Abstract
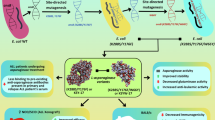
Pantoea piersonii IIIF1SW-P2T (basionym Kalamiella piersonii IIIF1SW-P2T), a bacterial species isolated from the International Space Station, was predicted to be non-pathogenic to humans, unlike its clinical counterpart P. piersonii YU22, which induces struvite crystallization and causes toxicity to Human Embryonic Kidney cell line (HEK 293 T). Here, we identify L-asparaginase-mediated cytotoxicity of IIIF1SW-P2T on HEK 293 T in vitro and demonstrate strain’s colonization ability in the reproductive tract in vivo using Caenorhabditis elegans as a model system. The recombinant L-asparaginase of IIIF1SW-P2T (Kp_AnsA, ~ 37 kDa) generated significant amounts of NH4+ (4.1‒15.5 μM, P < 0.0001) and exerted cytotoxicity to HEK 293 T (29.1‒36.0%, P < 0.0001). NH4+-driven cytotoxicity of HEK 293 T was validated through the introduction of nearly equimolar amounts of standard NH4+ (12.7 μM; 31% cytotoxicity, P < 0.0001). Kp_AnsA found to be a halotolerant enzyme, highly active in alkaline pH (optimum pH 9), and exhibited Km, Vmax and Kcat values of 5.4 mM, 8.4 U/mg and 135.6 μmoles s-1, respectively, in the presence of L-asparagine. Kp_AnsA displayed absolute amino acid sequence identity with that of YU22 and formed a tight phyletic association with that of marine bacteria. Analysis in vivo revealed the rapid and sustained colonization of green fluorescent protein (gfp)-tagged IIIF1SW-P2T in the reproductive tract with 5% mortality of C. elegans. Gfp-tagged IIIF1SW-P2T were localized at the vulvar and luminal regions, embryos and interembryonic space of C. elegans. The role of L-asparaginase in the colonization of IIIF1SW-P2T at the reproductive tract of C. elegans merits further investigation. Nonetheless, this study demonstrated cytotoxicity of bacterial L-asparaginase on HEK 293 T and discovered the potential ability of IIIF1SW-P2T to colonize the reproductive tract of a eukaryote.
Similar content being viewed by others
Introduction
Pantoea piersonii IIIF1SW-P2T (basionym Kalamiella piersonii IIIF1SW-P2T) is a multidrug-resistant bacterial species affiliated with the family Erwiniaceae, isolated and characterized from the Port panel of the Cupola, an observation deck for the crew inhabiting at the International Space Station (ISS)1. The present taxon was predicted to be a non-pathogen to humans based on the PathogenFinder1. However, P. piersonii YU22, a closely related strain that was isolated and described from the urine sample originating from a kidney stone patient in India, was found to stimulate struvite crystallization and exert cytotoxicity to the Human Embryonic Kidney 293 cell line (HEK 293 T) in vitro2. A series of subsequent reports indicated the widespread occurrence of the present taxon, including human saliva3, blood4,5, and tissue6. Kalamiella piersonii IIIF1SW-P2T was reclassified as Pantoea piersonii IIIF1SW-P2T based on phylogenomic analysis7. Detailed molecular studies may shed more light on the complex ecology and help us understand the persistence and prevalence of the present taxon at contrasting niches across the globe and Space.
L-asparaginase is an enzyme involved in the catalytic conversion of L-asparagine into aspartic acid and NH4+. Asparaginase has been isolated and characterized from various bacteria, such as Stenotrophomonas maltophilia EMCC22978, Salmonella paratyphi9, Burkholderia pseudomallei10, as well as from other resources11,12. The enzyme is also marketed as an antineoplastic drug since it constrains amino acid supply to tumour cells and blocks cell proliferation by interrupting asparagine-dependent protein synthesis13. Asparaginase has been tested extensively for its anticancer efficacy in general and acute lymphoblastic leukaemia in particular8,9,10,11,12. Recently, asparagine-driven NH4+ formation was identified and attributed to struvite crystallization by YU222,14. However, the mechanistic insights of cytotoxicity and the preferred niche for colonization remain poorly understood.
The human urine association, asparagine-driven NH4+ formation and direct involvement in struvite crystallization identified in YU22, which shared the highest genomic relatedness (99%) with IIIF1SW-P2T, prompted the present research. We hypothesized L-asparaginase-driven cytotoxicity and the ability of IIIF1SW-P2T to colonize the reproductive tract of a eukaryote and tested the hypothesis using HEK 293 T and Caenorhabditis elegans as in vitro and in vivo models, respectively.
Materials and methods
Bacteria and culture conditions
Pantoea piersonii IIIF1SW-P2T (= DSM 108198 T) was procured from Leibniz-Institut DSMZ, Braunschweig. Cells were grown in nutrient agar (HiMedia, India) at 37 °C for 24‒48 h. Competent cells of Escherichia coli DH5α and E. coli BL21 (DE3) were procured from Yeastern Biotech. Transformed DH5α and BL21 (DE3) were grown on Luria Bertani (LB) agar plates or liquid broth supplemented with ampicillin (50 μg ml−1) and kanamycin (30 μg ml−1) at 37 °C for 24 h. TA cloning vector, pET28a(+) overexpression vector and gfp-bearing pHC60 were procured from Yeastern Biotech, Novagen and NovoPro, respectively, and maintained in DH5α.
Engineering recombinant L-asparaginase overexpression system
Genomic DNA (gDNA) of IIIF1SW-P2T was isolated and purified using a DNA isolation kit (MolBio). The full-length gene (1014 bp) encoding L-asparaginase (Kp_ansA) was amplified from gDNA of IIIF1SW-P2T using NdeI restriction site-tagged forward (Kp_ansA_F: 5’-ATACATATGCAAAAGAAAAATATCTATG-3’) and XhoI restriction site-tagged reverse primer (Kp_ansA_R: 5’-GTGCTCGAGTTAGTCTTCGGTCAGTTCA-3’), respectively. The PCR conditions were as follows: an initial denaturation at 95 °C for 2 min, and 35 cycles of denaturation at 95 °C for 30 s, annealing at 57 °C for 30 s and extension at 72 °C for 1.5 min followed by a final extension at 72 °C for 7 min. The NdeI/XhoI tagged amplicon was cloned to the TA cloning vector (Yeastern Biotech) using the manufacturer’s protocol and the ligated product TA_Kp_ansA was introduced to DH5α through the heat shock method.
The positive transformants bearing TA_Kp_ansA appeared on ampicillin (25 μg/ml) supplemented LB agar were confirmed through colony PCR using M13F (5’-GTTTTCCCAGTCACGAC-3’) and M13R (5’-TTCACACAGGAAACAGCTATGAC-3’) primers. The PCR conditions used for the amplification were as follows: an initial denaturation at 95 °C for 2 min, and 35 cycles of denaturation at 95 °C for 30 s, annealing at 55 °C for 30 s and extension at 72 °C for 1.5 min, followed by a final extension at 72 °C for 7 min. The amplicon was sequenced to verify the orientation and base pair errors using M13 primers. The TA_Kp_ansA construct was isolated from the transformants using the NucleoSpin plasmid extraction kit (Takara) and subjected to restriction double digestion using NdeI and XhoI enzymes (New England Biolabs) as per the manufacturer’s protocol. The double digestion was verified through the agarose gel and the NdeI/XhoI-tagged gene was purified using NucleoSpin gel and PCR clean-up kit (Takara).
Simultaneously, pET28a(+) plasmid maintained at DH5α was extracted using NucleoSpin plasmid extraction kit (Takara) and subjected to NdeI/XhoI double digestion as per the manufacturer’s protocol. Linearized plasmid was extracted from agarose gel using NucleoSpin gel and PCR clean-up kit (Takara). A 1014 bp amplicon was ligated to linearized pET28a(+) using T4 DNA ligase (New England Biolabs) as per the manufacturer’s protocol. The pET28a(+)_Kp_ansA construct was introduced to competent BL21 (DE3) (Yeastern Biotech Co. Ltd) cells through heat shock transformation. Colony PCR was carried out to identify positive transformants using T7F (5’-TAATAGCACTCACTATAGGG-3’) and T7R (5’-GCTAGTTATTGCTCAGCGG-3’) primers. The PCR conditions were as follows: an initial denaturation at 95 °C for 2 min, and 35 cycles of denaturation at 95 °C for 30 s, annealing at 60 °C for 30 s and extension at 72 °C for 1.5 min followed by a final extension at 72 °C for 7 min.
Expression and purification of N-terminal His-tagged L-asaparaginase of IIIF1SW-P2T
E. coli BL21 (DE3) cells transformed with pET28a(+)_Kp_ansA were cultivated in kanamycin (30 μg/ml) supplemented LB broth at 37 °C and 120 rpm till attainment of OD600 0.5. Expression of protein was induced by 1 mM (final concentration) isopropyl β-D-thiogalactoside (IPTG, HighMedia) and cells were cultured for an additional 4 h. The culture was centrifuged (10,000 rpm, 10 min, 4 °C) and pellets were suspended in 2 ml sodium phosphate buffer containing 10 mM imidazole. The cells were disrupted by sonication (5 s pulse, 20 s rest over 30 min) and centrifuged (12,000 rpm, 4 °C, 20 min). Recombinant L-asparaginase of P. piersonii IIIF1SW-P2T (Kp_AnsA) was purified from supernatant using a Ni–NTA column chromatography with sodium phosphate buffer containing gradients of imidazole (20, 50, 100, 150, 200 and 250 mM) followed by dialysis (10 kDa cut-off).
Sodium dodecyl sulfate polyacrylamide gel electrophoresis and bradford assay
The purity and molecular mass of Kp_AnsA were assessed using Sodium Dodecyl Sulfate Polyacrylamide Gel Electrophoresis (SDS-PAGE). Denatured proteins were analyzed alongside a pre-stained protein ladder as a molecular mass reference. The target protein’s theoretical molecular mass, derived from its amino acid sequence in SwissProt, was 37.975 kDa. For SDS-PAGE, 15 µl of each eluate was mixed with 5 µl of loading buffer and denatured at 95 °C for 5 min. The samples were resolved on a gel comprising a 5% acrylamide stacking gel (0.5 M Tris–Cl, pH 6.8) and a 10% acrylamide resolving gel (1.5 M Tris–Cl, pH 8.8) using a Bio-Rad system, with electrophoresis conducted over 75 min. The gel was stained with Coomassie Brilliant Blue for 60 min on an orbital shaker, followed by destaining with methanol:water:acetic acid (5:4:1, v/v) solution.
The protein concentration of the samples was determined through the Bradford method15 using bovine serum albumin as a standard. The reaction mixtures were transferred to a 96-well microplate, and absorbance was measured at 595 nm using a microplate reader Asys UVM 340 (Biochrom, Holliston, MA, USA).
In silico prediction and molecular docking analysis
The theoretical isoelectric pH (pI) and molecular weight of the recombinant protein were predicted through pI and Mw tools of Expasy (https://web.expasy.org/compute_pi/). The domain search was carried out at UniProt using RKJ90748.1 as a query sequence. The 3D structure of L-asparaginase was predicted using Swiss-Model Software (http://swissmodel.expasy.org/), and the resulting models were further validated with ITASSER16. The Qualitative Model Energy Analysis (QMEAN) score and Ramachandran plot were used to analyse the quality of the predicted model. Protein structures were prepared using the UCSF chimera protein preparation wizard (version 1.10). The 2D structures of the selected ligands were sourced from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/). To explore the molecular interactions between the L-asparagine/glutamine and L-asparaginase, molecular docking using PyRx version 0.9.817 was performed with the default setting and the output files were analysed using Discovery Studio 2020 Client and Chimera version 1.10.
L-asparaginase assay
L-asparagine monohydrate was used as a substrate for the L-asparaginase assay. The NH4+ released was measured according to the method published elsewhere18 with the microplate-scale modification of the experimental protocol2. Briefly, 95 µl of 1 mM L-asparagine prepared in phosphate buffer of pH 7 was mixed with 5 µl of purified enzyme. The reaction mixture was incubated at 37ºC for 30 min and the reaction was terminated by the addition of 80 µl of sodium cocktail and sodium hypochlorite solution18. The plates were incubated for an additional 30 min to attain a stable color and read at 650 nm. The emerald green or blue coloration indicated the liberation of NH4+ and the amount of liberated NH4+ was calculated using a standard curve prepared for (NH4)2SO42,18.
Effect of pH, temperature, metal ions and substrate concentrations on L-asparaginase
The optimum pH was determined by carrying out the enzymatic reaction in the buffer with a pH range of 4‒9 (1 pH unit intervals). The optimum temperature for the catalytic activity was determined by maintaining the enzymatic reactions at 4, 25, 37, 50, 75 and 100ºC. The effect of metal ions such as manganese, cobalt, potassium, magnesium, sodium, zinc, calcium and mercury was studied by incubating the enzyme with chloride salts of respective metal ions (20 mM). Relative activity of Kp_AnsA at various pH, temperature and metal ions were determined by comparing its activity to respective controls maintained at pH 7, 37 °C and 20 mM NaCl, respectively. The effect of the substrate concentration was determined by carrying out reactions with increasing concentration of L-asparagine (0‒5 mM). The Michaelis constant (Km), maximum velocity (Vmax) and turnover number (Kcat) were calculated using GraphPad Prism version 8.
Phylogenetic analysis
The genes encoding cytoplasmic L-asparaginase I of IIIF1SW-P2T (Locus tag: D7S44_08820; Protein: RKJ90748.1) and YU22 (Locus tag: EKL29_18180 Protein: RTY55432.1) were retrieved from their corresponding whole genome shotgun sequence data available at NCBI (GCA_003612015.1 and GCA_003970755.1, respectively). Closest related sequences were retrieved from the NCBI database through the BLASTp search. Reference amino acid sequences corresponding to asparaginases were also retrieved from the in-house bacterial genome collections maintained at RAST for additional phylogenetic analysis. The NCBI and RAST originated sequences were aligned independently through Clustal_X19 and analysed via MEGA 5 (Molecular Evolutionary Genetics Analysis, version 5.020). The evolutionary history was inferred by using the Maximum Likelihood method based on the JTT matrix-based model21. The tree visualized tough MEGA with the highest log likelihood is shown. The percentage of trees in which the associated taxa clustered together is shown next to the branches. Initial tree(s) for the heuristic search were obtained by applying the Neighbor-Joining method to a matrix of pairwise distances estimated using a JTT model. The tree is drawn to scale, with branch lengths measured in the number of substitutions per site. All positions containing gaps and missing data were eliminated. Tree topology was evaluated by using bootstrap resampling based on 1,000 replications22.
In vitro cytotoxicity assay
HEK 293 T cell line procured from the National Centre for Cell Science in Pune, India, was cultured in Dulbecco’s Modified Eagle’s Medium supplemented with 10% fetal bovine serum and 1% antibiotic–antimycotic solution. Post-confluence of 70%, cells were trypsinized, centrifuged, and resuspended in cell line media (CLM), then seeded into a 96-well plate (5000 cells/well). After a 24 h incubation at 37 °C with 5% CO2, HEK 293 T cells were cultivated in CLM, CLM supplemented with 0.5% (w/v) L-asparagine (CLMA), CLMA containing Kp_AnsA (CLMA_Kp_AnsA), CLM containing Kp_AnsA (CLM_Kp_AnsA) and aqueous NH4OH (CLM_NH4OH). Following treatment, MTT reagent (1 mg/mL) was added to the wells, and the plate was incubated for 4 h. The formazan crystals were dissolved in dimethyl sulfoxide and the absorbance was measured at 570 nm using a multimode reader.
Live-dead cell staining and quantification of residual NH4 +
Cytotoxicity in the above-mentioned treatments was also evaluated through fluorescence microscopy using acridine orange-ethidium bromide (AO-EB) staining. HEK 293 T cells were plated (20,000 cells/well) and incubated for 24 h, followed by treatment as described under cytotoxicity assay for an additional 24 h. Cells were washed, and stained with 2 µg/ml of AO-EB for 15 min, excess stain was washed off using 1X PBS, cells were overlaid with PBS and examined under ZOE™ fluorescent cell imager, using green and red channels to assess differential viability. Results were compared to the untreated control. The NH4+ in the media supplied was quantified according to earlier descriptions2,18.
Engineering gfp-tagged P. piersonii IIIF1SW-P2T
Competent IIIF1SW-P2T cells were prepared using the calcium chloride method, transformed with pHC60 using heat shock and grown on an LB plate containing tetracycline (20 μg ml−1) for 24‒48 h. The gfp expression in IIIF1SW-P2T cells was achieved by adding 1 mM IPTG to LB agar. The expression of gfp was visualized through epifluorescence microscopy (excitation 475 nm; emission 509 nm).
Synchronisation of Caenorhabditis elegans
The worms were synchronized as per the earlier description23 and cultured in Nematode Growth Medium (NGM), with E. coli OP50 lawn at 21 °C until maturity as described earlier24. Worms were collected after reaching a dense population by pouring M9 buffer over the plate. The sample was centrifuged (1500 rpm, 25 °C, 1 min), the pellet was resuspended in sodium hypochlorite solution, and shaken for 3 min to remove contaminants and separate eggs from adults. After rinsing twice with M9 buffer, the eggs were incubated at 16 °C overnight to hatch, ensuring a uniform population of larvae for assays.
C. elegans survival assay
The synchronized eggs were incubated at 16 °C for 3 days. Once the nematodes reached the L4 stage, the synchronized individuals were washed with 1 ml of S-basal buffer three times until the supernatant became clear. Twenty L4 nematodes were transferred to a fresh NGM plate containing 1 mM IPTG, inoculated with gfp-tagged IIIF1SW-P2T (IIIF1SW-P2T_pHC60). The impacts of OP50 and P. aeruginosa PAO1 and wild type IIIF1SW-P2T on C. elegans were analysed in parallel. Bacterial suspensions (~ 1 × 108 CFU per ml each) were plated separately on NGM for the preparation of respective bacterial lawns. Dead worms were counted at 0, 2, 22 and 46 h of incubation.
Tracing gfp-tagged P. piersonii IIIF1SW-P2T at the reproductive tract of C. elegans
L4 stage C. elegans were maintained on the lawn of gfp-tagged IIIF1SW-P2T grown on LB supplemented with 1 mM IPTG for 2 h. The nematodes were picked up and observed under a fluorescence microscope (excitation 475; emission 509) for localization of gfp-tagged IIIF1SW-P2T in vivo.
Time course analysis of P. piersonii IIIF1SW-P2T colonization
Synchronized L4 nematodes (n = 20) were transferred to an NGM plate containing 1 mM IPTG, inoculated with gfp-tagged IIIF1SW-P2T (IIIF1SW-P2T_pHC60), and incubated at 21 °C. Worms were collected 0.5, 2, 22, and 46 h post-exposure for fluorescence imaging analysis. The worms were placed on the glass slide and immobilized using 5% (v/v) aqueous methanol. After mounting the coverslip, the worms were observed using the ZOE™ fluorescent cell imager.
Sustained colonization and fluorescence measurement
Synchronized L4 nematodes (n = 20) were transferred to an NGM plate containing 1 mM IPTG, inoculated with gfp-tagged IIIF1SW-P2T (IIIF1SW-P2T_pHC60). Worms were also simultaneously transferred to separate NGM plates containing wild-type IIIF1SW-P2T, OP50 and PAO1 in parallel. Worms were collected at 0.5, 2, 22, and 46 h post-exposure for fluorescence imaging analysis. The worms were placed on the glass slide and immobilized using 5% (v/v) aqueous methanol. After mounting the coverslip, the worms were observed using ZOE™ fluorescent cell imager. The vulvar region of C. elegans was examined for sustained colonization. Fluorescence images of C. elegans expressing gfp were analysed using ImageJ software. A fixed region of interest covering only the vulvar region, measuring an area of 2500 pixels2 was manually selected to measure total fluorescence. The integrated density was used to estimate the fluorescence intensity.
Statistical analysis
Statistical significance was determined by either ordinary one-way analysis of variance (ANOVA) with Tukey’s multiple comparisons test or t-test using GraphPad Prism (*P < 0.1, **P < 0.05, ***P < 0.01, ****P < 0.0001; ns, non-significant), unless specified otherwise.
Results
Molecular and phylogenetic characterization of L-asparaginase of IIIF1SW-P2T
The gene encoding cytoplasmic L-asparaginase I (Kp_ansA, 1014 bp; D7S44_08820) of IIIF1SW-P2T shared absolute nucleotide sequence similarity with that of YU22 (EKL29_18180). The theoretical pI and molecular weight of the L-asparaginase I of IIIF1SW-P2T were estimated to be 5.84 and 36,954.94 Da, respectively. Ni–NTA purified recombinant L-asparaginase of IIIF1SW-P2T (Kp_AnsA) complied with the theoretical molecular weight as evident through SDS PAGE analysis (Fig. 1a, Supplementary Fig. S1). The domain search at UniProt revealed the asparaginase domain (position 5‒188) at the N-terminus and glutaminase domain (position 213‒326) at the C-terminus. The QMEN score of the L-asparaginase model was 0.04, while the Global Model Quality Estimation score was 0.90. The Ramachandran plot analysis showed that 95.58% of the residues were within the allowed region. Binding energy scores of −6.0 kcal/mol and −5.5 kCal/mol, respectively, were obtained for Kp_AnsA while using L-asparagine and glutamine as ligands during the docking study. L-asparagine predicted to establish hydrogen bond with Thr14, Ser 60, Ser61, Thr91, Asp92, Lys163 and Asn246 of Kp_AnsA (Fig. 1b‒c). In contrast, glutamine established hydrogen bonds with Thr14, Thr91, Asp92, Ser117, Gln118 and Lys163. These two ligands also varied in terms of residues that established Van der Waals forces of attraction. These data suggested the possible involvement of Thr14, Thr91, Asp92 and Lys163 as common catalytic site residues for L-asparagine and glutamine.
Purification, characterization and phylogenetic analysis of recombinant L-asparaginase from Pantoea piersonii IIIF1SW-P2T (Kp_AnsA), overexpressed in Escherichia coli BL21 (DE3). SDS PAGE analysis of Ni–NTA affinity chromatography purified Kp_AnsA (a, arrowhead), its substrate (Asparagine) binding site (b), active site residues (c), enzyme kinetics (d), influence of pH (e), temperature (f) and metal ions (g) are shown. Additional details of SDS PAGE can be found in Fig. S1. Reactions maintained at pH 7, 37 °C and NaCl were used as controls for pH (e), temperature (f) and metal ion (g) assays, respectively. Error bar, mean (n = 2) ± SD. Statistical significance was determined through ordinary One-way ANOVA (Dunnett’s multiple comparison test). *P < 0.1, **P < 0.05, ***P < 0.01, ****P < 0.0001; ns, non-significant. Unrooted maximum likelihood tree of amino acid sequences depicting the phyletic lineage of P. piersonii IIIF1SW-P2T and other bacterial species/strains of closest sequence match (h). Bootstrap values of > 70% after 1,000 bootstrap replicates are shown at the branch points. Reference sequences were retrieved from NCBI. Bar, 0.01 substitutions per site.
Enzyme assays were carried out for Kp_AnsA using L-asparagine as a substrate. Michaelis constant (Km) was found to be 5.4 mM for Kp_AnsA. Enzymatic analysis revealed a maximum velocity (Vmax) and turnover number (Kcat) of 8.4 U/mg and 135.6 μmoles s−1, respectively (Fig. 1d). The enzyme was found to be highly active in pH 8‒10 (optimum pH 9) and 37 °C (Fig. 1e‒f, respectively). The enzyme exhibited statistically similar activity when provided with diverse metal ions, except for cobalt and mercury that suppressed the catalysis (Fig. 1g). Kp_AnsA formed a tight (83% bootstrap confidence of the node) coherent cluster with that of strains of P. piersonii during maximum likelihood phylogenetic analysis (Fig. 1h). The analysis using randomly chosen genome-sequenced bacterial species revealed a tight (99% bootstrap confidence of the node) phylogenetic neighborhood formed between Kp_AnsA and Idiomarina tyrosinivorans CC-PW-9 T, a marine bacterial species (Fig. S2). Several new bacterial species isolated from marine samples were also formed a tight adjacent clade. Thus, both IIIF1SW-P2T and YU22 found to produce structurally identical L-asparaginase, which is active at 37 °C, pH 8‒10 and in the presence of diverse metal ions.
L-asparaginase-mediated NH4 + generation is cytotoxic to HEK 293 T
The possible cytotoxicity of bacterial L-asparaginase on HEK 293 T was investigated. The impacts of pure Kp_AnsA (1.5 μg per well; final concentration) on HEK 293 T were tested in the presence/absence of 0.5% (w/v) L-asparagine as a substrate. The extent of cytotoxicity exerted by Kp_AnsA on HEK 293 T was assessed qualitatively and quantitatively using AO/EB staining. Fluorescence microscopy showed enhanced cytotoxicity in HEK 293 T cells in the presence of Kp_AnsA and L-asparagine (CLMA_Kp_AnsA) as compared to that of enzyme-free L-asparagine-amended media (CLMA) (Fig. 2a). Similarly, cytotoxicity was also found when HEK 293 T cells were treated with enzyme without L-asparagine (CLM_Kp_AnsA). HEK 293 T cells treated with standard NH4OH (CLM_NH4OH) also displayed cytotoxicity. Kp_AnsA enhanced cytotoxicity was substantiated through fluorescent measurements where significantly (P < 0.0001) high fluorescence was detected emitting from cells treated with CLMA_Kp_AnsA as compared to that of CLMA (Fig. 2b). Similarly, significant fluorescent intensity was detected in HEK 293 T cells when exposed to L-asparagine-free media (CLM_Kp_AnsA), suggesting L-asparagine-independent NH4+ formation. Fluorescence response of HEK 293 T cells treated exclusively with standard NH4OH (CLM_NH4OH) validated NH4+ driven cytotoxicity. Thus, fluorescence microscopy provided preliminary evidence for the susceptibility of HEK 293 T to NH4+, a by-product of enzymatic degradation of L-asparagine.
Cytotoxic impacts of purified recombinant L-asparaginase of Pantoea piersonii IIIF1SW-P2T (Kp_AnsA) on Human Embryonic Kidney 293 (HEK 293 T) cells (a-c) and chromogenic reactions showing differential NH4+ content (d). The response of HEK 293 T cells cultivated in cell line media (CLM), CLM supplemented with 0.5% asparagine (CLMA), CLMA containing Kp_AnsA (CLMA_Kp_AnsA), CLM containing Kp_AnsA (CLM_Kp_AnsA) and aqueous NH4OH (CLM_NH4OH) after live/dead assay done through acridine orange-ethidium bromide (a) and corresponding CLM-subtracted fluorescent intensity of merged image (b) are shown. Cytotoxicity (%) of HEK 293 T in CLMA, CLMA_Kp_AnsA, CLM_Kp_AnsA and CLM_NH4OH are shown (b). Chromogenic reactions displaying the differential presence of NH4+ in CLMA, CLMA_Kp_AnsA, CLM_Kp_AnsA and CLM_NH4OH are shown (b). CLMA was used as a control in (b) and (c) for statistical analysis. Error bar in (b), mean (n = 7) ± SD; Error bar in (c), mean (n = 3) ± SD; ****P < 0.0001; **P < 0.01. Red asterisks display a significant level based on t-test.
Kp_AnsA-driven cytotoxicity on HEK 293 T was assessed further using the MTT assay. Significant (P < 0.0001) cytotoxicity was detected in cells treated with CLMA_Kp_AnsA as compared to that of CLMA (Fig. 2c). Significant (P < 0.0001) cytotoxicity was also detected in HEK 293 T cells when treated with L-asparagine-free media (CLM_Kp_AnsA), substantiating L-asparagine-independent NH4+ formation. HEK 293 T cells treated with standard NH4OH (CLM_NH4OH) displayed significant (P < 0.0001) cytotoxicity, further validating the direct involvement of NH4+. The colorimetric analysis showed high amounts of NH4+ in asparaginase amended CLMA containing Kp_AnsA (CLMA_Kp_AnsA, 15.5 μM, P < 0.0001), followed by CLM_NH4OH (12.7 μM, P < 0.0001) and CLM_Kp_AnsA (4.1 μM, P < 0.0001) as compared to CLMA (Fig. 2d). These findings were in line with that of the live-dead staining assay, wherein L-asparagine-amended media in the presence of L-asparaginase and media with aqueous NH4OH induced cell death in HEK 293 T cells, as indicated by the presence of yellow fluorescing nuclei in the merge channel images.
Rapid and sustained colonization of IIIF1SW-P2T at the reproductive tract of C. elegans
The potential ability of IIIF1SW-P2T to colonize the eukaryotic reproductive tract in vivo was investigated using C. elegans as a model system. The worms were introduced to NGM plates containing a lawn of gfp-tagged cells of IIIF1SW-P2T and monitored under fluorescence microscopy. We found distinct colonization of IIIF1SW-P2T at the vulvar and adjacent luminal region of C. elegans (Fig. 3a‒b). The intense fluorescence corresponding to IIIF1SW-P2T cells was not evident at the mouth and anal region of the C. elegans, suggesting that IIIF1SW-P2T cells specifically colonize the reproductive tract of the nematode. The colonization of the reproductive tract was found to initiate as early as 2 h of grazing and was found consistent till the completion (46 h) of the experiments (Fig. 3c). The time course analysis revealed rapid, sustained and specific luminal colonization of IIIF1SW-P2T_pHC60 at the reproductive tract of C. elegans.
Localization and time course of colonization of gfp-tagged Pantoea piersonii IIIF1SW-P2T (IIIF1SW-P2T_pHC60) at the reproductive tract of Caenorhabditis elegans. Bright field (a) and fluorescent (b) microscopic images showing the colonization of IIIF1SW-P2T_pHC60 at the anterior reproductive tract of C. elegans. Scale bar, 100 μm. Bright-field and fluorescence micrographs showing the time course of colonization of IIIF1SW-P2T_pHC60 in the reproductive tract of hermaphrodite C. elegans as evident after 2 h, 26 h and 46 h of infection (c). Arrowhead shows colonization at the vulvar and luminal regions. Red scale bar, 100 μm; black/white scale bar, 25 μm.
The impacts of IIIF1SW-P2T on the survival of C. elegans was assessed. Synchronized worms were left to graze on the plates containing wild-type IIIF1SW-P2T, IIIF1SW-P2T_pHC60, OP50 and PAO1. C. elegans treated with wild-type IIIF1SW-P2T and IIIF1SW-P2T_pHC60 were highly motile due to enhanced undulatory movements as compared to the worms treated with OP50. IIIF1SW-P2T (with and without gfp-tagging) exhibited marginal killing capacity, with higher mean lifespan and survival over time when compared to PAO1 (LRT: P = 0.0079; GBWT: P = 0.0097) and slightly decreased mean lifespan and survival over time when compared to OP50 (LRT: P = 0.0253; GBWT: P = 0.0253) (Fig. 4a). Fluorescence signals emitted from hermaphrodite worms were assessed after treating with IIIF1SW-P2T, IIIF1SW-P2T_pHC60, OP50 and PAO1. The low and high magnification bright field and fluorescence micrographs of C. elegans treated with IIIF1SW-P2T (Fig. 4b), IIIF1SW-P2T_pHC60 (Fig. 4c), OP50 (Fig. 4d) and PAO1 (Fig. 4e) consistently showed the autofluorescence of varying intensity emitting from the worm body. However, elevated gut fluorescence was observed in C. elegans treated with IIIF1SW-P2T_pHC60. Extensive analysis of the vulvar region revealed fluorescence signals corresponding to IIIF1SW-P2T_pHC60 emitting from the embryos and interembryonic space apart from the vulvar/luminal regions (Fig. 4c). In contrast, such signals were absent in the worms treated with OP50 and PAO1. Integrated density measurement revealed significantly high fluorescence emitting from IIIF1SW-P2T_pHC60 followed by IIIF1SW-P2T (Fig. 4f). In contrast, weak fluorescence signals were detected in the vulvar regions of worms treated with OP50 and PAO1. These data indicated the potential ability of IIIF1SW-P2T to colonize the reproductive tract of C. elegans with mild lethality to the host.
Survival assay and demonstration of sustained colonization of P. piersonii IIIF1SW-P2T at the reproductive tract. Percent survival was averaged across experimental replicates, before the calculation of differences in mean lifespan and survival over time of gfp-tagged P. piersonii IIIF1SW-P2T (P. piersonii IIIF1SW-P2T_pHC60) compared to wild-type P. piersonii IIIF1SW-P2T, P. aeruginosa PAO1 (positive control) and E. coli OP50 (negative control). Statistical significance was measured through the Log-rank (Mantel-Cox) test and the Gehan-Breslow-Wilcoxon test. Low and high magnification bright field and fluorescence micrographs showing bacterial colonization in hermaphrodite C. elegans (b). The bright field and fluorescence responses of C. elegans treated with wild-type P. piersonii IIIF1SW-P2T (a), gfp-tagged P. piersonii IIIF1SW-P2T (IIIF1SW-P2T_pHC60) (b), OP50 (c) and PAO1 (d) are shown. The expression of gfp in (b) was achieved using 1 mM IPTG. The vulvar region is highlighted in a black/white arrowhead. Colonization in the vulvar region, including the lumen, embryos and extraembryonic space, are highlighted through yellow arrowhead, white asterisks and white arrow, respectively. Red scale bar, 100 μm; black/white scale bars 25 μm. (e), fluorescent intensity detected in C. elegans across various treatments. Statistical comparisons were made using one-way ANOVA followed by Tukey’s multiple comparisons test. Error bar, mean (n = 4) ± SD. *P < 0.1, **P < 0.05, ***P < 0.01, ****P < 0.0001; ns, non-significant.
Discussion
Bacterial L-asparaginases have been well studied for their cytotoxicity, particularly on cancer cells8,9,10,12,13. The constraint in the bioavailability of L-asparagine driven by L-asparaginase, as the latter readily converts L-asparagine to aspartic acid, is regarded as one of the mechanisms causing cancer cell death13. However, the role of NH4+, a by-product of the catalytic breakdown of L-asparagine, in cytotoxicity remained unexplored. The rapid formation of NH4+ and significant upregulation of the asparaginase gene in YU22 when exposed to L-asparagine2,14 prompted this investigation on the ISS counterpart. Asparaginase-driven NH4+ formation was demonstrated to trigger struvite crystallization in urease-negative YU222. A subsequent study showed an alternative ATP-dependent catabolic pathway driven by urea carboxylase and allophanate hydrolase acting upon urea, possibly contributing to NH4+ formation14. Asparaginase-driven NH4+ formation, which is an ATP-independent process, appears to be one of the important virulence factors that provides a competitive advantage to the pathogen to thrive under extreme conditions of the reproductive tract. The amino acids and small peptides present in the urine serve as carbon sources for the bacteria residing in the urinary tract25,26. Asparagine is identified as one of the lifespan-decreasing amino acids for C. elegans27. The enzyme kinetics data presented here validated the active enzymatic formation of NH4+ from a simple amino acid L-asparagine. While NCBI-based phylogenetic analyses confirmed the proximity of purified enzyme to Pantoea, RAST-based analysis revealed its similarity to that of bacterial isolates of marine origins, substantiating the halotolerance.
An earlier study has employed the HEK 293 T cell line to demonstrate the adhesion and cytotoxicity of YU222. In this study, the impacts of purified recombinant Kp_AnsA on HEK 293 T cell line were investigated to decode the mechanism of cytotoxicity. NH4+ generated through the catalytic activity of Kp_AnsA on L-asparagine resulted in cytotoxicity. In contrast, cytotoxicity observed in the absence of L-asparagine is most likely due to the activity of Kp_AnsA on an alternative substrate, possibly glutamine present in the cell culture media. Our analysis in silico predicted the glutaminase domain in the C-terminus of Kp_AnsA. However, further studies are warranted to validate Kp_AnsA-driven NH4+ formation from alternative substrates. Mild toxicity detected in enzyme-free L-asparagine amended media (CLMA) is most likely due to the residual NH4+ impurities originating from L-asparagine/or spontaneous degradation of L-asparagine. The assessment of possible aspartate-driven toxicity was not possible in the current experimental setup. Furthermore, the enzyme and substrate concentrations during in vitro study were optimized to get detectable signals in colorimetric and fluorimetric analyses. The physiological and pathophysiological relevance of the tested doses of enzyme, substrate and product warrants further investigation using in vivo model.
C. elegans shares key genetic and physiological similarities with humans and hence being used as one of the potential non-mammalian model organisms to investigate ageing and the onset, progression and pathophysiology of various human disorders, including Alzheimer’s and Parkinson’s diseases28,29,30. Hermaphrodite C. elegans shares some reproductive similarities with human females and hence is proposed as a powerful high-throughput model to study the female reproductive health31. In this study, green fluorescence was consistently observed in the C. elegans gut irrespective of the bacterial treatment. This is probably due to intestinal autofluorescence that arises due to fluorophores (age pigments) of gastrointestinal tract32. The autofluorescent molecules are spectrally heterogeneous with distinct biological properties, and can be used as a non-invasive biomarker to probe senescence and advanced glycation end products in vivo30,32. While red autofluorescence is a candidate marker for health as it correlates well with an individual’s remaining days of life, blue autofluorescence is proposed to be an indicator of an individual’s incipient or recent demise32. In contrast, green autofluorescence is an ill-suited probe either for life or death since it combines both properties and extreme caution needs to be taken to distinguish gfp expression near the time of death from full-body fluorescence32. In this study, live worms were taken for imaging after brief methanol treatment in order to probe the colonization of target organisms. The distinct emission of green fluorescence at the vulva, lumen, uterus, eggs and embryos substantiated the rapid and sustained colonization of IIIF1SW-P2T at the reproductive tract. However, further studies are needed to discriminate the gut colonization of IIIF1SW-P2T_pHC60 and the autofluorescence emitting from the gastrointestinal tract of C. elegans.
C. elegans is reported to be a facile and inexpensive model host for investigating several Gram-positive bacterial pathogens infecting humans33. The lethal impacts of Enterococcus faecalis, Streptococcus pneumoniae, and Staphylococcus aureus, but not Bacillus subtilis, Enterococcus faecium, or Streptococcus pyogenes, on C. elegans have been documented. Another study identified diminished capacity of aged C. elegans to control intestinal bacterial accumulation34. These two studies emphasized the differential compatibility of C. elegans to pathogenic bacteria and suggested the insufficiency of a single exposure to track colonisation. Although the bacteria are the essential nutritional source of C. elegans, some pathogenic bacteria may cause infection and death to the nematode35. For example, Microbacterium nematophilum was demonstrated to cause distinct modes of infection and host response since it adheres to the rectal and post-anal cuticle, mimicking the natural infection of C. elegans36. Therefore, monitoring the colonization and clearance dynamics alongside the survival was necessary33,34. Therefore, the time course of colonization of IIIF1SW-P2T at the reproductive tract and survival of worms were invetigated in parallel. Our study revealed rapid and sustained colonization of IIIF1SW-P2T at the reproductive tract of C. elegans with mild mortality to the host. gfp-tagged IIIF1SW-P2T were localized in the vulvar area, including the vulvar lumen, which connects the vulva with the uterus required for mating and egg-laying37. In contrast, significant mortality observed after PAO1 treatment aligned with the previous reports on Pseudomonas representatives that impart lethality in C. elegans38,39,40,41.
Conclusion
A recombinant L-asparaginase of IIIF1SW-P2T (Kp_AnsA) was produced using E. coli BL21 (DE3), and its cytotoxic attributes on HEK 293 T cell line were demonstrated. Kp_AnsA was found to be a halotolerant enzyme phylogenetically related to asparaginases of marine bacteria and found to be highly active at alkaline pH. The recombinant Kp_AnsA with a molecular weight of 37.975 kDa displayed Km, Vmax and Kcat values of 5.4 mM, 8.4 U/mg and 135.6 μmoles s−1 versus L-asparagine and was found to readily generate NH4+ ions in vitro. The catalytic formation of NH4+ from L-asparagine underpins cytotoxicity in HEK 293 T. Assays in vivo using gfp-tagged cells revealed the rapid and sustained colonizing ability of IIIF1SW-P2T at the vulva, luminal region, embryo and interembryonic space of Caenorhabditis elegans, with mild mortality on the host. Thus, C. elegans and IIIF1SW-P2T appear to be a potential infection model to investigate reproductive health. The role played by L-asparaginase in the colonization of IIIF1SW-P2T at the reproductive tract of C. elegans merits further investigation.
Data availability
The datasets used and/or analysed during the current study available from the corresponding authors Dr. Asif Hameed & Dr. Rajesh P. Shastry on reasonable request.
Abbreviations
- ANOVA:
-
Analysis of variance
- AO-EB:
-
Acridine orange-ethidium bromide
- BL21 (DE3):
-
E. coli BL21 (DE3)
- BLASTp:
-
Basic local alignment search tool for protein
- CFU:
-
Colony-forming units
- CLM:
-
Cell line media
- CLMA:
-
CLM supplemented with L-asparagine
- CLM_Kp_AnsA:
-
CLM containing Kp_AnsA
- CLMA_Kp_AnsA:
-
CLMA containing Kp_AnsA
- CLM_NH4OH:
-
CLM containing aqueous NH4OH
- DH5α:
-
E. coli DH5α
- gDNA:
-
Genomic DNA
- GBWT:
-
Gehan-breslow-wilcoxon test
- Gfp:
-
Green fluorescent protein
- HEK 293 T:
-
Human embryonic kidney cell line
- IIIF1SW-P2T :
-
Wild-type Pantoea piersonii IIIF1SW-P2T
- IIIF1SW-P2T :
-
pHC60 + gfp-tagged IIIF1SW-P2T
- IPTG:
-
Isopropyl β-D-thiogalactoside
- ISS:
-
International Space Station
- K cat :
-
Turnover number
- K m :
-
Michaelis constant
- Kp_ansA :
-
Gene encoding cytoplasmic L-asparaginase I of IIIF1SW-P2T
- Kp_AnsA:
-
Recombinant L-asparaginase I of P. piersonii IIIF1SW-P2T
- LB:
-
Luria Bertani
- L4:
-
Fourth larval stage
- LRT:
-
Log-rank (Mantel-Cox) test
- MEGA:
-
Molecular evolutionary genetics analysis
- MTT:
-
3-(4,5-Dimethylthiazol-2-yl)-2,5-diphenyl-2H-tetrazolium bromide
- NCBI:
-
National centre for biotechnology information
- NGM:
-
Nematode growth medium
- Ni–NTA:
-
Nickel-nitrilotriacetic acid
- OP50:
-
E. coli OP50
- PAO1:
-
Pseudomonas aeruginosa PAO1
- PBS:
-
Phosphate buffered saline
- PCR:
-
Polymerase chain reaction
- pET28a:
-
Protein overexpression vector pET28a( +)
- pHC60:
-
gfp Expression vector pHC60
- pET28a_Kp_ansA :
-
pET28a( +) vector carrying L-asparaginase gene of IIIF1SW-P2T
- QMEAN:
-
Qualitative model energy analysis
- RAST:
-
Rapid Annotation using subsystem technology
- SDS-PAGE:
-
Sodium dodecyl sulfate polyacrylamide gel electrophoresis
- TA_Kp_ansA :
-
TA cloning vector carrying L-asparaginase gene of IIIF1SW-P2T
- V max :
-
Maximum velocity
- YU22:
-
P. piersonii YU22
References
Singh, N. K., Wood, J. M., Mhatre, S. S. & Venkateswaran, K. Metagenome to phenome approach enables isolation and genomics characterization of Kalamiella piersonii gen. nov., sp. nov. from the International Space Station. Appl Microbiol Biotechnol 103, 4483–4497. https://doi.org/10.1007/s00253-019-09813-z (2019).
Rekha, P. D. et al. First report of pathogenic bacterium Kalamiella piersonii isolated from urine of a kidney stone patient: draft genome and evidence for role in struvite crystallization. Pathogens https://doi.org/10.3390/pathogens9090711 (2020).
McDonagh, F. et al. First complete genome of a multidrug-resistant strain of the novel human pathogen Kalamiella piersonii (GABEKP28) identified in human saliva. J Glob Antimicrob Resist 32, 31–34. https://doi.org/10.1016/j.jgar.2022.12.003 (2023).
Howard, M., Maki, J. J., Connelly, S., Hardy, D. J. & Cameron, A. Complete genome sequence of a human bacteremia isolate of Kalamiella piersonii. Microb Resour Announc 12, e0029323. https://doi.org/10.1128/MRA.00293-23 (2023).
Sada, J. et al. Bacteremia caused by Kalamiella piersonii found in an infant during the course of gastrointestinal food allergy. Infect Drug Resist 16, 2647–2651 (2023).
Atilan, K., Ozdem, T., Aydogan, C. N. & Hosbul, T. A rare case report of tissue infection caused by Pantoea piersonii (basionym Kalamiella piersonii). Folia Microb (Praha) https://doi.org/10.1007/s12223-024-01203-x (2024).
Soutar, C. D. & Stavrinides, J. Phylogenomic analysis of the Erwiniaceae supports reclassification of Kalamiella piersonii to Pantoea piersonii comb. nov. and Erwinia gerundensis to the new genus Duffyella gen. nov. as Duffyella gerundensis comb. nov. Mol Genet Genomics 297, 213–225. https://doi.org/10.1007/s00438-021-01829-3 (2022).
Abdelrazek, N. A., Saleh, S. E., Raafat, M. M., Ali, A. E. & Aboulwafa, M. M. Production of highly cytotoxic and low immunogenic L-asparaginase from Stenotrophomonas maltophilia EMCC2297. AMB Express 14, 51. https://doi.org/10.1186/s13568-024-01700-9 (2024).
Abdullah, E. M. et al. Expression, characterization and cytotoxicity of recombinant l-asparaginase II from Salmonella paratyphi cloned in Escherichia coli. Int J Biol Macromol 279, 135458. https://doi.org/10.1016/j.ijbiomac.2024.135458 (2024).
Darwesh, D. B. et al. Anticancer activity of extremely effective recombinant L-asparaginase from Burkholderia pseudomallei. J. Micro. Biotechnol. 32, 551–563. https://doi.org/10.4014/jmb.2112.12050 (2022).
Borges, G. A. et al. Asparaginase induces selective dose- and time-dependent cytotoxicity, apoptosis, and reduction of NFκB expression in oral cancer cells. Clin. Exp. Pharmacol. Physiol. 47, 857–866. https://doi.org/10.1111/1440-1681.13256 (2020).
Serravalle, S., Bertuccio, S. N., Astolfi, A., Melchionda, F. & Pession, A. Synergistic Cytotoxic Effect of L-Asparaginase Combined with Decitabine as a Demethylating Agent in Pediatric T-ALL, with Specific Epigenetic Signature. Biomed Res International 2016, 1985750. https://doi.org/10.1155/2016/1985750 (2016).
Narta, U. K., Kanwar, S. S. & Azmi, W. Pharmacological and clinical evaluation of L-asparaginase in the treatment of leukemia. Crit Rev Oncol Hematol 61, 208–221. https://doi.org/10.1016/j.critrevonc.2006.07.009 (2007).
Yuvarajan, S., Hameed, A., Arun, A. B., Saptami, K. & Rekha, P. D. Urease-negative uropathogen Kalamiella piersonii YU22 metabolizes urea by urea carboxylase and allophanate hydrolase enzyme system. Microb Res 263, 127142. https://doi.org/10.1016/j.micres.2022.127142 (2022).
Hammond, J. B. & Kruger, N. J. The bradford method for protein quantitation. Methods Mol Biol 3, 25–32. https://doi.org/10.1385/0-89603-126-8:25 (1988).
Yang, J. Y. et al. The I-TASSER Suite: protein structure and function prediction. Nat. Methods 12, 7–8. https://doi.org/10.1038/nmeth.3213 (2015).
Dallakyan, S. & Olson, A. J. Small-molecule library screening by docking with PyRx. Methods Mol Biol 1263, 243–250. https://doi.org/10.1007/978-1-4939-2269-7_19 (2015).
Baethgen, W. E. & Alley, M. M. A manual colorimetric procedure for measuring ammonium nitrogen in soil and plant Kjeldahl digests Commun Soil Sci. Plant Anal. 20(961), 969 (1989).
Thompson, J. D., Gibson, T. J., Plewniak, F., Jeanmougin, F. & Higgins, D. G. The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25, 4876–4882. https://doi.org/10.1093/nar/25.24.4876 (1997).
Tamura, K. et al. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28, 2731–2739. https://doi.org/10.1093/molbev/msr121 (2011).
Jones, D. T., Taylor, W. R. & Thornton, J. M. The rapid generation of mutation data matrices from protein sequences. Comput Appl Biosci 8, 275–282. https://doi.org/10.1093/bioinformatics/8.3.275 (1992).
Felsenstein, J. Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39, 783–791. https://doi.org/10.1111/j.1558-5646.1985.tb00420.x (1985).
Bajire, S. K., Jain, S., Johnson, R. P. & Shastry, R. P. 6-Methylcoumarin attenuates quorum sensing and biofilm formation in Pseudomonas aeruginosa PAO1 and its applications on solid surface coatings with polyurethane. Appl Microb Biotechnol 105, 8647–8661. https://doi.org/10.1007/s00253-021-11637-9 (2021).
Bajire, S. K., Prabhu, A., Bhandary, Y. P., Irfan, K. M. & Shastry, R. P. 7-Ethoxycoumarin rescued from infection of COPD derived clinical isolate Pseudomonas aeuginosa through virulence and biofilm inhibition via targeting Rhl and Pqs quorum sensing systems. World J Microbiol Biotechnol 39, 208. https://doi.org/10.1007/s11274-023-03655-8 (2023).
Alteri, C. J., Himpsl, S. D., Shea, A. E. & Mobley, H. L. T. Flexible Metabolism and Suppression of Latent Enzymes Are Important for Adaptation to Diverse Environments within the Host. J Bacteriology 201, e00181. https://doi.org/10.1128/JB.00181-19 (2019).
Mann, R., Mediati, D. G., Duggin, I. G., Harry, E. J. & Bottomley, A. L. Metabolic Adaptations of Uropathogenic in the Urinary Tract. Frontiers Cellular Infection Microbiology 7, 241. https://doi.org/10.3389/fcimb.2017.00241 (2017).
Edwards, C. et al. Mechanisms of amino acid-mediated lifespan extension in Caenorhabditis elegans. BMC Genet 16, 8. https://doi.org/10.1186/s12863-015-0167-2 (2015).
Alvarez, J., Alvarez-Illera, P., Santo-Domingo, J., Fonteriz, R. I. & Montero, M. Modeling Alzheimer’s Disease in. Biomedicines 10, 288. https://doi.org/10.3390/biomedicines10020288 (2022).
Cooper, J. F. & Van Raamsdonk, J. M. Modeling Parkinson’s Disease in C. elegans. J Parkinson Dis 8, 17–32. https://doi.org/10.3233/Jpd-171258 (2018).
Komura, T., Yamanaka, M., Nishimura, K., Hara, K. & Nishikawa, Y. Autofluorescence. as a noninvasive biomarker of senescence and advanced glycation end products in. Npj Aging Mech Dis 7, 12. https://doi.org/10.1038/s41514-021-00061-y (2022).
Athar, F. & Templeman, N. M. C. elegans as a model organism to study female reproductive health. Comp Biochem Phys A 266, 111152. https://doi.org/10.1016/j.cbpa.2022.111152(2022) (2022).
Pincus, Z., Mazer, T. C. & Slack, F. J. Autofluorescence as a measure of senescence in: look to red, not blue or green. Aging -Us 8, 889–898. https://doi.org/10.18632/aging.100936 (2016).
Garsin, D. A. et al. A simple model host for identifying Gram-positive virulence factors. Proc Natl Acad Sci U S A 98, 10892-10897, https://doi.org/10.1073/pnas.191378698 (2001).
Portal-Celhay, C., Bradley, E. R. & Blaser, M. J. Control of intestinal bacterial proliferation in regulation of lifespan in Caenorhabditis elegans. BMC Microb 12, 49. https://doi.org/10.1186/1471-2180-12-49 (2012).
Schulenburg, H. & Felix, M. A. The Natural Biotic Environment of Caenorhabditis elegans. Genetics 206, 55–86. https://doi.org/10.1534/genetics.116.195511 (2017).
Hodgkin, J., Kuwabara, P. E. & Corneliussen, B. A novel bacterial pathogen, Microbacterium nematophilum, induces morphological change in the nematode C. elegans. Curr Biol 10, 1615–1618. https://doi.org/10.1016/s0960-9822(00)00867-8 (2000).
Schindler, A. J. & Sherwood, D. R. Morphogenesis of the Caenorhabditis elegans vulva. Wiley Interdiscip Rev Dev Biol 2, 75–95. https://doi.org/10.1002/wdev.87 (2013).
Darby, C., Cosma, C. L., Thomas, J. H. & Manoil, C. Lethal paralysis of Caenorhabditis elegans by Pseudomonas aeruginosa. Proc Natl Acad Sci U S A 96, 15202–15207. https://doi.org/10.1073/pnas.96.26.15202 (1999).
Kirienko, N. V. et al. Pseudomonas aeruginosa disrupts Caenorhabditis elegans iron homeostasis, causing a hypoxic response and death. Cell Host Microbe 13, 406–416. https://doi.org/10.1016/j.chom.2013.03.003 (2013).
Mahajan-Miklos, S., Tan, M. W., Rahme, L. G. & Ausubel, F. M. Molecular mechanisms of bacterial virulence elucidated using a Pseudomonas aeruginosa-Caenorhabditis elegans pathogenesis model. Cell 96, 47–56. https://doi.org/10.1016/s0092-8674(00)80958-7 (1999).
Tan, M. W., Mahajan-Miklos, S. & Ausubel, F. M. Killing of Caenorhabditis elegans by Pseudomonas aeruginosa used to model mammalian bacterial pathogenesis. Proc Natl Acad Sci U S A 96, 715-720, https://doi.org/10.1073/pnas.96.2.715 (1999).
Acknowledgements
This research was funded in part by the National Science and Technology Council, Taiwan (Grant 113-2321-B-005-006), and the Innovation and Development Center of Sustainable Agriculture from The Featured Areas Research Center Program within the framework of the Higher Education Sprout Project by the Ministry of Education (MOE) in Taiwan. Asif Hameed acknowledges Yenepoya (Deemed to be University) for the Seed Grant (YU/Seed grant/139–2023). We thank Dr. Manjunatha Thondamal, Department of Biotechnology, GITAM School of Technology, Visakhapatnam, for providing C. elegans N2 strains and OP50. We also thank Ms. Krithika and Ms. Malathi for their help in optimizing protein purification and enzyme assays.
Funding
Yenepoya (Deemed to be University), India, YU/Seed grant/139–2023; National Science and Technology Council, Taiwan, 113-2321-B-005-006
Author information
Authors and Affiliations
Contributions
A.H. conceptualization, supervision, methodology, investigation, data analysis and curation, drafting original manuscript, C.C.Y. resources, investigation, data analysis and curation, K.V.S. investigation, visualization, data analysis and curation, A.P. methodology, investigation and manuscript editing, H.R.D. investigation, visualization, data analysis and curation, R.P.S. conceptualization, methodology, investigation, data analysis and curation, manuscript editing. All the authors have reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hameed, A., Young, CC., Suchithra, K.V. et al. Pantoea piersonii IIIF1SW-P2T triggers cytotoxicity through L-asparaginase-driven ammonium secretion and colonizes reproductive tract of Caenorhabditis elegans. Sci Rep 15, 35295 (2025). https://doi.org/10.1038/s41598-025-19368-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-19368-x