Abstract
Celiac disease etiopathogenesis requires genetic predisposition and exposure to gluten, yet these factors alone are not sufficient. Larger longitudinal studies are needed to determine the role of time-varying infections and gut microorganisms. The aim was to design a celiac disease case-cohort longitudinal study using The Environmental Determinants of Diabetes in the Young (TEDDY) study. By age 3-years, persistent tissue transglutaminase autoantibodies (tTGA), i.e., celiac disease autoimmunity (CDA), was confirmed in 704 of the 6132 genetically at-risk TEDDY children. Celiac disease onset (CD-onset) was defined as the age CDA developed when followed by a biopsy-proven diagnosis. A competing risk analysis on CD-onset and CDA children with no diagnosis (CDA-only) revealed female-sex, HLA and non-HLA genes and higher gluten-consumption correlate with an increased risk of both outcomes. However, reports of virus-related respiratory infections from August to October correlate consistently with an increased risk of CD-onset and not CDA-only. A sub-cohort of 561 children (9% sampling fraction) has been randomly selected to represent the TEDDY cohort. All incident CD-onset cases (N = 306) were included. The case-cohort will be utilized to analyze virus antibodies and bacteriome from longitudinal plasma and stool samples (the Microbial Associations and Viruses on the Risk of Celiac disease study, MAVRiC).
Similar content being viewed by others
Introduction
Celiac disease is a complex, primarily T-cell, immune mediated1 systemic disorder that manifests due to damage caused to the lining of the intestinal mucosa in the small bowel2. Without treatment by gluten free diet, absorption of nutrients is often impaired, leading to malnutrition and possibly both gastrointestinal and non-gastrointestinal symptoms. Globally, the prevalence of celiac disease exceeds 1.4% with an increasing trajectory and an impact that is highest for the child3.
The age of development of celiac disease in childhood is highly variable as revealed by cohort studies4. The onset of disease is known to require two components, the ingestion of gluten to elicit an autoimmune response5 and the presence of HLA-DR3-DQ2 or DR4-DQ8 haplo-genotypes6,7. Although factors like family history, female sex, and non-HLA loci contribute to the risk of celiac disease, genetics account for only about 50% of the risk and gluten exposure alone is insufficient to trigger the disease. Epidemiological studies suggest that environmental factors, such as infections and gut microbiota, may trigger with celiac disease autoimmunity (CDA) (i.e. seroconversion to tissue transglutaminase autoantibodies, tTGA) or accelerate progression to intestinal damage (i.e. celiac disease)4,8 . However, the identification and characterization of microbial associations is difficult without studies that frequently collect biological samples and without a defined longitudinal outcome indicating when mucosal damage is truly initiated over time. A biopsy has been considered the “gold standard” for diagnosis, but a positive histology alone is not sufficient, and a final diagnosis should include a combination of positive serology and histology, with or without classical clinical features and a response to a gluten free diet (GFD)9. These time-varying criteria are unlikely to be measured continuously over time or even develop simultaneously. Nevertheless, cohort studies consistently show that peak incidences of both CDA seroconversion and celiac disease diagnosis occur after 1-year and before school age7,10,11. Thus, the prodromal period between positive serology and histology is short enough at this young age to link disease initiation close to CDA seroconversion. The early peak incidences further suggest that environmental factors may act early in life to promote inflammation, reduce tolerance, or condition an aggressive immune response to the rising gluten consumption. Although a recent study showed that measurement of IL-2 may define an even earlier disease onset than that of time to seroconversion to tTGA12, the present study of incident CDA by age 3-years with a subsequent biopsy-proven diagnosis still offers an excellent opportunity to study factors such as early viruses and bacteria as candidate triggers of the onset of celiac disease.
The Environmental Determinants of Diabetes in the Young (TEDDY) prospective study13,14,15,16 has screened CDA annually and followed up for celiac disease among HLA-DR3-DQ2 or HLA-DR4-DQ8 risk children up to 15 years of age at six clinical sites in four countries across two continents. TEDDY has identified over 113,000 parental reported infectious episodes in early childhood while at risk of CDA and has reported correlations between CDA with reported gastro-intestinal infections (GIE)17, season of birth18 and enteroviruses19. Additionally, TEDDY has collected monthly stool samples and quarterly plasma samples from all children, between 3 and 48 months of age, making it possible to study whether viruses and/or bacteria are associated with subsequent disease risk.
The primary aim of this study was to select a longitudinal outcome defining celiac disease onset using the TEDDY study and design an ancillary study, Microbial Association and Viruses on the Risk of Celiac disease (MAVRiC), to examine the role of bacterial and viruses with risk of celiac disease. A secondary aim is to re-examine reported infections in the TEDDY cohort with the risk of CD-onset to generate hypothesis as to the time-dependent role of microorganisms on risk of celiac disease. In this study, we hypothesize that infections reported early in life will associate with risk of celiac autoimmunity differently for children who are subsequently diagnosed with celiac disease as compared to children who are not.
Methods
Study designs
Cohort study (TEDDY): As an observational birth cohort study, TEDDY is following children for type 1 diabetes (T1D) until 15 years of age14. The study aims to identify environmental risk factors of islet autoimmunity (IA) and CDA as primary outcomes and will follow for T1D and celiac disease as secondary outcomes. Between September 2004 and February 2010, 424,788 newborns across three clinical centers in the US (Colorado, Georgia/Florida, and Washington state) and three centers in the EU (Sweden, Finland, Germany) were screened for HLA-DR-DQ haplo-genotypes and 21,583 children were deemed eligible. At the age of 3 to 4 months, 8,676 of these families enrolled in the study of which 6,555 (76%) participated in annual screening for CDA by age 3-years. Nearly all enrolled children (96%) had one of the following four HLA-DR-DQ haplo-genotypes: DR3-DQ2/DR3-DQ2 (20.7%), DR3-DQ2/DR4-DQ8 (39.0%), DR4-DQ8/DR4-DQ8 (19.5%), DR4-DQ8/DR8-DQ4 (16.7%). Additional characteristics at the time of enrollment are shown elsewhere7. The follow-up of children consisted of quarterly clinical visits and blood-draws until the age of 48 months. Stool samples were collected monthly starting at enrollment until age 48 months. To preserve priority samples, children positive for IA before 4 years of age (n = 423/6555) were excluded at the request of the TEDDY study. The final Celiac Cohort study included 6132 children. This ancillary study was approved by the TEDDY ancillary committee and is being led by several members of the TEDDY Infectious Agents and Celiac Disease committees.
Case-Cohort study (MAVRiC): To combine the flexible advantages of a cohort study with the efficiency of a nested case–control (NCC), a case-cohort design20 was chosen for the MAVRiC study to address the role of virus infections and bacteriome in the development of celiac disease in the TEDDY study. Unlike an NCC study that first identifies incident cases during follow-up before selecting 1 to 3 controls to match with each case, the case-cohort design involves first selecting a sub-cohort of children from the original cohort as a representative baseline population20,21. All incident cases outside this sub-cohort are later included to complete the case-cohort study design. Cases chosen into the sub-cohort by random chance represent time-varying controls while they remain at risk of the case event. They are weighed differently in analysis to account for the fact that cases are oversampled. The selection of the sub-cohort allows for its flexible re-use as a comparison group both in stratified analysis and if the study needs to be extended in the future to include additional cases.
The sub-cohort consisted of 600 randomly selected children from the original cohort of 6555 (9.2% sampling fraction) before IA positive cases were excluded. Of the children selected, 39/600 developed IA before 4-years of age and were omitted from the remaining analysis. The random selection was performed before excluding the IA cases so that sensitivity analyses could be performed at a later stage to study the impact of their exclusion.
Onset of celiac disease as a primary outcome
The inclusion into the MAVRiC case-cohort study of all children developing any CDA was cost prohibitive. Moreover, any biopsy conducted to determine celiac disease was often a year after CDA development and after the disease had likely developed. Therefore, the primary outcome of interest in the MAVRiC study was the onset of celiac disease (CD-onset) defined as the age when CDA develops before age 4-years and celiac disease is subsequently confirmed with a positive biopsy. Children developing CDA without a subsequent celiac disease diagnosis (CDA-only) were examined in the full cohort only and were not included in the MAVRiC case-cohort study. A comparison of the outcome definitions in TEDDY and MAVRiC studies are presented in Table 1.
The TEDDY protocol was designed for timely and thorough ascertainment of the development of CDA and subsequent celiac disease7. The annual screening for tTGA started at 24 months of age using radiobinding assays as described previously7. Children with tTGA levels > 1.3 U/ml were defined as being tTGA positive and had their subsequent sample tested to determine if the child had developed CDA. Two consecutive positive tTGA results at least 3 months apart defined any CDA. If a child missed their annual visit or had no sample available, tTGA screening results were collected at the next visit. Any child positive for tTGA had their remaining available samples tested retrospectively to determine CDA seroconversion. Nearly all the children developing a CDA up to the age of the 27-month visit tested positive on their first annual screen, while CDA between 30- and 48-month visits were initially screened negative. The incidence proportion of any CDA by age 3-years in the TEDDY Celiac Cohort was 11.5% (704/6132). Tissue TGA was monitored frequently in the entire cohort after CDA seroconversion, with prompt referral of those positive for further medical evaluation. An intestinal biopsy showing a Marsh score 2 or greater was indicative for celiac disease. Diagnosis of celiac disease was established if children with Marsh score > 1 had significant reduction of tTGA levels and relief of eventual symptoms after treatment with gluten free diet at follow-up in accordance to the NASPGHAN criteria22.
The primary outcome of the MAVRiC study is CD-onset defined from the TEDDY outcomes as CDA before 4 years of age with a celiac disease diagnosis before age 14 years, or before August 31st ,2020, whichever came first. In all, 306 of the 704 children developing any CDA had CD-onset. Because the identification of CD-onset was determined through retrospective analysis of the TEDDY cohort, it was necessary to have a secondary CDA phenotype (CDA-hi) with a higher specificity for mucosal damage during CDA development and not after (Supplemental Fig. 1). This was to ensure microbial correlations found with risk of CD-onset also occurred with early evidence of disease progression at the time of CDA development and not on factors occurring only after CDA development. A secondary outcome in the MAVRiC study is CDA-hi and is defined as any CDA before 4-years of age with TGA levels of the persistent sample ≥ 60U/ml. Tissue TGA patterns during CDA development were examined for correlation with a subsequent celiac disease diagnosis. All but seven of the 704 children were followed in TEDDY after CDA development. The median (IQR) number of years from the first sample positive for tTGA positive to the diagnosis of biopsy-proven celiac disease was 1.1 (0.8, 1.7) years. Compared to the first positive sample, the tTGA level of the persistent positive sample more clearly correlated with celiac disease diagnosis (Supplemental Fig. 2). Furthermore, tTGA level distributions of the persistent positive sample compared to the first positive samples showed stronger correlation (r = 0.86, p < 0.001) with the child’s maximum average tTGA levels across any two consecutive samples during following (r = 0.54, p < 0.001, Supplemental Fig. 3). Of the children with CDA-hi (n = 202) with tTGA level > 60 U/ml in the persistent positive sample, 10% (20/202) were not subsequently determined for celiac disease (i.e. CD-onset). These 20 CDA-only children were still added to the MAVRiC study as a baseline for secondary outcome analysis using only serological information; 11/20 had a follow-up biopsy that was negative and 9/20 had max average tTGA level in follow-up samples between 40 and 100 U/ml with no biopsy.
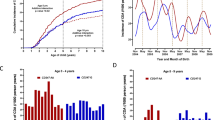
Age-specific incidence reports of infectious episode categories (Panel A), mean ± 95%CI gluten intake by age (Panel B) and age-specific incidence of CDA and celiac disease before four years of age (Panel C).
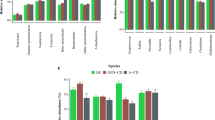
Number of monthly (Panel A) and age-seasonal (Panel B) specific reports of virus related RIEs during the first year of life on the risk of CD-onset before the age of 2.5 years.
TEDDY cohort followed for CD-onset and CDA-only until age 3-years and MAVRiC CD-onset case-cohort study.
Reported infectious episodes
Symptoms or diagnosis of any illnesses were recorded by the parents in a TEDDY diary-book or at each clinic visit23. Infections reported from the previous visit were collected by study nurses who translated illness reports into diagnosis codes according to the ICD-10 classification. Infectious disease data processing and categorization in the TEDDY used an infectious episode (RIE) approach reduces the possibility of overestimation of reported exposure due to multiple symptoms and/or diagnosis reports during a single microbial infection23. In the present study, four common categories of respiratory infectious episodes were of interest: (1) virus related respiratory type of infections that included mostly of common colds but also laryngitis and tracheitis, specific indication of influenza, enterovirus, chicken pox (varicella), measles, mumps, rubella herpes simplex virus infection and other virus infections not elsewhere classified; (2) infections of ear and mastoid process; (3) bronchitis and lower respiratory infections and (4) conjunctivitis. Two common categories of GIE were (1) gastroenteritis with report of infection and (2) gastrointestinal symptoms. All infections and categories were identified as having a record of an ICD-10 code since the last scheduled clinic visit. Given the season of birth18 and enteroviruses19 was of particular interest, the common virus-related respiratory infection was examined every 3-months starting with infections reported in the fall between August and October when enterovirus prevalence was expected to peak (Fig. 1A).
Gluten intake was estimated from 3-day food records collected at ages 6, 9, and 12 months and biannually thereafter. TEDDY has previously reported on the gluten intake during the first 5 years of life11. Higher intake correlates with an increased risk of CDA and celiac disease, whereas the age of introduction to gluten is not associated with these outcomes. From the age of 6 until 18 months, gluten intake rose rapidly (Fig. 1B). Thereafter, the rate of increase was more gradual. Since only 44 CDA cases were observed before the age of 18 months (Fig. 1C), the rapid rise in daily intake was captured by a linear slope trajectory between the age of 6 and 18 months while at risk of CDA and was calculated by fitting a linear mixed model with a random slope. If a child missed a 6-month visit, gluten intake at the 3 month-visit was included. The extracted slopes describing the rise in gluten intake for each child were accounted in every correlation model examining reported infections with risk of CDA. The average slope for a child during this age year was + 3.8 per average daily grams. Child on a gluten free diet was rare before CDA development and common after CDA development (Supplemental Fig. 4).
A polygenic risk score (PRS) for celiac disease was created to account for any non-HLA single nucleotide polymorphisms (SNPs) previously listed to be associated with CDA or celiac disease in TEDDY.24 SNPs previously reported and available from the ImmunoChip or TEDDY-T1DExomeChip data were re-examined for their association with CD-onset before the age of 4 years. Each SNP was examined in relation to the risk of CD-onset using Cox proportional hazard models adjusting for country of residence, sex, HLA-DR-DQ haplo-genotype and the first three principal components accounting for population stratification (ancestral heterogeneity). The PRS was created that summed the product of log beta coefficient (log hazard ratio = log HR) from the proportional hazard model and number of minor alleles of the SNPs (Supplemental Table 1). The polygenic score was included in the final model to account for genetic risk of the non-HLA loci.
Statistical analysis
A competing risk analysis of CD-onset, CDA-hi and CDA-only was performed on the whole cohort (n = 6132). Multivariate proportional hazard models were used to characterize associations with genetic risk factors7,24, sex, demographic and socioeconomic factors17, exposure to gluten11,25, month of birth18 and infectious episodes17 all of which were previously reported associated with any CDA or celiac disease. Sensitivity analysis also considered model adjustments for vaccines17 and antibiotics26 that were not associated with risk of CDA overall, as well as model adjustment for factors related to early or later withdrawal that may contribute to selection bias in the TEDDY study27,28. All children developing CDA were censored at the time of seroconversion including the CDA group of interest and competing CDA groups. Effect sizes were described by outcome-specific hazard ratios with 95% confidence intervals. Categories of infectious episodes were examined as time-invariant factors during the first year of life and as time-varying factors during any year of follow-up. The age of infection was modeled as a step function and infections during follow-up prior to CDA were modeled as a lag function. The same analysis on risk of a CD-onset was repeated in the MAVRiC case-cohort study. However, CD-onset (and CDA-hi) cases were intentionally over-represented in a case-cohort design which necessitated an appropriate adjustment to the proportional hazard models20,29. Instead of using the partial likelihood, a pseudolikelihood estimator was implemented. This approach incorporates weights to produce estimates of hazard ratios comparable to those observed in the cohort. Prentice weights were used to correct for oversampling of cases in the case-cohort design and confidence intervals were calculated using robust standard errors30. Unless otherwise stated, p-values less than 0.05 were considered significant. All statistical analyses were performed using R, version 4.4.2 (www.R-project.org), SAS, Version 9.4 (SAS Institute Inc. Cary, NC, USA), and figures generated using GraphPad PRISM-10.4.1 (GraphPad Software Inc., San Diego, CA).
Results
Factors with specific risk for CD-onset in the whole cohort
Factors previously reported associated with any CDA in TEDDY4,14 up to age 3-years were examined together in a multivariate analysis on the specific risk of CD-onset, CDA-hi and CDA-only. Female-sex, number of HLA-DR3-DQ2 haplo-genotypes, first degree relative with celiac disease, an increase in the non-HLA PRS, a rapid rise in gluten intake after 3 to 18 months, born April to July compared to other months were independently associated with an increased risk of CD-onset (Table 2). In addition, the number of virus-related respiratory infectious reported August to October during first year of life correlated with a significant increased risk of CD-onset (/additional RIE reported Aug to Oct, HR = 1.28, 95%CI = 1.12 – 1.48, p = 0.0005). After accounting for these factors, the country of residence, increase in fiber intake during the first year31, mother’s education, age mother stopped breastfeeding, age child started gluten25 were not associated with risk of CD-onset (not all data shown in Table 2). Except for family history of celiac disease, all the factors correlating with CD-onset also correlated significantly with increased risk of CDA-hi that was highly predictive of a celiac diagnosis shortly after CDA seroconversion.
However, it was noted that the risk of CD-onset and not CDA-hi, (i.e. CDA development with tTGA levels < 60U/ml in the persistent sample and subsequent biopsy proven celiac disease) was more strongly correlated with a family history of celiac disease (HR = 2.93, 95%CI = 1.46 – 5.88, p = 0.002), a more rapid rise in gluten intake between 6 and 18 months (HR = 1.43, 95%CI = 1.26 – 1.63, p < 0.001), and the risk was marginally reduced with a 3 to 12 month rise in fiber intake (HR = 0.93, 95%CI = 0.87 – 1.00, p = 0.07). Birth month and virus-related RIE reports from August to October during the first year of life did not correlate with this CDA group. Similarly, month of birth from April to July and RIE reports during the first year of life were not associated with the risk of CDA-only (i.e. risk of CDA with no subsequent celiac disease diagnosis, Table 2).
Age and timing of infectious episodes on risk of CD-onset in the whole cohort
The impact of RIEs and GIEs overall and by category on the risk of CD-onset was examined in more detail. In any year during follow-up, RIEs but not GIEs correlated with a subsequent increased risk of CD-onset (/additional infection, HR = 1.14, 95%CI = 1.02 – 1.26 p = 0.02). An examination of the common category of RIEs showed more reports of virus-related respiratory infections reported August to October in any year during follow-up explained this increased risk of CD-onset (/additional infection, HR = 1.24, 95%CI = 1.08 – 1.43, p = 0.002). Given the similar result with August to October reported infections during the first year, Table 2, a stratification analysis by age of CD-onset was conducted (< 2.5 years vs. 2.5—< 4.0 years). The age ranges were indicative of whether the child was identified with CDA on their first annual screened at age 2-years or in later years.
Each additional virus related infections reported between August and October in the first year of life correlated with an increased risk of CD-onset before 2.5 years (additional fall infection, HR = 1.56, 95%CI = 1.30 – 1.88, p < 0.0001), but not from 2.5 to < 4 years (/additional fall infection, HR = 1.02, 95%CI = 0.81 – 1.27, p = 0.89, age*interaction p = 0.007). No other reported infectious episode categories were associated with CD-onset before or after 2.5 years.
These infection reports were examined in greater detail between quarterly visits on the risk of CD-onset before 2.5 years. Virus-related RIE reported August to October in the first year of life correlated with strong consistency both by quarter month and by quarter age (3 to 6, 6 to 9 and 9 to 12 months) on the risk of CD-onset, Fig. 2.
The MAVRiC CD-onset case cohort study
The sub-cohort random selection from the TEDDY Celiac Cohort included 483 children who were followed in TEDDY through age 3-years, and 78 children who were followed until they developed CDA. By chance, these children were chosen to serve as time varying controls representative of the whole TEDDY cohort over time before their seroconversion. The inclusion of CDA children will allow for analysis of factors associated with the hazard-risk of disease. The representative CDA group included 37 children who subsequently developed CD-onset by age 3-years and 41 children with CDA-only, Fig. 3. All incident CD-onset cases are included in the MAVRiC case-cohort designed study. A description of the sub-cohort and TEDDY celiac cohort by country, sex, HLA, CD-onset and by CDA-only are shown in Table 3. Similarly, the 202 CDA-hi cases are characterized by country, sex and HLA, Supplemental Table 2.
Risk factors of CD-onset were re-examined on the case-cohort study. As expected, the month of birth between April and July as compared to other months (HR = 1.52, 95%CI = 1.11 – 2.08, p = 0.008) and number of virus-related RIE reported August to October during the first year (/additional report, HR = 1.23, 95%CI = 1.03 – 1.46, p = 0.02) remained significantly associated with CD-onset after adjusting for the other risk factors. Similarly, April to July month of birth (HR = 1.93, 95%CI = 1.32 – 2.82, p = 0.0006) as well as virus-related respiratory infections reported August to October (/additional report, HR = 1.32, 95%CI = 1.08 – 1.63, p = 0.008) were more strongly associated with CDA-hi.
Discussion
Infections and gut microorganisms have been commonly implicated in the etiology or pathogenesis of celiac disease. However, the lack of large longitudinal studies among genetically at-risk populations has made it challenging to link exposures that vary with age and season, with a time-dependent specific risk of disease. TEDDY is the largest longitudinal study that has prospectively screened genetically at-risk populations in the US and Europe for the incidence of celiac disease autoimmunity before a celiac disease diagnosis. A retrospective examination of children with autoimmunity, with and without a subsequent biopsy-proven celiac disease diagnosis, has revealed virus-related respiratory infections during the months of August to October correlate specifically with a chronic form of autoimmunity leading to disease. Using these findings, we have presented a newly designed celiac disease case-cohort study that will provide an excellent opportunity to investigate the role of microbes prospectively by analyzing longitudinally collected blood and stool samples from enrolment at age 3-months until celiac autoimmunity and disease develop. Analysis is currently ongoing to identify the celiac disease associated viruses by serology at regular 6-month intervals, by RT-PCR from 18,214 monthly stool samples and through the analysis of the bacteriome to identify celiac disease changes associated in the whole microbiome.
Previous work in TEDDY has examined how reported infections correlate with the risk of a child developing celiac disease autoimmunity at an early age. However, only GIEs and not RIEs correlated with the risk of CDA17. Despite studying 732 incident CDA cases in a total cohort population of 6,327, only GIEs reported in any 3-month period were significantly associated with an increased risk of CDA by the next quarterly visit. However, Kemppainen et al.17, did observe RIEs trending upwards 0–3 and 3–6 months prior to CDA seroconversion, and observed that the magnitude of the correlation between GIE and risk of CDA by the next visit was strongly dependent on other factors during the first year of life. This suggested the possibility of disease heterogeneity. In this study we excluded families whose child had developed diabetes autoimmunity before the age of 4 years. Additionally, we re-examined RIE and GIEs for correlations specifically for risk of celiac disease, i.e. CDA followed by biopsy-proven celiac disease diagnosis and, separately, for risk of CDA-only with no subsequent disease diagnosis. We found RIEs in any year, particularly when reported between August to October at any age during the first year of life, correlated with the onset of celiac disease before 3 years of age. This difference suggests that RIEs as early as the first year of life conditions a more severe type of autoimmunity which may make it easier to identify respiratory type of viruses as a significant trigger of celiac disease among younger children. Our findings adjusted for other strong TEDDY risk factors of CDA including HLA-DR3-DQ27, female predominance, non-HLA single nucleotide polymorphisms24 and a higher gluten intake during the first five years of life11. A small, nested metagenomics case–control study of fecal virome showed preliminary evidence of a cumulative effect of enterovirus infections and gluten on risk of CDA19. These results together suggest that the cumulative influence of infections during the early Autumn may be specific to a certain virus and may involve other factors that dictate how quickly CDA, and celiac disease develop.
Viruses that cause respiratory infections typically replicate on the respiratory mucosa. For example, the most common respiratory viruses, rhinoviruses, replicate mostly in the upper respiratory track, and some rhinovirus types also in lower respiratory track. However, some respiratory viruses can also replicate in the intestinal mucosa. The prime example is enteroviruses, which have associated with increased risk of celiac disease in previous studies19,32,33. These viruses replicate both in the respiratory and intestinal mucosa, and can spread to the submucosal immune system, including gut-associated lymphoid tissue (GALT). Thus, they may interact with gluten in the intestinal mucosa and GALT, which offers one plausible mechanism how they could contribute to the initiation of CD-onset at an early age when gluten consumption is rapidly increasing. The most common symptom of enterovirus infection is common cold while they only rarely cause gastroenteritis. Thus, in the present study, which was based on questionnaire data, enterovirus infections have mostly been classified as respiratory infection. They also peak at autumn time, in contrast to many other respiratory viruses. Enterovirus in the gut microbiome may present as pathogen-derived antigens and provoke inflammation early in life34. Another picornavirus, parechovirus, can also replicate in both respiratory and intestinal mucosa, and one prospective study has shown that detection of parechoviruses in stools was associated with increased risk of celiac disease35. A similar mechanism has been observed in Epstein-Barr virus infection and its association with multiple sclerosis, where persistent infection acts as a driver of autoimmune reactions and, subsequently, disease development36.
The need to add additional CDA cases in the future contributed to the decision to choose a case-cohort over a nested case–control study design. Nearly all children who develop celiac disease had started a GFD after CDA development. Moreover, none of the remaining CDA children with no diagnosis for celiac disease had started a GFD, Supplemental Fig. 4. Most parents were not aware of their child’s CDA development until at least after the persistent sample. Still, a small percentage of CDA children who started GFD were neither biopsied nor diagnosed with celiac disease. The reasons for starting a GFD diet were unknown. Furthermore, it was possible for CD-onset children to be determined with low to medium tTGA levels (< 60 U/ml) at CDA development and to be diagnosed with celiac diagnosis years later. It is less likely these children had celiac disease at the time of CDA development, and our preliminary analysis suggests diet and genetic factors may play an additional role after seroconversion. For this reason, we included a secondary outcome (CDA-hi), high tTGA levels on the persistent sample, for sensitivity analysis and shall use the outcome to better ensure any microbial associations found can be linked specifically and subsequently with onset of disease in young children and not because of reverse causation. Results associated with CD-onset will be confirmed with CDA-high.
As mentioned above, children developing IA before 4 years of age were excluded. Most of these children developed IA during the first two years of life and parents learned of the results by the next quarterly visit. Of the 423 who developed IA, 50 (11.8%) developed CDA and 20/50 were confirmed with celiac disease. Similar viral and bacterial studies have been conducted in TEDDY and results may help investigate the impact of their exclusion from MAVRiC.
The first objective of the MAVRiC longitudinal study will be to identify enteroviruses and bacteria that correlate especially with the onset of celiac disease and determine if they validate the infectious episodes reported from August to October and explain the correlations observed in this study. Of importance is discovering these infections independent of known risk factors. Because of the large scale of TEDDY, its international scope across four countries on two continents, its standardized methods for screening CDA and for collection of samples, this project is expected to produce universal findings that will clarify the inconsistencies in the literature, enhance understanding of the celiac disease process, and provide compelling data for future mechanistic studies of viral or microbial triggers in autoimmunity. We hypothesize that the longitudinal impact of infections and microorganisms on the risk of disease will likely depend on the child’s age of exposure, time of disease onset and timing relative to age of gluten introduction. Furthermore, identification of infectious triggers (or microbial dysbiosis) may lead to precise prevention strategies with vaccines and microbiome-enhancers.
In conclusion, evidence supports that the number of risk factors add up to increase risk of an earlier onset of celiac disease and the MAVRiC study will investigate the role of viruses as triggers at an early age with gut microbiome and diet as a time-dependent contribution.
Data availability
The datasets generated and analyzed during this current study are available at the NIDDK Central Repository (https:/doi.org/https://doi.org/10.58020/y3jk-x087).
Abbreviations
- CDA:
-
Celiac Disease Autoimmunity
- CI:
-
Confidence Interval
- GIE:
-
Gastrointestinal Infectious Episode
- HLA:
-
Human Leukocyte Antigen
- HR:
-
Hazard Ratio
- MAVRiC:
-
Microbial Associations and Viruses on the Risk of Celiac disease study
- RIE:
-
Respiratory Infectious Episode
- TEDDY:
-
The Environmental Determinants of Diabetes in the Young
- tTGA:
-
Tissue Transglutaminase Autoantibodies
References
Abadie, V., Sollid, L. M., Barreiro, L. B. & Jabri, B. Integration of genetic and immunological insights into a model of celiac disease pathogenesis. Annu. Rev. Immunol. 29, 493–525. https://doi.org/10.1146/annurev-immunol-040210-092915 (2011).
Parra-Medina, R. C., AC. . Celiac Disease In: Autoimmuity: From Bench to Bedside [Interet]. (El Rosario Universiy Press, 2013).
Singh, P. et al. Global Prevalence of Celiac Disease: Systematic Review and Meta-analysis. Clinical gastroenterology and hepatology : The official clinical practice journal of the American Gastroenterological Association 16, 823–836 e822 (2018). https://doi.org/10.1016/j.cgh.2017.06.037
Stahl, M. et al. Coeliac disease: What can we learn from prospective studies about disease risk?. Lancet Child Adolesc Health 8, 63–74. https://doi.org/10.1016/S2352-4642(23)00232-8 (2024).
Caio, G. et al. Celiac disease: A comprehensive current review. BMC Med 17, 142. https://doi.org/10.1186/s12916-019-1380-z (2019).
Stahl, M. et al. Incidence of Pediatric Celiac Disease Varies by Region. Am J Gastroenterol 118, 539–545 (2023). https://doi.org/10.14309/ajg.0000000000002056
Liu, E. et al. Risk of pediatric celiac disease according to HLA haplotype and country. N. Engl. J. Med. 371, 42–49. https://doi.org/10.1056/NEJMoa1313977 (2014).
Stordal, K. et al. Review article: Exposure to microbes and risk of coeliac disease. Aliment Pharmacol Ther 53, 43–62. https://doi.org/10.1111/apt.16161 (2021).
Raiteri, A. et al. Current guidelines for the management of celiac disease: A systematic review with comparative analysis. World J Gastroenterol 28, 154–175. https://doi.org/10.3748/wjg.v28.i1.154 (2022).
Meijer, C. R. et al. Prediction Models for Celiac Disease Development in Children From High-Risk Families: Data From the PreventCD Cohort. Gastroenterology 163, 426–436. https://doi.org/10.1053/j.gastro.2022.04.030 (2022).
Andren Aronsson, C. et al. Association of Gluten Intake During the First 5 Years of Life With Incidence of Celiac Disease Autoimmunity and Celiac Disease Among Children at Increased Risk. JAMA 322, 514–523 (2019). https://doi.org/10.1001/jama.2019.10329
Moscatelli, O. G. et al. Blood-Based T-Cell Diagnosis of Celiac Disease. Gastroenterology https://doi.org/10.1053/j.gastro.2025.05.022 (2025).
Group, T. S. The Environmental Determinants of Diabetes in the Young (TEDDY) Study. Ann N Y Acad Sci 1150, 1–13 (2008). https://doi.org/10.1196/annals.1447.062
Lernmark, A. et al. Looking back at the TEDDY study: Lessons and future directions. Nat Rev Endocrinol 21, 154–165. https://doi.org/10.1038/s41574-024-01045-0 (2025).
Rewers, M. et al. The Environmental Determinants of Diabetes in the Young (TEDDY) Study: 2018 Update. Curr Diab Rep 18, 136. https://doi.org/10.1007/s11892-018-1113-2 (2018).
Rewers, M. et al. Unfolding the Mystery of Autoimmunity: The Environmental Determinants of Diabetes in the Young (TEDDY) Study. Diabetes Care https://doi.org/10.2337/dc24-2886 (2025).
Kemppainen, K. M. et al. Factors That Increase Risk of Celiac Disease Autoimmunity After a Gastrointestinal Infection in Early Life. Clinical gastroenterology and hepatology : The official clinical practice journal of the American Gastroenterological Association 15, 694–702 e695 (2017). https://doi.org/10.1016/j.cgh.2016.10.033
Euren, A. et al. Risk of celiac disease autoimmunity is modified by interactions between CD247 and environmental exposures. Sci Rep 14, 25463. https://doi.org/10.1038/s41598-024-75496-w (2024).
Lindfors, K. et al. Metagenomics of the faecal virome indicate a cumulative effect of enterovirus and gluten amount on the risk of coeliac disease autoimmunity in genetically at risk children: The TEDDY study. Gut 69, 1416–1422. https://doi.org/10.1136/gutjnl-2019-319809 (2020).
Sharp, S. J., Poulaliou, M., Thompson, S. G., White, I. R. & Wood, A. M. A review of published analyses of case-cohort studies and recommendations for future reporting. PLoS ONE 9, e101176. https://doi.org/10.1371/journal.pone.0101176 (2014).
Kim, R. S. A new comparison of nested case-control and case-cohort designs and methods. Eur J Epidemiol 30, 197–207. https://doi.org/10.1007/s10654-014-9974-4 (2015).
Hill, I. D. et al. Guideline for the diagnosis and treatment of celiac disease in children: Recommendations of the North American Society for Pediatric Gastroenterology, Hepatology and Nutrition. J Pediatr Gastroenterol Nutr 40, 1–19. https://doi.org/10.1097/00005176-200501000-00001 (2005).
Lonnrot, M. et al. A method for reporting and classifying acute infectious diseases in a prospective study of young children: TEDDY. BMC Pediatr 15, 24. https://doi.org/10.1186/s12887-015-0333-8 (2015).
Sharma, A. et al. Identification of Non-HLA Genes Associated with Celiac Disease and Country-Specific Differences in a Large. International Pediatric Cohort. PLoS One 11, e0152476. https://doi.org/10.1371/journal.pone.0152476 (2016).
Aronsson, C. A. et al. Age at gluten introduction and risk of celiac disease. Pediatrics 135, 239–245. https://doi.org/10.1542/peds.2014-1787 (2015).
Kemppainen, K. M. et al. Association Between Early-Life Antibiotic Use and the Risk of Islet or Celiac Disease Autoimmunity. JAMA Pediatr 171, 1217–1225. https://doi.org/10.1001/jamapediatrics.2017.2905 (2017).
Johnson, S. B. et al. Predicting Later Study Withdrawal in Participants Active in a Longitudinal Birth Cohort Study for 1 Year: The TEDDY Study. J Pediatr Psychol 41, 373–383. https://doi.org/10.1093/jpepsy/jsv092 (2016).
Johnson, S. B. et al. The Environmental Determinants of Diabetes in the Young (TEDDY) study: Predictors of early study withdrawal among participants with no family history of type 1 diabetes. Pediatr Diabetes 12, 165–171. https://doi.org/10.1111/j.1399-5448.2010.00686.x (2011).
Prentice, R. L. On the design of synthetic case-control studies. Biometrics 42, 301–310 (1986).
Barlow, W. E. Robust variance estimation for the case-cohort design. Biometrics 50, 1064–1072 (1994).
Hard Af Segerstad, E. M. et al. Early Dietary Fiber Intake Reduces Celiac Disease Risk in Genetically Prone Children: Insights From the TEDDY Study. Gastroenterology 168, 1185–1188 e1182 (2025). https://doi.org/10.1053/j.gastro.2025.01.241
Oikarinen, M. et al. Enterovirus Infections Are Associated With the Development of Celiac Disease in a Birth Cohort Study. Front Immunol 11, 604529. https://doi.org/10.3389/fimmu.2020.604529 (2020).
Kahrs, C. R. et al. Enterovirus as trigger of coeliac disease: Nested case-control study within prospective birth cohort. BMJ 364, l231. https://doi.org/10.1136/bmj.l231 (2019).
Iversen, R. & Sollid, L. M. The Immunobiology and Pathogenesis of Celiac Disease. Annu Rev Pathol 18, 47–70. https://doi.org/10.1146/annurev-pathmechdis-031521-032634 (2023).
Tapia, G. et al. Parechovirus Infection in Early Childhood and Association With Subsequent Celiac Disease. Am J Gastroenterol (2020). https://doi.org/10.14309/ajg.0000000000001003
Bjornevik, K. et al. Longitudinal analysis reveals high prevalence of Epstein-Barr virus associated with multiple sclerosis. Science 375, 296–301. https://doi.org/10.1126/science.abj8222 (2022).
Funding
This study was funded by the National Institute of Health (NIH/NIDDK 1 R01 DK124581-01) and the Swedish Research Council (grant 2022–00537). The TEDDY Study is funded by U01 DK63829, U01 DK63861, U01 DK63821, U01 DK63865, U01 DK63863, U01 DK63836, U01 DK63790, UC4 DK63829, UC4 DK63861, UC4 DK63821, UC4 DK63865, UC4 DK63863, UC4 DK63836, UC4 DK95300, UC4 DK100238, UC4 DK106955, UC4 DK112243, UC4 DK117483, U01 DK124166, U01 DK128847, and Contract No. HHSN267200700014C from the National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK), National Institute of Allergy and Infectious Diseases (NIAID), Eunice Kennedy Shriver National Institute of Child Health and Human Development (NICHD), National Institute of Environmental Health Sciences (NIEHS), Centers for Disease Control and Prevention (CDC), and Breakthrough T1D (formerly JDRF). This work is supported in part by the NIH/NCATS Clinical and Translational Science Awards to the University of Florida (UL1 TR000064) and the University of Colorado (UL1 TR002535). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
KFL: Concept and Design, statistical analysis, interpretation of the data, drafting the manuscript, final approval of the version to be published. EWT: Concept and Design, interpretation of the data, drafting the manuscript, critical revision of the manuscript for important intellectual content, final approval of the version to be published. HH: Concept and Design, interpretation of the data, drafting the manuscript, critical revision of the manuscript for important intellectual content, final approval of the version to be published. APA: Concept and Design, interpretation of the data, drafting the manuscript, critical revision of the manuscript for important intellectual content, final approval of the version to be published. JEL: Interpretation of the data, drafting the manuscript, critical revision of the manuscript for important intellectual content, final approval of the version to be published. JFP: Interpretation of the data, critical revision of the manuscript for important intellectual content, final approval of the version to be published. REL: Concept and Design, interpretation of the data, drafting the manuscript, critical revision of the manuscript for important intellectual content, final approval of the version to be published. DA: Concept and Design, interpretation of the data, drafting the manuscript, critical revision of the manuscript for important intellectual content, final approval of the version to be published.
Corresponding author
Ethics declarations
Competing interests
DA receives consultant fees as member of Sanofi´s scientific advisory board. HH is a stock owner and member of the board of Vactech Oy which develops vaccines against picornaviruses. KFL, EWT, APA, JEL, JFP and REL do not have any conflict of interest to declare.
Ethical approval
Written informed consent was obtained from all families. All research was performed in accordance with relevant regulations, and the study was performed in accordance with the Declaration of Helsinki. The study design and recruitment was approved by local U.S. Institutional Review Boards and European Ethics Committee Boards in Colorado’s Colorado Multiple Institutional Review Board 04–0361, Georgia’s Medical College of Georgia Human Assurance Committee (2004–2010), Georgia Health Sciences University Human Assurance Committee (2011–2012), Georgia Regents University Institutional Review Board (2013–2016), Augusta University Institutional Review Board (2017-present) HAC 0405380, Florida’s University of Florida Health Center Institutional Review Board IRB201600277, Washington state’s Washington State Institutional Review Board (2004–2012) and Western Institutional Review Board (2013–2019), WCG IRB (2020-present) 20130211, Finland’s Ethics Committee of the Hospital District of Southwest Finland Dnro168/2004, Germany’s Bayerischen Landesärztekammer (Bavarian Medical Association) Ethics Committee 04089, Sweden’s Regional Ethics Board in Lund, Sect. 2 (2004–2012) and Lund University Committee for Continuing Ethical Review (2013–2021), Swedish Ethical Review Authority (2022-present) 217/2004 and the study is monitored by External Advisory Board formed by the National Institutes of Health.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lynch, K.F., Triplett, E.W., Hyöty, H. et al. Microbial associations and viruses on the risk of celiac disease (MAVRiC): a longitudinal post-hoc case-cohort study. Sci Rep 15, 42704 (2025). https://doi.org/10.1038/s41598-025-26700-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-26700-y