Abstract
Follicular development is closely related to poultry egg production. Granulosa cells (GCs) are important components of follicles and vital for follicular selection and hierarchical development. Previous studies demonstrated that miRNAs play a key role in controlling follicular development and atresia. However, biological effects and mechanism of action of miR-726-3p on GC proliferation and differentiation of hen ovaries remain poorly understood. In this study, we compared miRNA profiles in larger white follicles (LWF), along with large (LYF) and small yellow follicles (SYF), to identify target genes related to follicular development. Nine cDNA libraries were constructed, in which 61 miRNA-linked genes were significantly differentially expressed (DE). MiR-726-3p was the most crucial co-DE miRNA and, therefore, was chosen for further evaluation. Using bioinformatics and the luciferase reporter assay, we identified CYB5A as the target of miR-726-3p. Over-expression of miR-726-3p significantly negatively regulated GC proliferation, differentiation and knocking down CYB5A expression supported miR-726-3p overexpression. Overall, miR-726-3p suppresses GC proliferation and differentiation by targeting CYB5A, thus providing new insight into the mechanism that underlies hen follicle development.
Similar content being viewed by others
Introduction
Follicular development is closely related to poultry egg production which is economically important. In growing follicles, there are three key developmental stages: (1) pre-hierarchical 3.5–5.5 mm slow larger white follicles (LWF) and (2) 6.0–8.0 mm small yellow follicles (SYF), and (3) pre-ovulatory/hierarchical 8.5–10.5 mm differentiated large yellow follicles (LYF)1. Follicles with varied sizes occur at different stages and possess distinct genetic characteristics. Most ovarian follicles experience atresia, because only about 5% progress to the final hierarchical stages of maturation and ovulation2.
Follicles are primarily composed of granulosa cell layers (GCs), theca cell layers, and oocytes. Granulosa cells (GCs) play an important role in folliculogenesis, follicular development, and atresia3. Although both GCs and oocytes can cause a degenerative process of follicles known as “atresia”, mostly involving GC apoptosis4. In the early stage of atresia, apoptosis occurs in GCs, but not in cumulus cells, oocytes, or cells of internal or external theca layers5. Therefore, GCs are vital for studying follicular selection and development.
MicroRNA (miRNA) is a kind of endogenous short-strand non-coding RNA having 19 to 24 nucleotides, that are expressed and can control various biological processes by regulating development and cellular homeostasis6. Pairing the complementary 3’ Untranslated Region (3’-UTR) in targeted mRNAs, miRNAs trigger degradation or inhibition of gene expression on the transcriptional, translational, or epigenetic levels7,8. Chickens (Gallus gallus domesticus) are an important model and provide a major source of protein worldwide. Some miRNAs have been identified in chickens and play an important role in chicken follicle development in terms of steroidogenesis, ovulation, apoptosis and gene regulation9,10,11. Kang et al. (2013) first reported miRNA expression profiles of sexually immature and mature ovaries in chickens12. Several miRNAs have been shown to regulate granulosa cell function in chickens and mammals13,14,15. However, understanding how miRNAs are involved in controlling follicular development by regulating GCs remains to be studied.
By using high-throughput sequencing of small-RNA cDNA libraries in the eyes of zebrafish, miR-726-3p was first discovered as one of the 66 new miRNAs16. The expression of miR-726-3p was most abundant in 5-day-old larvae in five developmental stages: 24 h, 48 h, 72 h, 5 days and dropped significantly in adulthood. MiR-726-3p thermally sensitive in zebrafish ovaries due to its temperature- repressed expression. However, the effects of miR-726-3p on the proliferation and differentiation of GCs have not been reported.
Follicle development is also closely associated with proliferation and differentiation of GCs and theca cells, oocyte maturation, interactions of oocyte and GC. It is strictly controlled by various reproductive endocrine hormones, paracrine and autocrine regulatory factors, as well as regulation of the hypothalamic-pituitary-gonadal axis17,18. The most representative factors include steroidogenic acute regulatory protein (STAR), cytochrome P450 side-chain cleavage (P450scc/CYP11A1), cell proliferation-related factors PCNA and apoptosis-related factors Bcl-2 and caspase-3 (CASP3) have been extensively reported19,20,21,22,23. The genes of STAR and CYP11A1 are not only involved in steroidogenesis, but they also facilitate GC differentiation24.
Cytochrome b5 type A (CYB5A) plays a pivotal role in steroidogenesis where cholesterol produces pregnenolone catalyzed by P450scc as encoded by CYP11A1. In the delta-4 pathway, pregnenolone is converted to progesterone and then to androstenedione, a process that is catalyzed by CYP17A1. CYB5A affects this reaction as a regulator of androstenedione production25,26. CYB5A interacts allosterically with the CYP17A1-P450 oxidoreductase complex to stimulate lyase activity25,27. In the human adrenal gland, CYP17 has high 17,20-lyase activity only in the delta-5 pathway28. The 17,20-lyase activity occurs via its association with CYB5A29. In the △5 pathway, pregnenolone is synthesized via DHEA into androstenedione. The efficient androstenedione production appears to require both 3βHSD and CYB5A26. Androstenedione is a common precursor of sex steroids, which are converted and aromatized into estradiol (E2) by CYP19A130.
In the present study, we identified the key miRNAs associated with follicle selection by using small RNA transcriptome analysis on LWF, SYF and LYF of Jilin black chickens. Key miRNAs associated with follicle selection, i.e. miR-726-3p, were identified, and the miRNA-targets between miR-726-3p and CYB5A were determined by using bioinformatics and the dual-luciferase assay. We verified the effects of miR-726-3p on granulosa cell proliferationand differentiation. Overall, this study lays the foundation for further research on the exact role and regulatory mechanism of miRNAs in ovarian development and provides a basis to understand chicken egg-laying traits for breeding and genetic improvement.
Materials and methods
Institutional review board statement
The animal study protocol was approved by Jilin Agricultural University’s Institutional Animal Careand Use Committee (IACUC) (Changchun, China)(Permission No. GR (J) 18 − 010). All animal experiments were performed in accordance with recommendations in the Guide for the Care and Use of Laboratory Animals of the National Institutes of Health. All animal studies complied with the ARRIVE (Animal Research: Reporting of In Vivo Experiments) guidelines. Thirty 210-day-old Jilin Black (JB) hens from the breeding base of Jilin Agricultural University were used in this study. These hens were bred and maintained under standard conditions, including controlled temperature, humidity, and light cycles, to ensure optimal health and welfare. They were fed a balanced diet formulated specifically for their age and breed, and their care was overseen by experienced animal husbandry staff to ensure compliance with ethical standards.
Sample collection and RNA isolation
The Jilin Black chicken, an indigenous Chinese breed (dual-purpose type for egg and meat production), is characterized by high quality meat, but a low rate of egg production, low laying peak, and strong broodiness31.Jilin Black hens were reared in laying battery cadges until they were 21 weeks old24. Their average egg production rate at that age was 11%. Ten hens of each lineage were euthanized. Following the completion of behavioral experiments, chicks were deeply anesthetized via intraperitoneal injection of sodium pentobarbital (150 mg/kg). After confirming loss of consciousness and absence of corneal reflex, euthanasia was completed by rapid cervical dislocation. This method involved firmly stabilizing the head at the occipital joint and applying immediate, forceful traction to the body to ensure severance of the spinal cord. Death was verified by cessation of vital signs and confirmed via thoracotomy. All procedures complied with the AVMA Guidelines (2020)32. After chickens are euthanized, their follicles are collected. Three different types of follicles sized 3.5–5.5, 6.0–8.0, and 8.5–10.5 mm in diameter, i.e. large white follicles (LWF), small yellow follicles (SYF) and large yellow follicles (LYF) were chosen. The surrounding vascular and connective tissues of the follicles were removed by fine forceps and a scalpel and the remaining follicles were immediately frozen in liquid nitrogen and preserved at −80℃ until RNA extraction. All experiments and methods were performed according to the ARRIVE guidelines.
After pre-cooling the mortar in liquid nitrogen, take an appropriate amount of follicular tissue and quickly grind it into a powder. Using sterilized forceps, transfer the powdered follicular tissue sample quickly into a 1.5 mL EP tube containing 1 mL of Trizol reagent (Invitrogen Life Technologies, Carlsbad, CA, USA). Mix thoroughly and allow to lyse for 10 min.The concentration, quality and integrity were tested by a Thermo NanoDrop 8000 Spectrophotometer (Thermo Fisher Scientific, Wilmington, Delaware, USA). Three micrograms were used for RNA sample preparations, and the remaining materials were stored for later validation.
Library preparation and sequencing
Small RNA-Seq libraries were prepared by using the TruSeq® Small RNA Library Prep Kit according to the manufacturer’s protocol. In this study, 9 hens were selected. Based on the different stages of follicular development, 3 follicles were sampled at each stage, resulting in a total of 9 follicles. Nine cDNA libraries of the LWF (named JB1, including JB11、JB12、JB13), SYF (named JB2, including JB21、JB22、JB23) and LYF (named JB3, including JB31、JB32、JB33) from Jilin Black (JB) hens were constructed, respectively. Library quality was assessed on the Agilent Bioanalyzer 2100 system using DNA High Sensitivity Chips. Construction and sequencing of miRNA libraries were sent to Shanghai Personal Biotechnology Co., Ltd. (China) for miRNA-omics analysis.
Raw data, consisting of 18–36 nt sequence adaptors, first went through a data cleaning process to remove low quality adaptors, having quality values less than 20, 5’ and 3’ primer contaminants, N adaptors, polyA adaptors, sequences without insert tags and adaptors shorter than 18 nt or longer than 36 nt. All of the clean reads were aligned with the reference genome (ftp://ftp.ensembl.org/pub/release-86/fasta/gallusgallus/dna/Gallusgallus.Gallusgallus5.0.dna.toplevel.fa.gz 5.0.dna.toplevel.fa.gz) by using sequence alignment program HISAT 2.1.033. And rRNA, tRNA, miscRNA, snRNA and snoRNA were discarded from the small RNA sequences; the remaining sequences were searched against the miRBase 21.0 database of known Gallus gallus miRNA sequences (http://www.mirbase.org/)34,35,36. The sequences matching Gallus gallus miRBase by Bowtie were considered known miRNA sequences. The remaining sequences matching chicken genome sequences (except for one to three 5’- or 3’-end nt) were used to predict novel miRNAs by MIRDEEP2(https://www.mdcberlin.de/8551903/en/research/research_teams/systems_biology_of_gene_regulatory_elements/projects/miRDeep) using default parameters37,38. These sequences were considered as a potential novel miRNA, and expression of all miRNAs was assayed.
MiRNA transcriptomic analysis
Reads of miRNAs were calculated and reads per million miRNA mapped values were used to represent expression levels (RPM, which is the ratio of the count of miRNA to the total count of clean reads multiplied by 1,000,000). p < 0.05 and |log2fold change| >1 were used to screen differentially expressed miRNAs using the DESeq R package39 for 6 groups, i.e. LWF vs. SYF, SYF vs. LYF, LWF vs. LYF in JB hens, respectively. Hierarchical cluster analysis was performed with Cluster 3.0 and TreeView1.6 programs (http://rana.lbl.gov/eisen, accessed on 19 November 2019). Volcano plots to visualize up- and down-regulated genes of each sample were generated by using the R package “ggplot2” (version 3.5.0). A Venn diagram was constructed to visualize the co-expressed differentially expressed miRNA.
MiRNA target gene prediction and functional annotation
The miRNA target prediction software miRDB (http://mirdb.org/miRDB/)40 and online software programs in the TargetScan (http://www.targetscan.org/) website were used to predict the binding sites of the differentially expressed miRNA. MiRanda 0.8.441 was also used to predict the target relationships with miRNAs, which needed an alignment score of N ≥ 140, and required energy less than − 10 kcal/mol. The intersection of these three prediction programs was selected. The main functions of the predicted target genes regulated by differentially expressed miRNAs were determined using GO (gene ontology) and KEGG (Kyoto Encyclopedia of Genes and Genomes) functional classifications by Blast2GO program42. The P-value cut-off was 0.05 for GO and KEGG terms.
Cell culture
Granulosa cell (GCs) from the 6–8 mm follicles of Jilin Black (JB) hens were isolated with a needle and rinsed with Medium199 (M199; Gibco, Waltham, MA, USA) with 10% (v/v) fetal calf serum (Gibco) at 37 °C in a humidified incubator under 5% CO2 according to our published method43,44. The cultured GCs used in this experiment have been purified and quantified in our laboratory. The specificity of the GCs has been identified by the H & E staining procedure and fluorescence staining analysis45,46.
Plasmid (vector) construction
The fragment miR-726-3p (MIMAT0050086) was cloned into the pcDNA3.1-3xFlag-C vector between the kpn I and Xho I sites, using the In-Fusion® HD Cloning Kit (Takara, Dalian, China). The miR-726-3p expression vector pcDNA3.1-miR-726-3p were verified by sequencing technology (Fig. S1).
Small interfering RNA (siRNA) transfection
Three small interfering (siRNA) sequences targeting CYB5A gene were designed and the most effective siRNA, i.e. siRNA406 was screened out (Table S1). GCs were then transfected with the miR-726-3p expression vector pcDNA3.1-miR-726-3p or CYB5A-siRNA in 24-well plates (2 × 105 cells/well; Corning Costar®, catalog no. 3524, Corning, NY, USA) and incubated at 37℃ with 5% CO2. After culturation for 24 h or longer, the GCs were collected for EdU cell proliferation, CCK-8 proliferation and cell apoptosis assays, and lysed for Western blotting, RT-qPCR and E2 concentration analysis.
Data validation by quantitative real-time RT-PCR
Transcription levels of the selected gene regions were assessed by quantitative real-time reverse transcriptase PCR (RT-qPCR) as the method described43. The primers utilized for amplification were listed in Table S2. The 2−△△CT method was used to calculate the relative expression47. In addition, 18 S rRNA was used as an endogenous control to normalize the RT-qPCR results for each miRNA24. All reactions were run in three technical replicates and included negative controls without template.
Western blotting
Western blotting was performed using total cellular extracts following our previously description43,44. The affinity-purified antibodies were used (Table S3). Proteins were separated by 10% SDS-polyacrylamide gel. The gels were then electro-transferred to Protran nitrocellulose membranes (Whatman, Dassel, Germany). After electrophoresis of the protein samples in a mini gel apparatus, a pre-stained protein molecular weight marker was loaded to monitor the target proteins in the gels. At the approximated protein size position, the gel was directly cut and transferred to the nitrocellulose membrane for western blotting. Anti-β-actin antibody acted as a loading control. Diluted primary antibodies were incubated with the membranes overnight at 4 ◦C. The horseradish peroxidase-conjugated anti-rabbit or anti-mouse IgG secondary antibody was incubated for 2 h at 22–25 °C. Blots were subsequently performed with ECL western blotting agent (Rockford, IL, USA) for 5 min and exposed to X-ray film for 1–5 min. The signals were detected using the ECL Plus Western blotting detection system (Thermo Fisher Scientific, Waltham, MA, USA; catalog no. 32132). Briefly, membranes were incubated with a 1:1 mixture of ECL reagent A and freshly prepared reagent B for 5 min at 22–25 °C in the dark. Chemiluminescent signals were captured using a ChemiDoc XRS + imaging system (Bio-Rad, Hercules, CA, USA) with exposure times optimized between 10 s and 5 min to avoid signal saturation. Band intensities were quantified using Image Lab software (version 6.0.1, Bio-Rad) and normalized to the corresponding β-actin bands. The images show the original gels of western blot in the Fig. S2-20.
Luciferase reporter assay
293 T cells, sourced from ATCC (Manassas, VA, USA, cat. no. CRL-3216), were seeded in 24-well plates in triplicate and co-transfected with each dual-luciferase (Firefly luciferase and Renilla luciferase) reporter gene plasmid (pmirGLO) containing the wild-type or the mutant plasmid, and with miR-726-3p mimics or miR-NC mimics (negative control) using Lipofectamine™ 2000 transfection reagent (Invitrogen). At 48 h post-transfection, luciferase activity was measured using the Dual-Luciferase Reporter Assay System (Promega, Madison, WI, USA).
Cell counting Kit-8 proliferation assay
Cell proliferation was measured using the CCK-8 assay (Cell Counting Kit-8, Biyuntian, Shanghai, China; catalog no. C0038). GCs were seeded into 96-well culture plates (Corning Costar®, catalog no. 3599, Corning, NY, USA) at a density of 1 × 105cells/mL and cultured at 37 ℃ for 24 h. GCs were transfected with plasmid pcDNA3.1-miR-726-3p. Twenty-four and 48 h after transfection, we measured the absorbance at 450 nm on a microplate reader.
EdU cell proliferation assay
Cell proliferation was detected after the shRNA transfection of the GCs with CYB5A-siRNA406 by EdU (5’-Ethynyl-2’-deoxyuridine) incorporation assay using the Cell-Light EdU imaging kit (RiboBio, Guangzhou, China; catalog no. C10310-3). Briefly, at 48 h post-transfection, GCs were incubated with 50 µM EdU working solution (diluted in complete culture medium) at 37℃ with 5% CO₂ for 2 h. After removing the EdU solution, cells were fixed with 4% paraformaldehyde for 15 min at 22–25 °C, permeabilized with 0.5% Triton X-100 for 20 min, and then incubated with the Click-iT reaction mixture (containing Alexa Fluor 488 azide, copper sulfate, and sodium ascorbate) for 30 min in the dark. Nuclei were counterstained with 4’,6-diamidino-2-phenylindole (DAPI) for 10 min. Images were captured using a fluorescence microscope (Olympus, Tokyo, Japan; model IX73) with a 20× objective, and the proliferation rate was calculated as the percentage of EdU-positive cells (green fluorescence) relative to the total number of DAPI-stained cells (blue fluorescence) in five randomly selected fields per well.The number of EdU positive cells was divided by the total cells as previously described43. We performed each experiment in triplicate and repeated five times.
Flow cytometry assay
The rate of apoptosis of GCs was measured by Annexin V-FITC/PI apoptosis kit (KeyGen Biotech Co., Ltd., Jiangsu, China). GCs were digested using 0.25% trypsin (without EDTA) after transfection with vector with miR-726-3p or CYB5A -siRNA406 and washed twice with cold PBS. The cells were then stained with FITC and PI, and detected the apoptosis rate by flow cytometry (BD Biosciences, USA). The cell cycle was also tested by flow cytometry. GCs were digested by trypsin, washed with PBS and suspended, added cold 80% alcohol. The cells were fixed for 4 h at 4 ℃, and incubated 0.5 h at 37 ℃ with 1 mg/mL Rnase 10 µL. After stained with 400 µg/mL PI (10 µL), the cells during the cycle were counted.
Statistical analysis
Statistical analyses were performed using GraphPad Prism 8.0 and SPSS 22.0 software (IBM Corp, Armonk, NY, USA). Comparison between two groups was analyzed using the Student’s t-test after confirmation of normal distributions. Values are displayed as the mean ± SEM with asterisk(s) that indicate the significant difference compared with control group at ** p < 0.01, * p < 0.05.
Results MiRNA transcriptomic analysis of the Hen ovarian follicles
In total, 52.90, 50.05, 53.81 million clean reads in LWF, SYF, LYF of JB hens were used for analysis (Table S4). The percentage of clean reads with Q20 and Q30 were greater than 99.40 and 98.20% (Table S4). Differentially expressed genes (DEGs) in the follicles of LWF vs. SYF, SYF vs. LYF, LWF vs. LYF in JB hens were identified. As a result, 22 miRNA-linked genes were significantly differentially expressed (DE) miRNAs between JB1 and JB2, in which 13 were significantly up-regulated and 9 were down-regulated (Fig. 1a and c, Table S5). Nine miRNA-linked genes were significantly DE between JB1 and JB3, in which four were significantly up-regulated and five were down-regulated (Fig. 1a and e, Table S6). Likewise, 30 miRNA- linked genes were significantly DE miRNAs between JB2 and JB3, in which 13 were significantly up-regulated and 17 were down-regulated (Fig. 1a and d, Table S7).
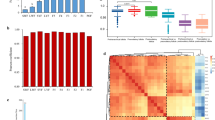
Sequencing data correlation analysis. (a) Venn diagram analysis of distribution of annotated genes of different follicle groups of hens. JB1, LWF of hens; JB2, SYF of hens; JB3, LYF of hens. (b) Common differentially expressed miRNAs in comparison groups. (c, d, e) Inter group differential expression of miRNAs Volcano map. Red dots represent up-regulated genes, blue dots represent down-regulated genes and gray dots indicate non-differentially expressed genes (p-value adjusted for multiple testing < 0.05). B1, LWF of hens; B2, SYF of hens; B3, LYF of hens.
Ten miRNAs were co- differentially expressed between JB1 vs. JB2 and JB2 vs. JB3 groups (Fig. 1b). One miRNA, i.e. miR-726-3p, was co- differentially expressed between JB1 vs. JB2 and JB1 vs. JB3 (Fig. 1b). Two miRNAs were between JB1 vs. JB3 and JB2 vs. JB3. No miRNAs were co- differentially expressed between all of those three groups (Fig. 1b).
MiR-726-3p was not only co-differentially expressed between JB1 vs. JB2 and JB1 vs. JB3, but it also was the only miRNAs that is consistent with co-differentially expressed miRNAs between 3 pairs in another breed, i.e. Lohmann Brown (LB) (data not shown). Therefore, it was selected for further analysis of molecular regulatory mechanisms underlying follicle development.
MiRNA target gene prediction and functional enrichment analysis
Using the principle of complementary base pairing between miRNA and target gene mRNA 3’UTR region (especially seed region complementarity), potential target genes can be predicted through bioinformatics tools.
Because miRNAs affect various biological activities by regulating target genes, the target genes for functional enrichment were analyzed. The GO and KEGG pathway analyses were used to investigate the possible roles of DE genes in follicle development. GO enrichment analysis was performed and a total of 206 significantly enriched GO terms were obtained with p < 0.05 in JB1 vs. JB2 (Table S8). There were 118 biological process (BP) terms, 65 cellular components (CC) terms and 23 molecular function (MF) terms. Likewise, a total of 329 significantly enriched GO terms were obtained in JB2 vs. JB3 with 228 BP terms, 62 CC terms and 39 MF terms in detail (Table S9). A total of 80 significantly enriched GO terms were in JB1 vs. JB3 with 31 BP terms, 41 CC terms and 8 MF terms (Table S10).
Among the 3 pairs, most of the BP terms were related to metabolic processes. The most enriched GO terms include cellular metabolic process, nitrogen compound metabolic process, organic substance metabolic process. The most enriched GO terms in the CC include cytoplasm, cytoplasmic part, intracellular membrane-bounded organelle, and intracellular part. The most enriched GO terms in the MF include protein binding, catalytic activity, identical protein binding, enzyme binding (Fig. 2a, b,c).
GO functional enrichment analysis of target genes. (a) GO functional enrichment analysis of target genes in LWF vs. SYF. (b) GO functional enrichment analysis of target genes in SYF vs. LYF. (c) GO functional enrichment analysis of target genes in LWF vs. LYF.
Eleven significantly enriched KEGG pathways in JB1 vs. JB2 group were focal adhesion, fatty acid degradation, AMPK signaling pathway, cellular senescence, SNARE interactions in vesicular transport, Toll and Imd signaling pathways, cell cycle, endocytosis, PI3K-Akt, chemokine, and Rap1 signaling pathways (p < 0.05) (Table S11)48,49. There were 17 significantly enriched pathways in the JB2 vs. JB3 group, including focal adhesion, cellular senescence, mitophagy-animal, endocytosis, the apelin signaling pathway, apoptosis, SNARE interactions in vesicular transport, Toll and Imd, and Prolactin signaling pathways, fatty acid degradation, oxytocin, AMPK, chemokine, Rap1, and p53 signaling pathways, cell cycle, progesterone-mediated oocyte maturation (Table S12)48,49. Five significantly enriched pathways in the JB1 vs. JB3 group were endocytosis, SNARE interactions in vesicular transport, focal adhesion, the chemokine signaling pathway, and cell cycle (Table S13)48,49.
Notably, there are 5 co-enriched pathways across the 3 groups, i.e. SNARE interactions in vesicular transport (ko04130), focal adhesion (ko04510), cell cycle (ko04110), endocytosis (ko04144), and chemokine signaling pathways (ko04062), which may be key pathways relevant to ovarian follicular development regulated by miRNAs.
Prediction and verification of specific target genes of miR-726-3p
Characterization of specific target genes
Targets for miR-726-3p were predicted by using the programs MIRDB, TargetScan, and MiRanda. The intersection of these three predictions was 37 miRNA target genes (Fig. 3a). To reveal the most important DE genes involved in follicle development across 3 stages, i.e. LWF, SYF and LYF, CYB5A and NSF, the two most promising candidates associated with egg-laying performance were chosen from the 37 target genes. The NSF gene is involved in SNARE interactions in the vesicular transport pathway (ko 04130), and CYB5A is involved in the estradiol hormone synthesis pathway.
Potential binding sites for miR-726-3p within CYB5A and NSF was analyzed using the Targetscan database (Fig. 3b and c). The potential physical interaction network of proteins encoded by CYB5A and NSF was done using the String database as shown in Fig. 3d and e and suggested that NSF and CYB5A are involved in the regulation of vesicular transport and steroid synthesis, respectively, thus laying a foundation to study the genetic mechanism underlying chicken follicle development.
Prediction of miR-726- 3p specific target genes. (a) Three prediction programs was 37 miRNA target genes. (b) Binding sites for miR-726-3p within the CYB5A sequence. (c) Binding sites for miR-726-3p within the NSF sequence. (d) Inferred CYB5A was involved in estradiol hormone synthesis pathway (e) Presumed regulatory network of NSF in SNARE interactions in vesicular transport by using the String database.
Effect of miR-726-3p on mRNA and protein of NSF and CYB5A
To test whether miR-726-3p regulates the expression of CYB5A and NSF, we overexpressed miR-726-3p in hen granulosa cells (GCs), that significantly decreases transcript and expression levels of CYB5A and NSF (Fig. 4a, b,c). To explore whether miR-726-3p directly affects expression of CYB5A and NSF, we employed the luciferase assay. This demonstrated that miR-726-3p-mimics + CYB5A -WT was significantly lower than NC-mimics + CYB5A-WT (p < 0.01), with no significant difference observed between NC-mimics + CYB5A-MT and MiR-726-3p-mimics + CYB5A-MT. The result of NSF was consistent with that of CYB5A, indicating that CYB5A and NSF are direct targets of miR-726-3p (Fig. 4d and e).
miR-726- 3p target gene validation. OE-miR-726-3p (overexpression of miR-726-3p), GCs being transfected with overexpression miR-726-3p; BC (blank control), no overexpression miR-726-3p as a vehicle (blank control); OE-NC, scrambled overexpression miR-726-3p (negative control). (a) The CYB5A and NSF mRNA expression in the GCs was analyzed using RT-qPCR. (b, c) Expression of CYB5A and NSF proteins in the GCs with or without interference using the overexpression of miR-726-3p was analyzed by western blotting; β-actin was used as a loading control. (d) Dual luciferase validation of miR-726-3p and CYB5A targeting relationship. (e) Dual luciferase validation of miR-726-3p and NSF targeting relationship.
Effects of miR-726-3p and CYB5A on the granulosa cell (GC) proliferationand differentiation
To confirm the role of miR-726-3p in follicular development, we transfected GCs from the 6–8 mm follicles using the miR-726-3p expression vector pcDNA3.1-miR-726-3p. Overexpression of miR-726-3p significantly decreased mRNA and protein expression levels of STAR, CYP11A1, CYP19A1,PCNA, and BCL2 in GCs (OE-miR-726-3p group) (p < 0.01, Fig. 5a, b,c, d, e, f), whereas those of CASP3 were upregulated (p < 0.01, Fig. 5d, e,f), as shown by RT-qPCR and western blotting. As expected, cell viability in the GCs transfected with pcDNA3.1-miR-726-3p in 24 and 48 h decreased significantly in the CCK-8 assay (p < 0.05, p < 0.01, Fig. 6e). The GC apoptosis rate was significantly increased after transfection (P < 0.01, Fig. 6a, b,c, d). This suggests that miR-726-3p inhibits GC proliferation, but facilitates apoptosis, thereby suppressing follicular development by enhancing expression of CASP3 and decreasing PCNA and BCL2 expression levels. By attenuating expression of STAR, CYP11A1, steroidogenesis is likely suppressed.
Overexpression of miR-726-3p in the GCs. (a) The STAR, CYP11A1, CYP17A1 and CYP19A1 mRNA expression in the GCs was analyzed using RT-qPCR. (b, c) Expression of STAR, CYP11A1, CYP17A1 and CYP19A1 proteins in the GCs with or without interference using the overexpression of miR-726-3p was analyzed by western blotting. (d) The PCNA, BCL2 and CASP3 mRNA expression in the GCs was analyzed using RT-qPCR. (e, f) Expression of PCNA, BCL2 and CASP3 proteins in the GCs with or without interference using the Overexpression of miR-726-3p was analyzed by western blotting.
Overexpression of miR-726-3p in the GCs. (a) Effect of OE-miR-726-3p (overexpression of miR-726-3p) on estradiol concentration. (b) Cell viabillity of GCs at 24 h and 48 h. (c, d, e, f) OE-miR-726-3p on cell apoptosis. Total number of apoptotic cells comprises the number of cells in the right upper quadrant (Q2) and number of cells in the right lower quadrant (Q3).
We also performed loss-of-function studies by CYB5A siRNA406. Transcript and protein expression levels of CYB5A showed that the levels in the CYB5A-siRNA group were significantly lower than the control group. The mRNA and protein expression levels of STAR, CYP11A1, CYP19A1, PCNA and BCL2 in GCs of CYB5A-siRNA group were significantly lower than the control group (p < 0.05, p < 0.01, Fig. 7a, b,c, d,e, f), whereas those of CASP3 were significantly higher than the control group (p < 0.01, Fig. 7d, e,f). In addition, the EdU assay showed that the cell proliferation ratio of transfected GCs was remarkably decreased compared with the negative control (p < 0.01, Fig. 7g and h). The GC apoptosis rate was significantly increased after transfection (p < 0.01, Fig. 8a, b,c, d). Flow cytometry also showed that CYB5A siRNA promoted GCs to proceed to the G0/G1 stage (p < 0.01, Fig. 8e, f,g, h), yet inhibited them from proceeding to S and G2/M stages. This suggests that CYB5A siRNA suppresses cell differentiation (p < 0.01, Fig. 8e, f,g, i,j). Overall, our findings show that repression of GC differentiation and proliferation results from inhibition of CYB5A.
Silence of CYB5A in the GCs. CYB5A siRNA, GCs being transfected with CYB5A-specific siRNA; NC, scrambled siRNA (negative control); BC (blank control), no siRNA as a vehicle (blank control). (a) The STAR, CYP11A1, CYP17A1 and CYP19A1 mRNA expression in the GCs was analyzed using RT-qPCR. (b, c) Expression of STAR, CYP11A1, CYP17A1 and CYP19A1 proteins in the GCs with or without interference using the CYB5A siRNA was analyzed by western blotting. (d) The PCNA, BCL2 and CASP3 mRNA expression in the GCs was analyzed using RT-qPCR. (e, f) Expression of PCNA, BCL2 and CASP3 proteins in the GCs with or without interference using the CYB5A siRNA was analyzed by western blotting. (g) Effect of CYB5A siRNA on estradiol concentration. (h, i) CYB5A silence on the GC proliferation.
Silence of CYB5A in the GCs. (a, b, c, d) CYB5A silence on cell apoptosis. Total number of apoptotic cells comprises the number of cells in the right upper quadrant (Q1-UR) and number of cells in the right lower quadrant (Q1-LR). (e, f, g, h, i, j) Silencing of CYB5A on cell cycle G0/G1, S and G2/M.
Discussion
MicroRNA (miRNA) functions in a specific manner on the post-transcription level to regulate gene expression in various organisms. Being easily secreted and absorbed by tissues and organs, miRNAs can have an impact on multiple target genes and thereby regulate normal physiological function and development of diseases50. Many miRNAs have been shown in chicken and other animal gonads indicating their important roles in female reproductive physiology. In this study, 22, 9, 30 miRNAs linked genes were significantly DE miRNAs between JB1 and JB2, JB1 and JB3, and JB2 and JB3, respectively. A previous study showed seventeen significantly DE miRNAs identified in a Luhua chicken breed at 300 days of age with a high rate of egg production compared with the Luhua chicken with a low rate of egg production, including miR-34c-5p, miR-34b-5p, miR-1744-3p, which are consistent with our results51. The gga-miR-34c-5p and gga-miR-34b-5p were up-regulated significantly in chickens with high-rate of egg production compared to those with a low-rate. Furthermore, those two miRNAs were significantly down-regulated in JB3 compared with JB2, suggesting that down-regulation may decrease egg production. Previous studies indicated that the expression of miR-34a, b and c appears to be related to p53 and serves as a potential tumor suppressor for regulating processes, such as proliferation, cell cycle, apoptosis and metastasis in mice52,53. Our results indicate that miR-34c-5p, miR-34b-5p and miR-34a-5p are DE miRNAs, suggesting that gga-miR-34 may play an important role in ovulatory function in chicken follicles. gga-miR-135a was one of the 15 differentially expressed miRNAs between sexually mature and immature ovary tissues of Single Comb White Leghorn chickens, an observation that is consistent with our results when comparing JB1 and JB212.
In this study, five co-enriched pathways across 3 groups were SNARE interactions in vesicular transport (ko04130), focal adhesion (ko04510), cell cycle (ko04110), endocytosis (ko04144), and chemokine signaling pathways (ko04062). Here, SNARE refers to N-ethylmaleimide-sensitive factor attachment protein receptor. DE miRNAs were also significantly enriched in the SNARE interactions in vesicular transport in egg-type ducks compared with egg-meat-type ducks and chickens with high-rate egg production compared to those with low-rates51,54. SNARE proteins play a role in fertilization in mice55. Soluble N-ethylmaleimide-sensitive factor (NSF) attachment protein (α-SNAP) is a multifunctional protein that takes part in SNARE-mediated membrane fusion events. α-SNAP is vital to the balance between follicular development and atresia in mice56. Focal adhesion is also significantly enriched in chicken pituitary tissue with high-rate egg production51. Focal adhesions (FA) are large macromolecular assemblies, functioning as dynamic signaling hubs within the cell. They connect intracellular actin to the extracellular matrix (ECM) and respond to environmental signals57. The injection of focal adhesion kinase (FAK) inhibitor suppressed not only ovulation but also luteinization of granulosa cells and cumulus expansion in mice58. Chemotactic chemokines are a family of small cytokines. Chemokines work in immune cells by trafficking and regulating cell proliferation, migration, activation, differentiation, and homing59. Our study found that the chemokine signaling pathway is influenced by cytokine-cytokine receptor interactions between JB1 and JB2, JB1 and JB3, and JB2 and JB3. The reason for this remains to be further explored. The significantly enriched pathways, i.e. cell cycle, endocytosis, and chemokine signaling pathways, indicate that DE miRNAs are involved in cell development, cell nutrition, signal transduction and metabolic and immune processes. The pathway “Progesterone-mediated oocyte maturation” of target genes for differentially expressed miRNAs is enriched between JB2 and JB3, which is like that of the target genes for miRNAs between mature and immature ovaries in chickens and between egg-type and egg-meat-type ducks12, suggesting that DE miRNAs regulate follicle development by changing ovarian steroids.
Regulation of GC proliferation, differentiation and apoptosis is the vital step in determining follicular fate. Some miRNAs have been reported to be involved in regulating proliferation and apoptosis of ovarian GCs60,61,62. It was found that miR-10b downregulates brain-derived neurotropic factor (BDNF) expression in goat granulosa cells by targeting BDNF and inhibits granulosa cell proliferation60. Results suggest that miR-34c-5p promotes GC apoptosis by targeting Bcl2 in the broody goose61. MiR-202-5p was shown to suppress proliferation and facilitate apoptosis of goose GCs by targeting BTBD 1062. In this study, given the fact that miR-726-3p is co-DE miRNAs and consistent with another breed, i.e. Lohmann Brown (LB) chicken, a commercial egg-laying breed (data not shown), this was chosen to detect its role in regulating the development of GCs. Genetic regulation of 6–8 mm diameter follicles is generally involved in follicle selection18,63,64 and may possess a pivotal influence on hierarchy of undifferentiated prehierarchical follicles. In the GCs from the 6–8 mm follicles cultured in vitro, over-expression of miR-726-3p significantly negatively regulated GC proliferation and differentiation by reducing the mRNA and protein expression levels of PCNA, STAR, CYP11A1 and CYP19A1 as well as by CCK-8 assay. It facilitated cell apoptosis by increasing mRNA and protein expression levels of CASP3 while decreasing those of BCL2, as also shown by flow cytometry. BCL2, an apoptosis suppression protein, is an important component of the apoptotic pathway. The BCL2 family of proteins is the key regulator of apoptosis and promote a number of pathological disorders65.
By using miRNA sequencing, we discovered significantly higher levels of miR-726-3p in JB2, JB3, i.e. small yellow follicles (SYF) and large yellow follicles (LYF) than those in JB1, i.e. large white follicles (LWF), suggesting a negative effect on GC proliferation, differentiation and estradiol synthesis, as well as the possibility of promoting GC apoptosis. Given the fact that previous studies revealed that GC apoptosis may cause follicular atresia66,67, the development of new targeted drugs that modulate the levels of miR-726-3p may prevent the occurrence of follicular atresia and thereby ovulation failure.
However, by which miR-726-3p regulates GC function remains unclear. Putative miRNA targets of miR-726-3p were predicted using MIRDB, TargetScan, and MiRanda. This confirmed that miR-726-3p can inhibit NSF and CYB5A expression by targeting their 3’UTR regions. CYB5A and NSF were identified by bioinformatics, miR-726-3p overexpression, and luciferase reporter assays as the direct targets of miR-726-3p in chickens, which is negatively regulated by miR-726-3p. The predicted regulatory network of CYB5A shows the interaction of CYB5A and other proteins, such as PGRMC1,2, CYB5R2,3, CYP17A1 andCYP19A1, which engage in steroid synthesis. Likewise, the regulatory network involves the interaction of NSF and other proteins, such as NAPB, YKT6, SEC22B, VAMP3 and so on, involved in SNARE related vesicle movement.
Ovaries have dual functions in folliculogenesis and steroidogenesis in chickens. Sex steroids play an important role in the processes of follicle development and atresia and influence the quality of follicles and oocytes. In this study, the genes associated with steroidogenesis, including STAR, CYP11A1, CYP19A1 and CYB5A are necessary for the study of follicle development. Because CYB5A was confirmed to be the direct target of miR-726-3p, we used RNA interference to knock down the expression levels of CYB5A. Inhibition of CYB5A expression was shown to suppress GC proliferation, differentiation and promote GC apoptosis by the key gene analyses of transcripts and expression levels, EdU assay, flow cytometry. Specifically, this is consistent with results of miR-726-3p overexpression as discussed above. Prior to our study, little was known about the involvement of CYB5A in follicular development in chickens. Therefore, in this paper we present initial results of the first analysis of the role of CYB5A in GC proliferation and differentiation. CYB5A was previously identified as a regulator of androstenedione production.in pigs; CYB5A and CYB5B can interact with porcine CYP17A1 to increase production of DHEA68. It was highly expressed in goat oocytes and recognized as one of the marker genes indicating goat fertility and ovarian reserve ability69. There was a significant upregulation of CYB5A as human follicles transition from antral stage to the pre-ovulatory stage, thus contributing to enhanced progesterone production. Indeed, when GCs begin to express HSD3B2, progesterone production increased, which was supported by upregulation of CYB5A for ovarian steroidogenesis23. Thus, the result obtained in the knockdown of CYB5A extended and supported observations of miR-726-3p over-expression, showing the same trend and implying the mechanism of changes in GC proliferation, apoptosis, differentiation and estradiol synthesis induced by miR-726-3p.
Discoveries of the past years have showed that lncRNAs could act as endogenous miRNA sponges, competing with the transcript for the miRNAs, which may reduce the negative regulatory effect of miRNAs on its target genes70. The future directions for this research are finding the corresponding lncRNA which could be the miR-726-3p target and confirming the co-expression networks of NSF or CYB5A and the co-regulated lncRNAs, which could follow the ceRNA hypothesis involved in the regulation of fecundity71.
In conclusion, our results suggest that miR-726-3p negatively regulates GC proliferation, apoptosis and differentiation, and positively regulates cell apoptosis by targeting the 3’ untranslated regions of CYB5A. This, in turn, indicates the potential mechanism of action of miR-726-3p underlying follicle development in chickens, providing an effective strategy to improve egg production. This finding provides a theoretical reference for understanding chicken egg-laying traits, as well as improving marker-assisted breeding and chicken egg production.
Data availability
Our raw RNA sequencing data is available at BioProject under accession numbers PRJNA670553.
References
Onagbesan, O., Bruggeman, V. & Decuypere, E. Intra-ovarian growth factors regulating ovarian function in avian species: a review. Anim. Reprod. Sci. 111, 121–140. https://doi.org/10.1016/j.anireprosci.2008.09.017 (2009).
Zhao, J. et al. Untargeted metabolomics revealed potential biomarkers of small yellow follicles of chickens during sexual maturation. Metabolites 13, 176. https://doi.org/10.3390/metabo13020176 (2023).
Rong, Y. et al. MiR-181a-5p inhibits Goose granulosa cell viability by targeting SIRT1. Br. Poult. Sci. 62, 373–378. https://doi.org/10.1080/00071668.2020.1870660 (2021).
Manabe, N. et al. Regulation mechanism of selective Atresia in Porcine follicles: regulation of granulosa cell apoptosis during Atresia. J. Reprod. Dev. 50, 493–514. https://doi.org/10.1262/jrd.50.493 (2004).
Gioia, L. et al. Abundances of autophagy-related protein LC3B in granulosa cells, cumulus cells, and oocytes during Atresia of pig antral follicles. Anim. Reprod. Sci. 211, 106225. https://doi.org/10.1016/j.anireprosci.2019.106225 (2019).
Gebert, L. F. R. & MacRae, I. J. Regulation of MicroRNA function in animals. Nat. Rev. Mol. Cell. Biol. 20, 21–37. https://doi.org/10.1038/s41580-018-0045-7 (2019).
Salim, U., Kumar, A., Kulshreshtha, R. & Vivekanandan, P. Biogenesis, characterization, and functions of Mirtrons. Wiley Interdiscip Rev. RNA. 13, e1680. https://doi.org/10.1002/wrna.1680 (2022).
Toury, L. et al. miR-140-5p and miR-140-3p: key actors in Aging-Related diseases? Int. J. Mol. Sci. 23, 11439. https://doi.org/10.3390/ijms231911439 (2022).
Zhang, G. et al. miR-122-5p regulates proliferation and apoptosis of chicken granulosa cells of hierarchal follicles by targeting MAPK3. Gene 824, 146397. https://doi.org/10.1016/j.gene.2022.146397 (2022).
Wei, Q. et al. miR-23b-3p inhibits chicken granulosa cell proliferation and steroid hormone synthesis via targeting GDF9. Theriogenology 177, 84–93. https://doi.org/10.1016/j.theriogenology.2021.10.011 (2022).
Zhou, Y. et al. Transcriptome Analysis of the Chicken Follicular Theca Cells with miR-135a-5p Suppressed. G3 (Bethesda, Md.). 10, 4071–4081 (2020). https://doi.org/10.1534/g3.120.401701
Kang, L., Cui, X., Zhang, Y., Yang, C. & Jiang, Y. Identification of MiRNAs associated with sexual maturity in chicken ovary by illumina small RNA deep sequencing. BMC Genom. 14, 352. https://doi.org/10.1186/1471-2164-14-352 (2013).
Ru, M. et al. Chicken ovarian follicular atresia: interaction network at organic, cellular, and molecular levels. Poult. Sci. 103, 103893. https://doi.org/10.1016/j.psj.2024.103893 (2024).
Gong, Z., Yang, J., Bai, S. & Wei, S. MicroRNAs regulate granulosa cells apoptosis and follicular development - A review. Asian-Australas J. Anim. Sci. 33, 1714–1724. https://doi.org/10.5713/ajas.19.0707 (2020).
Zhang, J., Xu, Y., Liu, H. & Pan, Z. MicroRNAs in ovarian follicular Atresia and granulosa cell apoptosis. Reprod. Biol. Endocrinol. 17, 9. https://doi.org/10.1186/s12958-018-0450-y (2019).
Kloosterman, W. P. et al. Cloning and expression of new MicroRNAs from zebrafish. Nucleic Acids Res. 34, 2558–2569. https://doi.org/10.1093/nar/gkl278 (2006).
Padmanabhan, V., Karsch, F. J. & Lee, J. S. Hypothalamic, pituitary and gonadal regulation of FSH. Reprod. Suppl. 59, 67–82. https://doi.org/10.1530/rep.0.1230067 (2002).
Johnson, A. L. & Woods, D. C. Dynamics of avian ovarian follicle development: cellular mechanisms of granulosa cell differentiation. Gen. Comp. Endocrinol. 163, 12–17. https://doi.org/10.1016/j.ygcen.2008.11.012 (2009).
Johnson, A. L., Solovieva, E. V. & Bridgham, J. T. Relationship between steroidogenic acute regulatory protein expression and progesterone production in Hen granulosa cells during follicle development. Biol. Reprod. 67, 1313–1320. https://doi.org/10.1095/biolreprod67.4.1313 (2002).
Sechman, A., Pawlowska, K. & Hrabia, A. Effect of 3,3’,5-triiodothyronine and 3,5-diiodothyronine on progesterone production, cAMP synthesis, and mRNA expression of STAR, CYP11A1, and HSD3B genes in granulosa layer of chicken preovulatory follicles. Domest. Anim. Endocrinol. 41, 137–149. https://doi.org/10.1016/j.domaniend.2011.05.007 (2011).
Huang, J. et al. The association between splenocyte apoptosis and alterations of Bax, Bcl-2 and caspase-3 mRNA expression, and oxidative stress induced by dietary nickel chloride in broilers. Int. J. Environ. Res. Public. Health. 10, 7310–7326. https://doi.org/10.3390/ijerph10127310 (2013).
Heinosalo, T., Saarinen, N. & Poutanen, M. Role of hydroxysteroid (17beta) dehydrogenase type 1 in reproductive tissues and hormone-dependent diseases. Mol. Cell. Endocrinol. 489, 9–31. https://doi.org/10.1016/j.mce.2018.08.004 (2019).
Zheng, M. et al. Expression of genes and enzymes involved in ovarian steroidogenesis in relation to human follicular development. Front. Endocrinol. (Lausanne). 14, 1268248. https://doi.org/10.3389/fendo.2023.1268248 (2023).
Sun, X. et al. Transcriptome comparative analysis of ovarian follicles reveals the key genes and signaling pathways implicated in Hen egg production. BMC Genom. 22, 899. https://doi.org/10.1186/s12864-021-08213-w (2021).
Squires, E. J., Gray, M. A. & Lou, Y. Effect of mutations in Porcine CYB5A and CYP17A1 on the metabolism of pregnenolone. J. Steroid Biochem. Mol. Biol. 195, 105469. https://doi.org/10.1016/j.jsbmb.2019.105469 (2019).
Nakamura, Y. et al. 3βHSD and CYB5A double positive adrenocortical cells during adrenal development/aging. Endocr. Res. 40, 8–13. https://doi.org/10.3109/07435800.2014.895377 (2015).
Grindflek, E., Berget, I., Moe, M., Oeth, P. & Lien, S. Transcript profiling of candidate genes in testis of pigs exhibiting large differences in Androstenone levels. BMC Genet. 11, 4. https://doi.org/10.1186/1471-2156-11-4 (2010).
Flück, C. E., Miller, W. L. & Auchus, R. J. The 17, 20-lyase activity of cytochrome p450c17 from human fetal testis favors the delta5 steroidogenic pathway. J. Clin. Endocrinol. Metab. 88, 3762–3766. https://doi.org/10.1210/jc.2003-030143 (2003).
Akhtar, M. K., Kelly, S. L. & Kaderbhai, M. A. Cytochrome b(5) modulation of 17{alpha} hydroxylase and 17–20 lyase (CYP17) activities in steroidogenesis. J. Endocrinol. 187, 267–274. https://doi.org/10.1677/joe.1.06375 (2005).
Gui, H. et al. miR-27a-3p targets NR5A2 to regulate CYP19A1 expression and 17-β estradiol synthesis in ovine granulosa cells. Anim. Reprod. Sci. 248, 107160. https://doi.org/10.1016/j.anireprosci.2022.107160 (2023).
Chen, X. et al. Transcriptome analysis of ovarian follicles reveals potential pivotal genes associated with increased and decreased rates of chicken egg production. Front. Genet. 2, 622751. https://doi.org/10.3389/fgene.2021.622751 (2021).
Kollias, N. S., Hess, W. J., Johnson, C. L., Murphy, M. & Golab, G. A literature review on current practices, knowledge, and viewpoints on pentobarbital euthanasia performed by veterinarians and animal remains disposal in the united States. J. Am. Vet. Med. Assoc. 261, 733–738. https://doi.org/10.2460/javma.22.08.0373 (2023).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915. https://doi.org/10.1038/s41587-019-0201-4 (2019).
Ambros, V. et al. A uniform system for MicroRNA annotation. RNA (New York N Y). 9, 277–279. https://doi.org/10.1261/rna.2183803 (2003).
Griffiths-Jones, S. The MicroRNA registry. Nucleic Acids Res. 32, D109–D111. https://doi.org/10.1093/nar/gkh023 (2004).
Griffiths-Jones, S., Grocock, R. J., van Dongen, S., Bateman, A. & Enright, A. J. MiRBase: MicroRNA sequences, targets and gene nomenclature. Nucleic Acids Res. 34, D140–D144. https://doi.org/10.1093/nar/gkj112 (2006).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25. https://doi.org/10.1186/gb-2009-10-3-r25 (2009).
Mackowiak, S. D. Identification of novel and known MiRNAs in deep-sequencing data with miRDeep2. Curr Protoc. Bioinformatics Chapter. 12, 12.10.1–12.10.15 (2011).
Zhou, L. et al. Integrated profiling of MicroRNAs and mRNAs: MicroRNAs located on Xq27.3 associate with clear cell renal cell carcinoma. Plos One. 5, e15224. https://doi.org/10.1371/journal.pone.0015224 (2010).
Wong, N. & Wang, X. MiRDB: an online resource for MicroRNA target prediction and functional annotations. Nucleic Acids Res. 43, D146–D152. https://doi.org/10.1093/nar/gku1104 (2015).
Enright, A. et al. MicroRNA targets in drosophila. Genome Biol. 5, R1. https://doi.org/10.1186/gb-2003-5-1-r1 (2003).
Conesa, A. et al. Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21, 3674–3676. https://doi.org/10.1093/bioinformatics/bti610 (2005).
Xu, R. et al. Inhibitory effect of SLIT2 on granulosa cell proliferation mediated by the CDC42-PAKs-ERK1/2 MAPK pathway in the prehierarchical follicles of the chicken ovary. Sci. Rep. 8, 9168. https://doi.org/10.1038/s41598-018-27601-z (2018).
Zhu, H. et al. Synergistic Inhibition of csal1 and csal3 in granulosa cell proliferation and steroidogenesis of Hen ovarian prehierarchical development. Biol. Reprod. 101, 986–1000. https://doi.org/10.1093/biolre/ioz137 (2019).
Qin, N. et al. Cooperative effects of FOXL2 with the members of TGF-β superfamily on FSH receptor mRNA expression and granulosa cell proliferation from Hen prehierarchical follicles. PloS One. 10, e0141062. https://doi.org/10.1371/journal.pone.0141062 (2015).
Lyu, Z. et al. The Hippo/MST pathway member SAV1 plays a suppressive role in development of the prehierarchical follicles in Hen ovary. PloS One. 11, e0160896. https://doi.org/10.1371/journal.pone.0160896 (2016).
Rao, X., Huang, X., Zhou, Z. & Lin, X. An improvement of the 2ˆ(-delta delta CT) method for quantitative real-time polymerase chain reaction data analysis. Biostat Bioinforma Biomath. 3, 71–85. https://doi.org/10.1016/S0920-5489(99)92176-1 (2013).
Kanehisa, M., Sato, Y., Kawashima, M., Furumichi, M. & Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 44, D457–D462. https://doi.org/10.1093/nar/gkv1070 (2016).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30. https://doi.org/10.1093/nar/28.1.27 (2000).
Mishra, S., Yadav, T. & Rani, V. Exploring MiRNA based approaches in cancer diagnostics and therapeutics. Crit. Rev. Oncol/Hematol. 98, 12–23. https://doi.org/10.1016/j.critrevonc.2015.10.003 (2016).
Wu, N. et al. Expressed MicroRNA associated with high rate of egg production in chicken ovarian follicles. Anim. Genet. 48, 205–216. https://doi.org/10.1111/age.12516 (2017).
Hermeking, H. The miR-34 family in cancer and apoptosis. Cell. Death Differ. 17, 193–199. https://doi.org/10.1038/cdd.2009.56 (2010).
Ji, Q. et al. MicroRNA miR-34 inhibits human pancreatic cancer tumor-initiating cells. PloS One. 4, e6816. https://doi.org/10.1371/journal.pone.0006816 (2009).
Qiu, M. et al. High-throughput sequencing analysis identified MicroRNAs associated with egg production in ducks ovaries. PeerJ 8, e8440. https://doi.org/10.7717/peerj.8440 (2020).
Choi, S. S., Jung, J. Y., Lee, D. H., Kang, J. Y. & Lee, S. H. Expression and regulation of SNAP-25 and synaptotagmin VII in developing mouse ovarian follicles via the FSH receptor. J. Mol. Histol. 44, 47–54. https://doi.org/10.1007/s10735-012-9434-y (2013).
Arcos, A. et al. Publisher Correction: α-SNAP is expressed in mouse ovarian granulosa cells and plays a key role in folliculogenesis and female fertility. Sci. Rep. 10, 5691. https://doi.org/10.1038/s41598-020-61863-w (2020).
Murphy, K. N. & Brinkworth, A. J. Manipulation of focal adhesion signaling by pathogenic microbes. Int. J. Mol. Sci. 22, 1358. https://doi.org/10.3390/ijms22031358 (2021).
Kitasaka, H. et al. Inductions of granulosa cell luteinization and cumulus expansion are dependent on the fibronectin-integrin pathway during ovulation process in mice. PloS One. 13, e0192458. https://doi.org/10.1371/journal.pone.0192458 (2018).
Zhou, W., Guo, S., Liu, M., Burow, M. E. & Wang, G. Targeting CXCL12/CXCR4 axis in tumor immunotherapy. Curr. Med. Chem. 26, 3026–3041. https://doi.org/10.2174/0929867324666170830111531 (2019).
Peng, J. Y. et al. MicroRNA-10b suppresses goat granulosa cell proliferation by targeting brain-derived neurotropic factor. Domest. Anim. Endocrinol. 54, 60–67. https://doi.org/10.1016/j.domaniend.2015.09.005 (2016).
Hou, L. et al. MiR-34c-5p promotes granulosa cells apoptosis by targeting Bcl2 in broody Goose (Anser cygnoides). Anim. Biotechnol. 33, 1280–1288. https://doi.org/10.1080/10495398.2021.1886943 (2022).
Ran, M. et al. MiR-202-5p regulates geese follicular selection by targeting BTBD10 to regulate granulosa cell proliferation and apoptosis. Int. J. Mol. Sci. 24, 6792. https://doi.org/10.3390/ijms24076792 (2023).
Johnson, A. L. Ovarian follicle selection and granulosa cell differentiation. Poult. Sci. 94, 781–785. https://doi.org/10.3382/ps/peu008 (2015).
Son, W. Y., Das, M., Shalom-Paz, E. & Holzer, H. Mechanisms of follicle selection and development. Minerva Ginecol. 63, 89–102, http://dx.doi.org/ (2011).
Siddiqui, W. A., Ahad, A. & Ahsan, H. The mystery of BCL2 family: Bcl-2 proteins and apoptosis: an update. Arch. Toxicol. 89, 289–317. https://doi.org/10.1007/s00204-014-1448-7 (2015).
Hillier, S. G. Gonadotropic control of ovarian follicular growth and development. Mol. Cell. Endocrinol. 179, 39–46. https://doi.org/10.1016/s0303-7207(01)00469-5 (2001).
Matsuda, F., Inoue, N., Manabe, N. & Ohkura, S. Follicular growth and Atresia in mammalian ovaries: regulation by survival and death of granulosa cells. J. Reprod. Dev. 58, 44–50. https://doi.org/10.1262/jrd.2011-012 (2012).
Billen, M. J. & Squires, E. J. The role of Porcine cytochrome b5A and cytochrome b5B in the regulation of cytochrome P45017A1 activities. J. Steroid Biochem. Mol. Biol. 113, 98–104. https://doi.org/10.1016/j.jsbmb.2008.11.012 (2009).
Li, S. et al. Transcriptome profile of goat folliculogenesis reveals the interaction of oocyte and granulosa cell in correlation with different fertility population. Sci. Rep. 11, 15698. https://doi.org/10.1038/s41598-021-95215-z (2021).
Hu, H. et al. Long non-coding RNA TCONS_00814106 regulates Porcine granulosa cell proliferation and apoptosis by sponging miR-1343. Mol. Cell. Endocrinol. 520, 111064. https://doi.org/10.1016/j.mce.2020.111064 (2021).
Miao, X., Luo, Q., Zhao, H. & Qin, X. Ovarian transcriptomic study reveals the differential regulation of MiRNAs and LncRNAs related to fecundity in different sheep. Sci. Rep. 6, 35299. https://doi.org/10.1038/srep35299 (2016).
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (Grant No. 32172714 and 31902145); the China Agriculture Research System of MOF and MARA (Grant No. CARS-41) and the Project of Science and Technology Development Plan of Jilin Province (Grant No. 20220101306JC).
Author information
Authors and Affiliations
Contributions
Rifu Xu was responsible for the project design, sample collection and the data analysis. Yuhan Sun, Chunchi Yan, Chang Ma coordinated the sample collection, Xue Sun coordinated the experimental design. Jinghua Zhao, Ning Qin were responsible for the experimental design, data analysis and manuscript preparation. All authors approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhao, J., Sun, Y., Sun, X. et al. Micro RNA miR-726-3p targets CYB5A in Hen ovaries to modulate granulosa cell proliferation and differentiation. Sci Rep 15, 45717 (2025). https://doi.org/10.1038/s41598-025-28318-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-28318-6