Abstract
Crohn’s disease (CD) is a chronic, relapsing inflammatory bowel disease (IBD). Recent studies have revealed that intestinal tissues of CD patients exhibit progeroid-like features, suggesting that cellular senescence may contribute to CD progression. However, the specific roles of senescence-related genes in CD and their impact on prognosis remain unclear. This study identified CD-associated senescence genes through differential expression analysis. Machine learning algorithms were employed to further screen for hub genes. Gene Set Enrichment Analysis (GSEA) was performed to explore the potential functions of these hub genes, and their association with immune cell infiltration was assessed using the CIBERSORT algorithm. Independent external datasets were utilized to validate the diagnostic model’s robustness, with additional validation of hub gene expression through mouse models and clinical samples. Furthermore, single-cell RNA sequencing (scRNA-seq) data analysis was conducted to examine senescence heterogeneity at the single-cell level in CD patients. This study identified five senescence-associated hub genes (STAT1, S100A11, FILIP1L, F3, and HPS5) significantly correlated with CD. GSEA revealed that these genes were closely associated with tissue inflammatory responses, nutrient absorption, and energy metabolism. Immune infiltration analysis further demonstrated a strong correlation between the expression levels of these five genes and the degree of immune cell infiltration. scRNA-seq analysis indicated significant differences in senescence levels between affected and unaffected areas in CD patients. Senescence-related genes are closely associated with disease progression in CD patients.
Similar content being viewed by others
Introduction
Crohn’s disease (CD) is a chronic relapsing inflammatory bowel disease (IBD) that can affect any segment of the gastrointestinal tract from the mouth to the anus, with a predilection for the terminal ileum and colon1. The global incidence of CD continues to rise, with the highest rates observed in North America and Europe, while Asian countries have also shown a significant increasing trend in recent years2. The pathogenesis of CD is complex, involving interactions among genetic susceptibility, gut microbiota dysbiosis, environmental factors, and aberrant immune system activation3. Its typical clinical manifestations include abdominal pain, chronic diarrhea, weight loss, and fatigue. The inflammation is characterized by transmural and segmental distribution, suggesting histopathological heterogeneity across different intestinal segments in CD patients4,5. Although the use of biologics has improved outcomes for some patients, a considerable proportion still face treatment failure or postoperative recurrence, highlighting the urgent need to explore novel pathogenic mechanisms and therapeutic targets6.
Cellular senescence is a state of cell cycle arrest induced by factors such as DNA damage, telomere shortening, or oxidative stress, often accompanied by the senescence-associated secretory phenotype (SASP)7. This phenotype exacerbates local inflammation and tissue damage through the secretion of pro-inflammatory factors and matrix metalloproteinases (MMPs)8,9. Studies have shown that chronic inflammation accelerates cellular senescence, while senescent cells, in turn, amplify inflammatory responses via SASP, creating a self-reinforcing vicious cycle10. This self-perpetuating interaction between chronic inflammation and cellular senescence is likely to drive disease progression in CD. Recent research has observed abnormal expression of senescence markers in the intestinal stem cell niche of CD patients, suggesting that cellular senescence may be a potential mechanism underlying CD pathology11.
Given that CD predominantly affects young adults and cellular senescence has traditionally been viewed as an aging-related process, its role in CD pathogenesis may have long been underestimated. Although abnormal expression of senescence markers has been detected in the intestines of CD patients, systematic investigations into how cellular senescence influences CD progression remain insufficient. Elucidating the impact of cellular senescence on CD progression will not only deepen our understanding of the disease but may also provide a theoretical basis for developing novel anti-senescence therapeutic strategies to overcome current treatment limitations.
This study systematically analyzed the expression patterns and heterogeneous distribution of cellular senescence-related genes in intestinal tissues of CD patients by integrating RNA-seq and scRNA-seq data. A diagnostic prediction model based on CD-associated senescence genes was successfully constructed using machine learning algorithms, and its predictive efficacy was validated in multiple independent external datasets. Mouse models and clinical samples further confirmed the differential expression of key genes from the prediction model in CD, providing direct evidence for the involvement of cellular senescence in CD progression. Notably, at single-cell resolution, we observed significantly higher senescence scores in fibroblasts and epithelial cells from lesional areas compared to non-lesional areas in CD patients, suggesting that cellular senescence may promote CD progression through intestinal fibrosis and disruption of the epithelial barrier function.
Aiming to leverage existing sequencing data and bioinformatic approaches, this study seeks to elucidate the role of senescence-related genes in CD at both bulk transcriptome and single-cell transcriptome levels, with the goal of identifying new potential therapeutic targets for CD treatment.
Materials and methods
Data acquisition
The datasets used in this study were obtained from the Gene Expression Omnibus (GEO) database (https://www.ncbi.nlm.nih.gov/geo/). We analyzed four datasets, including three bulk RNA-seq datasets (GSE75214, GSE36807, and GSE102133) and one single-cell RNA-seq (scRNA-seq) dataset (GSE134809). The GSE75214 dataset was utilized to identify senescence related differentially expressed genes (SR-DEGs) in CD and to construct a diagnostic model. The GSE36807 and GSE102133 datasets served as independent validation cohorts to assess the robustness of the diagnostic model. The scRNA-seq dataset (GSE134809) was employed to investigate senescence-related gene expression signatures and cellular heterogeneity in CD at single-cell resolution. Additionally, 77 senescence-related genes were obtained from the MsigDB database (C2 collection, https://www.gsea-msigdb.org/gsea/msigdb) for subsequent analyses. Detailed sample information for each dataset is provided in Additional file: Table S1.
Data process and differential expression analysis
It was performed principal component analysis (PCA) using the R package “FactoMineR” to evaluate sample clustering patterns in the GSE75214 dataset. Subsequently, differential expression analysis was conducted using the “limma” R package. The obtained differentially expressed genes (DEGs) were intersected with a predefined senescence-associated gene set, ultimately identifying CD SR-DEGs. Visualization of the results was achieved using the following R packages: “ggplot2”, “heatmap”, “ggvolcano”, and “ggVennDiagram”.
Diagnostic model construction
Feature selection was employed a multi-dimensional machine learning approach incorporating Least Absolute Shrinkage and Selection Operator (LASSO) regression, Random Forest (RF), and Support Vector Machine (SVM). LASSO regression analysis was performed using the R package “glmnet” with 10-fold cross-validation to determine the optimal λ value for feature dimension reduction. Additionally, the Random Forest (RF) algorithm from the R package “randomForest” was utilized to calculate gene importance scores, while variable selection was implemented via Support Vector Machine (SVM) using the R package “e1071”. The R package “UpSetR” was applied to visualize overlapping genes identified by these three distinct machine learning methods. Finally, Receiver Operating Characteristic (ROC) curves were generated with the R package “pROC” in both the training cohort (GSE75214) and independent validation cohorts (GSE102133 and GSE36807) to systematically evaluate the predictive performance of candidate biomarkers.
Functional enrichment and PPI network analysis
Functional annotation and pathway enrichment analysis were performed using the R package “clusterProfiler” based on the Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) databases12. The STRING database was employed to predict protein-protein interaction (PPI) networks of SR-DEGs, with subsequent visualization of key pathway-gene associations conducted using Cytoscape software. Pearson correlation analysis was applied to calculate correlations between core genes in the diagnostic model and other genes. Single-gene Gene Set Enrichment Analysis (GSEA) was further performed based on correlation results to explore potential functional roles of the core genes.
Immune infiltration analysis
The CIBERSORT algorithm was employed to perform deconvolution analysis of gene expression data for quantifying the relative abundances of 22 immune cell subsets in tissue samples13. Based on the algorithm results, comparative analysis of immune cell infiltration profiles between the CD group and healthy controls was conducted to evaluate differences in immune microenvironment. Pearson correlation analysis was performed to assess associations between hub gene expression levels in the diagnostic model and infiltration degrees of various immune cell subsets14.
Single-cell transcriptome analysis
Analysis of the GSE134809 single-cell dataset was performed using the R package “Seurat”. Stringent data preprocessing and quality control were first implemented, retaining cells expressing 500–5000 genes while rigorously excluding low-quality cells with > 35% mitochondrial gene content. The SCTransform algorithm was then applied for data normalization and variance stabilization, with potential batch effects corrected via the IntegrateLayers function. Nonlinear dimensionality reduction was achieved through RunUMAP, followed by cell type identification using FindNeighbors (dims = 1:15) and FindClusters (resolution = 0.1). Gene set activity scores were calculated with AddModuleScore, and pathway enrichment analysis was conducted against the Hallmark gene sets using the GSVA package. Finally, intergroup differential gene expression was assessed by Wilcoxon rank-sum test.
Animal experiments and clinical samples
CD-like murine model was established by administering 3% dextran sulfate sodium (DSS) in drinking water to 6-week-old C57BL/6 mice for 7 consecutive days. At the end of the experiment, the mice were euthanized by cervical dislocation. No anesthetics were used at any point during the experiment. Animal care and use throughout the experiments were approved by the Institutional Animal Care and Use Committee of the Center for Medical Genetics at Central South University (Animal 2025-2-50). All animal experiments were conducted in accordance with the relevant designated guidelines and regulations and in compliance with the ARRIVE Guidelines. The collection of human samples was approved by the local Ethics Committee and the Review Board of Xiangya Hospital of Central South University (Changsha, China, human 2025-1-51). All study participants provided their informed consent, the study protocol was conducted in accordance with the Declaration of Helsinki. CD was diagnosed through the combined analysis of clinical manifestations and pathological, endoscopic and imaging examinations by experienced clinical doctors according to published guidelines. All volunteers provided informed consent for the collection of colon. Total RNA was extracted from colonic tissues using TRIzol reagent, with cDNA synthesized via PrimeScript RT reagent kit. qPCR was performed with SYBR Green Master Mix. The primer information is provided in Additional file: Table S2.
Statistical analysis
All statistical analyses in this study were performed using the R (Version 4.3.0).
Results
Identification of SR-DEGs in CD
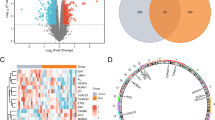
To identify genes significantly associated with cellular senescence in CD, we performed differential expression analysis on the GSE75214 dataset. Prior to differential expression analysis, PCA was conducted to identify samples with clear grouping patterns, ensuring the validity of subsequent analyses. Since inactive CD samples could not be clearly distinguished from the other two groups, comparisons involving the inactive group were temporarily excluded from this differential analysis. Ultimately, 59 samples from active CD patients (8 colon tissues and 51 ileal tissues) and 22 healthy control samples were included for differential expression analysis (Fig. 1A).
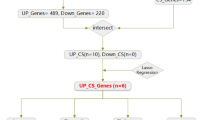
The differential expression analysis identified a total of 1,237 DEGs (P.Value < 0.05, |logFC| > 0.58, 1.5-fold expression changes), comprising 828 upregulated and 409 downregulated genes (Fig. 1B). Functional enrichment analysis of these DEGs revealed significant associations with cytokine-cytokine receptor interactions, leukocyte migration, collagen-containing extracellular matrix, and inflammatory bowel disease pathways (Fig. S1), suggesting abnormal immune system activation and intestinal fibrosis due to ECM deposition, consistent with previous reports3.
By intersecting the DEGs with senescence-associated gene sets, we identified 20 CD SR-DEGs, including 18 upregulated and 2 downregulated genes (Fig. 1C, D). Pearson correlation analysis demonstrated significant co-expression relationships among these SR-DEGs (Fig. 1E).
Functional characterization of SR-DEGs
To elucidate the functional characteristics of SR-DEGs, we conducted GO and KEGG enrichment analyses along with PPI network construction. The GO analysis revealed: biological processes (BP) were significantly enriched in the regulation of epithelial cell proliferation/differentiation and positive regulation of cytokine production; molecular functions (MF) were primarily associated with extracellular matrix structural constituents; and cellular components (CC) showed strong correlation with collagen-containing extracellular matrix (Fig. 1F). KEGG pathway analysis demonstrated that the AGE-RAGE signaling pathway in diabetic complications exhibited the most significant alterations (Fig. 1G, H). Notably, several immune-inflammatory pathways were markedly enriched, including IBD, IL-17 signaling pathway, TGF-β signaling pathway, HIF-1 signaling pathway, and Th17 cell differentiation. The protein-pathway interaction network constructed using Cytoscape software unveiled intricate regulatory relationships between SR-DEGs and associated pathways (Fig. 1I). These findings suggest that SR-DEGs may be involved in: Senescence-inflammation positive feedback loop, impaired epithelial cell repair, fibrogenic drive and immunometabolic reprogramming.
Identification of hub genes
To identify core genes associated with CD, we employed three machine learning approaches for feature selection from SR-DEGs. The LASSO regression analysis identified 5 candidate genes (Fig. 1J, K), while SVM analysis demonstrated optimal diagnostic performance when retaining all 20 variables (Fig. 1L). Random Forest (RF) analysis selected 11 genes exceeding the mean precision threshold (Fig. 1M). Through integration of these three analytical methods, we ultimately identified five core genes: STAT1, S100A11, FILIP1L, F3, and HPS5 (Fig. 1N).
These genes exhibited consistently upregulated expression in active CD samples (Fig. 1D) and showed positive intercorrelations (Fig. 1O, P). ROC curve analysis confirmed that these core genes, either individually or as a combined diagnostic model, possess excellent potential as diagnostic biomarkers for CD (Fig. 1Q, R).
Construction of a senescence-related signature for CD. (A) PCA plot showing sample clustering in GSE75214 dataset. (B) Volcano plot showing DEGs between the active CD group and the normal control group in the GSE75214 dataset. (C) Venn diagram illustrating the overlap between DEGs and senescence-related genes, identifying 20 CD SR-DEGs. (D) Expression heatmap of the 20 SR-DEGs between active CD and control groups. (E) Correlation heatmap of SR-DEGs. (F) GO and (G) KEGG analysis of SR-DEGs. (H) Network diagram of the top five KEGG pathways. (I) Protein-pathway interaction network constructed by Cytoscape. (J) Coefficient distribution profile of variables in the LASSO regression model. (K) Trajectory plot of parameter coefficients in the LASSO regression analysis. (L) Constructing variables for optimal model selection based on SVM. (M) Selection of optimal variables above the average precision threshold based on RF. (N) Three machine learning methods select the common intersection of variables. (O, P) Heat map and circle chart of the correlations of five key genes. (Q) The AUC area under the ROC curve constructed by single hub gene. (R) The AUC area under the ROC curve of the overall model construction. The closer the AUC is to 1, the better the effect. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
GSEA of hub genes
Through GSEA of hub gene correlations with other genes (Fig. S2), we systematically characterized their functional profiles. The results demonstrated that all hub genes showed significant negative correlations with oxidative phosphorylation (OXPHOS) pathways but positive correlations with viral protein-cytokine receptor interaction pathways (Figs. 2A-E, S3). Further analysis revealed that most hub genes (≥ 3) exhibited significant negative associations with both the tricarboxylic acid (TCA) cycle and multiple nutrient absorption/metabolic pathways, while displaying positive correlations with hematopoietic cell lineage, osteoclast differentiation, rheumatoid arthritis pathogenesis, and various disease-related pathways (Fig. S3). These findings suggest that mitochondrial dysfunction may contribute to CD pathogenesis, wherein hub genes potentially suppress OXPHOS activity, induce cellular senescence, and promote the secretion of SASP factors, consequently recruiting immune cell infiltration.
The correlation between hub gene expression and immunological characteristics
To investigate the differences in immune cell infiltration between CD patients and healthy controls, we performed immune infiltration analysis. Comparative analysis demonstrated significant alterations in 12 immune cell subtypes in CD patients compared to healthy controls. Specifically, CD patients exhibited decreased infiltration levels of 7 immune cell populations, such as regulatory T cells (Tregs), CD8 + T cells, and M2 macrophages, while showing significantly increased infiltration of M1 macrophages, CD4 + T cells, and neutrophils etc. (Fig. 2F). These immune cell infiltration patterns displayed significant correlative variations among samples (Fig. 2G).
To further explore the relationship between hub genes and immune infiltration in CD, we performed correlation analysis. The results revealed that the majority of hub genes (≥ 4) showed significant positive correlations with infiltration levels of activated dendritic cells (DCs), M0/M1 macrophages, neutrophils, resting natural killer (NK) cells, and memory CD4 + T cells, while exhibiting negative correlations with resting mast cells, monocytes, activated NK cells, CD8 + T cells, and Tregs (Fig. 2H). Notably, the correlation between hub genes and immune infiltration highly coincided with the altered immune infiltration profiles in CD patients, suggesting that these hub genes may drive intestinal immune dysregulation, thereby promoting disease progression in CD.
Hub gene GSEA enrichment analysis and immune infiltration analysis. (A-E) GSEA was performed based on correlation results between STAT1, F3, S100A11, HPS5, FILIP1L and other genes respectively, the top 10 most significantly positively and negatively correlated pathways were visualized using ridge plots. (F) The differences in immune infiltration between Active CD and normal intestinal segments. (G) Heatmap of immune cell correlation in CD patients. (H) The correlation between key genes and various immune cells in CD patients. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
Diagnostic model performance validation and disease model construction
To validate the robustness of our constructed model, we performed external validation using independent dataset. PCA and cluster analysis of the GSE102133 dataset identified 32 CD samples (including both newly diagnosed and long-standing cases) along with 12 normal controls for subsequent analyses (Fig. S4). ROC curve analysis demonstrated excellent predictive performance for all five hub genes individually, with the integrated model exhibiting outstanding diagnostic efficacy (AUC = 0.995) (Fig. 3A, B). Differential expression analysis revealed significantly elevated expression levels of all hub genes in the CD group compared to normal controls (Fig. 3C). This predictive capacity was consistently replicated in the independent GSE36807 dataset, where the model maintained equally robust performance (Fig. 3D, E), while hub genes also showed upregulated expression patterns in CD (Fig. 3F).
To validate the expression of hub genes in vivo, we established a mouse model of CD, dextran sulfate sodium (DSS) was administered to induce colitis (Fig. 3G). Comparative analysis revealed that DSS-treated mice exhibited significant body weight loss starting from day 3, with a pronounced reduction compared to the control group by day 7 (Fig. 3H). Additionally, the DSS-treated group displayed marked shortening of the colon length relative to controls (Fig. 3I).
Furthermore, we performed a comparative analysis of hub gene expression between CD and normal control groups in both clinical samples and the mouse model. In the DSS-induced mouse model, the expression of F3 and S100a11 was significantly upregulated in the CD group, while the remaining genes showed an increasing trend without reaching statistical significance (Fig. 3J-N). In contrast, all hub genes were significantly overexpressed in CD patients compared to healthy controls in clinical samples (Fig. 3O-S).
Validation of diagnostic model efficacy and hub gene expression. (A-C) GSE102133. (D-F) GSE36807. (G) Construction of mouse models. (H) Body weight changes in mice over time. (I) Collection of mouse colon tissue samples. (J-N) Expression levels of hub genes in control and CD mice (N = 3). (O-S) Expression levels of hub genes in CD patients (involved areas, N = 5), with normal human colon tissues as controls. *p < 0.05, **p < 0.01.
Identification of 9 cell types in CD patients
To further investigate senescence heterogeneity at single-cell resolution in CD patients, we analyzed the scRNA-seq dataset GSE134809. Following stringent quality control measures, we retained 65,368 high-quality cells and 22,631 genes for subsequent analysis (Fig. S5). Based on established gene markers, we performed comprehensive cell annotation and ultimately classified 22 initial cell clusters into 9 distinct cell populations: fibroblasts, dendritic cells, T cells, B cells, macrophages, endothelial cells, epithelial cells, plasma cells, and NK cells (Fig. 4A-D). Analysis of matched colon samples (involved vs. uninvolved regions) from 11 CD patients (total n = 22) revealed a two-fold reduction in epithelial cell proportion in involved areas compared to uninvolved regions, suggesting potential compromise of intestinal mucosal integrity (Fig. 4E).
Senescence-associated gene expression patterns in CD patients from scRNA-seq data
To investigate cellular senescence characteristics in Crohn’s disease (CD) at single-cell resolution, we performed feature scoring analysis using both senescence-associated gene sets and hub genes across cell populations. The senescence-associated gene scoring revealed endothelial cells and fibroblasts as the predominant cell types exhibiting elevated scores, while hub gene scoring identified fibroblasts and macrophages as the highest-scoring populations (Fig. 4F, G). Comparative analysis between involved and uninvolved regions demonstrated significantly higher scores for both senescence-associated genes and hub genes in involved CD lesions (Fig. 4H-K). At the individual cell type level, fibroblasts showed the most pronounced differences in both scoring systems, with epithelial cells ranking second in hub gene score differentials (Fig. 4L, M).
In the T cell subpopulation analysis, we stratified T cells into three principal subtypes: tissue-resident CD8 + T cells, CD4 + T cells, and effector CD8 + T cells. The analysis revealed increase in CD4 + T cell proportion coupled with decrease in tissue-resident CD8 + T cell frequency in involved regions compared to uninvolved lesions (Fig. S6E), these findings demonstrate concordance with prior bulk RNA-seq-based immune infiltration profiling results.
Heterogeneity of cell annotation and senescence score in single-cell sequencing data of CD patients. (A) Cell annotation marker genes. (B, C) UMAP visualization of the cells were clustered into 22 distinct subclusters and subsequently annotated into 9 major cell types. (D) UMAP visualization of cell types stratified by involved vs. uninvolved regions in CD patients. (E) Pie chart showing cell proportions in involved vs. uninvolved groups (F, G) UMAP plot, single-cell senescence scores based on senescence-associated gene sets and hubgene scores based on hub genes. (H, I) Differential senescence-associated gene scores between involved and uninvolved groups. (J, K) Differential hubgene scores between involved and uninvolved groups. (L) Comparison of senescence scores between involved and uninvolved regions for each individual cell type in CD patients. (M) Comparison of hubgene scores between involved and uninvolved regions for each individual cell type in CD patients. Red indicates higher scores in involved group, blue represents higher scores in uninvolved group, Circle size represents the cell count per population (×1000). *p < 0.05, **p < 0.01, ***p < 0.001.
Analysis of fibroblast and epithelial cell subpopulations in CD patients
Given that fibroblasts exhibited the most significant differences in hub gene scores among all cell populations, we performed subpopulation analysis on 2,159 fibroblasts to further investigate their heterogeneity. Dimensionality reduction clustering revealed six distinct fibroblast subclusters (Fig. 5A), among which Subcluster 1 and Subcluster 3 showed marked proportional differences between involved and uninvolved regions (Fig. 5B). Comparative analysis of DEGs between Subcluster 1 and Subcluster 3 demonstrated significantly elevated expression of multiple collagen-related molecules and senescence markers (CDKN1A, CDKN2A) in Subcluster 1, whereas the stemness-associated gene SOX2 was substantially downregulated (Fig. 5C, D). Notably, Subcluster 3 exhibited high expression of neuroglial marker genes including S100B, PLP1 and NRXN1 (Fig. 5C). Functional enrichment analysis indicated that these DEGs were primarily involved in extracellular matrix collagen formation and associated with various neurological disorders (Fig. 5E).
Based on enrichment analysis results (Fig. 2) and differential gene set scores (Fig. 4L, M), which suggested a potential critical role of senescence-associated genes in regulating intestinal epithelial cell proliferation and differentiation in CD, we performed further subpopulation analysis of epithelial cells. This analysis identified four distinct subclusters (Fig. 5F), with Subcluster 0 and Subcluster 3 showing significant proportional differences between involved and uninvolved groups. Transcriptomic profiling revealed that Subcluster 0 exhibited a unique expression pattern characterized by upregulated genes involved in nutrient absorption and metabolism, including FABP6, SLC15A1, APOA4, APOA1, and APOC3, likely representing terminally differentiated intestinal absorptive epithelial cells. Notably, the proportion of this subpopulation was decreased in involved regions (Fig. 5G, H). In contrast, Subcluster 3 demonstrated higher abundance in involved regions compared to uninvolved controls, along with elevated expression of CDKN2A and an enhanced SASP (Fig. 5I, J).
Identification of core senescence-associated signaling pathways by GSVA
To further evaluate senescence-associated pathways at single-cell resolution across different cell types, we computed single-cell senescence scores based on a senescence-associated gene set and quantified Hallmark pathway activities using GSVA. Correlation analysis between these two metrics was performed for all cell populations. The results demonstrated that the epithelial-mesenchymal transition (EMT) pathway showed significant positive correlations with senescence scores consistently across all cell types. Moreover, other pathways including INFLAMMATORY RESPONSE, TNFA SIGNALING VIA NFKB, APOPTOSIS, and HYPOXIA also demonstrated strong positive correlations with senescence scores in Multiple cell types (Fig. 5K).
Analysis of fibroblast and epithelial subpopulation. (A) UMAP dimensionality reduction clustering of 2,159 fibroblasts (6 subclusters). (B) Proportional differences of fibroblast subclusters between involved and uninvolved regions. (C) Heatmap of differentially expressed genes between fibroblast Subcluster 1 and Subcluster 3: red dots represent upregulated genes in Subcluster 1, blue dots indicate upregulated genes in Subcluster 3. (D) Differential expression of senescence markers (CDKN1A, CDKN2A) between Subcluster 1 and Subcluster 3. (E) Functional enrichment analysis of differentially expressed genes: (left) GO biological process analysis; (right) KEGG pathway enrichment. (F) UMAP dimensionality reduction clustering of epithelial cells (4 subclusters). (G) Proportional differences of epithelial subclusters between involved and uninvolved regions. (H) Violin plot showing significantly enriched signature genes in Subcluster 0. (I) Expression distribution of senescence marker CDKN2A across epithelial subclusters, with high-expression cells predominantly localized in Subcluster 3; (right) Differential CDKN2A expression between involved and uninvolved regions within Subcluster 3. (J) Expression profiles of SASP factors in Subcluster 3 across different groups. (K) Integrated analysis of cellular senescence and pathway activity at single-Cell resolution. *p < 0.05, **p < 0.01, ***p < 0.001.
Discussion
Crohn’s disease (CD), as a multifactorial complex disorder, poses significant challenges for clinical management, typically requiring lifelong treatment15. Current research indicates its pathogenesis involves genetic susceptibility, gut microbiota dysbiosis, Western dietary patterns, and environmental triggers, among other factors3. While conventional therapies targeting these mechanisms have achieved certain therapeutic effects, the disease remains associated with a high postoperative recurrence rate for reasons that are not yet fully understood16. Since CD predominantly affects younger populations, research exploring the link between cellular senescence and CD has been relatively scarce. However, emerging evidence shows significant upregulation of cellular senescence markers (e.g., p16 and p21) in the intestinal stem cell niche of CD patients, suggesting a potential role of senescence in disease pathogenesis11. Furthermore, clinical data indicate that elderly CD patients have a poorer prognosis compared to their younger counterparts, further underscoring the critical role of cellular senescence in CD17. Therefore, investigating the potential connections between senescence-associated genes and CD could enhance our understanding of novel key mechanisms driving CD progression and provide new therapeutic targets.
The senescence-associated secretory phenotype (SASP), through the sustained release of pro-inflammatory factors (e.g., IL-6, IL-8, and matrix metalloproteinases)18,19,20, may mediate persistent activation of mucosal immunity – a mechanism highly consistent with the characteristic chronic inflammation pathology of CD. Additionally, given the high turnover rate of intestinal epithelial cells, senescence or exhaustion of intestinal stem cells could significantly impair mucosal repair capacity and lead to compromised epithelial barrier function21,22. This study successfully identified five key hub genes (STAT1, S100A11, F3, FILIP1L, and HPS5) closely associated with cellular senescence and established a high-performance diagnostic prediction model for CD. Using single-cell transcriptomic data, we computed senescence-associated scores and hub gene-based scores for individual cells. The results revealed that both senescence scores and hub gene scores were significantly higher in fibroblasts and epithelial cells from CD-involved areas compared to non-involved areas. Concurrently, the expression level of the stemness-related gene SOX2 was markedly reduced. These findings collectively suggest that senescent fibroblasts and epithelial cells are key drivers of CD progression. Senescent fibroblasts likely contribute to fibrosis (stricturing) in CD. By persistently secreting pro-inflammatory and pro-fibrotic factors (e.g., MMPs) via SASP, they directly drive tissue remodeling and scar formation, potentially representing a primary cause of intestinal obstruction. Senescence of epithelial cells directly explains the core issue of barrier function defect in CD. The diminished renewal and repair capacity of senescent epithelial cells leads to a “leaky” gut barrier, allowing intestinal bacteria and antigens to penetrate the intestinal wall, thereby persistently activating the mucosal immune system and initiating/sustaining chronic inflammation. SOX2 is a key transcription factor maintaining the self-renewal and multilineage differentiation capacity of intestinal stem cells; its downregulation directly indicates the exhaustion or functional failure of these stem cells. Future therapeutic strategies combining anti-senescence therapies and stem cell-based treatments may represent a promising avenue for CD management.
The five identified CD-related hub genes may play significant roles in CD progression: As a core regulator of the interferon (IFN) signaling pathway, STAT1 can be persistently activated by chronic inflammatory stimuli, thereby accelerating cellular senescence23. Mechanistic studies indicate that TNF-α and IFN-γ establish a positive feedback loop via hyperactivation of the JAK/STAT1 cascade, ultimately amplifying the SASP effect24. The pro-inflammatory calcium-binding protein S100A11 may sustain chronic inflammation by activating the NF-κB pathway25. Research reveals it can competitively disrupt the SIRT6-Smad2/3 interaction, subsequently activating TGF-β signaling to drive fibrotic progression26, providing new insights into intestinal fibrosis development in CD patients. Beyond its classical role in coagulation, emerging evidence suggests F3 can trigger macrophage activation and TNF-α secretion via PAR2 receptor signaling27, offering a theoretical framework for understanding F3’s role in CD intestinal inflammation. HPS5 encodes a core component of the Biogenesis of Lysosome-related Organelles Complex 2 (BLOC-2). Mutations in HPS5 are associated with Hermansky-Pudlak syndrome (HPS) – patients with this rare genetic disorder (particularly HPS1 and HPS4 subtypes) often exhibit gastrointestinal manifestations highly similar to CD28, suggesting HPS5 may play a role in maintaining intestinal homeostasis. The actin-binding protein FILIP1L is closely involved in cytoskeletal dynamics, cell motility, and signal transduction, and has been demonstrated to regulate epithelial-mesenchymal transition (EMT) – its knockdown enhances EMT while its overexpression reverses this phenotype29.
Notably, this study found that HPS5 and FILIP1L exhibit unique expression patterns in CD: although bulk RNA sequencing showed their overall upregulation in CD tissues, single-cell transcriptomic analysis revealed a higher expression trend in non-lesional areas. These findings suggest that HPS5 and FILIP1L might participate in compensatory protective mechanisms in relatively uninvolved intestinal segments (Fig. S7A, B). Analysis of the GSE75214 dataset confirmed that HPS5 expression was significantly higher in non-active areas compared to active areas, and both HPS5 and FILIP1L expression levels in non-active areas were significantly higher than in normal controls (Fig. S7C, D). These results confirm the spatial expression heterogeneity of the hub genes: HPS5 and FILIP1L may exert compensatory protective functions in non-lesional bowel segments. This does not detract from the predictive performance of our model; on the contrary, it underscores the robust predictive power of these hub genes when considered collectively. This could also explain the lack of statistical significance for HPS5 and FILIP1L in the mouse model: unintended inclusion of uninvolved segments during tissue sampling, where compensatory upregulation occurs, might have diluted measurable differences. Overall, the expression trends of these hub genes in the mouse model align with the transcriptomic analysis results, while the excellent validation using clinical specimens likely stems from precise identification of lesional areas during endoscopic/pathological sampling.
In the single-cell transcriptomic analysis section, we unveiled the heterogeneous landscape of cellular senescence within the CD tissue microenvironment: fibroblasts exhibited the most prominent senescent phenotype, with pro-fibrotic subpopulations specifically enriched in lesional areas, whereas neuron-like S100B + subpopulations dominated non-lesional areas. This spatial distribution pattern highly corresponds to the characteristic “skip lesions” pathology of CD30,31, suggesting that senescence-associated fibroblasts might be key effector cells driving the regional heterogeneity of intestinal wall fibrosis. Among epithelial cells, we observed significant enrichment of senescent epithelial cells with high CDKN2A expression in lesional areas, providing direct evidence for the progressive decline in epithelial regenerative capacity during CD progression. Notably, metabolically reprogrammed epithelial cluster 0 showed significant upregulation of FABP6 and apolipoproteins (APOA1/APOA4), and the proportion of this cluster was downregulated in lesional areas. This finding aligns with emerging evidence on the gut microbiota-bile acid-lipid metabolism axis in CD pathogenesis, indicating that senescent epithelia might promote disease progression by remodeling the local metabolic microenvironment32.
This study has several limitations. The model was constructed using untreated and normal intestinal mucosa samples, which excludes local genetic variation differences caused by biologic therapy before and after treatment, making it more suitable for diagnostic prediction in treatment-naïve CD patients. Consequently, it lacks a systematic analysis of senescence-associated gene expression profiles before and after treatment, or between recurrent and non-recurrent CD tissues. Such analyses could provide crucial insights into recurrence mechanisms and guide clinical treatment strategies, representing an important direction for future research. Secondly, current single-cell RNA sequencing technology cannot distinguish between the mucosal layer and full-thickness intestinal wall tissue, thereby limiting the spatial resolution of senescence-associated gene expression patterns. Furthermore, substantial evidence indicates that the gut microbiome interacts closely with host immunity and epithelial function, and its dysbiosis may directly or indirectly regulate the senescence-related pathways under investigation. Therefore, the extent to which the observed gene expression variations are driven by underlying microbial differences cannot be excluded. Future studies utilizing prospective cohorts with comprehensive clinical metadata and paired metagenomics are essential to further validate the stability of these senescence-associated hub genes and deeply explore the role of host-microbiota interactions in driving cellular senescence.
In summary, this study, based on bulk and single-cell transcriptomics, systematically delineated a expression landscape of senescence-associated genes in CD, identified a novel diagnostic prediction model for CD comprising five senescence-associated genes, provided new insights into the relationship between cellular senescence and CD, offered a fresh perspective on understanding disease heterogeneity, and laid a molecular foundation for developing therapeutic strategies targeting cellular senescence.
Data availability
The datasets used in this study were obtained from open-access repositories. Detailed information regarding the storage platforms and corresponding accession numbers can be found in the article.
References
Torres, J., Mehandru, S., Colombel, J. F. & Peyrin-Biroulet, L. Crohn’s disease. Lancet 389 (10080), 1741–1755 (2017).
Ananthakrishnan, A. N. Epidemiology and risk factors for IBD. Nat. Rev. Gastroenterol. Hepatol. 12 (4), 205–217 (2015).
Heerasing, N. & Kennedy, N. A. Interaction between NOD2 and smoking in the pathogenesis of crohn’s disease. EBioMedicine 21, 49–50 (2017).
Allen, P. B. et al. Review article: moving towards common therapeutic goals in crohn’s disease and rheumatoid arthritis. Aliment. Pharmacol. Ther. 45 (8), 1058–1072 (2017).
Gravina, A. G. et al. Crohn’s disease and skin. United Eur. Gastroenterol. J. 4 (2), 165–171 (2016).
Gisbert, J. P. & Chaparro, M. Anti-TNF agents and new biological agents (Vedolizumab and Ustekinumab) in the prevention and treatment of postoperative recurrence after surgery in crohn’s disease. Drugs 83 (13), 1179–1205 (2023).
Zhang, L. et al. Cellular senescence: a key therapeutic target in aging and diseases. J. Clin. Invest. 132 (15), e158450 (2022).
McHugh, D., Durán, I. & Gil, J. Senescence as a therapeutic target in cancer and age-related diseases. Nat. Rev. Drug Discov. 24 (1), 57–71 (2025).
Singh, A. et al. Aging and inflammation. Cold Spring Harb Perspect. Med. 14 (6), a041197 (2024).
Li, X. et al. Inflammation and aging: signaling pathways and intervention therapies. Signal. Transduct. Target. Ther. 8 (1), 239 (2023).
Wang, X., Bootsma, H., Kroese, F., Dijkstra, G. & Pringle, S. Senescent stem and transient amplifying cells in crohn’s disease intestine. Inflamm. Bowel Dis. 26 (2), e8–e9 (2020).
Kanehisa, M., Furumichi, M., Sato, Y., Matsuura, Y. & Ishiguro-Watanabe, M. KEGG: biological systems database as a model of the real world. Nucleic Acids Res. 53 (D1), D672–D677 (2025).
Newman, A. M. et al. Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods. 12 (5), 453–457 (2015).
Liu, J. et al. Screening of crosstalk and pyroptosis-related genes linking periodontitis and osteoporosis based on bioinformatics and machine learning. Front. Immunol. 13, 955441 (2022).
Bachour, S. P. & Click, B. H. Clinical update on the prevention and management of postoperative crohn’s disease recurrence. Curr. Gastroenterol. Rep. 26 (2), 41–52 (2024).
Petagna, L. et al. Pathophysiology of crohn’s disease inflammation and recurrence. Biol. Direct. 15 (1), 23 (2020).
Singh, S., Poulsen, G. J., Bisgaard, T. H., Bonfils, L. & Jess, T. Epidemiology of elderly onset IBD: a nationwide Population-Based cohort study. Clin. Gastroenterol. Hepatol. 23 (7), 1204–1215e11 (2025).
Wei, L. et al. H3K18 lactylation of senescent microglia potentiates brain aging and alzheimer’s disease through the NFκB signaling pathway. J. Neuroinflammation. 20 (1), 208 (2023).
Wang, Y. et al. The role of cellular senescence in neurodegenerative diseases. Arch. Toxicol. 98 (8), 2393–2408 (2024).
Guccini, I. et al. Senescence reprogramming by TIMP1 deficiency promotes prostate cancer metastasis. Cancer Cell. 39 (1), 68–82e9 (2021).
Malagola, E. et al. Isthmus progenitor cells contribute to homeostatic cellular turnover and support regeneration following intestinal injury. Cell 187 (12), 3056–3071e17 (2024).
van der Flier, L. G. & Clevers, H. Stem cells, self-renewal, and differentiation in the intestinal epithelium. Annu. Rev. Physiol. 71, 241–260 (2009).
Clark, D. N., Begg, L. R. & Filiano, A. J. Unique aspects of IFN-γ/STAT1 signaling in neurons. Immunol. Rev. 311 (1), 187–204 (2022).
Kandhaya-Pillai, R. et al. TNF-α/IFN-γ synergy amplifies senescence-associated inflammation and SARS-CoV-2 receptor expression via hyper-activated JAK/STAT1. Aging Cell. 21 (6), e13646 (2022).
Li, W. et al. Inhibiting NF-κB-S100A11 signaling and targeting S100A11 for anticancer effects of demethylzeylasteral in human colon cancer. Biomed. Pharmacother. 168, 115725 (2023).
Zhu, T. et al. The S100 calcium binding protein A11 promotes liver fibrogenesis by targeting TGF-β signaling. J. Genet. Genomics. 49 (4), 338–349 (2022).
Wang, J., Chakrabarty, S., Bui, Q., Ruf, W. & Samad, F. Hematopoietic tissue factor-protease-activated receptor 2 signaling promotes hepatic inflammation and contributes to pathways of gluconeogenesis and steatosis in obese mice. Am. J. Pathol. 185 (2), 524–535 (2015).
O’Brien, K. J. et al. Inflammatory bowel disease in Hermansky-Pudlak syndrome: a retrospective single-centre cohort study. J. Intern. Med. 290 (1), 129–140 (2021).
Jing, R. et al. FILIP1L-mediated cell apoptosis, epithelial-mesenchymal transition and extracellular matrix synthesis aggravate posterior capsular opacification. Life Sci. 286, 120061 (2021).
Herren, R. & Geva-Zatorsky, N. Spatial features of skip lesions in crohn’s disease. Trends Immunol. 45 (6), 470–481 (2024).
Feuerstein, J. D. & Cheifetz, A. S. Crohn disease: epidemiology, diagnosis, and management. Mayo Clin. Proc. 92 (7), 1088–1103 (2017).
Lv, Y. et al. The impact of exclusive enteral nutrition on the gut Microbiome and bile acid metabolism in pediatric crohn’s disease. Clin. Nutr. 42 (2), 116–128 (2023).
Funding
This work was supported by the National Natural Science Foundation of China (Grant No. 82173107 and 82372703), the Natural Science Foundation of Hunan Province (Grant No. 2024JJ5580), and the Fundamental Research Funds for the Central Universities of Central South University (Grant Nos. CX20240300).
Author information
Authors and Affiliations
Contributions
M. L, Z. Y, X. Luo, and M. Zhao conceived and designed the study; M. L, X. L, C. C, and M. G performed data analysis; X. L, Z. Y and M. Z conducted the experiments; X. L, C. C and M. G drafted the manuscript; M. L, Z. Y and M. Z revised and edited the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Luo, X., Zhao, M., Cui, C. et al. Senescence-related gene signatures in Crohn’s disease: integrating bulk and single-cell RNA sequencing analysis. Sci Rep 15, 45269 (2025). https://doi.org/10.1038/s41598-025-29727-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-29727-3