Abstract
Wound healing is a critical aspect of modern medicine, impacting patient health, quality of life, and healthcare resource allocation. Okanin, a flavonoid from the Asteraceae family, has shown potential in promoting wound healing. This study investigates okanin’s key molecular targets, binding affinity, and mechanisms of action using network pharmacology, molecular docking, molecular dynamics simulations, and in vivo experimental validation. Okanin’s potential targets were identified using the Comparative Toxicogenomics Database (CTD) and SwissTargetPrediction, while wound healing-related targets were sourced from GeneCards and DrugBank. Overlap analysis of these datasets revealed common targets. Key target proteins were filtered through protein-protein interaction (PPI) analysis using the STRING database. Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses were conducted using Metascape to build a drug-target-pathway-disease network. Molecular docking was performed with AutoDockTools, and binding affinity was evaluated through energy scores, particularly with AURKA and HDAC1. Molecular dynamics simulations with GROMACS confirmed the stability of okanin-target complexes. ADME/T properties were assessed using SwissADME and ProTox-3.0 to evaluate pharmacokinetics and toxicity. In vivo quantitative real-time PCR (qRT-PCR) was performed to assess the expression of selected target genes in a mouse wound model following topical okanin treatment. A total of 72 common targets were identified between okanin and wound healing. PPI network analysis highlighted 17 key targets, with molecular docking revealing the highest binding affinity for AURKA and HDAC1 (ΔG = − 8.8 kcal/mol for both). GROMACS were then run on the top complexes. Target-ligand stability was quantified by convergence of RMSD/Rg, sustained hydrogen-bond counts, and MM/GBSA binding free energies (AURKA, − 24.27 ± 3.65 kcal/mol; HDAC1, − 47.7 ± 1.60 kcal/mol), confirming robust interactions. SwissADME predicted good drug-likeness (MW = 288.25 g/mol; logP = 1.69; high GI and moderate skin permeability) and no P-gp liability, while ProTox-3.0 indicated low systemic toxicity (LD₅₀ = 2500 mg/kg). qRT-PCR results demonstrated that okanin treatment significantly downregulated AURKA and PIK3R1, while upregulating HDAC1, in wounded skin, supporting the predicted molecular interactions and regulatory functions. Okanin promotes wound healing through multiple molecular targets and pathways, including antioxidant, anti-inflammatory, and cell proliferation mechanisms. Its high binding affinity for AURKA and HDAC1, along with modulation of the IL-17 and AMPK signaling pathways, underscores its therapeutic potential. This study provides a comprehensive theoretical and experimental framework for the development of okanin as a topical agent for wound healing, with future research focusing on formulation development and translational applications.
Similar content being viewed by others
Introduction
In contemporary medicine, wound healing is a domain of critical importance, as it not only directly influences patients’ health and quality of life, but also significantly affects the efficient allocation of healthcare resources and the overall management of medical costs1. The wound healing process is highly intricate, involving multifaceted interactions among various cell types, growth factors, and signaling pathways. AURKA initiates the transcription of autophagy-related genes, thereby enhancing cell survival while concomitantly increasing microvascular density and shortening the overall healing interval2,3. HDAC1 mitigates excessive inflammation during the inflammatory phase by modulating innate immune responses and, in the early proliferative phase, epigenetically regulates keratinocyte activity4,5. MMP3 drives dynamic remodeling of the extracellular matrix, creating a permissive migratory track for keratinocytes and fibroblasts and liberating matrix-sequestered EGF and TGF-β, which collectively accelerate wound closure6. During the inflammatory stage, PIK3R1 governs macrophage adhesion and polarization. Subsequently, activation of the PI3K–Akt–mTOR axis facilitated by PIK3R1 promotes angiogenesis and fibroblast-mediated collagen deposition, further advancing tissue repair7,8,9 (Fig. 1). Despite the recent proliferation of wound healing-related therapeutic products, the treatment of chronic non-healing wounds remains a major clinical challenge, particularly in diabetic patients and bedridden individuals.
Schematic representation of the roles of AURKA, HDAC1, MMP3, and PIK3R1 in cutaneous wound healing.
As the number of surgical procedures increases and the aging population grows, wound healing in postoperative wounds has become a focal point for surgeons. Wounds that fail to heal after surgery can become chronic. Furthermore, chronic wounds, such as diabetic foot ulcers and pressure sores, pose significant therapeutic challenges due to their resistance to treatment and high recurrence rates10. The risk of foot ulcers in diabetic patients ranges from 19–34%11, with a recurrence rate as high as 65% within five years, and severe cases may lead to amputation12. Additionally, between 3.5 and 69% of hospitalized patients are affected by pressure injuries, affecting up to 2.5 million patients annually13,14,15, with complications that may lead to wound-related infections and fatalities (> 55%)16,17. At present, the principal therapeutic modalities for chronic wounds are categorized into two main approaches: surgical debridement and growth factor therapy18. Surgical debridement does not result in a cure for chronic conditions, and repeated surgical interventions worsen patient suffering and diminish compliance. Therapies involving growth factors face challenges such as short half-life, instability, rapid dispersion, inhibition of signal control, and potential systemic adverse effects, thereby imposing physiological burdens on patients19,20. Consequently, the development of novel wound healing agents continues to be a critical area of research, offering additional therapeutic options for patients with wounds.
Okanin (C₁₅H₁₂O₆, Fig. 2), a flavonoid derived from the Asteraceae family of plants, is primarily obtained from the traditional Chinese medicinal herb Bidens pilosa L. The earliest records of Bidens pilosa L. can be traced back to the Tang Dynasty medical text Bencao Shiyi, which has been historically renowned for treating contusions and strains, as well as alleviating inflammatory responses. Okanin is one of the main active constituents of Bidens pilosa L., and its chemical structure is known as 3,4,2’,3’,4’-pentahydroxychalcone. The molecular structure of okanin contains two benzene rings, one substituted with three hydroxyl (-OH) groups and the other with two hydroxyl groups and an α,β-unsaturated ketone group. This unique molecular architecture contributes to its distinct physicochemical properties and underpins its various biological activities. Okanin has recently garnered interest due to its pharmacological potential21. It has been shown to significantly suppress LPS-induced expression of iNOS, as well as the production and mRNA expression of IL-6 and TNF-α in LPS-stimulated BV-2 cells. Its pharmacological effects include antioxidant22, anti-inflammatory23, vascular endothelial cell repair24, hepatoprotective25, and anticancer activities26. Notably, okanin exhibits substantial protective effects in the wound healing process, with mechanisms involving multiple signaling pathways such as anti-inflammatory responses, antioxidant actions, and endothelial repair27,28,29. However, the precise roles and mechanisms of okanin in wound healing remain incompletely understood. This study aims to elucidate the potential therapeutic targets and relevant signaling pathways of okanin in wound healing using network pharmacology, molecular docking, and molecular dynamics simulations.
The structural formula of okanin.
Network pharmacology, which integrates bioinformatics and network-based analytical tools, plays a pivotal role in elucidating interactions between drugs and their biological targets within the context of Traditional Chinese Medicine (TCM)30. This approach uncovers how active ingredients engage with molecular targets in TCM, thereby elucidating the underlying mechanisms and potential therapeutic effects31. By constructing interaction networks, researchers can predict novel targets and gain a deeper understanding of the multi-target actions and holistic therapeutic potential of TCM formulations⁽²⁶⁾. This methodology facilitates both TCM applications and novel drug discovery efforts, offering valuable insights into multifaceted therapeutic strategies. Specifically, network pharmacology enabled us to identify the potential targets of okanin and provided insights into the biological pathways associated with wound healing, thereby offering a theoretical basis for subsequent experimental validation.
Molecular dynamics (MD) simulation is an atomistic-level computational technique that models biomolecular behavior and provides detailed insights into dynamic changes in protein structure and function, predicts protein–ligand interactions, and offers a faster, more cost-effective alternative to wet-lab experiments—thereby reducing reliance on animal models and supporting preclinical validation and accelerated drug discovery32. When combined with enhanced sampling, MD can reveal previously unknown intermediate conformations and transition pathways, strengthening rational drug design33. Building on network pharmacology, molecular docking was used to identify high-affinity targets implicated in wound healing, and MD then tested whether okanin maintains stable, biologically meaningful engagement with these targets under near-physiological conditions, enabling a more comprehensive understanding of binding stability and mechanistic behavior34.
Collectively, the integration of network pharmacology, molecular docking, and molecular dynamics simulations allows for a comprehensive analysis of the potential mechanisms of okanin in wound healing. The complementary application of these three methods not only uncovered the possible targets and pathways of okanin but also provided theoretical guidance for subsequent experimental validation.
Methods and materials
Common targets prediction of Okanin and wound healing
Potential targets of okanin were retrieved from the Comparative Toxicogenomics Database (CTD)35 and SwissTargetPrediction36. Wound healing–related targets were obtained from GeneCards37 and DrugBank38 databases. After eliminating duplicate entries, the overlap between the two datasets was evaluated to determine the potential shared targets of okanin in the context of wound healing.
Protein-protein interaction network construction and key target screening
PPI analysis was performed on the predicted common targets using the STRING database (https://www.string-db.org)39. To ensure data reliability and reduce the incidence of false-positive interactions, a confidence score threshold of ≥ 0.7, as recommended by STRING, was applied in the network analysis40. The resulting data were subsequently imported into Cytoscape 3.9.1 for network visualization. Node size was adjusted according to degree centrality, and edge thickness was proportional to the interaction confidence score. Ultimately, a network map was constructed to illustrate the protein–protein interactions among the identified targets.
Key targets associated with okanin in the wound healing process were identified using topological centrality metrics, specifically by comparing the median Degree and Closeness centrality values. The resulting interaction network was visualized using Cytoscape 3.9.1 software. The Degree value represents the number of direct connections associated with a node, and an increased number of connections correlates with enhanced node influence. Betweenness centrality (BC) refers to the shortest paths between all pairs of nodes within the network, where a lower value suggests the node is less central. Closeness centrality (CC) is defined as the inverse of the average shortest path length between a particular node and all other nodes in the network, with lower values indicating reduced efficiency in signal propagation among the nodes.
ADME/Toxicity analysis of Okanin
The absorption, distribution, metabolism, and excretion (ADME) profile of okanin was predicted using SwissADME, a freely accessible web tool designed to evaluate pharmacokinetics, drug-likeness, and medicinal chemistry properties41. The molecular structure of okanin was provided as a canonical SMILES string (C1 = CC(= C(C = C1/C = C/C(= O)C2 = C(C(= C(C = C2)O)O)O)O)O), which was used to predict key physicochemical properties, such as molecular weight (MW), lipophilicity (LogP), and topological polar surface area (TPSA). Furthermore, SwissADME assessed okanin’s compliance with common drug-likeness filters, including Lipinski’s Rule of Five.
ProTox-3.0, an online platform for toxicity prediction, was employed to estimate the compound’s potential for organ-specific toxicities42. Utilizing structural similarity and machine learning models, ProTox-3.0 predicted various toxicological endpoints, including hepatotoxicity, neurotoxicity, nephrotoxicity, and respiratory toxicity, and provided the estimated LD50 values for okanin.
Enrichment analysis of key targets in GO functions and KEGG pathways
The key targets identified were then imported into Metascape (https://www.metascape.org) for further functional enrichment analysis. Enrichment parameters were set to select “Homo sapiens” as the species of interest, with a minimum gene count of three, a significance threshold of P ≤ 0.05, and an enrichment fold change cutoff of at least 1.5. These criteria were applied for both GO terms and KEGG pathways. A bubble chart was generated to visualize the enrichment results for GO terms and KEGG pathways. Additionally, the enrichment scores were sorted in descending order, and a bar chart was generated to highlight the top 10 GO terms.
Establishing a ‘drug-target-pathway-disease’ network
Using Cytoscape 3.9.1, a network map was constructed to illustrate the relationships among “drug-target-pathway-disease.” This map elucidates the connections between key targets and their associated pathways. Only the top 10 KEGG pathways, identified by the lowest P-values, were included in the analysis. Consequently, targets not associated with these pathways were excluded.
Molecular Docking of Okanin to key targets
The protein structures for molecular docking were obtained from the Protein Data Bank (PDB) (http://www.rcsb.org)43. Target proteins were selected based on criteria such as protein sequence, resolution, and the location of the active binding pocket. Prior to docking, the protein structures were prepared using PyMOL, which included the addition of hydrogen atoms and the removal of water molecules to ensure the proteins were ready for docking simulations. The okanin compound was prepared in AutoDockTools, with flexible bonds adjusted to facilitate optimal fitting during the docking process. Molecular docking of okanin with key targets was performed using the vina module of AutoDockTools 1.5.7, and the grid box was adjusted to ensure it encompassed the entire target protein. The binding affinity (kcal/mol) was used to represent the binding free energy between okanin and the target proteins involved in wound healing; a higher absolute value of binding free energy indicates a more stable ligand-receptor interaction.
Molecular dynamic simulations of okanin-target protein complexes
Molecular dynamics simulations were performed using GROMACS version 2022.344. The small molecule ligand was prepared using AmberTools22, with the GAFF force field applied using the tleap module. Hydrogen atom addition and RESP charge calculations were performed using Gaussian 16 W. The preparation process was carried out under neutral pH (pH 7) and 298 K temperature conditions, with the ligand in its neutral ionization state, as per the default settings of AmberTools22. The resulting charge and parameter data were incorporated into the topology file to enable subsequent simulations. All simulations employed the Amber99SB-ILDN force field, with the TIP3P water model used as the solvent. The system was neutralized by adding Na+ ions to counterbalance any net charge.
The simulation began with energy minimization via the steepest descent method, followed by equilibration under NVT (constant number, volume, and temperature) and NPT (constant number, pressure, and temperature) ensembles, each for 100,000 steps. Equilibration was conducted with a coupling constant of 0.1 ps over a 100 ps period. Production simulations were run for 100 nanoseconds with a time step of 2 femtoseconds, totaling 5,000,000 steps.
Post-simulation analysis was performed using integrated GROMACS tools, including the calculation of root-mean-square deviation (RMSD), root-mean-square fluctuation (RMSF), and radius of gyration (Rg) for each amino acid residue. Additional analyses included binding free energy (MM/GBSA), free energy landscapes, and other relevant metrics. Binding free energy was considered a key indicator of drug activity; a more negative value indicates a stronger and more stable ligand-receptor complex.
Excisional wound model in mice and topical Okanin treatment
Twelve SPF-grade male Kunming mice (8–10 weeks old) were obtained from Sipeifu (Beijing) Biotechnology Co., Ltd. (License No. SCXK [Jing] 2019-0010). Animals were randomly assigned to three groups (n = 4 per group): Control (wounded, no treatment), Vehicle (wounded, blank emulsion), and Okanin (wounded, 2% okanin emulsion). All animal procedures were commissioned to Ampry Biosciences Co., Ltd. (Fujian, China) under a service agreement. Experiments were performed by Ampry personnel under protocols approved by its Institutional Animal Care and Use Committee (IACUC) on 20 May 2025 (Approval No. IACUC-FJABR-2025052001). The study was designed and supervised by the corresponding author.
Under anesthesia with 1% sodium pentobarbital (40–50 mg/kg, i.p.), the dorsal fur was shaved and disinfected. A full-thickness circular wound (8 mm diameter) was created on the mid-dorsal region using a sterile biopsy punch. Mice in the Control group received no topical treatment. The Vehicle group received topical application of the blank emulsion every other day, while the Okanin group received the emulsion containing 2% (w/v) okanin with the same dosing schedule. Each application evenly covered the wound surface (0.2 g/cm2)45. On day 8 post-injury, mice were sacrificed with 1% sodium pentobarbital (70–80 mg/kg, i.p.), and wound tissues were collected and stored at − 80 °C for subsequent analysis. Animals were monitored twice daily (three times daily during the first 72 h post-operation). Prespecified humane endpoints included ≥ 20% body-weight loss, persistent lethargy/hunched posture or inability to eat/drink, severe wound necrosis/ulceration or uncontrolled infection/bleeding, and respiratory distress. Animals meeting any endpoint were euthanized by intraperitoneal sodium pentobarbital (70–80 mg/kg); death was confirmed by apnea and loss of pedal reflex.
The emulsion base was prepared as an oil-in-water formulation. The oil phase contained 1 g glycerol monostearate, 4 g stearic acid, 4 g liquid paraffin, and 5 g white petroleum jelly. The aqueous phase contained 1 g sodium dodecyl sulfate, 10 g glycerol, and 25 g distilled water. Both phases were heated and emulsified under continuous stirring. For the treatment group, okanin (purity ≥ 98%) was purchased from Shanghai Yuanye Bio-Technology Co., Ltd. (Shanghai, China; Cat. No. B26397) and incorporated into the emulsion at a final concentration of 2% (w/v).
Quantitative real-time PCR (qRT-PCR)
Total RNA was extracted from wound tissues using RNAiso Plus (Takara, Japan), and reverse-transcribed into cDNA using a commercial kit (Novoprotein, China). qPCR was performed with F488 SYBR Green qPCR Mix (Yugong Biotech, China) on an ABI 7300 system (Applied Biosystems, USA). GAPDH was used as the internal control. Each sample was measured in triplicate, and relative mRNA expression levels were calculated using the 2^–ΔCt method. Primer sequences are listed in Table S3. Raw Ct values, primer sequences, calculations and statistics are provided in Table S4-S7.
Results
ADME/Toxicity profile of Okanin
The ADME/Toxicity profile of okanin was predicted using SwissADME and ProTox-3.0. The compound has a molecular weight of 288.25 g/mol, which is favorable for absorption. It contains 21 heavy atoms, 12 of which are aromatic. okanin’s logP value suggests moderate lipophilicity, which is beneficial for membrane permeability and absorption. The TPSA of 118.22 Å2 indicates a balanced hydrophilic-lipophilic profile. Additionally, okanin is predicted to have high gastrointestinal (GI) absorption, a crucial feature for its oral bioavailability. The compound is also soluble in water and is not a substrate for P-glycoprotein (P-gp), further enhancing its potential for efficient absorption (Table S1).
Toxicity predictions indicate a low risk for hepatotoxicity (inactive, probability: 0.62) and neurotoxicity (inactive, probability: 0.87). However, moderate risks are predicted for nephrotoxicity (active, probability: 0.60) and respiratory toxicity (active, probability: 0.52). The predicted LD50 for okanin is 2500 mg/kg, classifying it in toxicity class V (possibly hazardous) (Table S2 and Figure S1). These results suggest that while okanin is generally safe, further studies are needed to evaluate its safety concerning kidney and lung toxicity.
Common targets prediction
From the CTD database, three protein targets that appeared in multiple entries were identified, while 103 protein targets were retrieved from the SwissTargetPrediction database. After removing duplicates, a total of 105 okanin-associated targets were obtained.
From the GeneCards database, 3,349 wound healing-related targets were identified based on a Relevance Score threshold set at the median value. Additionally, 35 targets were retrieved from the DrugBank database. Following deduplication, 3,359 non-redundant wound healing-related targets were compiled.
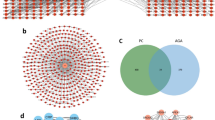
By intersecting the okanin-related and wound healing-related target sets, 72 common protein targets were finally obtained, suggesting a potential role of okanin in the regulation of wound healing processes.
Protein-protein interaction network analysis
The 72 potential shared targets associated with okanin and wound healing were subjected to PPI analysis using the STRING platform, and were subsequently visualized using Cytoscape 3.9.1. A network diagram was constructed to illustrate the interconnections between these shared targets.
In the resulting network, node size and color represent the degree values of the proteins, while the width and shade of the edges correspond to the STRING combined interaction scores. To improve the clarity of target connectivity, nodes with fewer than three edges were excluded, yielding a refined network containing 65 nodes (i.e., predicted common targets) and 145 edges representing the interactions among them (Fig. 3a).
a Interaction network of shared targets for okanin and wound healing. b: Interaction network of key targets for okanin against wound healing.
Key targets
We employed three topological network metrics: degree value, CC, and BC. To comprehensively identify key nodes and reduce potential statistical biases due to distribution skewness, the median value was set as the primary screening criterion46. The selection criteria were as follows: (1) degree value ≥ 3, (2) BC ≥ 0.01, and (3) CC ≥ 0.33. Seventeen targets meeting these criteria were identified, indicating their significant role in the network and potential involvement in wound healing. The gene names, degree values, BC, and CC for the key targets are summarized in Table 1. The interaction network of the key targets is shown in Fig. 3b.
The five key targets with the highest degree values are HSP90AA1 (degree = 23), ESR1 (degree = 14), IGF1R and PTGS2 (each with a degree of 10), and CDK1 and PIK3R1 (each with a degree of 9).
GO function analysis
Seventeen key targets were subjected to GO enrichment analysis via the Metascape platform. GO functional annotations are classified into three domains: BP, CC, and MF. The analysis revealed a total of 258 enriched biological processes, 21 cellular components, and 26 molecular functions. The top 10 terms from each category, based on enrichment scores, were selected and visualized using a histogram (Fig. 4) and a bubble chart (Fig. 5a).
Histogram of top 10 degree values from gene GO function enrichment analysis.
Bubble diagram of top 10 degree entries in enrichment analysis results a: GO function enrichment analysis. b: KEGG pathway enrichment analysis.
The top 10 enriched biological processes were: positive regulation of protein import into nucleus, insulin like growth factor receptor signaling pathway, regulation of protein import into nucleus, positive regulation of nucleocytoplasmic transport, G2/M transition of mitotic cell cycle, regulation of phospholipase activity, cell cycle G2/M phase transition, egative regulation of muscle cell apoptotic process, response to amyloidbeta, insulin receptor signaling pathway.
The top 10 enriched cellular components were: growth cone, site of polarized growth, neuronal cell body, distal axon, cell body, membrane raft, membrane microdomain, ransferase complex, transferring phosphorus containing groups, endoplasmic reticulum lumen, secretory granule lumen.
The top 10 enriched molecular functions were: nuclear receptor activity, ligandactivated transcription factor activity, 8Ebox binding, transcription coregulator binding, heat shock protein binding, proteinfolding chaperone binding, histone modifying activity, ubiquitin protein ligase binding, ubiquitinlike protein ligase binding, RNA polymerase II specific DNAbinding transcription factor binding.
KEGG pathway enrichment analysis
Seventeen key targets were subjected to KEGG pathway enrichment analysis via the Metascape platform, which led to the identification of 91 distinct signaling pathways. Based on enrichment scores, the top 10 KEGG pathways were selected and visualized in a bubble chart (Fig. 5b)47,48,49. The identified pathways and the corresponding annotations for the associated targets are provided in Table 2.
The top 10 enriched KEGG pathways were: hsa04213: Longevity regulating pathway - multiple species, Prostate cancer, Progesterone-mediated oocyte maturation, Longevity regulating pathway, AMPK signaling pathway, Thyroid hormone signaling pathway, IL-17 signaling pathway, Endocrine resistance, Estrogen signaling pathway, Lipid and atherosclerosis.
The ‘drug-target-pathway-disease’ network
A network diagram illustrating the relationships among okanin, the 17 key targets, the top 10 enriched signaling pathways, and wound healing was constructed via Cytoscape 3.9.1 software (Fig. 6).
Relationship network among okanin, key targets, Top10 KEGG pathways, and wound healing.
Molecular docking
Molecular docking techniques are employed to investigate the binding interactions between small molecules and their potential targets, thereby elucidating the therapeutic effects of these compounds on diseases. The binding energy, or affinity value, reflects the strength of the interaction between the ligand and the receptor target protein. A more negative binding energy indicates a stronger binding affinity between the active compound and the target protein. The molecular docking results for okanin with the top three key targets, ranked by binding energy, are presented in Fig. 7. The atomic coordinates of the targets were obtained from the RCSB-PDB. The binding energies between okanin and each key target are shown in Table 3. The findings reveal that okanin demonstrates a strong binding affinity for all key targets, with particularly notable affinities for AURKA and HDAC1, each exhibiting a binding energy of -8.8 kcal/mol. Molecular docking suggests that okanin may promote wound healing by interacting with these targeted proteins.
Molecular docking between okanin and key targets. a: okanin- AURKA; b: okanin-HDAC1; c: okanin-MMP3; d: okanin-PIK3R1.
Molecular dynamics simulations
RMSD and RMSF analysis
Molecular dynamics simulations were performed to validate the binding affinity of okanin with key target proteins identified through molecular docking. Specifically, the complexes with the highest binding energies to okanin—AURKA, HDAC1, MMP3, and PIK3R1 (with AURKA and HDAC1 sharing identical binding energies)—were selected for the simulations. The stability of these complexes was evaluated using RMSD values. As shown in Fig. 8, stabilization of the okanin-AURKA, okanin-HDAC1, okanin-MMP3, and okanin-PIK3R1 complexes occurred at 42 ns, 37 ns, 82 ns, and 17 ns, respectively.
RMSD of molecular dynamics simulation between okanin and key targets.a: okanin- AURKA; b: okanin-HDAC1; c: okanin-MMP3; d: okanin-PIK3R1.
Additionally, RMSF values, depicted in Fig. 9, were used to assess the conformational flexibility of each residue within the complexes. Simulation trajectories spanning 42–50 ns, 37–45 ns, 82–90 ns, and 17–25 ns were selected for MM/GBSA calculations. The resulting binding free energies were as follows: -24.27 ± 3.65 kcal/mol for the okanin-AURKA complex, -47.71 ± 1.60 kcal/mol for the okanin-HDAC1 complex, -30.52 ± 2.26 kcal/mol for the okanin-MMP3 complex, and − 44.46 ± 1.22 kcal/mol for the okanin-PIK3R1 complex. These results suggest that okanin can directly bind to these target proteins, exerting pharmacological effects.
RMSF of molecular dynamics simulation between okanin and key targets.a: okanin- AURKA; b: okanin-HDAC1; c: okanin-MMP3; d: okanin-PIK3R1.
For AURKA, the RMSD ranged from a minimum of 0.0005 nm at 0 ns to a maximum of 0.27 nm at 45.76 ns, with an average value of 0.22 nm. The RMSF ranged from 0.05 nm (residue 316) to 0.75 nm (residue 394), with an average of 0.13 nm.
For HDAC1, the RMSD ranged from 0.0005 nm at 0 ns to 0.373 nm at 28.61 ns, with an average of 0.29 nm. The RMSF ranged from 0.05 nm (residue 172) to 0.61 nm (residue 270), with an average of 0.13 nm.
For MMP3, the RMSD ranged from 0.0005 nm at 0 ns to 0.56 nm at 99.80 ns, with an average of 0.35 nm. The RMSF ranged from 0.07 nm (residue 118) to 1.195 nm (residue 173), with an average of 0.19 nm.
For PIK3R1, the RMSD ranged from 0.0005 nm at 0 ns to 0.39 nm at 14.54 ns, with an average of 0.31 nm. The RMSF ranged from 0.06 nm (residue 359, chain A) to 0.71 nm (residue 439, chain B), with an average of 0.17 nm.
SASA analysis
The solvent-accessible surface area (SASA) of the protein was monitored throughout the simulation, as shown in Fig. 10. Fluctuations in SASA reflect dynamic changes in the protein’s surface exposure to the solvent. For the okanin-AURKA complex, the average SASA value was 142.19 nm2, with a maximum of 150.18 nm2 at 54.29 ns and a minimum of 135.08 nm2 at 23.27 ns. For the okanin-HDAC1 complex, the average SASA value was 240.43 nm2, with a maximum of 252.90 nm2 at 73.37 ns and a minimum of 227.43 nm2 at 88.64 ns. For the okanin-MMP3 complex, the average SASA value was 102.99 nm2, with a maximum of 112.68 nm2 at 78.69 ns and a minimum of 93.62 nm² at 7.13 ns. For the okanin-PIK3R1 complex, the average SASA value was 594.79 nm2, with a maximum of 618.48 nm² at 0.26 ns and a minimum of 575.01 nm2 at 99.49 ns. These results suggest that okanin binding reduces the protein’s surface area exposure, particularly in the binding regions, indicating a stable interaction.
SASA of molecular dynamics simulation between okanin and key targets.a: okanin- AURKA; b: okanin-HDAC1; c: okanin-MMP3; d: okanin-PIK3R1.
Rg analysis
The Rg was calculated to assess the compactness of the protein-ligand complexes throughout the simulation, as shown in Fig. 11. Rg provides insight into the overall size and stability of the complex, with smaller values indicating a more compact structure. Fluctuations in Rg reflect dynamic changes in the tightness of the protein-ligand interaction over time.
Rg of molecular dynamics simulation between okanin and key targets. a: okanin- AURKA; b: okanin-HDAC1; c: okanin-MMP3; d: okanin-PIK3R1.
During the simulation, the overall Rg for the okanin-AURKA complex had an average value of 1.96 nm, fluctuating between a maximum of 2.00 nm at 35,790 ps and a minimum of 1.91 nm at 94,640 ps. For the okanin-HDAC1 complex, the average Rg was 2.44 nm, with a minimum of 2.40 nm at 0 ps, a maximum of 2.47 nm at 65,820 ps, and a final decrease to 2.43 nm. The okanin-MMP3 complex exhibited an average Rg of 1.61 nm, ranging from a minimum of 1.55 nm at 40,900 ps to a maximum of 1.74 nm at 99,510 ps. The okanin-PIK3R1 complex showed an average Rg of 3.53 nm, with a minimum value of 3.48 nm at 0 ps, reaching a maximum of 3.57 nm at 4,490 ps, and then decreasing to 3.51 nm.
Smaller Rg values indicate a more compact structure, typically associated with stable protein-ligand interactions. These results suggest that, despite fluctuations, the protein-ligand complexes remained relatively compact and stable throughout the simulation. The observed changes in Rg further suggest that the complexes did not undergo significant structural expansion or contraction, thereby supporting the stability of the protein-ligand binding.
H-bond analysis
Hydrogen bonds between okanin and the target proteins were monitored throughout the simulation to assess the stability of the protein-ligand interaction. The number of hydrogen bonds fluctuated, with the maximum number indicating periods of stable interaction. The formation and persistence of hydrogen bonds during the simulation suggest that okanin maintains a stable and significant interaction with the target proteins, contributing to the overall stability of the complex (Fig. 12).
H-bond of molecular dynamics simulation between okanin and key targets.a: okanin- AURKA; b: okanin-HDAC1; c: okanin-MMP3; d: okanin-PIK3R1.
For the okanin-AURKA complex, the maximum number of hydrogen bonds (6) was observed at 64.88 ns and 64.97 ns, with a minimum of 0 hydrogen bonds. The average number of hydrogen bonds throughout the simulation was 1.84.
For the okanin-HDAC1 complex, the maximum number of hydrogen bonds (9) occurred at 25.48 ns, with a minimum of 1 hydrogen bond. The average number of hydrogen bonds throughout the simulation was 5.32.
For the okanin-MMP3 complex, the maximum number of hydrogen bonds (7) was observed at 84.51 ns, with a minimum of 1 hydrogen bond. The average number of hydrogen bonds throughout the simulation was 3.54.
For the okanin-PIK3R1 complex, the hydrogen bond count consistently reached a maximum of 5 hydrogen bonds, with a minimum of 0 hydrogen bonds observed. The average number of hydrogen bonds throughout the simulation was 2.29, suggesting a relatively stable interaction.
The fluctuations in the number of hydrogen bonds indicate that the protein-ligand complexes maintained a dynamic yet stable interaction throughout the simulation.
qPCR validation of molecular simulation–predicted targets
To validate the potential targets predicted by molecular docking and molecular dynamics simulations, the mRNA expression levels of AURKA, HDAC1, PIK3R1, and MMP3 were quantified by qPCR using wound tissues collected on day 8 post-treatment.
AURKA expression was significantly downregulated in the okanin treatment group compared to both the control (P < 0.01) and vehicle (P < 0.01) groups, with no significant difference between control and vehicle.
HDAC1 expression was significantly elevated in the treatment group compared to both the vehicle (**P < 0.0001) and control (*P < 0.0001) groups, while no significant change was observed between control and vehicle.
For PIK3R1, expression was significantly decreased in the treatment group compared to both the control and vehicle groups (P < 0.01), with no difference between the latter two.
Similarly, MMP3 expression was significantly reduced in the treatment group compared to control and vehicle (P < 0.05), with no difference between vehicle and control (Fig. 13).
qPCR analysis of AURKA (A), HDAC1 (B), PIK3R1 (C), and MMP3 (D) mRNA expression in wound tissues from the control, vehicle, and okanin treatment groups. Data are presented as mean ± SD (n = 4). *P < 0.05, **P < 0.01, ****P < 0.0001.
These findings indicate that okanin modulates the in vivo expression of the predicted binding targets in a manner that aligns with the simulations, suggesting that these genes may be functionally involved in its wound-healing mechanism. Across targets, converged RMSD and a compact Rg with persistent hydrogen-bond occupancy indicate stable pocket engagement, and MM/GBSA estimates support favorable ΔG_ bind. Together, these trajectory features and qRT-PCR results suggest modulation of wound-healing pathways.
Discussion
Okanin is a bioactive compound found in Bidens pilosa L., a plant in the Asteraceae family, known for its various biological activities, including anti-inflammatory, antioxidant, anti-tumor, and antiviral effects. Over a thousand years ago, ancient Chinese practitioners used Bidens pilosa L. to treat a range of clinical diseases. As a result, both Bidens pilosa L. and its extract, okanin, show significant potential for development as clinical drugs for treating inflammatory diseases, oxidative stress-related disorders, tumors, and viral infections. Currently, okanin has emerged as a promising subject in pharmacological and TCM research29,30,31,32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47,48,49,50,51.
Wound healing has long been a critical focus for surgeons. Effective wound healing not only alleviates patient suffering but also reduces hospital stays and eases the financial burden on patients. Thus, developing safe and effective alternatives is a pressing concern and an urgent need within the medical community.
Network pharmacology, molecular docking, and molecular dynamics simulations are powerful scientific methodologies for drug discovery, screening, and preclinical validation. Molecular dynamics simulation uses computational methods based on classical mechanics principles to study the dynamic behavior and physical properties of biomolecular systems, such as proteins, nucleic acids, and lipids. It is widely applied in structural biology, drug design, and materials science52,53,54. Through network pharmacology screening, we identified 17 protein targets that may play a crucial role in okanin’s efficacy in wound healing. Among these, the top five, ranked by degree value, are HSP90AA1, ESR1, IGF1R, PTGS2, CDK1, and PIK3R1. An increased degree value suggests a higher likelihood that the protein target contributes to the pharmacological effects of okanin in promoting wound healing.
ADME/Toxicity and topical use
Okanin demonstrates promising multi-target mechanisms for wound healing; however, its therapeutic potential is influenced by pharmacokinetic and safety considerations. Computational profiling using SwissADME confirms okanin’s drug-likeness, with favorable compliance to Lipinski’s rules (MW = 288.25 g/mol, logP = 1.69). The compound exhibits moderate transdermal permeability, suggesting its potential for effective skin penetration when used topically. Despite a high Bioavailability Score (0.55), its oral bioavailability may be limited by first-pass metabolism, similar to other compounds like quercetin55.
Okanin’s non-substrate activity for P-gp and lack of blood-brain barrier (BBB) permeability indicate minimal systemic absorption when applied locally, reducing concerns about central nervous system effects. While ProTox-3.0 predicts moderate risks for nephrotoxicity (0.60) and respiratory toxicity (0.52), these risks are less relevant for topical use, as the compound is expected to remain localized at the site of application, thereby limiting systemic exposure.
To overcome bioavailability barriers, localized delivery strategies such as hydrogel-encapsulated nanocrystals or microneedle patches could enhance cutaneous retention and optimize okanin’s molecular weight and logP for efficient dermal penetration56,57. Future studies will aim to validate okanin’s modulation of IL-17 and AMPK in vivo, optimize nanoformulations for sustained release, and benchmark against clinical standards (e.g., EGF) to assess cost-effectiveness.
Binding stability and flexibility
In order to comprehensively evaluate the dynamic stability of the protein-ligand complexes, molecular dynamics simulations were analyzed based on RMSD, RMSF, SASA, Rg, and hydrogen bond number over a 100 ns simulation period.
The RMSD profiles indicated that the okanin-AURKA, okanin-HDAC1, and okanin-PIK3R1 complexes maintained relatively stable trajectories, with deviations fluctuating between 0.1 and 0.4 nm throughout the simulation, suggesting stable protein-ligand binding. Minor fluctuations were observed, particularly in the okanin-HDAC1 and okanin-PIK3R1 complexes, where RMSD values occasionally approached 0.37 nm and 0.39 nm, respectively. In contrast, the okanin-MMP3 complex exhibited a gradual increase in RMSD values after approximately 80 ns, reaching up to 0.56 nm, indicating greater structural flexibility.
RMSF analysis further revealed that the okanin-MMP3 complex displayed higher atomic fluctuations compared to the other complexes, particularly in specific regions of the protein. The average RMSF of the MMP3 complex was 0.19 nm, with maximum fluctuations reaching 1.195 nm at residue 173, suggesting increased local flexibility near the binding pocket. In contrast, the okanin-AURKA, okanin-HDAC1, and okanin-PIK3R1 complexes exhibited lower average RMSF values (ranging from 0.13 to 0.17 nm), indicating relatively more rigid structures.
The SASA and Rg values supported these observations. The okanin-AURKA, okanin-HDAC1, and okanin-PIK3R1 complexes maintained relatively constant SASA and Rg values throughout the simulation, indicating compact and stable structures. Specifically, their average SASA values were 142.19 nm², 240.43 nm², and 594.79 nm², respectively. In contrast, the okanin-MMP3 complex demonstrated a slight increase in SASA over time, with an average of 102.99 nm², suggesting a moderate expansion of the protein surface exposure. Similarly, the Rg of the MMP3 complex showed greater variability, ranging from 1.55 nm to 1.74 nm, indicative of a slight loosening of the overall structure.
Despite the greater fluctuations observed, hydrogen bond analysis demonstrated that intermittent but persistent interactions were maintained between okanin and MMP3 during the simulation. The average number of hydrogen bonds in the MMP3 complex was 3.54, with a maximum of 7 hydrogen bonds, indicating that okanin maintained notable interactions with the target even under dynamic conditions. In comparison, the okanin-HDAC1 complex exhibited the highest hydrogen bond stability, with an average of 5.32 hydrogen bonds.
It is important to recognize that biological macromolecules inherently exhibit dynamic fluctuations, and moderate flexibility at the binding interface may be essential for enabling conformational adaptability, functional modulation, and allosteric regulation58,59. Therefore, the dynamic behavior observed in the okanin-MMP3 complex may not necessarily denote instability but could instead represent an alternative binding mode characterized by flexibility and biological relevance.
Overall, while the okanin-MMP3 complex showed greater dynamic behavior compared to the okanin-AURKA, okanin-HDAC1, and okanin-PIK3R1 complexes, the ligand remained associated with MMP3 during the simulation, supporting the notion that flexible binding modes may coexist with more rigid ones in biologically functional states. In contrast, the strong and stable binding patterns observed in the okanin-AURKA, okanin-HDAC1, and okanin-PIK3R1 complexes validate the robustness of the simulation approach and highlight the potential of these targets for further development.
Mechanism interpretation
During wound healing, inflammation and biological processes such as autophagy and apoptosis frequently accompany the reparative process. The expression level of PTGS2 increases during inflammation and tissue damage; the associated signaling pathways are widely recognized as pro-inflammatory drivers and primary inducers of inflammation60. PIK3R1 modulates autophagic flux via the PIK3R1-AKT-mTOR pathway, while HSP90AA1 plays a key role in the regulation of autophagy, acting as a crucial regulatory factor61,62. It interacts with TFEB, leading to the nuclear translocation of TFEB and the initiation of autophagy63. In the early stages of inflammation, autophagy exerts an anti-infectious effect64, modulates the inflammatory response in an inverse manner, and prevents excessive inflammation that could lead to tissue damage65,66,67. CDK1 orchestrates the proliferation and migration of vascular smooth muscle cells, promoting wound angiogenesis, while autophagy in endothelial cells also contributes to wound healing. Autophagy in keratinocytes enhances their differentiation, proliferation, and migration, which facilitates wound re-epithelialization68,69. Concurrently, ESR1 acts as a central regulator of OLFM4-induced keratinocyte signaling, mobilizing both dermal and epidermal cell compartments to enhance wound healing70. During the proliferative phase, local hypoxia at the wound site can induce autophagy, which is implicated in anti-apoptotic mechanisms and oxidative stress resistance, thereby promoting cell survival71. IGF1R, a cell surface receptor, interacts with numerous intracellular second messengers and plays a pivotal role in cell proliferation, primarily mediating anti-apoptotic and pro-survival effects23,29. Suppression of IGF1R leads to dysfunction in HUVECs and impedes wound healing72. These findings suggest that HSP90AA1, ESR1, IGF1R, PTGS2, CDK1, and PIK3R1 may be involved in the wound healing process. Therefore, further investigation into the impact of okanin on these protein targets is warranted.
In this study, GO enrichment analysis revealed that okanin-associated therapeutic targets are significantly enriched in several BPs, including the positive regulation of protein import into the nucleus, insulin-like growth factor receptor signaling pathway, regulation of protein import into the nucleus, positive regulation of nucleocytoplasmic transport, and the G2/M transition of the mitotic cell cycle. Additionally, CCs were enriched in the growth cone, site of polarized growth, neuronal cell body, distal axon, and cell body. MFs identified include nuclear receptor activity, ligand-activated transcription factor activity, E-box binding, transcription co-regulator binding, and heat shock protein binding. Consequently, okanin’s role in promoting wound healing is likely mediated through the modulation of these BPs, CCs, and MFs.
The significantly enriched KEGG pathways are linked to a variety of human diseases, as well as to signaling pathways and their underlying pathophysiological mechanisms. Notably, the IL-17 signaling pathway and the AMPK signaling pathway rank among the top 10 enriched KEGG pathways, both playing crucial roles in diverse biological and disease-related processes.
Both the IL-17 and AMPK signaling pathways have the capacity to activate the mTOR signaling pathway73,74. The IL-17 signaling pathway significantly influences the pathophysiological processes of wound healing, including inflammatory responses, scar formation, cell proliferation, antibacterial activity, and antioxidant defense75,76,77. The AMPK signaling pathway is crucial for regulating cellular growth, autophagy, and metabolic processes. Activation of the AMPK pathway promotes endothelial cell migration to the wound site, facilitates tube formation, and enhances angiogenesis78. Furthermore, it downregulates mTORC1 activity within the mTOR signaling pathway, alleviates cellular hypoxia, stimulates keratinocyte movement and lateral migration79, modulates cell growth via mTORC180, induces autophagy81, and maintains mitochondrial homeostasis via Ulk182. Consequently, cell growth, autophagy, and metabolism, as regulated by the AMPK signaling pathway, are pivotal to the wound healing process. Network pharmacology analysis suggests that okanin’s impact on wound healing is, at least in part, mediated through the AMPK signaling pathway. This indicates that okanin’s mechanism of action primarily involves the promotion of cell proliferation, autophagy, and metabolism; however, the complexities of this network require further investigation.
Molecular docking and molecular dynamics simulations have elucidated the molecular mechanisms through which okanin contributes to wound healing. Molecular docking analysis revealed that okanin exhibits diverse binding affinities toward key protein targets, with the highest affinities observed for AURKA and HDAC1. Molecular dynamics simulation results showed that okanin maintains stability when complexed with AURKA and HDAC1, suggesting that these proteins play a crucial role in okanin-mediated wound healing. To further validate these in silico predictions, we examined the mRNA expression levels of AURKA, HDAC1, and related targets in vivo using qPCR. The results showed that okanin treatment significantly downregulated AURKA and upregulated HDAC1 expression in wound tissue, supporting their functional relevance.
AURKA is essential for the assembly and proper function of bipolar spindles during mitosis and meiosis, thereby controlling cell cycle progression and acting as a key protein kinase in regulating cell proliferation under conditions of early oxidative stress83,84. Additionally, AURKA can upregulate the expression of FOXO3a, promoting autophagy and modulating cellular stress responses. In diabetic wound models, AURKA protects cells under hyperglycemic conditions by significantly increasing the levels of wound epidermal growth factor, the anti-inflammatory cytokine IL-10, and reducing the levels of the pro-inflammatory cytokine IL-6, thereby facilitating skin regeneration in diabetic wounds3,85. Moreover, AURKA markedly enhances the formation of CD31-positive vascular-like structures, collagen levels, and VEGF factor levels in wounds, leading to improved angiogenesis, increased capillary density and hemoglobin content, enhanced wound blood supply, and accelerated healing86,87. HDAC1 plays a critical role in inflammation88, tumorigenesis regulation89, angiogenesis90, and cellular proliferation and differentiation91.
HDAC1 exhibits both pro-inflammatory and anti-inflammatory effects in response to distinct environmental stimuli and can modulate inflammation through the regulation of Fas-mediated T cell apoptosis92. Furthermore, HDAC1 regulates the expression of antioxidant enzymes by deacetylating histones and transcription factors, thereby controlling the redox balance in wounds and endothelial cells93. By deacetylating endothelial nitric oxide synthase 3, HDAC1 modulates endothelial cell function, influences nitric oxide production, inhibits platelet aggregation, maintains vascular tone, and promotes the proliferation of endothelial cells and neovascularization, thereby enhancing wound perfusion and facilitating wound healing94,95. It is clear that the signaling pathways mediated by AURKA and HDAC1 are crucial in wound healing; however, the complexity of this regulatory network exceeds current understanding and requires further investigation.
Comparison with prior studies
As a first-line clinical agent for wound repair, epidermal growth factor (EGF) exerts therapeutic effects primarily through direct activation of the EGFR-mediated MAPK/PI3K-AKT signaling axis, thereby stimulating keratinocyte proliferation and epithelial regeneration96,97. However, its clinical application is limited by pharmacokinetic constraints (half-life < 5 min in vivo), high production costs, and potential fibrotic complications98,99. In contrast, our computational findings reveal that okanin is predominantly associated with IL-17-mediated immune responses and AMPK-dependent metabolic pathways, suggesting dual therapeutic modalities that combine immunomodulation with energy homeostasis regulation. Molecular dynamics simulations demonstrate high-affinity binding to AURKA/HDAC1 (ΔG < -10 kcal/mol), indicating epigenetic modification and mitotic regulation capacities during tissue repair. Although empirical validation of okanin’s pathway interactions is needed, the observed signaling modulation aligns with established anti-inflammatory therapies that enhance macrophage M2 polarization—a mechanism distinct from quercetin’s (clinically validated in diabetic wounds) PI3K-AKT-centric action, which involves TNF-α suppression, MMP9 downregulation, and angiogenesis promotion100.
Clinically, EGF demonstrates superior therapeutic efficacy in acute wound healing but shows limited effectiveness in chronic wound microenvironments. In contrast, okanin exhibits computationally predicted cell cycle-regulatory capacity and shares a multitarget anti-inflammatory profile with QCT (though without structural analogy), suggesting complementary applications for metabolic disorders and diabetic ulcers, with potential for tri-agent synergistic therapy. The therapeutic integration of EGF-mediated epithelial activation with okanin-driven metabolic modulation could address the multifactorial complexity of wound healing, while QCT-okanin coadministration may establish interconnectivity within the PI3K-AMPK pathway. Future investigations should prioritize experimental validation of okanin’s modulation of the IL-17 pathway and the development of advanced formulations to overcome pharmacokinetic limitations, collectively emphasizing the need for pathophysiology-specific therapeutic strategies in precision wound management.
This research employs an integrated computational framework combining network pharmacology, molecular docking, and molecular dynamics simulations to systematically investigate okanin’s wound-healing mechanisms. The multi-stage strategy utilizes molecular dynamics simulations as a preclinical validation platform, offering cost-efficiency, rapid analysis, and molecular-level insights while eliminating the need for animal experimentation—aligning with ethical principles aimed at reducing animal use101. Despite some predicted systemic toxicity risks, okanin’s favorable antibacterial activity, suitability for topical application, and low predicted systemic toxicity position it as a promising candidate for development as an adjunctive therapy for wound healing. Although this computational approach provides mechanistic hypotheses and theoretical foundations for subsequent studies, the current findings are limited by the inherent constraints of in silico analyses. Acknowledging these limitations, future research will incorporate systematic experimental validation through established in vitro and in vivo models to confirm okanin’s biological activity and accurately characterize its therapeutic mechanisms. This sequential methodology ensures both methodological rigor and alignment with evolving standards in translational research.
Limitations
This study lacks protein-level validation (e.g., Western blotting and immunohistochemistry), and the in vivo design currently does not include a clinical positive control, limiting direct benchmarking against standard care. Statements regarding the safety of okanin rely on in silico/network-toxicology screening; no experimental dermal toxicology was performed. Our MD analyses used a classical MM/GBSA framework and finite trajectory lengths, which are appropriate for comparative ranking but do not yield absolute binding free energies. Finally, we focused on a single compound (okanin), and potential multi-component synergy typical of herbal formulations was not assessed.
Future directions
We plan to validate key targets at the protein level and map their tissue localization; include a clinical positive control (e.g., silver sulfadiazine or allantoin) for benchmarking; implement a stepwise dermal safety program guided by current network-toxicology signals; optimize topical formulations, with particular emphasis on hydrogel-based delivery, and evaluate long-term functional outcomes; and strengthen the computational arm with replicated/extended MD simulations and, where appropriate, alchemical free-energy calculations.
Conclusion
This study reveals the role of okanin in wound healing through multiple targets and signaling pathways, utilizing methods such as network pharmacology, molecular docking, molecular dynamics simulations and experimental validation. Its mechanism of action involves antioxidant responses, anti-inflammatory activities, and the promotion of cell proliferation. AURKA and HDAC1 exhibit the highest binding affinity for okanin and are likely central players in its pharmacological pathways. The IL-17 and AMPK signaling pathways are critical to the wound healing process, serving as key mediators of okanin’s effects. Additionally, downstream signaling pathways, such as the mTOR signaling pathway, may also be implicated.
Data availability
The datasets used and/or analysed during the current study available from the corresponding author on reasonable request.
Abbreviations
- AHR:
-
Aryl hydrocarbon receptor
- AMPK:
-
AMP-activated protein kinase
- APP:
-
Amyloid precursor protein
- AURKA:
-
Aurora kinase A
- BP:
-
Biological process
- BV-2:
-
Microglial cell line
- CC:
-
Cellular component
- CTD:
-
Comparative toxicogenomics database
- CXCR4:
-
C-X-C Motif Chemokine Receptor 4
- ESR1:
-
Estrogen receptor 1
- GAFF:
-
Generalized amber force field
- GO:
-
Gene Ontology
- HDAC1:
-
Histone deacetylase 1
- HSP90AA1:
-
Heat Shock Protein 90 kDa Alpha Family Class A Member 1
- HSPA8:
-
Heat Shock Protein Family A (Hsp70) Member 8
- IGF1R:
-
Insulin-like Growth Factor 1 Receptor
- IGFBP3:
-
Insulin-like Growth Factor Binding Protein 3
- IL:
-
Interleukin
- iNOS:
-
Inducible nitric oxide synthase
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- LPS:
-
Lipopolysaccharide
- MF:
-
Molecular function
- MM/GBSA:
-
Molecular Mechanics/Generalized Born Surface Area
- MMP3:
-
Matrix metalloproteinase 3
- mTOR:
-
Mechanistic target of rapamycin
- PDPK1:
-
3-Phosphoinositide-Dependent Protein Kinase-1
- PIK3R1:
-
Phosphoinositol-3-kinase Regulatory Subunit 1
- PPARG:
-
Peroxisome Proliferator-Activated Receptor Gamma
- PPI:
-
Protein-protein interaction
- PTGS2:
-
Prostaglandin-endoperoxide synthase 2
- RCSB-PDB:
-
Research Collaboratory for Structural Bioinformatics Protein Data Bank
- RMSD:
-
Root-mean-square deviation
- RMSF:
-
Root-mean-square fluctuation
- SNCA:
-
Alpha-synuclein
- STRING:
-
Search Tool for the Retrieval of Interacting Genes/Proteins
- TCM:
-
Traditional Chinese Medicine
- TNF-α:
-
Tumor necrosis factor-alpha
- AHR:
-
Aryl hydrocarbon receptor
References
Nussbaum, S. R. et al. An economic evaluation of the impact, cost, and medicare policy implications of chronic nonhealing wounds. Value Health J. Int. Soc. Pharmacoecon. Outcomes Res. 21, 27–32 (2018).
Varshney, N. et al. Integrating insights into cancer, inflammation, and infectious diseases. Gut Microbes Rep. 1, 2419069 (2024).
Yin, Y. et al. AURKA enhances autophagy of adipose derived stem cells to promote diabetic wound repair via targeting FOXO3a. J. Invest. Dermatol. 140, 1639–1649e4 (2020).
Cabanel, M., da Costa, T. P., EL-Cheikh, M. C. & Carneiro, K. The epigenome as a putative target for skin repair: the HDAC inhibitor trichostatin A modulates myeloid progenitor plasticity and behavior and improves wound healing. J. Transl Med. 17, 247 (2019).
Kida, M. et al. Inhibition of the CoREST repressor complex promotes wound Re-Epithelialization through the regulation of keratinocyte migration. J. Invest. Dermatol. 144, 378–386e2 (2024).
Zheng, L. et al. Matrix Metalloproteinase-3 accelerates wound healing following dental pulp injury. Am. J. Pathol. 175, 1905–1914 (2009).
Jiang, B. H. & Liu, L. Z. PI3K/PTEN signaling in angiogenesis and tumorigenesis. Adv. Cancer Res. 102, 19–65 (2009).
Luyendyk, J. P. et al. Genetic analysis of the role of the PI3K-Akt pathway in Lipopolysaccharide-Induced cytokine and tissue factor gene expression in Monocytes/Macrophages. J. Immunol. Baltim. Md. 1950. 180, 4218–4226 (2008).
Misiura, M., Baszanowska, W., Ościłowska, I., Pałka, J. & Miltyk, W. Prolidase stimulates proliferation and migration through activation of the PI3K/Akt/mTOR signaling pathway in human keratinocytes. Int. J. Mol. Sci. 21, 9243 (2020).
Bowers, S. & Franco, E. Chronic wounds: evaluation and management. Am. Fam Physician. 101, 159–166 (2020).
Armstrong, D. G., Boulton, A. J. M. & Bus, S. A. Diabetic foot ulcers and their recurrence. N Engl. J. Med. 376, 2367–2375 (2017).
Chang, M. & Nguyen, T. T. Strategy for treatment of infected diabetic foot ulcers. Acc. Chem. Res. 54, 1080–1093 (2021).
Fogerty, M., Guy, J., Barbul, A., Nanney, L. B. & Abumrad, N. N. African Americans show increased risk for pressure ulcers: a retrospective analysis of acute care hospitals in America. Wound Repair. Regen Publ Wound Heal Soc. Eur. Tissue Repair. Soc. 17, 678–684 (2009).
Manley, M. T. Incidence, contributory factors and costs of pressure sores. Afr. Med. J. Suid-Afrik Tydskr Geneeskd. 53, 217–222 (1978).
Shannon, M. L. & Skorga, P. Pressure ulcer prevalence in two general hospitals. Decubitus 2, 38–43 (1989).
Gould, L. J. et al. WHS guidelines for the treatment of pressure ulcers-2023 update. Wound Repair. Regen Publ Wound Heal Soc. Eur. Tissue Repair. Soc. 32, 6–33 (2024).
Velnar, T., Bailey, T. & Smrkolj, V. The wound healing process: an overview of the cellular and molecular mechanisms. J. Int. Med. Res. 37, 1528–1542 (2009).
Powers, J. G., Higham, C., Broussard, K. & Phillips, T. J. Wound healing and treating wounds: chronic wound care and management. J. Am. Acad. Dermatol. 74, 607–625 (2016).
Boulton, A. J. M., Vileikyte, L., Ragnarson-Tennvall, G. & Apelqvist, J. The global burden of diabetic foot disease. Lancet Lond. Engl. 366, 1719–1724 (2005).
Legrand, J. M. D. & Martino, M. M. Growth factor and cytokine delivery systems for wound healing. Cold Spring Harb Perspect. Biol. 14, a041234 (2022).
Rodríguez-Mesa, X. M. et al. Immunomodulatory properties of natural extracts and compounds derived from bidens Pilosa L. Literature Rev. Pharmaceutics. 15, 1491 (2023).
Kabanda, M. M., Gbashi, S. & Madala, N. E. Proportional coexistence of Okanin chalcone glycoside and Okanin Flavanone glycoside in bidens Pilosa leaves and theoretical investigation on the antioxidant properties of their aglycones. Free Radic Res. 55, 53–70 (2021).
Hou, Y. et al. Okanin, effective constituent of the flower tea coreopsis tinctoria, attenuates LPS-induced microglial activation through Inhibition of the TLR4/NF-κB signaling pathways. Sci. Rep. 7, 45705 (2017).
Liu, Y., Xiong, B., Qiu, X., Hao, H. & Sha, A. Study on the antithrombotic effect and physiological mechanism of Okanin. Biomed. Pharmacother Biomed. Pharmacother. 153, 113358 (2022).
Abdurehman, D. et al. Optimization of Preparation method of hepatoprotective active components from coreopsis tinctoria Nutt. And its action mechanism in vivo. Biomed. Pharmacother Biomed. Pharmacother. 167, 115590 (2023).
Guo, L., Zhou, Y. & Ma, R. Exploring the anti-gastric cancer mechanism of action of bidentis bipinnatae herba based on network pharmacology, molecular docking, and cellular experimental validation. Naunyn-Schmiedebergs Arch. Pharmacol. 397, 8681–8690 (2024).
Mi, Y. et al. Okanin from coreopsis tinctoria Nutt. Alleviates cognitive impairment in bilateral common carotid artery occlusion mice by regulating the miR-7/NLRP3 axis in microglia. Food Funct. 14, 369–387 (2023).
Wang, W. et al. New phenolic compounds from coreopsis tinctoria Nutt. And their antioxidant And angiotensin i-converting enzyme inhibitory activities. J. Agric. Food Chem. 63, 200–207 (2015).
Peng, A., Lin, L., Zhao, M. & Sun, B. Classification of edible chrysanthemums based on phenolic profiles and mechanisms underlying the protective effects of characteristic phenolics on oxidatively damaged erythrocyte. Food Res. Int. Ott. Ont. 123, 64–74 (2019).
Nogales, C. et al. Network pharmacology: curing causal mechanisms instead of treating symptoms. Trends Pharmacol. Sci. 43, 136–150 (2022).
Li, X. et al. Network Pharmacology approaches for research of traditional Chinese medicines. Chin. J. Nat. Med. 21, 323–332 (2023).
Bernetti, M., Cavalli, A. & Mollica, L. Protein-ligand (un)binding kinetics as a new paradigm for drug discovery at the crossroad between experiments and modelling. MedChemComm 8, 534–550 (2017).
Hertig, S., Latorraca, N. R. & Dror, R. O. Revealing Atomic-Level mechanisms of protein allostery with molecular dynamics simulations. PLoS Comput. Biol. 12, e1004746 (2016).
Burley, S. K. et al. RCSB protein data bank: powerful new tools for exploring 3D structures of biological macromolecules for basic and applied research and education in fundamental biology, biomedicine, biotechnology, bioengineering and energy sciences. Nucleic Acids Res. 49, D437–D451 (2021).
Davis, A. P. et al. Comparative toxicogenomics database (CTD): update 2023. Nucleic Acids Res. 51, D1257–D1262 (2023).
Daina, A., Michielin, O. & Zoete, V. SwissTargetPrediction: updated data and new features for efficient prediction of protein targets of small molecules. Nucleic Acids Res. 47, W357–W364 (2019).
Stelzer, G. et al. The genecards suite: from gene data mining to disease genome sequence analyses. Curr. Protoc. Bioinform. 54, 1301–13033 (2016).
Wishart, D. S. et al. DrugBank: a comprehensive resource for in Silico drug discovery and exploration. Nucleic Acids Res. 34, D668–672 (2006).
Szklarczyk, D. et al. The STRING database in 2023: protein-protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res. 51, D638–D646 (2023).
Szklarczyk, D. et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 47, D607–D613 (2019).
SwissADME. A free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci. Rep. (2025). https://www.nature.com/articles/srep42717.
Banerjee, P., Kemmler, E., Dunkel, M. & Preissner, R. ProTox 3.0: a webserver for the prediction of toxicity of chemicals. Nucleic Acids Res. 52, W513–W520 (2024).
Luger, K., Mäder, A. W., Richmond, R. K., Sargent, D. F. & Richmond, T. J. Crystal structure of the nucleosome core particle at 2.8 A resolution. Nature 389, 251–260 (1997).
Van Der Spoel, D. et al. GROMACS: fast, flexible, and free. J. Comput. Chem. 26, 1701–1718 (2005).
Sun, S. et al. Anti-photoaging effect and the mechanism of coreopsis tinctoria Okanin against UVB-induced skin damage in mice. Int. Immunopharmacol. 139, 112657 (2024).
Barabási, A. L., Gulbahce, N. & Loscalzo, J. Network medicine: a network-based approach to human disease. Nat. Rev. Genet. 12, 56–68 (2011).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. Publ Protein Soc. 28, 1947–1951 (2019).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Kanehisa, M., Furumichi, M., Sato, Y., Kawashima, M. & Ishiguro-Watanabe M. KEGG for taxonomy-based analysis of pathways and genomes. Nucleic Acids Res. 51, D587–D592 (2023).
Mu, Y., Zeng, H. & Chen, W. Okanin in coreopsis tinctoria nutt is a major quorum-sensing inhibitor against chromobacterium violaceum. J. Ethnopharmacol. 260, 113017 (2020).
Chia, W. T. et al. Okanin inhibits cell growth and induces apoptosis and pyroptosis in oral cancer. Cancers 16, 3195 (2024).
Hu, X. et al. Molecular dynamics simulation of the interaction of food proteins with small molecules. Food Chem. 405, 134824 (2023).
Wu, J. et al. Application of molecular dynamics simulation for exploring the roles of plant biomolecules in promoting environmental health. Sci. Total Env. 869, 161871 (2023).
Hildebrand, P. W., Rose, A. S. & Tiemann, J. K. S. Bringing molecular dynamics simulation data into view. Trends Biochem. Sci. 44, 902–913 (2019).
Liga, S., Paul, C. & Péter, F. Flavonoids overview of biosynthesis, biological Activity, and current extraction techniques. Plants Basel Switz. 12, 2732 (2023).
Zhang, W. et al. Double-Layered microneedle patch integrated with multifunctional nanoparticles and live bacteria for Long-Term treatment of atopic dermatitis. Small Weinh Bergstr Ger. 21, e2409121 (2025).
Zhang, L. et al. Recombinant collagen microneedles for transdermal delivery of antibacterial copper-DNA nanoparticles to treat skin and soft tissue infections. J. Controlled Release. 379, 191–201 (2025).
Henzler-Wildman, K. & Kern, D. Dynamic personalities of proteins. Nature 450, 964–972 (2007).
Boehr, D. D., Nussinov, R. & Wright, P. E. The role of dynamic conformational ensembles in biomolecular recognition. Nat. Chem. Biol. 5, 789–796 (2009).
Martín-Vázquez, E., Cobo-Vuilleumier, N., López-Noriega, L., Lorenzo, P. I. & Gauthier, B. R. The PTGS2/COX2-PGE2 signaling cascade in inflammation: pro or anti? A case study with type 1 diabetes mellitus. Int. J. Biol. Sci. 19, 4157–4165 (2023).
Ji, X. et al. GSTP1-mediated S-glutathionylation of Pik3r1 is a redox hub that inhibits osteoclastogenesis through regulating autophagic flux. Redox Biol. 61, 102635 (2023).
Qiang, L., Yang, S., Cui, Y. H. & He, Y. Y. Keratinocyte autophagy enables the activation of keratinocytes and fibroblastsand facilitates wound healing. Autophagy 17, 2128–2143 (2021).
Yang, S. et al. Regulation of TFEB nuclear localization by HSP90AA1 promotes autophagy and longevity. Autophagy 19, 822–838 (2023).
Skendros, P., Mitroulis, I. & Ritis, K. Autophagy in neutrophils: from granulopoiesis to neutrophil extracellular traps. Front. Cell. Dev. Biol. 6, 109 (2018).
Liu, K. et al. Impaired macrophage autophagy increases the immune response in obese mice by promoting Proinflammatory macrophage polarization. Autophagy 11, 271–284 (2015).
Kawano, A. et al. Docosahexaenoic acid enhances M2 macrophage polarization via the p38 signaling pathway and autophagy. J. Cell. Biochem. 120, 12604–12617 (2019).
Das, L. M., Binko, A. M., Traylor, Z. P., Peng, H. & Lu, K. Q. Vitamin D improves sunburns by increasing autophagy in M2 macrophages. Autophagy 15, 813–826 (2019).
Zhang, J. et al. Author correction: involvement of autophagy in hypoxia-BNIP3 signaling to promote epidermal keratinocyte migration. Cell. Death Dis. 10, 295 (2019).
Wang, J. et al. Autophagy plays a positive role in induction of epidermal proliferation. FASEB J. Publ Fed. Am. Soc. Exp. Biol. 34, 10657–10667 (2020).
Klaas, M. et al. Olfactomedin-4 improves cutaneous wound healing by promoting skin cell proliferation and migration through POU5F1/OCT4 and ESR1 signalling cascades. Cell. Mol. Life Sci. CMLS. 79, 157 (2022).
Jeong, I. H., Bae, W. Y., Choi, J. S. & Jeong, J. W. Ischemia induces autophagy of endothelial cells and stimulates angiogenic effects in a hindlimb ischemia mouse model. Cell. Death Dis. 11, 624 (2020).
Wang, J. et al. Diabetic macrophage small extracellular vesicles-associated miR-503/IGF1R axis regulates endothelial cell function and affects wound healing. Front. Immunol. 14, 1104890 (2023).
Lee, S. Y. et al. IL-17 induces autophagy dysfunction to promote inflammatory cell death and fibrosis in keloid fibroblasts via the STAT3 and HIF-1α dependent signaling pathways. Front. Immunol. 13, 888719 (2022).
Kim, J., Kundu, M., Viollet, B. & Guan, K. L. AMPK and mTOR regulate autophagy through direct phosphorylation of Ulk1. Nat. Cell. Biol. 13, 132–141 (2011).
Shan, K. et al. IL-17-triggered downregulation of miR-497 results in high HIF-1α expression and consequent IL-1β and IL-6 production by astrocytes in EAE mice. Cell. Mol. Immunol. 14, 909–923 (2017).
Gabbai-Armelin, P. R. et al. A systematic review and meta-analysis of the effect of thymol as an anti-inflammatory and wound healing agent: a review of thymol effect on inflammation and wound healing: a review of thymol effect on inflammation and wound healing. Phytother Res. PTR. 36, 3415–3443 (2022).
Khaire, K. et al. Site-specific variation in the activity of COX-2 alters the pattern of wound healing in the tail and limb of Northern house gecko by differentially regulating the expression of local inflammatory mediators. Zool. Jena. 148, 125947 (2021).
Han, B. et al. Exosomal EPHA2 derived from highly metastatic breast cancer cells promotes angiogenesis by activating the AMPK signaling pathway through Ephrin A1-EPHA2 forward signaling. Theranostics 12, 4127–4146 (2022).
Yan, T. et al. Hypoxia regulates mTORC1-Mediated keratinocyte motility and migration via the AMPK pathway. PLoS One. 12, e0169155 (2017).
Kalender, A. et al. Metformin, independent of AMPK, inhibits mTORC1 in a rag GTPase-dependent manner. Cell. Metab. 11, 390–401 (2010).
Hardie, D. G. AMPK and autophagy get connected. EMBO J. 30, 634–635 (2011).
Nakada, D., Saunders, T. L. & Morrison, S. J. Lkb1 regulates cell cycle and energy metabolism in Haematopoietic stem cells. Nat 468, 653–658 (2010).
Wang, G. F. et al. Oxidative stress induces mitotic arrest by inhibiting Aurora A-involved mitotic spindle formation. Free Radic Biol. Med. 103, 177–187 (2017).
Damodaran, A. P., Vaufrey, L., Gavard, O. & Prigent, C. Aurora A kinase is a priority pharmaceutical target for the treatment of cancers. Trends Pharmacol. Sci. 38, 687–700 (2017).
Tang, H. et al. AURKA facilitates the psoriasis-related inflammation by impeding autophagy-mediated AIM2 inflammasome suppression. Immunol. Lett. 240, 98–105 (2021).
Meng, J. et al. Upregulation of Aurora kinase A promotes vascular smooth muscle cell proliferation and migration by activating the GSK-3β/β-catenin pathway in aortic-dissecting aneurysms. Life Sci. 262, 118491 (2020).
Bai, T. et al. The promotion action of AURKA on post-ischemic angiogenesis in diabetes-related limb ischemia. Mol. Med. 29, 39 (2023).
Zhang, X., Lu, J., Zhang, Q., Luo, Q. & Liu, B. CircRNA RSF1 regulated ox-LDL induced vascular endothelial cells proliferation, apoptosis and inflammation through modulating miR-135b-5p/HDAC1 axis in atherosclerosis. Biol. Res. 54, 11 (2021).
Winter, M. et al. Divergent roles of HDAC1 and HDAC2 in the regulation of epidermal development and tumorigenesis. EMBO J. 32, 3176–3191 (2013).
Ding, X. et al. Suppression of the SAP18/HDAC1 complex by targeting TRIM56 and Nanog is essential for oncogenic viral FLICE-inhibitory protein-induced acetylation of p65/RelA, NF-κB activation, and promotion of cell invasion and angiogenesis. Cell. Death Differ. 26, 1970–1986 (2019).
Zhu, X. et al. HDAC1/2 control proliferation and survival in adult epidermis and Pre–Basal cell carcinoma through p16 and p53. J. Invest. Dermatol. 142, 77–87e10 (2022).
Zimmerman, M. A. et al. Butyrate suppresses colonic inflammation through HDAC1-dependent Fas upregulation and Fas-mediated apoptosis of T cells. Am. J. Physiol. Gastrointest. Liver Physiol. 302, G1405–1415 (2012).
Zelko, I. N. & Folz, R. J. Regulation of oxidative stress in pulmonary artery Endothelium. Modulation of extracellular superoxide dismutase and NOX4 expression using histone deacetylase class I inhibitors. Am. J. Respir Cell. Mol. Biol. 53, 513–524 (2015).
Hyndman, K. A., Ho, D. H., Sega, M. F. & Pollock, J. S. Histone deacetylase 1 reduces NO production in endothelial cells via lysine deacetylation of NO synthase 3. Am. J. Physiol. Heart Circ. Physiol. 307, H803–809 (2014).
Gan, Y. et al. Role of histone deacetylation in cell-specific expression of endothelial nitric-oxide synthase. J. Biol. Chem. 280, 16467–16475 (2005).
Hardwicke, J., Schmaljohann, D., Boyce, D. & Thomas, D. Epidermal growth factor therapy and wound healing–past, present and future perspectives. Surg. J. R Coll. Surg. Edinb. Irel. 6, 172–177 (2008).
An, S. J. et al. Regulation of EGF-stimulated activation of the PI-3K/AKT pathway by exocyst-mediated exocytosis. Proc. Natl. Acad. Sci. U. S. A. 119, e2208947119 (2022).
Romero Prada, M. et al. Cost-effectiveness analysis of the human Recombinant epidermal growth factor in the management of patients with diabetic foot ulcers. Diabet. Foot Ankle. 9, 1480249 (2018).
Coentro, J. Q., Pugliese, E., Hanley, G., Raghunath, M. & Zeugolis, D. I. Current and upcoming therapies to modulate skin scarring and fibrosis. Adv. Drug Deliv. Rev. 146, 37–59 (2019).
Zhang, Z., Wang, L., Li, X., Miao, Y. & Li, D. Integrating network Pharmacology, molecular Docking and experimental validation to explore the Pharmacological mechanisms of Quercetin against diabetic wound. Int. J. Med. Sci. 21, 2837–2850 (2024).
Wu, X., Xu, L. Y., Li, E. M. & Dong, G. Application of molecular dynamics simulation in biomedicine. Chem. Biol. Drug Des. 99, 789–800 (2022).
Funding
The study was supported by the Clinical Research Center for Traditional Chinese Medicineon Anorectal and Perianal Wound Repair of Fujian Province (2022Y2011).
Author information
Authors and Affiliations
Contributions
ZCZ and YL conceptualized and designed the study. ZCZ drafted and wrote the manuscript. ZCZ and YL performed the experiments. JML and YC performed data curation. ZCW, YTC, and ZXY participated in part of the data analysis. JWZ, XXW, and YMX handled the visualization. JH and WHZ corrected the manuscript. RS supervised the experimentators. JW supervised the study and approved the final version. All data were generated in-house, and no paper mill was used. All authors agree to be accountable for all aspects of work ensuring integrity and accuracy.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
All animal experiments were conducted in compliance with the ARRIVE guidelines (https://arriveguidelines.org) and the editorial policies of Scientific Reports. Experimental procedures were approved by the IACUC of Ampry Biosciences Co., Ltd. on 20 May 2025 (IACUC-FJABR-2025052001). Mice were sourced from Sipeifu (Beijing) Biotechnology Co., Ltd. (License No. SCXK [Jing] 2019-0010). All procedures adhered to national regulations on the care and use of laboratory animals. Animals were anesthetized for surgery and humanely euthanized at study endpoints with a terminal dose of sodium pentobarbital to minimize pain and distress.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhao, C., Li, Y., Lin, J. et al. Mechanisms of Okanin against wound healing based on network pharmacology, molecular docking and molecular dynamics simulation. Sci Rep 15, 43034 (2025). https://doi.org/10.1038/s41598-025-25557-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-25557-5