Abstract
Tens of millions of people worldwide travel to high-altitude regions (> 2500 m) every year for short-term work, tourism and pilgrimage. Hypoxia, resulting in a range of gastrointestinal disorders, is a significant physiological challenge in high-altitude environments. The gastrointestinal microbiota is closely linked to the mucosal barrier and our previous research found that hypoxia-induced mucosal injury mainly occurs in stomach and small intestine. This study aimed to characterize the temporal dynamics of gastric and small intestinal microbiota by directly analyzing the luminal contents to elucidate site-specific microbial alterations and their potential role in mucosal damage under hypoxic conditions. C57BL/6 J mice were randomly assigned to four groups: normoxic control, 1-day hypoxic exposure, 3-day hypoxic exposure and 12-day hypoxic exposure. Fresh gastric and small intestinal contents were collected for 16S rRNA sequencing. The operational taxonomic unit (OTU) counts, α/β-diversity and temporal microbial abundance shifts across different durations of hypoxia exposure were analyzed. Our results found that short-term hypoxia (1–3 days) significantly reduced the OTU counts and diminished α/β-diversity in the stomach and small intestine. However, prolonged hypoxia (day 12) reversed the diminished OTU counts and α/β-diversity to near-normoxic levels. In the stomach, short-term hypoxia increased the abundances of the phylum Firmicutes and genus Lactobacillus and suppressed the levels of the phyla Verrucomicrobia, Candidate_division_TM7, Tenericutes, Actinobacteria, and Bacteroidetes and genera S24-7_unclassified and Bifidobacterium. However, at 12 days of hypoxia, these phyla and genera reverted to near-normoxic levels. In the small intestine, the relative abundances of the phylum Firmicutes and genus Lactobacillus increased and those of the phyla Actinobacteria and Verrucomicrobia and genera Bifidobacterium and Akkermansia decreased during short-term hypoxia. However, these altered bacteria returned to near-normoxic levels by day 12 of hypoxia. Temporal clustering revealed persistent taxon shifts; the relative abundances of the phyla Gemmatimonadetes and Acidobacteria and genera Gemmatimonas, Kocuria, Parasutterella, and Saccharopolyspora declined steadily, while Staphylococcus increased consistently in the stomach. In the small intestine, the relative abundance of the genus Stomatobaculum consistently increased in response to prolonged hypoxia. This study provided a time-resolved overview of gastric and small intestinal microbiota adaptations to hypoxia, identified potential candidate taxa involved in mucosal injury caused by hypoxia.
Similar content being viewed by others
Introduction
Tens of millions of people worldwide travel to high-altitude regions (> 2500 m) every year for short-term work, tourism and pilgrimage. Hypoxia represents a significant physiological challenge in high-altitude environments1. Hypoxia can cause gastrointestinal disorders, including anorexia, nausea, diarrhea, vomiting and gastrointestinal bleeding, which predominantly affect the upper gastrointestinal tract2,3,4,5. The gastrointestinal symptoms induced by hypoxia may be caused by mucosal barrier damage, which can lead to bacterial translocation5,6. Hypoxia influences the structure and function of the microbial community7,8,9. The microbiota influences the structural and functional integrity of the mucosal barrier via increased mucus secretion, inhibition of pathogenic bacteria, altered immune responses, and changes in intercellular tight junctions10,11,12.
Recently, the role of gut microbiota in adapting to the hypoxic environment at high altitudes has garnered increasing interest. In Mus musculus domesticus, the levels of obligate anaerobes positively correlate with altitude, and non-anaerobes negatively correlate with altitude13. The gut microbiota of Tibetan antelope at high altitudes exhibits a higher ratio of Firmicutes to Bacteroidetes than the gut microbiota of Tibetan sheep at low altitudes14. The α-diversity in humans and pigs at high altitudes is significantly lower than in their low-altitude counterparts15. Additionally, population-specific studies demonstrated that the abundance of aerobic bacteria is significantly diminished and the abundance of anaerobic bacteria is augmented in high-altitude populations16,17. A cohort study including 45 healthy individuals has shown that during high-altitude hypoxic exposure, the species and functional diversity of the gut microbiota significantly decreases, including specific changes in the community composition at the phylum and genus levels18. The gut microbiota is significantly different between Han individuals living in the plains and Tibet; the abundance of Firmicutes was significantly enriched in Han individuals living at high altitudes, and the abundance of Bacteroidetes was higher in Han individuals living at low altitudes19. Faecalibacterium, Bacteroides, and Bifidobacterium positively correlate with altitude exposure, and Ruminococcaceae, Prevotella, and Lachnospiraseae negatively correlate with altitude in Tibetan populations living at altitudes between 2800 and 4500 m20.
Gastrointestinal symptoms gradually diminish to a certain extent as individuals adapt to high-altitude hypoxic environments21,22. Interestingly, similar adaptations to high-altitude hypoxic environments are present in the gut microbiota. Gut microbiota α- and β-diversity are significantly reduced at the onset of high-altitude hypoxic exposure but subsequently recover during prolonged exposure; the genus Blautia A may be a key bacterium for this adaptation to high-altitude hypoxia18. Lactobacillus abundance increases on the first 2 days and decreases on the eighth day of exposure to high-altitude hypoxia in healthy males21. Our preliminary study demonstrated that hypoxia caused mucosal damage in the upper gastrointestinal tract, primarily in the stomach and small intestine, rather than in the large intestine23. Due to the difficulty and low acceptability of samples collected from the stomach and small intestine, most microbiota studies are currently conducted by collecting feces. This inevitably incorporates the influence of the large intestinal microbiota, complicating efforts to obtain accurate samples and analyses of gastric and small intestinal microbiota. Therefore, we constructed a hypoxic mouse model and collected the contents of the stomach and small intestine at different time points for microbiota analysis to explore the microbial changes at the site of hypoxic gastrointestinal mucosal injury and to investigate the potential role of microbiota in this context.
Materials and methods
Mice and hypoxic models
Male and female C57BL/6 J mice (6–8 weeks old, 18–22 g) were obtained from Charles River Laboratories (Beijing, China) and maintained in the Tsinghua University Animal Facility under specific pathogen-free conditions with a 12 h dark/light cycle and free access to food and water. The mice were fed a standard rodent breeding diet comprising 20% crude protein, 3.7% crude fiber, and approximately 52.7% carbohydrates. All experimental protocols were approved by the Animal Care and Use Committee of Tsinghua University (protocol #24-CZJ4), and all methods were performed in accordance with the relevant guidelines and regulations. Also, the study is reported in accordance with ARRIVE guidelines. Following a 1-week acclimation period, age- and sex-matched mice were randomly assigned to four experimental groups (n = 6 per group): normoxic control (N), 1-day hypoxic exposure (HD1), 3-day hypoxic exposure (HD3), and 12-day hypoxic exposure (HD12). All hypoxia-exposed mice were euthanized immediately after their designated exposure period. Hypoxic conditions were generated using a ventilated hypoxic chamber system (LP-1500, Shanghai Yuyan Instruments, Shanghai, China) to simulate an altitude of 5500 m above sea level.
Sample collection
Mice were placed in a closed induction chamber and exposed to medical oxygen containing 5% isoflurane (v/v) until deep anesthesia was achieved. Following this, cervical dislocation was performed immediately. After euthanizing the mice, the fresh gastric and small intestinal contents (including the entire duodenum, jejunum and ileum) were collected in sterile tubes, snap-frozen in liquid nitrogen and stored at − 80 °C for 16S rRNA sequencing.
16S rRNA sequencing
Total DNA was extracted from the samples using the Omega DNA Extraction Kit (M5635-02, Omega Bio-Tek, Norcross, GA, USA) according to the manufacturer’s instructions. The quantity and quality of extracted DNA were assessed using a Nanodrop NC2000 spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA) and 1.2% agarose gel electrophoresis, respectively. Forward primer 338F (5′-ACTCCTACGGGGAGGCAGCA-3′) and reverse primer 806R (5′-GGACTACHVGGGTWTCTAAT-3′) were used for amplification of the V3-V4 region of the bacterial 16S rRNA gene. The amplified PCR products were recovered using magnetic beads and quantified by fluorescence.
Sequencing libraries were prepared using the TruSeq Nano DNA LT Library Prep Kit (Illumina, San Diego, CA, USA), and index codes were incorporated. The quality of the libraries was assessed using an Agilent Bioanalyzer 2100 system (Agilent Technologies Inc., Santa Clara, CA, USA), and the libraries were quantified using a Promega QuantiFluor system (Promega Corporation, Madison, WI, USA) before sequencing. Sequencing of the 16S rRNA V3-V4 region (~ 480 bp amplicon) was conducted on an Illumina NovaSeq 6000, producing final reads of approximately 430 bp following the trimming of primer and barcode sequences.
Data analysis
Sequence data were processed and filtered using the Mothur pipeline. Low-quality sequences with an average Phred score ≤ 20 or containing excessively long homopolymers were removed. Further filtering excluded sequences with primer mismatches, as well as those shorter than 200 bp or longer than 500 bp. Chimeric sequences were identified and eliminated using UCHIME in reference to the gold dataset. The resulting high-quality sequences were subsequently used for operational taxonomic unit (OTU) clustering and analysis. Sequence clustering was performed using USEARCH (sequence similarity was set to 97%), and the representative OTU sequences were used for taxonomic assignment, referencing the Silva database. The community composition of each sample was determined across various taxonomic levels of species classification. Meanwhile, all raw paired-end sequences were processed using the QIIME2 pipeline (version 2025.4). Barcodes were trimmed with Cutadapt, and reads were merged using Vsearch. Low-quality sequences were filtered out with Trimmomaticall. Then clean reads were used to generate amplicon sequence variant (ASV) and ASV table by the DADA2 plugin with default parameters, followed by taxonomic annotation using the SILVA 132 (16S_V34) database. Based on the results of the OTU and ASV analysis, the Shannon and Chao1 α-diversity indices were calculated to reflect the species abundance and diversity of the community, and the principal components and principal coordinate analyses (PCoA) were conducted to illustrate the β-diversity of the community. To characterize temporal microbial abundance shifts across different hypoxia exposure durations, OTUs from the N, HD1, HD3 and HD12 groups were classified into distinct clusters at the phylum and genus levels. Differentially abundant taxa between groups were identified using Metastats, which applied a non-parametric permutation-based statistical test tailored for sparse microbial compositional data. Differences in OTU counts across the four groups were assessed by one-way analysis of variance, followed by Bonferroni’s post hoc test. A P value < 0.05 was considered statistically significant.
Results
Alterations in OTU counts in the stomach and small intestine over time
Samples were obtained as illustrated in Fig. 1A. The gastric and small intestinal contents were extracted and subjected to sequencing. No significant differences in the mean sequence counts (approximately 10,000) in the stomach and small intestine were detected among the four groups. However, the OTU counts in the stomach and small intestine decreased significantly in the HD1 and HD3 groups compared with the N group, reflecting a decreased microbial richness during short-term hypoxia. By day 12 of hypoxia, the OTU counts in the hypoxia groups rebounded to levels similar to the N group (Figs. 1B,D, S1). As shown in the Venn diagrams, the OTU counts in the stomach were 1157 for the normoxic control group, 222 for the 1-day hypoxic exposure group, 116 for the 3-day hypoxic exposure group, and 1423 for the 12-day hypoxic exposure group. The maximum number of overlapping OTUs in the stomach between the N and HD12 groups was 933 (Fig. 1C). In the small intestine, the OTU counts in the N, HD1, HD3 and HD12 groups were 459, 100, 101 and 421, respectively. The maximum number of overlapping OTUs between the N and HD12 groups was 285 (Fig. 1E).
Operational taxonomic unit (OTU) counts were significantly reduced on hypoxic days 1 and 3 but recovered by day 12. Mice were placed in hypoxic chambers to simulate an altitude of 5500 m above sea level for 1, 3, or 12 days (n = 6 per group). (A) Schematic of sample collection. OTUs and sequence counts reflecting the microbiota communities in the stomach (B) and small intestine (D). Venn diagrams of OTUs counts reflecting the microbiota communities in the stomach (C) and small intestine (E). HD hypoxic exposure days; N = Normoxic control; S = Stomach; and SB small bowel.
Alterations in α- and β-diversity in response to short-term and prolonged hypoxia
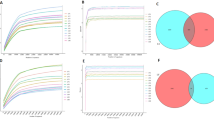
To assess the microbial community diversity, α- and β-diversity were analyzed. Both gastric and small intestinal samples exhibited progressive declines in α-diversity in response to short-term hypoxia (1–3 days), followed by a significant recovery by day 12 of hypoxia (Fig. 2A,C, S4A, S4C). Furthermore, the unweighted and weighted PCoA revealed distinct clustering of the 1-day hypoxic exposure group compared with the 3-day hypoxic exposure group. However, the 12-day hypoxic exposure samples converged with the normoxic control clusters (Fig. 2B,D, S4B, S4D).
Alpha and beta diversity of microbiota significantly decreased on days 1 and 3 but recovered by day 12 of hypoxic conditions. The α-diversity of the microbiota communities in the stomach (A) and small intestine (C). The β-diversity of the microbiota communities in the stomach (B) and small intestine (D). HD hypoxic exposure days; N = Normoxic control; S = Stomach; and SB small bowel.
Alterations in the microbial community in short-term versus prolonged hypoxia
The composition of the microbial community at the phylum and genus levels was analyzed. The microbiota in the stomach was predominantly composed of Firmicutes (60%), Bacteroidetes (22%), and Actinobacteria (10%) at the phylum level. Short-term hypoxia (1–3 days) induced increases in Firmicutes abundance and suppression of Bacteroidetes and Actinobacteria. By day 12 of hypoxia, these altered phyla reverted to near-normoxic levels. At the genus level, the relative abundance of Lactobacillus within Firmicutes was significantly elevated by day 3 of hypoxia, and the abundance of Bifidobacterium within Actinobacteria and S24-7_unclassified within Bacteroidetes was significantly decreased during short-term hypoxia; as the duration of hypoxia increased, the abundance of these two genera recovered (Fig. 3A, S2). Heatmap analysis showed a reduction in the phyla Verrucomicrobia, Candidate_division_TM7, Tenericutes, Actinobacteria, and Bacteroidetes during short-term hypoxia, followed by recovery by prolonged hypoxia. At the genus level, the abundance of most bacteria also recovered on the 12th day of hypoxia (Fig. 3B, S2). In the small intestine, the phyla Firmicutes, Actinobacteria, and Verrucomicrobia exhibited the highest relative abundance under normoxia. On the first and third days of hypoxia, Firmicutes increased and Actinobacteria and Verrucomicrobia declined in the small intestine; abundances of these phyla rebounded to levels comparable to normoxia by day 12. At the genus level, the relative abundance of Lactobacillus increased to over 90% on the third day of hypoxia but returned to approximately 30% on the twelfth day of hypoxia. The relative abundances of the genera Bifidobacterium and Akkermansia decreased consistently on days 1 and 3 of hypoxia but recovered to levels approaching normoxia after prolonged hypoxia (Fig. 3C, S3). The abundance of most bacteria changed during the initial days of hypoxia (days 1 and 3) but recovered on day 12 at the phylum and genus levels (Fig. 3D, S3).
Composition of the microbial community was significantly altered on hypoxic days 1 and 3 but largely recovered by day 12. The relative abundances of the microbial community composition in the stomach (A) and small intestine (C) at the phylum and genus levels. Heatmaps of the microbiota communities in the stomach (B) and small intestine (D) at the phylum and genus levels. HD hypoxic exposure days; N = Normoxic control; S = Stomach; and SB small bowel.
Two bacterial phyla and six bacterial genera changed during long-term hypoxia
As the duration of hypoxia increased, most microbiota in the stomach and small intestine adapted to the hypoxic conditions. Therefore, to determine if some bacteria exhibited unidirectional trends in response to hypoxia, temporal clustering was analyzed. The microbiota from the stomach and small intestine were grouped into four distinct clusters at the phylum and genus levels (Fig. 4A). In the stomach, the abundances of Gemmatimonadetes and Acidobacteria progressively declined at the phylum level in response to prolonged hypoxia (Fig. 4B). This reduction was primarily driven by decreases in the genera Gemmatimonas (belonging to the phylum Gemmatimonadetes) and Kocuria (belonging to the phylum Actinobacteria). Additionally, the relative abundance of Staphylococcus (belonging to the phylum Firmicutes) increased, and the relative abundances of Parasutterella (belonging to the phylum Proteobacteria) and Saccharopolyspora (belonging to the phylum Actinobacteria) decreased. In the small intestine, the relative abundance of Stomatobaculum (belonging to the phylum Firmicutes) consistently increased during prolonged hypoxia (Fig. 4B).
Two bacterial phyla and six bacterial genera exhibited continuous upward or downward trends during prolonged hypoxia. (A) Analysis of the time series clustering of microbiota in the stomach and small intestine at the phylum and genus levels. (B) Relative abundances of Gemmatimonadetes and Acidobacteria at the phylum level and Staphylococcus, Parasutterella, Kocuria, Gemmatimonas, Saccharopolyspora, and Stomatobaculum at the genus level (marked in red). HD hypoxic exposure days; N = Normoxic control.
Discussion
Hypoxia-induced damage to the gastrointestinal mucosa occurs in the stomach and small intestine. To determine the changes in the microbiota, the contents were collected directly from the stomach and small intestine to avoid large intestinal microbiota interferences. Our results revealed a biphasic pattern of initial microbial diversity suppression under hypoxic conditions, followed by a recovery upon prolonged hypoxia. Additionally, it elucidated the shifts in the composition of gastric and small intestinal microbiota corresponding to varying durations of hypoxia. Hypoxia leads to mucosal damage in the gastrointestinal tract23 and affects the structure and function of the microbial community7,8,9. Our results demonstrated that short-term hypoxia (1 and 3 days) induced significant reductions in OTU counts and α- and β-diversity in both the gastric and small intestinal microbial communities; these results are consistent with previous studies8,24. Extended exposure to hypoxia (12 days) restored microbial diversity and community structure to levels approaching those of the normoxia group, as confirmed by Bai et al.8. Short-term hypoxia increased the phylum Firmicutes and genus Lactobacillus and decreased the phylum Bacteroidetes similar to the pattern of changes observed in Tibetan antelope and populations adapted to high-altitude environments19,21. However, these changes in phyla and genera were attenuated by day 12 of hypoxia. The recovery of microbial diversity by day 12 of hypoxia correlates with clinical reports of intestinal permeability recovery and reduction of gastrointestinal symptoms in humans during prolonged high-altitude acclimatization21,22.
The microbial dynamics illustrated a hypoxic-driven succession. Initially (1–3 days), hypoxia acted as a strong selective pressure, promoting facultative anaerobes like Lactobacillus to dominance. Their rapid acid production suppressed diversity and inhibited competitors. This initial phase was followed by a restructuring (day 12), where the early dominants receded and a new, stable community adapted to prolonged hypoxia emerged. A notable observation from Supplemental Figs. S2 and S3 was the initial increase (not significant) in Clostridium_sensu_stricto_1 (an obligate anaerobe) under 1-day hypoxia in both the stomach and small intestine, which contrasted sharply with its subsequent complete disappearance at the 3-day hypoxia time point. The initial increase of Clostridium_sensu_stricto_1 under 1-day hypoxia was consistent with its character as an obligate anaerobe. Its subsequent disappearance by day 3 was likely attributed to the massive proliferation of Lactobacillus (relative abundance > 90%). The organic acids (e.g. lactic acid, acetic acid) and bacteriocins produced by Lactobacillus are known to effectively suppress clostridial species25,26. Additionally, an inverse correlation was observed between Lactobacillus and Bifidobacterium, suggesting possible competitive exclusion driven by their physiological differences. Lactobacillus, with its rapid growth and acid tolerance, outcompeted other taxa by rapidly acidifying the environment upon hypoxia induction. This acid-driven suppression might initially limit Bifidobacterium, which is typically more acid-sensitive27. The subsequent recovery of Bifidobacterium by day 12 pointed to niche partitioning upon community stabilization, likely resulting from a shift in resources and reduced competitive pressure from Lactobacillus. It is important to note that Lactobacillus constitutes a highly complex and diverse genus encompassing numerous species, some of which are well-known for their beneficial roles, while others are either poorly characterized or of uncertain significance. Therefore, it is imperative that subsequent experiments pinpoint the exact species within this genus that were elevated.
Certain bacteria failed to recover during prolonged hypoxia. In the stomach, the relative abundances of the phyla Gemmatimonadetes and Acidobacteria and genera Gemmatimonas, Kocuria, Parasutterella and Saccharopolyspora were diminished, and the relative abundance of Staphylococcus steadily increased during prolonged hypoxia. In the small intestine, the relative abundance of the genus Stomatobaculum consistently increased during prolonged hypoxia. However, multiple sets of zero values exist for these bacteria. Thus, more research is needed to determine whether they contribute to chronic gastrointestinal discomfort associated with hypoxia.
Conclusions
This study provided a time-resolved overview of microbiota adaptations to hypoxia in the stomach and small intestine and identified taxa that may be involved in mucosal injury caused by hypoxia.
Data availability
The 16S rRNA sequencing data in this study are available in GenBank under SRA accession: PRJNA1249210.
Abbreviations
- OTU:
-
Operational taxonomic unit
- PCoA:
-
Principal coordinates analyses
- ASV:
-
Amplicon sequence variant
References
Richalet, J.-P., Hermand, E. & Lhuissier, F. J. Cardiovascular physiology and pathophysiology at high altitude. Nat. Rev. Cardiol. 21, 75–88. https://doi.org/10.1038/s41569-023-00924-9 (2024).
Anand, A. C., Sashindran, V. K. & Mohan, L. Gastrointestinal problems at high altitude. Trop. Gastroenterol. 27, 147–153 (2006).
Fruehauf, H. et al. Evaluation of acute mountain sickness by unsedated transnasal esophagogastroduodenoscopy at high altitude. Clin. Gastroenterol. Hepatol. 18, 2218-2225.e2212. https://doi.org/10.1016/j.cgh.2019.11.036 (2020).
Wu, T. Y. et al. High-altitude gastrointestinal bleeding: an observation in Qinghai-Tibetan railroad construction workers on Mountain Tanggula. World J. Gastroenterol. 13, 774–780. https://doi.org/10.3748/wjg.v13.i5.774 (2007).
McKenna, Z. J. et al. High-altitude exposures and intestinal barrier dysfunction. Am. J. Physiol. Regul. Integr. Comp. Physiol. 322, R192-r203. https://doi.org/10.1152/ajpregu.00270.2021 (2022).
Khanna, K. et al. Effects of acute exposure to hypobaric hypoxia on mucosal barrier injury and the gastrointestinal immune axis in rats. High. Alt. Med. Biol. 20, 35–44. https://doi.org/10.1089/ham.2018.0031 (2019).
Han, N., Pan, Z., Liu, G., Yang, R. & Yujing, B. Hypoxia: The “Invisible Pusher” of gut microbiota. Front. Microbiol. 12, 690600. https://doi.org/10.3389/fmicb.2021.690600 (2021).
Bai, X. et al. Changes in the gut microbiota of rats in high-altitude hypoxic environments. Microbiol. Spectr. 10, e0162622. https://doi.org/10.1128/spectrum.01626-22 (2022).
Liu, D. et al. High-altitude-induced alterations in intestinal microbiota. Front. Microbiol. 15, 1369627. https://doi.org/10.3389/fmicb.2024.1369627 (2024).
Fukuda, S. et al. Bifidobacteria can protect from enteropathogenic infection through production of acetate. Nature 469, 543–547. https://doi.org/10.1038/nature09646 (2011).
Ma, J., Piao, X., Mahfuz, S., Long, S. & Wang, J. The interaction among gut microbes, the intestinal barrier and short chain fatty acids. Anim. Nutr. 9, 159–174. https://doi.org/10.1016/j.aninu.2021.09.012 (2022).
Ghosh, S., Whitley, C. S., Haribabu, B. & Jala, V. R. Regulation of intestinal barrier function by microbial metabolites. Cell. Mol. Gastroenterol. Hepatol. 11, 1463–1482. https://doi.org/10.1016/j.jcmgh.2021.02.007 (2021).
Suzuki, T. A., Martins, F. M. & Nachman, M. W. Altitudinal variation of the gut microbiota in wild house mice. Mol. Ecol. 28, 2378–2390. https://doi.org/10.1111/mec.14905 (2019).
Ma, Y. et al. Gut microbiota adaptation to high altitude in indigenous animals. Biochem. Biophys. Res. Commun. 516, 120–126. https://doi.org/10.1016/j.bbrc.2019.05.085 (2019).
Zeng, B. et al. Gut microbiota of Tibetans and Tibetan pigs varies between high and low altitude environments. Microbiol. Res. 235, 126447. https://doi.org/10.1016/j.micres.2020.126447 (2020).
Jia, Z. et al. Impacts of the plateau environment on the gut microbiota and blood clinical indexes in Han and Tibetan individuals. mSystems https://doi.org/10.1128/mSystems.00660-19 (2020).
Adak, A., Maity, C., Ghosh, K., Pati, B. R. & Mondal, K. C. Dynamics of predominant microbiota in the human gastrointestinal tract and change in luminal enzymes and immunoglobulin profile during high-altitude adaptation. Folia Microbiol. (Praha) 58, 523–528. https://doi.org/10.1007/s12223-013-0241-y (2013).
Su, Q. et al. Gut microbiota contributes to high-altitude hypoxia acclimatization of human populations. Genome Biol. 25, 232. https://doi.org/10.1186/s13059-024-03373-w (2024).
Li, L. & Zhao, X. Comparative analyses of fecal microbiota in Tibetan and Chinese Han living at low or high altitude by barcoded 454 pyrosequencing. Sci. Rep. 5, 14682. https://doi.org/10.1038/srep14682 (2015).
Lan, D. et al. Correlations between gut microbiota community structures of Tibetans and geography. Sci. Rep. 7, 16982. https://doi.org/10.1038/s41598-017-17194-4 (2017).
Karl, J. P. et al. Associations between the gut microbiota and host responses to high altitude. Am. J. Physiol. Gastrointest. Liver Physiol. 315, G1003-g1015. https://doi.org/10.1152/ajpgi.00253.2018 (2018).
Chen, Y. et al. Gastrointestinal syndrome encountered during a train voyage to high altitudes: A 14-day survey of 69 passengers in China. Travel Med. Infect. Dis. 59, 102718. https://doi.org/10.1016/j.tmaid.2024.102718 (2024).
Wang, D. et al. Oral administration of Robinia pseudoacacia L. flower exosome-like nanoparticles attenuates gastric and small intestinal mucosal ferroptosis caused by hypoxia through inhibiting HIF-1α- and HIF-2α-mediated lipid peroxidation. J. Nanobiotechnol. 22, 479. https://doi.org/10.1186/s12951-024-02663-6 (2024).
Wang, F., Zhang, H., Xu, T., Hu, Y. & Jiang, Y. Acute exposure to simulated high-altitude hypoxia alters gut microbiota in mice. Arch. Microbiol. 204, 412. https://doi.org/10.1007/s00203-022-03031-4 (2022).
Anjana, T. & Tiwari, S. K. Bacteriocin-producing probiotic lactic acid bacteria in controlling dysbiosis of the gut microbiota. Front. Cell. Infect. Microbiol. 12, 851140. https://doi.org/10.3389/fcimb.2022.851140 (2022).
Spehar, I. D. et al. Antimicrobial activity of bacteriocins of lactic acid bacteria on listeria monocytogenes, Staphylococcus aureus and Clostridium tyrobutyricum in cheese production. Mljekarstvo 70, 135–149. https://doi.org/10.15567/mljekarstvo.2020.0301 (2020).
Boylston, T. D., Vinderola, C. G., Ghoddusi, H. B. & Reinheimer, J. A. Incorporation of bifidobacteria into cheeses: Challenges and rewards. Int. Dairy J. 14, 375–387. https://doi.org/10.1016/j.idairyj.2003.08.008 (2004).
Acknowledgements
We would like to thank Zhijie Chang and Jianqiu Sheng for providing experimental resources.
Funding
Not applicable.
Author information
Authors and Affiliations
Contributions
[XCL], [DZW] and [JQS], designed the research. [XCL], [XXW] and [DZW] performed the experiments. [XCL], [XXW] and [DZW] analyzed and visualized the results. [XW], [MJZ] and [FLR] prepared animal resources. [XCL] drafted manuscript. [DZW] and [JQS] reviewed the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
All experimental protocols were approved by the Animal Care and Use Committee of Tsinghua University (protocol #24-CZJ4).
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liao, X., Wang, X., Wang, X. et al. Alterations in the gastric and small intestinal microbiota of mice exposed to short-term and long-term hypoxia. Sci Rep 15, 42615 (2025). https://doi.org/10.1038/s41598-025-26859-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-26859-4