Abstract
One such prevalent kind of cancer among women is cervical cancer (CC). Fatality rates and incidence are progressively increasing, mainly in developing countries, due to a lack of experienced specialists, inadequate public awareness, and limited screening facilities. Nevertheless, CC cells exhibit composite textural features, and smaller changes among dissimilar cell subcategories result in greater challenges for the higher-accuracy screening of CC. This systematic analysis aims to assess the predictive value of artificial intelligence (AI) technologies for diagnosing, screening, and predicting CC and precancerous lesions. Deep learning (DL) and AI generally have a positive impact on computer-aided clinical diagnosis, particularly with the increasing accessibility of larger amounts of medical data that can aid AI methods in achieving high performance on various medical tasks. In this paper, a Fusion of Advanced Feature Reduction and Deep Representation Learning Approaches for Cervical Precancerous Lesion Classification (FAFRDRL-CPLC) technique using biomedical image analysis is proposed. The primary purpose of the FAFRDRL-CPLC technique is to serve as a valuable tool for assisting clinicians in the initial study and treatment planning of cervical precancerous lesions. Initially, the FAFRDRL-CPLC approach applies an anisotropic diffusion filtering (ADF) method for pre-processing to reduce noise while preserving crucial edges and lesion details. Furthermore, the fusion of advanced feature reduction models, such as the maximally scalable vision transformer (MaxViT-v2), the simple framework for contrastive learning of visual representations (SimCLR), and the Twins-spatially separable vision transformer (Twins-SVT) models, is employed to capture diverse and complementary representations from the pre-processed images. Finally, the stacked auto-encoder (SAE) classifier is utilized for the precancerous lesion detection process. The FAFRDRL-CPLC method is examined through experimentation using the Malhari dataset. The comparison study of the FAFRDRL-CPLC method demonstrated a superior accuracy value of 98.62% over existing approaches.
Similar content being viewed by others
Introduction
CC is the 2nd most prevalent cancer, which affects the reproductive system of females1. Even now, it remains to result in high illness and death rates in developing nations like China. This has a significant impact on the overall well-being and quality of life of women. CC begins in cells lining the cervix, which is the lower segment of the womb or uterus2. The cervix links the uterus’s upper portion, in which a fetus forms, to the birth canal or vaginal passage. Cancer arises when body cells start multiplying without control. To better understand how cancer begins and spreads, additional investigation and research may be required3. The cervix consists of two distinct regions, each lined by a separate set of cell types4. The endocervix is the cervical opening that leads to the uterus and is lined with gland-forming cells. The exocervix, is the cervix’s outer surface, seen at a pelvic inspection, and is lined with squamous cells5. It shifts position over a woman’s lifespan, particularly during ageing or after childbirth, and CC arises from cells located in this area. These cells don’t instantly turn into cancerous ones. Instead, cervical cells usually undergo a slow transition from normal to abnormal changes, known as precancerous6. In several developing nations, colposcopy-directed biopsy is frequently employed for prognosis, with people demonstrating positive cytology or human papillomavirus (HPV) testing outcomes preferred for colposcopy7.
Precise colposcopic prognosis needs doctors to accurately detect acetowhite epithelium features, an ability that deeply depends on medical practice. In areas with inadequate healthcare resources, a lack of skilled doctors and extensive screening processes poses a significant difficulty8. Computer-aided diagnosis (CAD) has made substantial progress due to advancements in AI, enhancing symptomatic reliability and precision while reducing the workload of clinical staff. Investigators are developing novel tools to achieve more precise, faster, and automated CC diagnosis9. Currently, DL has become increasingly prevalent in the medical domain. After its introduction, it has demonstrated strong potential in domains such as natural language processing, image recognition, and speech recognition10. Currently, numerous researchers have focused on examining the use of DL to address medical issues11. From tumour to retinal illnesses, from tissue pathology image classification to cancer metastasis recognition, convolutional neural networks (CNNs) have witnessed notable accomplishments in the medical image processing domain2.
In this paper, a Fusion of Advanced Feature Reduction and Deep Representation Learning Approaches for Cervical Precancerous Lesion Classification (FAFRDRL-CPLC) technique using biomedical image analysis is proposed. The key contributions of this article are summarised as demonstrated:
-
The ADF method is applied in the pre-processing stage to reduce the image noise while preserving crucial edge data effectively. This model also facilitates accurate feature extraction by enhancing the clarity of cervical cell structures. The technique enhances the quality of input data for downstream analysis by eliminating irrelevant discrepancies, thereby contributing to improved model performance.
-
The feature reduction model is proposed by integrating three DL methods, namely MaxViT-v2, SimCLR, and the Twins-SVT model. This fusion enables the extraction of compact yet highly discriminative features, while also improving the model’s capability to capture multi-scale and contrastive representations, thereby enhancing classification accuracy while mitigating computational complexity.
-
The SAE classifier is employed for detecting the precancerous lesions, enabling effective learning of hierarchical feature representations from mitigated feature vectors. Its deep architecture helps capture intrinsic data patterns, thereby enhancing classification precision. The model also improves the technique’s capability to distinguish subtle variations between lesion classes, supporting early and accurate diagnosis.
-
The novelty of the FAFRDRL-CPLC model lies in the integration of advanced ViT models, namely MaxViT-v2, SimCLR, and Twins-SVT, for effective feature reduction, combined with ADF for noise-free pre-processing. Unlike conventional methods, the framework captures both spatial and contrastive representations while reducing redundancy. The use of an SAE for final classification additionally improves diagnostic accuracy by learning deep, nonlinear feature hierarchies.
Literature survey on cervical precancerous lesion detection
Nurmaini et al.12 proposed an automated cervical precancerous lesion detection system utilizing You Only Look Once version 8 (Yolov8) and Slicing Aided Hyper Inference (SAHI) techniques. Wubineh, Jeleń, and Rusiecki13 improved cervical cancer cell image classification by utilizing the Deep Convolutional Generative Adversarial Network (DCGAN) technique for synthetic image generation and augmentation. Various DL techniques comprising CNN, VGG16, MobileNet, ResNet50V2, InceptionV3, and Xception, are used for improving the accuracy of classification on Pap smear images. Khan et al.14 employed a metaheuristic technique, particle swarm optimization (PSO), to enhance hierarchical feature learning in the Xception approach for cervigram image classification into normal and cancerous classes. With the utilization of PSO, this method improves the feature extraction process, whereas Xception’s DL abilities enable complex feature extraction, facilitating accurate distinction between normal and cancerous cervigrams. The authors leverage residual blocks alongside the Xception pre-trained model to enable faster convergence in training owing to its capability for capturing and refining intricate hierarchical features. Himabindu et al.15 presented a Leveraging Swin Transformer with an Ensemble of DL Methods for CC Screening (LSTEDL-CCS) approach. This approach intends to identify and classify CC. Then, the ST approach is deployed for extracting features. For cancer identification, the ensemble learning technique is implemented using a DBN, BiGRU, and AE. Allogmani et al.16 presented an Enhanced Cervical Precancerous Lesions Detection and Classification employing the Archimedes Optimisation Algorithm with TL (CPLDC-AOATL) model. This model focuses on diagnosing CC via medical imaging. Moreover, this model utilizes the InceptionResNet-v2 model for feature extraction, and AOA selects the hyperparameters. Abinaya and Sivakumar17 proposed a cervical cancer classification system that integrates a 3D CNN with a Vision Transformer (ViT), followed by a 3D feature pyramid network (FPN), a 3D squeeze-and-excitation (SE) block, and a kernel extreme learning machine (KELM) for accurate multi-class classification. Sholik et al.18 presented a cervical cancer classification technique that utilizes a CNN model for feature extraction, a linear discriminant analysis (LDA) technique for feature reduction, and classification using support vector machines (SVM), multi-layer perceptrons (MLP), and K-nearest neighbours (K-NN) across diverse datasets. Attallah19 proposed an effective CAD model named “CerCanNet” for automated CC diagnosis. This model utilizes three lightweight CNNs with small parameters and fewer deep layers, namely ResNet18, DarkNet19, and MobileNet, to decrease classification complexity. Additionally, it involves transfer learning (TL) to extract deep features from the final three CNN layers. It also merges the acquired features through multiple CNN layers. Song et al.20 suggested an African Vulture Optimiser Algorithm with DL-based CC Detection (AVOADL-CCD) method. Additionally, this method employs the Capsule Networks (CapsNet) paradigm to derive feature vectors. The AVOA is used for the parameter-tuning procedure. For the automatic CC detection and classification, an ANFIS methodology is applied.
Chauhan et al.21 proposed a model by utilizing the Hybrid Deep Feature Concatenated Network (HDFCN) approach integrated with fine-tuned DL models such as Visual Geometry Group 16 (VGG-16), Residual Network 152 (ResNet-152), and Densely Connected Convolutional Networks 169 (DenseNet-169) models. Chauhan et al.22 presented a model by utilizing a hybrid learning network (HLN) model. This technique integrates progressive resizing, transfer learning with fine-tuned ResNet-152 and VGG-16 models, and principal component analysis (PCA) for efficient extraction and dimensionality reduction. ML classifiers, such as support vector machines (SVMs) and random forests (RFs), are also used in conjunction with majority voting to enhance classification accuracy and reliability. Chauhan and Singh23 evaluated the classification performance of CNN models with varying convolutional channel depths for the multi-class classification of liquid-based cytology (LBC) whole-slide images (WSI). Bilal, Hekmat, and Khan24 improved the accuracy of cervical cancer cell classification by utilizing an optimized ensemble method that integrates multiple pre-trained CNN models, including DenseNet169, MobileNetV2, and DenseNet201. Grid search is utilized to optimally weight each base model, improving overall diagnostic performance on the Mendeley Liquid-Based Cytology (LBC) dataset. Chauhan and Singh25 evaluated and compared the efficiency of eight ML methods, including SVM, naïve bayes (NB), LDA, logistic regression (LR), kNN, decision tree (DT), MLP, and RF, for accurate detection of malignant cervical cancer. Raza et al.26 proposed a model that integrates a Neural Feature Extractor (NFE) based on the pre-trained VGG16 model with the automatic feature interaction learning (AutoInt) technique. The kNN, Light Gradient Boosting Machine (LGBM), and Extra Trees (ET) approaches are utilized for feature extraction. Chauhan and Singh27 introduced a model by utilizing the Synthetic Minority Oversampling Technique (SMOTE) technique, comprising various ML models such as RF, SVM, DT, and Recursive Feature Elimination (RFE) techniques. Gangrade et al.28 proposed a model by utilizing an ensemble of DL models comprising CNN, AlexNet, and SqueezeNet to form a robust ensemble model that improves diagnostic accuracy. Abd-Alhalem et al.29 introduced a technique that integrates bilinear pooling within CNNs and addresses the high computational complexity through a Random Projection-based Bilinear CNN (RP-BCNN). This approach integrates dyadic feature extraction and dimensionality reduction to achieve both improved accuracy and faster processing times in multi-class and dual-label classification tasks. Borse and Yadav30 presented a technique by integrating advanced image processing with ML techniques. By utilizing segmentation, feature extraction, and classification, the approach employs ML models to differentiate between normal and cancerous cervical tissues during colposcopic evaluation. Comparison analysis of the existing Cervical Precancerous Lesion Classification in Table 1.
Limitations and research gaps in clinical deployment and model efficiency
Although existing studies are effective in diagnosing cancerous cells, they are ineffective in diverse areas, such as dependence on intrinsic DL models, resource-constrained environments, and the need for extensive computational resources. Furthermore, various models focus on classification accuracy but overlook the importance of real-time applicability and interpretability, which are essential for clinical adoption. Diverse techniques suffer from imbalanced data issues that are insufficiently addressed, resulting in biased predictions. There is also a lack of standardized datasets and consistent evaluation metrics, which complicates performance comparisons across studies. Moreover, various studies do not adequately explore the potential of lightweight models or multimodal data integration, which is particularly suitable for mobile or edge devices. The research gap lies in addressing these issues to achieve efficient, explainable, and scalable solutions that balance accuracy with clinical deployment feasibility and robustness in real-world scenarios.
Algorithm and model design
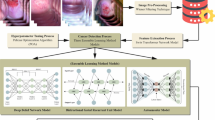
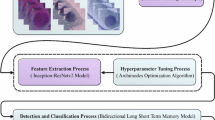
In this paper, an FAFRDRL-CPLC approach using biomedical image analysis is proposed. The primary purpose of the FAFRDRL-CPLC approach is to serve as a valuable device for supporting clinicians in the initial analysis and treatment planning of cervical precancerous lesions. It involves three distinct processes: pre-processing, feature selection, and precancerous lesion detection. Figure 1 exemplifies the complete procedure of the FAFRDRL-CPLC technique.
Complete process of FAFRDRL-CPLC technique.
Noise reduction using ADF technique
At the initial stage, the FAFRDRL-CPLC approach applies ADF to cervical images for pre-processing, effectively reducing noise while preserving essential edges and lesion details31. This model effectively smooths biomedical images while preserving crucial filters, such as Gaussian or median filtering. This technique prevents the fine structural data loss and performs selective smoothing based on image gradients. This methodology is also beneficial in cervical cytology images, where subtle features are significant for diagnosis. ADF mitigates irrelevant pixel-level discrepancies that could mislead DL techniques, thereby improving the quality of the feature extraction. Its edge-preserving nature ensures that significant lesion contours remain intact, assisting in better classification accuracy. Overall, ADF presents a balanced trade-off between denoising and structural preservation, making it a more suitable choice than conventional filtering techniques.
ADF is employed to enhance the image, and this pre-processing organizes the image for subsequent stages in analysis, such as tumour classification and detection. ADF is utilized to improve the image with specific parameters, such as iteration counts, Conductance parameter, Diffusion scheme, and Time step, thereby managing the details of the filtering process.
Employing AD to suppress or enhance specific features in the image contributes to removing features by denoising the image and improving the edge. It maintains edges while decreasing noise. Here, the image intensity modifies through time \(\:\frac{\partial\:,\:I}{\partial\:t}\) as a function of the gradient of the image \(\:\nabla\:I\) and diffusion coefficient \(\:c.\) \(\:I\) represent the input image, \(\:t\) denotes time, \(\:r\) depicts the gradient operator, and \(\:c(x,y,t)\) refers to the diffusion coefficient that manages the rate of diffusion depending on the local image gradient. Additionally, the discretized equation for ADF, employed in restricted variances for enhancing image denoising and edge-preserving smoothing, is given in Eq. (3).
Now \(\:\nabla\:t\) indicates the time step, \(\:\nabla\:\) denotes the discrete gradient operator, and \(\:{I}_{i,j}^{n}\) refers to the pixel value at position \(\:(i,j)\) in the image at time \(\:n\), typically utilizing central differences. \(\:c\) represents the diffusion coefficient function that manages the rate of diffusion depending on the gradient magnitude.
Fusion of feature dimensionality reduction models
Next, the fusion of advanced feature reduction models, such as MaxViT-v2, SimCLR, and Twins-SVT, is deployed to capture diverse and complementary representations from the pre-processed images. The fusion model is chosen for its strengths in capturing diverse image representations. The MaxViT-v2 method outperforms in extracting multi-scale spatial features with high scalability. At the same time, the SimCLR model utilizes a contrastive learning approach for learning robust and invariant feature embeddings without requiring labelled data. Additionally, the Twins-SVT technique effectively models both local and global dependencies; hence, incorporating these techniques improves the capability of capturing fine-grained and global contextual data, resulting in richer and more discriminative feature sets. This fusion model also mitigates redundancy and enhances the quality of the features, compared to conventional single-model techniques, thus improving the overall computational efficiency and classification performance. This multi-model integration addresses limitations in capturing intrinsic biomedical image discrepancies effectively.
MaxViT-v2 model
Transformers are DNNs primarily applied in the natural language processing (NLP) field32. Due to their strong capabilities, transformer-based methods have frequently been implemented as superior to or on par with other types of networks, such as RNNs and CNNs, in visual comparative analysis. The ViT splits all images into smaller patches. These patches are expanded based on their position order, and the particular dimensions characterize all patches. A position encoder is applied in \(\:ViT\) to maintain the patch sequence, allowing the method to learn the relationship between all patches. This method utilizes the self-attention mechanism (SAM) to recognize the context amongst removed pixels within the image.
CNNs utilize convolutional layers to seize spatial orders in images. CNNs have a fat order, which means higher, global features are characterized by progressing to the top layers. Therefore, CNNs are capable of working quickly and efficiently on small datasets and are primarily effective at identifying local patterns. They are extensively applied using their structures enhanced for TL. On the other hand, this model can learn more complex relationships and reach comparatively higher precisions after being trained on larger datasets with higher computing capacity.
In this work, an image classification method is advanced to implement the classification of PCa utilizing this structure. Unlike conventional CNN-based classification techniques, this method splits the images into smaller patches, utilizes all patches as input elements, and models the relationships between all patches using one another. Initially, the image data are normalized and rescaled to the stated dimensions of \(\:150\text{x}150\) pixels and then separated into patches. It navigates through layers, which consist of MLP and attention layer blocks for all patches. The Transformer block primarily consists of two significant elements: the MLP and the multi-head self-attention block. Initially, SA learned the relations among dissimilar areas (patches) of the image, thus seizing global features. Later in this phase, the block of MLP also augments the model of every patch by handling the attention mechanism (AM) output. During the development of the \(\:ViT\) classifier method, blocks of transformers, accompanied by residual links and multi-layer attention, are constructed using a patch-based image representation.
MaxViT, a multi-axis ViT, is a hybrid approach that incorporates CNN and ViT structures. In this way, it achieves higher performance by successfully capturing either local details or global context in the image processing task. This block structure incorporates a group of convolution and attention mechanisms specifically designed to capture both global and local features effectively. While block attention and depth convolutions remove local features, channel attention and grid attention learn the importance of features and global context. The multi-axis AM enables the method of learning relationships through dimensions concurrently. Normalization and Hopping methods increase the training procedure and raise the model’s depth while maintaining its performance. This hybrid model enables MaxViT to achieve higher precision and efficacy in computer vision tasks.
SimCLR method
SimCLR uses twin networks. These networks learn from unlabeled data in datasets that require a sufficient quantity of data, surpassing standard data augmentation in four phases33. During the initial phase of SimCLR, unlabeled data is augmented to create an improved data pool for every sample, utilizing standard data augmentation methods (such as adding Gaussian blur noise, random resizing, and random cropping). During the second phase, the encoding network \(\:f(.)\) receives feature vectors for the augmented images \(\:{x}_{i}\) and \(\:{x}_{j}.\) During the third phase, a projection operation \(\:g(.)\) is used. Finally, the contrastive loss value for the vectors \(\:{z}_{i}\) and \(\:{z}_{j}\) is computed. The contrastive loss is specified by the succeeding Eq. (4).
Now, \(\:N\) refers to sample counts, and \(\:\tau\:\) denotes the temperature parameter. The indicator function is described as \(\:\left\{E\breve{g}er\right[k\ne\:i\left]ise1\right\}\). After computing the loss, the contrastive loss is calculated according to the cosine similarity among the chosen number of positive data samples. The presented method is trained based on the backpropagation (BP) model, which captures the partial derivatives of the contrastive loss approach.
Twins-SVT technique
Twins-SVT incorporates a global attention mechanism and local self-attention, achieving image feature processing through the overview of Spatially Separable Self-Attention (SSSA)34. SSSA contains Global Subsampling Attention (GSA) and Local Spatial Attention (LSA). This incorporation enables the method to capture either global or local information simultaneously. Figure 2 depicts the framework of the Twins-SVT model.
Structure of the Twins-SVT method.
By combining global attention with local self-attention, this model addresses the issue of reducing receptive fields while achieving outstanding performance in forecasting tasks. Its prominent benefits lie in its efficacy and simplicity of execution, offering important possibilities for application across various visual tasks. Formally, SSSA was formulated as Eqs. (5)-(8):
.
SAE-based precancerous lesion detection process
Finally, the SAE classifier is utilized for the precancerous lesion detection process35. This model effectively learns the hierarchical and nonlinear feature representations from high-dimensional data. This model demonstrates excellence in extracting relevant features while mitigating noise and redundancy, a crucial aspect in detecting subtle discrepancies in lesion patterns. The deep architecture of the model also enables the capture of intrinsic data associations, thus enhancing the generalization of unseen samples. This technique also offers better robustness and adaptability compared to shallow models, making it particularly suitable for medical image analysis, where data variability is high. Moreover, the unsupervised pre-training followed by supervised fine-tuning improves the accuracy and stability of the classification, providing a reliable approach for early and precise lesion detection. Figure 3 specifies the structure of the SAE approach.
Architecture of the SAE method.
AE is an unsupervised ANN that comprises dual major elements: decoder and encoder. The decoder and encoder perform basic units of SAE methodology. Let \(\:{\alpha\:}^{\left(l\right)}=({\alpha\:}_{1}^{\left(l\right)},\:{\alpha\:}_{2}^{\left(l\right)},\:\cdots\:,\:{\alpha\:}_{{m}_{l}}^{\left(l\right)}{)}^{T}\) depict the latent feature vector taken from the lth encoder layer inside the SAE structure, where \(\:{m}_{l}\) signifies the overall hidden unit counts in the specified encoder layer. The \(\:l\)th encoder \(\:{\phi\:}_{\check{\theta}}^{\left(l\right)}(\bullet\:)\) compress input \(\:{\alpha\:}^{(l-1)}\) into hidden representation specified \(\:{\alpha\:}^{\left(l\right)}\), controlled by succeeding equation:
Here, \(\:{W}_{e}^{\left(l\right)}\) and \(\:{b}_{e}^{\left(l\right)}\) refer to the weighted matrix and bias of the encoder, and encoder \(\:{\phi\:}_{\check{\theta}}^{\left(l\right)}(\bullet\:)\) is parameterized by \(\:\check{\theta}=\left[{W}_{e}^{\left(l\right)},{b}_{e}^{\left(l\right)}\right]\), correspondingly, and \(\:L\) represents overall layer counts in the stacked encoder (SE). The original input \(\:{\alpha\:}^{\left(0\right)}\) is standardized snapshot data \(\:\stackrel{\sim}{u}\in\:{\mathbb{R}}^{{N}_{x}}\). The final latent feature \(\:{\alpha\:}^{\left(L\right)}\in\:{\mathbb{R}}^{{m}_{L}}\) depicts the “code” in subsequent discussions, denoted as \(\:\alpha\:\); its dimensionality \(\:{m}_{L}\) is specified by \(\:m.\) The \(\:lth\) decoder refers to \(\:{\psi\:}_{\widehat{\theta\:}}^{\left(l\right)}(\bullet\:)\), alters unseen representation \(\:{\widehat{\alpha\:}}^{\left(l\right)}\) into input space \(\:{\widehat{\alpha\:}}^{(l-1)}\) over the succeeding formula:
Now the decoder \(\:{\psi\:}_{\check{\theta}}^{\left(l\right)}(\bullet\:)\) is parameterized by \(\:\widehat{\theta\:}=\left[{W}_{d}^{\left(l\right)},{b}_{d}^{\left(l\right)}\right]\), and \(\:{W}_{d}^{\left(l\right)}\), and \(\:{b}_{d}^{\left(l\right)}\) refers to the weighted matrix and bias of the decoder correspondingly. Concerning the activation functions \(\:g(.)\), the ReLU function is accepted for in-between HLs, the \(\:tanh\) function is an alternate possibility, and the Sigmoid function is employed for the last resultant layer:
Train an SAE methodology to decrease the discrepancy between the first input \(\:{\stackrel{\sim}{u}}_{i}\) and the last output \(\:{\widehat{\stackrel{\sim}{u}}}_{i}\). It is accomplished by optimally modifying the parameters of a neural network that contains a weight, W, and a bias, \(\:b\). The above-mentioned discrepancy is quantified by a cost function containing three terms.
Here \(\:\beta\:\) and \(\:\gamma\:\) refer to constant coefficients. The 1st term of the loss function is the mean sum of square error for the discrepancy between input \(\:{\stackrel{\sim}{u}}_{i}\) and outcome \(\:{\widehat{\stackrel{\sim}{u}}}_{i}\) through the overall dataset comprising the data of snapshots \(\:n\):
A sparsity cost might be combined with a loss function to penalize the activation of HL, thereby preventing some hidden neurons from being activated without a cooperative outcome. This is depicted by the 2nd term in Eq. (14) that describes the cost of sparsity (here \(\:m\) depicts neuron counts in HL and index \(\:j\) iterates through every hidden neuron inside the system):
Now, \(\:{\widehat{\kappa\:}}_{j}\) depicts the average activation of hidden neuron values \(\:j\) in the entire dataset, and \(\:\kappa\:\) denotes the wanted value of average activation. The 3rd term in Eq. (14) represents the regularisation cost, which serves to minimize the magnitude of weights and thereby helps stop overfitting
Now, \(\:\varPhi\:(\cdot\:)\) indicates the SE function. A standardized snapshot of data \(\:{\stackrel{\sim}{u}}_{i}\) significantly compressed to a low-dimensional code \(\:{\alpha\:}_{i}\) over SE, here \(\:{N}_{X}"\:m\). Besides, the SD function retrieves the initial input \(\:{\widehat{\stackrel{\sim}{u}}}_{i}\) from code \(\:{\alpha\:}_{i}\):
Here \(\:\varPsi\:(\bullet\:)\) refers to the SD function. Algorithm 1 describes the SAE technique.
SAE technique.
Model performance analysis
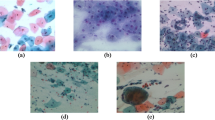
The performance evaluation of the FAFRDRL-CPLC technique is examined under the Malhari dataset36. The method runs on Python 3.6.5 with an i5-8600k CPU, 4GB GPU, 16GB RAM, 250GB SSD, and 1 TB HDD, using a 0.01 learning rate, ReLU, 50 epochs, 0.5 dropout, and batch size 5. Table 2 describes the dataset and Fig. 4 indicates the sample images.
Sample images.
Evaluation metrics for classification models: \(\:\varvec{A}\varvec{c}\varvec{c}\varvec{u}{\varvec{r}}_{\varvec{y}}\), \(\:\varvec{P}\varvec{r}\varvec{e}\varvec{c}{\varvec{i}}_{\varvec{n}}\), \(\:\varvec{S}\varvec{e}\varvec{n}\varvec{s}{\varvec{i}}_{\varvec{y}}\), \(\:\varvec{S}\varvec{p}\varvec{e}\varvec{c}{\varvec{i}}_{\varvec{y}}\), and \(\:{\varvec{F}}_{\varvec{S}\varvec{c}\varvec{o}\varvec{r}\varvec{e}}\)
The performance of the FAFRDRL-CPLC model is computed under diverse metrics. Equation (20) depicts the metric \(\:acc{ur}_{y}\), which measures the overall proportion of correct predictions. Equation (21) computes \(\:pre{ci}_{n}\), the ratio of correct positive predictions. Equation (22) defines \(\:sen{si}_{y}\), which evaluates the model’s ability to detect true positives correctly. Lastly, Eq. (23) defines \(\:spe{ci}_{y}\), which measures the model’s ability to correctly classify negative instances, providing a balanced evaluation of performance. Equation (24) illustrates the \(\:F{1}_{\:score}\), a metric that integrates \(\:pre{ci}_{n}\) and \(\:sen{si}_{y}\) into a single value to balance their trade-offs, specifically in cases with imbalanced classes.
Here, \(\:TP\) represents True Positives, \(\:TN\) indicates True Negatives, \(\:FP\) denotes False Positives, and \(\:FN\) stands for False Negatives.
Experimental validation evaluation metrics
Figure 5 displays the confusion matrices generated by the FAFRDRL-CPLC model for the 80% TRAPA/20% TESPA and 70% TRAPA/30% TESPA scenarios. Under 80% of TRAPA, NILM (127) and LSIL (58) were classified with high accuracy, while HSIL and SSC had minor misclassifications. Likewise, under 20:30, NILM remained the most accurately predicted class. LSIL and HSIL illustrated some confusion, specifically with each other, and SSC had lower accuracy due to class imbalance. The 70% training phase also showed consistent performance, with NILM (108) and LSIL (57) maintaining robust outputs. The outcomes represent that the FAFRDRL-CPLC method is capable of identifying four classes.
Confusion matrices (a-b) 80%TRAPA and 20%TESPA and (c-d) 70%TRAPA and 30%TESPA.
In Table 3; Fig. 6, the entire cancer recognition results of the FAFRDRL-CPLC technique, using 80% TRAPA and 20% TESPA, are displayed. The results showed that the FAFRDRL-CPLC technique can accurately determine the samples. On 80%TRAPA, the FAFRDRL-CPLC technique achieves an average \(\:acc{ur}_{y}\), \(\:pre{ci}_{n}\), \(\:sens{i}_{y}\), \(\:spec{i}_{y}\), and \(\:{F}_{score}\:\)of 98.62%, 97.07%, 95.39%, 98.92%, and 96.17%. Besides, on 20%TESPA, the FAFRDRL-CPLC approach reaches an average \(\:acc{ur}_{y}\), \(\:pre{ci}_{n}\), \(\:sens{i}_{y}\), \(\:spec{i}_{y}\), and \(\:{F}_{score}\:\)of 96.88%, 93.68%, 89.39%, 97.50%, and 90.53%.
Average of FAFRDRL-CPLC approach with 80%TRAPA and 20%TESPA.
In Fig. 7, the TRA \(\:acc{ur}_{y}\) (TRAAY) and validation \(\:acc{ur}_{y}\) (VLAAY) solutions of the FAFRDRL-CPLC technique with 80%TRAPA and 20%TESPA are revealed. The outcome emphasized that the TRAAY and VLAAY outcomes show an increasing trend, reporting the proficiency of the FAFRDRL-CPLC technique with greater outcomes through numerous iterations. Furthermore, the TRAAY and VLAAY remain closer throughout the epochs, which minimizes overfitting and yields higher outcomes of the FAFRDRL-CPLC, thereby safeguarding reliable predictions on unseen instances.
\(\:Accu{r}_{y}\) graph of FAFRDRL-CPLC technique with 80%TRAPA and 20%TESPA.
In Fig. 8, the TRA loss (TRALO) and VLA loss (VLALO) outcome of the FAFRDRL-CPLC technique with 80%TRAPA and 20%TESPA is shown. It is depicted that the TRALO and VLALO values demonstrate minimal tendencies, indicating the FAFRDRL-CPLC technique’s ability to balance a trade-off between data fitting and generalization. The constant decrease in loss outcomes also ensures a more favourable outcome for the FAFRDRL-CPLC method, adjusting the forecast outcomes over time.
Loss graph of FAFRDRL-CPLC technique with 80%TRAPA and 20%TESPA.
A brief precision-recall inspection of the FAFRDRL-CPLC approach with 80%TRAPA and 20%TESPA is represented in Fig. 9. By identifying the figure, it is noted that the FAFRDRL-CPLC method has achieved the highest precision-recall outcome under HSIL, LSIL, NILM, and SSC classes.
PR graph of FAFRDRL-CPLC technique with 80%TRAPA and 20%TESPA.
A detailed ROC study of the FAFRDRL-CPLC technique with 80% TRAPA and 20% TESPA is described in Fig. 10. The results indicate that the FAFRDRL-CPLC model has demonstrated proficiency in classifying four diverse classes, including HSIL, LSIL, NILM, and SSC, on the test dataset.
ROC graph of FAFRDRL-CPLC technique with 80%TRAPA and 20%TESPA.
In Table 4; Fig. 11, the entire cancer recognition outcomes of the FAFRDRL-CPLC technique under 70% TRAPA and 30% TESPA are depicted. The outcomes indicated that the FAFRDRL-CPLC technique can skillfully identify the samples. Based on 80%TRAPA, the FAFRDRL-CPLC model achieves average \(\:acc{ur}_{y}\), \(\:pre{ci}_{n}\), \(\:sens{i}_{y}\), \(\:spec{i}_{y}\), and \(\:{F}_{score}\:\)of 97.75%, 94.01%, 93.34%, 98.38%, and 93.33%. Moreover, on 20%TESPA, the FAFRDRL-CPLC model achieves average \(\:acc{ur}_{y}\), \(\:pre{ci}_{n}\), \(\:sens{i}_{y}\), \(\:spec{i}_{y}\), and \(\:{F}_{score}\:\)of 96.88%, 94.06%, 90.64%, 97.55%, and 91.41%.
Average of FAFRDRL-CPLC approach with 70%TRAPA and 30%TESPA.
In Fig. 12, the TRAAY and VLAAY results of the FAFRDRL-CPLC approach with 70% TRAPA and 30% TESPA are presented. The outcome indicates that the TRAAY and VLAAY graphs show an upward trend, reflecting the potential of the FAFRDRL-CPLC approach, which provides superior outcomes across numerous iterations. Additionally, the TRAAY and VLAAY remain close throughout the epochs, indicating marginal overfitting and demonstrating the efficacy of the FAFRDRL-CPLC method, which promises continuous prediction on unobserved samples.
\(\:Accu{r}_{y}\) graph of FAFRDRL-CPLC methodology with 70%TRAPA and 30%TESPA.
In Fig. 13, the TRALO and VLALO graphs of the FAFRDRL-CPLC approach with 70% TRAPA and 30% TESPA are illustrated. The TRALO and VLALO graphs indicate downward trends, highlighting the effectiveness of the FAFRDRL-CPLC approach in striking a balance between data fitting and generality. The persistent drop further promises a more accurate result from the FAFRDRL-CPLC model, ultimately improving prediction outcomes.
Loss graph of FAFRDRL-CPLC methodology with 70%TRAPA and 30%TESPA.
A brief precision-recall investigation of the FAFRDRL-CPLC method at 70%TRAPA and 30%TESPA technique is exhibited in Fig. 14. From the figure, it is noted that the FAFRDRL-CPLC technique attained maximal precision-recall performance on HSIL, LSIL, NILM, and SSC classes.
PR graph of FAFRDRL-CPLC methodology with 70%TRAPA and 30%TESPA.
A detailed ROC analysis of the FAFRDRL-CPLC model with 70% TRAPA and 30% TESPA is presented in Fig. 15. The results indicate that the FAFRDRL-CPLC technique has demonstrated its capability to categorize four different classes, namely HSIL, LSIL, NILM, and SSC, on the test dataset.
ROC graph of FAFRDRL-CPLC methodology with 70%TRAPA and 30%TESPA.
Comparison and computational time analysis of the FAFRDRL-CPLC approach with existing models under \(\:\varvec{A}\varvec{c}\varvec{c}\varvec{u}{\varvec{r}}_{\varvec{y}}\), \(\:\varvec{P}\varvec{r}\varvec{e}\varvec{c}{\varvec{i}}_{\varvec{n}}\), \(\:\varvec{S}\varvec{e}\varvec{n}\varvec{s}{\varvec{i}}_{\varvec{y}}\), \(\:\varvec{S}\varvec{p}\varvec{e}\varvec{c}{\varvec{i}}_{\varvec{y}}\), and \(\:{\varvec{F}}_{\varvec{S}\varvec{c}\varvec{o}\varvec{r}\varvec{e}}\) metrics
In Table 5; Fig. 16, a detailed comparison analysis of the FAFRDRL-CPLC method is reported12]– [13,37,38,39. The results showed that the Yolov8, SAHI, DCGAN, Adam-based CNN, ViT with SPT, U-Net, GoogLeNet, and W-Net methodologies provided ineffective detection outcomes with \(\:acc{ur}_{y}\) of 90.76%, 91.73%, 97.90%, 90%, 91%, 95.14%, 96.17%, and 97%, respectively. Whereas, the ensemble model has achieved reasonable outcomes with \(\:acc{ur}_{y}\) of 97.83%, \(\:pre{ci}_{n}\) of 96.05%, \(\:sens{i}_{y}\) of 92.11%, and \(\:spec{i}_{y}\) of 90.02%. Furthermore, the RF hierarchical algorithm has shown considerable performance with \(\:acc{ur}_{y}\) of 98.42%, \(\:pre{ci}_{n}\) of 90.95%, \(\:sens{i}_{y}\) of 91.43%, and \(\:spec{i}_{y}\) of 95.57%. Finally, the FAFRDRL-CPLC technique exhibits higher performance with an\(\:\:acc{ur}_{y}\) of 98.62%, \(\:pre{ci}_{n}\) of 97.07%, \(\:sens{i}_{y}\) of 95.39%, and \(\:spec{i}_{y}\) of 98.92%.
Comparative study of the FAFRDRL-CPLC model with existing approaches.
Table 6; Fig. 17 present the computational time (CT) analysis of the FAFRDRL-CPLC technique in comparison to existing models. The FAFRDRL-CPLC technique illustrates a CT of 10.37 s, thus outperforming the existing models. Additionally, the Ensemble, RF hierarchical, ViT with SPT illustrates CTs of 14.16 s, 17.14 s, and 17.31 s. Other models such as SAHI and U-Net attains CT of 21.39 and 20 s, respectively, while Yolov8 and Adam-based CNN exhibit longer CTs of 29.67 and 29.85 s. The results indicate that the FAFRDRL-CPLC model presents a significant improvement in processing efficiency while maintaining high classification performance.
CT analysis of the FAFRDRL-CPLC technique with existing models.
Comprehensive evaluation of the FAFRDRL-CPLC technique: performance Metrics, component Impact, and computational efficiency
The error analysis of the FAFRDRL-CPLC approach with recent techniques is shown in Table 7; Fig. 18. The FAFRDRL-CPLC approach attains the lowest error rates across \(\:acc{ur}_{y}\), \(\:pre{ci}_{n}\), \(\:sens{i}_{y}\), and \(\:spec{i}_{y}\) with values of 1.38%, 2.93%, 4.61%, and 1.08%, respectively, illustrating greater accomplishment. Comparatively, models like Adam-based CNN and Yolov8 illustrate higher error rates, with \(\:acc{ur}_{y}\) errors of 10% and 9.24%, respectively. W-Net and GoogLeNet exhibit relatively low \(\:pre{ci}_{n}\) and sensitivity \(\:sens{i}_{y}\) but higher errors in other metrics. This analysis highlights the robustness of the FAFRDRL-CPLC model in minimizing classification errors across all key metrics.
Error assessment of the FAFRDRL-CPLC approach with recent techniques.
Table 8; Fig. 19 clearly specify the ablation study of the FAFRDRL-CPLC methodology. The baseline ADF technique achieved an \(\:acc{ur}_{y}\) of 95.51%, \(\:pre{ci}_{n}\) of 93.57%, \(\:sens{i}_{y}\) of 92.36%, and \(\:spec{i}_{y}\) of 95.63%, each successive addition of deep feature extraction models led to noticeable performance enhancements. MaxViT-v2, SimCLR, and Twins-SVT contributed progressively to accuracy and generalization, with Twins-SVT attaining an \(\:acc{ur}_{y}\) of 97.43% and \(\:sens{i}_{y}\) of 94.14%. Incorporating the SAE model additionally enhanced the outputs by attaining an \(\:acc{ur}_{y}\) of 97.99% and \(\:pre{ci}_{n}\) of 96.35%. The FAFRDRL-CPLC model, integrating all key components, illustrated the highest \(\:acc{ur}_{y}\) of 98.62%, \(\:pre{ci}_{n}\) of 97.07%, \(\:sens{i}_{y}\) of 95.39%, and \(\:spec{i}_{y}\) of 98.92%, highlighting the overall strength and complementary behaviour of the chosen methods in improving cervical lesion classification performance.
Result analysis of the ablation study of the FAFRDRL-CPLC methodology.
Table 9 demonstrates the computational efficiency of the FAFRDRL-CPLC model40. The FAFRDRL-CPLC model demonstrates the lowest computational demand with 15.09 FLOPs and the least GPU memory consumption at 893 MB, indicating its insubstantial nature. In contrast, other models such as ESOD-YOLO and YOLOv3-tiny require higher FLOPs and significantly more GPU resources, with values attaining up to 29.62 FLOPs and over 2100 MB of GPU memory. This highlights the efficiency, making it appropriate for deployment in resource-constrained environments without losing performance.
Conclusion
In this article, the FAFRDRL-CPLC approach, utilizing biomedical image analysis, is proposed. The primary purpose of the FAFRDRL-CPLC approach is to serve as a valuable tool for assisting clinicians in the initial study and treatment planning of cervical precancerous lesions. Initially, the FAFRDRL-CPLC approach applies ADF for pre-processing. Furthermore, the fusion of advanced feature reduction models, such as MaxViT-v2, SimCLR, and Twins-SVT, is employed to capture diverse and complementary representations from the pre-processed images. Finally, the SAE classifier is utilized for the precancerous lesion detection process. The FAFRDRL-CPLC method is examined through experimentation using the Malhari dataset. The comparison study of the FAFRDRL-CPLC method demonstrated a superior accuracy value of 98.62% over existing approaches. The limitations of the FAFRDRL-CPLC method comprise a relatively small dataset, which may lead to ineffective generalization in the results. The performance may also be affected by discrepancies in image quality and acquisition protocols. Furthermore, the resource requirements and computational complexity may also pose a threat to deployment in low-resource clinical settings. The study also lacks a thorough exploration of potential clinical integration scenarios that require seamless compatibility with the existing healthcare infrastructure. Deployment feasibility in real-world environments also requires additional validation, including robustness against noise and artefacts. Moreover, challenges such as patient privacy, data security, and obtaining regulatory approvals must be addressed before widespread adoption can occur. Future work may focus on addressing these practical constraints to ensure the effective and reliable clinical use of these tools.
Data availability
The data that support the findings of this study are openly available in the Kaggle repository at https://www.kaggle.com/datasets/jocelyndumlao/malhari-dataset/data, reference number [37].
References
Allahqoli, L. et al. Diagnosis of cervical cancer and precancerous lesions by artificial intelligence: a systematic review. Diagnostics 12(11), 2771. (2022).
Fang, S., Yang, J., Wang, M., Liu, C. & Liu, S. An improved image classification method for cervical precancerous lesions based on ShuffleNet. Comput. Intell. Neurosci., 2022(1), 9675628. (2022).
Nancy, G. & Bhuvaneswari, E. A deep learning-driven multi-layer digital twin framework with Miot for precision oncology in cancer diagnosis. Journal Intell. Syst. & Internet Things, 17(1), 16–26 (2025).
Mahmood, H. et al. Use of artificial intelligence in diagnosis of head and neck precancerous and cancerous lesions: a systematic review. Oral Oncology, 110, 104885. (2020).
Wang, J. et al. Raman spectroscopic study of cervical precancerous lesions and cervical cancer. Lasers Med. Science, 36(9), 1–10. (2021).
Mansouri, R. & Ragab, M. Equilibrium optimization algorithm with ensemble learning based cervical precancerous lesion classification model. Healthcare 11(1), 55). (MDPI, 2022).
Agustiansyah, P. et al. Automated precancerous lesion screening using an instance segmentation technique for improving accuracy. Sensors 22(15), 5489. (2022).
Albayrak, A. et al. A whole-slide image grading benchmark and tissue classification for cervical cancer precursor lesions with inter-observer variability. Med. Biol. Eng. Comput. 59, 1545–1561 (2021).
Ephrem Dibisa, K., Tamiru Dinka, M., Mekonen Moti, L. & Fetensa, G. Precancerous lesion of the cervix and associated factors among women of West Wollega, West Ethiopia, 2022. Cancer Control, 29, 10732748221117900. (2022).
Masri, A. N. A. & Mokayed, H. An efficient machine learning-based cervical cancer detection and classification. J. Cybersecur. Inf. Manag. 2 (2), 8–58 (2021).
Wang, J. et al. Raman spectroscopic study of cervical precancerous lesions and cervical cancer. Lasers Med. Sci. 36 (9), 1855–1864 (2021).
Nurmaini, S. et al. Robust assessment of cervical precancerous lesions from pre-and post-acetic acid cervicography by combining deep learning and medical guidelines. Inform. Med. Unlocked 52, 101609. (2025).
Wubineh, B. Z., Jeleń, Ł. & Rusiecki, A. DCGAN-based cytology image augmentation for cervical cancer cell classification using transfer learning. Procedia Comput. Sci. 256, 1003–1011 (2025).
Khan, Z., Khan, S. U. R., Bilal, O., Raza, A. & Ali, G. Optimizing cervical lesion detection using deep learning with particle swarm optimization. In 2025 6th International Conference on Advancements in Computational Sciences (ICACS) (pp. 1–7). (IEEE, 2025).
Himabindu, D. D., Lydia, E. L., Rajesh, M. V., Ahmed, M. A. & Ishak, M. K. Leveraging swin transformer with ensemble of deep learning model for cervical cancer screening using colposcopy images. Sci. Rep. 15(1), 7900. (2025).
Allogmani, A. S., Mohamed, R. M., Al-Shibly, N. M. & Ragab, M. Enhanced cervical precancerous lesions detection and classification using Archimedes Optimization Algorithm with transfer learning. Sci. Rep. 14(1), 12076. (2024).
Abinaya, K. & Sivakumar, B. A deep learning-based approach for cervical cancer classification using 3D CNN and vision transformer. J. Imaging Inform. Med. 37(1), 280. (2024).
Sholik, M., Fatichah, C. & Amaliah, B. July. Classification of cervical cell images into healthy or cancer using convolution neural network and linear discriminant analysis. In 2023 IEEE International Conference on Industry 4.0, Artificial Intelligence, and Communications Technology (IAICT) (pp. 383–389). (IEEE, 2023).
Attallah, O. Cercan· net: cervical cancer classification model via multi-layer feature ensembles of lightweight Cnns and transfer learning. Expert Syst. Appl. 229, 120624 (2023).
Song, J., Wang, L., Yan, J., Feng, Y. & Zhang, Y. Enhancing cervical precancerous lesion detection using African vulture optimization algorithm with deep learning model. Biomed. Signal Process. Control 97, 106665. (2024).
Chauhan, N. K., Singh, K., Kumar, A. & Kolambakar, S. B. HDFCN: A robust hybrid deep network based on feature concatenation for cervical cancer diagnosis on WSI pap smear slides. BioMed Res. Int. 2023(1), 4214817. (2023).
Chauhan, N. K. et al. A hybrid learning network with progressive resizing and PCA for diagnosis of cervical cancer on WSI slides. Sci. Rep., 15(1), 12801. (2025).
Chauhan, N. K. & Singh, K. September. Impact of variation in number of channels in CNN classification model for cervical cancer detection. In 2021 9th international conference on reliability, infocom technologies and optimization (Trends and future directions)(ICRITO) (pp. 1–6). (IEEE, 2021).
Bilal, O., Hekmat, A. & Khan, S. U. R. Automated cervical cancer cell diagnosis via grid search-optimized multi-CNN ensemble networks. Netw. Model Anal Health Inform. Bioinform., 14(1), p.67. (2025).
Chauhan, N. K. & Singh, K. February. Diagnosis of cervical cancer with oversampled unscaled and scaled data using machine learning classifiers. In 2022 IEEE Delhi Section Conference (DELCON) (pp. 1–6). (IEEE, 2022).
Raza, M. A. et al. Advanced feature extraction for cervical cancer image classification: integrating neural feature extraction and autoint models. Sensors 25(9), 2826. (2025).
Chauhan, N. K. & Singh, K. Performance assessment of machine learning classifiers using selective feature approaches for cervical cancer detection. Wireless Pers. Commun. 124 (3), 2335–2366 (2022).
Gangrade, J., Kuthiala, R., Gangrade, S., Singh, Y. P. & Solanki, S. R, M. and A deep ensemble learning approach for squamous cell classification in cervical cancer. Sci. Rep., 15(1), 7266. (2025).
Abd-Alhalem, S. M. et al. Cervical cancer classification based on a bilinear convolutional neural network approach and random projection. Eng. Appl. Artif. Intell., 127, p.107261. (2024).
Borse, M. & Yadav, M. Cervical cancer detection using image processing and machine learning. Cuestiones De Fisioterapia. 54 (2), 3575–3581 (2025).
Nadeem, A., Ashraf, R., Mahmood, T. & Parveen, S. Automated CAD system for early detection and classification of pancreatic cancer using deep learning model. Plos One. 20 (1), e0307900 (2025).
Sarıateş, M. & Özbay, E. A Classifier model using fine-tuned convolutional neural network and transfer learning approaches for prostate cancer detection. Appl. Sci., 15(1), 225. (2024).
Fırıldak, K., Çelik, G. & Talu, M. F. SimCLR-based Self-Supervised learning approach for limited brain MRI and unlabeled images. Bitlis Eren Üniversitesi Fen Bilimleri Dergisi. 13 (4), 1304–1313 (2024).
Zhu, K. et al. FCFormer: fish density estimation and counting in recirculating aquaculture system. Front. Mar. Sci. 11, 1370786. (2024).
Zhu, C., Fu, J., Xiao, D. & Wang, J. Nonlinear model order reduction of engineering turbulence using data-assisted neural networks. Comput. Phys. Commun. 309, 109501. (2025).
https://www.kaggle.com/datasets/jocelyndumlao/malhari-dataset/data
Darwish, M., Altabel, M. Z. & Abiyev, R. H. Enhancing cervical precancerous classification using advanced vision transformer. Diagnostics 13(18), 2884. (2023).
Chatterjee, P., Siddiqui, S., Kareem, R. S. A. & Rao, S. Attention-enhanced lightweight architecture with hybrid loss for colposcopic image segmentation. Cancers 17(5), 781. (2025).
Diniz, D. N. et al. A hierarchical feature-based methodology to perform cervical cancer classification. App. Sci. 11(9), 4091. (2021).
Luo, J. et al. Efficient small object detection you only look once: a small object detection algorithm for aerial images. Sensors 24(21), 7067. (2024).
Funding
None.
Author information
Authors and Affiliations
Contributions
S. S. Saranya: Conceptualization, methodology development, experiment, formal analysis, investigation, writing. Santhanakrishnan C: Formal analysis, investigation, validation, visualization, writing. K. Pradeep Mohan Kumar: Formal analysis, review and editing. P. Anil Kumar: Methodology, investigation. Bibhuti Bhusan Dash: Review and editing.Kanchan Bala: Discussion, review and editing. Saroja Kumar Rout: Conceptualization, methodology development, investigation, supervision, review and editing.All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
This article does not contain any studies with human participants performed by any of the authors.
Consent to participate
Not applicable.
Informed consent
Not applicable.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Saranya, S.S., Santhanakrishnan, C., Kumar, K.P.M. et al. Advancements in fusion-based deep representation learning for enhanced cervical precancerous lesion classification using biomedical image analysis. Sci Rep 15, 36705 (2025). https://doi.org/10.1038/s41598-025-20693-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-20693-4