Abstract
Rehabilitation exercise assessment plays a crucial role in patient recovery, particularly for individuals recovering from injuries, surgeries, or illnesses affecting mobility. In this paper, we propose a novel approach for the assessment of skeleton-based data recorded from rehabilitation exercise, where we present the data as points on symmetric positive definite (SPD) manifold. Our method addresses the limitations of traditional Euclidean-based approaches by leveraging the SPD manifold’s ability to preserve motion variations and spatial relationships. We propose a novel framework leveraging SPD manifold to preserve the intrinsic geometry of human motion and capture nonlinear variations in complex movements. By embedding motion data into SPD manifolds, we integrate unsupervised K-Nearest Neighbors (KNN) with Riemannian geometry for precise classification of correct and incorrect movements. We further develop a Tangent Space Linear SPD Support Vector Machine (SVM), optimized via stochastic gradient descent (SGD) in the tangent space at the identity matrix. Additionally, a tailored neural network architecture with multi-scale feature extraction enhances movement assessment by capturing hierarchical patterns in vectorized SPD data. Our specialized neural network, designed for vectorized SPD data, outperforms state-of-the-art methods on three benchmark datasets: Kimore, UI-PRMD, and EHE. In cross-subject evaluations, accuracy improves to 92.40% (UI-PRMD), 85.18% (Kimore), and 87.59% (EHE), with even greater improvements in random train-test splits. Although the proposed method involves a high parameter count while reducing computational complexity in terms of floating-point operations, literature suggests that certain concepts and objects recur across diverse mathematical domains, often carrying significant implications. Additionally, manifold transformations in data representation effectively capture the intrinsic geometric structure. Furthermore, our training process is faster than state-of-the-art methods, leading to quicker model convergence and reduced computational overhead without compromising accuracy. These results highlight the potential of SPD manifolds for accurate, reliable rehabilitation assessment.
Similar content being viewed by others
Introduction
Rehabilitation plays a crucial role in improving the quality of life of individuals affected by injuries, medical conditions, or age-related decline. It aims to restore independence, optimize functional ability and reduce disability, enabling participation in daily activities, work, and education. According to the World Health Organization (WHO)1, approximately one in three people worldwide would benefit from rehabilitation, a need that is growing due to increasing life expectancy and the prevalence of chronic diseases. However, the effectiveness of rehabilitation can be compromised by low patient compliance and incorrect execution of exercises without therapist supervision2,3, resulting in longer treatment times and increased healthcare costs.
To address these challenges, there has been growing interest in AI-driven virtual rehabilitation systems4,5,6, which leverage motion sensors and machine learning techniques to analyze patients’ movements and provide automated feedback. These systems offer a cost-effective and scalable solution, enabling home rehabilitation while allowing clinicians to remotely monitor patient progress and intervene when necessary. Central to these systems is the ability to accurately assess rehabilitation exercises, ensuring that patients perform movements correctly and effectively.
Assessment of rehabilitation exercises has been explored through various approaches, including rule-based, template-based, and machine learning methods. Rule-based methods rely on predefined criteria, such as joint angles and movement trajectories, but struggle with generalization among individuals7,8. Template-based techniques, such as Dynamic Time Warping (DTW), improve robustness by aligning motion sequences9,10, yet remain sensitive to temporal distortions. More recently, deep learning methods—particularly Convolutional Neural Networks (CNNs)11,12 and Recurrent Neural Networks (RNNs)13,14—have shown significant promise in motion analysis tasks. Among these, Graph Convolutional Networks (GCNs)15,16 have emerged as powerful tools by modeling skeletal data as structured graphs, effectively capturing spatial and temporal dependencies in human movement.
Despite their strong performance, these deep learning methods are fundamentally rooted in Euclidean geometry, which constrains their capacity to fully capture the non-linear, non-Euclidean structure of complex human motion data. Specifically, when motion data lie on Riemannian manifolds—such as the SPD manifold—Euclidean-based techniques may distort intrinsic geometry, leading to suboptimal performance. This geometric mismatch limits generalization, interpretability, and robustness, especially in real-world rehabilitation settings involving high inter-subject variability and nuanced motion patterns.
To transcend the limitations inherent in traditional Euclidean-based motion analysis, we propose a novel framework that operates within the rich geometry of SPD manifolds. This formulation not only preserves the intrinsic spatial relationships of human motion, but also captures the nonlinear variations that characterize complex movement patterns. By embedding motion data into the SPD manifold, we introduce a sophisticated classification paradigm that combines unsupervised KNN with Riemannian geometry, enabling accurate distinction between correct and incorrect movements. Extending this, we develop a Tangent Space Linear SPD SVM that optimizes performance by mapping data to the tangent space at the identity matrix, leveraging stochastic gradient descent (SGD) for efficient training. To further enhance movement assessment, we introduce a specialized neural network architecture tailored to vectorized SPD data, augmented by a multi-scale feature extraction strategy that captures hierarchical movement patterns. This holistic approach redefines the potential for human motion analysis, offering an elegant and computationally efficient solution grounded in advanced geometric principles.
This paper makes the following key contributions:
-
SPD-Based Classification Framework: We present an innovative framework for assessing skeleton-based rehabilitation exercises, integrating KNN, a Tangent Space Linear SPD SVM, and a specialized neural network. By leveraging Log-Euclidean mapping, our approach preserves the geometric integrity of skeletal motion, ensuring a more precise and robust evaluation of movement patterns.
-
Tangent Space Linear SPD SVM: We propose a geometry-aware Tangent Space Linear SPD SVM, optimized via stochastic gradient descent (SGD) with an Log-Euclidean metric, enabling robust and accurate classification of SPD data.
-
Tailored Neural Network Architecture: We design a specialized neural network for vectorized SPD data, integrating convolutional and fully connected layers to effectively capture both local and global movement features. Our method achieves state-of-the-art performance, improving accuracy from 89.95% to 92.40% on UI-PRMD, from 83.56% to 85.18% on KIMORE, and from 86.38% to 87.59% on EHE, outperforming existing methods on these benchmark datasets.
-
Multi-Scale Feature Extraction: We introduce a multi-scale feature extraction strategy applied across all methods, which enhances classification performance by capturing hierarchical movement patterns, with extensive experiments demonstrating significant improvement over Euclidean baselines and competitiveness with state-of-the-art SPD methods.
The remainder of this paper is organized as follows. The Related Work section reviews prior research on skeleton-based representation learning and classification methods. Preliminaries introduce key mathematical concepts and background knowledge related to SPD manifolds. The Methodology section details our proposed approach, including KNN on the SPD manifold, Tangent Space Linear SPD SVM, Log-Euclidean mapping, vectorization strategy, and neural network architecture. Experiments and Analysis present experimental results and evaluations demonstrating the effectiveness of our method. Finally, the Conclusion summarizes the paper and discusses potential future directions.
Related work
Rehabilitation exercise assessment methods can be broadly categorized into three approaches: rule-based methods, template-based approaches, and machine learning techniques. Rule-based methods involve manually defined rules but struggle with generalizability. Template-based approaches compare observed actions to standard templates, but their accuracy can be impacted by individual variations. Machine learning techniques, especially those using GCNs, offer promising solutions by capturing complex movement dynamics. The following sections will explore each approach in detail.
Rule-based methods involve manually defined rules to evaluate actions, such as joint angles or movement trajectories. While these methods provide a structured framework for movement assessment, they are highly dependent on expert knowledge and can be time-consuming to develop. For example, Clark et al.7 utilized trunk flexion angles and joint distances for postural control, as well as trunk tilt angles for gait retraining17. Zhao et al.8 expanded on this approach by introducing dynamic, static, and invariance rules to assess real-time motion during rehabilitation exercises. This method was later enhanced with real-time feedback18, allowing patients to perform exercises correctly without direct clinical supervision. Additionally, its application to Total Knee Replacement rehabilitation in home settings19 demonstrated its effectiveness in real-world scenarios. However, despite their structured nature, rule-based methods often struggle with generalizability, as predefined rules may not account for individual movement variations, ultimately limiting their accuracy across diverse rehabilitation contexts.
Template-based approaches rely on comparing measured movements to a reference template, typically derived from correct performances by healthy subjects. Benetazzo et al.20 employed Euclidean distance to evaluate joint positions and velocities, offering a straightforward metric for quantifying movement discrepancies; however, its sensitivity to even minor time-axis distortions makes it less effective for comparing sequences of differing durations. To address this limitation, Su et al.9 and Hu et al.10 adopted Dynamic Time Warping (DTW), which non-linearly aligns time sequences to provide more robust comparisons despite temporal misalignments. Building on these methods, Osgouei et al.21 proposed using Hidden Markov Modeling (HMM) to compare a user’s performance against a reference, offering advantages in later rehabilitation phases where detecting subtle inconsistencies becomes crucial. More recently, Li et al.22 introduced a Mahalanobis distance-based DTW (MDDTW) technique; unlike the original DTW that relies on Euclidean distance, MDDTW leverages the Mahalanobis distance to account for variable correlations and scale differences, potentially providing a more sensitive measure for comparing complex movement patterns.
Deep Learning-based methods have gained significant attention in recent years due to their remarkable ability to learn complex patterns from large datasets and model intricate relationships in data. For example,23 employed strided one-dimensional convolutional filters to capture spatial dependencies in human movements, followed by LSTM layers to model temporal correlations in the learned representations. Similarly,24 utilized LSTM networks to extract features from skeleton data, while CNNs were used to process foot pressure images, with the extracted features subsequently fused for further analysis. The human skeleton can be viewed as a graph in non-Euclidean space, with joints as vertices and bones as edges. In this context, using Convolutional Neural Networks (CNNs) may result in the loss of crucial spatial information, as CNNs are designed for grid-like (Euclidean) data and may not fully capture the structure of skeletal data. To address this limitation, Yan et al.25 proposed Spatial-Temporal GCNs (ST-GCN), which model the human body as a graph, applying GCNs for spatial feature extraction and temporal convolutions for motion analysis. Building on this, Bruce16 introduced a two-stage training approach: first, training on joint positions for Human Activity Recognition to identify key neurons; then, refining the model for action correctness assessment using both joint positions and angles, ensuring relevant neurons influence evaluation. Further, He et al.26 proposed an expert-knowledge-based GCN, where adjacency matrices are designed based on rehabilitation action insights. Their model processes skeleton sequences through two streams—one for general spatio-temporal features and another for rehabilitation-specific features—combining both for improved assessment accuracy.
While rule-based, template-based, and deep learning methods, including GCNs, have all been explored for rehabilitation assessment, they primarily operate in Euclidean space or approximations of it. Although GCNs better capture spatial relationships in skeletal data, they still rely on message-passing operations rooted in Euclidean assumptions and are not inherently designed to respect the geometric properties of data on curved spaces. This limits their capacity to model the underlying Riemannian structure of motion data, such as geodesic distances and affine invariance, which are crucial for understanding subtle movement variations in rehabilitation tasks.
To better address these geometric complexities, recent work has explored manifold-aware approaches such as hyperbolic embeddings27, which are well-suited for capturing hierarchical relationships, and Grassmannian manifolds28, which model dynamic actions through subspace representations. However, these methods often lack the ability to encode second-order statistics, such as covariance matrices of motion features, that are critical for fine-grained motion assessment because they capture the variability and correlations between skeletal joint movements over time. In contrast, our approach explicitly relies on covariance matrices to represent skeletal motion by embedding them in the SPD manifold. This manifold provides a natural Riemannian framework for covariance matrices, preserving the intrinsic geometric structure of motion data while effectively encoding second-order statistics, resulting in a more robust, interpretable, and generalizable assessment framework for rehabilitation tasks.
Preliminaries
This section introduces the fundamental concepts and mathematical tools necessary for the methodology. We provide formal definitions of the SPD manifold and the eight distance metrics employed in our classification approach.
SPD manifold
The Symmetric Positive Definite (SPD) manifold consists of symmetric matrices with strictly positive eigenvalues. Formally, the SPD matrix space is defined as:
where \(A^T\) denotes the transpose of the matrix A, and \(\lambda _{min}(A)\) is the smallest eigenvalue of A. In other words, the SPD manifold includes all symmetric matrices whose eigenvalues are strictly positive.
The SPD manifold is also a Riemannian manifold, meaning that it has a curved geometry. A Riemannian manifold allows us to define a distance function that respects the structure of the space.
Distance metrics on the SPD manifold
Measuring distances between SPD matrices using traditional Euclidean distance is inadequate, as it does not account for the manifold’s geometric structure. Instead, specialized metrics have been developed to capture the intrinsic geometry of the SPD manifold, providing a more meaningful measure of dissimilarity.
In this work, we employ eight distance metrics to quantify differences between SPD matrices A and B, including the Euclidean distance to highlight its limitations. Each metric offers a unique perspective on matrix dissimilarity, playing a crucial role in our analysis.
The distance metrics we use are as follows:
-
1.
Cholesky distance:
$$\begin{aligned} \text {dist}_{\text {chol}}(A, B) = \Vert \textbf{L}_A - \textbf{L}_B \Vert _F \end{aligned}$$where \(\textbf{L}_A\) and \(\textbf{L}_B\) are the lower triangular matrices obtained from the Cholesky decompositions of \(A\) and \(B\), respectively, and \(\Vert \cdot \Vert _F\) denotes the Frobenius norm. This metric quantifies the difference between the Cholesky factors of the two matrices29.
-
2.
Log-Euclidean distance:
$$\begin{aligned} \text {dist}_{\text {logeuclid}}(A, B) = \Vert \log (A) - \log (B) \Vert _F \end{aligned}$$where \(\log (A)\) and \(\log (B)\) are the matrix logarithms of \(A\) and \(B\), respectively. This distance captures the difference between the matrices in the log-domain, which is suitable for SPD matrices30.
-
3.
Affine-invariant distance:
$$\begin{aligned} \text {dist}_{\text {riemann}}(A, B) = \Vert \log (A^{-1/2} B A^{-1/2}) \Vert _F \end{aligned}$$where \(A^{-1/2}\) and \(B^{-1/2}\) are the matrix square roots of the inverses of \(A\) and \(B\), and \(\log (\cdot )\) denotes the matrix logarithm. This metric measures the distance between \(A\) and \(B\) in the Riemannian geometry of the SPD manifold31.
-
4.
Euclidean distance:
$$\begin{aligned} \text {dist}_{\text {euclid}}(A, B) = \Vert A - B \Vert _F \end{aligned}$$This is the standard Frobenius norm between the matrices \(A\) and \(B\). While simple, this metric does not respect the geometry of the SPD manifold.
-
5.
Kullback–Leibler divergence:
$$\begin{aligned} \text {dist}_{\text {kullback}}(A, B) = \text {Tr}(A^{-1} B) - \log \det (A^{-1} B) - d \end{aligned}$$where \(\text {Tr}(\cdot )\) denotes the trace of a matrix, and \(\det (\cdot )\) denotes the determinant. This metric is based on the Kullback–Leibler divergence, a popular measure of dissimilarity between probability distributions32.
-
6.
Symmetric Kullback–Leibler divergence:
$$\begin{aligned} \text {dist}_{\text {kullback\_sym}}(A, B) = \frac{1}{2} \left( \text {Tr}(A^{-1} B) - \log \det (A^{-1} B) - d \right) + \frac{1}{2} \left( \text {Tr}(B^{-1} A) - \log \det (B^{-1} A) - d \right) \end{aligned}$$This is a symmetric version of the Kullback–Leibler divergence, which considers both directions of divergence between \(A\) and \(B\)33.
-
7.
Log-Cholesky distance:
$$\begin{aligned} \text {dist}_{\text {logchol}}(A, B) = \Vert \log (\textbf{L}_A) - \log (\textbf{L}_B) \Vert _F \end{aligned}$$where \(\textbf{L}_A\) and \(\textbf{L}_B\) are the Cholesky decompositions of \(A\) and \(B\), respectively. This distance measures the difference between the logarithms of the Cholesky factors of the two matrices34.
-
8.
Log-determinant distance:
$$\begin{aligned} \text {dist}_{\text {logdet}}(A, B) = \Vert \log (\det (A)) - \log (\det (B)) \Vert \end{aligned}$$where \(\det (A)\) and \(\det (B)\) are the determinants of \(A\) and \(B\), respectively. This metric focuses on the difference in the log-determinants of the matrices35.
These diverse metrics allow us to explore SPD matrices from multiple geometric and statistical perspectives. By comparing their behaviors, we gain a deeper understanding of the structure and variability within the manifold, enabling more informed analysis and modeling in applications where SPD matrices arise, the relevant comparison results of above distance metrics can be found in Table 2.
Methodology
In this section, we outline our approach for classifying rehabilitation exercises using the SPD manifold. We begin by describing the embedding of tabular skeleton data into the SPD manifold using a multiscale representation. Subsequently, we present three classification strategies that reflect different levels of model complexity and design philosophy: (1) a KNN method leveraging distance metrics defined on the SPD manifold, offering a parameter-free baseline based purely on geometric similarity; (2) a Tangent Space Linear SPD SVM, introducing supervised learning under a linear decision boundary assumption; and (3) a neural network–based approach tailored to SPD-structured data, designed to learn task-adaptive and expressive representations in a geometry-aware framework.
This progression, from non-parametric to linear to deep modeling, enables a structured exploration of the SPD manifold’s suitability for classification tasks. Rather than simply comparing classifiers, our aim is to investigate the generality, adaptability, and complementarity of SPD representations across fundamentally different learning paradigms. The overall framework is illustrated in Fig. 1.
Overview of multiscale SPD manifold classification framework.
Data embedding into the SPD manifold
In this work, we utilize skeleton-based rehabilitation exercise data structured as multivariate time-series signals. Each record contains \(f\) features (e.g., 3D joint coordinates or orientation data) recorded over \(T\) time steps. This yields a data matrix \(X \in \mathbb {R}^{f \times T}\), where each row represents a feature and each column a time step.
Constructing the covariance matrix
To capture the statistical dependencies between features, we compute a covariance matrix. Covariance matrices are symmetric and positive definite by construction, making them well-suited for embedding into the SPD manifold.
Given two feature vectors \(X_i\) and \(X_j\), the sample covariance is computed as:
where \(X_i = (x_{i1}, x_{i2}, \dots , x_{iT})\) and \(X_j = (x_{j1}, x_{j2}, \dots , x_{jT})\) are the time series for features i and j, respectively, and \(\bar{x}_i = \frac{1}{T} \sum _{k=1}^{T} x_{ik}\), \(\bar{x}_j = \frac{1}{T} \sum _{k=1}^{T} x_{jk}\) are their means.
From the full data matrix \(X\), the complete covariance matrix \(C \in \mathbb {R}^{f \times f}\) is constructed as:
Each element captures the co-movement of two features over time. The resulting matrix is symmetric and positive definite, placing it naturally within the SPD manifold — a space of matrices that generalizes Euclidean geometry to account for curvature and structure in data.
Multi-scale covariance embedding
Rather than using a single covariance matrix computed over all features, we adopt a multi-scale strategy to better capture the structured nature of human motion. Specifically, we divide the body into three anatomical regions: arms, which include joints such as the wrist, elbow, and shoulder; legs, encompassing the ankle, knee, and hip joints; and the full body, which combines all features. For each of these regions, we compute a separate covariance matrix—denoted as \(C_{\text {arms}} \in \mathbb {R}^{f_a \times f_a}\), \(C_{\text {legs}} \in \mathbb {R}^{f_l \times f_l}\), and \(C_{\text {full}} \in \mathbb {R}^{f \times f}\), respectively. These three covariance matrices are then combined into a block-diagonal matrix defined as
This block-diagonal structure ensures that each sub-region contributes independently, preserving both local movement characteristics within the arms and legs as well as the global correlations across the entire body. Importantly, the resulting matrix \(C_{\text {multi}}\) remains symmetric positive definite, thus maintaining compatibility with Riemannian manifold operations.
KNN for SPD manifold
KNN is a widely used classification algorithm that assigns a label to a new data point based on the majority vote of its K-nearest neighbors in the feature space. However, when working with SPD matrices, traditional distance metrics like Euclidean distance fail to capture the manifold’s intrinsic geometric structure. To address this, we use distance metrics specifically designed for SPD matrices, which respect their positive definiteness and symmetry.
Given two SPD matrices A and B, the distance between them is computed using an SPD-specific metric described in subsection Distance Metrics on the SPD Manifold (e.g., Affine-invariant, Log-Euclidean, etc.). For each test sample, we compute its distance to all training samples in the SPD manifold, selecting the K-nearest neighbors based on the chosen metric. The details of the algorithm are described below in Algorithm 1.
KNN Classification on the SPD Manifold
Tangent space linear SVM
The manifold of symmetric positive definite (SPD) matrices, denoted \(S^{++}_n\), comprises all \(n \times n\) real symmetric matrices with strictly positive eigenvalues. This set forms a curved Riemannian manifold, meaning standard linear classification techniques such as SVM, which operate in flat Euclidean space, are not directly applicable.
In Euclidean space (\(\mathbb {R}^n\)), a linear SVMs defines a hyperplane of the form
where \(w\) is the weight vector and \(b\) is the bias. This model separates data points while maximizing the margin between classes. However, SPD matrices are \(n \times n\) objects that do not form a vector space, and \(S^{++}_n\) has positive curvature. This curvature means flat hyperplanes are undefined across the manifold, as the non-linear structure and lack of global linearity complicate the direct application of traditional SVMs. Specifically, there are no consistent, flat separators that can achieve linear separation.
To address this, we adopt a tangent space approach. Specifically, we fix the base point at the identity matrix \(I \in S^{++}_n\), and map each SPD matrix \(X_i \in S^{++}_n\) to the tangent space \(T_I S^{++}_n\) via the matrix logarithm:
where \(\log (X_i)\) denotes the matrix logarithm. This transformation flattens the manifold locally, converting each SPD matrix into a symmetric matrix in a Euclidean space.
The tangent space consists of symmetric matrices, which naturally supports the Frobenius inner product:
enabling a standard linear SVM formulation. In this space, we optimize a tangent vector \(V\) to define a hyperplane that maximizes the margin between classes. This approach simplifies the classification task by transforming it into a linear problem while still leveraging the structure of the SPD manifold.
The algorithm, which operates with the identity matrix as the base point, is outlined in Algorithm 2 and details the process of finding the tangent space weight vector V. Once V is learned, it is used for classification. The decision function for a new SPD matrix \(X'\) is given by:
The final class prediction is then obtained via:
This formulation simplifies SVM classification on \(S_n^{++}\) by working in the tangent space, achieving effective performance with reduced computational complexity.
This tangent space mapping, using the identity matrix as the base point and the Log-Euclidean metric, provides a straightforward and effective framework for classification. Besides this approach, we also explored alternative mappings and base points to evaluate their influence on classification performance. In particular, we considered the Affine-Invariant Riemannian Metric (AIRM), where the base point \(\mu\) is set to the Riemannian(Fréchet) mean36 of the training data, and the logarithm map is defined as
The Riemannian mean \(\mu\) under the affine-invariant metric is defined as:
where \(\mathcal {S}_{++}^n\) denotes the set of \(n \times n\) symmetric positive-definite matrices, and \(\log (\cdot )\) is the matrix logarithm. This formulation follows31.
Additionally, we included a Euclidean baseline where SPD matrices are centered by the element-wise mean
and used directly as features without any manifold mapping, i.e.,
Tangent Space Linear SPD SVM
Neural network-based classification for SPD manifold
After utilizing KNN and SVM for classification on the SPD manifold, we adopt a neural network-based approach to capture complex, high-dimensional patterns more effectively. While KNN and SVM can capture local patterns and decision boundaries by considering the proximity of SPD matrices in the manifold, they struggle with capturing complex, high-dimensional relationships. Additionally, traditional neural network architectures are typically designed for Euclidean spaces, making them unsuitable for directly processing SPD matrices.
To overcome these challenges, we use a Log-Euclidean mapping, which projects SPD matrices onto a Euclidean space while preserving their geometric properties. This transformation enables us to leverage the expressiveness and flexibility of neural networks, which are particularly well-suited for high-dimensional data. The overall neural network pipeline for SPD manifold data processing is illustrated in Fig. 2.
Logarithmic transformation for SPD matrices
Given an SPD matrix \(A\), the logarithmic map is defined as:
where \(A = U \Sigma U^T\) is the eigenvalue decomposition of \(A\), \(\Sigma\) is the diagonal matrix of eigenvalues, \(U\) is the orthogonal matrix of eigenvectors, and \(\log (\Sigma )\) is the element-wise logarithm of the eigenvalues in \(\Sigma\).
The result, \(\log (A)\), is a symmetric matrix representing the tangent space at the identity matrix, mapping the SPD matrix onto a Euclidean space, thereby facilitating the application of traditional neural network models.
Matrix to vector conversion
Once the logarithmic transformation is applied, the next step is to convert the resulting symmetric matrix into a vector. This conversion is necessary to enable the use of machine learning techniques that expect vectorized inputs. Let \(A\) be a symmetric positive definite (SPD) matrix of size \(d \times d\). After applying the logarithmic transformation, we obtain the matrix \(L(A) = \log (A)\), which is also symmetric and of size \(d \times d\). We vectorize the matrix \(L(A)\) as follows:
where \(y_{i,j}\) denotes the elements of \(L(A)\), with off-diagonal elements scaled by \(\sqrt{2}\) to maintain symmetry in the vectorized representation.
This transformation efficiently maps the SPD matrix to a one-dimensional vector suitable for neural network input, enabling the model to capture the complex relationships present in the original SPD data.
Neural network model proposed
After applying the Log-Euclidean mapping and vectorizing the tangent space representations, we employ a neural network architecture for classification. The vectorized covariance features are processed through two one-dimensional convolutional layers. The first layer uses 16 filters with a kernel size of 3, and the second uses 32 filters with the same kernel size. Each layer is followed by ReLU activation and batch normalization. This configuration was selected based on common practice and empirical tuning, offering a balance between representational capacity and computational efficiency. A max pooling layer with a kernel size of 2 and stride of 2 follows, reducing spatial dimensions while retaining salient features.
The resulting features are passed through three fully connected layers with 1024, 256, and 2 neurons, respectively. These layer sizes were chosen to progressively compress the feature space, allowing the network to model complex patterns while avoiding over parameterization. ReLU activations are used throughout to introduce non-linearity, and dropout with a rate of 0.3 is applied after the first fully connected layer to improve generalization. This architecture was iteratively refined to capture both local and global structure in the data effectively, enabling robust performance on the classification task.
Neural network pipeline for SPD manifold data processing.
Experiments and analysis
Dataset overview
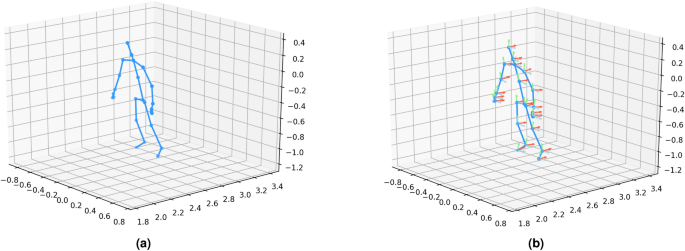
This study evaluates our method using three publicly available skeleton-based datasets: UI-PRMD37, KIMORE38, and EHE39. All three datasets provide two primary types of features: position and orientation of skeleton joints. These features are crucial for understanding the exercise performance of different populations, each presenting unique challenges in rehabilitation assessments. The position and orientation features are visualized in Fig. 3. An overview of the key characteristics of these datasets is provided in Table 1. The details of each dataset are described below.”
UI-PRMD Dataset contains 1326 exercise repetitions from 10 healthy subjects performing 10 rehabilitation exercises (e.g., side lunge, sit to stand, deep squat). Data was captured using Kinect v2 and Vicon motion capture. The datasets provide position and orientation features of skeleton joints. Subjects perform both correct and incorrect versions of exercises, simulating common errors in musculoskeletal rehabilitation. We use the Kinect v2 data for our experiments, following the consistent subset used in prior work to minimize errors.
KIMORE Dataset includes 2806 exercise repetitions from 78 subjects divided into three groups: Control Group - Experts (CG-E), Control Group - Non-Experts (CG-NE), and Group with Pain and Postural Disorders (GPP). Subjects perform five exercises (e.g., lifting arms, squatting) captured using the Kinect v2 sensor. The dataset includes RGB videos, depth videos, and skeleton joint positions and orientation information. It also includes clinical performance scores based on physician evaluations. This dataset offers a valuable benchmark for rehabilitation assessment, especially for subjects with motor dysfunctions.
EHE Dataset consists of 869 exercise repetitions from 25 elderly subjects performing six daily morning exercises (e.g., wave hands, hands up and down). Data was collected using Kinect v2 in a natural elderly home setting. The subjects had an average age of 68.4 years, with 10 diagnosed with Alzheimer’s disease (AD) at varying severity levels. The dataset provides position and orientation features of skeleton joints and is useful for analyzing exercise performance in elderly individuals, particularly those with AD.
KIMORE dataset visualization.
Experimental setup
Data division strategies
To evaluate the generalization capability of the proposed models, we adopt two data split strategies: cross-subject division and random division. In the cross-subject division, the training and testing sets are constructed from disjoint subject identities—i.e., actions performed by one group of subjects are used for training, and actions from different subjects are used for testing. This simulates real-world deployment on previously unseen individuals. In the random division, all samples are randomly partitioned into training and testing sets without considering subject identities. This setting evaluates the average-case performance of the model. Both strategies are commonly used in skeleton-based action recognition and are designed to test model robustness under different generalization conditions.
Parameter settings
We evaluate the performance of three classification methods: KNN on the SPD manifold, SVM on the SPD manifold, and a Neural Network in Euclidean space. The KNN and Neural Network models are applied to two types of covariance matrix representations. The first is a full-body covariance matrix, computed using all skeleton joints. The second is a multi-scale covariance matrix, which includes separate covariance matrices for arms, legs, and full-body movements, later combined into a larger representation (as detailed in the Methodology section). Due to computational constraints, SVM is evaluated only with the full-body covariance matrix. We next describe the specific hyperparameters and configurations used for each of the three methods.
KNN classification on the SPD manifold parameters We assessed a range of distance metrics across multiple categories for KNN classification on the SPD manifold. Cholesky-based metrics included the Cholesky distance29 and the Log-Cholesky distance34. Log-Euclidean metrics comprised the Log-Euclidean distance30 and the Log-det distance35. Additionally, we evaluated the Affine-invariant distance31, the Euclidean distance, and information-theoretic metrics, such as the Kullback-Leibler divergence32 and its symmetric variant33. We selected \(k=3\) neighbors for KNN, as preliminary testing across \(k \in {1, 3, 5, 7}\) showed that \(k=3\) consistently yielded competitive performance under both the cross-subject and random division protocols, effectively balancing bias and variance.
Tangent space SPD SVM parameters For the SVM classification in the tangent space of the SPD manifold, we configured the model with the following hyperparameters: a regularization parameter \(C=0.1\), a training duration of 150 epochs, a learning rate of 0.01, and a batch size of 8. These values were selected based on standard practices in prior work and preliminary tuning on a validation set, balancing training stability and generalization. To optimize convergence and prevent overfitting, we employed an early stopping patience of 20 epochs and implemented a learning rate reduction strategy with a step factor of 5.0, reducing the learning rate until a minimum value of \(1 \times 10^{-4}\) was reached. We explored three distance metrics for mapping the SPD manifold to the tangent space: the affine-invariant metric, the Log-Euclidean metric, and the Euclidean distance, evaluating their impact on classification performance.
Neural Network-Based Method Parameters For our neural network-based approach, we adopted a network architecture detailed in the methodology section. The model was trained using the Adam optimizer with a learning rate of 0.001 and a batch size of 8 over 50 epochs. These values were chosen based on commonly used defaults in similar settings and verified to provide stable convergence during initial experiments. To regularize the optimization process, we applied a weight decay of \(1 \times 10^{-4}\) and configured the optimizer’s beta parameters to [0.9, 0.999], balancing the influence of past gradients and squared gradients for stable convergence.
Results and comparison
This section presents the classification results for KNN on the SPD manifold, Tangent Space SVM, and a neural network in Euclidean space, using two covariance matrix representations: full-body and multi-scale (arms, legs, full-body). Experiments were conducted on three datasets: UI-PRMD, KIMORE, and EHE. The following subsections present the results for each method in detail.
SPD manifold KNN with different distance metrics
To assess the impact of multi-scale covariance matrices, we evaluated the KNN classifier using various distance metrics. The results are presented in Table 2. The goal of this experiment is to determine whether the multi-scale representation enhances the distinctiveness of covariance matrices, thereby improving classification performance.
The findings reveal the following trends:
-
The multi-scale representation consistently outperforms the full-body representation across all datasets, except when using Euclidean-based metrics, where it shows no advantage. This suggests that the multi-scale approach improves the distinctiveness of covariance matrices for most metrics.
-
Cholesky29 and Kullback-Leibler divergence32 metrics achieve higher accuracy in 5 out of 6 cases when using the multi-scale representation, indicating these distance metrics benefit from the finer granularity of motion captured by the multi-scale matrices.
-
The symmetric variant of the Kullback-Leibler divergence33 metric improves accuracy in 4 out of 6 cases with the multi-scale representation, further reinforcing the advantages of this approach for capturing motion patterns.
-
Log-Euclidean30, Log-det35, and Affine-invariant31 metrics achieve higher accuracy in all 6 cases with multi-scale representation, suggesting that these metrics are highly compatible with the multi-scale approach.
-
The Euclidean-based metric does not benefit from multi-scale representation, likely due to the inherent mismatch between Euclidean geometry and the SPD manifold, where Euclidean distance fails to capture the complex structure of the covariance matrices.
These results confirm that the multi-scale covariance representation better captures fine-grained motion patterns across different body parts, enhancing distinctiveness and improving classification accuracy.
We also observed that the random split outperformed the cross-subject split across all datasets. This can be attributed to the nature of the KNN classifier, which relies on distance-based measurements. In the random split scenario, parts of the same subject’s data can end up in both the training and test sets, resulting in more similar data for both training and testing. This similarity improves KNN’s performance. In contrast, the subject cross-split ensures that training and testing data come from different subjects, which introduces more variation and makes the task more challenging, thus leading to lower performance.
KNN, while effective with certain distance metrics, has inherent limitations. It relies on predefined distance measures, struggles with high-dimensional data, and cannot model complex, non-linear relationships. These weaknesses make it less suitable for capturing the intricate patterns in multi-scale covariance matrices.
For the KNN classifier, we evaluated all three feature types—position (Pos), orientation (Ori), and their combination (PO)—and reported the averaged results across all action classes. Due to space constraints and the large number of configurations involved (3 datasets \(\times\) 8 distance metrics), we present only the Pos-based results in the main paper. Detailed modality-specific results (Pos, Ori, PO) under the Log-Euclidean metric, which serves as a representative choice due to its consistent performance across datasets, are provided in Supplementary Tables S1.
Tangent space linear SPD SVM under different metrics
To further analyze the effectiveness of different metrics on SPD matrix classification, we evaluated a Tangent Space Linear SPD SVM. Unlike KNN, which relies on local distance comparisons, SVM is a more powerful classifier that can find complex decision boundaries, particularly when operating in the tangent space of SPD manifolds. By mapping SPD matrices to a vector space, SVM can leverage linear separation techniques to improve classification accuracy.
Table 3 presents the classification accuracy for the UI-PRMD, KIMORE, and EHE datasets under cross-subject and random division protocols. The Log-Euclidean metric consistently achieved the highest accuracy across all settings, demonstrating its effectiveness in capturing the intrinsic structure of SPD matrices while maintaining computational tractability. In contrast, the AIRM, despite its theoretical appeal, produced the lowest performance in nearly all cases, possibly due to sensitivity to class imbalance or numerical instability. The Euclidean baseline, lacking geometric modeling, sometimes outperformed AIRM, particularly under random division, indicating that simpler approaches can occasionally be more robust.
These findings suggest that geometry-aware mappings, especially Log-Euclidean, generally enhance classification performance when paired with discriminative classifiers such as SVM. However, method selection should be guided by the data and application context, motivating further investigation into metric suitability and robustness.
We report SVM results using the multi-scale covariance representation with position (Pos) features, as its superiority has been clearly demonstrated in the KNN experiments. Due to the computational expense of training SVMs across multiple settings, we did not include separate evaluations using the single-scale representation and instead focused on the multi-scale variant, which offers better discriminative power.
For completeness, we provide additional SVM results using all three input types: multi-scale Pos, multi-scale Ori, and their combination multi-scale PO. These results, computed under the Log-Euclidean metric, are included in Supplementary Table S2.
Neural network approach in Euclidean space
While KNN and our Tangent Space Linear SPD SVM have shown effectiveness, they face inherent limitations. KNN relies on predefined distance metrics and struggles in high-dimensional spaces, while the Tangent Space Linear SPD SVM assumes that SPD matrices can be well-separated using a linear decision boundary. This assumption may not hold for complex SPD data, restricting its ability to capture intricate patterns. To address these challenges, we turn to neural networks, which can learn non-linear patterns, handle high-dimensional data more effectively, and automatically extract relevant features. This makes them a more robust alternative for classifying SPD matrices, leading to improved performance.
As demonstrated in Table 4, for the UI-PRMD dataset, the classification results across both cross-subject and random division protocols highlight the superior performance of the PO method (Position and Orientation-based information), which achieves the highest average accuracy of 92.40% in the cross-subject protocol and 96.15% in the random division protocol. The Ori method (Orientation-based information) ranks second, with 90.84% accuracy in the cross-subject protocol and 95.61% in the random division protocol. The Pos method (Position-based information) shows slightly lower performance at 86.12% in the cross-subject protocol. Our proposed methods consistently outperform the state-of-the-art methods MLE-O and MLE-PO, with Ori and PO achieving the best results. Notably, our methods surpass the MLE methods in 8 out of 10 exercises for the cross-subject protocol and in 7 out of 10 exercises for the random division protocol.
As demonstrated in Table 5, for the KIMORE dataset, our methods consistently outperform the previous state-of-the-art methods, MLE-PO and MLE-O, across both the cross-subject and random division protocols. Our approach achieves an average accuracy of 85.18% in the cross-subject protocol and 96.69% in the random division protocol, surpassing MLE-PO and MLE-O in both protocols. Specifically, under the cross-subject protocol, our methods outperform MLE-PO and MLE-O in 5 out of 8 exercises, while in the random division protocol, our methods achieve superior performance in all 8 exercises compared to MLE-PO and MLE-O.
As demonstrated in Table 6, for the EHE dataset, the classification results indicate that while the Pos method performs well, the PO method achieves the best overall performance, particularly in the random division protocol (95.79% average accuracy). In the cross-subject protocol, Pos leads with the highest average accuracy (87.59%), followed closely by MLE-PO and MLE-O, with MLE-O showing slightly lower accuracy than both Pos and MLE-PO. In the random division protocol, PO outperforms all other methods, achieving the best results in five out of six exercises. While Pos also performs well, it falls short of PO, with MLE-PO and MLE-O showing similar performance, and MLE-O trailing slightly behind.
In conclusion, across the UI-PRMD, KIMORE, and EHE datasets, our methods consistently outperform the state-of-the-art MLE-PO and MLE-O methods. The PO method, which combines position and orientation information, proves to be more robust and generally achieves the best results across both protocols in all three datasets. Even in cases where it doesn’t provide the absolute best accuracy, it still delivers strong performance. Overall, our approach demonstrates significant improvements over MLE-PO and MLE-O, confirming its effectiveness and superiority in human activity recognition.
Comparison with state-of-the-art
To evaluate the effectiveness of our proposed method, we compare it against several state-of-the-art approaches on the UI-PRMD, KIMORE, and EHE datasets using the cross-subject evaluation protocol. The results, presented in Table 7, demonstrate the superior performance of our approach in various settings.
GCN 25 leverage spatial dependencies in skeleton-based action recognition by applying graph convolutions to model joint interactions. This method achieved moderate accuracy across datasets, with performance varying between position-based (Pos) and orientation-based (Ori) features. AGCN 41 Adaptive GCNs (AGCN) enhance GCN by introducing adaptive graph structures that can dynamically learn relations between joints. While this method improves upon standard GCN, it struggles with orientation-based representations, particularly on the EHE dataset. MS-G3D 42 Multi-Scale Graph 3D Convolution (MS-G3D) incorporates multi-scale graph convolutions to capture both local and global motion patterns. This approach outperforms AGCN and GCN, particularly in orientation-based features, but still falls short in fully leveraging positional and orientation cues. CTR-GCN 43 Channel-wise Topology Refinement GCNs (CTR-GCN) introduce an adaptive channel-wise topology refinement strategy to better capture motion dynamics. This approach yields significant improvements over previous methods, particularly in the UI-PRMD and EHE datasets, demonstrating strong generalization capabilities. In contrast to these graph-based methods, SPDNet 44 learns directly from Symmetric Positive Definite (SPD) matrices using Riemannian geometry-based layers, allowing the network to operate on manifold-structured data while preserving its non-Euclidean properties.
MLE-O 40 employs ensemble learning in a two-stage process: first, training on joint positions for action recognition to identify key neurons, then refining the model for action correctness using orientation information, with pointwise weight multiplication ensuring HAR-relevant neurons influence correctness evaluation. This strategy enhances integration of spatial and orientation cues, improving performance over standard graph-based methods. MLE-PO 16 builds upon MLE-O by further refining the second learning stage, incorporating both position and orientation information for action correctness assessment. This enhancement allows the model to leverage spatial and orientation cues more effectively, leading to improved accuracy over its predecessor.
Our method significantly outperforms existing approaches across all datasets. In the position and orientation (Pos+Ori) category, our model achieves the highest accuracy of 92.40% on UI-PRMD and 85.05% on KIMORE, surpassing MLE-PO 16 in each case. For the EHE dataset, while MLE-PO achieves a higher accuracy (86.38%), our method achieves the best performance in the position (Pos) category with 87.59%, highlighting the robustness of our approach in integrating both spatial and orientation-based features and leading to improved generalization across datasets.
As we observe, the state-of-the-art methods for this task are GCN-based, as these methods are particularly well-suited for modeling the geometric structures inherent in human motion data. This makes them the natural choice for comparison, as they achieve state-of-the-art performance on tasks involving manifold-aware representations. While attention-based models, such as Transformers, have shown strong performance in other contexts, they operate in Euclidean space and do not account for the underlying SPD manifold geometry. As such, they are not directly comparable in this setting. Adapting attention-based models to non-Euclidean domains is a promising direction for future work but is beyond the scope of this study.
Computational cost comparison
In this subsection, we evaluate the computational efficiency of our proposed CNN-based model in comparison to state-of-the-art GCN-based methods on the UI-PRMD dataset. When measuring training time for one fold of 5-fold cross-validation on a Tesla V100-PCIE GPU, our model demonstrates superior efficiency. With a batch size of 8, the MLE-O40 model, a graph-based approach leveraging both position and orientation (Pos+Ori) features, requires approximately 77 seconds to train a single fold. In contrast, our CNN-based model, incorporating the same Pos+Ori features, completes the training in just 46 seconds under identical conditions, highlighting its computational advantage.
As shown in Table 8, our model has a significantly higher parameter count (e.g., 267.2M for Pos+Ori features compared to 3.1M–6.2M for GCN-based methods). This is primarily due to the dense fully connected layers in our Conv + MLP model, where each connection has a separate weight, leading to a quadratic increase in parameters with input size. In contrast, GCNs share parameters across nodes and typically avoid large fully connected layers, making them more parameter-efficient, especially for graph-structured data. Additionally, as noted in27, certain concepts or objects recur across various mathematical domains and often carry substantial implications. For example, a system involving three identical globes in robotics: originally represented in a nine-dimensional Euclidean space, can alternatively be modeled on a three-dimensional orthogonal group manifold. In our approach, we transform the data onto a SPD manifold to better capture its intrinsic geometric structure while also reducing computational complexity through fewer floating-point operations.
Despite the higher parameter count, our model achieves substantially lower FLOPs—0.6G compared to 1.9G–12.2G for GCN-based methods. This difference stems from the complexity of graph-based operations. In GCNs, each node aggregates features from its neighbors, which involves multiple matrix multiplications or neighbor-to-node feature propagation. This process becomes computationally expensive, particularly in large or dense graphs, where node degree and graph size both affect the cost. On the other hand, our Conv + MLP model operates on fixed-size local regions (e.g., image patches) with more localized operations, resulting in fewer FLOPs per layer despite having more parameters. The global aggregation in GCNs increases computational cost significantly, making them more FLOP-intensive, even with fewer parameters.
Ablation studies
The contributions of each model component, as evaluated on the UI-PRMD, KIMORE, and EHE datasets, are detailed in Table 9.
-
Baseline Model: Our full approach embeds skeleton data onto the SPD manifold, employs a multi-scale strategy (arms, legs, whole body) to form a combined covariance matrix, and applies a Log-Euclidean mapping to project the data onto the tangent space. This configuration achieves a mean accuracy of 86.28%.
-
Without Multi-Scale: Removing the multi-scale component reduces the mean accuracy to 85.50% (a drop of 0.78%), demonstrating the benefit of capturing localized dynamics.
-
Without SPD Embedding: Excluding the SPD embedding significantly degrades performance, lowering the mean accuracy to 81.95% (a drop of 4.33%), with a dramatic decline in the UI-PRMD position data (from 86.12% to 69.17%).
-
Without Log-Euclidean Mapping: Omitting the Log-Euclidean mapping results in a moderate decrease to 84.78% (a drop of 1.5%), confirming its importance in linearizing the manifold structure for effective classification.
These ablation studies clearly demonstrate that each module contributes to the overall performance of the model. The SPD embedding is particularly critical, as evidenced by the substantial drop in accuracy when it is removed. The multi-scale strategy and log-Euclidean mapping, while offering relatively smaller improvements individually, still play a significant role in refining the discriminative power of the learned representations. Collectively, these components synergize to enhance the network’s ability to evaluate rehabilitation exercise correctness effectively.
Discussion
General-purpose manifold learning methods, such as t-SNE, Isomap, and UMAP, are widely used for nonlinear dimensionality reduction but often fail to model the underlying geometry of the data, treating it as generic point clouds in Euclidean space. This can lead to suboptimal representations for data with specialized structures. To address this, our method leverages the Riemannian geometry of SPD manifolds, which is well-suited for modeling covariance matrices prevalent in motion and sensor-based data. Equipped with geometric tools like logarithmic and exponential maps, the SPD manifold preserves the intrinsic structure of second-order statistics, making it ideal for capturing the variability of rehabilitation movement patterns. Unlike other non-Euclidean spaces, such as Grassmannian manifolds (suited for linear subspaces) or hyperbolic manifolds (designed for hierarchical data), the SPD manifold provides a more precise and effective framework for this application45,46.
Although this study focuses on performance in terms of computational complexity and accuracy, real-world rehabilitation applications demand models that can run efficiently on resource-constrained devices. When using position features from the UI-PRMD dataset, the current model requires approximately 2086 MB of GPU memory during evaluation, as reported by nvidia-smi, suggesting it is feasible for deployment on modern edge devices with CUDA support. However, the SPD manifold representation in this work is based on complete action sequences, which may limit its responsiveness in real-time scenarios. A promising direction for future work is to incrementally accumulate the covariance matrix over shorter time windows, enabling early-stage classification and progressive evaluation as the movement unfolds. This would allow for more interactive, real-time feedback, which is particularly valuable in rehabilitation contexts.
In this work, the multi-scale covariance embedding is constructed using three intuitive partitions: arms, legs, and full body. This grouping reflects a straightforward anatomical structure, where limbs are naturally treated as coherent units in human motion. The choice was based on simplicity and interpretability, rather than dataset constraints or clinical guidelines. However, we acknowledge that this is just one of many possible partitioning strategies. Future work could explore alternative groupings, such as data-driven clustering or functional segmentation, to assess their potential impact on model performance and insight generation. A potential direction for future work is to analyze which joints or covariance components contribute most to the model’s decisions, in order to enhance transparency and support more informed use in rehabilitation settings.
We adopt the Log-Euclidean Riemannian metric to map SPD matrices to the tangent space, as it strikes a balance between mathematical rigor and computational tractability. Unlike AIRM—which offers desirable invariance properties but at high computational cost—the Log-Euclidean approach treats the space of SPD matrices as a Lie group under the matrix logarithm, resulting in a vector space structure that simplifies both implementation and optimization. Prior work has demonstrated its empirical effectiveness across various domains30, and we find it especially suitable for our setting, where fast and stable optimization is critical for learning from high-dimensional covariance descriptors.
In our Tangent Space Linear SVM experiments, the Log-Euclidean mapping consistently outperformed both AIRM and the Euclidean baseline. Although AIRM respects the manifold geometry more precisely, its use of the Riemannian mean as the base point may introduce bias in imbalanced datasets and lead to numerical instability. Interestingly, even on the balanced dataset, AIRM still underperformed, suggesting that other factors such as data variability or curvature sensitivity may also contribute. The Euclidean baseline, which ignores manifold geometry entirely, performed less effectively overall, further supporting the value of geometry-aware mappings in SPD classification.
The three classification pipelines explored in this study—KNN, tangent-space SVM, and neural network—are intentionally selected to reflect a range of modeling capacities and assumptions. KNN relies purely on manifold geometry and does not involve any training, SVM introduces supervised learning with linear decision boundaries in the tangent space, and the neural network approach enables hierarchical and nonlinear feature learning directly on the SPD manifold. While we do not fuse these models in this work, they provide complementary perspectives on how SPD representations support classification tasks, highlighting the manifold’s versatility across different learning paradigms. Future work could investigate hybrid strategies that combine these models, such as ensemble learning or multi-branch architectures, to further enhance robustness and generalization.
Beyond these three paradigms, other prototype-based strategies have also been developed. For example, Tang et al.47 proposed generalized learning Riemannian space quantization (GLRSQ), which learns discriminative prototypes directly on the SPD manifold using Riemannian distances. Compared with our Riemannian KNN classifier, which is instance-based and requires no training, GLRSQ produces a compact set of prototypes, yielding stronger generalization and more efficient inference once trained. A promising future direction is to combine these two paradigms—for instance, by initializing prototypes from nearest neighbors or refining them adaptively—thereby balancing the simplicity of non-parametric methods with the discriminative power of prototype learning.
Conclusion
In this study, we presented a novel approach for assessing rehabilitation exercises using an SPD manifold-based framework. By integrating multi-scale features and leveraging manifold-based learning techniques, our method achieved significant performance improvements over traditional approaches. Extensive experiments across three benchmark datasets (Kimore, UI-PRMD, and EHE) demonstrated the robustness and reliability of our model, with notable accuracy improvements in both cross-subject and random train-test protocols.
From a clinical perspective, our method could improve patient outcomes by providing clinicians with timely feedback on exercise performance. This would not only help ensure correct execution of rehabilitation exercises but also support personalized rehabilitation plans. Ultimately, integrating such a system into clinical practice could enhance patient recovery, reduce the burden on healthcare providers, and improve the efficiency of rehabilitation programs.
Looking ahead, future work will focus on deploying the model on mobile devices for timely assessments, making it more accessible and practical for everyday rehabilitation. Further optimizations will also target efficient deployment, expanding clinical applications, and enhancing the interpretability of the model to foster greater trust among clinicians and patients.
Data availability
No new data were generated or analysed during this study. All data used are publicly available from the following GitHub repositories: https://github.com/bruceyo/EGCN and https://github.com/bruceyo/egcnplusplus. Further details are provided in the Methods and Supplementary Information.
References
Organization, W. H. Rehabilitation. (accessed 20 Feb 2025). https://www.who.int/health-topics/rehabilitation (2025).
Nguyen, S. M., Devanne, M., Remy-Neris, O., Lempereur, M. & Thepaut, A. A medical low-back pain physical rehabilitation dataset for human body movement analysis. arXiv preprint arXiv:2407.00521 (2024).
Kim, D.-W. et al. Automatic assessment of upper extremity function and mobile application for self-administered stroke rehabilitation. IEEE Trans. Neural Syst. Rehabil. Eng. 32, 652–661 (2024).
Sardari, S. et al. Artificial intelligence for skeleton-based physical rehabilitation action evaluation: A systematic review. Comput. Biol. Med. 158, 106835 (2023).
Kryeem, A. et al. Action assessment in rehabilitation: Leveraging machine learning and vision-based analysis. Comput..Vis. Image Underst. 251, 104228 (2025).
Kourbane, I., Papadakis, P. & Andries, M. Optimized assessment of physical rehabilitation exercises using spatiotemporal, sequential graph-convolutional networks. Comput. Biol. Med. 186, 109578 (2025).
Clark, R. A. et al. Validity of the microsoft kinect for assessment of postural control. Gait Posture 36, 372–377 (2012).
Zhao, W., Lun, R., Espy, D. D. & Reinthal, M. A. Rule based realtime motion assessment for rehabilitation exercises. In 2014 IEEE Symposium on Computational Intelligence in Healthcare and e-health (CICARE), 133–140 (IEEE, 2014).
Su, C.-J., Chiang, C.-Y. & Huang, J.-Y. Kinect-enabled home-based rehabilitation system using dynamic time warping and fuzzy logic. Appl. Soft Comput. 22, 652–666 (2014).
Hu, M.-C. et al. Real-time human movement retrieval and assessment with Kinect sensor. IEEE Trans. Cybern. 45, 742–753 (2014).
Mukherjee, P. & Roy, A. H. A deep learning-based comprehensive robotic system for lower limb rehabilitation. Biomed. Signal Process. Control 100, 107178 (2025).
Zhou, C., Feng, D., Chen, S., Ban, N. & Pan, J. Portable vision-based gait assessment for post-stroke rehabilitation using an attention-based lightweight cnn. Expert Syst. Appl. 238, 122074 (2024).
Zaher, M., Ghoneim, A. S., Abdelhamid, L. & Atia, A. Unlocking the potential of rnn and cnn models for accurate rehabilitation exercise classification on multi-datasets. Multimed. Tools Appl. 84(3), 1261–1301 (2025).
Carneros-Prado, D. et al. Synthetic 3d full-body skeletal motion from 2d paths using rnn with lstm cells and linear networks. Comput. Biol. Med. 180, 108943 (2024).
Deb, S., Islam, M. F., Rahman, S. & Rahman, S. Graph convolutional networks for assessment of physical rehabilitation exercises. IEEE Trans. Neural Syst. Rehabil. Eng. 30, 410–419 (2022).
Bruce, X., Liu, Y., Chan, K. C. & Chen, C. W. Egcn++: A new fusion strategy for ensemble learning in skeleton-based rehabilitation exercise assessment. IEEE Trans. Pattern Anal. Mach. Intell. 46(9), 6471–85 (2024).
Clark, R. A., Pua, Y.-H., Bryant, A. L. & Hunt, M. A. Validity of the Microsoft Kinect for providing lateral trunk lean feedback during gait retraining. Gait Posture 38, 1064–1066 (2013).
Zhao, W., Reinthal, M. A., Espy, D. D. & Luo, X. Rule-based human motion tracking for rehabilitation exercises: realtime assessment, feedback, and guidance. IEEE Access 5, 21382–21394 (2017).
Zhao, W., Yang, S. & Luo, X. Towards rehabilitation at home after total knee replacement. Tsinghua Sci. Technol. 26, 791–799 (2021).
Benetazzo, F. et al. Low cost rgb-d vision based system for on-line performance evaluation of motor disabilities rehabilitation at home. In Proceedings of the 5th Forum Italiano on Ambient Assisted Living For ItAAL (IEEE, 2014).
Osgouei, R. H., Soulsbv, D. & Bello, F. An objective evaluation method for rehabilitation exergames. In 2018 IEEE games, entertainment, media conference (GEM), 28–34 (IEEE, 2018).
Li, Q. et al. A motion recognition model for upper-limb rehabilitation exercises. J. Ambient Intell. Humaniz. Comput. 14, 16795–16805 (2023).
Liao, Y., Vakanski, A. & Xian, M. A deep learning framework for assessing physical rehabilitation exercises. IEEE Trans. Neural Syst. Rehabil. Eng. 28, 468–477 (2020).
Jun, K., Lee, S., Lee, D.-W. & Kim, M. S. Deep learning-based multimodal abnormal gait classification using a 3d skeleton and plantar foot pressure. IEEE Access 9, 161576–161589 (2021).
Yan, S., Xiong, Y. & Lin, D. Spatial temporal graph convolutional networks for skeleton-based action recognition. In Proceedings of the AAAI conference on artificial intelligence, vol. 32 (2018).
He, T., Chen, Y., Wang, L. & Cheng, H. An expert-knowledge-based graph convolutional network for skeleton-based physical rehabilitation exercises assessment. IEEE Trans. Neural Syst. Rehabil. Eng. 32, 1916–1925 (2024).
Snášel, V., Kong, L. & Das, S. From constraints fusion to manifold optimization: A new directional transport manifold metaheuristic algorithm. Inf. Fusion 113, 102596 (2025).
Bendokat, T., Zimmermann, R. & Absil, P.-A. A Grassmann manifold handbook: Basic geometry and computational aspects. Adv. Comput. Math. 50, 6 (2024).
Dryden, I. L., Koloydenko, A. & Zhou, D. Non-Euclidean statistics for covariance matrices, with applications to diffusion tensor imaging. Ann. Appl. Stat. 3(3), 1102–1123 (2009).
Arsigny, V., Fillard, P., Pennec, X. & Ayache, N. Geometric means in a novel vector space structure on symmetric positive-definite matrices. SIAM J. Matrix Anal. Appl. 29, 328–347 (2007).
Moakher, M. A differential geometric approach to the geometric mean of symmetric positive-definite matrices. SIAM J. Matrix Anal. Appl. 26, 735–747 (2005).
Kullback, S. & Leibler, R. A. On information and sufficiency. Ann. Math. Stat. 22, 79–86 (1951).
Jeffreys, H. An invariant form for the prior probability in estimation problems. Proc. R. Soc. Lond. Ser. Math. Phys. Sci. 186, 453–461 (1946).
Lin, Z. Riemannian geometry of symmetric positive definite matrices via Cholesky decomposition. SIAM J. Matrix Anal. Appl. 40, 1353–1370 (2019).
Dhillon, I. S. & Tropp, J. A. Matrix nearness problems with Bregman divergences. SIAM J. Matrix Anal. Appl. 29, 1120–1146 (2008).
Barachant, A., Bonnet, S., Congedo, M. & Jutten, C. Multiclass brain-computer interface classification by Riemannian geometry. IEEE Trans. Biomed. Eng. 59, 920–928 (2011).
Vakanski, A., Jun, H.-P., Paul, D. & Baker, R. A data set of human body movements for physical rehabilitation exercises. Data 3, 2 (2018).
Capecci, M. et al. The kimore dataset: Kinematic assessment of movement and clinical scores for remote monitoring of physical rehabilitation. IEEE Trans. Neural Syst. Rehabil. Eng. 27, 1436–1448 (2019).
Bruce, X., Liu, Y., Chan, K. C., Yang, Q. & Wang, X. Skeleton-based human action evaluation using graph convolutional network for monitoring Alzheimer’s progression. Pattern Recognit. 119, 108095 (2021).
Yu, B. X., Liu, Y., Zhang, X., Chen, G. & Chan, K. C. Egcn: An ensemble-based learning framework for exploring effective skeleton-based rehabilitation exercise assessment. EGCN: An Ensemble-based Learning Framework for Exploring Effective Skeleton-based Rehabilitation Exercise Assessment 3681–3687 (2022).
Shi, L., Zhang, Y., Cheng, J. & Lu, H. Two-stream adaptive graph convolutional networks for skeleton-based action recognition. In Proceedings of the IEEE/CVF conference on computer vision and pattern recognition, 12026–12035 (2019).
Liu, Z., Zhang, H., Chen, Z., Wang, Z. & Ouyang, W. Disentangling and unifying graph convolutions for skeleton-based action recognition. In Proceedings of the IEEE/CVF conference on computer vision and pattern recognition, 143–152 (2020).
Chen, Y. et al. Channel-wise topology refinement graph convolution for skeleton-based action recognition. In Proceedings of the IEEE/CVF international conference on computer vision, 13359–13368 (2021).
Huang, Z. & Van Gool, L. A riemannian network for spd matrix learning. In Proceedings of the AAAI conference on artificial intelligence, vol. 31 (2017).
Wang, R., Wu, X.-J., Chen, Z., Xu, T. & Kittler, J. Dreamnet: A deep riemannian manifold network for spd matrix learning. In Proceedings of the Asian conference on computer vision, 3241–3257 (2022).
Wang, R., Wu, X.-J., Chen, Z., Hu, C. & Kittler, J. Spd manifold deep metric learning for image set classification. IEEE Trans. Neural Netw. Learn. Syst. 35(7), 8924–38 (2024).
Tang, F., Fan, M. & Tiňo, P. Generalized learning Riemannian space quantization: A case study on Riemannian manifold of spd matrices. IEEE Trans. Neural Netw. Learn. Syst. 32, 281–292 (2020).
Acknowledgements
This scientific result is part of the CLARA project that has received funding from the European Union’s HORIZON EUROPE research and innovation programme under Grant Agreement No 101136607. The authors gratefully acknowledge financial support ROBOPROX of No. CZ.02.01.01/00/22 008/0004590 by Ministry of Education, Youth, and Sports and REFRESH of No. CZ.10.03.01/00/22_/0000048 by European Union.
Author information
Authors and Affiliations
Contributions
Z.B. and L.K. conceived the study and developed the methodology. V.S. supervised the work and revised the manuscript. C.G. and S.M. contributed to analysis and review. B.V. and J.-S.P. assisted with conceptualization and manuscript editing. All authors reviewed and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Bai, Z., Snášel, V., Grosan, C. et al. Multiscale SPD manifold learning for rehabilitation exercise evaluation. Sci Rep 15, 39403 (2025). https://doi.org/10.1038/s41598-025-19495-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-19495-5