Abstract
The CRISPR-Cas systems are supposed to be associated with antibiotic susceptibility. Klebsiella pneumoniae (K. pneumoniae) is a major multidrug-resistant (MDR) pathogen that may cause severe infections. This study aimed to detect the antimicrobial resistance (AMR) of K. pneumoniae isolates in addition to determine the association between the existence of CRISPR-Cas systems and the presence of AMR in Egypt. The antibiotic susceptibility patterns of 100 K. pneumoniae isolates were determined using the Kirby Bauer disc diffusion and broth microdilution methods. The frequency of carbapenem resistance encoding genes (blaKPC, blaOXA, blaIMP, blaNDM, and blaVIM), ESBLs encoding genes (blaTEM), aminoglycoside resistance encoding genes (aac(3)-Ia, aac(3)-IIa, colistin resistance encoding genes (mcr-1, mcr-2) and tetracycline resistance encoding genes (tetB) were determined using polymerase chain reaction (PCR). The presence or absence of CRISPR-Cas systems was determined by detection of Cas genes (Cas1 or Cas3) in conjunction with one of CRISPR arrays 1, 2 or 3. Kirby Bauer disc diffusion revealed that 95% of isolates were MDR. The resistance rates of K. pneumoniae isolates to amikacin, meropenem, and colistin were 76%, 67%, and 41% respectively by broth microdilution assay. Among selected 41 K. pneumoniae, the frequency of ESBLs; blaTEM was 92.7%, while the frequency of blaNDM blaOXA blaVIM blaIMP and blaKPC was 95.1%, 95.1%, 39%, 19.5% and 14.6% respectively. The frequency of mcr-1 and mcr-2 was 70.7% and 65.9%. Additionally, the frequency of aac(3)-Ia was 12.2%, and aac(3)-IIa was 87.8%, while the frequency of tetB was 100%. Our isolates exhibited varied profiles for CRISPR-Cas systems, where 65.9% were positive for CRISPR-Cas system. No general significant positive correlation between AMR and the presence ofCRISPR-Cas system was detected; however, a significant difference is present for imipenem, colistin and chloramphenicol phenotypic resistance andfor aac(3)-IIa and mcr-1 genes (P value > 0.1). A significant positive correlation was detected between AMR for imipenem, colistin and chloramphenicol and for aac(3)-IIa and mcr-1 genes and thepresence of CRISPR-Cas system.
Similar content being viewed by others
Introduction
The clustered regularly interspaced short palindromic repeats (CRISPRs) and CRISPR-associated (Cas) genes/proteins, are part of the adaptive immune system in prokaryotes and found in roughly 50% of bacterial genomes and 87% of archaea1. CRISPR-Cas adaptive immune systems are one of the most prevalent defenses in bacteria but their role in limiting the spread of antimicrobial resistance genes (ARGs) is not well-understood2.
Makarova and colleagues in 2011 introduced a system for categorizing CRISPR-Cas systems into three primary types: type I (identified by the cas3 gene), type II (characterized by the cas9 gene), and type III (distinguished by the cas10 gene), which has since become the standard terminology in the field3. Cas enzymes utilize CRISPR sequences to identify and cleave the complementary strands of target DNA. CRISPR arrays enable bacteria or archaea to retain information about previous viral encounters, with viral DNA copies being integrated into their genomes as “spacers” between the short DNA repeats of CRISPR arrays when encountering virus4.
Several studies have established a correlation between CRISPR-Cas systems and increased susceptibility to antibiotics, as well as reduced plasmid acquisition in various bacterial species. Some research also suggests that these systems may play a role in regulating bacterial pathogenicity5. For instance, the CRISPR-Cas system influences the contents of prophages in Streptococcus pyogenes, impacting its virulence6. Additionally, Mackow et al. found that Klebsiella pneumoniae with CRISPR-Cas systems exhibited heightened sensitivity to carbapenems7.
Conversely, some studies indicate that bacteria can evolve mechanisms to bypass the CRISPR-Cas system’s effects, such as acquiring mutations that disable its function8,9. Therefore, there is significant potential for further research into the impact of CRISPR-Cas systems on the virulence and antibiotic resistance of various bacterial pathogens.
Antimicrobial resistance carries significant social and economic repercussions, leading to prolonged sickness, extended hospitalization, and the requirement for more expensive medications, with a disproportionate impact on lower- and middle-income nations10. The primary catalysts behind the rise of MDR in bacteria stem from the inappropriate and excessive use of antibiotics11. Many microbial elements contribute to the development of AMR. Microbial factors such as genomic plasticity, genetic exchanges (e.g., horizontal genetic transfer [HGT]), mutations, enzyme modifications, degradative enzymes, changes in pathways, and activation of efflux pumps play a crucial role in the emergence and rise of AMR12. A pan-genome refers to a collection of genes that collectively make up a certain bacterial species13. A pan-genome is divided into three sections: the core (persistent) genome, which is shared by at least 95% of bacterial strains, the shell (accessory) genome, which is shared by at least 15% but less than 95% of strains, and the cloud (strain-specific/unique/singleton) genes, which are present in less than 15% of bacterial strains13,14. According to genomic studies, K. pneumoniae’s pan-genome is around five to six Mbp in size, with five to six kilogenes to be encoded. Of these encodable genes, around seventeen hundred are classified as core genes. However, the remaining genetic pool is the accessory genome which differs between Klebsiella species15.
Currently, K. pneumoniae has demonstrated resistance to major effective antibiotic categories like carbapenems, cephalosporins, and aminoglycosides, complicating antibiotic treatments and resulting in treatment failures16. Several AMR genes, such as blaKPC, blaOXA−48, and blaNDM−1, were initially identified in K. pneumoniae before dissemination to other pathogens17. Multidrug-resistant bacteria evade the effects of most antibiotics through successive genetic changes and the transfer of mobile genetic elements, highlighting the urgent necessity to investigate resistance development and prevention strategies18. The CRISPR-Cas system has been proposed as a novel tool to tackle antibiotic resistance, presenting a promising avenue for developing next-generation antimicrobials to address infectious diseases19,20.
In the current study, we aimed to detect the prevalence of AMR detected phenotypically and genotypically and CRISPR-Cas encoding genes in MDR K. pneumoniae clinical isolates and analyze the correlation between the presence of CRISPR-Cas system and prevalence of antimicrobial resistance to different antibiotics as of β-lactam, aminoglycoside, tetracycline and colistin particularly addressing a research gap in the Middle Eastern context regarding this association.
Methods
Study design and identification of bacterial isolates
All experiments were carried out at the research lab of the Microbiology & Immunology Department, Faculty of Pharmacy, Al-Azhar University, Cairo, Egypt, and the Genomics Lab at Biotechnology Department, Faculty of Agriculture, Al-Azhar University, Cairo, Egypt, during the period from October 2020 to June 2023. A total number of 100 clinical isolates of K. pneumoniae isolated from different clinical specimens (presented in supplementary file in Fig.S1) were obtained from the Microbiology lab at Al-Demerdash Hospital, Ain Shams University, Cairo-Egypt.
K. pneumoniae isolates were identified biochemically using standard microbiological tests as growth on triple sugar iron agar, Simmon’s citrate agar, indole test, urease tests21. etc. Identification of K. pneumonia was confirmed using the Vitek-2 automated system (Biomérieux, Marcy-LÉtoile, Paris, France) following the manufacturer’s instructions. All K. pneumoniae isolates were preserved at − 20 °C in brain–heart infusion broth with 20% glycerol (Oxoid, London, UK) until use22.
Antimicrobial susceptibility testing using Kirby–Bauer disc diffusion method
The Kirby–Bauer disc diffusion method was used to detect the antibiotic susceptibility of all identified K. pneumoniae isolates on Muller–Hinton agar (LAB M Limited, UK) according to the Clinical and Laboratory Standards Institute (CLSI 2023)23. All antibiotic discs were supplied by Oxoid, London, UK and included different categories amoxicillin/clavulanic acid (20/10 µg), piperacillin/tazobactam (100/10 µg) [penicillins and beta-lactamase inhibitors], ceftriaxone (30 µg), ceftazidime (30 µg), cefuroxime (30 µg), cefoxitin(30 µg) [cephalosporins], imipenem(10 µg), meropenem (10 µg) [carbapenem], ciprofloxacin (5 µg) [fluoroquinolones], gentamicin (10 µg), amikacin (30 µg) [aminoglycosides], chloramphenicol (30 µg)[glycols], and colistin (10 µg) [polymyxin]24. K pneumoniae ATCC700603 was used as positive control. The results of the disc diffusion assay were interpreted according to CLSI guidelines23. MDR K. pneumoniae isolates were identified as non-susceptibility to at least one agent in three antimicrobial classes25.
Antimicrobial susceptibility testing using broth microdilution assay
The MIC for amikacin, meropenem, and colistin was determined using the broth microdilution assay according to the guidelines of the CLSI 202323. Briefly, a 96-well microtiter plate was filled with 100 µL of double-strength MHB medium (LAB M Limited, UK). Then 100 µL of working antibiotic solution was added in each well of the first row of the microtiter plate. Two-fold serial dilution (from 128 to 0.5 µg/mL) was performed by transferring 100 µL of antibiotic from the first well to the subsequent wells of the plate. Fresh bacterial colonies were adjusted to a turbidity equivalent of 0.5 McFarland standard, and each well was inoculated with 7 µL of fresh K. pneumoniae inoculum (1.5 × 108) to reach final inoculum density of 105 CFU ml− 1 per wells26. The plate was incubated for 24 h at 37 °C. After incubation, the MIC was determined as the lowest concentration of antibiotic which showed no visible growth. The results of amikacin and meropenem were interpreted according to the guidelines of CLSI, the breakpoints for AMC were (Sensitive(S) ≤ 16 µg/mL, Intermediate (I) = 32, and Resistant (R) ≥ 64 µg/mL), and for MEM were (S ≤ 1 µg/mL, I = 2, R ≥ 4 µg/mL), while the results of colistin were interpreted according to the European Committee on Antimicrobial Susceptibility Testing27. The breakpoints for COL were (S ≤ 2 µg/mL, and R > 2 µg/mL).
DNA extraction and primer design
Total genomic DNA was extracted using G-spinTM Total DNA Extraction Kit, (Intron Biotechnology, Keonggi-do, South Korea) according to the manufacturer’s instructions. The DNA was stored at − 20 °C. Primers for ESBLs encoding genes; blaTEM (the most commonly found in community and livestock environments)28, carbapenem resistance encoding genes (blaKPC, blaOXA, blaIMP, blaNDM and blaVIM), aminoglycosides resistance encoding genes (aac(3)-Ia and aac(3)-IIa), colistin resistance encoding genes (mcr-1 and mcr-2) and tetracycline resistance encoding genes (tetB), were designed in the current study according to conserved region of the Klebsiella pneumoniae subsp. pneumoniae HS11286. The primers specificity was checked using Primer BLAST tools in NCBI https://www.ncbi.nlm.nih.gov/tools/primer-blast/.
The CRISPR-Cas system encoding genes including CRISPR1, CRISPR2, CRISPR3, Cas1, and Cas3 genes were chosen using the previously published specific primers29. All primers are listed in Table 1.
Molecular detection of resistance genes and CRISPER-CAS system by the polymerase chain reaction
PCR reaction mixture was performed in volume of 25 µL, including 10 µL of PCR master mix (Williford, Nottinghamshire NG, England), 1 µL of each specific primer (25 nanomoles), 5 µL (250 ng) of DNA template and 8 µL of RNAse free distilled water30. The timetable and thermal schedule for each gene are presented in Supplementary Table 1. The amplified products were run on 1% agarose gel (Genetix Biotech, New Delhi, India) stained with ethidium bromide (Bioshop, Ontario, Canada) and photographed under UV illumination. A 100–1000 base-pairs standard DNA ladder (Bengaluru, Karnataka, India) was used for sizing the PCR products31.
Statistical analysis
Statistical analysis was performed using IBM SPSS® Statistics version 26 (IBM® Corp., Armonk, NY, USA). Qualitative data were expressed as frequency and percentage. Pearson’s Chi-square test or Fisher’s exact test was used to examine the relation between qualitative variables. All tests were two-tailed. A p-value < 0.05 was considered significant. Flow chart that clarifies all performed methods is presented in Fig. 1.
Flow chart showing study design and methodology.
Results
Results of antimicrobial susceptibility using Kirby–Bauer disc diffusion method
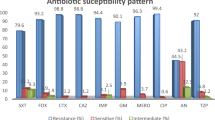
Regarding individual antibiotic susceptibility testing, 98% of isolates were resistant to amoxicillin/clavulanic acid and cefuroxime, while 97% of isolates were resistant to ceftriaxone followed by ceftazidime (94%), piperacillin-tazobactam (91%), and cefoxitin (90%). Also, 82% of K pneumoniae isolates were resistant to ciprofloxacin and gentamicin. The resistance rate to meropenem, imipenem, and colistin were 78%, 75%, and 67% respectively. On the other hand, only 46% of K. pneumoniae isolates showed resistance to chloramphenicol as shown in Fig. 2 and Supplementary Table 2. The results of the Kirby–Bauer disc diffusion method revealed that 95% of k. pneumoniae isolates were MDR.
The frequency of antimicrobial resistance of k. pneumoniae isolates by Kirby–Bauer disc diffusion method.
Results of antimicrobial susceptibility using broth microdilution assay
The MIC results of amikacin, meropenem, and colistin for the 100 k. pneumoniae isolates are shown in (Table 2). The resistance rates to amikacin, meropenem and colistin were 76%, 67%, and 41% respectively while 8% and 2% of isolates have intermediate sensitivity to meropenem and amikacin, respectively.
Molecular detection of antimicrobial resistance encoding genes via PCR
Forty-one isolates were selected for molecular identification including 18 isolates that showed MDR profile by Kirby–Bauer disc diffusion method and broth microdilution assay, 5 isolates that were sensitive to amikacin, meropenem, and colistin by Kirby–Bauer disc diffusion method, 6 isolates were sensitive to 2 antibiotics by broth microdilution assay, 7 isolates were sensitive to only 1 antibiotic by broth microdilution assay and five isolates were sensitive to most antibiotic groups.
The analysis of resistance encoding genes revealed a high prevalence of ESBLs, blaTEM gene which was detected in 92.7% isolates. Furthermore, genes conferring carbapenem resistance, specifically the blaNDM and blaOXA, were predominant, identified in 95.1% of isolates. Other carbapenemase genes, including blaVIM, blaIMP and blaKPC were present in 39%, 19.5%, and 14.6% of the total isolates, respectively. Regarding colistin resistance, the mcr-1 and mcr-2 genes were observed in 70.7% and 65.9% of isolates, respectively. The prevalence of aminoglycoside resistance-encoding genes, aac(3)-Ia and aac(3)-IIa, was 12.2% and 87.8%, respectively. Notably, the tetracycline resistance-encoding gene tetB was ubiquitously detected across all isolates. All PCR results are presented in Table 3 and all PCR gel pictures are presented in supplementary file (figures S2-S17).
Genotypic profiles of AMR genes among K. pneumoniae isolates are presented in Table 4 and revealed that blaOXA, blaTEM, blaVIM, mcr-1, mcr-2, aac(3)-IIa- tetB genetic profile was harbored by 21.9% of K. pneumoniae isolates as shown in Table 4.
The distribution of AMR-encoding genes with corresponding AMR phenotypes among K pneumoniae isolates is demonstrated in Table 5. The matching revealed similar phenotypic susceptibility patterns of amoxicillin-clavulanate, ceftriaxone, cefoxitin, cefuroxime, ceftazidime, piperacillin tazobactam, imipenem, meropenem, gentamycin, and amikacin the isolates versus coexisting resistance encoding genes (blaNDM, blaOXA, blaIMP, blaKPC, blaVIM, and aac(3)-Ia) and resistance pattern. while there was a different distribution pattern for blaTEM, and resistance pattern to amoxicillin–clavulanate, ceftriaxone, cefoxitin, cefuroxime, ceftazidime, piperacillin tazobactam, imipenem, meropenem. There was a different distribution pattern for mcr-1 and mcr-2 resistance encoding genes and resistance pattern to colistin and for aac(3)-IIa and resistance pattern to gentamycin and amikacin.
Genotypic profiles of CRISPER-CAS system encoding genes among K. pneumoniae isolates
PCR for detecting CRISPR1, CRISPR2, CRISPR3, Cas1 and Cas3, revealed that 21(51.2%) harbored the CRISPR1 gene, 19(46.3) harbored the CRISPR2 gene, 26(63.4%) harbored the CRISPR3 gene, 28(68.3%) of isolates harbored the Cas1 gene and 23(56.1%) harbored the Cas3 gene (Table 6).
The presence or absence of CRISPR-Cas systems was shown where cas genes (cas1 or cas3) are present in conjunction with one of CRISPR arrays (CRISPR1, CRISPR2, CRISPR3). Out of 41 K. pneumoniae isolates, 27 (65.9%) were CRISPR-Cas-positive while 14 (34.1%) were.
CRISPR-Cas-negative. The distribution of different genotypes of CRISPR-Cas system is shown in Fig. 3.
The distribution of different genotypes of CRISPR-Cas system.
Genotypic profiles of CRISPR-Cas system encoding genes among K. pneumoniae isolates are presented in Table 7 and revealed that 21.9% of the isolates coharbored CRISPR1, CRISPR2, CRISPR3, Cas1, and Cas3.
Association between AMR detected by phenotypic Kirby–Bauer disc diffusion method and prevalence of the CRISPR-Cas system encoding genes
The AMR detected by Kirby–Bauer disc diffusion method among K. pneumoniae isolates was clearly higher in CRISPR-Cas-positive isolates compared to CRISPR-Cas-negative isolates. There was a non-significant positive correlation between the presence of CRISPR-Cas system and the existence of resistance in K. pneumoniae by Kirby–Bauer disc diffusion method (Pearson Chi-Square test, p- values = 0.1) (Table 8).
CRISPR-Cas-positive isolates exhibited resistance to a broader range of antimicrobial agents, including amoxicillin-clavulanate (100%), cefuroxime (100%), ceftriaxone (96.3%), ceftazidime (96.3%), piperacillin-tazobactam (88.9%), cefoxitin (88.9%), ciprofloxacin (88.9%), gentamicin (77.8%), amikacin (66.7%), meropenem (66.7%), imipenem (81.5%), colistin (85.1%), and chloramphenicol (81.5%). There was a significant difference of AMR of imipenem, colistin and chloramphenicol (Pearson Chi-Square test, p-values = 0.047, 0.005, and 0.012, respectively) among CRISPR-Cas-positive and CRISPR-Cas - negative K. pneumoniae isolates as shown in Fig. 4.
The prevalence of AMR detected by Kirby–Bauer disc diffusion method, among CRISPR-Cas-positive and -negative K. pneumoniae isolates.
Association between AMR encoding genes detected by PCR and prevalence of the CRISPR-Cas system encoding genes
The percentage of K. pneumoniae isolates harboring AMR encoding genes was clearly higher in the CRISPR-Cas-positive isolates than in the CRISPR-Cas-negative isolates except for blaNDM, blaOXA and blaIMP encoding genes. The percentage of presence of blaNDM,, blaOXA and blaIMP in CRISPR-Cas-positive isolates was 92.6%, 92.6%, and 14.8%, respectively while it was 100%, 100% and 28.6%, respectively in CRISPR-Cas-negative isolates.
For other AMR encoding genes CRISPR-Cas-positive K. pneumoniae isolates exhibited higher percentage of resistant genes compared to CRISPR-Cas-negative isolates. The presence of blaTEM, aac(3)-IIa, mcr-1, mcr-2, blaVIM, blaKPC, and aac(3)-Ia genes was observed in 100%, 96.3%, 81.5%, 66.7%, 44.4%, 22.2%, and 14.8% of isolates, respectively.
There was a significant difference (p-value < 0.05) of AMR encoding genes; aac(3)-IIa and mcr-1 (Pearson Chi-Square test, p-values = 0.039 and 0.036, respectively) among CRISPR-Cas-positive and CRISPR-Cas - negative K. pneumoniae isolates which were found in 96.3% and 81.5% of the CRISPR-Cas positive isolates, respectively versus 71.4% and 50% of the CRISPR-Cas negative isolate as shown in Fig. 5.
The prevalence of AMR encoding genes among the CRISPR-Cas-positive and CRISPR-Cas-negative K. pneumoniae isolates.
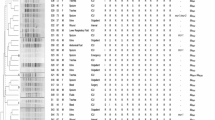
Phenotypic resistance and susceptibility to the tested antimicrobial agents, together with the presence or absence of the ARGs and their correlation with the presence of CRISPR-Cas system are displayed in a heatmap (Fig. 6); the isolates carrying CRISPR-Cas are in upper panel while CRISPR-Cas negative isolates are in lower panel.
A heatmap displaying the AMR profiles and genetic determinants among the isolates carrying CRISPR-Cas (Upper panel) versus CRISPR-Cas negative (Lower panel) Red and pink colors indicate the presence and absence of the ARGs, respectively; Dark and light blue colors indicate phenotypic resistance and susceptibility to the tested antimicrobial agents, respectively.
Discussion
K. pneumoniae is among the top pathogens causing nosocomial infections worldwide32. K. pneumoniae’s pan-genome is around five to six Mbp in size15, The size and fluidity of the accessory genome are key indicators of a species’ adaptability. Species with a large, “open” pan-genome (like E. coli or Klebsiella pneumoniae) are highly versatile and can rapidly acquire new traits, which is why they can be significant threats in clinical and environmental settings. In contrast, species with a smaller, “closed” pan-genome tend to be more specialized and have less genetic diversity33.
K. pneumoniae has become increasingly resistant to antibiotics, making antibiotic therapy more challenging, several antimicrobial resistance genes were discovered in K. pneumoniae for the first time before spreading to other pathogens e.g. blaKPC, blaOXA−48, and blaNDM−34.
The CRISPR-Cas systems are involved in limiting the entry of foreign DNA to bacteria and archaea and have also been related to the expression of virulence factors in bacteria32. In the current study, we aimed to detect the prevalence of CRISPR-Cas encoding genes in MDR clinical isolates of K. pneumoniae and analyze the correlation between the presence of CRISPR-Cas system and prevalence of the AMR detected by phenotypic and molecular techniques as limited researches are being carried out in the Middle East to address this correlation.
In the current study, antimicrobial susceptibility testing by Kirby–Bauer disc diffusion methods revealed that 95% of K. pneumoniae isolates were MDR. The results of this study were in accordance with two studies who found that 90.2% and 94% of K. pneumoniae isolates were MDR35,36. Previous studies conducted in Egypt found that the rate of MDR Gram-negative isolates was 85.6%, 66.5% and 93.6%37,38,39. Moreover, a recent study conducted in Egypt revealed that the rate of MDR Gram-negative isolates was 98.2% that was higher than the results of this study40.
K. pneumoniae isolates in the current study showed high AMR against amoxicillin-clavulanate (98%) and piperacillin-tazobactam (91%), 2nd and 3rd generation cephalosporins, cefoxitin (90%), cefuroxime (98%), ceftriaxone (97%), and ceftazidime (94%). High resistance rates to these antimicrobials were also observed in other studies41,42.
Also, considerable carbapenem resistance was detected in the current study, the resistance rate to meropenem and imipenem was 78% and 75%, respectively. This is higher than a previous study conducted in Egypt, which reported a resistance rate of 50% to meropenem43. Another study in Egypt showed that the resistance rate was 31.3% and 30% to imipenem and meropenem, which was lower than this study44. Likewise, Baral et al. (2024) from Nepal showed that 61.1% and 69.4% of K. pneumoniae isolates were resistant to meropenem and imipenem respectively, which is comparable to our results45.
Chloramphenicol is no longer the drug of choice for many infections. Its usage in the developed world is limited to life-threatening infections without safer alternatives. However, it is still commonly used in many parts of the developing world46.
In the current study, K. pneumoniae isolates showed moderate incidence of resistance to chloramphenicol (46%), this is consistent with the findings of a study from Egypt reported that the frequency of resistance to chloramphenicol was 54%36.
Microbroth dilution results of meropenem, amikacin reveled that the percentage of meropenem resistance was 67% which was lower than that found in other studies reported 100% resistance to meropenem by MIC47,48. The resistance rate to amikacin by MIC was 76%, this result was comparable to a study showed a resistance of 50% to amikacin49. The moderate resistance to amikacin may be due to its less frequent use as empirical therapy and the absence of considerable cross-resistance with β-lactam antimicrobials50.
Although colistin is now used as a “last resort” for treating infections caused by some of the most virulent MDR Gram-negative bacteria51. Determination of MIC provide more accurate results for colistin susceptibility testing compared to disc diffusion methods because colistin is a large molecule that does not diffuse well in agar, leading to inconsistent results52. The K. pneumoniae isolates in the current investigation had a significant incidence of colistin resistance (41%) by determination of MIC. Several studies conducted in Egypt reported lower rates of colistin resistance that were 4.9%, 7.5 and 17.2%53,54,55. This resistance rate could be because of colistin therapy that was discontinued many years ago, is now used as a last-resort antibiotic for carbapenem resistant K. pneumoniae infections.
The high MDR rate could be a result of misuse and abuse of antibiotics in Egypt. The increasing rate of these isolates emphasizes the importance of choosing an appropriate antimicrobial regimen based on antibiotic susceptibility testing56. Numerous causes, including the use of antibiotics in healthcare facilities, the community, farm animals, agriculture, and the environment, all contribute to the rise of antibiotic resistance. Antibiotics are overused because they may be purchased easily over the counter without a prescription57. Thus, antibiotic usage must be monitored closely to reduce antibiotic misuse, as antibiotic resistance is spreading quickly58.
In the current study, forty-one isolates were selected for molecular investigation where the prevalence of ESBL encoding gene; blaTEM was 92.7%. Other studies revealed that the frequency of blaTEM was 100% and 52%59,60. ESBL-producing Enterobacteriaceae carry a broad-spectrum beta-lactamase enzyme that enables them to become resistant to nearly all antibiotics in the penicillin and cephalosporin classes. In such cases, the remaining treatment option is an antibiotic from the carbapenem family, which contributes to the development of carbapenem resistance61.
The current study reported higher frequencies of carbapenem resistance encoding genes among K. pneumoniae isolates, including blaNDM and blaOXA−48 genes (95.1%). These results were significantly higher than several studies including a study from Egypt reported that the incidence of blaNDM and blaOXA−48 was 70% and 52%, respectively62. Another study found that 66.67% and 62.50% of carbapenem resistant Enterobacteriaceae isolates possessed blaOXA−48 and blaNDM, respectively63. In contrast, the prevalence of blaVIM, blaIMP, and blaKPC in this study was 39%, 19.5%, and 14.6%, respectively. Similar to our results a study in Egypt showed that the prevalence of blaVIM and blaIMP were 11.1% and 0%, respectively64. Another study in Egypt, revealed that blaOXA−48 was the prevalent gene (15.5%), followed by blaVIM (15%), blaIMP (7.5%), blaKPC (4%), and blaNDM (3.8%).
The rationale behind the expression-discrepancy of carbapenem resistance genes with crossponding phenotypes is not well-understood65. It is possible that the antibiotic itself may modulate the ARGs into low in vitro expression66, or that the heteroresistance phenomena associated with unstable tandem gene amplification, or rare mutation such as frameshift mutations associated with upregulation of the MexXY-OprM pump, loss of OprD expression67 and environmental modulation to the resistant genes, all these factors may explain the exhibited phenotypic carbapenem susceptibilities65.
Untreatable infections due to carbapenem resistant K. pneumoniae are on the rise among patients in medical facilities68. With the increasing use of carbapenems in clinical practice, the emergence of carbapenem-resistant pathogens now poses a great threat to human health. Currently, antibiotic options for the treatment of carbapenem resistant K. pneumoniae are very limited69.
In this study, aac(3)-IIa was the most prevalent aminoglycoside resistance encoding gene at 87.8%, followed by aac(3)-Ia at 12.2%. The combination of both genes was observed in four isolates. One study reported a lower frequency for the two aac (3)-II genes that was 79.3% and 64%, respectively70. In contrast, Kadkhoda et al. (2024) found that the percentage of aac(3)-Ia was 29.5% while aac(3)-IIa genes weren’t detected71.
The mcr-1 gene is the most commonly to cause colistin resistance in humans, also mcr-2, has a role in the development of colistin resistance72.
The results of the current study revealed that 70.7% of the K. pneumoniae isolates were harboring mcr-1 gene while 65.9% were harboring mcr-2 gene. This high prevalence rate of mcr-1 and mcr-2 genes was in accordance with the high resistance detected by the phenotypic determination of MIC (41%). One study from Egypt reported that 84.4% of their K. pneumoniae isolates harbored mcr-1 encoding gene72. On the other hand, many studies revealed lower rates of mcr-1 which were 4%, 4.2% and 5%, respectively and no one of their isolates had mcr-253,73,74. The mcr-1 and mcr-2 genes dissemination makes it necessary to search for alternatives to substitute ineffective antibiotics72.
Due to dissemination of tetracycline resistance in our region75,76 the tetracycline resistance gene (tet B) was included in our study as a key molecular marker to investigate the mobility of resistance determinants and their relationship with the CRISPR-Cas system that is separate from the phenotypic antimicrobial susceptibility testing as it is not a first-line or routinely used therapeutic option for treating infections caused by MDR K. pneumoniae isolates.
The tetB gene generally was reported as the most common gene among tetracycline-resistant Enterobacteriaceae77. In the present study, tetB gene was detected in 100% of isolates. Other studies detected tetB gene in 42% and 18.4% in their K. pneumoniae isolates78,79.
These findings show a remarkable frequency of tetB gene distribution and subsequent tetracycline resistance. This may be due to long-term and widespread use of tetracycline in animal farms and agriculture for growth promotion purposes. Also, the tet genes are located on highly mobile genetic elements as plasmids, transposons, and conjugative transposons which are responsible for the horizontal transfer of ARGs, contributing to the widespread distribution of tetracycline resistance in K. pneumoniae80.
Pearson’s Chi-square test or Fisher’s exact test are both used to compare observed and expected frequencies in a contingency table, determining if differences are due to chance or not, with Pearson’s being an approximation suitable for larger sample sizes and Fisher’s providing an exact calculation for low number samples81.
The results of this study indicated a significant agreement between resistance profile identified by Kirby–Bauer disc diffusion method and the presence of resistance encoding genes identified by PCR in K. pneumoniae. We found that the phenotypic resistance of amoxicillin-clavulanate, ceftriaxone, cefoxitin, cefuroxime, ceftazidime, piperacillin-tazobactam, imipenem, and meropenem among K. pneumoniae isolates correlate with the presence of blaNDM, blaOXA, blaTEM, blaVIM, blaIMP, and blaKPC genes. This correlation between phenotypes and genotypes were in agreement with recent studies82,83.
K. pneumoniae has developed antimicrobial resistance to many antibiotics due to high selection pressure from increasing use and misuse of antibiotics over the years. The transmission and acquisition of AMR occur primarily via a human-to-human interface within or outside of healthcare facilities84. In order to combat AMR, rational antibiotic prescription, limited use of prophylactic antimicrobials, patients’ education, compliance with antibiotic therapy, and appropriate hospital hygiene through effective antimicrobial stewardship are necessary85.
The CRISPR-Cas systems, derived from the adaptive immune system of prokaryotes, has been found in approximately 50% of bacterial genomes and 87% of archaea1. It is worth noting that pathogens with CRISPR-Cas systems were less likely to carry antibiotic resistance genes than those lacking this defense system2. However, recent studies have shown that the CRISPR systems are sometimes missed or inactivated and may not be an effective barrier to plasmid and drug resistance spread86. Therefore, the analysis of the relationship between the CRISPR-Cas system and antibiotic resistance will help to better understand the mechanism of bacterial resistance and provide new directions for the prevention and combating bacterial resistance87.
In this study, 65.9% (27/41) of K. pneumoniae isolates harbored CRISPR-Cas system encoding genes, which is considered a high percentage. This is higher than a previous report from Egypt that found CRISPR-Cas in 25.4% of their collection isolates88.
The high prevalence of the CRISPR-Cas systems in MDR K. pneumoniae in this study could be contrary to the capability of CRISPR-Cas to accurately eliminate drug resistance-related genes from bacterial strains in populations and to re-sensitize bacteria to antibiotics by deleting AMR encoding genes. The presence of CRISPR-Cas system encoding genes in resistant bacteria can indicate that these systems were present before but become inactive during evolution in order to give the bacteria the chance to host AMR genes and become resistant89.
In the current study, we found diverse profiles for CRISPR-Cas systems, the most prevalent one was CRISPR1, CRISPR2, CRISPR3, cas1, and cas3 (21.9%). This was in agreement with Makarova et al., who revealed that the CRISPR arrays are highly variable among bacterial species3.
Two cas genes were detected in this study. These genes included cas1 (68.3%), a universal cas gene that is found in most CRISPR-Cas types, and cas3(56.1%), the signature gene of the type I CRISPR-Cas system88,90.
No typical significant positive correlation between AMR and the presence of CRISPR-Cas system was detected (P value 0.1). The percentage of resistant K. pneumoniae isolates was clearly higher in the CRISPR-Cas-positive isolates than in the CRISPR-Cas-negative isolates.
The CRISPR system may be present as a remnant of a previous encounter with phages or foreign DNA, but it is inactive. Additionally, the CRISPR system requires the presence of specific triggering elements, such as protospacer adjacent motifs and foreign nucleic acids, to become active. Absence of these elements may render the CRISPR system inactive91,92. Additionally, mutations in genes encoding CRISPR-associated proteins or in the regulatory sequences of the CRISPR array could render the system inactive. Moreover, environmental conditions may play a role in regulating the activity of the CRISPR system. For example, nutrient availability or stress conditions could influence the activation of the system93,94.
Similar to our results Alkompoz et al. found a weak, non-significant positive correlation between the number of CRISPR system spacers and the number of resistance plasmids and ARGs in K. pneumoniae isolates88. Also two studies conducted in China found a negative correlation between the acquisition of ARGs and the presence of CRISPR-Cas systems87,95.
In the current study, the AMR detected by Kirby–Bauer disc diffusion method among K. pneumoniae isolates was higher in CRISPR-Cas-positive isolates compared to CRISPR-Cas-negative isolates indicating a potential association between the presence of the CRISPR-Cas system and increased antimicrobial resistance in K. pneumoniae.
A study from Egypt found that 25.4% of K. pneumoniae isolates carried the CRISPR-Cas system, and these isolates were more likely to be MDR or XDR compared to CRISPR-Cas-negative isolates. However, there was non-significant difference in susceptibility to the tested antimicrobial agents among the two groups of isolates88.
On the other hand, another study reported that resistance to various antibiotic classes, including β-lactams, quinolones, aminoglycosides, tetracyclines, and β-lactam/enzyme inhibitor combinations, was higher in the absence of the CRISPR-Cas system among K. pneumoniae isolates95. Similarly, a study in Iraq demonstrated an inverse association between the presence of the CRISPR-Cas system and antibiotic resistance, including ESBL production and carbapenem resistance in K. pneumoniae isolates and reported that multidrug resistance, extensive drug resistance, and pandrug resistance were more prevalent in CRISPR-Cas-negative strains29.
In the current study, the percentage of K. pneumoniae isolates harbored AMR encoding genes was clearly higher in the CRISPR-Cas-positive isolates than in the CRISPR-Cas-negative isolates except for blaNDM, blaOXA and blaIMP encoding genes. There was a significant difference (p-value < 0.05) of AMR encoding genes; aac(3)-IIa and mcr-1 (Pearson Chi-Square test, p-values = 0.039 and 0.036, respectively) among CRISPR-Cas-positive and CRISPR-Cas - negative K. pneumoniae isolates which were found in 96.3% and 81.5% of the CRISPR-Cas positive isolates, respectively versus 71.4% and 50% of the CRISPR-Cas negative isolate.
Generally, the CRISPR-Cas systems found in K. pneumoniae are not always correlated to the absence of ARGs; rather, an enormous number of ARGs and CRISPR-Cas systems have been found co-existing in the analyzed genomes96.
Alkompoz et al. revealed that the frequency of the genes, including blaVIM, blaNDM, and tetB were significantly higher in the presence of CRISPR-Cas systems. However, other genes such as blaTEM and blaKPC were significantly higher in the genomes of the CRISPR-Cas negative strains88. Also, Kadkhoda et al. found that the frequency of the blaTEM was significantly lower in the isolates with CRISPR-Cas- positive isolates in comparison to CRISPR-Cas- negative isolates. There was a significant inverse correlation between the presence of ESBL and some AME genes with CRISPR-Cas-positive isolates71.
The impact of the CRISPR-Cas system on limiting the dissemination of ARGs and, consequently, antimicrobial resistance was evident from some studies conducted on other bacterial species. The CRISPR-Cas system was previously found to be significantly associated with the absence of ARGs and high drug susceptibility in Enterococcus faecalis97,98and Pseudomonas aeruginosa99. On the other hand, several studies demonstrated that there was a highly significant inverse association between the prevalence of CRISPR-Cas system and drug resistance in carbapenem-resistant and ESBL-producing K. pneumoniae29,95,100. There are a lot of reasons explaining why the existence of the CRISPR-Cas systems on the bacterial genome does not always impede the dissemination of ARGs, starting from the adaptation stage. Point mutations and insertion sequence-mediated mutations in the adaptation genes correlated with the spread of MDR strains, as reported before in Shigella species101. In addition, strong selective pressure for antibiotic resistance may result in CRISPR repression and many CRISPR-harboring strains may be immunologically inactive owing to the existence of self-targeting spacers, which would be expected to induce an autoimmune response and host cell death102. On the other hand, phages expressing anti-CRISPR proteins (Acrs) may inactivate the CRISPR-Cas system, resulting in the dissemination of ARGs, as found in P. aeruginosa103.
Conclusion
The study revealed the high prevalence of MDR K. pneumoniae isolates in Egypt by phenotypic and molecular techniques. The study showed high resistance rates of K. pneumoniae to broad-spectrum β-lactam antibiotics, including third and fourth generation cephalosporins. Additionally, high resistance rates were noted for ciprofloxacin, gentamicin, amikacin, meropenem, and imipenem. In contrast, lower resistance rates were found for colistin and chloramphenicol. High prevalence of resistance encoding genes were detected, with a majority of isolates carrying the ESBLs encoding gene, carbapenem resistance encoding genes, blaNDM and blaOXA, aminoglycosides resistance encoding gene; aac(3)-IIa genes, tetB and the colistin resistance encoding genes mcr-1. There was no overall significant positive correlation between resistance detected byphenotypic and genotypic methods and the presence of CRISPR-Cas system except for imipenem, colistin and chloramphenicol phenotypic resistance andaac(3)-IIa and mcr-1 genes however, the percentage of resistant K. pneumoniae isolates was clearly higher in the CRISPR-Cas-positive isolates than in theCRISPR-Cas-negative isolates.
The results of this study challenge the belief that CRISPR-Cas systems are effective in eliminating drug resistance genes and sensitizing bacteria to antibiotics by deleting antimicrobial resistance encoding elements. Analysis of the relationship between the CRISPR-Cas system and AMR will help to better understand the mechanism of bacterial resistance and provide new strategies for the prevention and treatment of bacterial AMR. However, our investigation was limited by the small number of non-MDR isolates, as most Klebsiella pneumoniae isolates were MDR. Future studies with a larger range of isolates, including non-MDR strains, will be required to thoroughly confirm the association.
Data availability
All generated or analysed data during this study are included in this published article and supplementary materials.
Abbreviations
- AMR:
-
Antimicrobial resistance
- Cas:
-
CRISPR-associated genes/proteins
- CRISPR:
-
The clustered regularly interspaced short palindromic repeats
- K. pneumoniae :
-
Klebsiella pneumoniae
- MDR:
-
multidrug-resistant
- PCR:
-
Polymerase chain reaction
References
Makarova, K. S. et al. An updated evolutionary classification of CRISPR–Cas systems. Nat. Rev. Microbiol. 13, 722–736 (2015).
Pursey, E., Dimitriu, T., Paganelli, F. L., Westra, E. R. & van Houte, S. Crispr-cas is associated with fewer antibiotic resistance genes in bacterial pathogens. Philos. Trans. R Soc. B. 377, 20200464 (2022).
Makarova, K. S. et al. Evolution and classification of the CRISPR–Cas systems. Nat. Rev. Microbiol. 9, 467–477 (2011).
Bharathkumar, N. et al. Crispr/cas-based modifications for therapeutic applications: a review. Mol. Biotechnol. 64, 355–372 (2022).
Hatoum-Aslan, A., Maniv, I., Samai, P. & Marraffini, L. A. Genetic characterization of antiplasmid immunity through a type III-A CRISPR-Cas system. J. Bacteriol. 196, 310–317 (2014).
Louwen, R., Staals, R. H., Endtz, H. P., van Baarlen, P. & van der Oost, J. The role of CRISPR-Cas systems in virulence of pathogenic bacteria. Microbiol. Mol. Biol. Rev. 78, 74–88 (2014).
Mackow, N. A. et al. Crispr-cas influences the acquisition of antibiotic resistance in Klebsiella pneumoniae. PLoS One. 14, e0225131 (2019).
Tian, Q. et al. Application of CRISPR/Cas9 in rapeseed for gene function research and genetic improvement. Agronomy 12, 824 (2022).
Mayorga-Ramos, A. et al. Crispr-cas-based antimicrobials: design, challenges, and bacterial mechanisms of resistance. ACS Infect. Dis. 9, 1283–1302 (2023).
Havenga, B. et al. Exploring the antimicrobial resistance profiles of WHO critical priority list bacterial strains. BMC Microbiol. 19, 1–16 (2019).
Huang, L. et al. Bacterial multidrug efflux pumps at the frontline of antimicrobial resistance: an overview. Antibiotics 11, 520 (2022).
Algammal, A. M. & Behzadi, P. Antimicrobial resistance: a global public health concern that needs perspective combating strategies and new talented antibiotics. Discov Med. 36, 1911–1913 (2024).
Karampatakis, T., Tsergouli, K. & Behzadi, P. Carbapenem-resistant Pseudomonas aeruginosa’s resistome: pan-genomic plasticity, the impact of transposable elements and jumping genes. Antibiotics 14, 353 (2025).
Karampatakis, T., Tsergouli, K. & Behzadi, P. Pan-genome plasticity and virulence factors: a natural treasure trove for Acinetobacter baumannii. Antibiotics 13, 257 (2024).
Karampatakis, T., Tsergouli, K. & Behzadi, P. Carbapenem-resistant Klebsiella pneumoniae: virulence factors, molecular epidemiology and latest updates in treatment options. Antibiotics 12, 234 (2023).
Mulani, M. S. et al. Emerging strategies to combat ESKAPE pathogens in the era of antimicrobial resistance: a review. Front. Microbiol. 10, 539 (2019).
Gorrie, C. L. et al. Antimicrobial-resistant Klebsiella pneumoniae carriage and infection in specialized geriatric care wards linked to acquisition in the referring hospital. Clin. Infect. Dis. 67, 161–170 (2018).
Elshobary, M. E. et al. Combating antibiotic resistance: mechanisms, multidrug-resistant pathogens, and novel therapeutic approaches: an updated review. Pharmaceuticals 18, 402 (2025).
Olatunji, A. O. et al. Next-generation strategies to combat antimicrobial resistance: integrating genomics, CRISPR, and novel therapeutics for effective treatment. Eng. Sci. Tech. J. 5, 2284–2303 (2024).
Ahmed, M. M. et al. Crispr-cas systems in the fight against antimicrobial resistance: current status, potentials, and future directions. Infect. Drug Resist. 2024, 5229–5245 (2024).
Ramanan, P., Bryson, A. L., Binnicker, M. J., Pritt, B. S. & Patel, R. Syndromic panel-based testing in clinical microbiology. Clin Microbiol. Rev 31, 1–28 (2018).
Shoaib, M. et al. A mini-review on commonly used biochemical tests for identification of bacteria. Int. J. Res. Pub. 54, 1–7 (2020).
Weinstein, M. P. Performance standards for antimicrobial susceptibility testing. (2018).
Golpasand, T., Keshvari, M. & Behzadi, P. Distribution of chaperone-usher fimbriae and curli fimbriae among uropathogenic Escherichia coli. BMC Microbiol. 24, 344 (2024).
Odari, R. & Dawadi, P. Prevalence of multidrug-resistant Klebsiella pneumoniae clinical isolates in Nepal. J. Trop. Med. 2022, 5309350 (2022).
Zhuang, Q. et al. Advances in the detection of $\beta$-lactamase: a review. Int. J. Biol. Macromol. 251, 126159 (2023).
Giske, C. G., Turnidge, J., Cantón, R. & Kahlmeter, G. Update from the European committee on antimicrobial susceptibility testing (EUCAST). J. Clin. Microbiol. 60, e00276–e00221 (2022).
Effendi, M. H. et al. Molecular identification of BlaTEM gene of extended-spectrum beta-lactamase-producing Escherichia coli from healthy pigs in Malang district, East java, Indonesia. J. Adv. Vet. Anim. Res. 9, 447 (2022).
Jwair, N. A., Al-Ouqaili, M. T. & Al-Marzooq, F. Inverse association between the existence of CRISPR/Cas systems with antibiotic resistance, extended spectrum $\beta$-lactamase and carbapenemase production in multidrug, extensive drug and pandrug-resistant Klebsiella pneumoniae. Antibiotics 12, 980 (2023).
Asif, S., Khan, M., Arshad, M. W. & Shabbir, M. I. PCR optimization for beginners: a step by step guide. Res. Mol. Med. 9, 81–102 (2021).
Lee, P. Y., Costumbrado, J., Hsu, C. Y. & Kim, Y. H. Agarose gel electrophoresis for the separation of DNA fragments. J. Vis. Exp. : JoVE. 62, 3923 (2012).
Ostria-Hernández, M. L., Sánchez-Vallejo, C. J., Ibarra, J. A. & Castro-Escarpulli, G. Survey of clustered regularly interspaced short palindromic repeats and their associated Cas proteins (CRISPR/Cas) systems in multiple sequenced strains of Klebsiella pneumoniae. BMC Res. Notes. 8, 1–10 (2015).
Rouli, L., Merhej, V., Fournier, P. E. & Raoult, D. The bacterial pangenome as a new tool for analysing pathogenic bacteria. New. Microbes New. Infect. 7, 72–85 (2015).
Wyres, K. L. & Holt, K. E. Klebsiella pneumoniae as a key trafficker of drug resistance genes from environmental to clinically important bacteria. Curr. Opin. Microbiol. 45, 131–139 (2018).
Manjula, N. et al. Antibiotic susceptibility pattern of ES$\beta$L producing Klebsiella pneumoniae isolated from urine samples of pregnant women in Karnataka. J. Clin. Diagn. Res. : JCDR. 8, DC08 (2014).
Abdelhamid, S. M., Abd-Elaal, H. M., Matareed, M. O. & Baraka, K. Genotyping and virulence analysis of drug resistant clinical Klebsiella pneumoniae isolates in Egypt. J. Pure Appl. Microbiol. 14, 1967–1976 (2020).
Khalifa, H. O. et al. High prevalence of antimicrobial resistance in gram-negative bacteria isolated from clinical settings in egypt: recalling for judicious use of conventional antimicrobials in developing nations. Microb. Drug Resist. 25, 371–385 (2019).
Abdelaziz, S. M., Abdel-Rahman, M. A., El-Sheikh, H. H. & Azab, E. Correlation between the antibiotic resistance genes and susceptibility to antibiotics among the carbapenem-resistant gram-negative pathogens. Antibiotics 10, 255 (2021).
Azab, K. S. et al. Distribution of extended-spectrum $\beta$-lactamase (ESBL)-encoding genes among multidrug-resistant gram-negative pathogens collected from three different countries. Antibiotics 10, 247 (2021).
Afify, F. A. et al. Emergence of carbapenem resistant gram-negative pathogens with high rate of colistin resistance in egypt: a cross sectional study to assess resistance trends during the COVID-19 pandemic. J. Genet. Eng. Biotechnol. 22, 100351 (2024).
Moges, F. et al. High prevalence of extended-spectrum beta-lactamase-producing gram-negative pathogens from patients attending Felege Hiwot comprehensive specialized hospital, Bahir dar, Amhara region. PLoS One. 14, e0215177 (2019).
Al-Ouqaili, M. T., Khalaf, E. A. & Al-Kubaisy, S. H. DNA sequence analysis of BlaVEB gene encoding multi-drug resistant and extended-spectrum $\beta$-lactamases producer isolates of Enterobacteriaceae and Pseudomonas aeruginosa. Open Microbiol. J 14, 40–47 (2020).
Anwar, M. A. Detection of BlaKPC gene among carbapenem resistant Enterobacteriacae isolates from Ain Shams university hospital, Egypt. Egyp J. Med. Microbiol. 31, 151–156 (2022).
Taha, M. S. et al. Genotypic characterization of carbapenem-resistant Klebsiella pneumoniae isolated from an Egyptian university hospital. Pathogens 12, 121 (2023).
Baral, R., Tuladhar, R., Manandhar, S., Singh, A. & Sherchan, S. Detection of Bla Kpc gene among carbapenemase producing Klebsiella pneumoniae isolated from different clinical specimens at tertiary care hospital of Nepal. BMC Microbiol. 24, 144 (2024).
Eliakim-Raz, N. et al. Efficacy and safety of chloramphenicol: joining the revival of old antibiotics? Systematic review and meta-analysis of randomized controlled trials. J. Antimicrob. Chemother. 70, 979–996 (2015).
Mikhail, S. et al. Evaluation of the synergy of ceftazidime-avibactam in combination with meropenem, amikacin, aztreonam, colistin, or fosfomycin against well-characterized multidrug-resistant Klebsiella pneumoniae and Pseudomonas aeruginosa. Antimicrob. Agents Chemot1her. 63, (2019). https://doi.org/10.1128/aac.00779-00719
Mikhail, V. E. et al. Antimicrobial resistance of clinical isolates of Klebsiella pneumoniae and Escherichia coli in Russian hospitals: results of a multicenter epidemiological study. Clin. Microbiol. Antimicrob. Chemother. 26, 67–78 (2024).
Wang, M. et al. Clinical outcomes and bacterial characteristics of carbapenem-resistant Klebsiella pneumoniae complex among patients from different global regions (CRACKLE-2): a prospective, multicentre, cohort study. Lancet Infect. Dis. 22, 401–412 (2022).
Magalhães, M. L. & Blanchard, J. S. Aminoglycosides: mechanisms of action and resistance (ed. Mayers, D.L.) Antimicrobial Drug Resistance. Infectious Disease. 171–181 (Humana Press, 2009).
Richter, P. et al. Using colistin as a Trojan horse: inactivation of gram-negative bacteria with chlorophyllin. Antibiotics 8, 158 (2019).
Abu-El-Azayem, A. K., Dwedar, R. A., Nashaat, N. & Shash, R. Y. Comparative evaluation of colistin susceptibility testing using agar Dilution and broth microdilution in multidrug-resistant and extensively drug-resistant gram-negative isolates. Egyp J. Med. Microbiol. 32, 109–115 (2023).
Zafer, M. M. et al. Emergence of colistin resistance in multidrug-resistant Klebsiella pneumoniae and Escherichia coli strains isolated from cancer patients. Ann. Clin. Microbiol. Antimicrob. 18, 1–8 (2019).
Mahmoud, F. et al. Molecular mechanisms of colistin resistance among multi-drug resistant (MDR) Klebsiella pneumoniae and Escherichia coli isolated from ICU patients and their susceptibility towards Eravacycline. Microbes Infect. Dis. 4, 127–137 (2023).
Emara, M. M., Abd-Elmonsef, M. M., Elnasr, A., Elfeky, A. & L. M. & A. E.-E. Study of mcr-1 gene-mediated colistin-resistance in gram-negative isolates in Egypt. Egyp J. Med. Microbiol. 28, 9–16 (2019).
Farhadi, M. et al. High frequency of multidrug-resistant (MDR) Klebsiella pneumoniae harboring several $\beta$-lactamase and integron genes collected from several hospitals in the North of Iran. Ann. Clin. Microbiol. Antimicrob. 20, 1–9 (2021).
Prestinaci, F., Pezzotti, P. & Pantosti, A. Antimicrobial resistance: a global multifaceted phenomenon. Pathog Global Health. 109, 309–318 (2015).
Elkhawaga, A. A., Hetta, H. F., Osman, N. S., Hosni, A. & El-Mokhtar, M. A. Emergence of Cronobacter Sakazakii in cases of neonatal sepsis in upper egypt: first report in North Africa. Front. Microbiol. 11, 215 (2020).
Akinbami, O. R., Olofinsae, S. & Ayeni, F. A. Prevalence of extended spectrum beta lactamase and plasmid mediated quinolone resistant genes in strains of Klebsiella pneumonia, Morganella morganii, leclercia adecarboxylata and Citrobacter freundii isolated from poultry in South Western n2igeria. PeerJ 6, e5053 (2018).
Olowo-okere, A. et al. Phenotypic and genotypic characterization of clinical carbapenem-resistant Enterobacteriaceae isolates from sokoto, Northwest Nigeria. New. Microbes New. Infect. 37, 100727 (2020).
Shaikh, S. et al. Antibiotic resistance and extended spectrum beta-lactamases: types, epidemiology and treatment. Saudi J. Biol. Sci. 22, 90–101 (2015).
El-Domany, R. A., El-Banna, T., Sonbol, F. & Abu-Sayedahmed, S. H. Co-existence of NDM-1 and OXA-48 genes in carbapenem resistant Klebsiella pneumoniae clinical isolates in kafrelsheikh, Egypt. Afr. Health Sci. 21, 489–496 (2021).
ElBaradei, A. Co-occurrence of blaNDM-1 and blaOXA-48 among carbapenem resistant Enterobacteriaceae isolates causing bloodstream infections in alexandria, Egypt. Egyp J. Med. Microbiol. 31, 1–7 (2022).
Mohamed, T., Yousef, L. M., Darweesh, E. I. & Khalil, A. H. Detection and characterization of carbapenem resistant Enterobacteriacea in Sohag university hospitals. Egyp J. Med. Microbiol. 27, 61–69 (2018).
Urmi, U. L. et al. Genotypic to phenotypic resistance discrepancies identified involving β-betaβ-lactamase genes, Bla KPC, Bla IMP, Bla NDM-1, and Bla VIM in uropathogenic Klebsiella pneumoniae. Infect. Drug. Resist.,13 2863–2875 (2020).
Depardieu, F., Podglajen, I., Leclercq, R., Collatz, E. & Courvalin, P. Modes and modulations of antibiotic resistance gene expression. Clin. Microbiol. Rev. 20, 79–114 (2007).
Johnning, A. et al. The resistomes of six carbapenem-resistant pathogens–a critical genotype–phenotype analysis. Microb. Genom. 4, e000233 (2018).
Kulengowski, B. & Burgess, D. S. Imipenem/relebactam activity compared to other antimicrobials against non-MBL-producing carbapenem-resistant Enterobacteriaceae from an academic medical center. Pathog Dis. 77, ftz040 (2019).
Almangour, T. A. et al. Novel β-lactam-β-lactamase inhibitors as monotherapy versus combination for the treatment of drug-resistant Pseudomonas aeruginosa infections: a multicenter cohort study. J. Infect. Chemother. 30, 1008–1014 (2024).
Harir Foroush, M., Shokoohizadeh, L. & Mirzaee, M. Prevalence of genes encoding aminoglycoside modifying enzymes in clinical isolates of Klebsiella pneumoniae in the hospitals of Borujerd. Int. J. Med. Lab. 5, 35–41 (2018).
Kadkhoda, H. et al. Prevalence of the CRISPR-cas system and its association with antibiotic resistance in clinical Klebsiella pneumoniae isolates. BMC Infect. Dis. 24, 554 (2024).
Abozahra, R., Gaballah, A. & Abdelhamid, S. M. Prevalence of the colistin resistance gene MCR-1 in colistin-resistant Klebsiella pneumoniae in Egypt. AIMS Microbiol. 9, 177 (2023).
Zaki, M., ElKheir, N. & Mofreh, M. Molecular study of colistin resistant clinical isolates of Enterobacteriaceae species. J. Clin. Mol. Med. 1, 1–4 (2018).
Rabie, R. A. & Abdallah, A. L. Plasmid mediated colistin resistant genes mcr-1 and mcr-2 among Escherichia coli and Klebsiella pneumoniae isolates at zagazig university hospitals, egypt. Egyp. J. Med. Microbiol. 29, 61–66 (2020).
Abdelmalak, D. N., Hassan, M. G., Nasr, A. & Shaaban, M. T. A comprehensive analysis of antibiotic resistance pattern in nosocomial infections in Egypt. Egyp Acad. J. Biol. Sci. G Microbiol. 17, 15–25 (2025).
EL-Ganiny, A. M. et al. Phenotypic and genotypic detection of-lactams resistance in Klebsiella species from Egyptian hospitals revealed carbapenem resistance by OXA and NDM genes. Afr. J. Microbiol. Res. 10, 339–347 (2016).
Tao, R. et al. Detection of antibiotic resistance and Tetracycline resistance genes in Enterobacteriaceae isolated from the Pearl rivers in South China. Environ. Pollut. 158, 2101–2109 (2010).
Sheykhsaran, E. et al. The rate of resistance to tetracyclines and distribution of tetA, tetB, tetC, tetD, tetE, tetG, TetJ and TetY genes in Enterobacteriaceae isolated from azerbaijan, Iran during 2017. Physiol. Pharmacol. 22, 205–212 (2018).
Kashefieh, M., Hosainzadegan, H., Baghbanijavid, S. & Ghotaslou, R. The molecular epidemiology of resistance to antibiotics among Klebsiella pneumoniae isolates in azerbaijan, iran. J Trop. Med. , 12 9195184 (2021).
Li, Q. et al. Microbial degradation of Tetracycline antibiotics: mechanisms and environmental implications. J. Agric. Food Chem. 72, 13523–13536 (2024).
Kim, H. Y. Statistical notes for clinical researchers: chi-squared test and fisher’s exact test. Restor. Dent. Endod. 42, 152 (2017).
Tawfick, M. M., Elshamy, A. A. & Mohamed, K. T. & El Menofy, N. G. Gut commensal Escherichia coli, a high-risk reservoir of transferable plasmid-mediated antimicrobial resistance traits. Infect Drug Resist. 15, 1077–1091 (2022).
Elmanakhly, A. R. et al. Carbapenem-resistant Klebsiella pneumoniae: diversity, virulence, and antimicrobial resistance. Infect Drug Resist. 15, 6177–6187 (2022).
Salam, M. A. et al. Antimicrobial resistance: a growing serious threat for global public health. Healthcare, 1946 (2023).
Abushaheen, M. A. et al. Antimicrobial resistance, mechanisms and its clinical significance. Dis. -a-Mon. 66, 100971 (2020).
Gophna, U. et al. No evidence of Inhibition of horizontal gene transfer by CRISPR–Cas on evolutionary timescales. ISME J. 9, 2021–2027 (2015).
Tao, S., Chen, H., Li, N. & Liang, W. The application of the CRISPR-Cas system in antibiotic resistance. Infect Drug Resist. 15, 4155–4168 (2022).
Alkompoz, A. K. et al. Correlation of CRISPR/Cas and antimicrobial resistance in Klebsiella pneumoniae clinical isolates recovered from patients in Egypt compared to global strains. Microorganisms 11, 1948 (2023).
Pinilla-Redondo, R. et al. Crispr-cas systems are widespread accessory elements across bacterial and archaeal plasmids. Nucleic Acids Res. 50, 4315–4328 (2022).
Makarova, K. S. & Koonin, E. V. Annotation and classification of CRISPR-Cas systems. CRISPR: Methods And Protocols 47–75 (2015).
Papikian, A., Liu, W., Gallego-Bartolomé, J. & Jacobsen, S. E. Site-specific manipulation of Arabidopsis loci using CRISPR-Cas9 suntag systems. Nat. Commun. 10, 729 (2019).
Hong, J. K. et al. Multiplex CRISPR/Cas9 mutagenesis of BrVRN1 delays flowering time in Chinese cabbage (Brassica Rapa L. ssp. pekinensis). Agriculture 11, 1286 (2021).
Bilger, R. et al. Control of a type III-Dv CRISPR–Cas system by the transcription factor RpaB and interaction of its leader transcript with the DEAD-box RNA helicase CrhR. bioRxiv 2023, 2023.2012. 2007.570523. (2023).
Li, Y. et al. Quorum sensing inhibits type III-A CRISPR-Cas system activity through repressing positive regulators SarA and ArcR in Staphylococcus aureus. Biorxiv 2023, 2023.2001. 2017.524377 (2023).
Wang, G., Song, G. & Xu, Y. Association of CRISPR/Cas system with the drug resistance in Klebsiella pneumoniae. Infect Drug Resist. 13, 1929–1935 (2020).
Enany, S. et al. Whole genome sequencing of Klebsiella pneumoniae clinical isolates sequence type 627 isolated from Egyptian patients. PLoS One. 17, e0265884 (2022).
Gholizadeh, P. et al. Role of CRISPR-Cas system on antibiotic resistance patterns of Enterococcus faecalis. Ann. Clin. Microbiol. Antimicrob. 20, 1–12 (2021).
Alduhaidhawi, A. H. M. et al. Prevalence of CRISPR-cas systems and their possible association with antibiotic resistance in Enterococcus faecalis and Enterococcus faecium collected from hospital wastewater. Infect Drug Resist. 15, 1143–1154 (2022).
Soliman, M., Said, H. S., El-Mowafy, M., Barwa, R. & Novel PCR detection of CRISPR/Cas systems in Pseudomonas aeruginosa and its correlation with antibiotic resistance. Appl. Microbiol. Biotechnol. 106, 7223–7234 (2022).
AL-Yozbakee, Z. M. & Mohammad, K. O. Crispr-cas system in multi drugs resistant Klebsiella pneumoniae from different clinical samples and its correlation with antibiotic-resistant genes in Mosul city/iraq. J Appl. Nat. Sci 16, 820–829 (2024).
Ren, L. et al. Relationship between drug resistance and the clustered, regularly interspaced, short, palindromic repeat-associated protein genes cas1 and cas2 in Shigella from giant panda Dung. Medicine 96, e5922 (2017).
Shen, J., Lv, L., Wang, X., Xiu, Z. & Chen, G. Comparative analysis of CRISPR-Cas systems in Klebsiella genomes. J. Basic. Microbiol. 57, 325–336 (2017).
Sontheimer, E. J. & Davidson, A. R. Inhibition of CRISPR-Cas systems by mobile genetic elements. Curr. Opin. Microbiol. 37, 120–127 (2017).
Acknowledgements
The authors would like to acknowledge the Open access funding provided by The Science, Technology and Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB) and Springer Nature transformative agreement.
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB).
Author information
Authors and Affiliations
Contributions
NGE performed conceptualization, supervision, methodology, validation, investigation, data curation, writing, review and editing final draft. ANP conducted investigation, methodology, resources, formal analysis, and writing-original draft. MAE conducted supervision, validation, investigation, data curation, and reviewed the final draft. AE performed conceptualization, supervision, visualization, validation, and reviewed the final draft, all authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
The study obtained its approval from the ethical committees of Faculty of Pharmcy (Girls), Al-Azhar University (Approval number: Rec – CLAR8B).
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
El Menofy, N.G., Payoumi, A.N., Eissa, M.A. et al. Association of the existence of CRISPR-Cas system and antimicrobial resistance in multi-drug resistant Klebsiella pneumoniae in Egypt. Sci Rep 15, 41814 (2025). https://doi.org/10.1038/s41598-025-26706-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-26706-6