable performance (Table 7). However, based on regression coefficient (bi), hybrids namely, DPCHYB 2, DPCHYB 5 and DPCHYB 8 for both fruit number and fruit yield, DPCHYB 4, DPCHYB 6 and DPCHYB 8 for fruits per plant and DPCHYB 1, DPCHYB 10 and DPCHYB 12 for green fruit yield showed average response (bi=1) indicating stable performance. Further, DPCHYB 12 for green fruits per plant and DPCHYB 6 for green fruit yield with bi>1 indicating their suitability under favourable environmental conditions. On the other hand, four genotypes viz., DPCHYB 8, DPCHYB 10, DPCHYB 11 and Surajmukhi were suitable for all the environments with average responsiveness (bi = 1) and non-significant deviation from regression (S2di = 0).
AMMI biplot
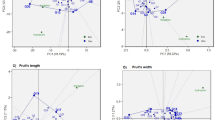
The AMMI analysis revealed highly significant effects of environment, genotype, and genotype × environment interaction (GEI) for all the traits (Table 8). Among the sources of variation, environment effects were markedly higher than that of genotype and interaction for days to flowering, days to first harvest, plant height, fruits per plant, fruit yield per plant and harvest duartion. The first two principal components (PC1 and PC2) explained major portion of the total GEI variation for all the traits. The environment and genotype effects were illustrated simultaneously by generating AMMI1 (IPCA1 vs. Mean) and AMMI2 (IPCA2 vs. IPCA1) biplots. The AMMI1 biplot revealed that genotypes namely, DPCHYB 88, DPCHYB 22, and DPCHYB 44 for fruits per plant (Fig. 2A), and DPCHYB 22 for fruit yield per plant (Fig. 2B), were the most resilient and highly productive across environments. In contrast, genotypes DPCHYB 99 and DPCHYB 1111 for fruits per plant and Him Palam Mirch 214, Surajmukhi15, and Him Palam Mirch 113 for yield per plant, though stable, exhibited low performance and are thus not recommended for selection. Genotypes DPCHYB 1212, DPCHYB 66 and DPCHYB 77 for fruits per plant and DPCHYB 1010, DPCHYB 55, DPCHYB 1212 and DPCHYB 88 for fruit yield showed high productivity but were positioned far from the origin and were unstable. DPCHYB 11 and Him Palam Mirch-113 for fruits per plant were located close to the IPCA1 origin, suggesting general stability across environments. Environments E1 and E6 for fruits per plant, and E1 and E4 for yield per plant showed high additive genotypic main effects, indicating their suitability for discriminating among genotypes.
Additive main effects and multiplicative interaction (AMMI) biplots illustrating G × E interactions of the 16 chilli genotypes in seven locations: (A) AMMI 1 biplot for fruits per plant (FPP) (B) AMMI 1 biplot for fruit yield per plant (YPP) (C) AMMI 2 biplot for fruits per plant (FPP) (D) AMMI 2 biplot for fruit yield per plant (YPP); where, green lines represented with 1–7 depicts environments at Palampur1, Conventional farming under poly house at Palampur2 and Natural farming under polyhouse at Palampur3, Bajaura4, Berthin5, Sundernagar6 and Dhaulakuan7; *Representation of genotypes as 1–16 in blue colour dotted lines where 1: DPCHYB 1, 2: DPCHYB 2, 3: DPCHYB 3, 4: DPCHYB 4, 5: DPCHYB 5, 6: DPCHYB 6, 7: DPCHYB 7, 8: DPCHYB 8, 9: DPCHYB 9, 10: DPCHYB 10, 11: DPCHYB 11, 12: DPCHYB − 12, 13: Him Palam Mirch 1, 14: Him Palam Mirch 2, 15: Surajmukhi, 16:CH-27.
The AMMI2 biplot for fruits per plant demonstrated that environments E6, E4, E3, E5, and E7 were more stable while E1 and E2 had greater interaction effects (unstable environments) (Fig. 2C). Notable genotype-environment associations included DPCHYB 22 and CH 2716 in E5, E6 and E7, DPCHYB 33 in E4, Surajmukhi15 in E2, DPCHYB 1212 in E3, and DPCHYB 77 in E1were observed. Similarly, AMMI2 biplot for yield per plot indicated that E1 and E6 are located farther away from the biplot origin than environments E3, E4, E5, E2, and E7. DPCHYB 33 and DPCHYB 1010 in E4 and E2, CH 2716 in E3 and E6, DPCHYB 11 and DPCHYB 77 in E1, and DPCHYB 1212 in E5 were found to be the highest yielding genotypes in the respective environments (Fig. 2D). These interactions aid to identify genotypes for targeted environments or recommend those with broader adaptability.
GGE biplot
A short vector indicates that the test environment is unrelated to other environments. The environment vector view of the GGE biplot explained 78.45, 76.57, 75.33, 87.60, 95.09, 95.06, 78.99, 78.63 and 92.67% of the overall GEI variance for days to flowering (Fig. 3A), days to first harvest (Fig. 3B), number of primary branches (Fig. 3C), plant height (Fig. 3D), fruit length (Fig. 3E), fruit diameter (Fig. 3F), green fruits per plant (Fig. 3G), harvest duration (Fig. 3H) and green fruit yield per plant (Fig. 4D), respectively. The seven environments were divided into two groups for fruit yield with E1, E5 and E3 in first group while E4, E6, E7 and E2 in the other (Fig. 4A). All the environments formed an acute angle indicating positive correlation wherein E4 and E6 were highly correlated with very close environment vector.
Which won where G + GE biplots for (A) Days to flowering (B) Days to first harvest (C) Primary braches per plant (D) Plant height (E) Fruit length (F) Fruit diameter (G) Fruits per plant (H) Harvest duration. Where, “E” represents the environments Palampur (E1), Conventional farming under polyhouse at Palampur (E2) and Natural farming under polyhouse at Palampur (E3), Bajaura(E4), Berthin(E5), Sundernagar, Mandi (E6) and Dhaulakuan (E7) *Representation of genotypes as 1–16 where 1: DPCHYB 1, 2: DPCHYB 2, 3: DPCHYB 3, 4: DPCHYB 4, 5: DPCHYB 5,6: DPCHYB 6, 7: DPCHYB 7, 8: DPCHYB 8, 9: DPCHYB 9, 10: DPCHYB 10, 11: DPCHYB 11, 12: DPCHYB − 12, 13: Him Palam Mirch 1, 14: Him Palam Mirch 2, 15: Surajmukhi, 16:CH-27.
G + GE biplots for yield per plant (A) An environment vector view biplot ; (B) Discrimitiveness vs. representativeness biplot ; (C) Means vs. Stability biplot . The single arrowed horizontal line represents the AEC abscissa and the direction of AEC abscissa arrowhead indicates increasing yield. The vertical projections on the AEC abscissa indicates stability (greater the projection, higher is the instability); (D) “Which won where” biplot . Where, “E” represents the environments i.e. Palampur (E1), Conventional farming under polyhouse at Palampur (E2) and Natural farming under polyhouse at Palampur (E3), Bajaura (E4), Berthin (E5), Sundernagar (E6) and Dhaulakuan (E7). *Representation of genotypes as 1–16 where 1: DPCHYB 1, 2: DPCHYB 2, 3: DPCHYB 3, 4: DPCHYB 4, 5: DPCHYB 5, 6: DPCHYB 6, 7: DPCHYB 7, 8: DPCHYB 8, 9: DPCHYB 9, 10: DPCHYB 10, 11: DPCHYB 11, 12: DPCHYB − 12, 13: Him Palam Mirch 1, 14: Him Palam Mirch 2, 15: Surajmukhi, 16:CH-27.
The discriminativeness and representativeness analysis (Fig. 4B) revealed the longest vectors for E1, E4, E5 and E2 for green fruit yield, E1, E3and E6 for days to flowering, E3 for days to first harvest, E5 for fruit length, E2 for fruit diameter, E2 and E3 for plant height and green fruits per plant, and E6 for harvest duration suggesting these environments as the most informative and discriminative which would help effectively in differentiating between genotypes. The environments E5 for green fruit yield and fruit length, E1 for days to flowering, E2 for fruit diameter, plant height and green fruits per plant and E7 for harvest duration were characterised by a longer vector that creates a smaller angle with the AEC abscissa line is considered optimal for the identification and selecting superior genotypes.
The mean vs. stability biplot analysis (Fig. 4C) construed the desirable performance of DPCHYB 10 and DPCHYB 5 for green fruit yield and fruit length, DPCHYB 5 and DPCHYB 4 for early flowering and tall plants, DPCHYB 2 and DPCHYB 5 for early harvesting, DPCHYB 10 for broad fruits, DPCHYB 12 and DPCHYB 5 for more green fruits per plant, and DPCHYB 5 and DPCHYB 6 with longer harvest duration. The genotypes which were located close to the AEC showed better stability e.g., DPCHYB 2 for early flowering, number of fruits per plant and green fruit yield per plant, DPCHYB 5 for early flowering, fruit length, and harvest duration and that of DPCHYB 10 for fruit length and number of primary branches.
'Which won where’biplot divided the environments into two sectors and that of genotypes into five sectors (Fig. 4D). Based on the position of hybrids on the polygon’s vertices, DPCHYB 10 was the winning genotype based on its better performance for fruit yield across the environments except E1 where DPCHYB 5 was the most responsive. In addition, DPCHYB 8 and DPCHYB 6 also showed average stability across the environments while DPCHYB 12 and DPCHYB 2 showed the similar response in E1. Further, GGE biplot for flowering (Fig. 3A) revealed that E1, E2 and E3 held in the sector with Surajmukhi15 as the most responsive genotype for these environments. Similarly, CH-2716 for flowering in E4, E5, E6 and E7 (Fig. 3A), CH-2716 and DPCHYB 1010 for early harvest in E6, E7, E5 and E4 (Fig. 3B), DPCHYB 1010 in E6, DPCHYB 1212 in E1, E7, E5 and E4 and Him Palam Mirch 113 in E2 and E3 for primary branches (Fig. 3C), DPCHYB 44 in E1, E2, E3 and E5 for plant height (Fig. 3D), DPCHYB 1010 for fruit length (Fig. 3E) and fruit diameter (Fig. 3F) in all environments, and DPCHYB 1212 across environments except E4 for fruits per plant (Fig. 3G) were clear winner and stable hybrids for the respective traits. Similarly, DPCHYB 55 in E2 and E7, DPCHYB 22 in E4 and E5 and DPCHYB 66 in E6, E1 and E3 for harvest duration (Fig. 3H) were the most responsive and winning genotypes.
Discussion
The chilli breeders have to plan their heterosis breeding programme as per the aspirations of the stakeholders for hybrids with medium to long fruits, high yield and wider adaptability. Keeping this in view, male sterility derived hybrids were synthesized involving diverse inbred lines. It is essential to evaluate these hybrids for stability and wider adaptability for which, multi-location assessment for yield and associated characteristics is important to release them for cultivation in the state or different zones with variable climatic conditions31. The climatic conditions in the north-western Himalayas range from humid subtropical to dry temperate and the development of hybrids with consistent yield performance across environments is necessary and is of utmost significance. G×E interaction is a serious challenge before breeders while evaluating genotypes through multiple environment trials32. To handle this issue, multiple statistical models viz., Eberhart-Russell (E-R) joint regression model, AMMI and GGE etc. support plant breeders to assess the performance of genotypes across variable environmental conditions that allow to identify the best genotypes for a specific or range of environments29,33. E-R model is broadly used to make decisions on advancement of genotypes, and the present study was aimed to compare the E-R with AMMI and GGE biplot analysis.
E-R joint regression model, AMMI and GGE analyses revealed substantial G×E effects for fruit yield and its attributes based on evaluation of hybrids across environments indicating the involvement of diverse inbred lines as parents along with variation among test environments. The earlier researchers have also found strong genotypic and G×E impact on fruit yield and associated characters34. Hence, it is necessary to use stability parameters to identify wider adaptive genotypes. Further, the significance of combined environment and G×E interaction [E(G×E)] components of variance (Table 5) demonstrated the impact of environments and their interactions with genotypes on the performance of hybrids. The linear and non-linear components of environments had a significant influence on majority of the traits. Tembhurne and Rao35, Datta and Jana36, Raghavendra et al.37, Barchenger et al.38 and Ozukum and Seyie39 have also observed the significance of linear component of GEI on different traits of chilli. A significant GEI affect the relative ranking/performance of genotypes across environment and hampers breeding progress during selection and advancing genotypes to successive selection steps40.
The regression coefficient variance for all the characters suggested heterogeneous response of genotypes to environmental fluctuations41. The commercially available hybrids should have improved mean performance vis-a-vis stability across environments15,29. However, variable response of different genotypes was recorded for mean fruit yield and yield attributes besides stability (Table 5). Eberhart and Russell18 define a stable genotype that has high mean performance, regression coefficient (bi) close to unity, and the deviation from regression (S2di) near to zero. High mean performance along with wider stability (bi = 1 and S2di = 0) of DPCHYB 10 for fruit yield and yield attributes in comparison to check Him Palam Mirch-2 categorized it as the most suitable hybrids across environments. Similarly, DPCHYB 2 and DPCHYB 5 also showed promise based on their superiority and stability with unit regression coefficient and non-significant deviation from regression (S2di = 0) for fruit yield, early flowering, number of fruits and harvest duration than check variety. Therefore, these hybrids can be considered as the most desirable with high yield potential and were insensitive to the variable environmental conditions. The earlier research findings using Eberhart and Russel model have also successfully identified chilli hybrids using variable inbred lines with wider adaptability and superior yield performance23,25,27,42,43. Eberhart and Russell method identified the most stable hybrids but could not reliably detect the high yielding hybrids.
The AMMI biplot is a valuable tool for dissecting genotype-by-environment interaction (GEI) along with identifying stable and high-performing genotypes across varying environments28. The magnitude of mean sum of squares due to environment was higher than genotype indicating strong environmental influence on different traits (Table 8). The first two principal components (PC1 and PC2) explained more than 70% of the total GEI variation for most of the traits that clearly demonstrated the suitability of the AMMI model in capturing the crossover interaction and identifying stable genotypes for specific and wider adaptation. Genotype-environment interactions are visually represented by the distance from the biplot origin that help to predict adaptability20. A short vector length and proximity to the origin in the biplot signify environmental stability and are considered ideal conditions for selecting consistent genotypes19. Accordingly, AMMI1 biplot revealed that genotypes DPCHYB 8, DPCHYB 2, and DPCHYB 4 for fruits per plant (Fig. 2A), and DPCHYB 2 for yield (Fig. 2B) were the most resilient across environments. The AMMI2 biplot further enhanced this interpretation by partitioning mega-environments and identifying genotypes with specific adaptations such as nutrient availability and climate influence on genotype performance27. In chilli, agronomic traits like fruit number, fruit length, and fruit diameter had significant impact on fruit yield and showed sensitivity to environmental fluctuations, emphasizing the relevance of GEI analysis26. Environments and genotypes close to the biplot origin were associated with low interaction effects and higher stability28, thus validating the AMMI2 model’s suitability for identifying genotypes with broad or specific adaptation across diverse environments e.g. DPCHYB 3 and DPCHYB 10 in E4 and E2, DPCHYB 1 and DPCHYB 7 in E1, and DPCHYB 12 in E5 for green fruit yield per plant (Fig. 2D).
GGE biplot has an edge over AMMI biplot on account of better graphical representation based on mega environment analysis and classify genotypes to specific environment or as wider adaptive genotype by describing G + GE. Further, the discriminating vs. representativeness of the GGE biplot is effective to evaluate test environments, which is otherwise not possible in AMMI analysis17,44. Accordingly, the most promising and highly stable genotypes may be recommended for large scale commercial cultivation in target environments45. These aspects favour GGE biplot to be very popular as a comprehensive tool in quantitative genetics and plant breeding. The environment-vector aspect of the GGE biplot (Fig. 4A) indicated positive correlation among majority of test environments by forming acute angle for fruit yield, fruit length, fruit diameter, plant height, green fruits per plant and harvest duration suggesting the suitability of expression of these traits across the environments15,46,47,48. Further, the ‘Discriminative vs. representative’ view of GGE biplots (Fig. 4B) described that the environment E5 for green fruit yield and fruit length, E1 for days to flowering, E2 for fruit diameter, plant height and green fruits per plant and E7 for harvest duration had a long vector and a small angle with the AEC abscissa that suggest these environments as the most suitable for the identification of superior genotypes according to the traits of interest. Further, the mean vs. stability GGE biplot (Fig. 4C) identified hybrids DPCHYB 22 as the most stable hybrid for fruit yield based on their closeness to the origin and short vector as depicted by a circle with an arrow on the AEC. Anilkumar et al.49 and Sahmat et al.26 have also observed differences in stability performance of genotypes for number of green fruits per plant in different environments.
‘Which won where’ is the most attractive feature of GGE biplot as it displays the winning genotypes along with interaction patterns between genotypes and environments in the polygon view of the GGE biplot50 in MET data analysis (Fig. 4D). This provides an opportunity to evaluate and understand the likely presence of various mega-environments in the manifestation of traits of interest and adaptation of genotypes under certain mega-environments15,51. The genotypes with superior performance and better adaptability would be the ones which are present at the vertices of the polygon along with environments aspect. Keeping this in view, DPCHYB 10 showed superiority for fruit yield in majority of the environments indicating better adaptability with stable performance except E1. DPCHYB 5 had the similar response only in E1. In contrary, DPCHYB 3, DPCHYB 7 and Surajmukhi are present at the vertices but do not represent any environment and hence, were poor performer across environments26. This pattern of genotype and environment interactions along with presence of different mega environment cannot be predicted by Eberhart and Russell model18. “Which won where” seems to be ideal for ‘exploiting’ the GE interactions, though recommendation of genotypes to specific locations must be made with caution as certain times, the variation explained by the PC1 and PC2 of the GGE model might be too low. Also, this two-dimensional model explains only two principal components at a time and there is probability that some of the projections might be placed outward in a third dimension13,52.
Conclusions
Based on the present investigation, it can be concluded that DPCHYB 10 was identified as the most stable hybrid across environments as predicted by both Eberhart and Russel model and ‘Which won where’ of GGE biplot analysis. AMMI depicted DPCHYB 2 as the most stable hybrid across environments which was also described by mean vs. stability biplot of GGE and E-R. Similarly, DPCHYB 5 revealed stability across environments based on E-R with the same expression in E1 based on GGE. DPCHYB 10 (627.68 g/plant) and DPCHYB 5 (583.50 g/plant) expressed yield advantage of 66.11 and 54.42% over the best check variety Him Palam Mirch 2 (377.86 g/plant). E3 and E7 identified as the most representative environments that provided supportive testing conditions for identifying hybrids with high stability for green fruit yield per plant. The hybrids DPCHYB 10 and DPCHYB 5 could be recommended to the stakeholders for cultivation after thorough evaluation by conducting onfarm trials at farmer’s field.
Data availability
The datasets generated and/or analyzed during the current study are presented in the main manuscript and as additional supporting files. Further, any additional information can be obtained from the corresponding author on sensible request.
References
Lata, H. & Sharma, A. Exploitation of heterosis in Chilli using genetic male sterile lines for red fruit yield with special reference to high rainfall conditions. Sci. Rep. 14, 23061. https://doi.org/10.1038/s41598-024-73728-7 (2024).
Kim, S. et al. Genome sequence of the hot pepper provides insights into the evolution of pungency in capsicum species. Nat. Genet. 46 (3), 270–278. https://doi.org/10.1038/ng.2877 (2014).
Lata, H. et al. Heterosis and combining ability vis-à-vis association for green fruit yield and component traits involving male sterile lines in Chilli (Capsicum annuum) under wet temperate zone of North Western Himalayas. Plant. Breed. 142, 547–562. https://doi.org/10.1111/pbr.13114 (2023).
Olatunji, T. L. & Afolayan, A. J. The suitability of Chili pepper (Capsicum annuum L.) for alleviating human micronutrient dietary deficiencies: A review. Food Sci. Nutr. 6 (8), 2239–2251. https://doi.org/10.1002/fsn3.790 (2018).
Manyam, P. Molecular characterization of virus associated with chilli (Capsicum annuum L.). M.Sc. Thesis. Department of Plant Pathology, College of Agriculture, University of Agricultural Sciences, Dharwad, p. 137 (2011).
FAO. World Food and Agriculture (2022). https://www.fao.org/faostat/en/#data/QCL/visualize -23).
Area And Production of Vegetables in India (National Horticulture Board, 2023).
Verma, S. K., Rai, D. P. & Verma, L. R. Knowledge and adoption of recommended Chilli production technology by the farmers of Raipur district. Agric. Update. 10, 120–125 (2015).
Export import data. (2024). https://www.exportimportdata.in/blogs/chilli-export-from-india.aspx
Wani, K. P. N. et al. Hussain. Comparative performance of various Chilli genotypes under temperate conditions of Kashmir. SKUAST J. Res. 15, 117–122 (2013).
Dhall, R. K. & Cheema, D. S. Use of male sterility in hybrid seed production of Chilli (Capsicum annuum L.): a review. J. Res. Punjab agric. Univ .46–52 (2010).
Danyali, S. F., Razavi, F., Ebadi-Segherloo, A., Dehghani, H. & Sabaghpour, S. H. Yield stability in Chickpea (Cicer arietinum L.) and study relationship among the univariate and multivariate stability parameters. Res. Plant. Biol. 2, 46–61 (2012).
Asilo, M. A. I. et al. Stability and G × E analysis of zinc-biofortified rice genotypes evaluated in diverse environments. Euphytica 215, 261. https://doi.org/10.1007/s10681-019-2384-7 (2019).
Anniciarico, P. Genotype × environment interaction: Challenges and opportunities for plant breeding and cultivar recommendation. Food and Agriculture Organization of the United Nations, Rome, 1–2 (2002).
Yan, W. & Tinker, N. A. Biplot analysis of multi-environment trial data: principles and applications. Can. J. Plant. Sci. 86, 623–645. https://doi.org/10.4141/P05-169 (2006).
Ahmed, M. S. et al. Country-wide, multi-location trials of green super rice lines for yield performance and stability analysis using genetic and stability parameters. Sci. Rep. 14, 9416. https://doi.org/10.1038/s41598-024-55510-x (2009).
Zewdie, Y. & Bosland, P. W. Evaluation of genotype, environment, and genotype-by-environment interaction for capsaicinoids in Capsicum annuum L. Euphytica 111, 185–190. https://doi.org/10.1023/A:1003837314929 (2000).
Eberhart, S. A. & Russel, W. A. Stability parameters for comparing varieties. Crop Sci. 6, 36–40. https://doi.org/10.2135/cropsci1966.0011183X000600010011x (1966).
Firew, A. M., Amsalu, B. & Tsegaye, D. Additive main effects and multiplicative interaction (AMMI) and genotype main effect and genotype by environment interaction (GGE) biplot analysis of large white bean (Phaseolus vulgaris L.) genotypes across environments in Ethiopia. Afr. J. Agric. Res. 14 (35), 2135–2145. https://doi.org/10.5897/AJAR2019.14180 (2019).
Abid, S. & Jahanzaib, M. Identification of promising groundnut genotypes using stability analysis. Pakistan J. Agricultural Res. 37 (3), 314–319. https://doi.org/10.17582/journal.pjar/2024/37.3.314.319 (2024).
Choudhary, M. et al. V. GGE biplot analysis of genotype× environment interaction and identification of mega-environment for baby corn hybrids evaluation in India. Indian J. Genet. Plant. Breed. 79 (04), 658–669. https://doi.org/10.31742/IJGPB.79.4.3 (2019).
Yan, W., Kang, M. S., Ma, B., Sheila, W. & Cornelius, P. L. GGE biplot vs AMMI analysis of genotype-by-environment data. Crop Sci. 47, 643–653. https://doi.org/10.2135/cropsci2006.06.0374 (2007).
Sran, T. S., Jindal, S. K. & Chawla, N. Genotype by environment interaction for quality traits in Chilli pepper (Capsicum annuum L). Genetika 53 (1), 23–49. https://doi.org/10.2298/GENSR2101023S (2021).
Baruah, J., Begum, T. & Lal, M. Estimation of environment stability for fruit yield and capsaicin content by using two models in Capsicum Chinense jacq. (Ghost Pepper) with multi-year evaluation. Peer J. 12, 17511. https://doi.org/10.7717/peerj.17511 (2024).
Degefa, G., Benti, G., Wakgari, G., Jafar, M. & Tadesse, F. Genotype by environment interaction and stability analysis for yield of Chili pepper (Capsicum annuum L.) in East hararghe, Ethiopia. Int. J. Agric. Sci. Technol. 4, 51–57. https://doi.org/10.22004/ag.econ.344721 (2024).
Sahmat, S. S. et al. Unravelling the dynamics of genotype and environment interactions on Chilli (Capsicum annuum L.) yield-related attributes in soilless planting systems. Sci. Rep. 14 (1), 1698. https://doi.org/10.1038/s41598-023-50381-0 (2024).
Subhavyuktha, S. et al. Employing empirical models to analyze stability of yield and quality traits in Chili peppers (Capsicum species). Crop Sci. 64 (6), 2977–2997. https://doi.org/10.1002/csc2.21390 (2024).
Gauch, H. G. A simple protocol for AMMI analysis of yield trials. Crop Sci. 53, 1860–1869. https://doi.org/10.2135/cropsci2013.04.0241 (2013).
Yan, W. & Kang, M. S. GGE Biplot Analysis: A Graphical Tool for Breeders, Geneticists, and Agronomists (CRC, 2003). https://doi.org/10.1201/9781420040371
R Development Core Team R. A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2022).
Rana, C. et al. Stability analysis of garden pea (Pisum sativum L.) genotypes under North Western Himalayas using joint regression analysis and GGE biplots. Genet. Resour. Crop Evol. 68, 999–1010. https://doi.org/10.1007/s10722-020-01040-0 (2021).
Hajjarpoor, A., Nelson, W. C. & Vadez, V. How process-based modelling can help plant breeding deal with G×E×M interactions. Field Crop Res. 283, 108554. https://doi.org/10.1016/j.fcr.2022.108554 (2022).
Cooper, M. et al. Predicting genotype× environment× management (G× E× M) interactions for the design of crop improvement strategies: integrating breeder, agronomist, and farmer perspectives. Plant. Breed. Rev. 46, 467–485. https://doi.org/10.1002/9781119874157.ch8 (2022).
Cabral, N. S. S. et al. Genotype × environment interaction on experimental hybrids of Chilli pepper. Genet. Mol. Res. 16, 1–9 (2017).
Tembhurne, B. V. & Rao, S. K. Stability analysis in Chilli (Capsicum annuum L). J. Spices Aromat. Crops. 22, 154–164 (2013).
Datta, S. & Jana, J. C. Stability of Chilli genotypes for different seasons at Terai zone of West Bengal. J. Spices Aromat. Crops. 21, 71–75 (2012).
Raghavendra, H., Puttaraju, T. B., Varsha, D. & Jodage, K. Stability analysis of different Chilli hybrids (Capsicum annuum L.) for their yield and yield attributing traits. J. Appl. Hortic. 19, 218–221 (2017).
Barchenger, D. W. et al. Stability of yield and yield components of pepper (Capsicum annuum), and evaluation of publicly available predictive meteorological data in East and Southeast Asia. HortScience 53, 1776–1783. https://doi.org/10.21273/HORTSCI13581-18 (2018).
Ozukum, C. & Seyie, K. Stability analysis of Naga King Chilli (Capsicum Chinense Jacq). Indian J. Hill Farming (2019). https://epubs.icar.org.in/index.php/IJHF/article/view/94727
Pham, H. N. & Kang, M. S. Interrelationship among and repeatability of several stability statistics estimated from international maize trials. Crop Sci. 28, 925–928. https://doi.org/10.2135/cropsci1988.0011183X002800060010x (1998).
Ortiz, R., Crossa, J., Vargas, M. & Izquierdo, J. Studying the effect of environmental variables on the genotype × environment interaction of tomato. Euphytica 153, 119–134. https://doi.org/10.1007/s10681-006-9248-7 (2007).
Basak, J. Identification of stable high-yielding hybrids in Chilli (Capsicum annuum L). J. Food Legumes. 32, 139–144 (2019).
Singh, T. N., Joshi, A. K., Vikram, A., Yadav, N. & Prashar, S. Mean performances, character associations and multi-environmental evaluation of Chilli landraces in North Western Himalayas. Sci. Rep. 14 (1), 769. https://doi.org/10.1038/s41598-024-51348-5 (2024).
Roostaei, M., Mohammadi, R. & Amri, A. Rank correlation among different statistical models in ranking of winter wheat genotypes. Crop J. 2 (2–3), 154–163. https://doi.org/10.1016/j.cj.2014.02.002 (2014).
Anputhas, M., Samita, S. & Abeysiriwardena, D. S. D. Z. Stability and adaptability analysis of rice cultivars using environment-centred yield in two-way ANOVA model. Commun. Biom Crop Sci. 6, 80–86 (2011).
Yan, W. & Tinker, N. A. An integrated biplot analysis system for displaying, interpreting, and exploring genotype × environment interaction. Crop Sci. 45, 1004–1016. https://doi.org/10.2135/cropsci2004.0076 (2005).
Farshadfar, E., Mohammadi, R., Aghaee, M. & Vaisi, Z. GGE biplot analysis of genotype × environment interaction in wheat-barley disomic addition lines. Aust J. Crop Sci. 6, 1074–1079 (2012).
Parihar, A. K. et al. Targeting test environments and rust-resistant genotypes in lentils (Lens culinaris) by using heritability-adjusted biplot analysis. Crop Pasture Sci. 69, 1113–1125. https://doi.org/10.1071/CP18259 (2018).
Anilkumar, C. et al. Yield stability of Chilli (Capsicum annuum L.) hybrids differing for fruiting habit traits. Int. J. Curr. Microbiol. Appl. Sci. 7, 1664–1674. https://doi.org/10.20546/ijcmas.2018.709.201 (2018).
Erdemci, I. Investigation of genotype × environment interaction in Chickpea genotypes using AMMI and GGE biplot analysis. Turkish J. Field Crops. 23, 20–26. https://doi.org/10.17557/tjfc.414846 (2018).
Alwala, S., Kwolek, T., McPherson, M., Pellow, J. & Meyer, D. A comprehensive comparison between Eberhart and Russell joint regression and GGE biplot analyses to identify stable and high yielding maize hybrids. Field Crop Res. 119, 225–230. https://doi.org/10.1016/j.fcr.2010.07.010 (2010).
Akter, A. et al. R. GGE biplot analysis for yield stability in multi-environment trials of promising hybrid Oryza sativa L. Bangladesh Rice J. 19, 18 (2015).
Acknowledgements
The authors are greatful to National Agricultural Higher Education Project (NAHEP) Centre for Advanced Agricultural Science and Technology (CAAST), New Delhi for support during the study.
Author information
Authors and Affiliations
Contributions
V.S. conducted the experiment and analyzed data and manuscript writing; A.S. conceptualized the research, planning and execution of experiments, data interpretation, writing of manuscript; N.K., N.R., and S. helped in material preparation and data collection.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Singh, V., Sharma, A., Kumar, N. et al. Stability models regulate the adaptation of male sterility-based chilli hybrids for agro-ecologically diverse regions of north-western Himalayas. Sci Rep 15, 34742 (2025). https://doi.org/10.1038/s41598-025-18393-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-18393-0