Abstract
Hepatocellular carcinomas (HCC) with multinodular morphology were typically polyclonal and exhibited extensive heterogeneity. We explored the multiple-layer heterogeneity of HCC utilizing single-cell multiomics sequencing together with multiregional sampling. We found that Confluent Multi-Nodular samples exhibit more heterogeneous immune landscapes characterized by increased transcriptome heterogeneity and more complex immune-related interactions compared to Single Nodular (SN) samples. We identified differential DNA methylation patterns among distinct tumor foci in a subset of liver cancer patients. Global DNA hypomethylation in cancer cells predominantly occurs in partially methylated domains. We predicted and validated two genes, GADD45A and SNHG6, as potential drivers of DNA hypomethylation. Lastly, we demonstrated that DNA methylation distance serves as a more robust metric than gene expression distance for reconstructing tumor evolutionary trajectories.
Similar content being viewed by others
Introduction
Liver cancer ranked as the seventh most diagnosed cancer and the third most common cause of cancer death in 20221. Hepatocellular carcinoma (HCC) is the main subtype, representing 75–85% of all clinical cases. Surgical treatments, such as hepatic resection and liver transplantation, have demonstrated substantial efficacy in enhancing the 5-year survival rate for HCC patients. However, there is a high recurrence rate of approximately 70% within the initial 5 years. Despite recent advances in immunotherapy for HCC, the problem of limited objective response rates persists2,3,4. The suboptimal effectiveness of current therapeutic methods is closely linked to both inter- and intra-tumor heterogeneity (ITH), which is marked by the accumulation of genetic and epigenetic changes, the development of new tumor clones, and metastatic advancement.
Several studies have reported epigenetic changes in HCC, particularly in the context of DNA methylation3,5,6,7,8,9. Hypermethylation of CpG island (CGI) promoters and global DNA hypomethylation are two well-defined epigenomic characteristics of cancer10. However, most studies primarily focused on analyzing DNA methylation at gene promoter regions, representing only about 6% of all the CpG sites across the human genome. It is widely acknowledged that DNA methylation also plays significant roles in noncoding regions, including enhancers11, partially methylated domains (PMDs)12, and repetitive elements13. PMDs, which are typically hypomethylated in cancer, cover more than one-third of the genome and are enriched by heterochromatin, also known as chromatin “B” compartment determined by Hi-C, and nuclear lamina-associated domains14,15,16. Conversely, the regions where methylation levels are maintained are called highly methylated domains (HMD)12,17. HCC is documented to exhibit the largest PMD-HMD difference among all TCGA cancer types18. DNA hypomethylation in PMD may contribute to chromosomal instability and oncogene activation19 It can also reactivate silenced retrotransposons, disrupting gene structure and function and further accelerating cancer progression20,21.
While extensive research has delved into intra-tumor heterogeneity (ITH) and the tumor microenvironment (TME) in HCC and other cancers, most studies have focused on heterogeneity across different tumor nodules or between primary and metastatic tumors. The exploration of spatial ITH within the same tumor, as well as ITH between primary and satellite tumors, is still insufficient. HCC is classified into Single Nodular (SN) and confluent multinodular (CMN) types based on morphology22. SN has one tumor nodule, a clear or unclear outline, and the possible presence of satellite nodules. On the contrary, CMN involves multiple fused nodules, each exhibiting diverse pathological features and clear tumor borders. SN generally has a better prognosis, while CMN, with its complex biological behavior, often indicates a poorer outcome22,23. The genomic heterogeneity underlying these two different types awaits further exploration. Moreover, previous single-cell studies have predominantly concentrated on dynamic transcriptional changes within the TME, missing the potential collaborative contribution of various ‘omics’ layers—including the genome, transcriptome, and epigenome—in shaping ITH in tumor patients.
Herein, we conducted single-cell RNA sequencing (scRNA-seq) on cells derived from primary regions within the same tumor and satellite nodules in three patients with SN HCC and six patients with CMN HCC. Additionally, we employed an advanced single-cell multi-omics sequencing strategy known as scTrio-seq224, enabling concurrent examination of transcriptomic profiles, DNA methylation levels, and genomic copy number alterations (CNAs) to comprehensively analyze tumor cells from diverse regions. We also performed whole-exon sequencing to delineate lineage relationships and clarify the clonal origins of tumor cells. Our study provides novel insights into tumorigenesis, evolution, and heterogeneity in HCC.
Methods
Human specimens ethical application
Nine patients who were pathologically diagnosed with liver cancer and treated with surgical resection, including five males and four females, were enrolled in this study after approved by the Ethics Committee of Sun Yat-sen University Cancer Center (B2022-700-01). The available clinical characteristics of these patients are summarized in Table S1. Their tumor and adjacent noncancerous liver tissues were obtained for the subsequent single cell isolation. Tumor tissues were dissected from nodule core, while the adjacent liver tissues were at least 2 cm from the matched tumors.
Sample collection and single-cell processing
Fresh liver tissues were delivered in with MACS Tissue Storage Solution on ice within 3 h after dissection. Liver tissues were cut into approximately 1 mm3 pieces and enzymatically digested with MACS tumor dissociation kit (Miltenyi Biotec, Cat. 130-095-929) using 37C_h_TDK_3 program in the gentleMACS Octo Dissociator with Heaters, according to the manufacturer’s instructions. Dissociated cells were filtered through a 70 μm cell strainer (BD) and then centrifuged at 400 g for 10 min at 4 °C. After removing the supernatant, the cell pellet was re-suspended by 1X PBS with 10% FBS to wash, and the red blood cells were removed using the red blood cell lysis buffer (Roche), according to the manufacturer’s instructions. The cell pellet was washed again with 1X PBS with 10% FBS and the stained with APC anti-human CD45 Antibody (Biolegend, Cat. 368512) and 7AAD (BD, Cat. 559925) for cell sorting to remove cell fragments and separate CD45+ and CD45- cells. Cells were then filtered through a 40 μm cell strainer (BD, Cat. 352340), and loaded on BD sorter (BD FACSAria III). Single cells were picked up by mouth pipetting from CD45- cell population and process scTrio-seq2. CD45+ and CD45- cells were collected and counted and mixed 1:1 to load on loaded onto the Chromium single cell controller (10 × Genomics) or Drop-Seq droplet generation device (Dolomite Bio) to generate single-cell gel beads in the emulsion according to the manufacturer’s protocol. ScRNA-seq libraries were constructed using Single Cell 3′ Library and Gel Bead Kit v3.1 or Drop-Seq (v1.8) and sequenced using the NovaSeq 6000 sequencer (Illumina).
scTrio-seq2 sequencing
We processed scTrio-seq2 as described before with minor modifications24. In brief, 0.2 μl magnetic beads (Invitrogen, Cat. 65011) were used to separate the nucleus and RNA of one single cell in lysis buffer. Each single cell was lysed in one tube and vortexed to release RNA to lysis buffer. Then, we collected nuclei using magnetic rack, because magnetic beads can aggregate on the surface of the nucleus and maintain the nucleus in the pellet. The supernatants containing RNA were transferred to a new tube for transcriptome sequencing. RNA seq libraries were constructed according to a multiplexed scRNA-seq method25, in which the poly T primers were combined with barcodes and unique molecule identifiers (UMIs). For nuclei, we performed the single-cell whole-genome bisulfite sequencing (scBS-seq)26, which covered the whole genome. The beads containing a single nucleus were re-suspended with lysis buffer of scBS-seq for DNA methylation sequencing. All the sequencing were conducted by NovaSeq 6000 sequencer (Illumina).
Processing of single-cell RNA-seq data
For scRNA-seq data generated on the 10 × Genomics platform, we employed the cellranger27 (v5.0.1) pipeline to process the raw sequencing data, which included basic quality control and subsequent reads alignment to the GRCh38 reference genome. In the case of scRNA-seq data obtained from the Drop-seq platform, we predominantly followed the dropEst28 (v0.8.6) pipeline for processing the raw sequencing data. For scRNA-seq data originating from scTrio-seq2, we utilized customized pipelines. These pipelines included the extraction of cell barcodes and unique molecular identifiers (UMIs) using ‘umi_tools’, the removal of potential template-switching oligos (TSOs) and polyA sequences, read alignment with STAR29, and read summarization with featureCounts30. As a result, we obtained the raw gene-barcode expression matrix for each HCC patient. The quality details of scRNA-seq libraries from all the samples were summarized in Table S4.
Subsequently, we conducted the main data analysis using the Seurat v431 package. We filtered out cells based on the following criteria: the number of expressed genes < 300 or > 5000, UMI count > 3 times the mean UMI count, and the percentage of mitochondrial genes > 20%. Genes expressed in fewer than 30 cells for 10 × Genomics, 10 cells for Drop-seq, and 5 cells for scTrio-seq224 were filtered out, respectively. We also utilized DoubletFinder32 to identify and exclude potential doublets. After preprocessing the raw expression matrix for each sample, we performed data integration for all HCC samples using the canonical correlation analysis (CCA) method implemented in the Seurat package. This process included data normalization, identification of variable features (n = 3000), determination of integration anchors using ‘FindIntegrationAnchors,’ and subsequent data integration. After data scaling, principal component analysis (PCA) dimension reduction (nPCs = 40), and Louvain clustering, we visualized cell embeddings with a UMAP plot. Cell type annotations were assigned based on the expression of canonical marker genes. Expression scores were calculated as the mean expression value of marker genes for each cell type. Finally, we removed undefined cell clusters and performed another round of clustering, as described above. Differentially expressed genes were identified using the ‘FindMarkers’ function with the following parameters: test.use = ‘wilcox,’ min.pct = 0.3, min.cells.feature = 3, and min.cells.group = 15. To eliminate batch effects across different sequencing platforms, we applied the Harmony33 software for data integration. Functional enrichment analysis was performed using the clusterProfiler34package.
Cellchat analysis
Cell–cell communication analysis was performed using the CellChat35 R package (version 2.1.2).The analysis was performed using all cells in the dataset, with cell types categorized. Communication probabilities were computed (raw.use = TRUE) and filtered with a minimum threshold of 3 cells per population. Pathway-level communication probabilities were then calculated, and the overall networks were aggregated using the aggregateNet function.
Processing of single-cell DNA methylation data from multi-omics sequencing.
The raw sequencing reads were trimmed to remove sequencing adapters, amplification primers, and low-quality bases in read ends. Then, the trimmed reads were mapped to the human reference genome (hg38) using Bismark36 (version 0.24.1) (duplication caused by cycles of random priming in the pre-amplification step of the scBS-seq method could not be completely removed in data processing step).
Through the CpG report provided by Bismark, we can calculate the bin-level methylation level by separately adding all the methylated and demethylated CpG cites counts together and dividing the summed counts. We mostly used 100 k as the bin size for further analysis.
The strt-seq clustering result allows us to identify the hepatocyte cell (HPC) among all CD45- cells for downstream analysis.
To analyze methylation patterns, we clustered cross bins and cells based on their methylation levels using correlation distance and visualized the clusters using pheatmap37. Only bins showing valid methylation levels across all samples were retained. The filtered bins were then intersected with the hg38 PMD annotation BED file using bedtools38 for further analysis. Hypomethylated regions were defined by comparing bin-level methylation in tumor nodules to the average in normal liver tissue. Bins differing by more than 0.25 were considered hypomethylated, while those within 0.25 were considered “normal-like”.
TCGA methylation array data re-analysis
Liver DNA methylation datasets from The Cancer Genome Atlas (TCGA)39, analyzed using the Illumina Infinium Human Methylation 450 K BeadChip platform, were obtained from the National Cancer Institute’s Genomic Data Commons (GDC) Data Portal (https://portal.gdc.cancer.gov). The datasets encompass 379 tumor samples and 50 normal samples. The CpG site-level methylation data were initially mapped to hg38 coordinates utilizing the illuminaMethyl450_hg38_GDC annotation provided by GDC.
For downstream analysis, the data were segmented into fixed 100 kb non-overlapping genomic windows, within which the average methylation fraction was calculated for each sample. Bins were excluded from the analysis if they lacked adequate probe coverage on the Illumina Infinium Human Methylation 450 K BeadChip array (less than 10).
Gene expression data and clinical information for TCGA Liver Hepatocellular Carcinoma (LIHC) samples were retrieved from UCSC Xena (https://xena.ucsc.edu/).
Over-expression of SNHG6 and GADD45A
Full-length human GADD45A and SNHG6 genes were amplified by PCR from HEK293T cDNA with gene-specific primers and Super-Fidelity DNA Polymerase (Vazyme, Cat. P525). For each gene, the forward primers contained a 20 nt vector homologous sequence at the 5 ends followed by a gene-specific amplification primer. The reverse primers contained a 20 nt vector homologous sequence followed by the Thosea asigna virus (T2A) peptide coding sequence, and then a gene-specific amplification primer. Gene cloning primers can be found in Table S2.
The PCR products were taken for 1.0% Agarose electrophoresis and were purified by gel extraction kit (Magen, DMA02). The purified PCR products were quantified by NanoDrop One (Thermo Fisher). 1ug of PB-TRE-EGFP-EF1a-rtTA empty plasmid (Addgene, 104454) was digested with NcoI-HF (NEB, Cat. R3193V) restriction-endonuclease at 37℃ for 1 h 1 × rCutSmart Buffer (NEB, B6004V). The digestion product was taken for 1.0% Agarose electrophoresis and was purified by gel extraction kit. The purified linearized vector were quantified by NanoDrop One.
For each gene, 10 ng of purified PCR product was mixed with 50 ng linearized vector, of which the molar concentration ratio was about 3:1. Equal volume of 2 × Hieff Clone Universal Enzyme Premix (Yeasen, Cat. 10922ES) is added to the mixture, gently mixed and incubated at 50 °C for 10 min. The reaction product was transformed into DH5α Chemically Competent Cell (Yeasen, Cat. 11802ES). A single positive colony was picked from the resistance culture plate with an inoculation loop, and was placed into 30 mL of liquid culture medium containing antibiotics, and then cultured overnight at 37 °C and 200 rpm. Plasmids were extracted with plasmid extraction kit (Omega, Cat. D6950-02). 2ug of plasmid was taken for Sanger sequencing to ensure the correctness of gene sequence and inframe with the following EGFP coding region.
HepG2 cells were plated in 6-well plate and cultured in DMEM (Cytiva, Cat. SH30243.01) supplemented with 10% FBS (BIOCHROM, Cat. S0615) and 1% penicillin–streptomycin (Gibco, 15140122) and incubated at 37 °C and 5% CO2. After two passages, 2 × 105 cells per well were plated in 6-well plate (Costar, Cat. RF1154-50) with 800 μL medium and transiently transfected with 2 μg of PB-TRE-GADD45A-T2A-EGFP-EF1a-rtTA plasmid and PB-TRE-SNHG6-T2A-EGFP-EF1a-rtTA plasmid with 0.5ug piggyBac transposase plasmid PB200PA (gift from Man Zhang) for one well using Lipofectamine 3000 reagent (Invitrogen, Cat. L3000015) according to the manufacturer’s instructions, respectively. When the cell density reached to 80% confluency, changed 2 mL fresh Medium per well, and added puromycin (MCE, Cat. HY-B1743A) at final concentration of 1 mg/mL to select positive cells, HepG2 cells without transfection as a control. When control cells all died (usually 2–3 days), there should be some survived colonies in the dishes with transfected HepG2 cells. Continuously culture the survived colonies after puromycin selection, passage the cells three generations (reduce the concentration of puromycin to final concentration of 0.5 mg/mL) to obtain the stable cell line. Cells from each stable cell line were plated in two wells of 6-well plate at 1 × 105 cells per well with 2 mL medium. Doxycycline (MCE, Cat. HY-N0565B) was added into one well to final concentration 2 mg/mL in one day, while the other well without Dox as a control. Check the cells under the fluorescence microscope in 48 h. At this point, the most of cells should be GFP positive, which indicated the success of generation of inducible expression stable cell lines.
For each stable cell line, the medium was removed in 48 h after Dox induction. The cells were washed once with 2 mL 1 × DPBS (Cytiva, Cat. SH30028.03) and incubated with 200μL 0.05% Trypsin–EDTA (Gibco, Cat. 25300062) at 37 °C and 5% CO(2) for 5 min. 1 mL of culture medium was added to the culture wells to block digestion. The cells were gently dislodged from the plate with pipette and transferred into a 1.5 mL Eppendorf tube, centrifuged at 500 × g for 5 min at room temperature. And then the supernatant was discarded. The RNA was extracted from the collected cells and purified by MolPure Cell/Tissue Total RNA Kit (Yeasen, Cat. 19221ES) according to the manufacturer’s protocol. The purified RNA was were quantified by NanoDrop One. 500 ng RNA was converted into oligo(dT)-primed first-strand cDNA using HiScript IV All-in-One Ultra RT SuperMix (Vazyme, Cat. R433) . ChamQ Universal SYBR qPCR Master Mix (Vazyme, Cat. Q711) was used for qPCR reaction according to the manufacturer’s protocol in Quantagene q225 Real-Time PCR System (KUBO Tech Co., Ltd). Gene expression was normalized to β-ACTIN. The delta-Ct method was used for analysis and to calculate fold changes based on Ct values. All primers used for qPCR are listed in Table S2.
Genomic DNA (gDNA) was isolated using the DNeasy Blood & Tissue Kit (Qiagen, Cat. 69506) following manufactures instructions. In total, 1 μg genomic DNA was used as input material, and the DNA was fragmented into around 300 bp by Covaris. DNA was bisulfite-converted using EZ-96 DNA Methylation-Direct™ MagPrep Kit (Zymo, Cat. D5044). Bisulfite-converted DNA was captured using Accel-NGS Methyl-Seq DNA Library Kit (Swift Biosciences, Cat. 30024) according to the manufacturer’s instructions. Library samples were subjected to Illumina Nova-seq 6000 sequencing system.
Knock-down of SNHG6 and GADD45A
We used shRNA to knock down SNHG6. pLKO constructs were used to make lentivirus in 293 T cells via PEI reagent (POLYSCIENCES INC, Cat. 24765-100). Viral media was collected and filtered with Syringe Filters (0.45 µm). C. Polybrene was added into the media (5 μg/mL final concentration) to improve the infection rate. Cells were incubated for 24 h with shRNA lentivirus before adding fresh growth media for another 24–48 h. The GFP-positive rate was used to estimate infection efficiency.
Lentiviral particles of SNHG6 shRNA were collected 36 h post-transfection in HEK293T cells. Huh7 cells were infected twice, with a one-day interval, using shRNA titers at an MOI of 0.7 to attain 90% infection efficacy. Post-infection, the cells were cultured for 15 days in DMEM (HyClone, Cat. SH30243) supplemented with 10% FBS (Sigma, Cat. F0193), 1% L-glutamine (Gibco, Cat. 35050-061), and 1% penicillin–streptomycin (Gibco, Cat. 15140-122).
We used the CRISPick tool1,2 to identify sgRNAs targeting the human GADD45A gene. These sgRNAs were then constructed into a lentiviral vector that concurrently expressed the dCas9-KRAB fusion protein (Addgene Cat. 71236). The lentiviral particles were efficiently packaged in HEK293T cells for 48 h post-transfection. Subsequently, HEK293T cells were infected with the lentivirus at a low MOI, followed by selection with 2 μg/ml of puromycin for three days. The surviving cells were then cultured in DMEM medium (Hyclone, Cat. SH30243.01), supplemented with 10% FBS (Cellmax, Cat. SA211.02) and Penicillin–Streptomycin (Gibco, Cat. 15140122), for an additional 12 days. Total RNA was extracted from these cells using TRIzol Reagent, and cDNA was synthesized employing the HiScript IV 1st Strand cDNA Synthesis Kit (Vazyme, Cat. R412-02). To generate an mRNA library for sequencing, we employed the VAHTS® Universal V8 RNA-seq Library Prep Kit for Illumina (Vazyme, Cat. NR605-01) with two biological replicates.
Lentiviral particles of GADD45A CRISPRi (using Addgene72136 as the plasmid backbone) were collected 36 h post-transfection in HEK293T cells. Fresh HEK293T cells were infected with GADD45A CRISPRi titers at an MOI of 0.2. After infection for 36 h, the cells were treated with 2 μg/mL puromycin for three days to kill the most uninfected cells. Subsequently, those survival cells were cultured for an additional 12 days in DMEM (HyClone, Cat. SH30243) supplemented with 10% FBS (Sigma, Cat. F0193) and 1% penicillin–streptomycin (Gibco, Cat. 15140-122).
All primers used for qPCR and sequences constructed for knockdown are listed in Table S3.
Total RNA was extracted in TRIzol reagent (Thermo, Cat. 15596018) and dissolved in RNase-free water (Thermo, Cat. 10977023).
For RT-qPCR, cDNA was synthesized using HiScript IV 1st Strand cDNA Synthesis Kit with gDNA wiper (Vazyme, Cat. R412). Relative expression of SNHG6 and GADD45A were measured with ChamQ Universal SYBR qPCR Master Mix (Vazyme, Cat. Q711) using GAPDH as control.
For RNA-sequencing, mRNA was captured with VAHTS® mRNA Capture Beads (Vazyme, Cat. N401), and mRNA library was constructed using VAHTS® Universal V8 RNA-seq Library Prep Kit for Illumina (Vazyme, Cat. NR605-01), and sequenced on the Illumina Novaseq 6000 platform.
For WGBS, ~ 10,000 cells were collected by centrifugation at 4 °C for 5 min at 300 × g, washed with 1 × PBS (Solarbio, Cat. P1022), and resuspend with 200 μL Lysis Buffer (10 mM Tris–HCl, pH 8.0; 10 mM EDTA, pH 8.0; 10 mM NaCl; 0.5% SDS; 50 ng/μL RNase A (Vazyme, Cat. DE111); 1 μg/μL Proteinase K (Vazyme, Cat. DE102-01)). The suspension was incubated at 55 °C for 40–60 min with max speed in a thermal mixer. 112 μL (0.5X) SPRIselect beads (Beckman Coulter, Cat. B23318) were used to extract gDNA and stored at − 80 °C before use.
Identification of differentially methylation regions
Sequencing data were aligned to the hg38 human genome using Bismark. We first performed a statistical test of the differentially methylated locus (DML) using the DMLtest function (smoothing = TRUE) in DSS(53), the results were then used to detect differentially methylated regions (DMRs) using the CallDMR function in DSS40, the p-value threshold for calling DMR is 0.01. The minimum length for DMR is 100 bps, and the minimum number of CpG sites for DMR is 3. The minimum methylation difference is 0.2. the maximum distance between two CpGs was set to 100 bp.
Whole exon sequencing (WES)
Tissues of HCC patients were rinsed thoroughly with pre-cold PBS and minced into small pieces. Genomic DNA was extracted using the DNeasy Blood & Tissue Kit (Qiagen, Cat. 69504). Subsequently, WES libraries were prepared and captured using VAHTS® Target Capture Hybridization and Wash Kit (Vazyme, Cat. NC103) and Universal Plus DNA Library Prep Kit for Illumina V2 (Vazyme, Cat. ND627) following the manufacturer’s instructions.
WES data processing
The raw sequencing reads were trimmed and then mapped to the human genome reference (hg38) using BWA41 (version 0.7.17-r1188). BAM files from different libraries and sequencing lanes of one sample were sorted using Samtools42 (version 0.1.18). BAM files were then preprocessed by GATK43 (version 4.3.0.0). SNVs were called using Mutect244,45 (version 4.3.0.0) and further filtered using custom scripts.
CNV deduction
For the CNV estimation of paired tumor-normal whole-exome sequencing (WES) data from bulk tissue samples, we utilized CNVkit46 (version 0.9.9). The CNVs were inferred using the standard procedure with default parameters.
For the CNV estimation in single-cell RNA-seq data, we used cellranger to align reads to the human genome and InferCNV47 to identify large-scale chromosomal copy number alterations. The HMM parameter was set to True, the analysis mode was set to subclusters, with a cutoff of 0.1, and other parameters were set to default values.
For the CNV estimation in single-cell whole-genome bisulfite sequencing (WGBS) data, we employed Samtools depth. For each window, we calculated the total sequence depth and subsequently normalized it based on the overall depth for each cell. The sequence depth for each window was then divided by the normalization factor, which was determined by averaging the depth values from normal HPC data.
Clustering and distance calculation of computational phylogenetics
All single-cell data used were preprocessed through pseudo-bulk aggregation. The samples were then grouped based on similarities calculated using the Euclidean distance method and average linkage.
Statistical analysis
Statistical analyses in this study were performed using R software (version 4.2). The data were analyzed by wilcox test, one-way ANOVA, Pearson’s correlation or Kaplan‒Meier analysis according to the actual conditions. R software were used to construct the graphs. p < 0.05 was considered to indicate statistical significance.
Data and code accessibility
All sequencing data reported in this paper have been deposited in the Genome Sequence Archive48 in National Genomics Data Center49, China National Center for Bioinformation / Beijing Institute of Genomics, Chinese Academy of Sciences, linked to BioProject: PRJCA031530. (https://ngdc.cncb.ac.cn/). The raw sequence data are publicly accessible at GSA-human (https://ngdc.cncb.ac.cn/gsa-human) under accession numbers HRA009266 and HRA009544 The processed data are publicly accessible at OMIX(https://ngdc.cncb.ac.cn/omix) under accession numbersOMIX007875 and OMIX007876.
The code used for data analysis and visualization in this paper is publicly available at https://github.com/RobinZzi/zhang_et_al_2024.
Results
Celltype heterogeneity identified in HCC by scRNA-seq
To investigate the heterogeneity of HCC, we conducted single-cell experiments on tumor specimens and adjacent normal liver tissues obtained from three patients with SN type HCC and six with CMN HCC. Multiple clearly demarcated regions were sampled from each tumor lesion, and satellite nodules were also included in the analysis. To fully capture the tumor cells, we performed flow cytometry sorting and selectively isolated CD45- non-immune cells and CD45+ immune cells, mixed them in a 1:1 ratio before performing microfluidics based scRNA-seq50. For the non-immune cells, we also performed scTrio-seq221,24 to obtain multiomics information(Fig. 1A).
Single-cell RNA seq Landscape of HCC. (A)Scheme of the overall study design including sample collection, single cell sequencing, and data analysis. (B, D-E) Uniform Manifold Approximation and Projection (UMAP) showing clustering of all cells from 9 HCC patients. Cells are colored by cell type (B) ,patient (D) and tissue type (E). (C) Violin plot showing show the canonical marker genes expression for cell type annotation.
We performed an integrated analysis of all the scRNA-seq data, identifying 37 distinct cell clusters (Sup Fig. 1A). These clusters were initially classified into four broad categories—epithelial (KRT8+), endothelial (PECAM1+), immune (PTPRC+), and fibroblast (COL1A2+) cell (Sup Fig. 1B). Subsequently, these categories were further subdivided into 16 cell types, including hepatocyte cell (HPC) (AMBP+), fibroblast (COL1A2+), endothelial cell (ENG+), neutrophil (S100A8+), B cell (MS4A1+), plasma B cell (MS4A1+, CD79A+), Proliferative T cell (CD3D+, MKI67+), CD8+ memory cell (CD8A+, LTB+), CD8+ exhausted cell (CD8Ahi, PDCD1+), CD8+ cytotoxic cell (CD8A+, CCL5hi), CD4+ memory cell (CD4+, LTB+, FOXP3-), CD4+ Treg cell (CD4+, FOXP3+), NK cell (SPON2+), mast cell (KIT+), Dendritic cell (FCN1+, LYZhi), Macrophage cell (CD63hi, CD168hi), (Fig. 1B, C). Notably, cells clustered primarily according to cell type rather than by patient or tissue type (Fig. 1D, E), indicating effective batch effect control. Given the use of three different scRNA-seq platforms (10 × Genomics, Drop-seq, and STRT-seq) in this study, we compared the data from all three platforms. Remarkably, the major cell types identified and the number of genes detected were consistent across platforms (Sup Fig. 1C, D). But in terms of the throughput, the 10 × platform yielded many more cells than the other two methods in our hands.
The tumor microenvironment plays multiple roles in tumorigenesis and progression. We first looked at the ratios of different immune cell types in the tumor ecosystem, compared to adjacent normal tissues. As expected, we observed a significant enrichment of CD4+ regulatory T cells expressing immune inhibitory genes such as CTLA4, FOXP3, IL2RA, and LAYN in tumor nodules compared to normal counterparts. Conversely, there was a notable reduction in infiltrating natural killer cells and CD8+ cytotoxic T cells, characterized by high expression of CST7, GZMB, KLRD1, and NKG7, suggesting a relative immune-suppressive state within the tumor nodules. Additionally, there was a significant enrichment of CD4+ memory T cells, accompanied by a decreased presence of CD8+ memory T cells in tumor nodules compared to normal tissues (Sup Fig. 1E). Of note, we identified an extremely high percentage (over 50%) of macrophages in tumor nodules from patients HCC8 and HCC9, and thus they were excluded from the analysis of cell type proportions (Sup Fig. 1F). These results collectively suggest that the balance between different immune cell populations is heavily disrupted during tumorigenesis, aligning with findings from prior research26.
Immune landscape and gene expression heterogeneity between CMN and SN, and primary and satellite nodules
In this study, patients were categorized into CMN and SN groups based on morphological differences. We sought to determine whether there are any differences in immune landscape and gene expression heterogeneity between these tumor subtypes. To ensure that cell type composition estimates were not biased by differences in sequencing platforms, we grouped the samples according to platform (10 × Genomics, Drop-seq) and compared the cell type proportions for tumors of the same type (Sup Fig. 2A). Our analysis demonstrated high consistency in cell type proportions across platforms, indicating that platform-specific effects—such as differences in encapsulation rates or cell-calling thresholds—did not meaningfully influence our results. Our analysis revealed that nodules in CMN samples exhibit a lower proportion of CD8+ cytotoxic T cells, and a higher proportion of Proliferative T cells than SN samples (Fig. 2A). To determine whether the cell type composition difference were caused by different single-cell sequencing platforms, we compared these cell type proportions under each platform. The CD8 cytotoxic T cells were still significantly more in the SN samples (Sup Fig. 2B). In addition, we performed cell communication analysis across all samples. To avoid the influence of cell numbers to cell–cell communication, we only use the Drop-Seq data as the cell numbers were consistent across samples. Compared to SN samples, CMN samples exhibited more cellular communication interactions (Fig. 2B, Sup Fig. 2B), particularly enriched in various immune-related pathways (Fig. 2C). We also explored transcriptomic consistency within individual nodules. As expected, the intra-nodule transcriptomic correlation was higher than the inter-lesion correlation within the same patient, and this difference holds true in both CMN and SN patients (Sup Fig. 2C). While when comparing the intra-nodule correlation difference between CMN and SN, in both primary tumor and satellite tumor from drop-seq dataset with the similar number of genes detected per cell, SN showed significantly higher transcriptomic correlation, indicating greater heterogeneity in CMN patients (Sup Fig. 2D).
Gene expression heterogeneity between CMN and SN, and primary and satellite lesions. (A) Box plot showing the immune cell type proportion comparison between Confluent Multi-Nodular (CMN) and Single Nodular (SN). P values represent results of Wilcoxon test. (B) Box plot showing the cell–cell interaction counts comparison between CMN and SN samples in 10x-excluded cells(left) and drop-seq-only cells (right).P values represent results of Wilcoxon test. (C) Box plot showing differential pathways of cell–cell interaction between CMN and SN samples.P values represent results of Wilcoxon test. (D) Box plot showing the immune cell type proportion comparison between primary tumor (pt) and satellite tumor (st).P values represent results of Wilcoxon test. (E–F) Box plot showing the cell–cell RNA expression level correlation comparison between pt and st in CMN (E) and SN (F) samples. P values represent results of Wilcoxon test. (G) Dot plot showing the gene ontology (biology process) term result of common down-regulated genes in satellite tissue of HCC1.and HCC6.
Tumor nodules were also classified as primary nodules (PT) or satellite nodules (ST) based on their spatial location. When comparing primary and satellite nodules, satellite nodules exhibit a higher proportion of CD8+ memory T cells, whereas no other significant differences were found (Fig. 2D). On the other hand, when examining ITH, primary nodules showed significantly higher transcriptomic correlation than satellite nodules In CMN patients (Fig. 2E), indicating greater heterogeneity in satellite nodules In CMN patients. Interestingly, this difference was not observed in SN patients, where heterogeneity within satellite and primary nodules was comparable (Fig. 2F).
Lastly, we compared the gene expression profiles of tumor cells from primary and satellite nodules in patients with satellite nodules (HCC1, HCC3, HCC5, HCC6, HCC7) to identify shared differential genes(Sup Fig. 2E, F). Our results revealed that common down-regulated genes in the satellite nodules of HCC1 and HCC6 were enriched in the angiogenesis pathway (Fig. 2G). This may be related to their smaller size, as smaller tumors typically require less extensive vascular networks for nutrient and oxygen supply, resulting in reduced expression of angiogenesis-related genes. No shared up-regulated genes were found in satellite nodules.
Single-cell multi-omics profiling reveals diverse methylation patterns of HPC
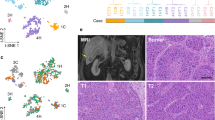
Trio-seq2 provided us with an opportunity to examine the DNA methylation landscape in individual tumor cells. At both single-cell and pseudo-bulk levels, compared with adjacent normal tissues, we observed hypomethylation within specific tumor nodules from patients HCC2, HCC6, HCC8, and HCC9 (4 of 6 patients with trio-seq data) (Fig. 3A, B, Sup Fig. 3A, 100 kb bin size). And these hypomethylated regions are enriched in PMDs (fisher exact test p < 2.22 × 10−16; odds ratio = 27.474). To further investigate these hypomethylated regions, we included liver cancer samples from The Cancer Genome Atlas (TCGA) in our analysis and identified a common trend of hypomethylation within these specific regions across multiple liver cancer patients (Fig. 3C). In addition, we asked what features are enriched in these hypomethylated regions. Our analysis indicated that compared to normally-methylated regions, hypomethylated regions are significantly enriched for heterochromatin-associated signals, including H3K9me3, H3K27me3, and L1, and depleted for H3K36me3, H3K27ac, and Alu (Fig. 3D). These findings align with previous studies that hypomethylated regions in cancer cells are typically gene-sparse and enriched for heterochromatin markers18.
Single-cell multi-omics profiling reveals diverse methylation patterns of HPCs. (A) Heatmap showing single-cell DNA-global-methylation level of all samples. Each col is the DNA methylation level of one single cell. The legend shows the origin factor of the cell. Line plot showing the average DNA methylation level of PMD, HMD and global area of each cell. (B) Flat-violin plot showing the cell-wise distribution of global DNA methylation level in each nodule. (C) Heatmap showing DNA-global-methylation level of all samples after pseudo-bulk, combined with TCGA-450 K array result. Line plot showing the average DNA methylation level of PMD, HMD and global area of each sample. Trio-seq result shows the same demethylated regions with TCGA-450 K array result. (D)Boxplot showing the overlapping length of hypomethylation or normally-methylated regions with histone modified regions or transposon elements of heterochromatin (left) and euchromatin (right) in each 100 kb bin. P values represent results of Wilcoxon test.
At the single-cell level, the methylation patterns vary considerably between patients and tumor nodules from the same patients. Dimensionality reduction analysis using single-cell level methylation data showed that cells from the same lesion clustered more closely with each other (Sup Fig. 3B). We also found that global hypomethylation also occurs in only a subset of patients, and a subset of nodules in some patients. For instance, patients HCC3 and HCC7 do not have any hypomethylated nodules, while patients HCC2 and HCC6 show distinct methylation patterns in different nodules (Fig. 3B, Sup Fig. 3B, C).
SNHG6 and GADD45A implicate in hypomethylation in HCC
To answer what genes might cause the hypomethylation phenotype in a subset of tumor nodules, we focused on tumor cells from HCC2, HCC8, and HCC9 as they displayed hypomethylation (Fig. 4A–C). Among these tumor nodules, only PT3 from HCC2 displayed a methylation pattern similar to the normal tissues (Fig. 3B). Therefore, we used HCC2-PT3 as the control group and performed differential gene expression analysis to PT1 and PT2, and all PTs from HCC8, and HCC9, respectively.
DNA methylation regulated transcriptome. (A–C) UMAP clustering of all cells from 3 patients(HCC2, HCC8, HCC9) containing hypo-methylated nodules, colored by patient(A), tissue type(B) and methylation level type(C). (D) Venn diagram showing the intersect result of the common up-regulated genes of hypomethylated nodules across three patients. (E) Venn diagram showing the intersect result of the up-regulated genes of hypomethylated nodules in HCC2 and the up-regulated genes of hypomethylated samples in TCGA LIHC dataset and top 1000 genes that exhibited a negative correlation with overall methylation levels in the TCGA LIHC dataset.(F–G) Sector graph showing the distribution of DMR in SNHG6 (F) and GADD45A (G) over-expressed cell lines. (H–I) Sector graph showing the distribution of DMR in SNHG6 (H) and GADD45A (I) knockdown cell lines.
Overlap analysis revealed that 24 genes were commonly up-regulated in all hypomethylated tumor nodules from three patients (Fig. 4D). Among them, APOE, APOC1, APOA2, APOB are associated with lipid metabolism51,52; NUPR1, SNHG6, IGFBP2 are associated with transcriptional regulation53,54,55. Additionally, we leveraged the TCGA data to perform a correlation analysis of gene expression and genome-wide methylation levels to identify genes whose expression levels are negatively associated with methylation. An intersection of the top 1000 genes whose expression levels are associated with hypomethylation in the TCGA samples and up-regulated genes in hypomethylated nodules in HCC2 revealed 36 common genes (Fig. 4E). Among them, DUXAP8, ZNF781, ZNF99, ZNF727, and GADD45A are closely associated with transcriptional regulation and the cell cycle. Among the 60 genes shortlisted from these two criteria, SNHG6 and GADD45A stand out because they were previously reported to be involved in the DNA demethylation process.
SNHG6 is a long non-coding RNA that influences the genome’s methylation composition by interacting with methyl-CpG-binding proteins, which identify and latch onto methylated DNA56. By swaying these proteins to modify DNA methylation patterns, SNHG6 may regulate gene activity and play a significant role in cellular processes. Increased SNHG6 expression is noted in various cancers, exhibiting oncogenic properties and leading to poorer patient outcomes55,57,58,59. We noted that in the TCGA liver cancer dataset, patients with higher SNHG6 expression levels have poorer survival outcomes. (Sup Fig. 4A).
GADD45A, a protein involved in the DNA damage repair pathway, contributes to cellular responses under environmental stress and is crucial for maintaining genomic stability. Notably, it has been reported that GADD45A can modulate methylation levels through interactions with Thymine-DNA Glycosylase (TDG), an enzyme involved in the base excision repair route60. Through this interaction, GADD45A can influence DNA methylation dynamics, potentially leading to changes in gene expression, and participate in intricate epigenetic regulation61,62,63,64 In the TCGA liver cancer dataset, the expression level of GADD45A is negatively correlated with the overall average DNA methylation level in patients (Sup Fig. 4B). However, patients with higher GADD45A expression have better survival outcomes (Sup Fig. 4C).
To further validate whether SNHG6 and GADD45A are truly involved in the DNA methylation process in cancer, we conducted overexpression experiments of SNHG6 and GADD45A in HepG2 and knockdown experiments of GADD45A in 293 T and SNHG6 in Huh7, respectively. The over-expression and knockdown efficiencies were validated by qPCR (Sup Fig. 4D–G) and RNA-Seq. We performed WGBS on all the edited cell lines to examine the DNA methylation changes. Although there is no global change in DNA methylation levels (Sup Fig. 4H), we found that there were quite a number of differentially methylated regions (DMRs) between the edited cell lines and their respective control. As expected, the DMRs in the overexpression experiments were predominantly hypomethylated (GADD45A 83.8%, SNHG6 67.3%) (Fig. 4F, G), , whereas the DMRs in the knockdown experiments were predominantly hypermethylated (GADD45A 59.4%, SNHG6 67.8%) (Fig. 4H, I). These results validated our prediction that these two genes are likely involved in regulating the DNA demethylation process.
DNA methylation patterns can discriminate the origins between tumor nodules
Lastly, since we collected multiple nodules from the same tumors, we wanted to investigate what is the most accurate strategy to infer cancer evolution trajectory using single-cell data. To achieve this, we mainly focus on HCC2 as an example because it has the most complete data. First of all, we performed whole-exome sequencing (WES) in tumor nodules from HCC2, and then we used copy number variants (CNVs) and single nucleotide variants (SNVs) from WES data to build the trees. Both results show that PT1 and PT2 shared more variants than PT3, suggesting that PT1 and PT2 may be from a common ancestor (Fig. 5A, B). CNVs inferred from scRNA-seq data and DNA methylation data are also consistent with WES data (Sup Fig. 5A, B). Interestingly, PT1 and PT2 also show a similar DNA hypomethylation profile, which is very distinct from PT3 (Fig. 5C). On the contrary, at the transcriptome level, PT2 and PT3 are more similar to each other and are most distant from PT1, as shown by correlation analysis (Fig. 5D). If we treat WES data as the gold standard, this result suggests that DNA methylation profile may be a more faithful representation of the clonal memory than transcriptional profiles, which is supported by a recent publication65. However, we note that this conclusion is primarily derived from the most complete multimodal dataset (patient HCC2), and thus should be considered as an observational result with limitations due to the sample size.
DNA methylation patterns can discriminate the origins between Tumor Nodules (A) Heatmap showing CNV clustering result called by CNYkit based on WES data. (B) Heatmap showing SNV allele frequency clustering based on WES data. (C) Heatmap showing DNA methylation level clustering result based on pseudo bulk scPBAT data. (D)Umap plot showing RNA expression level clustering result based on scRNA data.
Discussion
In the current study, we provide crucial insights into the epigenetic and immunological landscapes of distinct tumor nodules in liver cancer patients. Our result revealed that CMN samples exhibit distinct immune landscapes, characterized by greater transcriptome heterogeneity and more complex immune-related interactions compared to SN samples. This heightened complexity suggests that CMN nodules may be subjected to unique selective pressures, potentially fostering immune evasion or modulation66.
Furthermore, we observed global DNA hypomethylation in PMDs within some of the nodules. Notably, GADD45A and SNHG6 were identified as potential regulators of this hypomethylation, consistent with their established roles in DNA repair and epigenetic modification. Our data aligns with previous findings that hypomethylation in PMDs is a hallmark of tumorigenesis. The heterogeneity of division-induced DNA demethylation among patients may potentially originate from differential capabilities in maintaining nuclear laminarity under high cellular division rates. It is noteworthy that patients with high levels of DNA methylation generally demonstrate improved survival67. Conversely, patients exhibiting demethylation have shown a more significant response to immunotherapy67. This difference could potentially be linked to variations in the DNA damage repair process. Also, hypomethylation often targets immune-related loci or heterochromatin, suggesting a dual role for hypomethylation in promoting immune evasion as well as tumor growth68.
Further analysis demonstrated that DNA methylation distance serves as a more reliable feature for inferring cancer evolution trajectories compared to gene expression distance. This finding supports the notion that epigenetic alterations, particularly changes in DNA methylation patterns, may precede transcriptional changes during tumor progression. These selective pressures likely shape the evolutionary paths of distinct tumor nodules.
In conclusion, our findings highlight the critical role of DNA methylation dynamics, particularly within PMDs, in liver cancer progression by mediating the balance between immune evasion and tumor proliferation. Future studies focusing on the precise mechanisms governing these epigenetic alterations may uncover novel therapeutic targets aimed at restoring immune surveillance and inhibiting tumor growth.
Limitations
Our study also has certain limitations. The technical challenges associated with library construction limited the sample size, which may affect the generalizability of our findings. Additionally, the exclusion of certain samples due to macrophage over-representation may impact the completeness of the immune landscape characterization. Thirdly, although we identifies potential hypomethylation regulators, such as GADD45A and SNHG6, functional validation is limited to in vitro experiments using cancer cell lines. Further in vivo studies and validation in clinical samples are necessary to confirm the role of these genes in HCC progression and their potential as therapeutic targets.
Future directions
Further research should focus on validating our findings in larger, more diverse patient cohorts and exploring the therapeutic potential of targeting epigenetic regulators like SNHG6 and GADD45A, Developing epigenetic therapies that specifically target hypomethylated regions or modulate key regulatory genes holds promise for improving HCC treatment outcomes. Additionally, our results shows that multi-omics techniques, such as scTrio-seq2, could be extended to other cancer types to understand the interplay between DNA methylation, gene expression, and immune dynamics in shaping tumor heterogeneity.
References
Jokhadze, N., Das, A. & Dizon, D. S. Global cancer statistics: A healthy population relies on population health. CA Cancer J. Clin. 74, 224–226 (2024).
Llovet, J. M. et al. Hepatocellular carcinoma. Nat. Rev. Dis. Primer 7, 1–28 (2021).
Ishizawa, T. et al. Neither multiple tumors nor portal hypertension are surgical contraindications for hepatocellular carcinoma. Gastroenterology 134, 1908–1916 (2008).
Villanueva, A. Hepatocellular carcinoma. N. Engl. J. Med. https://doi.org/10.1056/NEJMra1713263 (2019).
Ma, L. et al. Tumor cell biodiversity drives microenvironmental reprogramming in liver cancer. Cancer Cell 36, 418-430.e6 (2019).
Gonçalves, E., Gonçalves-Reis, M., Pereira-Leal, J. B. & Cardoso, J. DNA methylation fingerprint of hepatocellular carcinoma from tissue and liquid biopsies. Sci. Rep. 12, 11512 (2022).
Fan, G. et al. DNA methylation biomarkers for hepatocellular carcinoma. Cancer Cell Int. 18, 140 (2018).
Cheng, J. et al. Integrative analysis of DNA methylation and gene expression reveals hepatocellular carcinoma-specific diagnostic biomarkers. Genome Med. 10, 42 (2018).
Yamada, N. et al. Genome-wide DNA methylation analysis in hepatocellular carcinoma. Oncol. Rep. 35, 2228–2236 (2016).
Baylin, S. B. & Jones, P. A. Epigenetic determinants of cancer. Cold Spring Harb. Perspect. Biol. 8, a019505 (2016).
Angeloni, A. & Bogdanovic, O. Enhancer DNA methylation: Implications for gene regulation. Essays Biochem. 63, 707–715 (2019).
Lister, R. et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462, 315–322 (2009).
Slotkin, R. K. & Martienssen, R. Transposable elements and the epigenetic regulation of the genome. Nat. Rev. Genet. 8, 272–285 (2007).
Hansen, K. D. et al. Increased methylation variation in epigenetic domains across cancer types. Nat. Genet. 43, 768–775 (2011).
Berman, B. P. et al. Regions of focal DNA hypermethylation and long-range hypomethylation in colorectal cancer coincide with nuclear lamina–associated domains. Nat. Genet. 44, 40–46 (2012).
Hon, G. C. et al. Global DNA hypomethylation coupled to repressive chromatin domain formation and gene silencing in breast cancer. Genome Res. 22, 246–258 (2012).
Salhab, A. et al. A comprehensive analysis of 195 DNA methylomes reveals shared and cell-specific features of partially methylated domains. Genome Biol. 19, 150 (2018).
Zhou, W. et al. DNA methylation loss in late-replicating domains is linked to mitotic cell division. Nat. Genet. 50, 591–602 (2018).
Hur, K. et al. Hypomethylation of long interspersed nuclear element-1 (LINE-1) leads to activation of proto-oncogenes in human colorectal cancer metastasis. Gut 63, 635–646 (2014).
Intragenic DNA methylation in transcriptional regulation, normal differentiation and cancer. Biochim. Biophys. Acta BBA Gene Regul. Mech. 1829, 1161–1174 (2013).
Tubio, J. M. C. et al. Extensive transduction of nonrepetitive DNA mediated by L1 retrotransposition in cancer genomes. Science 345, 1251343 (2014).
He, J. et al. The clinicopathologic and prognostic significance of gross classification on solitary hepatocellular carcinoma after hepatectomy. Medicine 94, e1331 (2015).
Shimada, M. et al. The role of macroscopic classification in nodular-type hepatocellular carcinoma. Am. J. Surg. 182, 177–182 (2001).
Bian, S. et al. Single-cell multiomics sequencing and analyses of human colorectal cancer. Science 362, 1060–1063 (2018).
Dong, J. et al. Single-cell RNA-seq analysis unveils a prevalent epithelial/mesenchymal hybrid state during mouse organogenesis. Genome Biol. 19, 31 (2018).
Smallwood, S. A. et al. Single-cell genome-wide bisulfite sequencing for assessing epigenetic heterogeneity. Nat. Methods 11, 817–820 (2014).
Zheng, G. X. Y. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Petukhov, V. et al. dropEst: Pipeline for accurate estimation of molecular counts in droplet-based single-cell RNA-seq experiments. Genome Biol. 19, 1–16 (2018).
Dobin, A. et al. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: An efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2024).
McGinnis, C. S., Murrow, L. M. & Gartner, Z. J. DoubletFinder: Doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 8, 329-337.e4 (2019).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with harmony. Nat. Methods 16, 1289–1296 (2019).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterProfiler: An R package for comparing biological themes among gene clusters. OMICS J. Integr. Biol. 16, 284–287 (2012).
Jin, S. et al. Inference and analysis of cell-cell communication using cell chat. Nat. Commun. 12, 1088 (2021).
Krueger, F. & Andrews, S. R. Bismark: A flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 27, 1571–1572 (2011).
Raivo Kolde. pheatmap: Pretty Heatmaps. 1.0.12 (2010).
Quinlan, A. R. & Hall, I. M. BEDTools: A flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Ally, A. et al. Comprehensive and integrative genomic characterization of hepatocellular carcinoma. Cell 169, 1327-1341.e23 (2017).
Wu, H. et al. Detection of differentially methylated regions from whole-genome bisulfite sequencing data without replicates. Nucleic Acids Res. 43, e141 (2015).
Li, H. & Durbin, R. Fast and accurate short read alignment with burrows–wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
McKenna, A. et al. The genome analysis toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Van der Auwera, G. A. & O’Connor, B. D. Genomics in the Cloud: Using Docker, GATK, and WDL in Terra (1st Edition) (O’Reilly Media, 2020).
Poplin, R. et al. Scaling accurate genetic variant discovery to tens of thousands of samples. 201178 Preprint at https://doi.org/10.1101/201178 (2018).
Talevich, E., Shain, A. H., Botton, T. & Bastian, B. C. CNVkit: Genome-wide copy number detection and visualization from targeted DNA sequencing. Plos Comput. Biol. 12, e1004873 (2016).
Tickle, T., Tirosh, I., Georgescu, C., Brown, M. & Haas, B. inferCNV of the Trinity CTAT Project. (2019).
Chen, T. et al. The genome sequence archive family: Toward explosive data growth and diverse data types. Genomics Proteomics Bioinform. 19, 578–583 (2021).
CNCB-NGDC Members and Partners et al. Database Resources of the National Genomics Data Center, China National Center for Bioinformation in 2024. Nucleic Acids Res. 52, D18–D32 (2024).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Ren, L. et al. Apolipoproteins and cancer. Cancer Med. 8, 7032–7043 (2019).
Basavaraju, P. et al. Genetic regulatory networks of apolipoproteins and associated medical risks. Front. Cardiovasc. Med. 8, 788852 (2022).
Liu, J. et al. NUPR1 is a critical repressor of ferroptosis. Nat. Commun. 12, 647 (2021).
Li, T. et al. IGFBP2: Integrative hub of developmental and oncogenic signaling network. Oncogene 39, 2243–2257 (2020).
Wang, H., Zhang, W., Zhu, H., Li, Q. & Miao, L. Long noncoding RNA SNHG6 mainly functions as a competing endogenous RNA in human tumors. Cancer Cell Int. 20, 219 (2020).
Guo, T. et al. SNHG6 acts as a genome-wide hypomethylation trigger via coupling of miR-1297–mediated S-adenosylmethionine–dependent positive feedback loops. Cancer Res. 78, 3849–3864 (2018).
Ansari, H. et al. Long noncoding RNAs in colorectal adenocarcinoma; an in silico analysis. Pathol. Oncol. Res. 25, 1387–1394 (2019).
Shao, Q. et al. SNHG 6 promotes the progression of colon and Rectal adenocarcinoma via miR-101-3p and Wnt/β-catenin signaling pathway. BMC Gastroenterol. 19, 163 (2019).
Wu, G., Ju, X., Wang, Y., Li, Z. & Gan, X. Up-regulation of SNHG6 activates SERPINH1 expression by competitive binding to miR-139-5p to promote hepatocellular carcinoma progression. Cell Cycle 18, 1849–1867 (2019).
Li, Z. et al. Gadd45a promotes DNA demethylation through TDG. Nucleic Acids Res. 43, 3986 (2015).
Zhou, L. et al. GADD45a promotes active DNA demethylation of the MMP-9 promoter via base excision repair pathway in AGEs-treated keratinocytes and in diabetic male rat skin. Endocrinology 159, 1172–1186 (2018).
Arab, K. et al. Long noncoding RNA TARID directs demethylation and activation of the tumor suppressor TCF21 via GADD45A. Mol. Cell 55, 604–614 (2014).
He, Y.-F. et al. Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science 333, 1303–1307 (2011).
Weber, A. R. et al. Biochemical reconstitution of TET1–TDG–BER-dependent active DNA demethylation reveals a highly coordinated mechanism. Nat. Commun. 7, 10806 (2016).
Li, L. et al. A mouse model with high clonal barcode diversity for joint lineage, transcriptomic, and epigenomic profiling in single cells. Cell 186, 5183-5199.e22 (2023).
Ueshima, E. et al. Gadoxetic acid-enhanced magnetic resonance imaging features can predict immune-excluded phenotype of hepatocellular carcinoma. Liver Cancer 14, 271–285 (2024).
Lakshminarasimhan, R. & Liang, G. The role of DNA methylation in cancer. Adv. Exp. Med. Biol. 945, 151–172 (2016).
Guo, H. et al. DNA hypomethylation silences anti-tumor immune genes in early prostate cancer and CTCs. Cell 186, 2765-2782.e28 (2023).
Acknowledgements
We thank the High-performance Computing Center of Westlake University for providing computational and storage resources. Our Flow Cytometry work was performed at the Advanced Cell Technology Core Facility, Guangzhou National Laboratory. This work is funded by the Major Project of Guangzhou National Laboratory (No: GZNL2024A03001 to X.F.); The National Natural Science Foundation of China (32370710 to Y.Z.); The Guangdong Provincial Pearl River Talents Program (2021QN02Y747 to X.F.); The Guangzhou Science and Technology Elite Pilot Project (SL2024A04J01788 to X.F.); National Key R&D Program of China ( 2022YFA1302700 to Y.Z.).
Author information
Authors and Affiliations
Contributions
X.F.,Y.Z. and J.Z. designed the research; D.H., and Y.Z. confirmed patients’ diagnoses, annotated clinical information and processed patient and donor samples; L.Z., D.S., L.C., Y.G., T.K., Q.H., L.X., X.X.and W.M. performed experiments; L.Z., X.P.,E.D. and Z.S. performed computational analyses; L.Z. wrote the manuscript, with inputs from other authors. All authors discussed the results and reviewed the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
This study was approved by the Ethics Committee of Sun Yat-sen University Cancer Center (approval number: B2022-700-01). All procedures involving human participants were conducted in accordance with the ethical standards of the institutional and national research committee and with the 1964 Declaration of Helsinki and its later amendments or comparable ethical standards. Written informed consent was obtained from all individual participants included in the study. All participants were legally adults (aged 18 years or older) and had full legal capacity to provide consent independently at the time of recruitment. This study did not involve any experiments on live vertebrates or higher invertebrates.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, L., Hu, D., Peng, X. et al. Interrogating subclonal heterogeneity of liver cancer with single-cell multi-omics analysis. Sci Rep 15, 39021 (2025). https://doi.org/10.1038/s41598-025-24732-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-24732-y