Abstract
Hepatocellular carcinoma (HCC) is one of the most prevalent malignant tumors worldwide, and its occurrence and development are intricately associated with the abnormal regulation of various genes and signaling pathways. Exosome Component 3(EXOSC3) is involved in the occurrence and growth of tumors. However, the exact mechanisms by which EXOSC3 influences HCC remain to be elucidated. Bioinformatics analysis method was used to detect the expression of EXOSC3 in HCC, qRT-PCR and Western blot were used to verify EXOSC3 expression in HCC cell lines.CCK-8 and colony formation assay was used to evaluate the effect of EXOSC3 on tumor proliferation, and validate the impact on cell migration and invasion through Transwell and Wound healing assay. Flow cytometry and Hochest staining were used to investigate the effect of EXOSC3 knockdown on the cell cycle and apoptosis of HCC. Western blot method was used to detect proteins expression. In HCC tissues, EXOSC3 mRNA and protein expression levels were notably higher than those in normal liver tissues and these levels correlated with poor prognosis. After knocking down EXOSC3, cell proliferation, colony formation, and metastasis were significantly inhibited in HCC. Flow cytometry and Western blot analyses showed that knockdown EXOSC3 promoted HCC cell apoptosis and inhibited cell cycle progression with reduced G1/S checkpoint protein expression. Furthermore, knockdown EXOSC3 activated P53 and decreased retinoblastoma protein (RB1) phosphorylation, suggesting that EXOSC3 functions via the P53 pathway in HCC. EXOSC3 has potential as a prognostic biomarker in HCC. Knockdown EXOSC3 leads to cell cycle arrest and consequently inhibits the proliferation of liver cancer cells by activating the P53 signaling pathway and concomitantly suppressing the expression of crucial regulatory proteins. These results not only establish EXOSC3 as a clinically relevant prognostic indicator but also highlight its therapeutic potential as a molecular target for HCC intervention strategies.
Similar content being viewed by others
Introduction
Hepatocellular carcinoma (HCC), ranking as one of the most prevalent cancers globally, exhibits the 6th leading cause of cancer worldwide and the 3rd leading cause of cancer-related death, thereby posing formidable challenges to the worldwide healthcare system1,2. Therapeutic modalities for hepatocellular carcinoma are stratified into curative approaches (including hepatic transplantation, surgical resection, or ablative therapies/selective internal radiation therapy) and palliative interventions (transarterial chemoembolization, systemic therapies). Where clinically feasible, transplantation represents the most definitive therapeutic option, though its implementation remains constrained by donor organ scarcity3. However, the precise molecular mechanisms underlying HCC progression remain poorly understood. Given that accurate prognostic assessment and identification of effective therapeutic targets are crucial for clinical decision-making in HCC management, there is an urgent need to elucidate the fundamental molecular pathways governing HCC initiation and progression.
EXOSC (exosome component) are ribonuclease complexes comprised of 10 evolutionarily conserved subunits, which are well known for degrading RNA4,5,and can protect cells from genomic instability by degrading DNA/RNA hybrids and restricting DNA strand mutational asymmetry6,7. Interestingly, Emerging evidence links multiple EXOSC components to cancer beyond their RNA degradation role. Notably, EXOSC1 promotes DNA damage and sensitizes renal clear cell carcinoma cells to DNA repair inhibitors8, it make also as a serve as a target protein for the anticancer activity of spider peptide toxin LVTX-8 in nasopharyngeal carcinoma9. Upregulated EXOSC2 expression, directly modulated by tRNA-GluUUC, significantly correlates with breast cancer metastasis and mediates WTAP’s oncogenic effects in breast cancer cells through Wnt/β-catenin signaling activation10. Moreover, upregulated EXOSC5 expression can enhance cell proliferation in colorectal cancer through the ERK and AKT signaling pathways11, underlining its significance in gastrointestinal cancers. Similarly, accumulating evidence suggests that other exosome complex components, particularly EXOSC8, EXOSC9, and EXOSC10, play significant roles in the pathogenesis and progression of multiple cancers12,13,14, whereas currently, there is no evidence suggesting an association between EXOSC 6/7 and cancer15.
EXOSC3 is one of the components of RNA exosomes, which responsible for 3′–5′ exoribonuclease activity and are involved in RNA degradation and post-transcriptional modifications. EXOSC3 has S1 and KH RNA‐binding domains16,17,18. Mutations in EXOSC3 have been reported to cause pontocerebellar hypoplasia, spinal motor neuron degeneration, denervation, and complex hereditary spastic paraplegia19,20,21. Masum et al. have shown that forced expression of EXOSC3 and CNOT4 in human colon epithelial cells increased the expression of IFNGR1, MYD88, NFκBIA, and STAT3 in 293 T cells, and activated ERK1/2 and JNK. These results suggest that EXOSC3- and CNOT4-mediated RNA stability, including the stability of MYD88 in inflamed mucosa, could trigger cancer development and serve as potential predictive markers and innovative treatments for controlling cancer development22. However, the role of EXOSC3 in HCC has been scarcely studied.
In this study, our investigations revealed that EXOSC3 was markedly upregulated in HCC tissues and was strongly inversely correlated with the disease prognosis. The results demonstrated that silencing EXOSC3 effectively inhibited the proliferation and growth of HCC cells through the activation of the P53 signaling pathway. These findings collectively suggest that EXOSC3 may serve as both a valuable prognostic indicator and a promising therapeutic target for HCC management.
Methods
Expression analysis of EXOSC3
Utilizing the online portal TIMER23(http://timer.cistrome.org) to analyzed the EXOSC3 expression profiles across 33 cancer types ,and the mRNA and protein expression levels of EXOSC3 in HCC were evaluated in the UALCAN24 database (https://ualcan.path.uab.edu) were sourced from the Cancer Genome Atlas (TCGA) database. The expression levels of EXOSC3 were normalized using Log2 transformation in these datasets.
The human protein atlas (HPA) database
The Human Protein Atlas (HPA) database (http://www.proteinatlas.org) is a remarkable initiative for mapping human proteins. It utilizes antibody—based imaging, data from the brainstem, transcriptomic analysis, mass spectrometry—based proteomics, and other methods to comprehensively map all human proteins. This database incorporates immunohistochemistry (IHC) analysis of numerous protein—coding genes in different cancer patients, employing a variety of techniques. For the purpose of identifying disparities, IHC images of EXOSC3 in both liver cancer and normal liver tissues were retrieved from the HPA database.
GO annotation enrichment and KEGG pathway enrichment analysis
The mRNA expression data and relevant original clinical data of TCGA-LIHC samples as well as their corresponding normal samples were obtained from the XENA website (http://xena.ucsc.edu)25. Analyze the co-expressed genes of EXOSC3 in TCGA-LIHC patients using R software with a correlation coefficient > 0.6 and P < 0.001 as the standard. Gene Function Ontology Annotation (GO) and Kyoto Gene Encyclopedia (KEGG) pathway enrichment analysis of selected co-expressed genes through the “clusterProfiler” package, and p < 0.05 and FDR < 0.1 were considered to be statistically significant.
Kaplan–Meier
KM-plotter (http://kmplot.com/analysis) analyzed datasets of over 30,000 samples from 21 different tumor types. Furthermore, the statistical analysis involved the use of a Cox proportional hazards regression and the calculation of false discovery rates. Two survival analyses were performed to assess the effect of RNA expression levels on patient overall survival (OS) and disease-specific survival (DSS). Furthermore, we obtained p-values from log-rank tests to determine significance in survival.
Cell culture
Liver cancer cell lines (Huh-7, HepG2, MHCC97H and HCCLM3) and normal human liver cell line LO2 were purchased from the typical cultured cell bank of Shanghai Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences, without mycoplasma contamination. Cells were cultured in DMEM supplemented with 10% FBS (fetal bovine serum) and 1% antibiotic / antifungal solution. Cultured in a 37℃, 5% CO2 air cell incubator.
Knockdown of EXOSC3
To knockdown EXOSC3 expression, small interfering RNA (siRNA) and negative control (NC) RNA targeting EXOSC3 were transfected into HCC cells using lipo8000 reagent (Lipo8000™ Transfection Reagent) following the manufacturer’s protocol. The following primers were used: NC (5′–3′, S′-UUCUCCGAACGUGUCACGUTT; AS′-ACGUGACACGUUCGGAGAATT); siEXOSC3-424 (5′–3′, S′-GGAGUGAGCCAGCUUCUUUTT; AS′-AAAGAAGCUGGCUCACUCCTT).
RNA extraction and RT-qPCR
Total RNA from HCC cells was extracted using MonzolTM Reagent Pro reagent (Monad (Suzhou) Biotech Co., Ltd) and reverse transcribed into cDNA (ToloScript RT EasyMix for qPCR # 22,107). The qRT-PCR analysis was performed using 2 Q3 SYBR qPCR Master Mix (Universal, # 22,204) according to the manufacturer’s protocol. The final relative expression level was quantified by the 2ΔΔCt method. Primers were synthesized by Sangon Biotech (Shanghai, China),Primer sequences: GAPDH-F: 5’-GGACCTGACCTGCCGTCTAG-3’, GAPDH-R: 5’-GTAGCCCAGGATGCCCTTGA-3’;
EXOSC3-F: 5’-CGTTGAGCCTGAATGCTAGAG-3’, EXOSC3-R: 5’-GCTGAGAGTCCACCCAGTAAA-3’.
CCK-8 assay
Transfected logarithmic growth phase cells were digested with trypsin, and after cell counting cells were seeded at a density of 2000 cells per well in 96-well plates and five complex wells were cultured in a 37℃ incubator. After cell transfection, 24 h, 48 h, 72 h and 96 h, 100 μL CCK-8 working solution (CCK-8: DMEM medium = 1:10) was added to each well, incubated at 37℃ for 1 h before OD at 450 nm, and the cells inhibition rate were quantified by spectrophotometry. As illustrated:
Wound healing assay
Transfected cells from the logarithmic growth phase were seeded in 6-well plates to achieve 90% confluence after 48H transfection. After suction medium and washing with PBS, the cell monolayer was gently scraped with a 10ul pipette tip, washed again twice with PBS, and detached cells were removed. The treated cells were cultured in DMEM medium without FBS. Samples were taken at 0 and 24H and photographed.
Transwell assay
Cells from the serum-free medium were seeded into the upper chamber of a 24-well transwell. DMEM containing 10% FBS was added to the lower chamber. After 24 h, cells were fixed with 4% paraformaldehyde for 30 min, stained with crystal violet and then cotton swabs to carefully remove non-migrated cells in the upper chamber. Migrating cells on the lower surface were photographed by a microscope in three randomly selected fields and migrated cells were counted.
Flow cytometry analysis
Cell cycle analysis
Cells were fixed in 75% ethanol at-20℃ for at least 24 h. After washing, the cells were incubated in propidium iodide staining solution (PI) for 30 min. Cell distribution was detected and analyzed on a NovoCyte flow cytometer (ACEA Biosciences).
Cell apoptosis analysis
Cells were washed and collected with PBS. Then, the cells were stained with propidium iodide (PI) staining solution and fluorescein isothiocyanate (FITC), incubated at room temperature for 20 min, and the results were determined by flow cytometry.
Clone formation experiments
The indicated transfected HCC cells were seeded on 6-well plates at a density of 1200 per well. Cells were cultured for approximately 12 days, and samples were washed with PBS, fixed using 4% paraformaldehyde, and stained with 0.1% crystal violet. Final photographs were taken and the community number was counted by image J.
Hoechst 33,342 assay
After cells were seeded in 6-well plates and transfection, cells were washed twice with 1 ml of precooled PBS buffer and 1 ml Hoechst33342 of staining solution was successively added to each well. The cells were incubated at room temperature for 30 min, the dye solution was washed with 1 × PBS buffer, observed and photographed using a fluorescence microscope, and the experiment was repeated three times.
Western blot
Western blotting was carried out according to the manufacturer’s instructions. HCC cells were lysed with RIPA lysis buffer on ice for 20 min. Subsequently, the cell lysates were centrifuged at 14,000 g for 10 min at 4℃ to collect the supernatant. Twenty to forty micrograms of protein samples were separated using 10% SDS-PAGE and transferred onto a polyvinylidene fluoride (PVDF) membrane. The membrane was blocked with skim milk at room temperature for 2 h, incubated with Primary Antis overnight at 4 ℃, and then washed 3 times with Tris–HCl solution + Tween-20 (TBST) for 10 min each time. After washing, the membrane was incubated with goat anti-rabbit or goat anti-mouse secondary antibodies for 1 h. Finally, the blots were detected using enhanced chemiluminescence (ECL Plus). The density of protein bands was evaluated by Image J software, and the protein levels were normalized to GAPDH. The dilution ratio of the antibody used is as follows:
EXOSC3(Cat no : 15,062–1-AP;1:3000; Proteintech, Wuhan, China), P53(Cat no:10442-1-AP;1:3000; Proteintech, Wuhan, China), p-P53(Cat no : 28,961-1-AP;1:3000; Proteintech, Wuhan, China), P21(Cat no : 10,355-1-AP; 1:3000; Proteintech, Wuhan, China), P-RB1(Cat no : 30,376-1-AP;1:500; Proteintech, Wuhan, China), CDK4(Cat no : 11,026-1-AP;1:2000; Proteintech, Wuhan, China), pCDK2(AP1364; 1:1500;ABclonal,Wuhan,China), CyclinD1(Cat no : 26,939-1-AP;1:6000; Proteintech, Wuhan, China), CyclinE1(Cat no : 11,554-1-AP;1:1500; Proteintech, Wuhan, China), β-catenin(Cat no : 51,067-2-AP;1:8000; Proteintech, Wuhan, China), C-myc(Cat no : 10,828-1-AP;1:4000; Proteintech, Wuhan, China), Bax (Cat No. 50599-2-Ig, 1:1000; Proteintech, Wuhan, China), Bcl-2 (Cat No.68103-1-Ig, 1:1000; Proteintech, Wuhan, China), Casepase3(Cat no : 19,677-1-AP, 1:1500; Proteintech, Wuhan, China), N-cadherin (Cat No.22018-1-AP, 1:5000; Proteintech, Wuhan, China), E-cadherin (Cat No.20874-1-AP, 1:5000; Proteintech, Wuhan, China), Vimentin(Cat no : 10,366-1-AP,1:20,000; Proteintech, Wuhan, China), GAPDH (Cat No.60004-1-Ig, 1:20,000; Proteintech, Wuhan, China), β-actin(Cat no : 20,536-1-AP,1:20,000; Proteintech, Wuhan, China), β-tubulin (Cat no : 10,094-1-AP,1:6000; Proteintech, Wuhan, China).
Statistical analysis
Data analysis was performed using R 4.4.2, Prism 10.1.2 (GraphPad Software, La Jolla, CA, USA) software and image J software. All data are expressed as the mean ± standard deviation (SD) from 3 experiments. Statistical differences between groups were calculated using Student t test and one-way ANOVA, followed by Dunnett post hoc test. Statistical significance was defined as p < 0.05. Significance levels are denoted as follows: *p < 0.05; **p < 0.01; ***p < 0.001;****p < 0.0001; NS: not significant (p > 0.05).
Results
EXOSC3 expression is elevated in HCC and is closely associated with patient survival
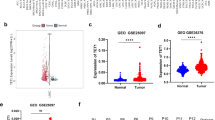
To investigate EXOSC3 expression patterns, we first performed comprehensive bioinformatics analysis of TCGA datasets. Using the TIMER platform, we systematically compared EXOSC3 mRNA expression between tumor tissues and matched normal tissues (n ≥ 3) across multiple cancer types. Significant EXOSC3 upregulation (P < 0.001) was observed in BLCA, CHOL, COAD, HNSC, LIHC, LUAD, READ, and STAD malignancies (Fig. 1A), suggesting its potential oncogenic role. Subsequent UALCAN-based validation in HCC confirmed elevated EXOSC3 expression at both transcriptional and translational levels (Fig. 1B, C). Importantly, EXOSC3 levels exhibited progressive increases with advancing tumor stage and grade (Fig. 1D, E), supporting its clinical relevance in HCC progression.
EXOSC3 Expression in HCC and Its Association with Patient Survival. (A) EXOSC3 expression across various tumor tissues versus normal tissues, analyzed using TCGA database. (B-C) Bioinformatics analysis via the UALCAN portal (http://www.example.com) revealed differential EXOSC3 expression at mRNA (B) and protein (C) levels between normal and primary HCC tissues. (D, E) Correlation of EXOSC3 expression with tumor metastasis stage (D) and malignancy grade (E) in HCC, as assessed using UALCAN. (F) Immunohistochemical techniques and the Human Protein Atlas (HPA) database were utilized to evaluate the local expression pattern of EXOSC3 in HCC tissue samples. (G) Comparison of overall survival and recurrence-free survival between HCC patients with high versus low EXOSC3 expression, using the online Kaplan–Meier plotter database. Statistical Significance levels: *p < 0.05, **p < 0.01, ***p < 0.001, NS: no significant difference (p > 0.05).
To ascertain whether EXOSC3 protein levels are elevated in hepatocellular carcinoma (HCC) compared to normal liver tissue, we analyzed the immunohistochemical staining results of EXOSC3 in 1 normal liver tissue sample and 12 liver cancer tissue samples from the HPA database. In normal liver tissue, no EXOSC3 protein was detected in hepatocytes or cholangiocytes. However, among all ovarian cancer samples, 10 showed varying degrees of cytoplasmic and membrane staining. Specifically, 2 samples exhibited 25–75% high-intensity staining, 6 samples showed 25–75% medium-intensity staining, and 2 samples had 25–75% weak staining, with representative microscopic images depicted in Fig. 1F. Furthermore, data from the online Kaplan–Meier plotter revealed that the overall survival and disease-specific survival of HCC patients with high EXOSC3 expression were markedly lower than those with low EXOSC3 expression. The analysis indicated that high mRNA expression of EXOSC3 correlates with a poor prognosis for HCC patients (Fig. 1G). In summary, these findings suggest that the significant overexpression of EXOSC3 in HCC tissues is an important prognostic maker for predicting the prognosis of HCC patients.
Knockdown of EXOSC3 inhibited the growth of HCC cells
To elucidate EXOSC3’s biological function in HCC, we first evaluated its expression in LO2 hepatocytes and four HCC cell lines (HCCLM3, Huh-7, HepG2, MHCC-97H) using qRT-PCR and Western blot. Both mRNA and protein levels were significantly upregulated in all malignant cell lines versus LO2 controls (p < 0.05, Fig. 2A, B), corroborating previous bioinformatics and IHC findings regarding EXOSC3’s association with HCC progression.
EXOSC3 knockdown suppresses HCC cell growth. (A, B) qRT-PCR (A) and Western blot (B) analysis of EXOSC3 expression in four HCC cell lines (Huh-7, HCCLM3, HepG2, MHCC97H) versus immortalized LO2 hepatocytes. (C) Fluorescence microscopy confirmed transfection efficiency. (D) EXOSC3 knockdown validation in Huh-7 cells (siEXOSC3 vs. siNC controls). (E, F) RT-qPCR and Western blot quantification of EXOSC3 suppression in Huh-7 and HCCLM3 cells post-knockdown. (G) CCK-8 assay demonstrating reduced proliferative capacity following EXOSC3 knockdown. (H) Impaired colony formation in siEXOSC3-treated cells. (I, J) Enhanced apoptosis evidenced by Hoechst 33,342 staining (200 ×) and flow cytometry. (K, L) Differential expression profiles of key proteins between control and EXOSC3-knockdown groups. Statistical significance levels:*p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
High-expressing Huh-7 and HCCLM3 cells were selected for functional studies. Successful transfection was confirmed by fluorescence microscopy (Fig. 2C, D), with subsequent qRT-PCR and Western blot validating significant EXOSC3 knockdown (p < 0.05, Fig. 2E, F). Functional assays revealed EXOSC3’s proliferative role: CCK-8 showed reduced 450 nm absorbance in siEXOSC3 groups (p < 0.05, Fig. 2G), while colony formation assays demonstrated significantly fewer clones (p < 0.05, Fig. 2H), collectively indicating EXOSC3’s critical role in maintaining HCC proliferative capacity. The observed suppression of cell growth may result from either proliferation inhibition, apoptosis induction, or both mechanisms. To investigate this, we systematically evaluated the impact of EXOSC3 knockdown on hepatocellular carcinoma (HCC) cell proliferation and apoptosis. Hoechst 33,342 staining demonstrated markedly enhanced nuclear condensation and brighter blue fluorescence in EXOSC3-depleted cells compared to negative controls (NC) (Fig. 2I), suggesting increased apoptotic activity. This finding was corroborated by flow cytometry analysis, which revealed significantly elevated apoptosis rates in both Huh-7 and HCCLM3 cell lines following EXOSC3 knockdown (Fig. 2J).
Mechanistically, EXOSC3 downregulation appears to suppress tumor growth through apoptotic pathway activation. Western blot analysis of key apoptosis regulators showed consistent alterations in protein expression profiles: decreased anti-apoptotic Bcl2 and Caspase3 levels accompanied by increased pro-apoptotic BAX expression in EXOSC3-knockdown cells (Fig. 2K, L). These molecular changes collectively indicate that EXOSC3 silencing promotes programmed cell death in HCC cells.
Our findings demonstrate that EXOSC3 plays a pivotal role in HCC progression by simultaneously regulating both proliferative and apoptotic pathways.
.
Downregulation of EXOSC3 inhibits the metastasis of HCC cells
To evaluate the role of EXOSC3 in hepatocellular carcinoma (HCC) metastasis, we performed comprehensive functional analyses of migration and invasion capacities in Huh-7 and HCCLM3 cell lines following EXOSC3 knockdown. Wound healing assays revealed that EXOSC3 depletion significantly impaired cell motility, with siEXOSC3-treated cells exhibiting markedly reduced wound closure rates compared to negative controls (NC) (Huh-7: 22.6% vs 55.2%, p < 0.05; HCCLM3: 31.9% vs 17.4%, p < 0.05) (Fig. 3A, B).Consistent with these findings, Transwell migration assays demonstrated a substantial decrease in membrane-traversing cells upon EXOSC3 knockdown (p < 0.05) (Fig. 3C, D). Parallel invasion assays using Matrigel-coated chambers yielded similar results, confirming EXOSC3’s regulatory role in both migration and invasion processes (Fig. 3E, F).
EXOSC3 downregulation suppresses HCC cell metastasis. (A, B) Wound healing assays demonstrated impaired lateral migration capacity in EXOSC3-knockdown HCC cells (100 ×). (C, F) Transwell migration and invasion assays revealed significantly reduced longitudinal migration potential following EXOSC3 depletion. (G, H) Western blot analysis of epithelial-mesenchymal transition (EMT) marker expression in HCC cells after EXOSC3 knockdown. Statistical significance levels: *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001,NS: no significant difference (p > 0.05).
Given the established link between epithelial-mesenchymal transition (EMT) and tumor metastasis, we investigated EMT marker expression profiles. Western blot analysis showed significant downregulation of mesenchymal markers (Vimentin and N-cadherin) in EXOSC3-depleted cells (Fig. 3G, H), indicating EMT pathway inhibition. Collectively, our data demonstrate that EXOSC3 promotes HCC cell motility and invasiveness through EMT activation, suggesting its potential as a therapeutic target for metastatic HCC intervention.
EXOSC3 inhibits HCC proliferation and is related to DNA replication and the cell cycle
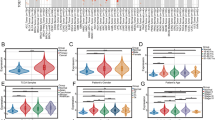
Building upon our previous investigation of EXOSC3-associated biological processes in HCC, we systematically explored the molecular mechanisms underlying EXOSC3-mediated regulation of HCC proliferation. Utilizing the LinkedIn Omics platform’s analytical modules, we performed comprehensive pathway enrichment analyses to identify EXOSC3-related functional networks. The LinkFinder module revealed EXOSC3 co-expression patterns in TCGA-LIHC data through Pearson correlation analysis (Fig. 4A), with heatmaps visualizing the top 50 positively and negatively correlated genes (Fig. 4B, C). GSEA via LinkInterpreter identified significant enrichment of DNA replication-related genes in a key GO module (Fig. 4D, E). KEGG analysis further demonstrated cell cycle pathway enrichment (Fig. 4F, G)26,27, particularly involving cyclin family members (A1, D2, D3, E1, E2). Notably, Wnt/β-catenin pathway components (β-catenin, C-myc) exhibited elevated expression in EXOSC3-high versus EXOSC3-low or normal tissues (Fig. 4H), consistent with cyclin D1 regulation. These findings substantiate the association between reduced EXOSC3 expression and impaired HCC proliferative capacity.
EXOSC3 inhibits HCC proliferation and is related to DNA replication and the cell cycle. (A) EXOSC3 association differential gene results (volcano plot). (B) Positive correlation of significant genes (heatmap). (C) Negatively correlated important genes (heat map). (D) Biological processes in the Gene Ontology (GO). (E) The KEGG pathway. (F, G) GO and KEGG pathways indicate that EXOSC3 is involved in a variety of signals, mainly cycle-and replication-related. (H)Relative mRNA expression levels of cell cycle-related proteins in the TCGA database of normal (n = 50) and HCC tissues (low EXOSC3, n = 185; high EXOSC3, n = 189). Statistical significance levels: *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
Knocking down EXOSC3 inhibits HCC cell proliferation by regulating the G1/S phase of the cell cycle
To elucidate EXOSC3’s mechanistic role in HCC proliferation, we employed EXOSC3-siRNA knockdown models to investigate cell cycle regulation. Flow cytometry revealed EXOSC3 knockdown significantly increased G1-phase populations in both Huh-7 and HCCLM3 cells, with concomitant S-phase accumulation and G2-phase reduction (Fig. 5A, D). Western blot analysis at 72 h post-transfection demonstrated downregulation of critical G1/S transition regulators (cyclin D1, CDK4, cyclin E1, p-CDK2 Thr160) (Fig. 5E, F), while cyclin B1 remained unaffected. These findings establish EXOSC3’s pivotal role in G1/S progression. Given Wnt/β-catenin’s known regulation of cyclin D1, we examined pathway components and observed significant β-catenin and C-myc downregulation following EXOSC3 knockdown (Fig. 5G, H), suggesting potential Wnt pathway modulation. While these data implicate EXOSC3 in coordinating cell cycle progression through both direct and Wnt-mediated mechanisms, additional validation studies are warranted.
Knocking down EXOSC3 inhibits HCC cell proliferation by regulating the G1/S phase of the cell cycle. Flow cytometry analysis of the cell cycle distribution of Huh-7 (A) and HCCLM3 (B) cells in G1, S, and G2 (C, D) in control and siEXOSC3 transfection treated groups. (E, F) Expression of cell cycle key proteins in the control and EXOS3C knockdown groups. (G-H) Protein expression levels of two markers of the Wnt/β-catenin pathway were determined in HCC cells after EXOSC3 knockdown by western blot analysis. Statistical significance levels:*p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
Silencing EXOSC3 causes HCC to arrest at the G1/S phase via P53/P21/Cyclin/CDK
To delineate EXOSC3-mediated signaling pathways in HCC, we performed R-based co-expression analysis using TCGA data, identifying 192 significantly correlated genes (r > 0.6, p < 0.05). KEGG and GSEA analyses revealed predominant enrichment in cell cycle and p53 signaling pathways, with GSEA specifically highlighting cell cycle checkpoint involvement (Fig. 6A, B)26,27. Focusing on p53 signaling due to its established role in proliferation regulation, we examined EXOSC3 knockdown effects. Silencing EXOSC3 upregulated both p53 and p21 expression while increasing p53 phosphorylation (Fig. 6C, D).
Silencing EXOSC3 causes HCC to arrest at the G1/S phase via P53/P21/Cyclin/CDK. (A, B) based on KEGG and GSEA analysis of co-expressed genes.(C, F) Key protein expression and the mean protein expression levels of the indicated proteins in the NC and EXOS3C knockdown groups. Statistical significance levels:*p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
The retinoblastoma protein (RB) forms inhibitory complexes with E2F transcription factors, suppressing G1/S transition genes. This regulation depends on RB phosphorylation status, which p21 modulates through CDK2 inhibition.As a p53 transcriptional target, p21 links p53 activity to cell cycle control28. Consistent with this mechanism, EXOSC3 knockdown reduced p-RB levels, demonstrating EXOSC3’s role in modulating the p53/p21/RB axis to promote HCC progression.
To investigate this mechanism, the protein level expression of p-RB was verified(Fig. 6D) . As anticipated, the protein expression level of p-RB in the siEXOSC3 group was significantly lower compared to the NC group (Fig. 6E).These results suggest that EXOSC3 may promote the progression of HCC by modulating the P53/P21/RB signaling pathway.
Discussion
Exosome complex (EXOC), originally identified as an RNA degradation machinery, has gained increasing attention due to its family members’ (EXOSCs) involvement in various human diseases, particularly cancer15. Hepatocellular carcinoma (HCC) is one of the most aggressive gastrointestinal malignancies. While HCC pathogenesis has been extensively studied, the specific roles of EXOSC components, particularly EXOSC3 in HCC, remain poorly understood. We adopted comprehensive bioinformatics analyses to identify EXOSC3 overexpression as a clinically significant prognostic biomarker in HCC tissues.
Based on GO and KEGG pathway analyses demonstrating EXOSC3’s involvement in cell cycle regulation (Fig. 4), we focused EXOSC3’s possible mechanistic investigation on this critical biological process, which is precisely coordinated by cyclin-dependent kinases (CDKs) and their regulatory cyclin partners29,30. These regulatory components (including CDK4/6-Cyclin D and CDK2-Cyclin E complexes) orchestrate progression through G1, S, G2, and M phases, with dysregulation frequently observed in cancers31. Our findings demonstrate that EXOSC3 knockdown induces G1/S phase arrest in HCC cells, and delaying the transition to the G2/M phase, accompanied by downregulation of critical G1/S transition regulators (cyclin D1, CDK4, cyclin E1, p-CDK2 Thr160). These results not only establish EXOSC3’s role in HCC cell cycle control but also suggest its potential as a therapeutic target through modulation of CDK-cyclin complexes.
To further investigate EXOSC3’s regulatory effects, KEGG analysis of its co-expressed DEGs revealed significant p53 pathway enrichment. This pathway can regulate cell cycle by p53 binding the P21 promoter to induce expression32,33. The cyclin-dependent kinase inhibitor p21 blocks CDK activity, maintaining RB in a hypophosphorylated state. As a transcriptional corepressor, RB plays a central role in cell cycle control. During G1-S transition, CDK-mediated RB phosphorylation releases E2F transcription factor, whereas hypophosphorylated RB binds E2F to induce cell cycle arrest28.Consistent with expectations, EXOSC3 silencing activated the P53 pathway, as demonstrated by elevated P53, p-P53 and p21 expression in HCC cells. Increased p21 expression correlated with reduced RB phosphorylation and decreased cyclin D1/E1 levels. These findings demonstrate that EXOSC3 knockdown triggers P53-p21-RB pathway activation, inducing G1/S phase arrest in HCC cells. Additionally, Wnt/β-catenin pathway can regulate cyclin D1 expression and influence cell cycle progression. Therefore, we analyzed two key downstream effectors of the Wnt/β-catenin pathway: β-catenin and c-Myc. Our results demonstrated a consistent downregulation of both proteins, implying involvement of Wnt/β-catenin signaling pathway in EXOSC3-mediated regulation of cyclin D1 expression.
EXOSC3 simultaneously regulates both the p53 and Wnt signaling pathways, suggesting its potential role as a convergence point in the oncogenic signaling network. EXOSC3 knockdown increases p53 phosphorylation (possibly via MDM2/WIP1) while decreasing β-catenin/c-Myc and cyclin D1. Potential crosstalk (interaction of p21 and β-catenin) requires validation through dual perturbation experiments (EXOSC3 silencing + p53 knockdown) to elucidate synergistic cell cycle arrest mechanisms and therapeutic implications.
While our in vitro findings demonstrate EXOSC3’s mechanistic role in HCC cell cycle arrest, we emphasize that these conclusions require validation in in vivo models and clinical cohorts. The current study’s reliance on cell-line data limits direct translation to human pathophysiology, as tumor microenvironmental factors and systemic influences remain unaddressed. Future studies employing orthotopic models and patient-derived tissues will be essential to confirm EXOSC3’s therapeutic potential.
In summary, this study reveals the critical involvement of EXOSC3 in hepatocellular carcinoma (HCC) pathogenesis. Comparative analysis demonstrated significantly elevated EXOSC3 expression in HCC tissues relative to normal liver counterparts, with its overexpression strongly correlating with reduced patient survival, supporting its putative oncogenic function. Mechanistically, EXOSC3 knockdown triggered activation of p53 tumor suppressor pathway and suppressed Wnt/β-catenin signaling cascade, resulting in G1 phase cell cycle arrest and consequent proliferation inhibition. Collectively, these findings position EXOSC3 as a promising molecular target for novel HCC therapeutic strategies. As a non-catalytic component of the human exosome complex (which possesses 3’-5’ exoribonuclease activity), EXOSC3 participates in diverse RNA processing and degradation pathways. Future studies to map EXOSC3’s target mRNAs will elucidate its transcript-selective regulatory mechanism in HCC, supporting its dual potential as both a prognostic biomarker and therapeutic target.
Data availability
The bioinformatics analysis concerned data originated from online TCGA and GEO databases. To access the data supporting this study’s findings, please contact the corresponding author with a reasonable request.
Abbreviations
- EXOSC3:
-
Human exosome component 3
- HCC:
-
Hepatocellular carcinoma
- qRT-PCR:
-
Real-time quantitative reverse transcription PCR
- CCK-8:
-
Cell counting kit-8
- GO:
-
Gene ontology
- KEGG:
-
Kyoto encyclopedia of genes and genomes
- TCGA:
-
The cancer genome atlas
- HPA:
-
Human protein atlas
- EMT:
-
Epithelial mesenchymal transition
- CDKs:
-
Cyclin-dependent kinases
- pRB, RB1:
-
Retinoblastoma-associated protein
References
Vogel, A. et al. Hepatocellular carcinoma. Lancet 400(10360), 1345–1362. https://doi.org/10.1016/S0140-6736(22)01200-4 (2022).
Pol, S. Carcinome Hépatocellulaire (CHC) [Hepatocellular carcinoma (HCC)]. Med. Trop. Sante. Int. 4(4), mtsi.v4i4.2024.614. https://doi.org/10.48327/mtsi.v4i4.2024.614 (2024).
Ganesan, P. & Kulik, L. M. Hepatocellular carcinoma: New developments. Clin. Liver. Dis. 27(1), 85–102. https://doi.org/10.1016/j.cld.2022.08.004 (2023).
Januszyk, K. & Lima, C. D. The eukaryotic RNA exosome. Curr. Opin. Struct. Biol. 24, 132–140. https://doi.org/10.1016/j.sbi.2014.01.011IF (2014).
Kilchert, C., Wittmann, S. & Vasiljeva, L. The regulation and functions of the nuclear RNA exosome complex. Nat. Rev. Mol. Cell. Biol. 17(4), 227–239. https://doi.org/10.1038/nrm.2015.15 (2016).
Lim, J. et al. Nuclear proximity of Mtr4 to RNA exosome restricts DNA mutational asymmetry. Cell 169(3), 523-537.e15. https://doi.org/10.1016/j.cell.2017.03.043 (2017).
Pefanis, E. & Basu, U. RNA exosome regulates AID DNA mutator activity in the B cell genome. Adv. Immunol. 127, 257–308. https://doi.org/10.1016/bs.ai.2015.04.002 (2015).
Liu, Q. et al. Exosome component 1 cleaves single-stranded DNA and sensitizes human kidney renal clear cell carcinoma cells to poly(ADP-ribose) polymerase inhibitor. Elife 23(10), e69454. https://doi.org/10.7554/eLife.69454 (2021).
Zhu, G. et al. Comparative proteomics identified EXOSC1 as a target protein of anticancer peptide LVTX-8 in nasopharyngeal carcinoma cells. J. Proteome Res. 23(6), 2028–2040. https://doi.org/10.1021/acs.jproteome.4c00031 (2024).
Lv, C. G. et al. EXOSC2 mediates the pro-tumor role of WTAP in breast cancer cells via activating the Wnt/β-catenin signal. Mol. Biotechnol. 66(9), 2569–2582. https://doi.org/10.1007/s12033-023-00834-8 (2024).
Pan, H. et al. EXOSC5 as a novel prognostic marker promotes proliferation of colorectal cancer via activating the ERK and AKT pathways. Front Oncol. 18(9), 643. https://doi.org/10.3389/fonc.2019.00643 (2019).
Cui, K. et al. EXOSC8 promotes colorectal cancer tumorigenesis via regulating ribosome biogenesis-related processes. Oncogene 41(50), 5397–5410. https://doi.org/10.1038/s41388-022-02530-4 (2022).
Quttina, M. et al. Exosc9 Initiates SUMO-dependent lncRNA TERRA degradation to Impact telomeric integrity in endocrine therapy insensitive hormone receptor-positive breast cancer. Cells 12(20), 2495. https://doi.org/10.3390/cells12202495 (2023).
Meng, Z. Y. et al. EXOSC10 is a novel hepatocellular carcinoma prognostic biomarker: a comprehensive bioinformatics analysis and experiment verification. PeerJ 8(11), e15860. https://doi.org/10.7717/peerj.15860 (2023).
Zhang, Y. et al. Integrated bioinformatic investigation of EXOSCs in hepatocellular carcinoma followed by the preliminary validation of EXOSC5 in cell proliferation. Int. J. Mol. Sci. 23(20), 12161. https://doi.org/10.3390/ijms232012161 (2022).
Houseley, J., LaCava, J. & Tollervey, D. RNA-quality control by the exosome. Nat. Rev. Mol. Cell Biol. 7(7), 529–539. https://doi.org/10.1038/nrm1964 (2006).
Lloret-Llinares, M. et al. The RNA exosome contributes to gene expression regulation during stem cell differentiation. Nucl. Acids Res. 46, 11502–11513 (2018).
Zinder, J. C. & Lima, C. D. Targeting RNA for processing or destruction by the eukaryotic RNA exosome and its cofactors. Genes Dev. 31(2), 88–100. https://doi.org/10.1101/gad.294769.116 (2017).
Baas F, van Dijk T. EXOSC3 pontocerebellar hypoplasia. 2014 Aug 21 [updated 2020 Sep 24]. In: Adam MP, Feldman J, Mirzaa GM, Pagon RA, Wallace SE, Amemiya A, editors. GeneReviews® [Internet]. Seattle (WA): University of Washington, Seattle; 1993–2025. PMID: 25144110.
Morton, D. J. et al. The RNA exosome and RNA exosome-linked disease. RNA 24(2), 127–142. https://doi.org/10.1261/rna.064626.117IF (2018).
Halevy, A. et al. Novel EXOSC3 mutation causes complicated hereditary spastic paraplegia. J. Neurol. 261(11), 2165–2169. https://doi.org/10.1007/s00415-014-7457-x (2014).
Tsuda, M. et al. Aberrant expression of MYD88 via RNA-controlling CNOT4 and EXOSC3 in colonic mucosa impacts generation of colonic cancer. Cancer Sci. 112(12), 5100–5113. https://doi.org/10.1111/cas.15157 (2021).
Li, T. et al. TIMER2.0 for analysis of tumor-infiltrating immune cells. Nucleic Acids Res. 48(W1), W509–W514. https://doi.org/10.1093/nar/gkaa407 (2020).
Chandrashekar, D. S. et al. UALCAN: An update to the integrated cancer data analysis platform. Neoplasia 25, 18–27. https://doi.org/10.1016/j.neo.2022.01.001 (2022).
Goldman, M. J. et al. Visualizing and interpreting cancer genomics data via the Xena platform. Nat Biotechnol. 38(6), 675–678. https://doi.org/10.1038/s41587-020-0546-8 (2020).
Kanehisa, M. et al. KEGG: biological systems database as a model of the real world. Nucleic Acids Res. 53(D1), D672–D677. https://doi.org/10.1093/nar/gkae909 (2025).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 28(11), 1947–1951. https://doi.org/10.1002/pro.3715 (2019).
Engeland, K. Cell cycle regulation: p53–p21-RB signaling. Cell Death Differ. 29(5), 946–960. https://doi.org/10.1038/s41418-022-00988-z (2022).
Satyanarayana, A. & Kaldis, P. Mammalian cell-cycle regulation: several Cdks, numerous cyclins and diverse compensatory mechanisms. Oncogene 28(33), 2925–2939. https://doi.org/10.1038/onc.2009.170 (2009).
Wang, Z. Cell cycle progression and synchronization: An overview. Methods Mol. Biol. 2579, 3–23. https://doi.org/10.1007/978-1-0716-2736-5_1 (2022).
Gao, X., Leone, G. W. & Wang, H. Cyclin D-CDK4/6 functions in cancer. Adv. Cancer Res. 148, 147–169. https://doi.org/10.1016/bs.acr.2020.02.002 (2020).
Jung, Y. S., Qian, Y. & Chen, X. Examination of the expanding pathways for the regulation of p21 expression and activity. Cell Signal. 22(7), 1003–1012. https://doi.org/10.1016/j.cellsig.2010.01.013 (2010).
Thangavelu, L. et al. Targeting the p53–p21 axis in liver cancer: Linking cellular senescence to tumor suppression and progression. Pathol Res Pract. 263, 155652. https://doi.org/10.1016/j.prp.2024.155652 (2024).
Acknowledgements
We thank all members of Professor li-he Jiang’ team for technical support suggestions. We would like to extend our gratitude to the Life Science Research Institute, Youjiang Medical University for Nationalities, for providing the equipment and facilities necessary for our experiments.
Funding
Guangxi Natural Science Foundation (Grant No. 2025GXNSFHA069094); Guangxi Key Research & Development Plan (Grant No.2023AB23075); Guangxi Natural Science Foundation Program (Grant No.2022JJA140478); Baise City Science and Technology Plan Project(Science and Technology Infrastructure Support Program) (Grant No. ZJ252812);Grant of research project on high-level talents of Youjiang Medical College for Nationalities (Grant No. YY2021SK02);Guangxi Key Laboratory of Tumor Immunology and Microenvironmental Regulation (2024KF013);Open Project Program of Key Laboratory of Early Prevention and Treatment for Regional High Frequency Tumor, Ministry of Education, Guangxi Medical University (Grant No. GKE-KF202006); Fujian Provincial Key Laboratory of Innovative Drug Target Research(Grant No. FJ-YW-2025KF06); Grant of Guangxi Provincial-Level Project of University Students’ Innovation and Entrepreneurship (Grant No. S202410599049, S202510599070). All authors declare that they have no financial, personal interests or beliefs that affect their objectivity, and have no economic or personal relationships with other people or organizations that improperly influence or prejudice their works.
Author information
Authors and Affiliations
Contributions
Yu-Mo Chen conceived and designed the experiments, performed the experiments, analyzed the data, prepared figures and/or tables, authored or reviewed drafts of the article.Yi-Xuan Huo, Wan-Shuo Wei, Yi-Xin Liu,Shu-Han Wang, Lin Jiang and Rong-He performed the experiments, authored or reviewed drafts of the article, and approved the final draft. Hong-liang Zhang conceived and designed the experiments, authored or reviewed drafts of the article, and approved the final draft. Li–He Jiang conceived and designed the experiments and, funded research, instructed students to conduct experiments, analyzed the data, prepared figures and/or tables, authored or reviewed drafts of the article, and approved the final draft.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Ethics approval and consent to participate
All procedures were performed according to the Ethical Guidelines for Human Genome/Gene Research and were approved by the Ethics Committee of Youjiang Medical College for Nationalities (2022090501).
Consent for publications
All listed authors have actively participated in the study and have read and approved the submitted manuscript.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chen, YM., Huo, YX., Wei, WS. et al. EXOSC3 knockdown induces G1/S phase arrest to suppress hepatocellular carcinoma cell proliferation. Sci Rep 15, 44684 (2025). https://doi.org/10.1038/s41598-025-28270-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-28270-5