Abstract
Plasma metabolomics offers valuable insights for identifying viral infection biomarkers with applications in early diagnosis, outcome prediction, and treatment monitoring. This study aimed to investigate metabolomic alterations in influenza A patients and identify potential biomarkers for disease severity. From March 2023 to March 2024, 339 influenza A patients were enrolled. After age‒sex matching, 54 patients and 20 healthy controls were selected for analysis. Untargeted metabolomic profiling of 74 plasma samples was conducted using ultrahigh-performance liquid chromatography coupled with tandem mass spectrometry. Comparative analysis revealed 60 differentially expressed metabolites between H1N1 patients and healthy controls, with 41 significantly upregulated and 19 downregulated. Pathway enrichment analysis revealed prominent disruptions in glycerophospholipid metabolism, with several metabolites showing alterations across the severity spectrum. Amino acid metabolism, particularly propionate metabolism, glycine-serine-threonine metabolism, and branched-chain amino acid biosynthesis, was also notably disturbed. Critical exhibited marked disturbances in taurine-hypotaurine metabolism compared to milder. This study identified glycerophospholipid metabolism dysregulation as a potential biomarker for influenza severity stratification. The progressive alteration of taurine pathway metabolites in critical suggests their pivotal role in severe H1N1 pathogenesis, highlighting their dual potential as diagnostic biomarkers and therapeutic targets.
Similar content being viewed by others
Introduction
Influenza is an acute respiratory infectious disease caused by the influenza virus. It is seasonally prevalent every year and is mostly self-limiting. However, high-risk individuals with chronic underlying diseases may develop severe influenza, and a small number of cases progress rapidly to acute respiratory distress syndrome (ARDS), acute necrotizing encephalopathy, or multiorgan dysfunction; severe cases may even lead to death1. Therefore, early diagnosis and treatment of influenza play important roles in reducing the incidence of severe illness and fatality rate, as well as further transmission of the disease.
Plasma metabolomics is a method used to study the metabolic state of an organism by analyzing metabolites in plasma. In influenza, COVID-19 and other infectious diseases, metabolic processes are closely related to disease pathogenesis, pathology and pathophysiological changes, as well as the host defense response to infection2,3. For example, a previous study4 demonstrated distinct alterations in acylcarnitine and sphingolipid metabolism in patients with severe influenza, highlighting their potential as prognostic markers.
Influenza infection can lead to changes in host metabolism, including energy metabolism, lipid metabolism, and amino acid metabolism5,6. Plasma metabolomic methods can detect these changes, and understanding the biological events that occur during the infection process by analyzing the changes in metabolite levels may help to identify biomarkers of influenza infection. Notably, machine learning approaches that integrate metabolomic data with clinical parameters have shown > 90% accuracy in distinguishing influenza severity grades7. These markers can be used for early diagnosis, prediction of disease severity and monitoring the efficacy of treatment.
Our study aimed to identify specific metabolites associated with influenza virus infection by comparing the plasma metabolic profiles of influenza patients with those of healthy controls and comparing patients with varying degrees of influenza severity. By identifying specific metabolites linked to critical illness, these findings provide a foundation for the early identification of critically ill patients and offer potential strategies for reducing mortality. Specifically, we state that this is the first metabolomic study in Chinese adult patients with confirmed H1N1 infection stratified by severity. We emphasize the discovery of LysoPC(20:0/0:0) and taurine pathway disruptions as progressive markers from mild to critical cases.
Methods
Study subjects and sample collection
A total of 339 patients with influenza A, admitted to the Second Affiliated Hospital of Xi’an Jiaotong University from March 2023 to March 2024, were included in the study. This group included 301 cases of mild influenza, 24 severe cases, and 14 critical cases. Additionally, 54 patients with influenza A (including 20 mild, 20 severe, and 14 critical) were enrolled in the study. A control group of 20 healthy adults, who were not infected with influenza, was also included.
The inclusion criteria required that the influenza patients be older than 18 years and meet the diagnostic criteria outlined in the Expert Consensus on Antiviral Therapy for Influenza in Adults (2022 edition)8. Mild were characterized by typical influenza symptoms, such as fever, cough, sore throat, and muscle aches, without significant respiratory distress. Severe involved more pronounced symptoms, including high fever, persistent cough, difficulty breathing, and chest tightness. Severe were also associated with underlying comorbidities, such as chronic cardiovascular or respiratory conditions. Critical were defined by life-threatening symptoms, including respiratory failure, shock, or multiorgan dysfunction. These patients required intensive care unit (ICU) admission, mechanical ventilation, or other advanced life support measures. The study was performed in accordance with the Declaration of Helsinki. The study protocol was approved by the Ethics Committee of the Xi’an Jiaotong University Second Affiliated Hospital, and all patients provided written informed consent prior to enrollment.
Peripheral blood collection and processing
All samples were collected at hospital admission, before treatment initiation, to ensure uniformity in sampling timepoints across severity groups. All patients underwent throat swab and blood tests, including complete blood count, renal function, liver function, electrolyte level, and myocardial enzyme tests. There were measured using standard automated clinical chemistry analyzers (e.g., Beckman Coulter AU5800) at the hospital’s central lab. Peripheral blood was also collected for untargeted metabolomic analysis. Influenza A was confirmed via RT-PCR on pharyngeal swab specimens using kits from Sansure Biotech Inc. manufacturer (Cat No. 20213400256), following CDC guidelines.
For all patients, fasting cubital venous blood (10 mL) was collected in sodium heparin anticoagulant tubes. The blood was then centrifuged at 2000 rpm for 10 min, and the plasma was extracted at 4 °C. The plasma samples were stored at -80 °C until further analysis.
Plasma sample preparation and LC–MS/MS analysis
Plasma metabolites were extracted by mixing 100 µL of sample with 400 µL of methanol: acetonitrile (1:1, v/v) containing isotope-labeled internal standards. The mixture was vortexed, sonicated in ice-water for 10 min, incubated at − 40 °C for 1 h, and centrifuged at 12,000 rpm for 15 min at 4 °C. Supernatants were collected for analysis, and pooled quality control (QC) samples were prepared from all samples. Metabolomic profiling was performed using a Vanquish UHPLC system coupled to an Orbitrap Exploris 120 mass spectrometer (Thermo Fisher Scientific). Separation was achieved on a Waters ACQUITY UPLC BEH Amide column (2.1 mm × 50 mm, 1.7 μm) with a mobile phase comprising 25 mM ammonium acetate and 25 mM ammonium hydroxide (pH 9.75) in water (A) and acetonitrile (B). The injection volume was 2 µL, and the autosampler was maintained at 4 °C. Mass spectrometry was conducted in both positive and negative ionization modes with a resolution of 60,000 for MS1 and 15,000 for MS2. Data acquisition used stepped collision energy (20/30/40 eV), and system performance was monitored via repeated QC injections. Raw data were processed using XCMS in R, and metabolites were annotated by matching MS/MS spectra to the BiotreeDB (v2.1) in-house database with a match score threshold of 0.3. Annotation confidence followed the Metabolomics Standards Initiative (MSI) guidelines.
Metabolomics statistical analysis
Metabolite peak intensities were log-transformed, filtered based on missing value threshold (> 80% present across all samples), and normalized using probabilistic quotient normalization (PQN). Data were then pareto-scaled prior to multivariate analysis. Orthogonal partial least squares discriminant analysis (OPLS-DA) was performed using SIMCA (v16.0) to visualize group separation and identify discriminative metabolites. Compared to PLS-DA, OPLS-DA removes variation unrelated to group classification, improving model interpretability. Variable importance in projection (VIP) scores were used to rank metabolite contributions, with VIP > 1 considered significant. Univariate analysis was performed using Student’s t-test or Mann–Whitney U test as appropriate, and p-values were corrected using the Benjamini–Hochberg method for controlling the false discovery rate (FDR < 0.05). Metabolic pathway enrichment analysis was conducted using MetaboAnalyst 5.0, with pathway libraries selected from KEGG (human) and over-representation analysis (ORA) applied using the hypergeometric test.
Primary and secondary mass spectrometry data were obtained using an Orbitrap Exploris 120 mass spectrometer under the control of Xcalibur software (version: 4.4, Thermo). Detailed parameters are as follows: sheath gas flow rate as 50 Arb, Aux gas flow rate as15 Arb, capillary temperature 320 C, full Ms resolution as 60,000, Ms/Ms resolution as15000, collision energy: SNcE 20/30/40, spray voltage as 3.8 kv (positive) or -3.4 kV(negative), respectively. After the original data were converted to mzXML format via ProteoWizard software, the R program package (XCMS kernel) was used for peak identification, peak extraction, peak alignment, and integration, and then, a self-built secondary databank with BiotreeDB (V2.1) was used. Matching was performed with the mass spectrum database for substance annotation, and the cutoff value for algorithm scoring was set to 0.3.
Metabolite annotations were assigned based on the Metabolomics Standards Initiative (MSI) guidelines, with most identifications at Level 2 (MS/MS and RT match). Structural isomers were not distinguished unless confirmed with reference standards.
Statistical analysis
SPSS 23.0 software was used for statistical analysis. The measurement data are expressed as x ± s or M (Q25, Q75), and comparisons were performed via t tests (normally distributed data) and Mann‒Whitney U tests (nonnormally distributed data). The count data were analyzed via the χ2 test with α = 0.05. Spearman correlation analysis was used to evaluate associations between metabolite abundance and clinical indicators. Significance was defined as p < 0.05.
Results
Clinical characteristics of the study population
A total of 339 patients with influenza A were eventually enrolled as study subjects. All cases were confirmed etiologically, with all nucleic acid tests (pharyngeal swabs) returning positive for H1N1. The influenza patients were divided into three groups: mild, severe, and critical. Among these patients, 301 had mild (88.79%), 24 had severe (7.08%), and 14 had critical (4.13%). In addition, a subset of 54 patients (20 mild, 20 severe, and 14 critical) was selected from the full cohort for metabolomic analysis. Additionally, 20 healthy individuals were enrolled during the same period. A total of 74 blood samples were collected for untargeted metabolomic analysis. The screening and classification of the study subjects are depicted in Supplementary Fig. 1.
Among the 339 patients with influenza A, with 170 males (50.15%) and 169 females (49.85%), with a median age of 27 (21, 37) years. Fever was the initial symptom in 338 patients (99.71%). Cough, sputum, and intolerance to cold were observed in 299 patients (88.20%), 225 patients (66.37%), and 225 patients (66.37%), respectively, making them the next most common mild clinical manifestations after fever. Nasal congestion was reported by 214 patients (63.13%), runny nose by 216 patients (63.72%), and sore throat by 94 patients (27.73%). Among the 303 patients enrolled who presented with notable systemic symptoms, 203 had headaches (59.88%), 194 experienced muscle soreness (57.23%), 95 reported fatigue (28.02%), and 83 experienced dizziness (24.48%). Chest tightness and shortness of breath were observed in 40 patients (11.80%). Gastrointestinal symptoms such as nausea (32 patients, 9.44%), vomiting (7 patients, 2.06%), abdominal pain (3 patients, 0.88%), and diarrhea (6 patients, 1.77%) were also noted. The demographic data and epidemiological characteristics of influenza patients in the common group (301 common) and the critical group (24 severe and 14 critical ) in Supplementary Table 1, and clinical characteristics of influenza patients (20 mild, 20 severe, and 14 critical) and healthy controls in Table 1.
Changes in the plasma metabolic profiles of influenza patients
Differences in the plasma metabolome between the influenza group and the healthy control group
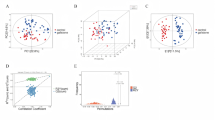
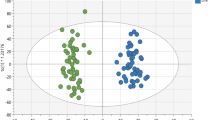
Plasma samples from 54 influenza patients and 20 healthy controls were subjected to untargeted metabolomic analysis. A total of 563 metabolites were detected and analyzed using orthogonal PLS-DA (R²Y = 0.882, Q²=0.605; P < 0.01), as shown in Fig. 1A and B, demonstrating clear separation between the influenza and control groups. Differentially abundant metabolites (n = 60) were identified on the basis of the following criteria: VIP > 1, P < 0.05, fold change > 1.5 or < 2/3, and M2 score > 0.7. Among these, 41 metabolites were upregulated, and 19 were downregulated (Fig. 1C). Putative exogenous metabolites, such as cytarabine and apigenin, were excluded from downstream biomarker analysis due to their likely dietary or medicinal origin.
Metabolomic analysis of the influenza group and healthy control group. OPLS-DA score diagram of influenza patients and healthy controls (A and B). Volcano plot of differentially abundant metabolites between the influenza group and the healthy control group (C). Top 20 metabolites with the highest VIP values (D). Heatmap of the top 30 differentially abundant metabolites between influenza patients and healthy controls (E). Enrichment analysis of the differentially abundant metabolic pathways in the influenza group vs. healthy controls (F). *p < 0.05, **p < 0.01, ***p < 0.001, based on Mann–Whitney U test.
Among the 60 differentially abundant metabolites, 11 (18.3%) were glycerophospholipids, 81.8% of which were significantly downregulated in influenza patients. The 20 metabolites with the highest VIP scores in the OPLS-DA model are depicted in Fig. 1D, whereas the top 30 differentially abundant metabolites are visualized in a heatmap in Fig. 1E.
KEGG pathway analysis revealed that the differentially abundant metabolites were enriched primarily in glycerophospholipid metabolism, primary bile acid biosynthesis, porphyrin metabolism, and linoleic acid metabolism (Fig. 1F, Supplementary Table 2).
Differences in plasma metabolism between mild and severe
Plasma samples from 20 mild and 20 severe were subjected to untargeted metabolomic analysis. OPLS-DA score plots (Supplementary Fig. 2A and B) revealed a cumulative R²Y of 0.825 and a Q² of 0.146, indicating minimal distinction between mild and severe. Among the 561 detected metabolites, 82 were significantly differentially abundant (P < 0.05), with 25 being further identified (17 upregulated, 8 downregulated, Supplementary Fig. 2C). Details of these differentially abundant metabolites are provided in Supplementary Table 3.
Differences in plasma metabolism between mild and critical
Plasma samples from 20 mild and 14 critical were subjected to untargeted metabolomic analysis. A total of 563 metabolites were detected and analyzed using orthogonal PLS-DA (R²Y = 0.968, Q²=0.704; P < 0.01), as shown in Fig. 2A and B, demonstrating clear separation between the mild and critical. Differentially abundant metabolites (n = 58) were identified on the basis of the following criteria: VIP > 1, P < 0.05, fold change > 1.5 or < 2/3, and M2 score > 0.7. Among these, 17 metabolites were upregulated, and 41 were downregulated (Fig. 2C).
Metabolomic analysis of mild and critical. OPLS-DA score diagram of mild and critical (A and B). Volcano plot of differentially abundant metabolites between mild and critical (C). Top 20 metabolites with the highest VIP values (D). Heatmap of the top 30 differentially abundant metabolites between mild and critical (E). Enrichment analysis of the differentially abundant metabolite pathways in mild vs. critical (F). *p < 0.05, **p < 0.01, ***p < 0.001, based on Mann–Whitney U test.
Among the 58 differentially abundant metabolites, 6 (10.3%) were glycerophospholipids, 83.3% of which were significantly downregulated in influenza patients. The 20 metabolites with the highest VIP scores in the OPLS-DA model are depicted in Fig. 2D, whereas the top 30 differentially abundant metabolites are visualized in a heatmap in Fig. 2E.
KEGG pathway analysis revealed that the differentially abundant metabolites were enriched primarily in glycerophospholipid metabolism, linoleic acid metabolism, primary bile acid biosynthesis, and taurine and hypotaurine metabolism (Fig. 2F, Supplementary Table 4).
Differences in plasma metabolism between mild and severe/critical
Plasma samples from 20 mild and 34 severe/critical were subjected to untargeted metabolomic analysis. A total of 563 metabolites were detected and analyzed using orthogonal PLS-DA (R²Y = 0.707, Q²=0.402; P < 0.05), as shown in Fig. 3A and B, demonstrating clear separation between the mild and severe/critical. Differentially abundant metabolites (n = 43) were identified on the basis of the following criteria: VIP > 1, P < 0.05, fold change > 1.5 or < 2/3, and M2 score > 0.7. Among these, 16 metabolites were upregulated, and 27 were downregulated (Fig. 3C).
Metabolomic analysis of mild and severe/critical. OPLS-DA score diagram of mild and severe/critical (A and B). Volcano plot of differentially abundant metabolites between mild patients and severe/critical (C). Top 20 metabolites with the highest VIP values (D). Heatmap of the top 30 differentially abundant metabolites between mild and severe/critical (E). Enrichment analysis of the differentially abundant metabolite pathways in mild vs. severe/critical (F). *p < 0.05, **p < 0.01, ***p < 0.001, based on Mann–Whitney U test.
The 20 metabolites with the highest VIP scores in the OPLS-DA model are depicted in Fig. 3D, whereas the 30 differentially abundant metabolites are visualized in a heatmap in Fig. 3E.
KEGG pathway analysis revealed that the differentially abundant metabolites were taurine and hypotaurine metabolism; glycerol phospholipid metabolism; glycine, serine and threonine metabolism; propanoate metabolism; and valine, leucine and isoleucine metabolism (Fig. 3F, Supplementary Table 5).
Association between glycerophospholipid metabolites and clinical indicators
The levels of 11 glycerophospholipid metabolites in influenza patients were significantly different from those in healthy controls, with 9 (81.8%) being markedly downregulated. Among the downregulated metabolites, LysoPC(20:0/0.0), PC(P-18:1(9Z)/16:1(9Z)), PC(P-18:1(9Z)/16:0) and PC(18:1(11Z)/14:1(9Z)) were negatively correlated with inflammatory markers (WBC, neutrophil count, CRP, and PCT). However, these metabolites were all positively correlated with lymphocyte counts (Fig. 4A).
Associations between differentially abundant glycerophospholipid metabolites and clinical indicators in patients with influenza. Associations between differentially abundant glycerophospholipid metabolites and clinical indicators in influenza patients and healthy controls (A). Associations between differentially abundant glycerophospholipid metabolites and clinical indicators in mild and critical (B). Association between differentially abundant glycerophospholipid metabolites and clinical indicators in mild and severe/critical (C). Correlation coefficients are based on Spearman’s rank test. *p < 0.05, **p < 0.01.
The levels of glycerophospholipid metabolites in critical were significantly different from those in mild. The downregulated metabolite LysoPC (20:0/0.0) was negatively correlated with lactate dehydrogenase (LDH). The upregulated metabolite PE(20:2(11Z,14Z)/14:0) was positively correlated with LDH (Fig. 4B).
The levels of glycerophospholipid metabolites in severe/critical were significantly different from those in mild. The upregulated metabolites LysoPA(16:0/0:0) and LysoPA(18:1(9Z)/0:0) were negatively correlated with lymphocyte counts and monocyte counts and positively correlated with the creatine kinase isoenzyme. The downregulated metabolite LysoPC (20:0/0.0) was positively correlated with inflammatory markers (lymphocyte and monocyte counts) and negatively correlated with inflammatory markers (WBC and neutrophil counts). (Fig. 4C).
Discussion
In this study, we performed untargeted metabolomic analysis on peripheral plasma samples from influenza patients and healthy controls. The results revealed significant differences in metabolites between the groups, particularly in glycerophospholipid metabolism and amino acid metabolism pathways. These findings suggest potential clinical correlations between specific glycerophospholipids and inflammatory/immunological markers, offering novel insights into metabolic reprogramming during influenza progression and highlighting potential biomarkers for severity stratification.
Influenza is a highly contagious acute respiratory infectious disease, with 3 to 5 million cases of severe illness and approximately 290,000 to 650,000 deaths occurring annually worldwide9. In addition to its direct health impact on patients, influenza imposes substantial pressure on global public health systems and healthcare resources, causing significant socioeconomic burdens. Influenza-related mortality is especially high in elderly individuals and patients with comorbidities, with severe often progressing to acute respiratory distress syndrome (ARDS), pneumonia, or other complications, leading to a fatality rate as high as 30% in ICU settings9. Early diagnosis and timely treatment of influenza are crucial for preventing severe outcomes. This study provides new insights for early detection and disease progression monitoring, offering new biomarkers to aid in the assessment of disease severity and personalized treatment strategies for influenza patients.
The pronounced downregulation of glycerophospholipid levels (81.8% in influenza patients vs. controls; 83.3% in critical vs. mild), a central factor in influenza pathogenesis, is consistent with emerging evidence linking lipid metabolism to viral replication and host defense10. As essential components of cell membranes and signaling molecules, glycerophospholipids such as phosphatidylcholine (PC) and lysophospholipids (LysoPCs) may play dual roles: their depletion could reflect both the viral exploitation of membrane components for assembly11 and host-directed anti-inflammatory responses. Notably, LysoPC(20:0/0.0) exhibited consistent negative correlations with proinflammatory markers (WBC, CRP) and positive associations with lymphocytes, suggesting its potential as a biomarker for immune homeostasis. This observation corroborates studies showing improved phospholipid profiles following antiviral therapy12,13, suggesting therapeutic avenues targeting lipid metabolism.
One study revealed that after influenza virus infection, the levels of PC and lysophospholipid (Lyso) in the blood significantly decrease11. Our study revealed that the expression of PC was downregulated in influenza patients, indicating the involvement of influenza. Therefore, there is a direct relationship between the induced immune response and lipid richness and metabolism in the body. This study revealed that the differences in the peripheral plasma metabolome between influenza patients and healthy controls were related mainly to primary bile acid biosynthesis, porphyrin metabolism and nitrogen metabolism. In addition, there were 11 differentially abundant metabolites. Glycerophospholipid metabolism among all the species, nine (81.8%) were significantly downregulated in the case group, indicating that in influenza patients, glycerophospholipid metabolism pathways may be downregulated. Our study revealed that, compared with those in patients with mild influenza, glycerophospholipid metabolism pathways in patients with critical influenza were significantly downregulated.
In the metabolomic study of influenza, glycerophospholipids are as one of the main components of the cell membrane; the germination of influenza virus is inseparable from the assembly of the virus on the host plasma membrane, and the viral assembly process is closely related to membrane rafts14. Therefore, metabolic changes in glycerophospholipids during influenza virus infection are important. Glycerophospholipids and their metabolites may participate in the regulation of inflammation and immune function13,15, which is particularly important for understanding the immunopathological mechanisms associated with influenza. Some studies13,16 have shown that after antiviral treatment, glycerophospholipid and phospholipid disorders are alleviated, mainly through the regulation of the metabolic pathways of glycerophospholipids, which may have an inhibitory effect on the assembly process of the influenza virus. The composition and abundance of glycerophospholipids may change during influenza infection, and these changes can serve as biomarkers of influenza infection or its severity.
The enrichment of amino acid-related pathways (taurine/hypotaurine, glycine/serine/threonine metabolism) in severe/critical revealed metabolic adaptations to increased immune activity. The dual role of taurine as an antioxidant and bile acid conjugate may explain its depletion in severe—a phenomenon previously linked to H1N1-associated ARDS17. The concurrent decrease in the levels of glycine and serine, amino acids critical for glutathione synthesis and viral replication, suggests a tug-of-war between host antioxidant defense and viral resource utilization18. These pathways offer discriminative potential beyond conventional biomarkers, as evidenced by their specificity for H1N1-associated ARDS compared with bacterial pneumonia17. Moreover, our study revealed that the differentially abundant metabolites in the peripheral blood of patients with severe and those with mild were enriched mainly in amino acid metabolism pathways, including propionate metabolism; glycine, serine and threonine metabolism; valine, leucine and isoleucine biosynthesis; and taurine and hypotaurine metabolism. Taurine is an important compound for bile acid binding in the liver. A previous study by Lu et al.19. revealed that the peroxidase type 2 multifunctional enzyme (HSD17B4) that converts dihydroxy-cholesterol-CoA to trihydroxy-β-cholesterol-CoA is regulated by NF-κB, suggesting that bile acid biosynthesis is closely related to inflammatory mechanisms. In addition, taurine is an important intracellular free amino acid that is considered an antioxidant and neuromodulator and participates in the regulation of the osmotic pressure in the neural retina and brain20. A previous 1 h-NMR plasma metabolomics study21 compared H1N1 pneumonia patients with control individuals, and the results revealed that the metabolism of glycine, serine and threonine can differentiate H1N1 patients from patients with bacterial pneumonia; a decrease in glycine, serine and threonine concentrations in H1N1 patients may be related to the metabolic consumption of these amino acids by the virus. Compared with H1N1-associated ARDS and bacterial pneumonia-associated ARDS, the metabolism of taurine and hypotaurine is increased in patients with H1N1-associated ARDS, indicating that the metabolism of taurine and hypotaurine may be involved in the development of severe influenza in patients with HINI17. Changes in the levels of taurine and hypotaurine metabolism may serve as biomarkers for the severity of influenza, and whether taurine and hypotaurine can be used as therapeutic targets for the prevention of severe influenza needs to be determined via further exploration.
The distinct glycerophospholipid profiles observed across influenza severity groups revealed metabolite-specific correlations with immune and cellular injury markers, providing mechanistic insights into disease progression. In influenza patients, the marked downregulation (81.8%) of glycerophospholipid metabolites—particularly LysoPC (20:0/0.0) and three PC species—and their negative correlation with systemic inflammation (WBC, neutrophils, CRP, PCT) suggest depletion of membrane-stabilizing lipids during viral infection. The positive associations of these metabolites with lymphocyte counts imply that these metabolites may support adaptive immune cell function, potentially by maintaining membrane integrity or modulating lipid-mediated signaling pathways22. This dual role positions LysoPCs as both biomarkers of immune dysregulation and therapeutic targets for mitigating inflammation22.
The upregulated LysoPA species (16:0/0:0, 18:1/0:0) exhibited proinflammatory tendencies through negative correlations with lymphocytes/monocytes and positive associations with creatine kinase isoenzyme, a marker of myocardial stress. This finding is consistent with the known role of LysoPA in promoting endothelial activation and cytokine release, potentially exacerbating multiorgan involvement in severe influenza23. Conversely, the downregulated LysoPC(20:0/0.0) displayed protective correlations (positive with lymphocytes/monocytes, negative with WBC/neutrophils), suggesting that its depletion may impair the resolution of inflammation22.
Critical revealed a striking link between LysoPC(20:0/0.0) depletion and elevated LDH levels, directly connecting glycerophospholipid homeostasis to the severity of cellular injury. The parallel positive correlation between upregulated PE(20:2/14:0) and LDH implies compensatory membrane remodeling during catastrophic cell lysis, potentially through increased phosphatidylethanolamine synthesis in stressed organelles24.
Notably, the 15 conserved differentially abundant metabolites (e.g., sphingolipids and alkaloids) across severity strata highlight the core metabolic pathways disrupted by influenza. Although glycerophospholipid alterations are observed in other infections, our stratification by confirmed H1N1 severity and the consistent depletion of LysoPC(20:0/0:0) from mild to critical supports its specific prognostic relevance to influenza A. Comparative studies with other viruses are indeed warranted. It is also plausible that influenza strain evolution and regional variation may influence host metabolic responses. Therefore, further validation across seasons and geographic locations is warranted to assess the stability and generalizability of these metabolic biomarker. Since influenza A virus evolves yearly, and our cohort is geographically restricted to one center in China, external validation in diverse cohorts and across different seasons is necessary to assess biomarker robustness.
Limitations
This study has several limitations. Firstly, it was conducted in a single center with a relatively limited number of severe and critical, which may restrict the generalizability of our findings. Secondly, although we identified potential biomarkers specific to influenza A, we did not include control groups with other viral or bacterial respiratory infections, making it difficult to fully assess the specificity of the observed metabolic changes. Thirdly while clinical covariates such as age and comorbidity were recorded, multivariate adjustment was not performed in the primary analysis due to limited sample size. These limitations warrant further validation in larger, multi-center cohorts and in comparative studies with other infectious diseases. What’s more, one limitation of this study is the relatively small sample size, which restricted our ability to perform additional targeted validation or generate a comprehensive heatmap to assess metabolite trends across different disease severity groups. Future studies with larger cohorts are warranted to systematically explore dynamic metabolite changes from mild to critical illness.
Although OPLS-DA was applied to distinguish metabolic profiles among groups, we acknowledge that in some comparisons (mild vs. severe), the Q² values were modest (< 0.5), suggesting limited predictive power. Future studies with larger cohorts and alternative models such as PLS-DA may provide more robust discrimination and account for within-group variation.
Conclusion
In summary, our findings establish glycerophospholipid metabolism as a central player in the pathogenesis and progression of influenza A. Specifically, LysoPC(20:0/0.0) and components of the taurine pathway have emerged as promising biomarkers for early severity prediction and personalized therapeutic intervention. Our study underscores the critical role of glycerophospholipids in viral replication and the host immune response, and these metabolic changes provide new insights into the biology of host‒virus interactions. Disruption of glycerophospholipid metabolism may serve as a valuable biomarker for the diagnosis and severity assessment of influenza infection, offering new avenues for future therapeutic strategies.
Data availability
Data is provided within the manuscript or supplementary information files.The de-identified raw LC–MS data and clinical metadata supporting the findings of this study are available from the corresponding author upon reasonable request. The processed metabolite dataset is being deposited in the MetaboLights database and will be publicly accessible upon publication.
References
Kalil, A. C. & Thomas, P. G. Influenza virus-related critical illness: pathophysiology and epidemiology. Crit. Care. 23 (1), 258 (2019).
Joshua, S. & Hoyle, K. M. Downard 1High resolution mass spectrometry of respiratory viruses: beyond MALDI-ToF instruments for next generation viral typing, subtyping, variant and sub-variant identification. Analyst 148 (18), 4263–4273. https://doi.org/10.1039/d3an00953j (2023).
Sindelar, M. et al. Longitudinal metabolomics of human plasma reveals prognostic markers of COVID-19 disease severity. Cell. Rep. Med. 17 (8), 100369doi. https://doi.org/10.1016/j.xcrm.2021.100369 (2021).
Cui, L. et al. Metabolomics investigation reveals metabolite mediators associated with acute lung injury and repair in a murine model of influenza pneumonia. Sci. Rep. 6, 26076. https://doi.org/10.1038/srep26076 (2016).
Keshavarz, M. et al. Metabolic host response and therapeutic approaches to influenza infection. Cell. Mol. Biol. Lett. 25, 15. https://doi.org/10.1186/s11658-020-00211-2 (2020).
Tian, X. et al. Metabolomic analysis of influenza A virus A/WSN/1933 (H1N1) infected A549 cells during first cycle of viral replication. Viruses 11 (11), 1007. https://doi.org/10.3390/v11111007 (2019).
Sumon, M. S. I., Hossain, M. S. A., Al-Sulaiti, H. & Yassine, H. M. and M. E. H. Chowdhury. Enhancing influenza detection through integrative machine learning and nasopharyngeal metabolomic profiling: A comprehensive study. Diagnostics (Basel) 14 (19), 2214. https://doi.org/10.3390/diagnostics14192214 (2024).
Consensus Expert Group on Antiviral Treatment of adult influenza. Expert consensus on antiviral therapy influenza in adults. Chin. J. Infect. Dis. 40 (11), 641–655 (2022).
Uyeki, T. M., Hui, D. S., Zambon, M., Wentworth, D. E. & Monto, A. S. Influenza Lancet ; 400(10353): 693–706. DOI: https://doi.org/10.1016/s0140-6736(22)00982-5. (2022).
Tanner, L. B., Chng, C., Guan, X. L., Lei, Z. & Rozen, S. G. Wenk. Lipidomics identifies a requirement for peroxisomal function during influenza virus replication. J. Lipid Res. 55 (7), 1357–1365. https://doi.org/10.1194/jlr.M049148 (2014).
Xue, M. et al. Effect of a functional phospholipid Metabolome-Protein association pathway on the mechanism of COVID-19 disease progression. Int. J. Biol. Sci. 18 (12), 4618–4628. https://doi.org/10.7150/ijbs.72450 (2022).
Tisoncik-Go, J. et al. Integrated omics analysis of pathogenic host responses during pandemic H1N1 influenza virus infection: the crucial role of lipid metabolism. Cell. Host Microbe. 19 (2), 254–266. https://doi.org/10.1016/j.chom.2016.01.002 (2016).
Ma, Q. et al. Corrigendum: investigating the effects of Liushen capsules (LS) on the metabolome of seasonal influenza: A randomized clinical trial. Front. Pharmacol. 13, 1036927. https://doi.org/10.3389/fphar.2022.1036927 (2022).
Gerl, M. J. et al. Quantitative analysis of the lipidomes of the influenza virus envelope and MDCK cell apical membrane. J. Cell. Biol. 196 (2), 213–212. https://doi.org/10.1083/jcb.201108175 (2012).
Xu, J. et al. Plasma metabolomic profiling of patients recovered from coronavirus disease 2019 (COVID-19) with pulmonary sequelae 3 months after discharge. Clin. Infect. Dis. 73 (12), 2228–2239. https://doi.org/10.1093/cid/ciab147 (2021).
Feng, J. et al. Mechanism of Jinzhen oral liquid against influenza-induced lung injury based on metabonomics and gut Microbiome. J. Ethnopharmacol. 303, 115977. https://doi.org/10.1016/j.jep.2022.115977 (2023).
Lee, C. H. et al. Using a targeted metabolomics approach to explore differences in ARDS associated with COVID-19 compared to ARDS caused by H1N1 influenza and bacterial pneumonia. Crit. Care. 28 (1), 63. https://doi.org/10.1186/s13054-024-04843-0 (2024).
Pohl, M. O. et al. Identification of Polo-like kinases as potential novel drug targets for influenza A virus. Sci. Rep. 7 (1), 8629. https://doi.org/10.1038/s41598-017-08942-7 (2017).
Lu, X. et al. NF-κB increased expression of 17β-hydroxysteroid dehydrogenase 4 promotes HepG2 proliferation via inactivating estradiol. Mol. Cell. Endocrinol. 401, 1–11. https://doi.org/10.1016/j.mce.2014.11.016 (2015).
Jakaria, M. et al. Taurine and its analogs in neurological disorders: focus on therapeutic potential and molecular mechanisms. Redox Biol. 24, 101223. https://doi.org/10.1016/j.redox.2019.101223 (2019).
Banoei, M. M. et al. Plasma metabolomics for the diagnosis and prognosis of H1N1 influenza pneumonia. Crit. Care. 21 (1), 97. https://doi.org/10.1186/s13054-017-1672-7 (2017).
Chang, Y. et al. A targeted metabolomics approach for sepsis-induced ARDS and its subphenotypes. Crit. Care. 27 (1), 263. https://doi.org/10.1186/s13054-023-04552-0 (2023).
Cummings, R., Parinandi, N., Wang, L., Usatyuk, P. & Natarajan, V. Phospholipase D/phosphatidic acid signal transduction: role and physiological significance in lung. Mol. Cell. Biochem. 234–235 (1–2), 99–109 (2002).
Naicker, T. et al. HIV associated preeclampsia: A multifactorial appraisal. Int. J. Mol. Sci. 22 (17), 9157. https://doi.org/10.3390/ijms22179157 (2021).
Acknowledgements
This study was approved by the Ethics Committee of the Xi’an Jiaotong University Second Affiliated Hospital (2022288). This study was supported by the Key Project of the Natural Science Basic Research Program of Shaanxi Province (No. 2025JC-QYXQ-046 to Li YP), Xi ‘an Jiaotong University “Basic-Clinical” Fusion Innovation Project (No. YXJLRH2022067 to Li YP); the Shaanxi Provincial College Student Innovation Training Program (No. S202310698650 to Wang Xinyu) and the Shaanxi Province Natural Science Basic Research Program Project (No. 2020JM-408 to Zhai Song).
Author information
Authors and Affiliations
Contributions
Yaping Li: Conceptualization, Methodology, Software, Validation, Formal analysis, Investigation, Resources, Data curation, Writing original draft preparation, Writing review and editing, Visualization. Ting Li: Conceptualization, Methodology, Software, Validation, Formal analysis, Investigation, Resources, Data curation, Writing original draft preparation, Writing review and editing, Visualization. Min Liu: Conceptualization, Methodology, Software, Validation, Formal analysis, Investigation, Resources, Data curation, Visualization. XinYu Wang: Conceptualization, Investigation, Resources, Data curation, Visualization. Song Zhai: Conceptualization, Methodology, Software, Validation, Formal analysis, Investigation, Data curation, Visualization. Ruihong Qin and Chenrui Liu: Conceptualization, Methodology, Software, Validation, Formal analysis, Visualization. Yuanyi Chen and Meng Zhang: Conceptualization, Methodology, Software, Validation, Formal analysis, Investigation, Resources, Data curation, Visualization. Xiaoli Jia: Conceptualization, Writing review and editing, Supervision, Project administration, Funding acquisition.All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, Y., Li, T., Liu, M. et al. Metabolomic and clinical feature analyses of plasma from influenza A patients. Sci Rep 15, 42616 (2025). https://doi.org/10.1038/s41598-025-26805-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-26805-4