Abstract
Pneumonia is a severe respiratory ailment that may be caused by viruses, fungus, and bacteria. Pneumonia causes the accumulation of water, purulent material, or other fluids in the air sacs (alveoli) of the lungs. A delay in the identification of pneumonia may be life-threatening. Nevertheless, the limited radiation levels used for diagnosis result in unreliable detection, posing a significant obstacle in the field of pneumonia detection in healthcare. Transfer Learning (TL) may revolutionize healthcare by offering an effective method for differentiating between normal and pneumonia patients. Therefore, we have proposed a TL utilizing model for predicting pneumonia. Our experiment uses 5856 highly imbalanced chest X-ray images to fit different vision models such as Xception, VGG16, and ResNet152V2 using TL approach. After training, our model performs exceptionally well on the X-ray dataset, achieving an accuracy of 80.45, 80.77, and 83.17% respectively. However, the results show that the ResNet152V2 performs exceptionally well as compared to other models. Also, it achieves a precision and recall score of 79.87 and 97.69% respectively. The results exemplify the potential of our framework to help pulmonologists and physicians make rapid diagnoses of pneumonia patients by providing accurate and fast pneumonia classification.
Similar content being viewed by others
Introduction
Respiratory infections remain a major clinical concern globally. Any difficulty with the lungs disrupts the regular process of breathing, and the individual may face a range of severity from minor to life-threatening consequences. Since these organs are vital for the healthy functioning of the human body, any issue in the lungs must be detected early, as it prevents patients from developing life-threatening diseases1,2. Pneumonia is one of the life-threatening respiratory infections, which affects a certain section of a population, especially children and elderly individuals with weak immune systems2. To this end, artificial intelligence (AI) may play an important role in the early and precise identification of pneumonia cases along with the analysis of chest X-rays3. Several advanced AI models based on deep learning (DL) and transfer learning (TL) have emerged as a promising tool in the healthcare sector4. These models are capable of identifying specific patterns and abnormalities that are indicative of pneumonia. By leveraging deep convolutional neural networks (DCNN) with multiple layers, healthcare professionals can achieve precise and rapid identification of pneumonia, decide the line of treatment, and improve the diagnostic process4. Moreover, integration of these technologies into the healthcare sector has the potential to automate diagnostics, reduce errors, and provide a cost-efficient framework.
UNICEF reports that pneumonia claims the life of a child in every 43 second 5. The research states that pneumonia is the leading cause of mortality among children, accounting for 700,000 deaths each year (or around 2,000 per day, including 190,000 infants). That works out to one case of pneumonia in every seventy-one kids. About 18 million more healthcare personnel will be required by 2030 to identify, prevent, and treat pneumonia. Individuals who are susceptible to pneumonia also include those who are 65 years of age or older and individuals with prior medical conditions especially taking chemotherapy.
Chest X-ray is a frequently used diagnostic tool for identifying pneumonia6,7,8. For example, computer-aided detection and diagnosis (CAD) systems can assist radiologists by improving diagnostic accuracy1. In this view, Computed Tomography (CT) scans are very effective in identifying pneumonia, in addition to chest X-rays, sputum cultures and biopsies9,10. CT scans use higher levels of radiation and are associated with higher costs10. The benefits of CT scans in identifying pneumonia must be considered with the hazards of excessive radiation exposure. An increased risk of cancer is associated with longer-term usage of higher-dose radiation or more CT scans. Sputum cultures are effective in identifying the specific microorganisms, but delay in culture findings may prolong therapeutic intervention. Biopsies or bronchoscopy is used for accurate pathogen and inflammatory pattern detection allows individualized treatment regimens for better patient outcomes, but lung biopsies are invasive and may cause infection and pneumothorax11.
Nevertheless, the minimal radiation levels used for diagnostic purposes result in imprecise detection. Deep learning and transfer learning are essential in identifying patterns in images to accurately categorize them as either normal or abnormal. With each additional layer, the system identifies patterns that span the whole picture, including the boundaries, edges and individual dots. TL’s capacity to apply information from pre-trained models on big datasets to smaller datasets makes it promising for pneumonia detection in medical imaging. TL can be used to develop strong and effective pneumonia detection models on small annotated data by transferring learnt characteristics from datasets with copious annotated pictures. This strategy reduces the requirement for large labeled datasets, which are limited and expensive in medical imaging, and improves model generalization and performance by capturing complex data patterns and representations. Thus, TL provides a compelling answer to pneumonia diagnosis issues, improving healthcare outcomes and resource-efficient diagnostic operations.
The following is a layout of the sections used to detail the entire study. Section II provides a thorough overview of prior studies that have investigated the use of DL and TL techniques in the diagnosis of pneumonia. In Section III, we explained each method and dataset used in this study. In section IV, the experimental setup is described. Section V delves into the measures used to evaluate the method’s performance. Section VI presents the synthesis of experimental results about chest X-ray data for pneumonia. In the end, section VII concludes the manuscript.
Related work
When applied to the medical field, AI has the potential to revolutionize patient care by facilitating rapid and precise diagnosis of illness. To handle the differences within and across classes, Peng et al.12 presented a multiscale residual network, where one channel receives the raw CT image, while the other receives the differential excitation component, allowing them to build a multi-scale residual network. They successfully manage inter-class variances by integrating multi-scale information from the CT scan picture. To further address intra-class variances, the model injects the multi-scale residual network with the CT scan image and differential excitation component. A remarkable 93.74% classification accuracy was attained by their method. The authors in1 investigated the effectiveness of a Quaternion Residual Network trained on a publicly available large Chest X-Ray dataset from the Kaggle repository. The study reported a classification accuracy of 93.75% and an F-score of 0.94, highlighting the model’s potential in diagnosing chest-related conditions1. A previous study examined the use of a hybrid deep learning model that combines quantum algorithms with neural networks to detect pneumonia from chest X-ray images. The proposed ZFNet-quantum neural network analyzed 5863 X-ray scans and achieved an accuracy of 96.5%, surpassing the traditional CNN model, which reached 94% accuracy, highlighting the potential of quantum-assisted deep learning in medical imaging2.
Shi et al.13 proposed radiographic imaging to distinguish COVID from pneumonia, which used the attention-transfer-based categorization model. The foundation of this model is the distillation network architecture, in which students learn from their teachers. The teacher’s network function is built using attention that maps by focusing on critical areas and extracting global characteristics from images using the deformable attention module. Finally, after integrating an image fusion module with the instructor network’s attention information, the student network begins to collect input. A new DCNN and InceptionV3 model was presented by Rajeashwari and Arunesh14 to address the issue of accurate pneumonia prediction. Following that, they noticed that the model’s accuracy was relatively low. To raise it, they employed a Neighborhood Component Analysis (NCA) technique that relies on entropy normalization. They also proposed Ensemble-Modified Classifiers (EMC), which included Random Forest, Naïve Bayes, and XGBoost for the final classification, to further enhance the model’s accuracy to 98.5%.
In their study, Bhandari et al.15 focused on improving feature extraction to improve the model’s accuracy. They did it in two stages. First, they utilized convolutional feature extraction to find high-level object information. Next, they employed the Shapley Additive exPlanation (SHAP), Local Interpretable Model-agnostic Explanation (LIME), and Gradient-weighted Class Activation Mapping (Grad-CAM) approaches. This helped them achieve test and validation accuracies of 94.31 ± 1.01% and 94.54 ± 1.33, respectively, using tenfold cross-validation. They used explainable AI to further validate the model, .
When CNN is trained on an unbalanced, overfitted, or noisy dataset, problems such as significant variations in training and validation occur. After preprocessing the input image, a hierarchical template-matching algorithm is applied to reduce noise and improve features, Kaya16 suggested an ideal feature fusion-based ensemble CNN model. The presented model used a pre-defined CNN with initial weights assigned using different vision models utilizing TL along with majority voting. Following these procedures, the model achieved a 98.94% accuracy and a 99.12% F1 score on a publicly accessible dataset.
To diagnose COVID-19 using an X-ray dataset and further distinguish it from pneumonia, Mohagheghi et al.17 suggested a DCNN model. They employed image retrieval techniques for distinction and a DCNN model for diagnosis. Also, they improved the accuracy using transfer learning and hashing methods, which resulted in a 97% accuracy. In a comparative investigation, Avola et al.18 applied 12 unique pre-trained ImageNet models to distinguish between healthy individuals and patients with pneumonia. Due to dataset scarcity, they pooled two datasets into a single one consisting of 6,330 images. They then trained the model using 100%, 50%, 20%, and 10% of the dataset, respectively, and documented the results.
An experiment was conducted with a dataset on pediatric pneumonia. Prakash et al.19 presented a DCNN architecture-based channel attention to extract image features. Next, they performed principal component analysis (PCA) on each feature to reduce dimensionality, and finally, they added all the features. Additionally, TL-based designs such as Xception, ResNet152V2, DenseNet169, and ResNet50V2 utilize the channel attention mechanism. For pneumonia prediction, an ensemble-based stacking method is used using traditional ML models, including Nu-SVC, XGBClassifier, K-Nearest Neighbour, Support Vector Classifier, and Logistic Regression. With scores of 96.15% for accuracy, 97.19% for precision, 95.00% for recall, 96.49% for AUC-score, and 96.24% for precision, the stacking mechanism produces outstanding results across the board.
In35 the authors use machine learning techniques to identify fibrosis, malignant lymph, metastases and normal lymphogram. Their comparative study demonstrates the capabilities of ML to enhance the accuracy of diagnostic tasks of complex medical imaging. In36 the authors present CovXNet, a multi-dilation multi-receptive field-optimized convolutional neural network-based COVID-19 and pneumonia detector based on the chest X-ray dataset. The model is very accurate and flexible in a variety of datasets in early pandemic diagnostics. The work in37 examined how the Internet of Medical Things (IoMT) can be used in healthcare monitoring and decision-making. It highlights the importance of inter-linked medical devices in enhancing patient care and distance healthcare provision. The authors in38 suggest a hybrid deep neural network framework called RVCNet to diagnose lung diseases. Their findings demonstrate a better robustness of results and diagnostic accuracy of these compared to conventional deep learning models. The paper39 used densely linked convolutional networks in the diagnosis of infectious diseases. The model offers more efficient and better representation of features by the process of rethinking with dense connections to enhance clinical diagnostics. Finally, authors in40 predicts the COVID-19 and ICU needs using predictive modeling. It gives practical information about the resource planning of the hospital and pandemic management.
Proposed framework
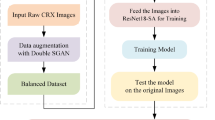
The proposed framework is described in detail below, along with technical details of the TL used in20. For a clear understanding, we will first understand the background details and then delve into the specifics of transfer learning techniques21. Our approach constructs a multi-step predictive tool that can accurately analyze the X-ray image and predict pneumonia with a faster response. Image data collection, data preparation and pre-processing, reconstruction of transfer learning architecture, and model refinement via the addition of layers like dense, batch normalization, and dropout layers are all part of the process22,23. Figure 1 presents the design of the proposed paradigm. Below are the series of procedures used in the proposed approach:
Workflow diagram for the Proposed Transfer Learning System for Pneumonia Classification.
Step 1: The analysis starts with a pneumonia dataset, which consists of 2 classes one is normal and another is pneumonia.
Step 2: During the pre-processing stage, to manage the uniformity we have taken the image size as (512, 512), To standardize the image we have used rescaling, for a clear visibility of the objects in the lungs we have adjusted the brightness of the images. For contrasting adjacent pixels, enhancing the visibility and definition of borders and features we have used sharpening filter.
Step 3: We have rebuilt the CNN model by adding some layer according to our need. After the architecture is ready for classifying the pneumonia, input the image augmentation right after the input layer.
Step 4: The layers like dropout, batch normalization, and dense layers are added to carry out the fine-tuning procedure. The purpose of this is to enhance the model for pneumonia classification.
Step 5: We have included three vision models, namely, Xception, VGG16, and ResNet152V2 for initial weight initialization and further image classification.
Step 6: After training of the models, we have compared the performance of all the transfer learning models and choose the best one using metrics like execution time, average memory utilization, accuracy, precision, recall, confusion matrix, and F1-score. Additionally, a thorough evaluation is conducted in relation to preexisting research.
Transfer learning framework
-
Xception: It is a convolutional neural network design based on the Inception model, uses a succession of depth-wise separable convolution layers with residual connections24. With the exception of the initial and final blocks, which are connected via linear residuals, the 36 convolutional layers are organized into 14 blocks25. These blocks make it famous for capturing picture characteristics at a fine granularity while keeping the computational cost down. The goal of this approach is to develop a parameter- and computationally-efficient model that represents data’s spatial and channel-wise relationships efficiently. Changing or understanding the architecture is a breeze. Just like VGG-16, it uses a high-level framework like Keras or TensorFlow Slim, which reduces the coding need to 30 to 40 lines. In comparison, advanced designs, such as Inception V2 or V3, are difficult to understand.
-
VGG16: VGG16, created in 2014 by the Visual Geometry Group (VGG) at Oxford University, is the name of one of the most recent dimensionality reduction algorithms26. With its 16 weight layers of uniform simplicity, this architecture is composed of 13 convolutional layers and 3 fully linked layers. By using 3 × 3 filters with a stride of 1, zero-padding, and max-pooling layers, this VGG16 design effectively decreases spatial dimensions24. Although VGG16 has a lot of parameters, it can still pull off impressive feature extractions. This architecture is ideal for use as a foundational model for picture categorization and object recognition due to its high efficacy in learning generic characteristics from photos.
-
ResNet152V2 Architecture: A novel deep neural network called ResNet was developed to address the disappearing gradient issue during training by allowing gradients to flow across the network22. With an astonishingly low error rate of just 3.57%, the ensemble achieved first place in the ILSVRC 2015 classification competition, proving beyond a reasonable doubt that this model is effective27. There are many variants that use the same basic idea but with a different number of layers. An alternative known as Resnet50 may function with 50 neural network layers. To make the model more accurate, deep residual networks use residual blocks. “Skip connections” are the fundamental idea behind residual blocks with bottleneck architecture, which is a resilient neural network topology. On the ImageNet dataset, ResNet152V2, a variant of ResNet, beats all previous versions of ResNet. To improve the transfer of information between blocks, ResNet152V2 features an upgraded inter-block connection topology. Training stability and computational efficiency are both improved by this design compared to its predecessor, using its optimized residual connections and “bottleneck” building blocks. The ResNet152V2 framework usually takes a tensor representation of an image as its input. The size of the input picture and the number of channels determine the dimensions of this tensor.
ResNet152V2 is made up of several convolutional layers. The input picture is passed through a series of filters by each convolutional layer, which extracts features at various abstraction levels. A convolutional layer is mathematically expressed as28:
In the above Eq. (1), δ denotes the activation function (such as ReLU), \({W}_{n,m,k}^{l}\) denote the convolutional filter’s weights, \({X}_{i+n-1,j+m-1,k}^{l-1}\) denotes the input activation at position \(i+(n-1,j+m-1,k)\) in the preceding layer \(l-1\), and \({b}_{i,j}^{l}\) indicates the bias term. \({H}_{i,j}^{l}\) is the output activation at position \(\left(i,j\right)\) in layer \(l\).
In order to facilitate the flow of gradients during training, ResNet topologies make use of residual blocks that include skip connections, also called identity mappings. The bottleneck block, with its three convolutional layers that use batch normalization and ReLU activations, is the fundamental building component of ResNet152V2. The mathematical representation of a residual block’s output is stated in Eq. (2) as below:
Hence, for every input \(x\), \(F\) stands for the block’s internal layers’ residual function.
As observed from Eq. (3), the weight matrices of the convolutional layers are denoted as \({(w}_{3},{w}_{2},{w}_{1}\)), the bias terms are \({b}_{1},{b}_{2}\).
In the latter stages of the network, the feature maps’ spatial dimensions are reduced to a vector by using global average pooling. To explain it, global average pooling takes all of the feature maps and averages them as given in Eq. (4) below:
The activation at location \(\left(i,j\right)\) in the \({k}^{th}\) feature map is represented as \({x}_{ij}^{k}\), while the feature maps’ height and breadth are denoted by \(H\) and \(W\), respectively.
The prediction for the classes is generated using a fully connected layer followed by a softmax activation. Logits \(z\) given in Eq. (5) below is calculated mathematically in this layer by first applying a linear transformation to the input and then using the Eq. (6) for softmax function:
where \({W}_{ijc}\) and \({b}_{ijc}\) are the weights and biases of the fully connected layer, respectively, and C is the number of classes. Exponentiation of logits is performed to make sure that the outputs are proper probability distributions with values between 0 and 1. The step-wise workflow of the proposed framework is depicted in Algorithm 1.

TL-based classification of pneumonia using chest X-ray images.
System model
The TL utilizing models are large and complex, having been pre-trained using a substantial amount of ImageNet data28. To exercise such models, we need to preprocess the data and fine-tune the model so that it can perform efficiently for the intended task29. Also, we need to reconstruct the output layer according to usage. There are two important procedures that we need to follow to use the TL:
Image augmentation
This is used in ML to modify or enrich images to expand the volume and diversity of training data14. Image augmentation is defined as the process of preparing image data for further processing by applying transformations such as rotation, cropping, shearing, flipping, scaling, and changing contrast or brightness to the original images30. This process will focus on the key image portion and omit the unwanted image part. In the process, the initial step adds the image enhancement layer, ensuring that the augmentation layer will be an integral part of the design. This method enables the modified system structure to use the GPU’s increased processing capability by using preprocessed pictures to augment the device in tandem with the other layers. Additionally, the pretreatment layers are kept together with the remaining model when it is extracted. This approach eliminates the need to re-implement this functionality on the server by automatically standardizing images according to the layer settings.
Adding layers for our use
After Augmentation and image transformations, we have used the original TL model as used except for the last layer, where we implement the layer according to our usage31. Incorporating more layers is made possible by this selective retention, which optimizes the design’s efficacy and enhances predictions in pneumonia classification32,33,34. However, overfitting may occur if there are too many additional layers.
To properly tune the model, we improved its architecture by adding layers tailored to the pneumonia dataset. Now the model is ready and can take in picture datasets and make predictions. We have included a portion of the dropout layer to prevent the model from overfitting. We are taking certain characteristics into account and ignoring the others by using the dropout layer. The improvements made to the suggested framework are detailed in the following sections: To integrate the basic model’s output, the Global Average Pooling2D Layer is used. Two Dense Layers, one with 512 neurons and the other with 64 neurons, are used in conjunction with Batch Normalization. Following the Global Average Pooling2D layer, we add a dense layer with 512 neurons, a dropout layer, and Batch Normalization. Relu is the activation function, and 1377 is the seed value for the kernel initializer, which is set to ‘glorot uniform’. By default, the bias initializer is set to zero. After that, we add another Batch Normalization layer. Next, we add a dense layer with 512 neurons and an activation function (we use ReLu) as well as batch normalization adjusted to ``Global Average Pooling2D” layer. Two neurons, one for each class, create the last dense layer for prediction. Because this is a multi-label classification task, the ‘softmax’ activation function is used. The bias initializer is set to ``zeros”, whereas the kernel initializer is set to “random uniform”.
We have taken into account the pre-trained trainable weights to utilize the learning strength of the current model. To calculate the step size at each iteration during training, the built architecture uses the ‘Adamax’ optimizer with a learning rate of 0.001. We use the ‘sparse categorical cross-entropy’ loss function, which finds the discrepancy between the expected probability distribution and the real category labels. To find the overall loss, it first determines the loss for each class individually in each iteration and then adds all losses together. Using a single integer instead of a whole vector to describe a class maximizes the efficiency of memory and computation.
Performance evaluation
To evaluate the performance of any ML or DL model there are basic evaluation matrices such as accuracy, F1-score, confusion matrix (CM), precision, recall, execution time, and average memory consumption. A brief description of these metrics is given below:
Confusion matrix
The CM is the basic fundamental technique matrix which is used to evaluate for haw many data the model predicts properly. It is a representation in tabular form that provide insights into four distinct combinations of actual and predicted values: True Positive (\({{\varvec{t}}}_{{\varvec{P}}}\)), False Positive (\({{\varvec{f}}}_{{\varvec{P}}}\)), True Negative (\({{\varvec{t}}}_{{\varvec{N}}}\)), and False Negative (\({{\varvec{f}}}_{{\varvec{N}}}\)). The CM, helps to evaluate the reliability of the model also from this we will be able to find the valuable information on accuracy, loss, recall, precision, and F1-score.
Accuracy
Accuracy is the first statistical measure for any ML models, which is the ratio of expected correct outcomes to actual observations. For the purpose of evaluating a model’s overall performance and dependability, the accuracy metric is used. The accuracy can be calculated as Eq. (7) below:
Precision
For the purposes of predictive modelling, precision is the ratio of anticipated positive cases to actually predicted positive cases. This ratio technically shows the model’s ability to recognize positive findings reliably. The precision can be calculated using the expression in Eq. (8) as follows:
Recall
To accurately identify all positive cases out of the total real positive instances by the model is called recall. It helps to identify proportion of actual predictions by the models. Models having high recall value indicates that the models don’t overlook real positive instances and it is minimizing the likelihood of false negatives. If the model having high recall value pulmonologists and clinicians rely on models for diagnose of the disease. Mathematically recall can be computed as Eq. (9):
F1-score
F1-score is another reliable statistic used to evaluate the performance of a model. It is computed by harmonically averaged of the accuracy and recall scores. By combining precision and recall, we may get a more complete picture of how effectively a model can detect positive examples with few false positives and negatives. The F1-score can be calculated as follows using Eq. (10):
Execution time
If results cannot be utilized within the specified time range, they are useless. Thus, the model’s execution time can guarantee its practical usefulness. From initial data loading and training to final prediction or classification, the whole time required for a model to process and analyze the provided data is known as execution time. The execution time \({T}_{EX}\) can be computed by the difference between the end time \({T}_{E}\) and the start time \({T}_{E}\) provided in Eq. (11):
Average memory utilization
Average memory utilization is a measurement of a model’s efficiency in amount of memory resource consumption during its execution. It can be computed from Eq. (12) by dividing the total memory resource consumed \({Total}_{mem}\) by the total number of data points or instances processed \(N\):
Loss
In DCNN, we used stochastic gradient descent optimization algorithm for training and updating the weights for each node we used backpropagation. The model then learns using those weights and try to make the predictions and it compares with the actual value and then each times the weights are updated using backpropagation and the error is calculated. This error is called as loss and the formula for the loss can be obtained from Eq. (13) as:
where NC is the number of classes and two distributions are, \(true {(T}_{j})\) and predicted \({(P}_{j})\).
Result and discussions
Dataset description and pre-processing
The dataset used for the experiment contains 5,856 chest X-ray images (both anterior and posterior) of two categories, labelled as normal or pneumonia. The dataset was collected from Kaggle32. The dataset was originally collected from a cohort of pediatric patients of one to five years old at the Guangzhou Medical Center, Guangzhou, China. These X-ray were performed as the routine clinical trials of children. Not all of the images include only the chest, but some have a lot more than merely the upper or lower part of it. The images classify all the bacterial and virus-infected lungs as pneumonia. These images were screened for quality control and also verified by the expert physicians. A more detailed information about the data collection and preparation process can be found in the dataset paper by Daniel et al.32. The limited scope of the current study is also the restriction to only pediatric patients because the manifestation of diseases and physiological reactions may vary greatly among the age groups. Although the findings provide important insights into patterns in pediatric settings, it is still not clear whether they can be generalized to the adult and the elderly populations. Subsequent studies must thus focus on the use of high-quality and large-scale datasets that capture larger demographic cohorts and allow assessment of age-related variation and guarantee the generalizability and strength of the constructed models to the entire range of patients. Such extension would not only confirm the relevance of the existing approach but increase its clinical applicability in a variety of healthcare settings to a greater extent.
In the preprocessing phase, we limit the batch size to 16 to process only 16 images in each epoch. Then, we applied an image transformation before providing it to the model to enhance the accuracy. Rescaling of the X-ray images to the interval [0,255] ranges from black to white, which standardizes the pixel intensity values. To improve consistency of the model’s train and test phases, it’s important to standardize the intensity between [0,1]. One may simulate a broad range of lighting conditions, from very dark to extremely light, by adjusting the brightness of the X-ray images. For this study, we specified the brightness within the range of [0.9, 1.3], which accounts for the variations in brightness. Some of the image information can be lost due to compression, which can be resolved with a blurred lens or noise. We have also included a sharpening filter to restore the distorted information, which can improve model predictability.
Experimental set-up
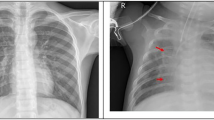
The computer that was used for this study has a 2.10 GHz power of CPU, 16 GB of RAM, and is associated with the i7 12th gen Intel processor. The simulation was carried out using Python 3.10.9, with libraries Tensorflow, Keras, Pandas, Numpy, and Matplotlib used for preprocessing, implementing the suggested transfer learning method, and for performance evaluation. Figure 2 represents some sample Chest X-ray images for normal and pneumonia patient. All the vision models that we leveraged in our analysis; we compared their performance using the widely-used confusion matrix (CM) as depicted in Fig. 3. We have shown the CM for Xception, VGG16, and ResNet152V2 in Fig. 3a–c respectively. It is evident from CM that ResNet152V2 produced a higher number of correct predictions.
Some Sample chest X-ray images for normal and pneumonia patient.
Confusion Matrices for (a) Xception (b) VGG16 and (c) ResNet152V2.
Table 1 shows performance parameters for three prominent TL models that we have used in our study: Xception, VGG16, and Resnet152V2. Reset152V2 earned the highest F1-score, accuracy, and precision as compared to VGG16 and Xception. ResNet152V2 is getting better than the others for accuracy, precision and f1-score matrix by showing 83.17, 79.87, and 80% respectively, but VGG16 is leading for recall which is of 99.23%. By observing the comparison, it turns out that Resnet152V2 has the impressive performance in all forms.
We measured the models’ performances after 50 epochs of training using the following metrics: training accuracy, precision, loss, recall, and F1-score. Knowing that, with the exception of loss, we tracked the rate of curve convergence to one for all of the evaluation matrices. A model is considered performing well if its curve either reaches one or approaches one. Figure 4a shows that ResNet152V2 converges to a local maximum faster than other models and even surpasses Xception and VGG16, which were evaluated considering training accuracy only. Regarding model assessment, the loss curve and the loss value are the most vital metrics, where loss is a numerical measure of the difference between the model’s predictions and the actual observations. As the number of epochs increases, the rate at which the ResNet152V2 loss value declines are faster than other models, as depicted in Fig. 4b. Such results confirm that the ResNet152V2 improves its predictive ability over time.
(a) Training accuracy curves for Xception, VGG16, and ResNet152V2 over different number of epochs (b) Training loss curves for Xception, VGG16, and ResNet152V2 over different number of epochs.
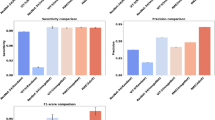
Precision and Recall curves is used to evaluate how accurately a model can identify or predict the true positives. From the Fig. 5a, b we have presented the comparison of precision and recall curve for Xception, VGG16, and ResNet152V2. From the both graph it is clearly shows that the ResNet152V2 converges to a local maximum more quickly than other models, and it even beats Xception and VGG16 with the growing number of epochs. Taking the F1-score into consideration was necessary due to the extreme imbalance in the image data that we have used for our study. The comparison of the F1-score for all the models is shown in Fig. 5c. The results demonstrate that ResNet152V2 outperforms Xception and VGG16 with respect to the F1-score.
(a)Training precision curves for Xception, VGG16, and ResNet152V2 over different epochs. (b) Training recall curves for Xception, VGG16, and ResNet152V2 over different epochs (c) Training F1-score curves for Xception, VGG16, and ResNet152V2 over different epochs.
After 50 epochs, we have measured the validation performance of each model using the following metrics: validation accuracy, precision, loss, recall, and f1-score. Knowing that, with the exception of loss, we need to track the rate of curve convergence to one for all of the evaluation matrices. We may say that a model is good if its curve either converges to one or is rapidly approaching towards one. Figure 6a shows that ResNet152V2 converges to a local maximum more quickly than other models, and it even surpasses Xception and VGG16, which were simply evaluated in terms of validation accuracy. The rate at which the ResNet152V2 method loss value declines is much faster comparing to the other models as the number of epochs increases as depicted in Fig. 6b. Such result confirms that the ResNet152V2 model is improving its predictive abilities over time.
(a) Validation accuracy curves for Xception, VGG16, and ResNet152V2 over different Epochs. (b) Validation loss curves for Xception, VGG16, and ResNet152V2 over different Epochs.
From the Fig. 7a, b we have presented the comparison of validation precision and recall curve for Xception, VGG16, and ResNet152V2. From the both graph it is clearly shows that the ResNet152V2 converges to a local maxima more quickly than other models, and it even beats Xception and VGG16 with the growing number of epochs. The comparison of the validation f1-score for all the models is shown in Fig. 7c. The results demonstrate that ResNet152V2 outperforms Xception and VGG16 with respect to the F1-score.
(a) Validation Precision for Xception, VGG16, and ResNet152V2 over different number of Epochs. (b) Validation Recall Curves for Xception, VGG16, and ResNet152V2 over different number of Epochs. (c) Validation F1 Score Curves for Xception, VGG16, and ResNet152V2 over different number of Epochs.
After all these observations we have consider an overall precision, accuracy, recall, and F1-scores for Xception, VGG16, and ResNet152V2, which will give a clear idea about the performance of each and every models. In Fig. 8a It is observe that the ResNet152V2 outperform all others in terms of all the performance metrics that we have considered. The execution time for building the Xception, VGG16, and ResNet152V2 is shown in Fig. 8c. It can be observed that the Xception takes the lowest execution time, followed by the ResNet152V2 and VGG16 model over the considered chest X-ray dataset. In Fig. 8, the average memory utilization for Xception, VGG16, and ResNet152V2 is presented. From the Fig. 8b, it is clearly observed that the Xception model consume the minimal space, followed by the ResNet152V2 and VGG16 models. As the time and space consumed by the ResNet152V2 is quite high, we can consider some more preprocessing technique to reduce the space and time consumption by this model. Table 2 provides a detailed summary of the space and time values for each vision model.
(a) Comparison of precision, accuracy, recall, and F1-score for Xception, VGG16, and ResNet152V2, (b) Comparison of mean space consumed (in bytes) for Xception VGG16, and ResNet152V2, (c) Comparison of execution time (in ms) for Xception, VGG16, and ResNet152V2.
The results of the pairwise t-test between AUC values of the three models, namely, Xception, VGG16, and ResNet152V2 are shown in Table 3 with a 95% confidence interval (CI). According to the results, there is statistical significance between the models. The comparison between Xception and VGG16 shows t-statistics of 10.9032 and p-value < 0.0001 to be the largest and the CI of the difference between the means was observed to be 0.0370 to 0.0557 does not intersect with zero, which indicates that Xception is significantly better than VGG16. The comparison between Xception and ResNet152V2 also shows a very high difference with a t-statistics of 4.4611, p-value of 0.0004, and the CI is positive, which lies between 0.0079 to 0.0264, indicating that Xception has higher AUC values than ResNet152V2. Furthermore, the mean difference between VGG16 and ResNet152V2 is negative with t-statistics of − 6.1725, and the p-value is 0.0001, also the CI is negative and lies between − 0.0352 and − 0.0232, meaning that ResNet152V2 is much better than VGG16.
Figure 9 provided below shows the ROC curves and the value of the AUC of the three considered transfer learning models namely Xception, VGG16, and ResNet152 V2 when applied to the same classification task. The ROC curve of the Xception model, with an AUC of 0.9700, is shown in sub Fig. 9a, which results in high discriminative performance and high ability to tell the difference between positive and negative classes. In Fig. 9b the VGG16 model with AUC of 0.9200 which corresponds to a good performance of classification, even though it is less than Xception. The trade-off between sensitivity and specificity has a moderate value. Figure 9c shows the ROC curve of ResNet152V2, the AUC of which is 0.9400, which is better than VGG16 but lesser than Xception, demonstrating that it has strong classification capacity.
The AUC-ROC Curve for (a) Xception, (b) VGG16, and (c) ResNet152V2.
Conclusion
Our study used Xception, VGG16, and ResNet152V2 to test the TL-based models for pneumonia patient classification. The presented model includes a preprocessing layer, TL-based framework , fine-tuning, and 5,856 chest X-ray datasets with significant imbalances. We evaluated the presented model using precision, recall, f1-score, accuracy, loss, confusion matrix, time, and space complexity. With an accuracy score of 83.17% for ResNet152V2, 80.77% for VGG16, and 80.45% for Xception, our tested models performed well. After analyzing performance and a thorough investigation, we found that ResNet152V2 performed better than the other methods and popular techniques. The experiment demonstrated the power of TL in tackling pneumonia detection difficulties and established that ResNet152V2 performed better than the other models. This contributes to better patient outcomes and more effective healthcare delivery by improving the early identification and classification of pneumonia patients. By reliably and quickly identifying cases of pneumonia, the ResNet152V2 model may help pulmonologists and doctors make correct diagnoses quickly. It also has the potential to properly categorize individuals as either normal or pneumonia patient.
Future works
Our study highlights the importance of using TL towards classification of Pneumonia using Chest X-ray dataset. However, only relying on a theoretical prediction framework may not resolve the issues associated with appropriate diagnosis and treatment plan for pneumonia, hence posing difficulty on healthcare providers. Thus, possible future research in this direction would be the design of an advanced healthcare monitoring platform which could keep track of the patients’ ailments and their relevant prescribed medications in conglomeration with healthcare professionals’ supervision. The accuracy of the transfer learning model can improve by using sophisticated strategies like attention mechanisms and adversarial training by concentrating the relevant regions of the chest X-ray instead of whole image. Furthermore, explainable AI techniques can be used to make deep learning model more transparent and easier to understand, which will give both doctors and patients greater faith in the diagnosis process. In light of ResNet152V2’s outstanding experimental performance, future work may focus on increasing the model’s optimization and fine-tuning, data preprocessing, and the inclusion of relevant regions within chest X-ray images in order to achieve even higher performance, improve diagnostic reliability, and reduce space and response time.
Data availability
The real datasets used in the current study was collected at the Guangzhou Medical Center, Guangzhou, China and available in the Kaggle repository. It was accessed using the following link: [https://www.kaggle.com/datasets/paultimothymooney/chest-xray-pneumonia] (Accessed on 5th March 2025).
Change history
14 December 2025
The original online version of this Article was revised: The original version of this Article erroneously assigned Dr Shakir Khan Affiliation 5: College of Computer and information Sciences, Imam Mohammad Ibn Saud Islamic University (IMSIU), Riyadh, Saudi Arabia. Additionally, the original version of the Article erroneously assigned Dr Mohd Fazil and Dr Abdullah M. Albarrak Affiliation 2: University Centre for Research and Development, Chandigarh University, Mohali, India, when they are associated only with Affiliation 5: College of Computer and Information Sciences, Imam Mohammad Ibn Saud Islamic University (IMSIU), Riyadh, Saudi Arabia.
References
Singh, S. & Tripathi, B. K. Pneumonia classification using quaternion deep learning. Multimedia Tools Appl. 81 (2), 1743–1764 (2022).
Shahwar, T., Mallek, F., Rehman, A. U., Sadiq, M. T. & Hamam, H. Classification of pneumonia via a hybrid ZFNet-quantum neural network using a chest X-ray dataset. Curr. Med. Imaging. 20(1), e15734056317489 (2024).
Chang, T. H. et al. Clinical characteristics of hospitalized children with community-acquired pneumonia and respiratory infections: Using machine learning approaches to support pathogen prediction at admission. J. Microbiol. Immunol. Infect.56(4), 772 – 81. (2023).
Liu, Y. N. et al. Infection and co-infection patterns of community-acquired pneumonia in patients of different ages in China from. to 2020: a national surveillance study. The Lancet Microbe. 4(5), e330-9. (2009).
https://data.unicef.org/topic/child-health/pneumonia/https://data.unicef.org/topic/child-health/pneumonia/
Ippolito, D. et al. Artificial Intelligence Applied to Chest X-ray: A reliable tool to assess the differential diagnosis of lung pneumonia in the emergency department. Diseases. 11(4), 171. (2023).
Sheu, R. K. et al. Interpretable classification of pneumonia infection using eXplainable AI (XAI-ICP). IEEE Access. 11, 28896–28919 (2023).
Akbulut, Y. Automated pneumonia based lung diseases classification with robust technique based on a customized deep learning approach. Diagnostics. 13(2), 260. (2023).
Wang, Y. et al. Prediction of viral pneumonia based on machine learning models analyzing pulmonary inflammation index scores. Comput. Biol. Med. 169, 107905. (2024).
Wang, W., Zhao, X., Jia, Y. & Xu, J. The communication of artificial intelligence and deep learning in computer tomography image recognition of epidemic pulmonary infectious diseases. Plos One. 19 (2), e0297578 (2024).
Dash, A. K., Mohapatra, P. & Ray, N. K. Pneumonia detection in children from chest X-ray images by executing network surgery of deep neural networks. SN Comput. Sci.. 5(2), 1–3. (2024).
Peng, L. et al. Classification and quantification of emphysema using a multi-scale residual network. IEEE J. Biomed. Health Inform. 23(6), 2526–2536. (2019).
Shi, W., Tong, L., Zhu, Y. & Wang, M. D. COVID-19 automatic diagnosis with radiographic imaging: explainable attention transfer deep neural networks. IEEE J. Biomedical Health Inf. 25 (7), 2376–2387 (2021).
Rajeashwari, S. & Arunesh, K. Enhancing pneumonia diagnosis with ensemble-modified classifier and transfer learning in deep-CNN based classification of chest radiographs. Biomed. Signal Process. Control. 93 ,106130. (2024).
Bhandari, M., Shahi, T. B., Siku, B. & Neupane, A. Explanatory classification of CXR images into COVID-19, pneumonia and tuberculosis using deep learning and XAI. Comput. Biol. Med. 150, 106156 (2022).
Kaya, M. Feature fusion-based ensemble CNN learning optimization for automated detection of pediatric pneumonia. Biomed. Signal Process. Control. 87, 105472 (2024).
Mohagheghi, S. et al. of CNN, CBMIR, and visualization techniques for diagnosis and quantification of covid-19 disease. IEEE J. Biomed. Health Inform. 25(6), 1873-80. (2021).
Avola, D. et al. Study on transfer learning capabilities for pneumonia classification in chest-X-rays images. Comput. Methods Program Biomed. 221, 106833. (2022).
Asswin, C. R. et al. Transfer learning approach for pediatric pneumonia diagnosis using channel attention deep CNN architectures. Eng. Appl. Artif. Intell. 123, 106416. (2023).
Ansari, N., Faizabadi, A., Motakabber, S. & Ibrahimy, M. Effective pneumonia detection using res net based transfer learning. Test Eng. Manag. 82, 15146-53. (2020).
Madhavan, M. V. et al. Res-CovNet: an internet of medical health things driven COVID-19 framework using transfer learning. Neural Comput. Appl. 35 (19), 13907–13920 (2023).
Kumar, N., Gupta, M., Gupta, D. & Tiwari, S. Novel deep transfer learning model for COVID-19 patient detection using X-ray chest images. J. Ambient Intell. Humaniz. Comput. 14 (1), 469–478 (2023).
Pham, T. A., Hoang, V. D. & Chest X-ray image classification using transfer learning and hyperparameter customization for lung disease diagnosis. J. Inform. Telecommun. 5, 1–5. (2024).
Luján-García, C., Villuendas-Rey, Y. & Camacho-Nieto, O. A transfer learning method for pneumonia classification and visualization. Appl. Sci. 10(8), 2908. (2020).
HashmiMF, Katiyar, S., Keskar, A. G., Bokde, N. D. & Geem, Z. W. Efficient pneumonia detection in chest Xray images using deep transfer learning. Diagnostics 10 (6), 417 (2020).
Jain, R., Nagrath, P., Kataria, G., Kaushik, V. S. & Hemanth, D. J. Pneumonia detection in chest X-ray images using convolutional neural networks and transfer learning. Measurement 165, 108046 (2020).
Elshennawy, N. M. & Ibrahim, D. M. Deep-pneumonia framework using deep learning models based on chest X-ray images. Diagnostics. 10(9), 649. (2020).
Mohamed, C., Mwangi, R. W. & Kihoro, J. M. Enhancing Pneumonia Detection in Pediatric Chest X-Rays Using CGAN-Augmented Datasets and Lightweight Deep Transfer Learning Models. J. Data Anal. Inform. Process. 12(1), 1–23. (2024).
Koul, A., Bawa, R. K. & Kumar, Y. An analysis of deep transfer learning-based approaches for prediction and prognosis of multiple respiratory diseases using pulmonary images. Arch. Comput. Methods Eng. 31(2), 1023-49. (2024).
Gu, C. & Lee, M. D. Transfer learning using real-world image features for medical image classification, with a case study on pneumonia X-ray images. Bioengineering. 11(4), 406. (2024).
Yoon, T. & Kang, D. Enhancing pediatric pneumonia diagnosis through masked autoencoders. Sci. Rep. 14 (1), 150 (2024).
Daniel, S. K., Michael, G., Wenjia, C., Anthony, L. Identifying medical diagnoses and treatable diseases by image-based deep learning. Cell. 172(5), 1122–1131 (2018).
Abou-Kreisha, M. T. et al. ElDahshan. Multisource smart computer-aided system for mining COVID-19 infection data. Healthcare. 10, 109 (2022).
Fathy, K. A., Humam, K., Yaseen, Mohammad, T., Abou-Kreisha & Kamal, A. ElDahshan. A novel meta-heuristic optimization algorithm in white blood cells classification. Comput. Mater. Continua. 75, 1527–1545. (2023).
Bharati, S., Robel, M. R. A. & Podder, M. A. and Niketa Gandhi. Comparative performance exploration and prediction of fibrosis, malign lymph, metastases, normal lymphogram using machine learning method. In: International Conference on Innovations in Bio-Inspired Computing and Applications, pp. 66–77. Cham: Springer International Publishing, (2019).
Mahmud, T. & Shaikh Anowarul, F. Md Awsafur Rahman, and CovXNet: A multi-dilation convolutional neural network for automatic COVID-19 and other pneumonia detection from chest X-ray images with transferable multi-receptive feature optimization. Comput. Biol. Med. 122, 103869. (2020).
Mahmud, T. Applications for the Internet of medical things. Int. J. Sci. Res. Arch. 10, 1247–1254. (2023).
Alam, F., Binte, Podder, Rubaiyat Hossain, M. & Mondal Prajoyand RVCNet: A hybrid deep neural network framework for the diagnosis of lung diseases. Plos one. 18, e0293125. (2023).
Podder, Prajoy, F. B., Alam, M. R. H. & Mondal Md Junayed Hasan, Ali Rohan, and Subrato Bharati. Rethinking densely connected convolutional networks for diagnosing infectious diseases. Computers. 12, 95. (2023).
Podder, P., Khamparia, A. & Mondal, M. R. H. Mohammad Atikur Rahman, and Subrato Bharati. Forecasting the spread of COVID-19 and ICU Requirements. (2021).
Acknowledgements
This work was supported and funded by the Deanship of Scientific Research at Imam Mohammad Ibn Saud Islamic University (IMSIU) (grant number IMSIU-DDRSP2501).
Funding
This work was supported and funded by the Deanship of Scientific Research at Imam Mohammad Ibn Saud Islamic University (IMSIU) (grant number IMSIU-DDRSP2501).
Author information
Authors and Affiliations
Contributions
Conceptualization, Biswajit Tripathy (B.T), Shakir Khan (S.K), Sujit Bebortta (S.B), Ashraf Kamal (A.K.), and Subhranshu Sekhar Tripathy (S.S.T.); Software, A.K. and A.M.A.; validation, B.T., S.B, S.S.T., Mohd Fazil (M.F.) data curation, S.K., S.B., A.K., writing—original draft preparation, B.T. , A.K. and A.M.A.; writing—review and editing, S.K., S.B, S.S.T., M.F.; visualization, B.T, S.B, S.S.T., A.K., S.K.; supervision, B.T.; project administration, S.K.; funding acquisition, S.K.,M.F.; All authors have read and agreed to the published this manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Tripathy, B., Khan, S., Bebortta, S. et al. TL-PneuNet: a transfer learning-based pneumonia classification framework. Sci Rep 15, 40307 (2025). https://doi.org/10.1038/s41598-025-24180-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-24180-8