Abstract
Peptide inhibitors represent a promising class of antiviral therapeutics, offering several advantages over traditional small-molecule drugs, including low toxicity, high specificity, and biocompatibility. However, rational and efficient design and optimization of inhibitor peptides remains a significant challenge to current methods. Here we show EvoPepFold, a genetic algorithm-based framework designed to generate inhibitory peptides. We evaluated EvoPepFold to design and optimize peptides targeting the SARS-CoV-2 main protease (Mpro). EvoPepFold was applied through two complementary strategies: molecular docking using the Rosetta suite, and peptide 3D modeling with ColabFold. The top candidates were further evaluated through molecular dynamics simulations to assess stability and interaction energy. Our results demonstrate that EvoPepFold successfully identified peptides with favorable binding affinities and stable protein-peptide interactions. These findings highlight the potential of evolutionary algorithms in guiding the rational design of peptide-based antivirals, contributing to ongoing efforts in peptide engineering for therapeutic applications.
Similar content being viewed by others
Introduction
Peptides are molecules composed of short chains of amino acids linked by peptide bonds, typically consisting of 2–50 residues1. In living beings, they play diverse roles in biological systems, acting as hormones, signaling molecules, neurotransmitters, and regulators of various physiological processes2. For example, in the case of infectious diseases, peptides can function as antiviral agents by interfering with key stages of the viral life cycle, such as entry, replication, or assembly. Additionally, peptide inhibitors represent a promising class of antiviral therapeutics, offering several advantages over traditional small-molecule drugs3.
Advances in synthetic approaches have enabled the modification of the biophysical and biochemical properties of peptides, making them promising candidates for drug development, mainly due to their low toxicity, high specificity, and biocompatibility4. Currently, more than 60 peptide-based drugs have been approved, with many others in clinical trials5. These compounds have demonstrated effectiveness in treating diseases such as cancer, type 2 diabetes, and autoimmune disorders, with exenatide derivatives being notable examples2. Another relevant example involves mirror-image peptides used in oncologic applications, which are composed of D-amino acid residues6. Their enhanced proteolytic stability and reduced immunogenicity confer significant advantages over conventional L-peptides. Consequently, mirror-image peptides and other oncolytic peptides are being actively explored for anticancer applications, with several in clinical trials7,8,9. Another example are the antiviral peptides. Antiviral peptides can be strategically developed to bind to vital viral enzymes or structural components, thereby blocking their activity. As a result, there is growing interest in the pharmaceutical industry in creating peptides that can interfere with essential protein–protein interactions required for viral replication10. These peptides emerged as promising allies in the fight against viral infections, such as COVID-19.
Threats to public health, such as the SARS-CoV-2 virus, highlight the need for methodologies to expedite the development of effective antiviral therapeutics11. While vaccination efforts have significantly mitigated the severity of the pandemic, the emergence of new variants and the need for treatments for infected individuals underscore the importance of antiviral drug discovery. In the particular case of SARS-CoV-2, the main protease (Mpro) is a crucial enzyme responsible for processing viral polyproteins. It has emerged as a prime target for therapeutic intervention due to its essential role in viral replication and its distinct substrate specificity compared to human proteases12. The unique preference of Mpro for a glutamine residue at the P1 position of its substrates presents an opportunity to design highly selective inhibitors13. However, the potential for drug resistance arising from mutations in the Mpro sequence highlights the need for innovative approaches to identify novel antiviral agents12.
Understanding protein–peptide interactions is essential for the rational design of new compounds with therapeutic and biotechnological potential5,14,15. As discussed, peptide inhibitors represent a promising class of antiviral therapeutics, offering several advantages over traditional small-molecule drugs3. Their inherent ability to interact with large protein surfaces with high specificity and potency makes them well-suited for targeting enzymes like Mpro3. However, designing peptides with high affinity to specific protein complexes is not trivial.
In this context, computational strategies have been widely employed to aid in designing peptides with enhanced affinity for the receptor. For example, molecular docking methods, such as HDOCK16,17, HPEPDOCK18, and Rosetta19,20, can help in identifying the binding poses of the peptide to the protein and predicting their binding energy. Although not explicitly designed for this purpose, AI-based structural modeling tools like AlphaFold Multimer21 have also been adopted to simulate protein-peptide interactions22. However, these tools do not perform regular molecular docking; instead, they model the entire complex using deep neural network techniques trained on large datasets of protein-protein interactions.
Furthermore, various computational methods can be employed to automate and optimize the generation of peptides with therapeutic properties. Genetic algorithms (GAs) are a class of optimization methods. They were developed inspired by evolution, which iteratively evolve a population of candidate solutions to improve their performance according to a defined fitness function. The use of these algorithms for sequence-based peptide optimization is already widely discussed in the literature23; however, to the best of our knowledge, the combination of molecular docking, AI-based modeling, molecular dynamics simulations, and genetic algorithms to optimize peptides has not yet been explored.
In this study, we aimed to identify peptides with the potential to bind to Mpro, thereby helping to develop new therapeutic candidates against SARS-CoV-2. To achieve this, we developed and evaluated the feasibility of an innovative computational pipeline composed of three main steps: (i) a genetic algorithm for the generation and optimization of peptide sequences based on evolutionary criteria; (ii) AlphaFold224, a deep learning-based tool used to predict the three-dimensional structures of the generated peptides; and (iii) the Rosetta software19,20 using its scoring function to estimate the binding energy between each peptide and Mpro. The function estimates this energy based on physicochemical criteria, serving as a relative indicator of complex stability and aiding in the selection of the most promising peptides. Lastly, we evaluated the best proposed peptides using molecular dynamics simulations.
Materials and methods
Data collection
We collected, from the Protein Data Bank (PDB)25, the 3D structure of the COVID-19 main protease (Mpro) in complex with an inhibitor N3 peptide (PDB ID: 6LU7) and defined this structure as the target. N3 is a peptide of six amino acid residues (sequence: “XAVLXX”, where X corresponds to a non-canonical amino acid).
Additionally, we collected 2355 peptides composed of 5 to 30 amino acids from the Propedia database26,27. These peptides were used to define the initial population for the genetic algorithm to improve on.
Mpro docking site definition
We used the binding position of the N3 peptide in the Mpro to define the binding region of interest (Fig. 1A, B). We selected residues with at least one atom within 5 Å of the N3 peptide, and described this region as the docking site contact interface (Fig. 1C). Finally, we removed the peptide from the complex (Fig. 1D) and used the remaining structure as a target for further analyses. The following 24 residues were selected from the Mpro interface of contact with the N3 peptide (Fig. 1E): T24, T25, T26, L27, H41, M49, F140, L141, N142, G143, S144, C145, H164, M165, E166, L167, P168, H172, D187, R188, Q189, T190, A191, and Q192.
Target structure. (A) COVID-19 main protease (Mpro – purple surface) complexed with the inhibitor N3 (green) (PDB ID: 6LU7). (B) Mpro residues directly interacting with the N3 peptide. (C) We selected residues up to 5Å from the N3 peptide (in orange). (D) Then, we removed the peptide from the complex. (E) MPRO docking target site residues: T24, T25, T26, L27, H41, M49, F140, L141, N142, G143, S144, C145, H164, M165, E166, L167, P168, H172, D187, R188, Q189, T190, A191, and Q192.
Our ultimate goal is to propose a peptide that binds to the Mpro structure with more affinity than the N3 peptide. Therefore, this hypothetical peptide should at least occupy the same binding site as the original peptide. To measure this, we define the residue occupancy (RO) parameter. The RO parameter indicates the percentage of the 24 amino acids of Mpro at least 5Å away from the original ligand. By default, the original peptide has RO = 100%. A docked peptide with an RO score equal to 0% is bound to a binding site different from the one targeted in this work.
Genetic algorithm overview
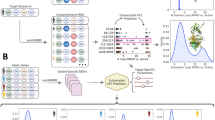
Figure 2 presents a general overview of the genetic algorithm (GA) proposed by this study. We implemented a GA designed to optimize peptide sequences to identify peptides with high binding affinity to the main protease (Mpro) of SARS-CoV-2. The initial dataset consisted of 2,355 peptides extracted from the Propedia database26,27. These peptides were docked to the Mpro structure, and the top 100 peptides were selected based on the Rosetta Energy Function score19,20 and the RO parameter. From this subset, 25 peptides were further selected to perform a grid search to determine the optimal parameters for the genetic algorithm.
Overview of the genetic algorithm. Peptides obtained from the Propedia database were initially docked to the Mpro structure. The top 100 results were selected as the initial population for the genetic algorithm. Two strategies were then employed: a docking-based approach (utilizing the Rosetta suite) and an AI-driven 3D modeling approach (utilizing ColabFold). A fitness function evaluated peptide candidates through a tournament selection scheme, and genetic operations were applied to peptide sequences to generate a new population. Each new set of peptide structures was modeled through docking (Rosetta) or 3D modeling (ColabFold), resulting in a population of 25 structures. These steps were iterated over 100 generations. Finally, the best peptides from each generation were selected based on the lowest docking scores.
The evolutionary process began with an initial population of 100 peptides. In each generation, new peptides were produced through two main genetic operators: crossover and mutation. The crossover operator recombined segments of parent peptides using variable crossover lengths, which were randomly chosen for each operation. The mutation operator introduced diversity by applying insertion, deletion, or substitution of amino acid residues at random positions in the peptide sequence.
Newly generated peptides were modeled using two strategies: docking protein-peptide using Rosetta and AI-modeling using ColabFold, an AlphaFold2-powered tool24 (further details are provided in the following subsections).
The fitness function used to evaluate each peptide was the docking score; however, the occupancy of the binding site (RO parameter) was used to filter out peptides that bind elsewhere. The highest-ranking peptides were selected to propagate the next generation. This iterative evolutionary process was repeated until convergence criteria were met, aiming to identify peptides with enhanced binding potential to Mpro.
Initial population
To define the initial population, docking was performed between the 2355 peptides collected from Propedia and the Mpro structure using PyRosetta4 - release 2024.3919,20. Due to the large number of peptides to be tested, we performed low-resolution ab initio docking, generating five poses for each peptide, followed by an energy minimization step. To accelerate computation, docking jobs were run in parallel on 50 CPUs (50 peptides processed concurrently, one peptide per CPU core). Peptides that did not achieve at least 30% RO were removed from further consideration. The remaining peptides were ranked by their best docking score across the five poses, and the top candidates were selected for downstream analysis.
Parameters definition
To define appropriate genetic algorithm parameters, a preliminary tuning run was performed using the top 25 peptides from the initial population. For each tested parameter combination, the GA was executed for 20 generations using a reduced-fidelity setup, where only one docking pose per peptide was computed using Rosetta. Although this approach does not adhere to best practices for docking-based scoring, it significantly reduces computational cost and enables the rapid comparison of parameter effectiveness. The goal of this experiment was not to estimate binding affinity, but to evaluate the relative impact of key parameters, including mutation rate, tournament size, crossover rate, and elitism, on optimization performance. The best-performing parameter set from this coarse search was: tournament size = 2, mutation rate = 10%, crossing over rate = 90%, elite size = 1, elite lifespan = 3. These parameters were then adopted for the full-scale pipeline, where higher-resolution scoring was applied.
Operations
The population was subjected to two types of genetic operations: crossover and mutation (Fig. 3). In the crossover operation, peptide segments were exchanged between two parent sequences, generating new peptide variants. In the mutation operation, diversity was introduced through random substitution, deletion, or insertion of amino acid residues. The type of operation applied at each step was chosen probabilistically, with 90% chance of performing crossover and a 10% chance of performing mutation.
Examples of mutations of the sequence KWGTSHVF: insertion, deletion, and substitution. Additionally, a crossing over between this sequence and QYADREMP is shown on the right.
Structural modeling
In the next phase, two modeling strategies were employed to evaluate peptide–Mpro binding. In the first approach, each sequence was modeled as a random conformation peptide and docked to the Mpro using Rosetta’s FlexPepDock19,20 high-resolution ab initio protocol, generating five poses per peptide. In the second approach, ColabFold28, an AlphaFold2-based tool24 (in multimer mode) was used to directly predict the structure of the peptide–Mpro complex. To ensure consistency, we used the parameter num_models = 5 and selected the top-ranked model according to AlphaFold2’s internal confidence metrics (pLDDT and TM-score) for relaxation and subsequent analysis (num_relax = 1).
Both experiments were executed on the same server — an AMD Ryzen Threadripper PRO 5995WX 64-Cores processor equipped with an 80GB NVIDIA A100 GPU. For each evaluation, the total time per generation and the time per peptide were recorded. In the AlphaFold2 run, multiple sequence alignments were generated using ColabFold28, and structure prediction was performed locally. Due to resource constraints, a 90-minute break was added between generations to avoid GPU saturation. In the Rosetta run, 25 CPU cores were used to perform docking in parallel, allowing batches of 25 peptides to be processed concurrently.
Fitness function
The top-performing peptides were identified using a tournament-based selection strategy. Complexes generated from both approaches were evaluated with Rosetta’s energy function, which estimates binding free energy at the peptide–protein interface. The PyRosetta library29 was used to assess binding energy and automate the score calculations. These scores guided the GA. In each variation, the best-performing peptide in each generation was retained via elitism. At the same time, the remaining population underwent tournament selection, mutation, and crossover to create the next generation of sequences. The best peptide can be maintained for up to three generations. This process was repeated for 100 generations, allowing the algorithm to converge on peptides with progressively improved predicted binding characteristics.
Molecular dynamics simulations
To evaluate the best results proposed by the case study and compare to other studies, we performed molecular dynamics (MD) simulation experiments. Five systems were built: (i) Mpro-AGVAKAKAV - obtained in this study; (ii) Mpro-VAKAKAV - obtained in this study; (iii) Mpro-WWTWTPFHLLV - obtained in30; (iv) Mpro-LTINWQKYFNT - obtained in31; and (v) Mpro-N3 - obtained in32. The simulations were performed using GROMACS33 with the CHARMM36 force field34,35 and a standard explicit water model on a workstation equipped with CUDA-enabled GPU acceleration (Nvidia A100 80GB). Protein–ligand complexes were placed in a cubic box, centered, solvated, and neutralized with counterions. Energy minimization was performed using the steepest-descent algorithm (50,000 steps). Equilibration proceeded in two stages: NVT for 100 ps at 300 K with a V-rescale thermostat, followed by NPT for 100 ps at 300 K and 1 bar with a V-rescale thermostat and a Parrinello–Rahman barostat, with protein atoms restrained. The production run lasted 100 ns (50,000,000 steps; 2 fs timestep) at 300 K and 1 bar, using the Verlet cutoff scheme and hydrogen-bond constraints.
Trajectories were centered and then least-squares-fitted on protein backbone atoms prior to analysis. RMSD (Root Mean Square Deviation) and RMSF (Root Mean Square Fluctuation) were computed with GROMACS Tools and the MDAnalysis Python library36,37; plots were generated with Matplotlib38.
Binding free energies (ΔG_bind) were estimated by MM-PBSA using gmx_MMPBSA39. Polar solvation energies were obtained by solving the Poisson–Boltzmann (PB) equation, and non-polar contributions were estimated from solvent-accessible surface area (SASA). Energies for complex, receptor, and ligand were evaluated over uniformly spaced frames extracted from the production trajectory, and ΔG_bind was computed as ΔG_complex − ΔG_receptor − ΔG_ligand, with ensemble averaging.
A per-residue free energy decomposition was also performed to identify the residues that contribute most to the binding affinity. The decomposition analysis followed the parameters defined in the input file, where the PB model was set to igb = 0 with a salt concentration of 0.150 M. The decomposition block was activated with idecomp = 1 and dec_verbose = 1, enabling the breakdown of total binding energy into individual residue contributions, including van der Waals, electrostatic, and solvation terms at the protein–peptide interface40.
Structures analysis
The best complexes were analyzed using PyMOL 3.0 (Schrödinger, LLC), ChimeraX 1.1041, and VMD tools42. Protein-peptide contacts were calculated using the COCαDA tool43,44. Additionally, we performed analysis to evaluate the physicochemical properties, toxicity, hemolytic potential, and anti-angiogenic activity of the proposed peptides, as well as the reference peptides, using ToxinPred45, HemoPI 2.046, and AntiAngioPep47.
Results and discussion
Peptides proposed to Mpro
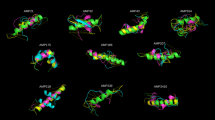
In the performed experiment, 50 peptides were proposed for each generation of the genetic algorithm. The best peptide is defined based on two metrics: docking energy score of Rosetta (the lower, the better) and occupancy (the higher, the better). Rosetta energies are given on a scale named Rosetta Energy Unit (REU), which is derived from a combination of physics-based and statistical potentials. Figure 4 shows the complexes formed by the binding of Mpro to the peptide that obtained the lowest docking score, and the best peptides from eight generations are shown: G1, G10, G25, G50, G75, G84, G96, and G100. Generations G84 and G96 were chosen because they presented the lowest overall values for the experiments with ColabFold and Rosetta, respectively. The other generations were chosen to illustrate how the algorithm evolved peptides.
Best protein-peptide complexes for generations: G1, G10, G25, G50, G75, G84, G96, and G100. The star indicates the generations that showed the best global complexes in the Rosetta (G96) and ColabFold (G84) experiments. Scores are given as REU (Rosetta Energy Unit) values.
Table 1 presents the top five results for the experiments using ColabFold and Rosetta.
For the experiment using ColabFold, the lowest score value was obtained for the peptide AGVAKAKAV obtained in generation 84: -634 REU. In the same generation, the average docking score was − 525 REU, with the worst result being − 96 REU. The occupancy rate was 79% (Table 1). In our experiment, the complete modeling and evaluation of the 50 peptides in the generation 84 required 10,104 s (~ 3 h). For the experiment with Rosetta, the lowest score value was obtained for the peptide PGGHSCC in generation 96 with a docking score of -611 REU and an occupancy of 70%. The modeling and evaluation of the 50 peptides in this generation required 1,672 s (~ 30 min). In the same generation, the average docking score value was − 468 REU, with the worst result being 448 REU. In all evaluated metrics for the top results, the Rosetta results were inferior to those of ColabFold (except for execution time). In this regard, it is important to highlight that the recorded times for Rosetta refer to the generation of five poses per peptide, a number far below the recommended number of poses for a comprehensive conformational search. However, we decided to use this value since ColabFold returns five models per run by default.
When analyzing the structure of the protein-peptide complex (Mpro-AGVAKAKAV), we can see that it makes a series of contacts between the different chains, calculated using the COCαDA tool43,44 (Fig. 5; Table 2). For example, we can cite the predicted hydrogen bonds between T26 (threonine 26 of the protein) and V9 (valine 9 of the peptide), or a salt bridge between E166 (glutamate 166 of the protein) and K7 (lysine 7 of the peptide).
Peptide AGVAKAKAV (sticks with surface displayed in green) complexed with Mpro (purple cartoon). Contacts are displayed with dashed blue lines. Figure generated using ChimeraX41.
Additionally, when considering all 100 generations, we observe that the protein-peptide binding energy of the models generated by ColabFold is generally lower (Fig. 6). It is essential to note that the lower the binding energy, the stronger the binding force.
Lowest value of docking energy score for each generation of Rosetta (blue line) and ColabFold (green line) experiments.
However, we can see that, in the first generations, the models proposed by Rosetta had a lower score (for example, generation 1, 4, 5, 6, 8, 11). This suggests that after a certain number of generations, the strategy using ColabFold consistently outperformed Rosetta, as it appeared to model the proposed peptide within the binding site in a more consistent and complementary pose (Fig. 7).
Difference in docking scores between Rosetta and ColabFold for the best results of each of the 100 generations. Red bars indicate that Rosetta’s approach obtained better performance. Blue bars indicate ColabFold approach obtained a better performance.
Regarding the occupancy metric, the best results of each generation varied between 60% and 80%. The occupancy defines which original residues of the Mpro binding site are likely to interact with a residue of the peptide. In the case of the Rosetta experiment, the occupancy values varied throughout the experiment. In the case of the ColabFold experiments, the values increased over the generations. The maximum occupancy observed was 79% (Supplementary Figure S1).
We also observed that the average size of the peptides with the best docking score decreased over the generations, as shown in the data from the experiment using Rosetta (Fig. 8A). For the experiment using ColabFold, the average peptide size was less than 10 amino acids in almost all generations. For over 40 generations, the best peptide had only five amino acids (Fig. 8B).
(A) Length distribution of the best peptide of each generation. (B) Peptide size frequency.
The evolution toward shorter peptides can be primarily explained by binding entropy. Shorter sequences exhibit reduced conformational flexibility, allowing more stable and compact interactions within the Mpro pocket. This trend was later confirmed by our molecular dynamics (discussed following) and contact analyses, in which a shorter variant (VAKAKAV) showed greater rigidity and improved binding energy compared to AGVAKAKAV.
Using manual curation to detect a better ligand
The results of the case study suggest that the AGVAKAKAV peptide is a potential binder for the Mpro protein. However, analysis of the peptide’s structure indicated that its size could be reduced by removing N-terminal residues, specifically alanine at position 1 and glycine at position 2. The literature has shown that shorter peptides tend to bind better to the receptor48,49. Wier & Beekman (2025) suggest that shortening the peptide sequence (truncation), allowing it to contain only residues essential for the interaction, can improve the efficiency of the ligand and may also simplify the synthesis process48. Furthermore, alanine and glycine are small, neutral amino acids that generally contribute little to specific interactions with the protein’s active site and may be dispensable for maintaining binding affinity and stability. Therefore, we hypothesized that the VAKAKAV peptide would bind more effectively to Mpro. To assess this, we performed molecular dynamics experiments to verify the binding of the Mpro-N3 complex (the original structure), the Mpro-AGVAKAKAV complex, and the Mpro-VAKAKAV complex. The molecular dynamics results are discussed in the next section.
Molecular dynamics simulations
To evaluate the structural stability and binding affinity of the complexes formed between SARS-CoV-2 Mpro and the proposed peptides, we performed 100-ns molecular dynamics simulations followed by RMSD, RMSF, hydrogen-bonding interactions, and free-energy calculations using the MM-PBSA method. These parameters enabled us to directly compare the dynamic behavior of the two defined systems (Mpro-AGVAKAKAV and Mpro-VAKAKAV) and relate it to the structural data previously described for the N3 (02JAVLPJE010) inhibitor in wild-type Mpro32 and two other peptides obtained from the literature (discussed in the next section).
RMSD analysis revealed distinct behaviors between the systems containing different types of ligands (Supplementary Fig. S2). The Mpro-N3 system, in which Mpro is associated with a ligand (non-peptide chemical structure), exhibited sharper oscillations and abrupt fluctuations throughout the simulation, suggesting a less stable interaction with the catalytic site. In contrast, the complexes formed with the proposed peptides (Mpro-AGVAKAKAV and Mpro-VAKAKAV) achieved more consistent values after the initial phase, indicating structural stabilization of the protein–ligand complex. Among them, Mpro-VAKAKAV exhibited a lower and more uniform plateau compared to MproAGVAKAKAV, suggesting greater cohesion and better accommodation of the peptide in the active site.
RMSF analysis along the Mpro residues showed apparent differences between the systems (Supplementary Fig. S3). The Mpro-N3 system, associated with the crystallographic ligand, consistently exhibited higher RMSF values across virtually the entire length of the protein, indicating greater residual mobility and less conformational restriction. In contrast, the complexes formed with the peptides (Mpro-AGVAKAKAV and Mpro-VAKAKAV) presented very similar profiles, with reduced RMSF values across most residues, suggesting that the binding to the peptide promoted local stabilization of the protein. Peaks with greater fluctuation were observed in specific regions, possibly corresponding to flexible loops, but without compromising overall stability. These results indicate that the peptides contribute to stiffening of the active site and adjacent regions, reinforcing the stabilizing role already suggested by the RMSD analyses.
Hydrogen bond interactions, as observed over the simulation, revealed differences between the systems (Supplementary Figure S4). Mpro-VAKAKAV shows a more persistent and continuously distributed hydrogen bond throughout the simulation, ensuring greater stability to the complex. Consistent interactions were observed with catalytic residues, including D153, C156, and D155, which remained throughout most of the simulation. Mpro-AGVAKAKAV also established important bonds, such as those involving E166, G143, and H164, but in a slightly less continuous manner compared to the other complex. The Mpro-N3 system presented a more diffuse pattern, with less regular interactions, mostly displaced to regions peripheral to the catalytic site, such as V157 ↔ K100 and S158 ↔ N151, which do not fully correspond to the hotspots described as critical for Mpro inhibition. These results reinforce that the proposed peptides favor greater stabilization of the active site compared to the N3 wild-ligand, with Mpro-VAKAKAV promoting the most consistent hydrogen bond interactions throughout the simulation.
Hydrogen bond pairs throughout the simulation reinforce the differences between the systems (Fig. 9). Mpro-AGVAKAKAV complex also exhibited persistent bonds, E166 ↔ K5 and K7 ↔ E166, in addition to interactions with G143 and H164, all of which are important residues of the S1 hotspot, part of the substrate-recognition pocket of the Mpro50. Peptides making interactions with such residues are thus highly likely to inhibit the pocket’s activity32. Mpro-VAKAKAV complex demonstrated consistency, with several residue pairs maintaining high occupancy throughout virtually the entire 100-ns runtime. The interactions with D153, D55, and C156 are notable, remaining active throughout, and demonstrate a continuous and well-distributed contact network within the catalytic site. This behavior suggests that Mpro-VAKAKAV complex promoted a good stabilization of Mpro. The Mpro-N3 system presented a more irregular pattern, with diffusely distributed contacts and less temporal continuity. The main pairs (such as V157 ↔ K100 and S158 ↔ N151) exhibited intermediate occupancy, but without the identical important active site residues observed in the peptides. Thus, the heatmaps corroborate that the proposed peptides interact more stably with the Mpro (Fig. 9).
Heatmaps of the ten most frequent hydrogen bonding interactions over 100 ns of simulation. Color intensity reflects bond occupancy at each time interval (dark blue = 100% presence; light yellow = absence).
The cumulative frequency analysis of hydrogen bonds over the 100 ns simulations revealed notable differences among the systems evaluated (Supplementary Figure S5). Mpro-AGVAKAKAV complex also exhibited relevant interactions, such as K7 ↔ E166, E166 ↔ K5, and G143 ↔ K7, which correspond to key residues of Mpro. Mpro-VAKAKAV complex presented the highest recurrence values, with emphasis on the interactions K7 ↔ E166, K100 ↔ D155, and D153 ↔ C156, which remained at high frequency throughout practically the entire trajectory. This pattern suggests that the peptide fosters a highly stable contact network, thereby contributing to firm anchoring at the catalytic site. On the other hand, the Mpro-N3 system presented lower frequencies distributed in pairs less central to the catalytic site, notably V157 ↔ K100 and S158 ↔ N151. Although these interactions were consistent, they do not fully reproduce the classic hotspots of protease inhibition, suggesting a less optimized docking pattern. Thus, the frequency data reinforce that the proposed peptides, especially Mpro-VAKAKAV, establish a more robust and persistent hydrogen bond network compared to the reference ligand.
The stability observed in the RMSD plots (Supplementary Fig. S2) indicated that both peptides conferred greater structural cohesion to Mpro compared to the crystallographic ligand N3, with Mpro-VAKAKAV standing out by reaching more stable plateaus. The RMSF results (Supplementary Fig. S3) showed that the peptide complexes significantly reduced residual flexibility, especially in regions close to the catalytic site, suggesting conformational constraint induced by peptide binding. Hydrogen bond analyses complemented these findings: Mpro-VAKAKAV established a more persistent and distributed network of interactions throughout the simulation, including stable contacts with critical residues such as D153, D155, and C156. In contrast, Mpro-AGVAKAKAV complex maintained relevant bonds with E166, G143, and H164, albeit less consistently. The original system, in turn, exhibited less regular interactions, with most displaced to regions peripheral to the active site, which was reflected in the less favorable free energy value.
Thus, by correlating all analyses, it is clear that the proposed peptides promote greater structural stability, reduce protease flexibility, and establish more robust interaction networks with catalytic residues. Among them, Mpro-VAKAKAV complex stands out as the most promising candidate, as it combines the lowest stable RMSD, reduced local fluctuations, greater hydrogen bond persistence, and the most favorable binding free energy. These results suggest that Mpro-VAKAKAV has greater potential for Mpro inhibition compared to Mpro-AGVAKAKAV and the reference complex.
Comparison to other studies
EvoPepFold is not a standalone modeling or docking algorithm, but rather a hybrid optimization pipeline that integrates established structural modeling tools (e.g., ColabFold and Rosetta) within a genetic algorithm (GA) framework. The closest approach to EvoPepFold is POTTER (peptide optimization tool to enhance receptor binding)51. POTTER implements a genetic algorithm-based approach with mutations allowed in the BLOSUM substitution matrix to design peptides with higher affinity. In its study51, populations are constructed using docking with the Rosetta tool, which is also responsible for assessing fitness. However, it was not used to propose mutations for Mpro. Therefore, it is not possible to directly compare their results with those of this study.
EvoPepFold presents a superior strategy to POTTER by implementing two distinct tools for constructing new populations: Rosetta and ColabFold. Our results indicate that the ColabFold-based approach proposed peptides with higher affinity. This difference can be attributed to the fundamental divergence between the two approaches. ColabFold, based on AlphaFold2, integrates deep learning with evolutionary and coevolutionary information derived from multiple sequence alignments (MSAs). This allows it to infer accurate residue-residue contacts and to predict stable peptide-protein conformations even when conformational sampling is limited to a few structures. Rosetta, in contrast, relies on stochastic conformational sampling guided by a combination of physics-based and statistical potentials, and, thus, its performance depends heavily on the appropriateness of the initial docking configuration and the number of sampled poses.
In this study, each peptide was positioned in the N3 inhibitor binding site to guide Rosetta’s local search. While proper convergence usually requires hundreds of poses, ColabFold’s superior performance at later generations reflects the advantage of its embedded evolutionary constraints and structural priors, which drive faster convergence toward low-energy states.
Comparing with peptides proposed in other studies with Mpro
A better way to evaluate the results proposed in this study would be to compare the peptides proposed here with peptides proposed in other studies that used Mpro as a target. Therefore, we selected two studies from the literature that sought to design peptides that bind to Mpro.
The first study was by Chan et al.31. This study investigates how the SARS-CoV-2 main protease recognizes its natural substrate peptides and, based on this knowledge, proposes new inhibitory peptides to act as competitive ligands for the enzyme. Among their proposed peptides, we selected the p15 peptide, whose sequence is LTINWQKYFNT.
Rossetto and Zhou30 used the GANDALF methodology52 to propose the peptide WWTWTPFHLLV as an Mpro inhibitor. Thus, we selected this peptide as a second comparative case study. Hence, through molecular dynamics experiments, we compared the two main peptides proposed in this work (AGVAKAKAV and VAKAKAV) with the peptide proposed by Chan et al.31, the peptide proposed by Rossetto and Zhou30, and the original peptide N332.
Free energy predicted using the MM-PBSA method revealed differences between the systems (Table 3). Mpro-VAKAKAV complex presented the most favorable value (− 23.97 kcal/mol), followed by Mpro-AGVAKAKAV complex (− 20.71 kcal/mol). Mpro-LTINWQKYFNT showed a binding energy of – 19.85 kcal/mol and Mpro-WWTWTPFHLLV of – 16.97 kcal/mol, while the original system (complex Mpro-N3) exhibited considerably lower affinity (− 8.88 kcal/mol). This corroborates the results of Chan et al.31 and Rossetto and Zhou30, indicating that they indeed proposed peptides with higher affinity than the original peptide. Furthermore, this result demonstrates how the peptides proposed by this methodology outperform results obtained in the literature.
Finally, we performed a residue-by-residue energy decomposition analysis for the four systems (Fig. 10). This analysis highlighted the importance of residue K7 interacting with residue E166 of the Mpro structure. This interaction occurs in both AGVAKAKAV (Fig. 10A) and VAKAKAV (Fig. 10B). For WWTWTPFHLLV, the strongest interaction is carried out by residue W4 (Fig. 10C), which makes polar interactions with R188 and Q189. For LTINWQKYFNT (Fig. 10D), the W5 residue occupies a position equivalent to W4 in the Mpro-WWTWTPFHLLV complex, also being the residue that contributes most to the interaction energy of the complex.
Per-residue free energy decomposition analysis for the four peptide–Mpro complexes. (A) AGVAKAKAV, (B) VAKAKAV, (C) WWTWTPFHLLV, and (D) LTINWQKYFNT. Bar plots represent the contribution of each peptide residue (ΔG_bind, kcal·mol⁻¹) to the total binding free energy obtained by MM-PBSA decomposition. Negative values indicate stabilizing contributions, whereas positive values indicate destabilizing effects. On the right side, we can see the last frame of each molecular dynamics simulation (100ns). The corresponding structural panels show each peptide (sticks) bound to Mpro (gray ribbon), with residues contributing most to binding highlighted. The recurrent electrostatic interaction between K7 and E166 is observed in panels (A) and (B), while aromatic and polar contacts involving tryptophan residues, W4 and W5, and Mpro residues R188 and Q189 characterize the interactions in panels (C) and (D), respectively. Since N3 has non-canonical amino acids, its complex cannot be displayed in this analysis.
These results reveal an interaction pattern among the peptides, centered on key catalytic-site residues E166, R188, and Q189, which are known from previous crystallographic and computational studies to participate in substrate recognition and stabilization of inhibitory ligands. The recurrence of the K7–E166 contact in both AGVAKAKAV and VAKAKAV suggests an electrostatic anchoring mechanism analogous to that observed in experimental Mpro complexes. Taken together, the decomposition profiles are consistent with the MM-PBSA results, indicating that the peptides proposed by EvoPepFold tend to stabilize catalytic residues and display more favorable local interaction energies compared with previously reported peptide inhibitors. Within the computational framework, these findings support the notion that the EvoPepFold pipeline and its GA-based approach overcomes latent space and can help to find sequence motifs capable of reproducing key electrostatic and aromatic interactions necessary for Mpro recognition. We hypothesize that this approach can be successfully used to identify peptides that bind to other sites.
Lastly, we evaluate the physicochemical properties, toxicity, hemolytic potential, and anti-angiogenic activity of all peptides analyzed here using ToxinPred45, HemoPI 2.046, and AntiAngioPep47 software. The results indicated that the designed peptides are predominantly non-toxic, non-hemolytic, and anti-angiogenic, exhibiting short amphipathic sequences with favorable solubility and stability profiles (details are provided in Table S1). Their physicochemical characteristics suggest a good membrane interaction potential, while future optimization may enhance proteolytic resistance and bioavailability through modifications such as cyclization or D-amino acid substitution. Thus, we conclude that the peptides proposed here are potential targets for in vitro experiments.
Limitations
Despite presenting promising results, this study has some limitations that must be acknowledged. First, the procedure for defining new populations within the genetic algorithm is computationally demanding, particularly when modeling complete protein-peptide complexes using ColabFold. In our experiments, running the genetic algorithm for 100 generations required more than 14 days, even on a server equipped with an Nvidia A100 80GB GPU, 786 TB RAM, and an AMD Ryzen Threadripper PRO 5995WX 64-core CPU. On the other hand, the docking-based experiments using the same system were completed in just over one day.
Second, the experiments indicated that the AGVAKAKAV peptide exhibited favorable binding to Mpro. However, manual inspection suggested that the two N-terminal residues contributed minimally to protein binding. Based on this observation, we proposed a shorter variant (VAKAKAV), which was subsequently supported by molecular dynamics simulations. This outcome highlights that, while the proposed method can generate promising candidate ligands, expert knowledge remains crucial for refining peptide design. Moreover, the algorithm could be further improved by incorporating the size, physicochemical properties, and chemical nature of amino acids, which could enhance the prioritization of residues contributing most to the interaction energy. Additionally, strategies such as systematically truncating nonessential residues at the peptide termini could be explored to optimize binding affinity and reduce peptide flexibility, ultimately leading to more potent and specific binders.
Finally, it should be noted that the N3 inhibitor contains non-canonical amino acids that are not supported by the Rosetta energy function employed in this work. As a result, N3 could not be properly re-evaluated under the same computational conditions as the designed peptides. This feature may have influenced the molecular dynamics simulations and binding energy calculations as well, potentially introducing bias into the evaluation process. In this study, the N3 peptide was used as a reference for defining the binding site, orienting the initial positioning of the peptides, and guiding the contact analyses. This approach ensured that all modeled peptides targeted the same functional pocket and adopted comparable binding orientations, while avoiding direct energetic comparisons with N3. Additionally, we must emphasize that the results presented here were evaluated through computational studies and corroborated by results in the literature. Therefore, future in vitro experiments should be conducted to confirm the results obtained here.
Conclusion
In this study, we employed a genetic algorithm (GA) approach, integrated with docking-based scoring and AI 3D structure modeling, to design peptide binders targeting the main protease (Mpro). Our results indicate that the approach, called EvoPepFold, was successful in finding peptides with a higher interaction energy between the protein-peptide complex. The optimization process identified promising candidates, including AGVAKAKAV and VAKAKAV, which exhibited improved predicted binding affinities compared to the initial peptide sequence (N3). These results suggest that GA-driven strategies can effectively explore sequence space to identify peptides with favorable interaction patterns at the protein interface. However, some limitations must be acknowledged. The current approach relies exclusively on docking scores (Rosetta energy) as a proxy for binding affinity, which may not fully capture the dynamics and solvation effects present in real biological environments. Future work will address experimental assays to validate the computational predictions and explore alternative score functions to evaluate complexes. We are confident that the insights gained in this study will help drive innovation in peptide engineering, opening new paths for their rational design and therapeutic use.
Data availability
Data and scripts are available at https://github.com/LBS-UFMG/evopepfold.
References
Frappier, V., Duran, M. & Keating, A. E. PixelDB: protein–peptide complexes annotated with structural conservation of the peptide binding mode. Protein Sci. 27, 276–285 (2018).
Angelova, A., Drechsler, M., Garamus, V. M. & Angelov, B. Pep-lipid cubosomes and vesicles compartmentalized by micelles from self-assembly of multiple neuroprotective building blocks including a large peptide hormone PACAP-DHA. ChemNanoMat 5, 1381–1389 (2019).
Nissan, N., Allen, M. C., Sabatino, D. & Biggar, K. K. Future perspective: Harnessing the power of artificial intelligence in the generation of new peptide drugs. Biomolecules 14, 1303 (2024).
Lee, A. C. L., Harris, J. L., Khanna, K. K. & Hong J.-H. A comprehensive review on current advances in peptide drug development and design. Int. J. Mol. Sci. 20, 2383 (2019).
Lau, J. L. & Dunn, M. K. Therapeutic peptides: historical perspectives, current development trends, and future directions. Bioorg. Med. Chem. 26, 2700–2707 (2018).
Qi, Y. K., Zheng, J. S. & Liu, L. Mirror-image protein and peptide drug discovery through mirror-image phage display. Chem 10, 2390–2407 (2024).
Yin, H. et al. Design, synthesis and anticancer evaluation of novel oncolytic peptide-chlorambucil conjugates. Bioorg. Chem. 138, 106674 (2023).
Yin, H. et al. The hybrid oncolytic peptide NTP-385 potently inhibits adherent cancer cells by targeting the nucleus. Acta Pharmacol. Sin. 44, 201–210 (2023).
Fu, X. Y. et al. Three rounds of stability-guided optimization and systematical evaluation of oncolytic peptide LTX-315. J. Med. Chem. 67, 3885–3908 (2024).
Hashemi, Z. S. et al. In Silico approaches for the design and optimization of interfering peptides against protein–protein interactions. Front Mol. Biosci 8, 669431 (2021).
Diakou, I. et al. Novel computational pipelines in antiviral structure–based drug design (Review). Biomed. Rep. 17, 1–5 (2022).
Ibrahim, M. et al. Why is the Omicron main protease of SARS-CoV-2 less stable than its wild-type counterpart? A crystallographic, biophysical, and theoretical study. hLife 2, 419–433 (2024).
Sacco, M. D. et al. Structure and Inhibition of the SARS-CoV-2 main protease reveal strategy for developing dual inhibitors against Mpro and cathepsin L. Sci. Adv. 6, eabe0751 (2020).
dos Santos, L., Mariano, D. & Minardi, R. ViPeC: Vision Transformer-Based Approach for Peptide-Protein Interface Classification. 8 (2024). https://doi.org/10.1109/CIBCB58642.2024.10702141
dos Santos, L., Mariano, D., Bastos, L., Cioletti, A. & Minardi, R. Peptide-protein interface classification using convolutional neural networks. in 112–122 (2023). https://doi.org/10.1007/978-3-031-42715-2_11
Yan, Y., Zhang, D., Zhou, P., Li, B. & Huang, S. Y. HDOCK: a web server for protein–protein and protein–DNA/RNA docking based on a hybrid strategy. Nucleic Acids Res. 45, W365–W373 (2017).
Yan, Y., Tao, H., He, J. & Huang, S. Y. The HDOCK server for integrated protein–protein docking. Nat. Protoc. 15, 1829–1852 (2020).
Zhou, P., Jin, B., Li, H. & Huang, S. Y. HPEPDOCK: a web server for blind peptide–protein docking based on a hierarchical algorithm. Nucleic Acids Res. 46, W443–W450 (2018).
Raveh, B., London, N., Zimmerman, L. & Schueler-Furman, O. Rosetta FlexPepDock ab-initio: simultaneous folding, docking and refinement of peptides onto their receptors. PloS One. 6, e18934 (2011).
Chaudhury, S. et al. Benchmarking and analysis of protein docking performance in Rosetta v3. 2. PloS One. 6, e22477 (2011).
Evans, R. et al. Protein complex prediction with AlphaFold-Multimer. biorxiv 2021–10 (2021).
Omidi, A., Møller, M. H., Malhis, N., Bui, J. M. & Gsponer, J. AlphaFold-Multimer accurately captures interactions and dynamics of intrinsically disordered protein regions. Proc. Natl. Acad. Sci. 121, e2406407121 (2024).
Fjell, C. D., Jenssen, H., Cheung, W. A., Hancock, R. E. W. & Cherkasov, A. Optimization of antibacterial peptides by genetic algorithms and cheminformatics. Chem. Biol. Drug Des. 77, 48–56 (2011).
Borkakoti, N. & Thornton, J. M. AlphaFold2 protein structure prediction: implications for drug discovery. Curr. Opin. Struct. Biol. 78, 102526 (2023).
Berman, H. M. et al. The protein data bank. Nucleic Acids Res. 28, 235–242 (2000).
Martins, P. M. et al. Propedia: a database for protein–peptide identification based on a hybrid clustering algorithm. BMC Bioinform. 22, 1 (2021).
Martins, P. et al. Propedia v2.3: A novel representation approach for the peptide-protein interaction database using graph-based structural signatures. Front Bioinforma 3, 1103103 (2023).
Mirdita, M. et al. ColabFold: making protein folding accessible to all. Nat. Methods. 19, 679–682 (2022).
Adolf-Bryfogle, J. & Jr, R. L. D. The pyrosetta toolkit: a graphical user interface for the Rosetta software suite. PLOS One. 8, e66856 (2013).
Rossetto, A. & Zhou, W. GANDALF: Peptide Generation for Drug Design using Sequential and Structural Generative Adversarial Networks. in Proceedings of the 11th ACM International Conference on Bioinformatics, Computational Biology and Health Informatics 1–10Association for Computing Machinery, New York, NY, USA, (2020). https://doi.org/10.1145/3388440.3412487
Chan, H. T. H. et al. Discovery of SARS-CoV-2 Mpro peptide inhibitors from modelling substrate and ligand binding. Chem. Sci. 12, 13686–13703 (2021).
Jin, Z. et al. Structure of Mpro from SARS-CoV-2 and discovery of its inhibitors. Nature 582, 289–293 (2020).
Abraham, M. J. et al. High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1. GROMACS, 19–25 (2015).
Huang, J. et al. CHARMM36m: an improved force field for folded and intrinsically disordered proteins. Nat. Methods. 14, 71–73 (2017).
Malde, A. K. et al. An automated force field topology builder (ATB) and repository: version 1.0. J. Chem. Theory Comput. 7, 4026–4037 (2011).
Michaud-Agrawal, N., Denning, E. J., Woolf, T. B. & Beckstein, O. MDAnalysis: a toolkit for the analysis of molecular dynamics simulations. J. Comput. Chem. 32, 2319–2327 (2011).
Gowers, R. J. et al. MDAnalysis: a python package for the rapid analysis of molecular dynamics simulations. (2019).
Hunter, J. D. & Matplotlib A 2D graphics environment. Comput. Sci. Eng. 9, 90–95 (2007).
Valdés-Tresanco, M. S., Valdés-Tresanco, M. E., Valiente, P. A. & Moreno, E. gmx_MMPBSA: a new tool to perform end-state free energy calculations with GROMACS. J. Chem. Theory Comput. 17, 6281–6291 (2021).
Kollman, P. A. et al. Calculating structures and free energies of complex molecules: combining molecular mechanics and continuum models. Acc. Chem. Res. 33, 889–897 (2000).
Meng, E. C. et al. UCSF chimerax: tools for structure building and analysis. Protein Sci. 32, e4792 (2023).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–38 (1996).
Lemos, R. P., Mariano, D., de Silveira, S., de Melo-Minardi, R. C. & A. & COCαDA-A fast and scalable algorithm for interatomic contact detection in proteins using Cα distance matrices. Front. Bioinforma. 5, 1630078 (2025).
Lemos, R., Mariano, D., Silveira, S. & Melo-Minardi, R. COCαDA - Large-Scale Protein Interatomic Contact Cutoff Optimization by Cα Distance Matrices. 70 (2024). https://doi.org/10.5753/bsb.2024.245545
Gupta, S. et al. In Silico approach for predicting toxicity of peptides and proteins. PloS One. 8, e73957 (2013).
Rathore, A. S., Kumar, N., Choudhury, S., Mehta, N. K. & Raghava, G. P. Prediction of hemolytic peptides and their hemolytic concentration. Commun. Biol. 8, 176 (2025).
Ettayapuram Ramaprasad, A. S., Singh, S., Gajendra, P. S., Venkatesan, S. & R. & AntiAngioPred: a server for prediction of anti-angiogenic peptides. PloS One. 10, e0136990 (2015).
van Wier, S. P. & Beekman, M. A. Peptide design to control protein–protein interactions. (2025). https://doi.org/10.1039/D4CS00243A.
Leenheer, D., ten Dijke, P. & Hipolito, C. J. A current perspective on applications of macrocyclic-peptide-based high-affinity ligands. Pept. Sci. 106, 889–900 (2016).
Verma, N., Henderson, J. A. & Shen, J. Proton-coupled conformational activation of SARS coronavirus main proteases and opportunity for designing small-molecule broad-spectrum targeted covalent inhibitors. J. Am. Chem. Soc. 142, 21883–21890 (2020).
Abreu, A. et al. An approach for engineering peptides for competitive Inhibition of the SARS-COV-2 Spike protein. Molecules 29, 1577 (2024).
Rossetto, A. M. & Zhou, W. GANDALF: A Prototype of a GAN-based Peptide Design Method. in Proceedings of the 10th ACM International Conference on Bioinformatics, Computational Biology and Health Informatics 61–66Association for Computing Machinery, New York, NY, USA, (2019). https://doi.org/10.1145/3307339.3342183
Acknowledgements
The authors would like to thank the Brazilian agencies CAPES, CNPQ, and FAPEMIG. This study was financed in part by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior – Brasil (CAPES) – Finance Code 001.
Author information
Authors and Affiliations
Contributions
FC developed the scripts and performed the experiments. FC and DM write the manuscript. SCA performed the molecular dynamics simulations. LB, APA, RPL, SCA, LMS, and RCMM edit the manuscript. All authors read and approved the final version of this manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
41598_2025_28061_MOESM1_ESM.pdf (download PDF )
Supplementary Material 1: Supplementary material is available at https://github.com/LBS-UFMG/evopepfold/blob/main/supplementary_material.pdf.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chaves Carvalho, F., Mariano, D., Bastos, L. et al. A hybrid evolutionary and structural method for AI-guided peptide inhibitor design using AlphaFold and Rosetta. Sci Rep 15, 44519 (2025). https://doi.org/10.1038/s41598-025-28061-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-28061-y