Abstract
Recurrent vulvovaginal candidiasis (RVVC) is a common, refractory fungal infection affectingwomen, primarily caused by Candida albicans. The interplay among fungal virulence factors, biofilm formation, and antifungal resistance is crucial in the pathogenesis of RVVC. This study compared 50 Candida albicans isolates from RVVC patients and 50 from asymptomatic vaginal colonizers. Antifungal susceptibility testing was performed using the broth microdilution method. Biofilm formation was assessed via crystal violet staining, and the expression levels of virulence factor hydrolases (SAP, PL, Lip) and cell wall protein genes (ALS1, ALS3, HWP1) were analyzed using phenotypic assays and quantitative real-time PCR (qRT-PCR). Pearson correlation analysis was used to evaluate the relationships among these parameters and antifungal resistance. RVVC isolates exhibited significantly higher MICs for fluconazole, voriconazole, and itraconazole. Biofilm formation ability and the expression levels of SAP, PL, Lip, ALS1, ALS3, and HWP1 were also significantly higher in RVVC isolates. A moderate correlation was observed between antifungal drug MIC values and biofilm OD, while a weak correlation existed between MIC values and ALS/HWP1 gene expression. Notably, hydrolase expression showed no significant correlation with resistance. Candida albicans from RVVC patients demonstrated enhanced biofilm formation, virulence factor expression, and antifungal resistance. Biofilm-mediated drug tolerance may be a key mechanism underlying the refractoriness of RVVC. Targeting biofilm formation and virulence factor genes may offer novel strategies for managing RVVC.
Similar content being viewed by others
Introduction
Recurrent vulvovaginal candidiasis (RVVC) is a refractory fungal infection prevalent among women, affecting approximately 5% of women globally, with Candida albicans accounting for over 80% of cases1. Characterized by recurrent episodes and therapeutic challenges, RVVC not only causes clinical symptoms such as curd-like vaginal discharge and pruritus but also correlates with psychological issues like anxiety and depression, significantly impairing quality of life2. Current evidence suggests that RVVC development involves multiple factors, including candidal virulence factors, host immune status, and genetic predispositions, with the interplay between fungal virulence and antifungal resistance serving as a core mechanism underlying disease refractoriness3.
The pathogenicity of C. albicans relies on a complex network of virulence factors. Secreted hydrolytic enzymes, such as aspartyl proteinases (SAP), phospholipases (PL), and lipases (Lip), degrade host mucosal proteins and cell membrane lipids, disrupting epithelial barriers and promoting fungal invasion4. Studies indicate that SAP family proteins weaken host defenses by degrading immunoglobulins and complement components5, while PL exacerbates tissue damage by inducing proinflammatory responses through arachidonic acid release6. Additionally, the dimorphic transition of C. albicans from yeast to hyphal forms is another pathogenic key: hyphae enhance adhesion and invasion via expressing cell wall adhesin genes (e.g., ALS1, ALS3, and HWP1)7. ALS3, a hypha-specific adhesin, mediates fungal internalization by binding to host epithelial integrins8, whereas HWP1 participates in hyphal cell wall cross-linking, contributing to maintain biofilm three-dimensional structure9.
Biofilm formation represents a critical mechanism of antifungal resistance in C. albicans. Composed of fungal communities encased in an extracellular matrix (including β-1,3-glucans and extracellular DNA), biofilms act as physical barriers to impede drug penetration, while reduced metabolic activity and overexpression of efflux pumps in biofilm cells further enhance resistance10. Clinical data indicate that biofilm-forming capacity is significantly higher in C. albicans isolates from RVVC patients than in asymptomatic colonizers, with biofilm density positively correlated with fluconazole resistance11. Notably, azole resistance is also associated with ERG11 mutations and overexpression of CDR1/CDR2 efflux pumps, though the direct link between virulence factors and resistance is still debated12.
Despite advances in understanding RVVC pathogenesis, key questions persist: (1) Do virulence factor expression and biofilm formation in RVVC patient isolates exhibit synergistic effects? (2) What is the mechanistic link between the expression of cell wall protein genes and antifungal resistance? This study aims to systematically analyze the correlation among virulence factors, biofilm formation, and resistance by comparing C. albicans isolates from RVVC patients and asymptomatic colonizers, seeking to provide new targets for precision therapy of RVVC.
Materials and methods
Study subjects
A total of 50 vaginal Candida albicans isolates were prospectively collected from adult female RVVC patients (age: 35.1 ± 5.8 years) and vaginal Candida albicans colonizers (age: 33.6 ± 6.1 years), with no statistically significant difference in age distribution between the two groups (t = 1.354, P = 0.198). All RVVC patients had a history of 3–4 episodes of candidal infection in the past year and were in acute attack at the time of the study. Routine leucorrhea examination showed mycelia of yeast-like fungi, and presented with clinical symptoms defined by the presence of at least two of the following signs and symptoms: curd-like leucorrhea, vaginal pruritus, obvious burning sensation, and dyspareunia. The enrolled patients had not received any antibacterial or antifungal treatment within one month before the study. Colonizers were women who had no history of candidal infection and no antifungal drug treatment in the past 2 years, had no obvious clinical symptoms, showed cleanliness grade I-II in routine leucorrhea examination, had yeast-like fungi but no mycelia found, and could isolate Candida albicans from secretions after separation and culture on Sabouraud agar medium. This study involving human subjects was approved by the Ethics Committee of Jiaying University School of Medicine (Approval No.: JYYXLL2025-09). All participants provided written informed consent prior to their involvement in the study. The experiments were performed in accordance with the Declaration of Helsinki and relevant guidelines and regulations.
Materials
The VITEK 2 Compact automatic microbial analysis system and yeast-like fungi drug susceptibility test kit were products of bioMérieux, France. The 7500 Real Time PCR System was from Thermo Fisher Scientific, USA. The Max250 multifunctional microplate reader was from Molecular Devices, USA. The yeast total RNA extraction kit, PrimeScript RT reagent Kit, and SYBR Premix Ex Taq were from TaKaRa, Japan. NaCl, KH₂PO₄, CaCl₂, glacial acetic acid, methanol, and absolute ethanol were purchased from Xi’an Taihua Medical Technology Co., Ltd. Calf serum, bovine serum albumin, peptone, and primers were synthesized by Shanghai Sangon Biotech Co., Ltd. Crystal violet, 50% egg yolk emulsion, CHROMagar Candida plates, Sabouraud agar medium, yeast extract peptone dextrose (YPD) liquid medium, tributyrin, and tributyrin plate agar base were all purchased from Hangzhou Binhe Microbial Reagent Co., Ltd. 96-well cell culture plates, RPMI-1640 culture solution, phosphate buffer, etc., were from Corning, USA. The standard strains, Candida albicans (ATCC 90028) and Candida parapsilosis (ATCC 22019), were both purchased from Shanghai Fuxiang Biotechnology Co., Ltd.
Fungal isolation, identificationand antifungal susceptibility testing
Vaginal swab specimens collected by gynecologists were inoculated onto CHROMagar Candida plates and cultured at 35 °C for 24–48 h. Emerald green colonies (diameter 2–3 mm) were picked, identified as Candida albicans by the VITEK 2 Compact automatic microbial analysis system, and preserved for subsequent use. Antifungal susceptibility testing was performed using the ATB Fungus 3 kit from bioMérieux, France. The minimum inhibitory concentrations (MICs) of five commonly used antifungal drugs (fluconazole, amphotericin B, 5-fluorocytosine, voriconazole, and itraconazole) against Candida albicans were determined by broth microdilution method. Results were interpreted as sensitive according to CLSI M59 and M60 breakpoints: 5-fluorocytosine ≤ 4 mg/L, amphotericin B ≤ 1 mg/L, fluconazole ≤ 8 mg/L, voriconazole ≤ 0.125 mg/L, and itraconazole ≤ 0.125 mg/L.Quality control for antifungal susceptibility testing was performed using standard strains: Candida albicans (ATCC 90028) and Candida parapsilosis (ATCC 22019). The MIC ranges of the five antifungal drugs for ATCC 90,028 were: fluconazole 0.25-1 mg/L, amphotericin B 0.25-1 mg/L, 5-fluorocytosine 0.125-0.5 mg/L, voriconazole 0.03–0.125 mg/L, and itraconazole 0.03–0.125 mg/L; for ATCC 22,019, the ranges were: fluconazole 0.5-2 mg/L, amphotericin B 0.5-2 mg/L, 5-fluorocytosine 0.25-1 mg/L, voriconazole 0.06–0.25 mg/L, and itraconazole 0.06–0.25 mg/L. All test results were within the recommended CLSI quality control ranges.
Detection of biofilm formation
Isolates were cultured on Sabouraud agar plates at 35 °C for 24 h. A Single colony of each isolate was inoculated into 5 mL of YPD liquid medium and cultured overnight at 35 °C with shaking at 200 rpm. The fungal culture was adjusted to 1 McFarland standard turbidity using RPMI-1640 medium with a McFarland turbidimeter. 100 µL of the fungal suspension was added to 96-well polystyrene microtiter plates and cultured at 35 °C with shaking for 2 h. The culture medium was discarded, and the wells were washed 3 times with phosphate buffer (pH 7.2), followed by adding 100 µL of fresh RPMI-1640 medium. The plates were continuously cultured at 35 °C, with medium replaced every 24 h after 3 washes with phosphate buffer. After two medium replacements, the culture medium was discarded, and the wells were washed 3 times with phosphate buffer. Each well was fixed with 10 µL of methanol for 15 min, and the supernatant was removed and air-dried. Each well was stained with 100 µL of 0.5% crystal violet solution for 15 min, washed 3 times with distilled water, and then 150 µL of 33% acetic acid solution was added to each well. The optical density (OD) was measured at 600 nm using a Max250 multifunctional microplate reader13. Candida albicans (ATCC 90028) was used as the positive control, and Candida parapsilosis (ATCC 22019) as the negative control. The evaluation of biofilm formation strength was performed according to the method described in the reference13. First, the OD cutoff value (ODc) for biofilm formation was defined as the mean absorbance value of the inoculum-free negative control plus three standard deviations (ODc = ODavg. of negative control + 3× standard deviation (SD) of negative control). Accordingly, isolates were classified as non-biofilm formers when OD ≤ ODc, weak biofilm formers when ODc < OD ≤ (2 × ODc), medium biofilm formers when 2 × ODc < OD ≤ (4 × ODc), and strong biofilm formers when OD > (4 × ODc).
Detection of three hydrolases
According to the formula reported in Reference14, bovine serum albumin plates for screening SAP, egg yolk agar plates for screening PL, and tributyrin plates for screening Lip were prepared. 10 µL of the fungal bacterial solution with a turbidity of 1 McFarland standard (as described above) was respectively spotted onto pre-applied circular filter papers (diameter 6 mm) on the bovine serum albumin plates, egg yolk agar plates, and tributyrin plates. The solution was spread on the filter paper with the blunt end of a 6-mm-diameter L-shaped rod, and the plates were placed in a 35 °C incubator for observation every 24 h. The optimal observation time was determined by the time when obvious precipitation rings or transparent rings first appeared around the colonies on the Candida albicans (ATCC 90028) plates. The optimal incubation times for the bovine serum albumin plate, egg yolk agar plate, and tributyrin plate were finally determined to be 7, 4, and 6 days, respectively. For other test strains, the colony diameter and the diameter of the colony plus the halo were observed and recorded at the optimal observation time, and the Pz value was calculated using the formula: Pz value = colony diameter / (colony diameter + halo diameter). Each strain was tested in triplicate on the three types of plates, and the average value was taken as the final result. A larger Pz value indicates less production of hydrolases. Currently, it is generally considered that a Pz value ≤ 0.59 indicates high hydrolase expression; a Pz value of 0.60–0.79 indicates moderate expression; a Pz value of 0.80-<1.00 indicates low expression; and a Pz value of 1 indicates no production of the hydrolase14.
Quantification of ALSI, ALS3, and HWP1 gene expression by qRT-PCR
Cells from overnight cultures grown in YPD broth (as described in Sect. Detection of Biofilm Formation) were harvested, and yeast cells were lysed with cell lysis solution. Total yeast RNA was extracted using a yeast total RNA extraction kit, washed, dissolved in RNase-free dH2O, and stored at -70 °C. The relative expression levels of ALS1, ALS3, and HWP1 under different conditions were detected by the SYBR Green I fluorescent dye method, using ACT1 and TEF1 as dual internal reference genes and the 2^-ΔΔCt method. Reverse transcription was performed according to the instructions of the PrimeScript RT reagent Kit. The reaction system, primer design for the target genes ALS1, ALS3, HWP1, ACT1, and TEF1, and the reaction system for real-time fluorescence quantitative PCR were all referred to Reference15. The calculation formulas for the relative expression levels of ALS1, ALS3, and HWP1 were: ΔΔCt = Ct (ALS1/ALS3/HWP1) - (CtACT1 + CtTEF1)/2, and Value = 2^-ΔΔCt, where Ct is the cycle threshold set according to the signal intensity of the PCR reaction.
Statistical methods
All data were statistically analyzed using SPSS 21.0 software. Enumeration data were expressed as rate (%), and the comparison of rates was performed by chi-square (χ²) test. Measurement data conforming to normal distribution were expressed as mean ± standard deviation (x̄ ± s), and t-test was used for comparison between groups. Data not conforming to normal distribution were described by minimum (min), 25th percentile (P25), 50th percentile (P50), 75th percentile (P75), and maximum (Max), and Mann-Whitney U test was adopted. The correlations among the expression levels of ALS1, ALS3, and HWP1 genes, biofilm formation indexes, the expression levels of three virulence factor hydrolases, and MIC values of various drugs were analyzed by Pearson correlation analysis. Effect sizes (odds ratios with 95% confidence intervals) were calculated for key comparative analyses to enhance result interpretation. A P < 0.05 was considered statistically significant.
Results
Antifungal susceptibility testing of Candida albicans from two sources
Thirteen strains of Candida albicans from RVVC patients were resistant to at least one of the five commonly used antifungal drugs, including 11 strains resistant to one drug and 2 strains resistant to two drugs. Ten strains of Candida albicans from vaginal colonizers were resistant to at least one of the five commonly used antifungal drugs, all of which were resistant to one drug. There was no statistically significant difference in the ratio of Candida albicans from the two sources resistant to at least one of the five commonly used antifungal drugs (χ²=0.508, P = 0.635). In terms of MIC, the MIC values of Candida albicans from RVVC patients to amphotericin B, 5-fluorocytosine, fluconazole, voriconazole and itraconazole were all higher than those from vaginal colonizers (all P<0.05), as shown in Table 1.
In vitro expression levels of three secreted hydrolases, ALS1, ALS3, HWP1 genes and biofilm formation by Candida albicans from different sources
Crystal violet staining showed that all 100 Candida albicans strains formed positive biofilms. The OD value of biofilm detection in strains from RVVC patients was significantly higher than that in strains from vaginal colonizers. In terms of biofilm formation ability, the biofilm-forming ability of strains isolated from RVVC patients is significantly stronger than that of strains from vaginal colonizers (P < 0.05). The Pz values of three secreted hydrolases (SAP, PL and Lip) in strains from RVVC patients were all significantly lower than those in strains from vaginal colonizers (all P < 0.05), indicating that the expression levels of three secreted hydrolases in strains from RVVC patients were significantly stronger than those in strains from vaginal colonizers (all P < 0.05). The in vitro relative mRNA expression levels of three cell wall protein genes (ALS1, ALS3 and HWP1) in strains from RVVC patients were all significantly higher than those in strains from vaginal colonizers (P < 0.05). See Tables 2 and 3.
In vitro expression levels of three secreted hydrolases, ALS1, ALS3 and HWP1 gene and biofilm formation by Candida albicans with different drug resistances
The OD value of biofilm detection in strains resistant to at least one of the five antifungal drugs was significantly higher than that in strains non-resistant to all five antifungal drugs (P < 0.05). However, there was no statistically significant difference in biofilm formation ability between the two groups (P > 0.05). There were no statistically significant differences in the Pz values of three secreted hydrolases (SAP, PL and Lip) between strains resistant to at least one of the five antifungal drugs and strains non-resistant to all five antifungal drugs (all P > 0.05), nor were there statistically significant differences in the expression levels of three secreted hydrolases between the two groups (all P > 0.05). The in vitro relative expression levels of three cell wall protein genes (ALS1, ALS3 and HWP1) in strains resistant to at least one of the five antifungal drugs were all higher than those in strains non-resistant to all five antifungal drugs, but the differences were not statistically significant (P > 0.05). See Tables 4 and 5.
Correlations among MIC values of antifungal drugs, Biofilm-OD, and relative expression levels of SAP, PL, Lip, ALS1, ALS3, and HWP1 genes in 100 vaginal Candida albicans isolates
The results showed that the MIC values of amphotericin B, 5-fluorocytosine, fluconazole, voriconazole, and itraconazole were moderately correlated with Biofilm-OD (all 0.4 < r < 0.6), and lowly correlated with the relative expression levels of ALS1, ALS3, and HWP1 genes (all 0.2 < r < 0.4), but showed no significant correlation with SAP, PL, and Lip (all |r| < 0.2). Biofilm-OD was lowly correlated with the relative expression levels of ALS1, ALS3, and HWP1 genes (all 0.2 < r < 0.4), and showed no significant correlation with SAP, PL, and Lip (all |r| < 0.2). The relative expression levels of ALS1, ALS3, and HWP1 genes were highly correlated with each other (all r > 0.6), but showed no significant correlation with SAP, PL, and Lip (all |r| < 0.2). See Table 6 for details.
Discussion
Our study revealed that the minimum inhibitory concentrations (MICs) of Candida albicans from RVVC patients against amphotericin B, 5-fluorocytosine, fluconazole, voriconazole, and itraconazole were significantly higher than those from colonizers (all P < 0.05). Notably, the maximum MIC of fluconazole reached ≥ 128 mg/L, indicating potential high-level resistance in some strains. Although the sensitivity rates showed no statistical difference between groups (e.g., fluconazole sensitivity rate: 88% vs. 96%, P = 0.269), the overall upward trend in MIC values reflected the evolutionary trend of drug resistance in Candida albicans from RVVC patients. This finding is consistent with previous studies16,17, suggesting that repeated infections may promote the selective colonization of drug-resistant strains. Importantly, 5-fluorocytosine maintained 100% sensitivity in both groups, indicating its potential as a salvage therapy for RVVC.
Regarding resistance mechanisms, the elevated MICs of azoles (e.g., fluconazole) may be associated with ERG11 gene mutations and overexpression of CDR1/CDR2 efflux pumps18. Although the expression of ALS1, ALS3, and HWP1 genes in drug-resistant strains trended upward, it did not reach significance, suggesting that cell wall protein genes may have an indirect rather than direct association with drug resistance. This indirect effect may be mediated by biofilm formation—drug-resistant strains exhibited significantly higher biofilm OD values than sensitive strains (P = 0.042), and biofilm structures enhance drug tolerance through physical barriers and metabolic heterogeneity19.
Biofilm formation is a key hallmark of Candida albicans pathogenicity. The biofilm OD values of strains from RVVC patients (0.31 ± 0.13) were significantly higher than those from colonizers (0.19 ± 0.11, P < 0.001), and 66% of RVVC strains showed strong biofilm formation ability, far exceeding 38% in colonizers. This directly confirms the central role of biofilms in RVVC pathogenesis—dense biofilm structures not only enhance fungal adhesion to host mucosa20 but also block drug penetration through the extracellular matrix21, leading to clinical treatment failure.
In terms of virulence factors, the expression levels of SAP, PL, and Lip hydrolases in RVVC strains were significantly higher than those in colonizer strains (all P < 0.05). Secreted aspartyl proteinase (SAP) degrades host mucosal proteins and disrupts the epithelial barrier, while phospholipase (PL) and lipase (Lip) promote fungal invasion by decomposing host cell membrane phospholipids and lipids. Pz value analysis showed that 82% of SAP and 94% of PL in RVVC strains were highly expressed, consistent with the high invasive phenotype of clinical isolates22. Notably, hydrolase expression was positively correlated with biofilm formation (e.g., r = -0.116 between SAP-Pz and Biofilm-OD), suggesting that virulence factors may synergize with biofilms to enhance the pathogenic potential of Candida albicans.
ALS1, ALS3, and HWP1, as hyphal-specific cell wall protein genes of Candida albicans, all showed high expression in RVVC strains (all P < 0.001). The relative expression level of ALS3 reached (4.32 ± 1.91)×10⁵, approximately 1.9 times that of colonizers, confirming its core role in adhesion and hyphal differentiation. Gene correlation analysis showed a high positive correlation among the three genes (r > 0.6), suggesting they may be regulated by common transcription factors such as Efg1 and Cph123. This coordinated expression pattern ensures the efficiency of morphological switching and biofilm construction during infection.
The high expression of cell wall protein genes and enhanced biofilm formation further support the pathogenic cascade of “adhesion-hyphalization-biofilm”24. ALS1 and ALS3 mediate initial adhesion to host cells, while HWP1 maintains the three-dimensional structure of biofilms by participating in hyphal cell wall cross-linking. The low correlation between ALS1/ALS3/HWP1 and Biofilm-OD (r = 0.324–0.343) in this study may be because transcriptional regulation was only the initial step of biofilm formation, with subsequent multi-level regulations involving protein translation, localization, and extracellular matrix assembly25.
Although biofilm OD values were significantly higher in drug-resistant strains, this study found no direct association between drug resistance and hydrolase expression (all P > 0.05), possibly due to the diversity of resistance mechanisms. For example, azole resistance is primarily driven by ERG11 mutations and efflux pump activation, while hydrolase expression is more regulated by environmental nutrient status26. Additionally, the upward trend in ALS gene expression in drug-resistant strains (e.g., ALS3: t = 1.897, P = 0.074), suggests a potential biological association that may not have reached statistical significance due to the limited sample size. This trend aligns with the role of ALS3 in biofilm stabilization, implying that enhanced ALS3 expression could indirectly contribute to drug tolerance by strengthening biofilm structure—an association worthy of validation in larger cohorts, suggests that cell wall proteins may indirectly contribute to drug resistance by enhancing the biofilm resistance barrier, which requires verification with a larger sample size.
Correlation analysis showed moderate correlation between antifungal drug MIC values and Biofilm-OD (r = 0.410–0.567) and low correlation with ALS genes (r = 0.215–0.347), further supporting biofilm as one of the main drivers of drug resistance. Clinically, therapeutic strategies targeting biofilms, such as combination therapy with farnesol (a quorum-sensing inhibitor that disrupts biofilm maturation2, patulin (a natural compound that inhibits extracellular matrix synthesis), or quercetin (a biofilm disruptor that enhances azole penetration10, (such as combined use of biofilm disruptors) may become a new direction for RVVC management. Moreover, the high correlation between ALS3 and HWP1 (r = 0.646) suggests that combined interventions targeting these two genes may more effectively inhibit biofilm formation.
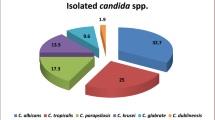
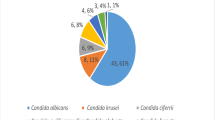
This study has several limitations: first, only Candida albicans was analyzed, while the proportion of non-albicans Candida (such as Candida glabrata and Candida tropicalis) is increasing in clinical cases, with recent reports indicating that these cases account for about 20%3. This limits the generalizability of our findings to the broader population of RVVC, and future studies should include these species for a comprehensive analysis; second, the regulatory mechanisms of gene expression, such as the activity changes of transcription factors Efg1 and Cph1, were not deeply explored; additionally, differences may exist between in vitro results and in vivo infection models, which need verification with animal experiments.
In conclusion, this study systematically revealed the drug resistance characteristics, biofilm formation ability, and virulence factor expression patterns of Candida albicans from RVVC patients, confirming the key roles of enhanced biofilm formation and upregulated expression of cell wall protein genes in RVVC recurrence. Our findings provide new insights into the pathogenic mechanisms of RVVC and lay an experimental foundation for developing antifungal strategies targeting biofilms and virulence factors. In clinical practice, personalized therapy combining drug susceptibility testing with biofilm/virulence factor detection may be an effective approach to improve RVVC prognosis.
Data availability
All data generated or analysed during this study are included in this published article.
Abbreviations
- RVVC:
-
Recurrent vulvovaginal candidiasis
- SAP:
-
Aspartyl proteinases (SAP)
- PL:
-
Phospholipases
- Lip:
-
Lipases
- ALS1:
-
Agglutinin-like sequence 1
- ALS3:
-
Agglutinin-like sequence protein 3
- HWP1:
-
Hyphal wall protein 1
- MIC:
-
Minimum inhibitory concentration
References
Vandecruys, P., Baldewijns, S., Sillen, M. & Genechten, W. V. Oteseconazole: a long-awaited diversification of the antifungal arsenal to manage recurrent vulvovaginal candidiasis (RVVC). Expert Rev. Anti-infe. 21 (8), 799–812. https://doi.org/10.1080/14787210.2023.2233696 (2023).
Nikoomanesh, F. et al. Combination of Farnesol with common antifungal drugs: inhibitory effect against Candida species isolated from women with RVVC. Medicina 59 (4), 743. https://doi.org/10.3390/medicina59040743 (2023).
Faustino, M., Ferreira, C. M. H., Pereira, A. M. & Carvalho, A. P. Candida albicans: the current status regarding vaginal infections. Appl. Microbiol. Biotechnol. 109 (1), 91. https://doi.org/10.1007/s00253-025-13478-2 (2025).
Gonçalves, B. et al. Vulvovaginal candidiasis: Epidemiology, microbiology and risk factors. Crit. Rev. Microbiol. 42 (6), 905–927. https://doi.org/10.3109/1040841X.2015.1091805 (2016).
Fernandes, Â. et al. Vulvovaginal candidiasis and asymptomatic vaginal colonization in portugal: Epidemiology, risk factors and antifungal pattern. Med. Mycol. 60 (5), myac029. https://doi.org/10.1093/mmy/myac029 (2022).
Talapko, J. et al. Candida albicans- the virulence factors and clinical manifestations of infection. J. Fungi. 7 (2), 79. https://doi.org/10.3390/jof7020079 (2021).
Fule, S. R., Das, D. & Fule, R. P. Detection of phospholipase activity of Candida albicans and Non albicans isolated from women of reproductive age with vulvovaginal candidiasis in rural area. Indian J. Med. Microbiol. 33 (1), 92–95. https://doi.org/10.4103/0255-0857.148392 (2015).
Bonfim-Mendonça, P. S. et al. Different expression levels of ALS and SAP genes contribute to recurrent vulvovaginal candidiasis by Candida albicans. Future Microbiol. 16, 211–219. https://doi.org/10.2217/fmb-2020-0059 (2021).
Roudbarmohammadi, S. et al. ALS1 and ALS3 gene expression and biofilm formation in Candida albicans isolated from vulvovaginal candidiasis. Adv. Biomed. Res. 5, 105. https://doi.org/10.4103/2277-9175.183666 (2016).
Gao, M., Wang, H. & Zhu, L. Quercetin assists fluconazole to inhibit biofilm formations of fluconazole-Resistant Candida albicans in in vitro and in vivo antifungal managements of vulvovaginal candidiasis. Cell. Physiol. Biochem. 40 (3–4), 727–742. https://doi.org/10.1159/000453134 (2016).
McCall, A. D., Pathirana, R. U., Prabhakar, A., Cullen, P. J. & Edgerton, M. Candida albicans biofilm development is governed by cooperative attachment and adhesion maintenance proteins. NPJ Biofilms Microbiomes. 5 (1), 21. https://doi.org/10.1038/s41522-019-0094-5 (2019).
Ardehali, S. H. et al. Molecular detection of ALS1, ALS3, HWP1 and SAP4 genes in Candida genus isolated from hospitalized patients in intensive care Unit, Tehran, Iran. Cell. Mol. Biol. 65 (4), 15–22. https://doi.org/10.1016/j.mimet.2018.02.010 (2019).
Sherif, M. M., Elkhatib, W. F., Khalaf, W. S. & Elleboudy, N. S. Abdelaziz N. A.Multidrug resistant acinetobacter baumannii biofilms: evaluation of Phenotypic-Genotypic association and susceptibility to cinnamic and Gallic acids. Front. Microbiol. 12, 716627. https://doi.org/10.3389/fmicb.2021.716627 (2021).
Lan, D. M. et al. H.A novel cold-active lipase from Candida albicans: cloning, expression and characterization of the Recombinant enzyme. Int. J. Mol. Sci. 12 (6), 3950–3965. https://doi.org/10.3390/ijms12063950 (2011).
Yazdanpanah, S. et al. Exploring the anti-biofilm and gene regulatory effects of anti-inflammatory drugs on Candida albicans. N-S Arch. Pharmacol. 398 (6), 7263–7272. https://doi.org/10.1007/s00210-024-03727-y (2025).
Tortorano, A. M., Prigitano, A., Morroni, G., Brescini, L. & Barchiesi, F. Candidemia: evolution of drug resistance and novel therapeutic approaches. Infect. Drug Resist. 19, 14:5543–5553. https://doi.org/10.2147/IDR.S274872 (2021).
Esfahani, A. et al. Molecular epidemiology, antifungal susceptibility, and ERG11 gene mutation of Candida species isolated from vulvovaginal candidiasis: comparison between recurrent and non-recurrent infections. Microb. Pathog. 170, 105696. https://doi.org/10.1016/j.micpath.2022.105696 (2022).
Navarro-Rodríguez, P., Martin-Vicente, A., López-Fernández, L., Guarro, J. & Capilla, J. Expression of ERG11 and efflux pump genes CDR1, CDR2 and SNQ2 in voriconazole susceptible and resistant Candida glabrata strains. Med. Mycol. 58 (1), 30–38. https://doi.org/10.1093/mmy/myz014 (2020).
Banerjee, A. et al. Structure, function, and Inhibition of catalytically asymmetric ABC transporters: lessons from the PDR subfamily[J]. Drug Resist. Update. 71, 100992. https://doi.org/10.1016/j.drup.2023.100992 (2023).
McKloud, E. et al. Recurrent vulvovaginal candidiasis: a dynamic interkingdom biofilm disease of Candida and Lactobacillus. Msystems. 6(4): e0062221. https://doi.org/10.1128/mSystems.00622-21 (2021).
Martorano-Fernandes, L., Goodwine, J. S., Ricomini-Filho, A. P., Nobile, C. J. & Cury, A. A. D. B. Candida albicans adhesins Als1 and Hwp1 modulate interactions with Streptococcus mutans. Microorganisms 11 (6), 1391. https://doi.org/10.3390/microorganisms11061391 (2023).
Freire, F. et al. Evaluation of gene expression SAP5, LIP9, and PLB2 of Candida albicans biofilms after photodynamic inactivation. Laser med sci. 30, 1511–1518. https://doi.org/10.1007/s10103-015-1747-0 (2015).
Freire, F., de Barros, P. P., Pereira, C. A., Junqueira, J. C. & Jorge, A. O. C. Photodynamic inactivation in the expression of the Candida albicans genes ALS3, HWP1, BCR1, TEC1, CPH1, and EFG1 in biofilms. Laser Med Sci. 33, 1447–1454. https://doi.org/10.1007/s10103-018-2487-8 (2018).
Nobile, C. J. et al. Complementary adhesin function in C. albicans biofilm formation. Curr. Biol. 18 (14), 1017–1024. https://doi.org/10.1016/j.cub.2008.06.034 (2008).
Liu, C., Xu, C., Du, Y., Liu, J. & Ning, Y. Role of agglutinin-like sequence protein 3 (Als3) in the structure and antifungal resistance of Candida albicans biofilms. FEMS microbiol. lett. 368 (14), fnab089. https://doi.org/10.1093/femsle/fnab089 (2021).
Fan, Y., He, H., Dong, Y., Pan, H. & Hyphae-specific genes HGC1, ALS3, HWP1, and ECE1 and relevant signaling pathways in Candida albicans. Mycopathologia. 176, 329–335 https://doi.org/10.1007/s11046-013-9684-6 (2013).
Acknowledgements
We sincerely thank the staff of the Department of Gynecology and Microbiology Laboratory at the Third Affiliated Hospital of Sun Yat-sen University (Yuedong Hospital) and Meizhou People’s Hospital. We also express our heartfelt gratitude to the staff of the Comprehensive Laboratory of the Medical College of Jiaying University.
Author information
Authors and Affiliations
Contributions
Experiments and research, Guangwen Xiao and Jing Zhang; Data analysis and statistics, Yanhua Yang; Writing—original draft, Caixia Yan and Guangwen Xiao; Case materials and strain providers, Xuehua Zeng and Caixia Yan; All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
This study was approved by the Ethics Committee of Jiaying College of Medicine.This study is not applicable to an informed consent statement.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yan, C., Zhang, J., Yang, Y. et al. Virulence factors, biofilm formation and antifungal resistance in Candida albicans from recurrent vulvovaginal candidiasis patients: a comparative study. Sci Rep 15, 37557 (2025). https://doi.org/10.1038/s41598-025-21846-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-21846-1