Abstract
Deep learning models leveraging electronic health records (EHR) for opportunistic screening of type 2 diabetes (T2D) can improve current practices by identifying individuals who may need further glycemic testing. Accurate onset prediction and subtyping are crucial for targeted interventions, but existing methods treat the tasks separately, thus limiting clinical utility. In this paper, we introduce a novel deep metric learning (DML) model that unifies both tasks by learning a latent space based on sample similarity. In onset prediction, the DML model predicts the onset of T2D 7 years later with an AUC of 0.754, outperforming logistic regression (AUC 0.706), clinical risk factors (AUC 0.693), and glycemic measures (AUC 0.632). For subtyping, we identify three subtypes with varying prevalences of obesity-related, cardiovascular, and mental health conditions. Additionally, the subtype with fewer comorbidities shows earlier metformin initiation and a greater reduction in HbA1c. We validated these findings using data from 300 U.S. hospitals in the All of Us program (T2D, n = 7567) and the Massachusetts General Brigham Biobank (T2D, n = 3298), demonstrating the transferability of our model and subtypes across cohorts.
Similar content being viewed by others
Introduction
Type 2 diabetes (T2D) is a complex, chronic disease affecting approximately 11% of the U.S. population as of 20211, with global cases projected to rise from 380 million in 2013 to 590 million by 20352. Despite its growing prevalence, screening relies on simple criteria like age and obesity3 while diagnostic tests depend on HbA1c and glucose levels4,5, failing to capture disease complexity. T2D manifests through distinct pathomechanisms, such as insulin resistance, beta-cell dysfunction, genetic predisposition, and environmental factors6,7,8. The current one-size-fits-all approach to prevention and management is inadequate, underscoring the need for subtyping to enable targeted interventions and precision medicine.
Many prior T2D subtyping efforts rely on features that are not commonly collected in routine practice9. A seminal study identified five subtypes with distinct disease trajectories, but replication requires specialized biomarkers such as beta-cell function and insulin resistance10. Genetic subtyping has also been explored11, though it excludes environmental influences and lacks clinical feasibility in the general population. Wagner et al. identified six prediabetic clusters with different risks of T2D complications and mortality, but require the collection of glycemic measures such as oral glucose tolerance tests and anthropometric traits, limiting applicability to clinical practice12.
To overcome these limitations, recent research uses readily available electronic health record (EHR) data to predict and subtype T2D. Anderson et al. demonstrated that machine learning models trained on comprehensive EHR data outperform those using limited risk factors13. Approaches using convolutional networks and Gaussian processes identify subtypes with distinct comorbidities and severity profiles14,15. A clustering method has also revealed subtypes, including a younger, non-obese, economically disadvantaged group16. These approaches, however, address either prediction or subtyping separately, lacking a unified framework for both tasks.
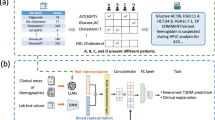
We propose a unified deep learning framework to predict T2D onset and subtypes using common EHR features. Leveraging patient similarity in deep metric learning (DML), the model identifies subtypes with distinct comorbidities, medication responses, and polygenic risk scores (PRS). Specifically, subtypes differ in comorbidity rates, such as obesity, depression, and hypertension. The Green subtype responds better to initial metformin treatment than the Red subtype, highlighting the potential for tailored interventions. Our developed model can be incorporated into existing EHR systems to simultaneously screen and subtype individuals, facilitating further diagnostics and personalized care while minimizing additional clinical workload (Fig. 1).
Our deep metric learning (DML) model uses routine EHR data to predict and subtype future T2D. Our model serves as a pre-screening tool, analyzing routine EHR data to identify individuals at risk for T2D without increasing clinical burden. The model also predicts future T2D subtypes, allowing personalized care and precision medicine.
Results
Cohort selection, data preprocessing, and model training
We utilized two datasets: the All of Us (AoU) dataset, which includes longitudinal electronic health records (EHR) and genetic data from a diverse cohort of over 400,000 participants across more than 340 centers in the United States17, and the Massachusetts General Brigham (MGB) Biobank, which contains EHR and genetic data from a large, integrated healthcare system in Massachusetts, encompassing over 1.5 million unique patients per year18. The MGB Biobank data were retrieved on 10/12/2022, while the AoU dataset contains data up to 01/01/2022 (Controlled Tier v6).
To identify people with type 2 diabetes (T2D), we applied the eMERGE algorithm19,20 to the AoU dataset (n = 7567) and the PheCap algorithm21 to the MGB dataset (n = 3298). The eMERGE algorithm defines T2D cases using condition codes, diabetes medication codes, and abnormal HbA1c values, excluding type 1 diabetes (T1D) codes. The PheCap algorithm, a machine learning-based method that utilizes both structured EHR data and unstructured clinical notes, internally developed and validated at MGH for T2D cohort selection. Additional preprocessing details are provided in Supplementary Note 2.
For robust model training, we selected high-risk controls who had not developed T2D, ensuring the model learns to distinguish subtle differences between cases and controls with similar risk factors. We define a population-matched control cohort (PopControl) by pairing each T2D case with a control matched on age, sex, and healthcare utilization. For test datasets during evaluation, we use the general population without T2D (GenControl) at natural disease prevalence.
In both the AoU and MGB datasets, we construct input features from EHR data, including conditions (n = 71), medications (n = 89), physical measurements (n = 6), laboratory values (n = 21), and demographic variables like age and sex. For each feature, we compute the mean, minimum, and maximum across three time windows: 6 months, 2 years, and the entire EHR history before the censor date. To avoid data leakage, we set the censor date at least 2 years before diagnosis, up to a maximum of 10 years. Missing values are imputed using the population mean.
For the onset prediction task, we train several models—Deep Metric Learning (DML), logistic regression (LR), deep learning models (SCARF22, TabTransformer23, CVAE24, ConvAE14, and dimensionality reduction methods (PCA25, UMAP26—using a consistent set of 698 preprocessed features. With LR, we also replicate established clinical models: a risk-factors model from Wilson et al.27 (Risk-Factors) and a glycemic-based model representing current diagnostic standards4,28 (Glycemic). Due to data preprocessing constraints in the MGB system, only the DML and LR models were applied to the MGB dataset. Model performance is evaluated using the area under the receiver operating characteristic curve (AUROC).
The core novelty of our method is the high-capacity DML encoder that learns a latent representation useful for both onset prediction and subtyping. For onset prediction, we apply a simple linear classifier on the learned representations. In our case, we use logistic regression as the classifier to allow for direct comparison with a baseline logistic regression trained on raw inputs. For subtyping, K-Means clustering is used on the latent representation of the T2D case cohort. In our analysis, we identify three subtypes (k = 3), characterized by their distance from the control group (see Supplementary Fig. 10 for justification of k = 3). Our approach differs from prior work by learning subtypes exclusively from general EHR data available up to two years before T2D diagnosis, without incorporating features like genetic information or advanced biomarkers. The clustering is based solely on latent distance metrics. To better understand the clinical characteristics of the identified clusters, we conducted post hoc analyses by examining differences in comorbidity rates10,14, medication effects29, and polygenic risk scores30. Although these features are valuable for enrichment and validation, they are not used in model training as they are not routinely collected in individuals at risk for diabetes.
DML prediction of T2D onset
First, we investigate whether the DML latent space can learn information from past EHR data to predict future T2D diagnoses. To quantify the impact of input features, we compare the DML model against logistic regression (LR)13,31,32 under three settings: full EHR (LR), a validated clinical risk-factors model (Risk-Factors)27, and glycemic measures alone (Glycemic)4,5. To compare against other latent space embedding methods, we further compare the DML model against deep learning (SCARF22, TabTransformer23, CVAE24, ConvAE14 and dimensionality reduction baselines (PCA25, UMAP26.
Models using full EHR data (DML and LR) consistently outperform limited feature models when using data from 2 to 7 years before diagnosis (Fig. 2a). This gap underscores the limitations of applying traditional clinical risk factor models in an EHR setting, where passively collected data often lacks key features (Supplementary Figs. 15–16). At 7 years prior to diagnosis, DML achieves an AUROC of 0.754, outperforming LR (0.706), Risk-Factors (0.693), and Glycemic (0.632). Beyond this time point, data quality generally deteriorates, and all models face increasing difficulty with predictions.
T2D onset prediction performance. (a) Temporal performance of the DML, LR, Glycemic, and Wilson models on AoU PopControl data with censor periods ranging from 10 years to 0 years before diagnosis (95% CIs over 500 bootstrap iterations). (b) Bar plot of 2-year T2D onset prediction. AUROC of the DML with LR-baselines (LR, Risk-Factors, Glycemic), deep learning baselines (SCARF, TabTransformer, CVAE, ConvAE), and dimensionality reduction baselines (PCA, UMAP). 95% CIs computed over 500 bootstrap iterations. (c) Transfer performance of MGB-trained DML and LR models evaluated on AoU data. (d) Latent Space Representations from the AoU DML model are visualized through dimensionality reduction with UMAP.
To further assess the DML model, we compare its 2-year T2D onset prediction against a range of baselines using the AoU dataset (Fig. 2b). The DML model achieves the highest AUROC (0.969), outperforming LR baselines (LR: 0.954, Risk-factors: 0.802, Glycemic: 0.773), deep learning baselines (SCARF: 0.918, TabTransformer: 0.909, CVAE: 0.795, ConvAE: 0.571), and dimensionality reduction baselines (PCA: 0.816, UMAP: 0.790). While the latent spaces of deep learning and dimensionality reduction baselines are plausible for subtyping as well as onset prediction (Supplementary Fig. 21), our DML model yields the most predictive latent space. This emphasizes the strength of the DML framework as a unified representation for both subtyping and onset prediction.
To evaluate generalizability, we trained the DML and LR models on the MGB dataset, with strong performance for 2-year prediction (AUROC: DML 0.908, LR 0.898). When applied directly to the AoU data, both models retained predictive power (AUROC: DML 0.829, LR 0.861), despite a noticeable performance drop (Fig. 2c, Supplementary Table 4). This suggests that general EHR features are predictive across cohorts. Lastly, we performed feature importance analysis and observed that the DML model prioritizes weight-related features (e.g., BMI, body weight)33,34 (Supplementary Table 3), while LR relies more on glycemic measures1,4,35 (Supplementary Table 2).
Defining DML subtypes along T2D risk continuum
Beyond T2D onset prediction, subtyping individuals based on future health trajectories enables targeted interventions. DML models create optimized latent spaces that cluster similar individuals and separate dissimilar ones, allowing subtypes to emerge naturally. Both MGB and AoU individuals who develop T2D form a continuum, with controls clustered at one end (Fig. 2d). Subtyping is performed on the full cohort of T2D-positive (case) individuals in each dataset, independent of the control group, to characterize variation within the T2D population. Using KMeans (k = 3), we define Green, Yellow, and Red subtypes based on proximity to controls. Using KMeans (k = 3), we define Green, Yellow, and Red subtypes based on proximity to controls. Projecting AoU individuals onto MGB subtypes shows strong alignment, confirming that our subtypes transfer across populations (Supplementary Fig. 2).
We analyze subtype demographics and key diagnostic markers (Table 1) and find no significant differences (Supplementary Table 5), suggesting that demographics do not drive subtype variation. Random blood glucose levels show no significant differences pre- or post-diagnosis (P = 0.745 before, P = 0.874 after, KS test; Supplementary Fig. 4). HbA1c levels differ significantly only post-diagnosis (P = 0.09 before, P = 0.014 after, KS test; Supplementary Fig. 3). These findings indicate subtypes emerge independently of pre-diagnostic demographic or diagnostic differences but remain relevant for future diagnostic and treatment strategies.
Comorbidity prevalences vary across DML subtypes
To validate subtype specificity, we compare the Green (closest to controls) and Red (farthest) subtypes using diagnostic codes with at least 5% prevalence, assessing T2D-associated conditions via binomial proportion tests (Table 2)36,37,38,39,40. Statistically significant p-values are indicated by * (significant at α = 0.05) or ** (significant after Bonferroni correction with 50 tests at α/50 = 0.001).
The Red subtype consistently shows higher obesity rates (AoU P = 7.2e−05**, MGB P = 2.8E−07**) with earlier divergence in MGB (5 years pre-diagnosis) than AoU (2 years). Obesity-related conditions, such as gastroesophageal reflux disease (GERD), sleep apnea, and hyperlipidemia, are more prevalent in Red, with gaps widening over time. Cardiovascular conditions (AoU) and mental health disorders like depression (AoU P = 2.3E−04**, MGB P = 2.4E−06*) and anxiety (MGB P = 2.3E−07**) are significantly elevated. Red also has higher neuropathy (MGB P = 7.1E−08**) and cataract rates (AoU P = 7.2E−04**). These Red and Green subtype trends are consistent across both the AoU and MGB datasets, illustrating the robustness and reproducibility of the subtype distinctions. See Supplementary Figs. 5–9 for visualizations.
Obesity alone does not fully explain these differences. After adjusting for BMI41 (Supplementary Table 6), cardiovascular42,43 and mental health comorbidities44,45 remain significantly different, suggesting that subtypes capture additional T2D-related variations.
Medication usage and effect vary across DML subtypes
Another significant difference between subtypes is their future medication usage post-diagnosis. We categorize medications into metformin, insulin, and other T2D-related drugs, examining initiation timing and HbA1c response. Metformin tends to be prescribed earlier in the disease course, while insulin is typically initiated later (Fig. 3a,b). While the Green subtype starts medication earlier than the Red subtype (Fig. 3a), the difference is not statistically significant. However, the time to achieve HbA1c control (< 6.5) is significantly shorter in the Green subtype (metformin: P = 2.4E−04; other T2D drugs: P = 2.4E−03, ANOVA), indicating better responsiveness (Fig. 3c). In AoU, the Green subtype shows a significantly larger HbA1c reduction after metformin initiation (P = 4.0E−03, ANOVA), with a mean decrease of − 0.64 vs. − 0.27 in the Red group (Fig. 3d), though responses to other medications do not differ.
T2D medication usage times and effect on HbA1c levels and across AoU PopControl subtypes. (a) Time from T2D diagnosis date to start of medication use (b) time from first measurement of HbA1c ≥ 6.5 to the start of medication (c), time from the start of medication to the first time when all HbA1c < 6.5 afterward (d), change in HbA1c levels in response to start of medication over 3 months (pre-medication: [− 12, 0] months; post-medication: [3, 15] months). The bar plot error bars represent 95% confidence intervals. Medication groups that present significant differences at \(\:\alpha\:=0.05\) are indicated by *.
Genetic contribution to T2D development across DML subtypes
To explore the contribution of genetics to T2D development across the identified subtypes, we calculated the T2D polygenic risk score (PRS)20,46, adjusted for key covariates of the top 10 principal components, age, and sex. We find that while the T2D cases show clear differences in PRS from controls (Supplementary Fig. 11), the DML T2D subtypes are largely similar genetically to each other in both datasets (Supplementary Tables 7–8; pairwise Tukey significance test). Specifically, the PRS for the control group is significantly lower than the T2D subtypes (AoU P = 1.45e−10, MGB P = 6.65E−55, ANOVA).
We further examined the genetic contribution using partitioned polygenic scores (pPS) derived from 12 genetic clusters identified in a previous study (Smith et al.47). These pPS were created by calculating a weighted sum of the genetic variants within each cluster. Our analysis revealed no significant differences between the three DML subtypes in either dataset (Supplementary Table 9, Supplementary Fig. 12). Thus, while the T2D group shows a clear genetic contribution to disease development compared to controls, no differences were observed among the DML subtypes.
Lastly, we compared the performance of polygenic risk score (PRS)-based models with our EHR-based models for predicting T2D onset and subtyping in the AoU PopControl cohort. PRS models (AUROC = 0.642–0.745) performed worse than our DML model (AUROC = 0.969) for predicting disease onset, and PRS subtypes revealed demographic biases that are not present in our DML subtypes. Additionally, PRS subtypes identified fewer significant comorbidities and showed no significant differences in medication effects on HbA1c, making them less effective for clinical insights compared to EHR-based models. See Supplementary Tables 10–12, Supplementary Figs. 13–14 for details.
Discussion
Our work demonstrates the power of deep metric learning (DML) in predicting T2D onset and identifying future T2D subtypes two or more years before diagnosis. Unlike prior studies, our model addresses both tasks with routinely available EHR data, enabling early intervention and precision prevention.
DML offers a key advantage by combining class-aware supervision with metric-based losses to directly shape the latent space around disease outcomes. In contrast, SCARF relies on self-supervised contrastive learning and does not use outcome labels to guide representation learning. TabTransformer is outcome-informed, but does not explicitly enforce structure in the latent space. CVAE enforces structure through a variational bottleneck, but relies on weaker classifier-based supervision. ConvAE’s temporal embeddings does not depend on outcomes. PCA and UMAP are fully unsupervised and ignore outcome information. Overall, DML’s combination of discriminative supervision and metric-based structure uniquely supports both accurate prediction and interpretable patient stratification.
Our model can integrate seamlessly into EHR systems as a non-invasive, automated screening tool, passively identifying high-risk individuals without adding clinical burden. It generalizes well to new cohorts and requires minimal preprocessing. By leveraging a broad range of EHR features, it surpasses standard diagnostic tests (HbA1c, glucose)3,4 in risk assessment, potentially benefiting the estimated 98 million U.S. adults with prediabetes48. While further clinical validation is needed, this approach offers a scalable solution for population-wide opportunistic screening49,50.
Our DML-derived subtypes show both overlap and distinction with existing T2D classifications. The Red subtype most closely resembles Ahlqvist’s SIRD group10, with high BMI and metabolic comorbidities. However, unlike the Ahlqvist model, our subtypes do not differ significantly by age of onset. Instead, they reflect a continuum of metabolic health and complication severity—ranging from the healthier Green to the higher-risk Red subtype. This gradient emerges within a population with similar baseline age and HbA1c, suggesting differences in disease progression and treatment response. The Red subtype also aligns with Wagner’s Cluster 5 (visceral obesity, high T2D and vascular risk), while the Green subtype resembles Cluster 2 (metabolically healthier)12. Compared to Landi et al.’s deep learning subtypes14, our Green group parallels Subtype I (milder complications) and the Red group aligns with Subtype III (severe cardiovascular complications). Unlike Landi’s model, our three subtypes form a clearer continuum of severity (Supplementary Figs. 17–18). In summary, when limited to subtyping using exclusively EHR data, our DML subtypes offer a clinically interpretable and data-driven framework for organizing patients by T2D risk and complication severity.
Our study also has several limitations. The DML model’s latent space shows smooth transitions in comorbidity severity rather than distinct clusters, requiring further trend analysis. The lack of family history data may constrain predictive performance. Lastly, our analysis may be affected by biases inherent in hospital-based retrospective studies, as data collection tends to overrepresent individuals with higher healthcare utilization.
In conclusion, DML offers a scalable, accurate approach for early T2D prediction and subtyping, supporting the advancement of precision medicine in diabetes care.
Methods
Dataset descriptions
All of Us dataset
The All of Us (AoU) program collects longitudinal EHR data from 400,000 participants across more than 340 centers in the United States17, emphasizing underrepresented groups (Supplementary Table 1). Participants are recruited through collaborating academic research centers, community health centers, and online self-recruitment. We use the Controlled Tier Dataset v6, with data up to 01/13/2023.
Ethical statement: All methods were carried out in accordance with relevant guidelines and regulations. This study used de-identified human data from the All of Us Research Program (Controlled Tier), accessed via the All of Us Researcher Workbench under an approved data use agreement. The research was approved by the All of Us Institutional Review Board (IRB Protocol 2021-02-TN-001). Due to the retrospective nature of the study, the requirement to obtain informed consent was waived by the All of Us IRB.
Massachusetts general Brigham dataset
Massachusetts General Brigham (MGB) is a major healthcare system serving over 1.5 million patients annually18. The protocol involving the sharing of deidentified data with MIT was reviewed by the Mass General Brigham. We use a biobank extracted on 10/12/2022, with 109,768 individuals.
Ethical statement: All methods were carried out in accordance with relevant guidelines and regulations. The study using Mass General Brigham (MGB) data was approved by the MGB Institutional Review Board (IRB) under protocol 2022P000611. Informed consent was obtained from all participants through the MGB Partners Biobank under protocol 2009P002312, which was approved by the MGB IRB on 01/17/2022.
Cohort construction
T2D cohort
In AoU, we identify T2D cases using the eMERGE algorithm (n = 7567)19. The T2D diagnosis date is determined by the earliest date out of: any T2D ICD code, any T2D medication code, and any HbA1c > 6.5. MGB cases (n = 3298) are identified using PheCAP21, a custom ML algorithm developed at MGB with 95% PPV.
Population matched controls (PopControl)
Controls are selected using a k-nearest neighbors51 algorithm based on age, sex, and healthcare utilization features, ensuring a 1:1 match with T2D cases (AoU n = 7567, MGB n = 3298). Healthcare utilization is approximated by the total number of EHR measurements per record. This cohort is used for model training to prevent the learning of shortcuts.
General controls (GenControl)
This broader cohort includes individuals without any T2D or T1D-related codes (AoU n = 77,567, MGB n = 81,787). Models are evaluated on this cohort to better reflect real-world disease prevalence and clinical population characteristics.
Data preprocessing
We construct input features from EHR conditions (m = 71), medications (m = 89), physical measurements (m = 6), labs (m = 21), and demographic factors (age, sex). Features are selected by prevalence (highest occurring) and association with T2D (known risk factors). Proxy indicators for T2D (e.g., “Complication due to type 2 diabetes”) are excluded to prevent label leakage. For continuous features, we compute mean, max, and min over the past 6 months, 2 years, and full history (Supplementary Fig. 20). Discrete features (conditions, medications) are binarized similarly. Sex is one-hot encoded, age normalized to [0,1]. All input features are concatenated into a 698-dimensional vector. We randomly split both the case and control cohorts into training (70%), validation (10%), and test (20%) sets, ensuring no overlap in patient records across splits. The validation set is used for hyperparameter tuning, and all final experimental results are reported on the independent hold-out test set. See Supplementary Fig. 1 for details.
Models
DML model
The goal of the deep metric learning (DML) model is to learn a projection \(\:\varphi\::\:\chi\:\to\:\varPhi\:\) that maps high-dimensional EHR features into a lower-dimensional metric space such that individuals with similar T2D status are closer together, while dissimilar individuals are farther apart. Formally, for two individuals \(\:{x}_{1\:},\:{x}_{2}\in\:\chi\:\), the learned distance \(\:d\left(\varphi\:\right({x}_{1}),\:\varphi\:({x}_{2}\left)\right)\) reflects meaningful clinical similarity. For example, the triplet DML loss52 based on distances between query \(\:x\), positive example \(\:{x}_{p}\) and negative example \(\:{x}_{n}\) would be computed as \(\:\underset{}{max}(0,\:\Vert\:\varphi\:(x)-\varphi\:{(x}_{p}){\Vert\:}^{2}+\Vert\:\varphi\:\left(x\right)-{\varphi\:(x}_{n}){\Vert\:}^{2}+m)\), with \(\:m\) being a hyperparameter for the margin distance between positive and negative pairs. We use a neural network encoder composed of 3–4 fully connected layers with ReLU activations and dropout (p = 0.2) to learn the DML projection. The encoder is trained on both T2D-positive and control (T2D-negative) individuals to capture relevant distinctions in the latent space. The learned representations for all individuals are used to train a logistic regression (LR) model to predict T2D onset two years in the future (3-fold cross-validation). For subtyping, we restrict analysis to the learned representations of T2D-positive cases, which are clustered to examine heterogeneity within the T2D population. For additional implementation details, see Supplementary Note 4. For visualization of the model architecture, see Supplementary Fig. 19.
Baseline models
For onset prediction, we compare DML with Logistic Regression (LR), implemented in scikit-learn53, using 3-fold cross-validation. Additional LR baselines include a risk-factors model27 (age, body mass index, blood pressure, high-density lipoprotein, triglyceride, glucose, HbA1c) and a glycemic model4,35 (HbA1c, glucose). For deep learning baselines, we compare against state-of-the-art latent embedding models with contrastive learning (SCARF22, transformers-based encoders (TabTransformer23, conditional variational autoencoders (CVAE24, and a prior T2D subtyping model (ConvAE) from Landi et al.14. ConvAE extracts EHR sequence patterns via convolutional autoencoding. Lastly, we compare against dimensionality reduction methods of PCA25 and UMAP26. A latent embedding dimension of 64 is used to maintain consistency across comparisons.
Training and evaluation
Training details
We train the DML model in PyTorch with triplet52, N-pair54, Lifted55, and ProxyNCA56 losses, varying dimension (d∈{32, 64}) and encoder layers (l∈{3,4}). Training runs for 50 epochs using Adam (learning rate 1e-4). We select the best DML loss and model hyperparameters based on validation AUROC.
Subtyping via clustering
T2D patients are embedded into a latent space, then clustered into three subtypes using KMeans57 (k = 3, see Supplementary Fig. 10 for choice of k). We designate the subtypes with colors based on the relative distance between the subtype and the control group in the representation space. The Green subtype is the closest and overlaps significantly with the control group, while the Yellow subtype is further along the gradient. The Red subtype is the furthest from the controls.
Polygenic risk scores (PRS)
Polygenic risk score (PRS) is generated with the PRS-CS software58 with input of meta-analyzed summary statistics from the European ancestry subset GWAS meta-analysis of T2D in Vujkovic et al.61 and the T2D genome-wide association study by the FINNGEN Consortium60. All PRS are corrected with the covariates of age, sex, and genetic principal components during significance testing.
Onset prediction evaluation
Models are assessed via the area under the receiver operating characteristic curve (AUROC)61, with 95% confidence intervals (CIs) quantified via 500 bootstrap iterations. Feature importance is evaluated via LR coefficients and permutation tests62. Model generalization is tested by transferring the MGB-trained model to AoU.
Clinical subtyping evaluation
Significant tests are used to quantify feature variations across subtypes. For demographic features, we perform the Pearson chi-squared test63 for binary features and the ANOVA test64 for continuous features. For binary features such as comorbidities, we perform the binomial proportion two-sample test. For continuous features such as laboratory values, we perform the two-sample Kolmogorov–Smirnov (KS) test65. Since we performed a total of 50 independent significance tests, we corrected the p-value threshold for significance by applying the Bonferroni correction66\(\:(0.05/50=0.001).\).
Data availability
The All of Us dataset is available for use by the research community once registered through their official research hub ([https://www.researchallofus.org/]). The MGB dataset is not publicly accessible. However, we will release a patient cohort of synthetic examples generated from the MGB cohort that yields similar performance with our trained models upon paper acceptance.
References
CDC. Type 2 Diabetes [Internet]. Centers for Disease Control and Prevention. 2023 [cited 2024 Feb 14]. https://www.cdc.gov/diabetes/basics/type2.html
Guariguata, L. et al. Global estimates of diabetes prevalence for 2013 and projections for 2035. Diabetes Res. Clin. Pract. 103 (2), 137–149 (2014).
US Preventive Services Task Force. Screening for prediabetes and type 2 diabetes: US preventive services task force recommendation statement. JAMA 326 (8), 736–743 (2021).
ElSayed, N. A. et al. 2. Classification and diagnosis of diabetes: standards of care in Diabetes—2023. Diabetes Care. 46 (Supplement_1), S19–40 (2022).
CDC. Diabetes Testing [Internet]. Centers for Disease Control and Prevention. 2023 [cited 2024 Feb 14]. https://www.cdc.gov/diabetes/basics/getting-tested.html
Hu, F. B. Metabolic profiling of diabetes: from Black-Box epidemiology to systems epidemiology. Clin. Chem. 57 (9), 1224–1226 (2011).
Dimas, A. S. et al. Impact of type 2 diabetes susceptibility variants on quantitative glycemic traits reveals mechanistic heterogeneity. Diabetes 63 (6), 2158–2171 (2014).
Franks, P. W., Pearson, E. & Florez, J. C. Gene-environment and gene-treatment interactions in type 2 diabetes: Progress, pitfalls, and prospects. Diabetes Care. 36 (5), 1413–1421 (2013).
Deutsch, A. J., Ahlqvist, E. & Udler, M. S. Phenotypic and genetic classification of diabetes. Diabetologia 65 (11), 1758–1769 (2022).
Ahlqvist, E. et al. Novel subgroups of adult-onset diabetes and their association with outcomes: a data-driven cluster analysis of six variables. Lancet Diabetes Endocrinol. 6 (5), 361–369 (2018).
Kim, H. et al. High-throughput genetic clustering of type 2 diabetes loci reveals heterogeneous mechanistic pathways of metabolic disease. Diabetologia 66 (3), 495–507 (2023).
Pathophysiology-based subphenotyping of. individuals at elevated risk for type 2 diabetes | Nature Medicine [Internet]. [cited 2024 Apr 23]. https://www.nature.com/articles/s41591-020-1116-9
Anderson, A. E. et al. Electronic health record phenotyping improves detection and screening of type 2 diabetes in the general united States population: A cross-sectional, unselected, retrospective study. J. Biomed. Inf. 60, 162–168 (2016).
Landi, I. et al. Deep representation learning of electronic health records to unlock patient stratification at scale. NPJ Digit. Med. 3 (1), 96 (2020).
Lou, J., Wang, Y., Li, L. & Zeng, D. Learning latent heterogeneity for type 2 diabetes patients using longitudinal health markers in electronic health records. Stat. Med. 40 (8), 1930 (2021).
Bej, S. et al. Identification and epidemiological characterization of Type-2 diabetes sub-population using an unsupervised machine learning approach. Nutr. Diabetes. 12 (1), 27 (2022).
Ramirez, A. H. et al. The all of Us research program: data quality, utility, and diversity. Patterns. 3(8). (2022).
Boutin, N. T. et al. The evolution of a large biobank at mass general Brigham. J. Pers. Med. 12 (8), 1323 (2022).
Type 2 Diabetes Mellitus | PheKB [Internet]. [cited 2024 Feb 13]. https://phekb.org/phenotype/type-2-diabetes-mellitus
Szczerbinski, L. et al. Algorithms for the identification of prevalent diabetes in the All of Us Research Program validated using polygenic scores—a new resource for diabetes precision medicine [Internet]. medRxiv; 2023 [cited 2024 Apr 12]. p. 2023.09.05.23295061. https://www.medrxiv.org/content/https://doi.org/10.1101/2023.09.05.23295061v1
Zhang, Y. et al. High-throughput phenotyping with electronic medical record data using a common semi-supervised approach (PheCAP). Nat. Protoc. 14 (12), 3426–3444 (2019).
Bahri, D., Jiang, H., Tay, Y. & Metzler, D. SCARF: Self-Supervised Contrastive Learning using Random Feature Corruption. arXiv; [cited 2025 Jun 9]. (2022). http://arxiv.org/abs/2106.15147
Huang, X., Khetan, A., Cvitkovic, M. & Karnin, Z. TabTransformer: Tabular Data Modeling Using Contextual Embeddings [Internet]. arXiv; [cited 2025 Jun 9]. (2020). http://arxiv.org/abs/2012.06678
Kingma, D. P., Rezende, D. J., Mohamed, S. & Welling, M. Semi-Supervised Learning with Deep Generative Models [Internet]. arXiv; [cited 2025 Jun 9]. (2014). http://arxiv.org/abs/1406.5298
Principal Component Analysis [Internet]. New York: Springer-Verlag. [cited 2025 Jun 9]. (Springer Series in Statistics). http://link.springer.com/ (2002). https://doi.org/10.1007/b98835
McInnes, L., Healy, J. & Melville, J. UMAP: Uniform manifold approximation and projection for dimension reduction. arXiv; [cited 2025 Jun 9]. (2020). http://arxiv.org/abs/1802.03426
Wilson, P. W. F. et al. Sr. Prediction of incident diabetes mellitus in Middle-aged adults: the Framingham offspring study. Arch. Intern. Med. 167 (10), 1068–1074 (2007).
Tests of Glycemia for the Diagnosis of Type 2 Diabetes Mellitus. | Annals of Internal Medicine [Internet]. [cited 2024 Feb 14]. https://www.acpjournals.org/doi/full/https://doi.org/10.7326/0003-4819-137-4-200208200-00011
Shepherd, M. H. et al. A UK nationwide prospective study of treatment change in MODY: genetic subtype and clinical characteristics predict optimal glycaemic control after discontinuing insulin and Metformin. Diabetologia 61 (12), 2520–2527 (2018).
DiCorpo, D. et al. Type 2 diabetes partitioned polygenic scores associate with disease outcomes in 454,193 individuals across 13 cohorts. Diabetes Care. 45 (3), 674–683 (2022).
Edlitz, Y. & Segal, E. Prediction of type 2 diabetes mellitus onset using logistic regression-based scorecards. eLife 11, e71862 (2022).
Brisimi, T. S. et al. Predicting Chronic Disease Hospitalizations from Electronic Health Records: An Interpretable Classification Approach [Internet]. arXiv; [cited 2025 Mar 1]. (2018). http://arxiv.org/abs/1801.01204
Ganz, M. L. et al. The association of body mass index with the risk of type 2 diabetes: a case–control study nested in an electronic health records system in the united States. Diabetol. Metab. Syndr. 6 (1), 50 (2014).
Lee, D. H. et al. Comparison of the association of predicted fat mass, body mass index, and other obesity indicators with type 2 diabetes risk: two large prospective studies in US men and women. Eur. J. Epidemiol. 33 (11), 1113–1123 (2018).
Barr, R. G., Nathan, D. M., Meigs, J. B. & Singer, D. E. Tests of glycemia for the diagnosis of type 2 diabetes mellitus. Ann. Intern. Med. 137 (4), 263–272 (2002).
Sevilla-González, M., del Quintana-Mendoza, R. & Aguilar-Salinas, B. M. Interaction between depression, obesity, and type 2 diabetes: A complex picture. Arch. Med. Res. 48 (7), 582–591 (2017).
Schlienger, J. L. Type 2 diabetes complications. Presse Medicale Paris Fr. 42 (5), 839–848 (2013).
The Mental Health Comorbidities. of Diabetes | Diabetes | JAMA | JAMA Network [Internet]. [cited 2024 Apr 7]. https://jamanetwork.com/journals/jama/article-abstract/1888681
Mechanisms of Disease. hepatic steatosis in type 2 diabetes—pathogenesis and clinical relevance | Nature Reviews Endocrinology [Internet]. [cited 2024 Apr 7]. https://www.nature.com/articles/ncpendmet0190
Changing epidemiology of type 2 diabetes mellitus. and associated chronic kidney disease | Nature Reviews Nephrology [Internet]. [cited 2024 Apr 7]. https://www.nature.com/articles/nrneph.2015.173
CDC. Defining Adult Overweight and Obesity [Internet]. Centers for Disease Control and Prevention. 2022 [cited 2024 Feb 13]. https://www.cdc.gov/obesity/basics/adult-defining.html
Ortega, F. B., Lavie, C. J. & Blair, S. N. Obesity and cardiovascular disease. Circ. Res. 118 (11), 1752–1770 (2016).
Khan, S. S. et al. Association of body mass index with lifetime risk of cardiovascular disease and compression of morbidity. JAMA Cardiol. 3 (4), 280–287 (2018).
Magallares, A. & Pais-Ribeiro, J. L. Mental health and obesity: A meta-analysis. Appl. Res. Qual. Life. 9 (2), 295–308 (2014).
Scott, K. M. et al. Obesity and mental disorders in the general population: results from the world mental health surveys. Int. J. Obes. 32 (1), 192–200 (2008).
Deutsch, A. J. et al. Type 2 diabetes polygenic score predicts the risk of Glucocorticoid-Induced hyperglycemia in patients without diabetes. Diabetes Care. 46 (8), 1541–1545 (2023).
Smith, K. et al. Multi-ancestry polygenic mechanisms of type 2 diabetes. Nat. Med. 30 (4), 1065–1074 (2024).
CDC. Prediabetes - Your Chance to Prevent Type 2 Diabetes [Internet]. Centers for Disease Control and Prevention. 2021 [cited 2024 Feb 15]. http://bit.ly/2hMpYrt
Jiang, L. Y. et al. Health system-scale language models are all-purpose prediction engines. Nature 619 (7969), 357–362 (2023).
van Leeuwen, K. G., Schalekamp, S., Rutten, M. J. C. M., van Ginneken, B. & de Rooij, M. Artificial intelligence in radiology: 100 commercially available products and their scientific evidence. Eur. Radiol. 31 (6), 3797–3804 (2021).
Fix, E. & Hodges, J. L. Discriminatory Analysis. Nonparametric discrimination: consistency properties. Int. Stat. Rev. Rev. Int. Stat. 57 (3), 238–247 (1989).
Ge, W. Deep metric learning with hierarchical triplet loss. In Proceedings of the European Conference on Computer Vision (ECCV), 269–285 (2018).
Pedregosa, F. et al. Scikit-learn: machine learning in python. J. Mach. Learn. Res. 12 (85), 2825–2830 (2011).
Sohn, K. Improved deep metric learning with multi-class n-pair loss objective. In Advances in Neural Information Processing Systems, 1857–1865 (2016).
Oh Song, H., Xiang, Y., Jegelka, S. & Savarese, S. Deep metric learning via lifted structured feature embedding. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 4004–4012 (2016).
Movshovitz-Attias, Y., Toshev, A., Leung, T. K., Ioffe, S. & Singh, S. No fuss distance metric learning using proxies. In Proceedings of the IEEE International Conference on Computer Vision (ICCV) (2017).
Lloyd, S. Least squares quantization in PCM. IEEE Trans. Inf. Theory. 28 (2), 129–137 (1982).
Ge, T., Chen, C. Y., Ni, Y., Feng, Y. C. A. & Smoller, J. W. Polygenic prediction via bayesian regression and continuous shrinkage priors. Nat. Commun. 10 (1), 1776 (2019).
Discovery of 318 new risk loci for type 2 diabetes and related vascular outcomes among 1.4 million participants in a multi-ancestry meta-analysis | Nature Genetics [Internet]. [cited 2024 Feb 16]. https://www.nature.com/articles/s41588-020-0637-y
FinnGen provides genetic. insights from a well-phenotyped isolated population | Nature [Internet]. [cited 2024 Feb 16]. https://www.nature.com/articles/s41586-022-05473-8
Hajian-Tilaki, K. Receiver operating characteristic (ROC) curve analysis for medical diagnostic test evaluation. Casp. J. Intern. Med. 4 (2), 627–635 (2013).
Altmann, A., Toloşi, L., Sander, O. & Lengauer, T. Permutation importance: a corrected feature importance measure. Bioinformatics 26 (10), 1340–1347 (2010).
Pearson, K. X. On the criterion that a given system of deviations from the probable in the case of a correlated system of variables is such that it can be reasonably supposed to have arisen from random sampling. Lond. Edinb. Dublin Philos. Mag J. Sci. 50 (302), 157–175 (1900).
Girden, E. R. ANOVA: Repeated Measures, 88 (SAGE, 1992).
Massey, F. J. The Kolmogorov–Smirnov test for goodness of fit. J. Am. Stat. Assoc. 46 (253), 68–78 (1951).
Armstrong, R. A. When to use the bonferroni correction. Ophthalmic Physiol. Opt. J. Br. Coll. Ophthalmic Opt. Optom. 34 (5), 502–508 (2014).
Acknowledgements
The All of Us Research Program is supported by the National Institutes of Health, Office of the Director: Regional Medical Centers: 1 OT2 OD026549; 1 OT2 OD026554; 1 OT2 OD026557; 1 OT2 OD026556; 1 OT2 OD026550; 1 OT2 OD 026552; 1 OT2 OD026553; 1 OT2 OD026548; 1 OT2 OD026551; 1 OT2 OD026555; IAA #: AOD 16037; Federally Qualified Health Centers: HHSN 263201600085U; Data and Research Center: 5 U2C OD023196; Biobank: 1 U24 OD023121; The Participant Center: U24 OD023176; Participant Technology Systems Center: 1 U24 OD023163; Communications and Engagement: 3 OT2 OD023205; 3 OT2 OD023206; and Community Partners: 1 OT2 OD025277; 3 OT2 OD025315; 1 OT2 OD025337; 1 OT2 OD025276. In addition, the All of Us Research Program would not be possible without the partnership of its participants. Supported in part by Quanta Computing, a National Science Foundation (NSF) 22-586 Faculty Early Career Development Award (#2339381), a Gordon & Betty Moore Foundation award, a Google Research Scholar award and the AI2050 Program at Schmidt Sciences.
Author information
Authors and Affiliations
Contributions
Q.J. and H.Z. performed the experiments, prepared the figures, and wrote the main manuscript text. M. G. and M.U. supervised the project. L.S. provided the main point of clinical support. J.Z., W.G., X.X., K.W., and T.H. contributed to manuscript writing. S.H., R.M., A.D., A.M., and J.M. provided valuable dataset support and technical guidance. All authors reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Jin, Q., Zhang, H., Szczerbinski, L. et al. Opportunistic screening of type 2 diabetes with deep metric learning using electronic health records. Sci Rep 15, 41892 (2025). https://doi.org/10.1038/s41598-025-25759-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-25759-x