Abstract
Since the recent application of metagenomic next-generation sequencing (mNGS) techniques to clinics, Torquetenoviruses (TTV) have received much attention due to their high positive rates. However, there is an insufficient understanding in clinical settings of the pathogen, especially in immunocompromised patients. This study explores the clinical characteristics of TTV infection in immunocompromised patients using mNGS. We enrolled a total of 120 TTV-infected patients in the study, including 81 immunocompromised and 39 immunocompetent individuals. The prevalence, diagnosis, treatment, and co-pathogens were compared between the two groups. The microbial diversity and presence of co-pathogens in patients infected with Torquetenovirus (TTV) were elucidated through comprehensive analysis. T-tests compared the normally distributed continuous data. The immunocompromised patients exhibited significantly elevated TTV loads, and a notable proportion of these patients also presented with hematopoietic disorders. Importantly, our investigation revealed that current treatments showed no efficacy against TTV infection.Furthermore, the presence of copathogens such as Staphylococcus, Bacillus, Mycobacterium, and Acinetobacter was observed in TTV-infected individuals. Immunocompromised patients exhibited a significantly higher abundance of Staphylococcus and Shewanella compared to immunocompetent patients (p < 0.05). Cautious use of antiviral therapy is recommended for patients with TTV mono-infection. However, greater attention should be given to co-pathogens, such as Staphylococcus spp. and Shewanella spp. This cohort study provides valuable insights into the clinical significance of TTV infection, particularly in immunocompromised patients. We found that TTV is frequently detected in this population, often with higher viral loads and an increased burden of co-pathogens. These findings suggest that TTV may serve primarily as a marker of immune dysfunction rather than as a sole pathogen. Incorporating TTV monitoring into mNGS-based diagnostics could help identify high-risk patients, support early intervention, and guide tailored management strategies in immunocompromised settings.
Similar content being viewed by others
Introduction
Torquetenovirus (TTV) is a small, single-stranded circular DNA virus that was first discovered in 19971. The virus has a genome size of 3.8 kilobases (kb) and contains at least four open reading frames (ORFs) and several smaller ORFs. The largest ORF spans from 389 to 2593 base pairs2,3,4. TTV is known to be highly diverse and can be divided into different genotypes based on more than 30% sequence divergence and different gene groups based on more than 50% sequence divergence5,6,7. It has been detected in various organs, tissues, and body fluids, suggesting that it can infect a wide range of cell types in humans. TTV can be transmitted through multiple routes, including blood transfusion, fecal-oral, and mother-to-child transmission. It is highly prevalent in the general population and is considered a part of the human virome8,9.
In general, traditional virus culture methods are time-consuming and labor-intensive, making them insufficient for timely clinical decision-making. Previous studies have reported cases such as TTV-associated aseptic meningitis, where immunological methods were used to detect IgM antibodies against the GP2 antigen in patient serum, providing indirect evidence of TTV infection10. This case was cited as a representative example to highlight the challenges in establishing TTV’s clinical relevance, particularly when its presence is detected in sterile body sites. However, immunological methods are virus-specific and cannot comprehensively detect all TTV genotypes. Due to the extensive genetic variability of TTV, traditional PCR methods targeting the ORF region may miss many strains, while the 5′ untranslated region is more conserved and more suitable for primer design. Despite this, the unclear pathogenicity of TTV has limited the development and clinical adoption of commercial detection kits11. In contrast, metagenomic next-generation sequencing (mNGS) offers an unbiased approach that does not require prior knowledge of the pathogen and is capable of identifying a wide spectrum of viral populations. Thus, mNGS serves as a promising alternative for investigating the presence and potential clinical impact of viruses like TTV, including in complex clinical presentations such as aseptic meningitis12.
With the increasing use of metagenomic next-generation sequencing (mNGS) in clinical settings, Torque teno virus (TTV) is being detected more frequently, particularly in immunocompromised individuals8. However, its pathogenic potential and clinical relevance remain uncertain. Current evidence suggests that TTV may be more prevalent in individuals with impaired immune function, yet its role in disease progression and outcomes has not been clearly established13. Moreover, the effectiveness of antiviral therapies against TTV remains unclear, raising concerns about appropriate clinical management strategies14,15,16,17. TTV is often detected alongside other bacterial, viral, or fungal pathogens, complicating interpretation and highlighting the need to consider its interactions within the broader microbial community. Therefore, this study aims to investigate the relationship between TTV load and immune status, evaluate the clinical outcomes of TTV infection—especially in immunocompromised patients—and assess the necessity and focus of antiviral treatment in cases of TTV mono-infection, as well as its potential utility as a biomarker for immune dysfunction.
Materials and methods
Patient and study design
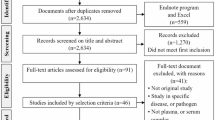
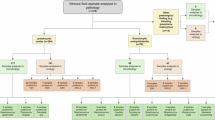
In this retrospective study, a total of 1,650 patients underwent mNGS testing at Zhujiang Hospital, Southern Medical University (Guangzhou, China) from June 2019 to June 2022. We enrolled 120 TTV-positive patients who met the following inclusion criteria: (1) age between 18 and 70 years, regardless of gender; (2) suspected lower respiratory tract or bloodstream infection; (3) complete testing data and case information. The study was approved by the ZhuJiang Hospital’s ethics committee (Southern Medical University,2019-KY-029-05).A small number of patients who did not sign the informed consent were granted a waiver by the ethics committee. In general, We enrolled 81 immunocompromised patients in total. Besides, there is also a control group of 39 immunocompetent patients with TTV infection enrolled. We set criteria for immunodeficiency: (1) rheumatic diseases, solid tumors, and hematologic malignancies; (2) long-term systemic corticosteroid use (0.3 mg/kg/day of prednisone equivalent for 3 weeks) and immunosuppressive medications (including chemotherapy for malignancies, but excluding corticosteroids); (3) solid organ transplantation; and (4) hematopoietic stem cell transplantation. The enrollment flowchart of this study is shown in Fig. 1.
Flow chart of TTV in immunocompromised and immunocompetent groups.
Sample collection
3 mL of blood was drawn from patients, prior to plasma separation, the blood sample was placed in a collection tube and kept at room temperature for 3–5 minutes. Subsequently, it was centrifuged at 4,000 rpm for 10 min at 4°C, with all processing completed within 8 h of collection. DNA was extracted from 300 uL of plasma using the TIANamp Micro DNA Kit (DP316, TIANGEN BIOTECH, Beijing, China) following the manufacturer’s operational manual.
Construction of DNA libraries and sequencing
All clinical samples were used to construct the sequence library after DNA fragmentation, end-repair, adapter-ligation, and PCR amplification. We used the QubitTM dsDNA HS Assay (Invitrogen, USA) to control the quality of the DNA libraries. Minimum of 1 ng/µL DNA was required to proceed to library preparation, and approximately 40–80 ng of DNA was used per sample for sequencing, depending on the sample yield.Single-stranded circular. DNA nanoballs were produced by rolling circle amplification (RCA) and then loaded on an MGISEQ-2000 platform for sequencing18.
Bioinformatic analysis
After high-throughput sequencing, the data were processed by quality control, adaptor filtering, and host sequence removal using Burrows–Wheeler Alignment algorithms. We first used fastp to filter the raw sequencing data19. Hisat2 (version 2.2.1), for mapping the reads to the human genome GRCh38 with default parameters, was used to remove the human reads20. Next, the remaining data were aligned to the Pathogens metagenomics Database (PMDB), consisting of approximately 16,000 species of bacteria, fungi, viruses, and parasites for pathogen identification. The classification reference databases were downloaded from NCBI (ftp://ftp.ncbi.nlm.nih.gov/genomes/), which provides RefSeqs for taxonomically diverse organisms, including archaea, bacteria, eukaryotes, and viruses. TTV were specifically quantified as reads per million (RPM) of the total reads in the library. Other microbial species were identified based on the analysis of the clean reads by Kraken221. A microorganism was considered positively detected with 2 than 2 reads remained after normalization of 120 samples. The detection was consistent with the patient’s clinical symptoms and presentation.
For microbial diversity analysis, alpha diversity (Shannon’s index, Simpson index and Richness), beta diversity (principal coordinate analysis, PCoA) based on Bray-Curtis metrics were generated by R (version 4.2.2). Venn diagram showing the number of common and unique OTUs between the two groups was made by online tool (http://bioinformatics.psb.ugent.be/webtools/Venn/).
Statistical analysis
Data were processed using SPSS 23.0 software version 9.4 (SAS Institute, Inc., Cary, NC, USA). Categorical variables were compared using the Chi-square or Fisher’s exact test. Normally distributed data were analyzed with the Student’s t test, and non-normally distributed data with the Mann–Whitney U test. P-values < 0.05 were considered statistically significant.
Results
Clinical characteristics, laboratory examination, and diagnosis of TTV immunocompromised patients
The clinical characteristics are presented in Table 1. The immunocompetent group was older, with no significant gender differences. In the routine blood tests, the leukocyte count in the immunocompetent group was significantly higher than that in the immunocompromised group (p < 0.001). However, the neutrophil count was higher in the immunocompetent group. There were no significant differences in lymphocyte count. Regarding TTV-associated conditions, we also focused on fever, pneumonia, bloodstream infections, and hepatitis. Pneumonia showed no significant differences between the groups, while hepatopathy diseases showed significant differences. To be noted, there were no significant differences in hepatic dysfunction or hepatitis between the groups.
Distribution and semiquantitative analysis in TTV -infected patients
A total of 120 TTV-infected patient samples were available for subsequent analysis. The number of TTV cases in immunocompromised patients was higher than in the immunocompetent group. Additionally, reads or reads per million mapped reads (RPM) is an important semi-quantitative index of viral load and is used for disease monitoring in NGS22,23. In both blood and BALF samples, the RPM of TTV infection in the immunocompromised group was significantly higher than that in immunocompetent patients (p < 0.001) (Fig. 2). Our data demonstrate that immunocompromised patients are more likely to carry the virus, and the RPM of TTV is significantly greater in this population.
Abundances of TTV in different samples. in immunocompromised and immunocompetent patients. (A) Log10(RPM) of TTV in BALF samples of immunocompromised (red) and immunocompetent patients (green). (B) Log10(RPM) of TTV in blood samples of immunocompromised (blue) and immunocompetent patients (yellow). RPM: The reads per million mapped.
Microbial diversity in TTV-infected patients
To evaluate the co-pathogens that may associate with TTV, we investigated the microbial diversity and composition across all samples. To ensure a consistent sequencing depth for microbial analysis, each sample was normalized to 12,785 reads(minimum number of reads for all samples), capturing the broadest range of microorganisms present. In the BALF samples, a Venn diagram illustrated that out of 2267 identified genera, 1265 were common to both immunocompromised and immunocompetent patients, with 810 genera uniquely identified in the former and 192 in the latter (Fig. 3A). In blood samples, 1195 out of 2164 total genera were shared between the two patient groups, with 686 genera unique to immunocompromised patients and 283 unique to immunocompetent patients (Fig. 3B). α diversity analysis at genus level showed no significant difference in microbial diverisity and richness between the two groups in either blood or BALF samples (p > 0.05, Figure S1,Table S1). Principal coordinate analysis (PCoA) using Bray-Curtis metrics indicated that β diversity at the genus level in blood samples were significantly different between immunocompromised and immunocompetent patients (p < 0.05, Fig. 3C), a pattern not observed in BALF samples (p > 0.05, Fig. 3D). This suggests that the blood microbiota composition may be more indicative of the immunocompromised conditions than BALF microbiota.
Summary of Taxonomic breakdown and β diversity in TTV-infected patients. (A) The Venn diagram based on the microbial composition at genus level in BALF samples of immunocompromised (purple) and immunocompetent patients (pink). (B) The Venn diagram based on the microbial composition at genus level in blood samples of immunocompromised (purple) and immunocompetent patients (pink). (C) PCoA based on Bray-Curtis indices of blood samples immunocompromised (red dot) and immunocompetent patients (green dot). (D) PCoA based on Bray-Curtis indices of BALF samples immunocompromised (red dot) and immunocompetent patients (green dot).
The composition of dominant microbial groups (top 20 abundant genera in different groups of patients. Samples groups: blood samples of immunocompromised and immunocompetent patients, BALF samples of immunocompromised and immunocompetent patients, average of all groups.
Co-pathogens in TTV-infected patients
We conducted a comparative analysis of the relative abundance of the top 20 genera between different patient groups, as depicted in Fig. 4. Collectively, Staphylococcus, Bacillus, Mycobacterium, Acinetobacter, Shewanella and Pseudomonas comprised a substantial portion, (averaging 52.4% across all samples) within the microbial composition. Notable differences were observed when comparing immunocompromised and immunocompetent patients in both blood and BALF samples. For blood samples, Staphylococcus and Shewanella were significantly more abundant in immunocompromised patients (25.6% vs. 8.4%, 10.4% vs. 0.3%, respectively; p < 0.05). Conversely, the presence of Bacillus (15.8% vs. 21.3%), Mycobacterium (7.2% vs. 12.2%), Acinetobacter (6.2% vs. 9.5%), Pseudomonas (3.0% vs. 4.4%), Escherichia (3.0% vs. 4.8%), Toxoplasma (2.6% vs. 4.1%), Streptococcus (0.8% vs. 1.1%), Campylobacter (1.4% vs. 2.4%), Klebsiella (1.5% vs. 2.2%), Enterococcus (1.5% vs. 2.2%), Nocardia (0.8% vs. 1.3%) and Candida (0.03% vs. 0.07%) were less in the blood samples of the immunocompromised patients (p < 0.05). In BALF samples of the immunocompromised patients, Staphylococcus and Shewanella also showed a significant increase in relative abundance (17.4% vs. 5.9%, 3.7% vs. 0.15%; p < 0.05; respectively). Bacillus, Prevotella, Rothia and Streptococcus were less abundant, however the differences were not statistically significant (p > 0.05).
Antiviral therapy statistics in TTV infection patients.
Impact of antimicrobial treatment of TTV infections
For patients with high TTV viral load and clinical manifestations, doctors will initiate antiviral treatment based on their clinical experience.Through retrospective analysis, it was found that 47% (38/81) of immunocompromised patients used antiviral drugs, which is higher than 15% (6/39) of immunocompetent patients, indicating that immunocompromised patients are more inclined to use antiviral therapy in clinical practice. Statistical analysis was conducted to compare the rates of improvement and deterioration between the two groups in anti-viral drug therapy. The result showed no statistical difference between the two groups (Fig. 5). Through retrospective analysis, a total of 13 patients underwent two mNGS tests before and after treatment, and no significant assistance was found in inhibiting TTV load among existing antiviral drugs (Table S2).
Discussion
In recent years, mNGS has emerged as an unbiased molecular diagnostic tool for detecting and studying microorganisms in biological samples, greatly improving the comparability and reliability of test results between different laboratories24,25,26. In this study, we used mNGS to detect TTV in immunocompromised and immunocompetent patients. We first compared TTV patients in different kinds of situations including immune deficiency disease, solid tumors, hematologic malignancies, long-term systemic corticosteroid use, solid organ transplantation, and so on. Our finding shows that the case number of immunocompromised patients is higher than that of immunocompetent patients. In addition, TTV appears inversely associated with immune competence. RPM is calculated as the number of reads per gene divided by the number of single-mapping reads per sample library times one million. We found that the RPM in immunocompromised patients appears higher than that in immunocompetent patients. The possible explanations are similar to the higher positive rate in immunocompromised patients.
The mechanisms underlying TTV co-infections have not yet been fully elucidated. However, studies have suggested that TTV may facilitate the infection of other pathogens by modulating the host immune system For example, TTV ORF2 protein may inhibit the NF-kappaB signaling pathway, thereby affecting the host’s immune response27,28. Earlier studies on TTV co-infection were mainly focused on TTV and hepatitis virus co-infection29,30. This idea was gradually reformed as new data on co-infection with other viruses became available. TTV has been detected in co-infections with many other viral species28,29,,29,31,32. In this study, the most notable differences in co-pathogen abundance were observed with Staphylococcus and Shewanella, which were more prevalent in immunocompromised patients. Staphylococcus a globally widespread gram-positive bacterium, is associated with significant mortality and morbidity, capable of causing severe infections such as skin and soft tissue infections (SSTIs), endocarditis, sepsis, pneumonia, osteomyelitis, bacteremia, abscesses, and toxic shock syndrome33. All analytical approaches and sequencing protocols employed in our study are well-established and supported by peer-reviewed publications.Our findings indicated that Staphylococcus was the dominant microorganism in both blood and BALF samples from immunocompromised patients, corroborating prior studies33,34. Given the increasing prevalence of antibiotic-resistant strains of Staphylococcus, such as those resistant to vancomycin and methicillin35,36, infection management should be adapted, for example, potentially incorporating monoclonal antibodies, either as monotherapy or in combination37. The genus Shewanella, a group of Gram-negative proteobacteria, includes several species known to cause serious community- and hospital-acquired infections38, leading to conditions such as SSTIs, sepsis, hepatobiliary diseases, otitis media, and their complications39. Currently, there is no consensus on antimicrobial therapy for Shewanella spp. infections in humans, with treatment decisions complicated by the patients’ underlying conditions and the diverse spectrum of diseases. Our data show that co-infections are more prevalent in immunocompromised individuals, which are particularly susceptible to opportunistic infections.
The top 20 genera represented 81.6% of the total microbial presence in blood samples from immunocompromised patients and 71.9% in BALF samples, respectively. These genera were predominantly recognized as pathogens in prior studies immune deficiency35,40. Although certain pathogenic genera appeared less abundant in immunocompromised patients, this may be attributed to the significant rise in the abundances of Staphylococcus and Shewanella. It is conceivable that the absolute abudances of co-pathogens had increased overall, a hypothesis that could be substantiated through absolute quantification methods such as real-time quantitative PCR. Furthermore, given that the composition of blood microbiota provides a more indicative reflection of immunocompromised conditions compared to BALF microbiota, we propose the utilization of blood samples as a preferable diagnostic medium.
Regarding clinical events, we found that the immunocompromised patients in our study seem younger than the immunocompetent patients, the former with lower neutrophil counts, lower incidence of pneumonia, and higher occurrence of hepatopathy/sepsis. TTV has been reported to be associated with major immune function determinants, including age41,42, while others have reported higher TTV in both young and older patients. We also found a higher incidence of pneumonia in immunocompetent patients with TTV infection, which is inconsistent with the literature43, possibly due to the differences in the study population. TTV can replicate in peripheral blood mononuclear cells, such as neutrophilic granulocytes. Our findings suggest that these agents had no apparent impact on TTV positivity or viral load, underscoring the insensitivity of TTV to available antivirals and the need to focus on managing co-infecting pathogens.
Importantly, although we referred to antiviral agents in the context of TTV infections, it is acknowledged that there is no approved or specific antiviral therapy targeting TTV. The term “antiviral therapy” in this study refers to commonly used broad-spectrum antivirals in clinical settings (e.g., ganciclovir, acyclovir), which are primarily administered for other viral infections in immunocompromised patients. We observed the improvement and deterioration in immunocompromised group. Anti-viral drug therapy showed no statistical difference. It might be that TTV is insensitive to current anti-viral therapy26,44,45,46.
This study has several limitations. First, it was conducted at a single center, and the findings need to be validated in larger, multi-center cohorts to enhance generalizability. Second, TTV load was evaluated using metagenomic next-generation sequencing (mNGS), which provides relative abundance rather than absolute quantification. As a result, we were unable to report viral load in standard units such as copies/mL. Additionally, there is currently no specific antiviral therapy for TTV, and its pathogenic role remains unclear. This limits the ability to assess treatment response or develop targeted strategies. Future research should apply standardized quantitative PCR and investigate potential antiviral approaches, with attention to co-infections that may influence clinical outcomes.
Conclusion
In conclusion, our study demonstrates that Torque teno virus (TTV) is frequently detected in immunocompromised patients, where it is often associated with higher viral loads, greater microbial diversity, and an increased burden of co-pathogens. Notably, current antiviral therapies did not significantly influence TTV carriage. These findings suggest that TTV may serve as an indicator of underlying immune dysfunction and interact with other microbial communities rather than acting as a sole pathogen. Its frequent co-detection with bacteria, viruses, and fungi underscores the importance of cautious interpretation of mNGS results. More importantly, monitoring TTV through mNGS may provide additional value for risk stratification, early recognition of immune suppression, and guiding targeted management strategies in immunocompromised patients. Further studies are warranted to validate its utility as a diagnostic or prognostic biomarker in this setting.
Data availability
The datasets generated during the current study are available in CNGB Sequence Archive (CNSA) of China National GeneBank DataBase (CNGBdb) with accession number CNP0005757.
References
Nishizawa, T. et al. A novel DNA virus (TTV) associated with elevated transaminase levels in posttransfusion hepatitis of unknown etiology. Biochem. Biophys. Res. Commun. 241, 92–97. https://doi.org/10.1006/bbrc.1997.7765 (1997).
Hino, S. TTV, a new human virus with single stranded circular DNA genome. Rev. Med. Virol. 12, 151–158. https://doi.org/10.1002/rmv.351 (2002).
Mankotia, D. S. & Irshad, M. Cloning and expression of N22 region of Torquetenovirus (TTV) genome and use of peptide in developing immunoassay for TTV antibodies. Virol. J. 11, 96. https://doi.org/10.1186/1743-422x-11-96 (2014).
Takahashi, K., Hijikata, M., Samokhvalov, E. I. & Mishiro, S. Full or near full length nucleotide sequences of TT virus variants (Types SANBAN and YONBAN) and the TT virus-like mini virus. Intervirology 43, 119–123. https://doi.org/10.1159/000025034 (2000).
Biagini, P. Human circoviruses. Vet. Microbiol. 98, 95–101. https://doi.org/10.1016/j.vetmic.2003.10.004 (2004).
Heller, F., Zachoval, R., Koelzer, A., Nitschko, H. & Froesner, G. G. Isolate KAV: a new genotype of the TT-virus family. Biochem. Biophys. Res. Commun. 289, 937–941. https://doi.org/10.1006/bbrc.2001.6089 (2001).
Reshetnyak, V. I., Maev, I. V., Burmistrov, A. I., Chekmazov, I. A. & Karlovich, T. I. Torquetenovirus in liver diseases: on the way towards unity of view. World J. Gastroenterol. 26, 1691–1707. https://doi.org/10.3748/wjg.v26.i15.1691 (2020).
Focosi, D. & Maggi, F. Torquetenovirus monitoring in transplantation: the quest for standardization. Am. J. Transpl. 19, 1599–1601. https://doi.org/10.1111/ajt.15194 (2019).
Luo, K. et al. Experimental infection of nonenveloped DNA virus (TTV) in rhesus monkey. J. Med. Virol. 61, 159–164. https://doi.org/10.1002/(sici)1096-9071(200005)61:1%3C159::aid-jmv26%3E3.0.co;2-h (2000).
Ikuta, Y. et al. Aseptic meningitis caused by Torquetenovirus in an infant: a case report. J. Med. Case Rep. 13, 302. https://doi.org/10.1186/s13256-019-2233-2 (2019).
Macera, L. et al. Comparative evaluation of molecular methods for the quantitative measure of Torquetenovirus viremia, the new surrogate marker of immune competence. J. Med. Virol. 94, 491–498. https://doi.org/10.1002/jmv.25488 (2022).
Janiak, M. et al. Next-Generation sequencing of hepatitis C virus (HCV) Mixed-Genotype infections in Anti-HCV-Negative blood donors. Adv. Exp. Med. Biol. 1096, 65–71. https://doi.org/10.1007/5584_2018_190 (2018).
Okamura, A. et al. Detection of TT virus sequences in children with liver disease of unknown etiology. J. Med. Virol. 62, 104–108. https://doi.org/10.1002/1096-9071(200009)62:1%3C104::AID-JMV16%3E3.0.CO;2-P (2000).
Garbuglia, A. R. et al. Detection of TT virus in lymph node biopsies of B-cell lymphoma and hodgkin’s disease, and its association with EBV infection. Int. J. Immunopathol. Pharmacol. 16, 109–118. https://doi.org/10.1177/039463200301600204 (2003).
Maggi, F. et al. TT virus in the nasal secretions of children with acute respiratory diseases: relations to viremia and disease severity. J. Virol. 77, 2418–2425. https://doi.org/10.1128/jvi.77.4.2418-2425.2003 (2003).
Miyamoto, M., Takahashi, H., Sakata, I. & Adachi, Y. Hepatitis-associated aplastic anemia and transfusion-transmitted virus infection. Intern. Med. 39, 1068–1070. https://doi.org/10.2169/internalmedicine.39.1068 (2000).
Bando, M. et al. Infection of TT virus in patients with idiopathic pulmonary fibrosis. Respir Med. 95, 935–942. https://doi.org/10.1053/rmed.2001.1151 (2001).
Jeon, Y. J. et al. The feasibility study of non-invasive fetal trisomy 18 and 21 detection with semiconductor sequencing platform. PLoS One. 9, e110240. https://doi.org/10.1371/journal.pone.0110240 (2014).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. Fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890. https://doi.org/10.1093/bioinformatics/bty560 (2018).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915. https://doi.org/10.1038/s41587-019-0201-4 (2019).
Wood, D. E., Lu, J. & Langmead, B. Improved metagenomic analysis with kraken 2. Genome Biol. 20, 257. https://doi.org/10.1186/s13059-019-1891-0 (2019).
Vasudevan, H. N. et al. Comparison of plasma metagenomic Next-generation sequencing and PCR methods for Epstein-Barr virus viral load monitoring in nasopharyngeal carcinoma. Anticancer Res. 44, 5445–5453. https://doi.org/10.21873/anticanres.17370 (2024).
Ai, J. W. et al. Dynamic and direct pathogen load surveillance to monitor disease progression and therapeutic efficacy in central nervous system infection using a novel semi-quantitive sequencing platform. J. Infect. 76, 307–310. https://doi.org/10.1016/j.jinf.2017.11.002 (2018).
Ramachandran, P. S. & Wilson, M. R. Metagenomics for neurological infections - expanding our imagination. Nat. Rev. Neurol. 16, 547–556. https://doi.org/10.1038/s41582-020-0374-y (2020).
Mitchell, A. B. & Glanville, A. R. Kinetics of TTV-DNA plasma load: A global measure of immune suppression? Transplantation 103, 660–661. https://doi.org/10.1097/tp.0000000000002437 (2019).
Wohlfarth, P. et al. Torquetenovirus dynamics and immune marker properties in patients following allogeneic hematopoietic stem cell transplantation: A prospective longitudinal study. Biol. Blood Marrow Transpl. 24, 194–199. https://doi.org/10.1016/j.bbmt.2017.09.020 (2018).
Handala, L. et al. No correlation between Torquetenovirus viral load and BK virus replication after kidney transplantation. J. Clin. Virol. 116, 4–6. https://doi.org/10.1016/j.jcv.2019.03.018 (2019).
Szládek, G. et al. High co-prevalence of genogroup 1 TT virus and human papillomavirus is associated with poor clinical outcome of laryngeal carcinoma. J. Clin. Pathol. 58, 402–405. https://doi.org/10.1136/jcp.2004.022103 (2005).
Webb, B., Rakibuzzaman, A. & Ramamoorthy, S. Torque teno viruses in health and disease. Virus Res. 285 https://doi.org/10.1016/j.virusres.2020.198013 (2020).
Zheng, H. et al. Torque teno virus (SANBAN isolate) ORF2 protein suppresses NF-kappaB pathways via interaction with IkappaB kinases. J Virol. 81 (21) 11917-24.x https://doi.org/10.1128/JVI.01101-07 (2007).
García-Álvarez, M. et al. Association of Torquetenovirus (TTV) and torque Teno mini virus (TTMV) with liver disease among patients coinfected with human immunodeficiency virus and hepatitis C virus. Eur. J. Clin. Microbiol. Infect. Dis. 32, 289–297. https://doi.org/10.1007/s10096-012-1744-1 (2013).
Borkosky, S. S., Whitley, C., Kopp-Schneider, A., zur Hausen, H. & de Villiers, E. M. Epstein-Barr virus stimulates Torquetenovirus replication: a possible relationship to multiple sclerosis. PLoS One. 7, e32160. https://doi.org/10.1371/journal.pone.0032160 (2012).
Chand, U., Priyambada, P. & Kushawaha, P. K. Staphylococcus aureus vaccine strategy: promise and challenges. Microbiol. Res. 271, 127362. https://doi.org/10.1016/j.micres.2023.127362 (2023).
Fayyaz, M. et al. Pathogens causing blood stream infections and their drug susceptibility profile in immunocompromised patients. J Coll Physicians Surg Pak. 23, 848–851 https://doi.org/12.2013/jcpsp.848851 (2013).
Dropulic, L. K. & Lederman, H. M. Overview of infections in the immunocompromised host. Microbiol. Spectr. 4 https://doi.org/10.1128/microbiolspec.DMIH2-0026-2016 (2016).
Cheng, X. et al. Resistance phenotype and genetic features of a heterogeneous Vancomycin intermediate-resistant Staphylococcus aureus strain from an immunocompromised patient. Braz J. Microbiol. 55, 323–332. https://doi.org/10.1007/s42770-023-01192-y (2024).
McNeil, J. C. Staphylococcus aureus - antimicrobial resistance and the immunocompromised child. Infect. Drug Resist. 7, 117–127. https://doi.org/10.2147/idr.S39639 (2014).
Yu, K., Huang, Z., Xiao, Y. & Wang, D. Shewanella infection in humans: Epidemiology, clinical features and pathogenicity. Virulence 13, 1515–1532. https://doi.org/10.1080/21505594.2022.2117831 (2022).
Yousfi, K., Bekal, S., Usongo, V. & Touati, A. Current trends of human infections and antibiotic resistance of the genus Shewanella. Eur. J. Clin. Microbiol. Infect. Dis. 36, 1353–1362. https://doi.org/10.1007/s10096-017-2962-3 (2017).
Duraisingham, S. S. et al. Immune deficiency: changing spectrum of pathogens. Clin. Exp. Immunol. 181, 267–274. https://doi.org/10.1111/cei.12600 (2015).
Jaksch, P. et al. Torquetenovirus as a novel biomarker targeting the efficacy of immunosuppression after lung transplantation. J. Infect. Dis. 218, 1922–1928. https://doi.org/10.1093/infdis/jiy452 (2018).
Brassard, J. et al. Association of age and gender with Torquetenovirus detection in stools from diarrheic and non-diarrheic people. J. Clin. Virol. 72, 55–59. https://doi.org/10.1016/j.jcv.2015.08.020 (2015).
Xie, Y. et al. Associations between sputum Torquetenovirus load and lung function and disease severity in patients with chronic obstructive pulmonary disease. Front. Med. (Lausanne). 8618757. https://doi.org/10.3389/fmed.2021.618757 (2021).
Pradier, A. et al. Torquetenovirus as a potential biomarker for complications and survival after allogeneic hematopoietic stem cell transplantation. Front. Immunol. 11, 998. https://doi.org/10.3389/fimmu.2020.00998 (2020).
Doberer, K. et al. Torquetenovirus for risk stratification of graft rejection and infection in kidney transplant recipients-A prospective observational trial. Am. J. Transpl. 20, 2081–2090. https://doi.org/10.1111/ajt.15810 (2020).
De Vlaminck, I. et al. Temporal response of the human Virome to immunosuppression and antiviral therapy. Cell 155, 1178–1187. https://doi.org/10.1016/j.cell.2013.10.034 (2013).
Acknowledgements
We are particularly grateful to all the people who have given us help with our article.
Funding
This work is supported by the Medical Talent Program for High-hroughput Sequencing Technology in Infectious Diseases, China (Grant No. MTP2022D027).
Author information
Authors and Affiliations
Contributions
Dingqiang Chen and Ke Yuan conceptualized and designed the experiments. Qike Zhang collected and analyzed data. Qike Zhang, Yihua Sun, Ke Yuan, and Dingqiang Chen drafted the initial manuscript. Yihua Sun contributed to data analysis and interpretation, and revisions to the manuscript. Wanting Liu and Qianwen Zhao provided theoretical support, conducted literature reviews, performed discussion analysis, and also revised manuscript revision. All authors reviewed and approved the final submitted version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
The study was conducted following the Declaration of Helsinki (revised in 2013). The study was approved by the Ethics Committee of the ZhuJiang hospital of Southern Medical University (Guangzhou, China).
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, Q., Sun, Y., Liu, W. et al. Clinical characteristics of Torquetenovirus infected immunocompromised patients explored by metagenomic next-generation sequencing. Sci Rep 15, 40857 (2025). https://doi.org/10.1038/s41598-025-24543-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-24543-1