Abstract
Ventilator-induced diaphragmatic dysfunction (VIDD) is characterized by diaphragmatic atrophy and contractile failure, leading to prolonged intensive care unit (ICU) stays and increases mortality. While phrenic nerve stimulation (PNS) has demonstrated efficacy in mitigating VIDD by preserving diaphragmatic activity, its underlying molecular mechanisms remain unclear. This study aimed to elucidate the role of competing endogenous RNA (ceRNA) networks in PNS-mediated protection against VIDD through integrated miRNA-Seq and RNA-Seq analyses in a rabbit model. Eleven adult male New Zealand white rabbits were divided into control (n = 4), MV (n = 3) and PNS groups (n = 4). MV and PNS groups underwent 24 h of MV, with intermittent bilateral transvenous PNS applied only to the PNS group. Differentially expressed (DE) analysis of mRNAs, miRNAs and circRNAs across pairwise group comparisons was performed via RNA-seq and miRNA-seq. Functional enrichment analyses (Gene Ontology and Kyoto Encyclopedia of Genes and Genomes) identified key pathways. Potential miRNA targets and interacting circRNAs were computationally predicted. An integrated ceRNA network was constructed using major DE RNAs to identify PNS-associated core regulators. CeRNA network was validated by quantitative real-time polymerase chain reaction (RT-qPCR) and dual-luciferase assays. High-throughput sequencing revealed significant dysregulation of miRNAs, circRNAs and mRNAs in the diaphragm following MV which was partially reversed by PNS. Bioinformatic screening identified a ceRNA network, wherein two key miRNAs emerged: miR-500-3p (targeting RAB37, an autophagy-related gene) and miR-133b-3p (targeting L-selectin, a cell adhesion molecule regulating immune responses and fibrosis). Both miRNAs were down-regulated after MV and restored by PNS. Computational prediction also identified five circRNAs (circRNA_12437, 24673, 14127, 14942, 12463) as putative sponges for these miRNAs, although this interaction lacks experimental confirmation. Dual-luciferase assays confirmed direct binding of miR-500-3p to RAB37 and miR-133b-3p to L-selectin, functionally linking them to PNS-mediated VIDD protection. Enrichment analyses indicated that DE genes were predominantly enriched in phagosome activity and cell adhesion molecule pathways. Collectively, these findings suggest that PNS preserves diaphragmatic function by modulating ceRNA networks to suppress excessive autophagy and immune cell infiltration. This study identifies the first PNS-responsive ceRNA network in VIDD pathogenesis. Our data highlight the potential critical roles of miR-500-3p-RAB37 and miR-133b-3p-L-selectin axes in regulating autophagy and immune responses. These results provide mechanistic insights and suggest potential therapeutic targets for diaphragm dysfunction.
Similar content being viewed by others
Introduction
Ventilator-induced diaphragmatic dysfunction (VIDD) represents a critical challenge in critical care medicine, characterized by progressive diaphragmatic atrophy and contractile failure during mechanical ventilation (MV)1. This condition significantly impedes ventilator weaning, prolongs intensive care unit (ICU) stays and increases risks of pulmonary complications and death. Understanding the mechanisms of VIDD and developing effective treatments are therefore crucial to mitigate these adverse outcomes2,3. In the context of VIDD, increased free reactive oxygen species (ROS) production activate four major proteolytic systems: ubiquitin–proteasome system (UPS); autophagy; caspase-3 and calpains, leading to diaphragm proteolysis and contractile dysfunction4,5. MV increasing mitochondrial ROS emission promotes autophagy and diaphragm atrophy by lysosomal degradation of cytosolic proteins and organelles in the diaphragm6,7. Indeed, the previous study also confirm that autophagy is required for VIDD, inhibition of autophagy may reduce ROS production to prevent diaphragm atrophy and contractile dysfunction in rats after 12 h of MV8. In a 24 h rabbit VIDD model, single-nucleus RNA sequencing (snRNA-seq) results showed that acute post mechanical ventilation diaphragm cell changes included an increase in the proportion of fibroblasts and a decrease in the proportion of myofibers, and immune cell infiltration may play a role in the early stage of driving diaphragm fibrosis during mechanical ventilation9. The findings suggest that myocyte atrophy related to autophagy and fibrosis associated with immune cell infiltration may jointly contribute to VIDD.
The weakening of diaphragmatic contractile movement is a key factor in the development of VIDD. Phrenic nerve stimulation (PNS) can maintain diaphragmatic activity and mitigate the adverse effects of MV on diaphragmatic function10. PNS preserves the contractile function of the rat diaphragm after 18 h of MV11. Another study confirmed that pigs receiving transvenous phrenic nerve pacing synchronized with MV had less diaphragm fiber atrophy than those receiving MV12. The RESCUE 2 study also demonstrated that temporary transvenous diaphragmatic pacing could correct diaphragmatic disuse atrophy and accelerate weaning in patients13. In addition, unilateral phrenic nerve stimulation in patients undergoing cardiothoracic surgery and receiving MV increased the mitochondrial respiratory rate of the diaphragm on the stimulated side14, this suggests that PNS can improve diaphragmatic energy metabolism, which is crucial for maintaining diaphragmatic contractile function. The merging evidences suggest that PNS can protect the diaphragm against VIDD through inhibits oxidative stress, improve mitochondrial oxygen utilization and enhance mitochondrial function, promote protein synthesis and strengthen diaphragmatic tone15. Therefore, using PNS to preserve diaphragmatic contractile activity may be a new approach in VIDD treatment and prevention. However, the effect of electrical stimulation on autophagy and immune cell infiltration which lead to VIDD is unclear.
A competing endogenous RNA (ceRNA) network plays a critical role in muscle cell differentiation and development. This complex regulatory system, comprising circular RNAs (circRNAs), microRNAs (miRNAs) and messenger RNAs (mRNAs), has been increasingly implicated in various muscle atrophy conditions16. For instance, distinct ceRNA networks have been identified across different pathological contexts: the RP11-253E3.3–miR-1207-5p–FOXO3 axis in spinal cord injury-induced muscle atrophy17, the Xist–miR-486-5p–Stk4 network during myoblast proliferation18, MALAT1-mediated networks in amyotrophic lateral sclerosis19. Within these networks, circRNAs function as molecular sponges by competitively binding to miRNA response elements (MREs), thereby sequestering miRNAs and preventing the suppression of their target genes. It has been found that overexpression of circTmeff1 induces muscle atrophy, whereas its knockdown produces protective effects20. Meanwhile, miRNAs themselves, as small endogenous RNAs of 19-23 nucleotides, typically bind to the 3′-untranslated regions (3′-UTRs) of target mRNAs to repress their translation. Several miRNAs regulating autophagy and fibrosis have emerged as potential key players in VIDD pathogenesis. Moroz et al.21 reported decreased expression of autophagy-related miRNAs (miR-106b, miR-20a, miR-101a, miR-376, miR-204 and miR-93) in diaphragm tissue during MV. Similarly, downregulation of miR-122-5p has been observed in skeletal muscle fibrosis, with its overexpression shown to attenuate fibrotic processes by targeting the TGF-β/Smad signalling pathway22. Previous RNA-seq data further revealed that MV-induced diaphragm atrophy and impaired energy metabolism are associated with upregulation of Myog, Trim63 and Fbxo32 alongside downregulation of Ppargc1a in rodent models23. In the context of electrical stimulation, Dong et al.24 demonstrated that transvenous PNS prevents VIDD by reducing oxidative stress through involvement of Aldh1a1 and Slc25a30. Additional findings indicate that 12-h MV upregulates muscle atrophy genes (Mstn, Trim63, Fbxo3) while downregulating regeneration-related genes (Trim72, Casq2, Myod1) and that PNS counteracts these changes by suppressing catabolic genes (Pnpla2, Samd4b, Tmem259, Pcbp4) while enhancing expression of genes involved in oxygen transport and mitochondrial respiration such as Ndufa225. Although certain miRNAs and mRNAs have been partially characterized in VIDD and PNS, the mechanism by which PNS prevents VIDD through ceRNA networks remains unexplored.
In this study, we analyze miRNA-Seq and RNA-Seq profiles from ventilated subjects with/without PNS to systematically investigate the differential expression levels of circRNAs, miRNAs and mRNAs by high-throughput RNA sequencing. Furthermore, we conducted a circRNA-miRNA-mRNA networks to explore the potential roles of autophagy and immune cell infiltration on the effect of electrical stimulation in a VIDD rabbit model. Additionally, we performed Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses to predict the biological functions and pathways annotated to DE mRNAs. The findings may help shed light on the roles of ceRNA networks in the pathophysiology underlying PNS preventing VIDD and provide potential new targets for the treatment of VIDD.
Methods
Animals
Eleven adult male New Zealand white rabbits (aged: six-eight months, weight 2.5–3.0 kg) were obtained from the Laboratory Animal Research Center at Shanxi Medical University in Taiyuan, China. These animals were selected due to their suitable body size and established utility in previous VIDD model investigations24. The subjects were maintained under controlled environmental conditions with 12-h light/dark cycles and unrestricted access to food and water throughout the study period. All experimental procedures strictly adhered to the National Institutes of Health guidelines for laboratory animal welfare and were approved by the Institutional Animal Care and Use Committee of Changzhi Medical College.
Mechanical ventilation and transvenous PNS
Rabbits were anesthetized by intraperitoneal injection of 3% sodium pentobarbital solution (40 mg/kg) and then continuous intravenous infusion via the right marginal ear vein using a pump (10 mg/kg/h). All the rabbits were randomly divided into the following three groups: control group (n = 4), MV group (n = 3) and PNS group (stimulation of the bilateral phrenic nerves for 10 min repeated every 2 h during MV, n = 4). The primary study endpoint was defined as completion of 24 h mechanical ventilation. No secondary endpoints were established. The trachea was connected to the mechanical ventilator (Inspira ASVV 55-7058; Harvard Apparatus, Cambridge, MA, USA) after tracheotomy. The ventilator settings were as follows: inspired fraction of oxygen, 0.40; tidal volume (Vt), 6–8 mL/kg; respiratory rate, 40 cycles per minute; and positive end-expiratory pressure, 2 cm H2O. Rabbits were kept warm with heating pads during the procedure (Fig. 1). Arterial blood was obtained every 12 h for determination of PO2, PCO2 and pH at 37 °C with a blood gas analyzer (ABL 800 FLEX blood gas analyzer, Radiometer, Denmark). For phrenic nerve stimulation, bilateral external jugular veins were surgically exposed and cannulated with 7.0F percutaneous sheath introducers (Arrow, Teleflex, Morrisville, NC, USA). Electrode catheters (Medtronic, Minneapolis, MN, USA) were advanced through the sheaths and positioned under tactile guidance by optimizing diaphragmatic contraction during low-current stimulation. Catheters were secured upon achieving maximal stimulation efficacy. A pulse generator (Department of Biomedical Engineering, Changzhi Medical College, China) delivered bilateral transvenous stimulation using parameters of 40 Hz frequency, 300 μs pulse duration, 40 cycles/min respiratory rate, 1:2 contraction-relaxation ratio and 2.5 mA peak current (Fig. 1). After 24 h of ventilation, deep anesthesia was maintained with intravenous 3% sodium pentobarbital (50 mg/kg), and rabbits were euthanized via exsanguination (species: New Zealand white rabbits, age: six-eight months, weight: 2.5–3.0 kg), with diaphragm tissues promptly frozen in liquid nitrogen for subsequent analysis.
Diagrammatic representation of the experimental setup. The rabbit was placed in the supine position and anesthetized. The endotracheal tube was connected to a mechanical ventilator. The electrical stimulation electrodes were connected to the pulse generator.
Small RNA sequencing
Total RNA was extracted from rabbit diaphragm samples using the mirVana miRNA Isolation Kit (Ambion) following the manufacturer’s protocol. RNA concentration was quantified using a Nanodrop 2000 spectrophotometer (Thermo Fisher Scientific, USA), and RNA integrity was verified through analysis on an Agilent 2100 Bioanalyzer (Agilent Technologies, USA). Small RNA libraries were prepared from 1 μg of total RNA per sample using the NEBNext Small RNA Library Prep Kit for Illumina (NEB #E7330S, USA). The procedure involved sequential 3′ and 5′ adapter ligation, reverse transcription to cDNA, and PCR amplification. Size selection of 140–160 bp fragments was performed using gel electrophoresis, followed by library quality verification on the Agilent 2100 Bioanalyzer. Final sequencing was conducted on an Illumina Novaseq 6000 platform (OE Biotech Co., Ltd., China) to generate 150 bp paired-end reads.
Raw sequencing reads were processed through the following quality control pipeline: (1) Removal of low-quality reads (Phred score < 20); (2) Trimming of 5′ adapter sequences and poly-A tails; (3) Exclusion of reads lacking 3′ adapters or containing insert sequences outside the 15–41 nt range. Clean reads were mapped to the reference genome using Bowtie and annotated against Rfam v10.1 to filter non-coding RNAs (rRNA, tRNA, snRNA, etc.). Potential repeat sequences were identified through alignment with the Repbase database. Known miRNAs were annotated using miRBase v22, with novel miRNAs predicted through miRDeep2 analysis26.
Differential miRNA expression analysis was performed using the DEseq2 algorithm in R with significance thresholds set at P < 0.05 and |log2FC|> 1. miRNA target prediction was conducted using miRanda with stringent parameters: alignment score ≥ 150, free energy ΔG ≤ −30 kcal/mol, and mandatory 5′ seed region complementarity. Novel miRNAs were further validated through identification of characteristic hairpin structures in precursor sequences and detection of corresponding miRNA-star sequences.
Whole RNA sequencing (RNA-Seq)
Total RNA was isolated from samples using TRIzol reagent (Invitrogen, CA, USA) following the manufacturer’s protocol. RNA integrity was evaluated using the Agilent 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA), and only samples with an RNA Integrity Number (RIN) ≥ 7 were retained for subsequent analysis. For mRNA library construction, 4 µg of DNase-treated total RNA was incubated with Oligo (dT)-conjugated magnetic beads to enrich polyadenylated mRNAs, followed by fragmentation, first-and second-strand cDNA synthesis, end repair, A-tailing, adapter ligation and PCR amplification. For circRNA enrichment, an aliquot of total RNA was treated with RNase R (Epicentre, 4 U/µg RNA) at 37 °C for 15 min to digest linear RNAs, and ribosomal RNA was subsequently removed using the Ribo-Zero Gold rRNA Removal Kit (Illumina).
Raw reads generated during high-throughput sequencing were fastq format sequences. In order to get high-quality reads that could be used for later analysis, raw reads needed to be further quality filtered. Trimmomatic27 software was first used for adopter removing, and then low-quality bases and N-bases or low-quality reads were filtered out. Finally, we got high-quality clean reads. Clean reads were aligned to the Oryctolagus cuniculus reference genome (Ocun2.0) using Hisat2 (v2.2.1)28. The result of alignment with the reference genome was stored in a binary file, called a bam file. Using the Stringtie29 software to assemble the reads, the new transcript was spliced. The gene expression levels (FPKM) were quantified via FPKM formula. Differential mRNA expression analysis was performed with DESeq2 (v1.30.1), applying an adjusted P-value < 0.05.
For circRNA analysis, we used BWA software to align the sequencing reads of each sample with reference genome in order to generate SAM file. Then CIRI software was used to scan for PCC (paired chiastic clipping) signals, and circRNA sequences were predicted based on junction reads and GT-AG cleavage signals. We uesd the RPM algorithm to quantify the circRNAs, normalized the number of junction reads counts and foldchange by DESeq. Enrichment of differentially expression (DE) circRNAs was analysised through the annotation information of circRNA source transcripts. CircRNAs can serve as miRNA target molecules. Miranda software was used to predict circRNA/miRNA interactions.
Gene ontology (GO) and Kyoto encyclopedia of genes and genomes (KEGG) pathway enrichment analysis
GO and KEGG pathway enrichment analyses were performed to functionally annotate the DE genes using the Database for Annotation, Visualization, and Integrated Discovery (DAVID, v6.8; https://david.ncifcrf.gov/). For GO analysis, functional annotations were categorized into three domains: biological processes (BP), cellular components (CC), and molecular functions (MF). Statistical significance was determined using the Benjamini–Hochberg method to control the false discovery rate (FDR), with an adjusted P-value < 0.05 set as the threshold. Visualization of the enrichment results, including GO term relationships and pathway-gene networks, was implemented using the GOplot R package (v1.0.2).
Construction of ceRNA networks
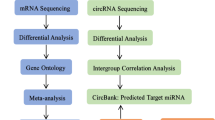
In this study, we constructed a short linear sequence by joining the 100 bp tail and 100 bp head sequences of circRNA. This chimeric sequence was then analyzed using miRanda software for miRNA interaction prediction. The prediction process integrated two key parameters: miRNA sequence complementarity and binding energy stability assessment. Through this comprehensive computational analysis, we identified miRNAs that exhibit potential binding interactions with both the circRNA-derived sequence and mRNA targets. A ceRNA network was constructed based on competitive endogenous RNA mechanism and DE genes. The workflow is shown in Fig. 2A.
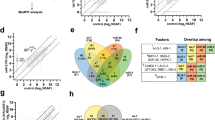
Identification of DE mRNAs, DE circRNAs and DE miRNAs. Study design flowchart (A). Principal component analysis (PCA) score plot showing sample grouping (B). Volcano plots presenting the differentially expressed miRNAs (C, D), circRNA (F, G) and mRNAs (I, J) between MV vs Control group and PNS vs MV group. The horizontal dashed line indicates the threshold for significance (P ≤ 0.05) and the vertical dashed line indicates the up-regulated (right side) and down-regulated (left side) RNAs. Up-regulated miRNAs were marked in red and down-regulated miRNAs are indicated in blue. Venn cluster of the DE miRNAs (E), DE circRNA (H) and DE mRNAs (K) in MV vs Control group and PNS vs MV group, with the orange section representing MV vs Control group and the blue representing PNS vs MV. C: control group, M: MV group, P: PNS group. The same as below.
Quantitative real-time polymerase chain reaction (RT-qPCR)
Total RNA was isolated using Trizol reagent (Seven, Beijing, China), and cDNA was synthesized using PrimeScript II RTase (Seven, Beijing, China), recombinant RNase inhibitor (Seven, Beijing, China) and oligo (dT) 18 primer (Seven, Beijing, China). ThePCR mixtures were prepared to a final volume of 20 µL using SYBR FAST qPCR Master Mix (Seven, Beijing, China). RT-qPCR was undertaken via a CFX Connect 96RT PCR detection system (Bio-Rad, Hercules, CA, USA). The comparative CT method (ΔΔCT) was utilized for the purpose of analyzing gene expression data, and GAPDH was regarded as an internal control. The presentation of results was undertaken in form of fold-change relative to control. Primer sequences are listed in Table S1.
Dual-luciferase reporter assays
Computational prediction of miR-500-3p, miR-133b-3p binding to the complementary seed matches within the RAB37, L-selectin genes was validated by luciferase reporter assay. In brief, the sequence harboring the predicted miRNA interaction site was cloned into the pEZX-MT06 vector. 293T cells were seeded into 96-well plates, and when the cell density reached 70–80%, the pEZX-MT06 vector containing either the wild type (WT) or mutant type (MUT) sequence was co-transfected with the synthetic miR-500-3p, miR-133b-3p mimic or negative control (NC) into 293T cells using Lipofectamine® 2000 Reagent (Invitrogen, Carlsbad, CA, USA). Firefly and Renilla luciferase activities were measured 48 h after transfection, using a dual-luciferase reporter system (Seven, Beijing, China) following the manufacturer’s protocol. The firefly luciferase activity was normalized by the Renilla luciferase activity. Each experiment was repeated at least three times.
Statistical analysis
A post-hoc power analysis was performed using the R package RNASeqPower to evaluate the statistical power of our study design (n = 11) to detect differential expression under the thresholds of a twofold change and a significance level (α) of 0.05. Data were analyzed using SPSS version 22.0 (IBM Corporation, Armonk, NY, USA). Normality of distribution was evaluated via the Kolmogorov–Smirnov test, with data presented as mean ± standard deviation (SD). For comparing differences across multiple groups, one-way analysis of variance (ANOVA) followed by post-hoc LSD test was performed. Statistical significance was defined as a two-tailed P < 0.05.
Results
Comparisons of physiological parameters between the three groups
No significant intergroup differences were observed in baseline hemodynamic parameters (heart rate, blood pressure), tidal volume(Vt), or arterial blood gas values (pH, PO2, PCO2) among control, MV and PNS groups (Table S2). These parameters showed no significant differences across all groups at both 12 and 24 h timepoints (Table S2), indicating that they did not act as significant confounding factors in subsequent analyses.
Quality control metrics for diaphragm miRNA-seq and RNA-seq data after MV and PNS
miRNA-seq data yielded an average of 20 million (M) clean reads per sample after quality control. The sequencing base quality scores (Q20) ranged from 93.46 to 97.02% (Table S3). Reads with a length of 22 bp were the most abundant, indicating well-constructed miRNA-seq libraries (Table S3). Transcriptome sequencing was performed on the Illumina platform, generating expression profiles for all 11 qualified RNA samples. The Q30 value (indicating bases with a base calling accuracy ≥ 99.9%) exceeded 95% for all samples. High-quality clean reads accounted for 96 to 97% of the raw data. These clean reads were aligned to the rabbit reference genome using HISAT2, resulting in 81.36 to 99.28 M genome-mapped reads. Uniquely mapped reads ranged from 74.12 to 90.32 M, corresponding to a mapping rate of 82 to 84% (Table S4). A post-hoc power analysis confirmed that our study design provided 81.7% statistical power to detect twofold changes in gene expression at a significance level of α = 0.05, supporting the robustness of subsequent differential expression findings. These results demonstrate high sequencing quality and sufficient depth for comprehensive transcriptome coverage.
To assess global transcriptome differences, principal component analysis (PCA) was performed on the samples from the Control, MV and PNS groups (Fig. 2B). The PCA revealed distinct clustering of the three sample groups. Furthermore, samples within the same group exhibited similar gene expression patterns, while significant differences were observed between the different groups. These results demonstrate clear separation based on experimental condition and support the reliability of the sample grouping for downstream analyses.
Differentially expressed miRNA-seq in the diaphragm after MV and PNS treatment
A total of 25 DE miRNAs were detected between the MV and Control group, with 13 up-regulated and 12 down-regulated (Fig. 2C). There were 47 DE miRNAs between the PNS and MV group, with 28 up-regulated and 19 down-regulated. The most significantly up-regulated and the most significantly down-regulated miRNAs based on the P-values were listed in Fig. 2D. There were three DE miRNAs overlapping in the MV vs Control group and the PNS vs MV group (Fig. 2E). Only two miRNAs (miR-500-3p and miR-133b-3p) were significantly down-regulated in the MV group but significantly up-regulated in the PNS group. Consequently, miR-500-3p and miR-133b-3p were selected for further investigation in this study.
Differentially expressed circRNAs in the diaphragm after MV and PNS treatment
To identify key circRNAs involved in PNS-mediated prevention of VIDD, comprehensive circRNA profiling was performed. Analysis of the MV vs Control groups revealed 1,328 DE circRNAs, with 551 up-regulated and 777 down-regulated (Fig. 2F). Similarly, profiling of the PNS vs MV groups identified 1,408 DE circRNAs, including 868 up-regulated and 540 down-regulated species (Fig. 2G). A total of 581 circRNAs were common to both comparisons, as shown in the Venn diagram (Fig. 2H). Notably, 576 (99.1%) of these shared circRNAs exhibited opposing expression patterns between the two experimental conditions: 319 were up-regulated during MV but down-regulated following PNS treatment, whereas 257 showed the converse pattern. These findings suggest these circRNAs may be functionally relevant to PNS-induced protection against VIDD.
Differentially expressed genes in the diaphragm after MV and PNS
A total of 1,359 DE genes were identified between the MV and Control groups, comprising 598 up-regulated and 761 down-regulated genes. These DE genes were displayed in a volcano plot (Fig. 2I). Comparatively, 641 DE genes were identified between the PNS and MV groups, with 229 up-regulated and 412 down-regulated (Fig. 2J). Among these comparisons, 175 genes were found to be significantly differentially expressed in both the MV vs Control and PNS vs MV groups (Fig. 2K). Seventy-one genes exhibited significant up-regulation during MV followed by down-regulation after PNS, while forty-five genes were significantly down-regulated during MV and subsequently up-regulated after PNS (Fig. 2K). Therefore, these 116 genes represent candidate genes potentially involved in the prevention of VIDD by intravenous PNS.
KEGG pathway and GO enrichment analyses of differential genes
KEGG pathway analysis revealed that DE mRNAs between the MV and control groups were predominantly enriched in phagosome, p53 signaling, NF-κB signaling and chemokine signaling pathways (up-regulated; Fig. 3A). Conversely, DE mRNAs between the PNS and MV groups showed significant enrichment in down-regulated pathways, including cell adhesion molecules (CAMs), chemokine signaling, NF-κB signaling and phagosome (Fig. 3B). These findings imply that MV activates autophagy and systemic inflammation, while PNS suppresses these processes through CAM-mediated mechanisms. GO analysis further demonstrated that the MV vs Control comparison exhibited significant up-regulation of inflammatory response, cytokine-mediated signaling, immune response and innate immune response. In contrast, the PNS vs MV group showed down-regulation of immune response, adaptive immune response, innate immune response and cell adhesion (Fig. 3C,D). Collectively, these results indicate that PNS mitigates MV-induced diaphragm dysfunction by reversing autophagy activation and immune dysregulation, suggesting therapeutic potential for VIDD management.
Gene set enrichment analysis and gene function analysis. Differentially expressed mRNAs. Bubble chart of KEGG pathways for DE mRNA between MV vs Control group and PNS vs MV group (A, B). Bar chart for (C, D) GO functional annotation analysis including biological process (BP), cellular component (CC) and molecular function (MF).
CircRNA, miRNA and mRNA co-expression network analysis
A total of 15 target genes that could combine with miR-500-3p were predicted. Among these, three genes (HK3, ENSOCUG00000000401; SIGLEC5, ENSOCUG00000005408; RAB37, ENSOCUG00000016341) exhibited binding energy values < −20 kcal/mol. Predicted binding sites between miR-500-3p and HK3, SIGLEC5, and RAB37 were shown in Fig. 4A. Expression analysis revealed inverse correlations between miR-500-3p and its target genes (HK3, SIGLEC5, RAB37) across experimental groups. Similarly, miR-133b-3p binding sites for XDH (ENSOCUG00000001230) and L-selectin (ENSOCUG00000017432) showed energy values < −20 kcal/mol (Fig. 4B). Their expression trends were inversely correlated with miR-133b-3p in both MV vs Control and PNS vs MV comparisons.
Specific binding sites showing the complementary situation of miRNA, circRNA and mRNA predicted by miRanda. The sequences show the complementary sites of miRNA: mRNA between miR-500-3p and rabbit genes (ENSOCUG00000000401; ENSOCUG00000005408; ENSOCUG00000016341) (A). miR-133b-3p and rabbit genes (ENSOCUG00000001230; ENSOCUG00000017432) (B). Specific binding sites showing the complementary situation of miR-500-3p and five circRNAs (circRNA_03891, circRNA_12435, circRNA_12437, circRNA_24673, circRNA_14127) (C). Specific binding sites showing the complementary situation of miR-133b-3p and five circRNAs (circRNA_12435, circRNA_12437, circRNA_12463, circRNA_14942, circRNA_19744) (D). The matched base pairs and seed sequences (indicated in red) are highlighted.
Based on interaction scores, 473 circRNAs were predicted to interact with miR-500-3p and 206 circRNAs with miR-133b-3p. Among these, 20 circRNAs showed expression positively correlated with RAB37, HK3 and SIGLEC5 but inversely correlated with miR-500-3p. Seven circRNAs exhibited expression positively correlated with XDH and L-selectin but inversely correlated with miR-133b-3p. The top five predicted circRNAs with the highest binding affinity for each miRNA were selected for further analysis (Fig. 4C,D). A putative ceRNA network, integrating these predictive circRNA-miRNA and the validated miRNA-mRNA interactions, was constructed to illustrate these potential regulatory relationships (Fig. 5A).
Construction and validation of circRNAs-miRNAs-mRNA Network based on the circRNA, miRNA and RNA-seq data. Blue nodes indicate miRNAs, purple nodes indicate circRNAs and yellow nodes indicate hub genes (A). Bar graphs showing the results of sequencing for miRNA (B), mRNA (C) and circRNA (D) during MV and after PNS. Bar graphs showing the results of RT-qPCR for two miRNAs (E), two mRNAs (F) and five circRNAs (G). Luciferase assay was performed by co-transfecting wild-type or mutant-type dual-luciferase reporter genes with miR-500-3p/miR-133b-3p mimic or mimic NC in 293T cells (H). *: P < 0.05; NC: negative control; WT1: The wild‐type sequence of L-selectin; WT2: The first wild‐type sequence of RAB37; WT3: The second wild‐type sequence of RAB37.
Validation of interaction pairs of ceRNA networks
In this study, the relative expression of miR-500-3p in the sequencing results was negatively correlated with the relative expression of RAB37 and three circRNAs in both the MV and PNS groups. Similarly, the expression of miR-133b-3p was negatively correlated with L-selectin and three circRNAs (Fig. 5B–D). The results of the RT-qPCR analysis were consistent with the expression trends shown in the sequencing results (Fig. 5E–G).
To validate the interactions between miR-500-3p/miR-133b-3p and RAB37/L-selectin, we performed dual-luciferase reporter assays for each miRNA-mRNA pair. The results demonstrated that miR-500-3p significantly suppressed luciferase activity via binding to two predicted sites in RAB37. Mutation of the first binding site abolished this suppression, whereas mutation of the second site did not affect miR-500-3p mimic-induced suppression. Similarly, miR-133b-3p significantly suppressed luciferase activity when bound to the WT L-selectin sequence and this suppression was effectively abolished by mutating its predicted binding site (Fig. 5H). These findings clearly confirmed that miR-500-3p directly targets the first functional site of RAB37 and miR-133b-3p directly targets L-selectin. Unfortunately, because it is currently impossible to amplify or synthesize the specific circRNA sequences investigated in this study, we were unable to experimentally validate the predicted interactions between the circRNAs and miRNAs.
Discussion
Mechanical ventilation can induce serious diaphragmatic dysfunction characterized by atrophy and contractile impairment, which prolongs intensive care stays and elevates mortality risks. While PNS shows promise in mitigating VIDD through diaphragm preservation, its precise protective mechanisms remain unclear. This study aimed to elucidate the ceRNA network mechanisms underlying the protective effects of transvenous PNS against VIDD using integrated miRNA-Seq and RNA-Seq analyses. We constructed a ceRNA network involving two axes: miR-500-3p and miR-133b-3p, targeting RAB37 and L-selectin, respectively. Furthermore, our study suggests a putative role for specific circRNAs (circRNA_12437, circRNA_24673, circRNA_14127, circRNA_14942, and circRNA_12463) in these axes, as they were computationally predicted to sponge the aforementioned miRNAs; however, their precise functions warrant further experimental validation. These two regulatory axes may offer novel therapeutic targets for combating VIDD.
miRNAs mediate post-transcriptional gene regulation by binding target mRNAs, a mechanism crucial for biological processes like muscle cell proliferation, differentiation and regeneration. In the study, miR-500-3p displayed inverse expression trends in response to MV and PNS (Fig. 5), and a direct regulatory interaction between miR-500-3p and its target gene RAB37 was experimentally validated through dual-luciferase reporter assays (Fig. 5). Notably, miR-500-3p has been identified as a key regulator of inflammation, exerting its effects by targeting transcription factors such as YY1, FOXO1, and PPARD, as well as signaling cascades including Wnt, chemokine and cytokine pathways, all of which are associated with age-related inflammatory process30. The inflammatory role of miR-500-3p is corroborated by its up-regulated expression in CD3 + T lymphocytes of collagen-induced arthritis (CIA)-prone mice31, indicating its conserved role in immune regulation. Beyond its inflammatory functions, miR-500-3p exhibits dynamic spatiotemporal expression patterns during development. For instance, murine cardiac tissue shows significant down-regulation of miR-500-3p at postnatal day 7 compared to earlier developmental stages32, suggesting its potential role in myocardial maturation through the modulation of proliferative and apoptotic pathways. Building on its established role in developmental regulation, our study extends the functional spectrum of miR-500-3p to pathological contexts by proposing a potential mechanism through which this miRNA may modulate autophagy in VIDD. Autophagosome membrane biogenesis is actually a membrane-trafficking process and the complex ATG12-ATG5-ATG16L1 is the core machinery of autophagy. RAB37 as a small GTPase could promote formation of the ATG12-ATG5-ATG16L1 complex in a GTP-dependent manner33. In our study, we found for the first time that there is a specific regulatory relationship between an miR-500-3p and its target gene RAB37. MV down-regulated miR-500-3p, leading to up-regulated RAB37 expression that increases autophagosome formation in the diaphragm. This autophagosome formation, responsible for engulfing and degrading cellular components, is associated with muscle weakness. Our study also suggests that electrical stimulation has the effect of inhibition of RAB37-mediated autophagy through targeting miR-500-3p. This observation aligns with prior studies demonstrating that RAB37 deficiency attenuates autophagy and mitigates ventilator-induced lung injury in geranylgeranyl diphosphate synthase large subunit 1(GGPPS1) knockout mice34. This regulatory shift was associated with enhanced autophagosome activity in diaphragmatic tissue during MV, potentially contributing to muscle weakness through excessive cellular component degradation. Furthermore, our data suggest that PNS may attenuate RAB37-mediated autophagic activity via miR-500-3p pathway modulation. Although the direct connection between electrical stimulation, miR-500-3p, RAB37 and autophagy is not well-established, this study provides a foundation on electrical stimulation mediated autophagy related signaling pathways in VIDD.
In the study, we identified miR-133b-3p as another important miRNA dynamically regulated during MV and PNS interventions. Dual-luciferase reporter assays confirmed the targeting of the L-selectin by this miRNA (Fig. 5). The miR-133 family demonstrates multi-organ anti-fibrotic properties, with miR-133a suppressing bladder fibrosis via transforming growth factor-β1 (TGF-β1: a key profibrotic cytokine) inhibition35, ameliorating myocardial fibrosis in hypertrophic contexts36, and potentially protecting against hepatic fibrosis by modulating collagen genes (e.g., Col1A1, Col5A3) and immune cell infiltration37. Additionally, miR-133b-3p attenuates renal fibrosis through EMT regulation in aging kidneys38. L-selectin, a cell surface adhesion molecule expressed on leukocytes, plays a vital role in the early stages of leukocyte recruitment to inflammatory sites and the immune response. It mediates the rolling of leukocytes along the endothelium, a critical step in their extravasation and infiltration into tissues. Previous studies demonstrate that L-selectin exacerbates pulmonary fibrosis by promoting leukocyte accumulation at inflammatory sites, while its loss of expression attenuates both inflammatory responses and subsequent fibrotic progression through reduced TGF-β1 production via diminishing leukocyte recruitment and aggregation39. Our study demonstrated that the miR-133b-3p/L-selectin regulatory axis may participate in PNS-mediated prevention of VIDD. miR-133b-3p could down-regulate L-selectin expression, thus reduce the accumulation of leukocytes and the production of profibrotic cytokines at inflammatory sites within the diaphragm. These findings suggest that the interaction between miR-133b-3p and L-selectin helps maintain diaphragmatic function and prevent the development of fibrosis through modulating the diaphragmatic immune dysregulation.
Our study further elucidates the role of circRNAs within the molecular network underlying PNS-mediated prevention of VIDD. CircRNAs have been shown to modulate pathophysiological processes such as immune cell infiltration and autophagy through MREs40,41. For instance, hsa_circ_0075341 exacerbates cerebral infarction by promoting immune cell recruitment42, while circCDYL drives breast cancer progression and circATG7 facilitates pancreatic cancer metastasis through autophagy dysregulation43. In the study, miRanda prediction analysis identified putative regulatory relationships between three distinct circRNAs (circRNA_12437, 24673, 14127) and miR-500-3p, as well as between three circRNAs (circRNA_12437, 12435, 12463) and miR-133b-3p. Subsequent RT-qPCR validation revealed a negative expression correlation between these circRNAs and their corresponding miRNAs (Fig. 5). Based on the negative correlation observed in our data, we hypothesize that the five circular RNAs (circRNA_14942, 12463, 12437, 24673, and 14127) function as molecular sponges for miR-500-3p and miR-133b-3p. Through these predicted interactions, they may orchestrate bidirectional regulation in ventilator-induced diaphragmatic injury and post-PNS muscle repair by modulating the RAB37 and L-selectin axes, consequently influencing immune cell infiltration and autophagy regulation to participate in PNS-mediated prevention of VIDD.
Pathway analysis further supports that PNS appears to restore diaphragmatic homeostasis and promote muscle repair/functional recovery, potentially through modulating excessive autophagy and mitigating immune-mediated tissue damage. In the study, MV pathologically activates autophagy and systemic inflammation through coordinated up-regulation of phagosome activity, NF-κB-mediated cytokine storms and chemokine-driven immune recruitment. This activation leads to diaphragm dysfunction and muscle atrophy. However, PNS counteracts the adverse effects of MV on diaphragm function by down-regulating the pathogenic signaling pathways including cell adhesion molecules, NF-κB signaling pathway, chemokine signaling pathway, phagosome pathway, immune response, adaptive immune response, innate immune response and cell adhesion. These findings highlight the therapeutic potential of PNS in VIDD, suggesting that it may be a useful tool to restore diaphragmatic homeostasis and promote muscle repair.
Limitations and future directions
While this study establishes a PNS-mediated regulatory network in VIDD, several limitations warrant consideration. Firstly, dynamic functional analyses of target genes and miRNAs in regulating diaphragm muscle function are lacking, future research should explore the therapeutic potential of targeting the miR-500-3p/RAB37 and miR-133b-3p/L-selectin axes in preclinical models. Secondly, we did not validate diaphragm dysfunction in the animal model caused by 24-h MV through diaphragm pathology and diaphragm muscle strength measurement in our study. However, previous studies have successfully established a 24-h VIDD animal model. Thirdly, the study focuses on a single time point (24 h of MV), which may not capture long-term effects or recovery processes. Fourthly, ceRNA networks are highly complex and involve numerous RNA molecules and interactions. Understanding the full extent and function of these networks is a challenging task. To date, there is a lack of direct evidence demonstrating an interaction between ceRNA networks and electrical stimulation. Fifthly, a key limitation of our ceRNA network analysis is the inability to experimentally validate the predicted circRNA-miRNA interactions, as mature circRNAs are currently not obtainable for rabbits. This remains an important goal for future research. Sixthly, while our power analysis demonstrates sufficient sensitivity (81.7%) for detecting twofold expression changes and thus confidence in the reported differentially expressed genes, the limited sample size likely prevented the identification of more subtle transcriptional differences. Future studies are needed to further elucidate the mechanisms underlying the immunomodulatory effects of PNS and to explore its potential clinical applications in patients with VIDD. At last, optimizing PNS parameters (e.g., frequency, duration) to maximize efficacy while minimizing side effects is critical.
Conclusion
This study is the first to delineate a PNS-mediated ceRNA network in VIDD, revealing the critical regulatory roles of miR-500-3p and miR-133b-3p in modulating autophagy processes and immune response functions through their respective targets, RAB37 and L-selectin. These findings establish a theoretical framework for developing RNA-targeted therapies to prevent VIDD.
Data availability
RNA-seq data generated in this study can be found under the SRA accession number: PRJNA1288756. Small RNA-seq generated in this study can be found under the SRA accession number: PRJNA1290043.
References
Li, L. F. et al. Reduction in ventilation-induced diaphragmatic mitochondrial injury through hypoxia-inducible factor 1α in a murine endotoxemia model. Int. J. Mol. Sci. 23, 1083 (2022).
Goligher, E. C. et al. Mechanical ventilation-induced diaphragm atrophy strongly impacts clinical outcomes. Am. J. Respir. Crit. Care Med. 197, 204–213 (2018).
Toledo, T. G. & Bacci, M. R. Value of diaphragm ultrasonography for extubation: a single-blinded randomized clinical trial. Crit. Care Res. Prac. 23, 8403971 (2023).
Zhou, X. L., Wei, X. J., Li, S. P., Ma, H. L. & Zhao, Y. Lung-protective ventilation worsens ventilator-induced diaphragm atrophy and weakness. Respir. Res. 21, 16 (2020).
Hyatt, H. W. et al. Calpains play an essential role in mechanical ventilation-induced diaphragmatic weakness and mitochondrial dysfunction. Redox Biol. 38, 101802 (2021).
Triolo, M. & Hood, D. A. Manifestations of age on autophagy, mitophagy and lysosomes in skeletal muscle. Cells 10, 1054 (2021).
Fu, W. et al. Ventilator-induced diaphragmatic dysfunction: pathophysiology, monitoring and advances in potential treatment and prevention. Eur. Respir. Rev. 34, 250069 (2025).
Xing, R. et al. Identification of key genes affecting ventilator-induced diaphragmatic dysfunction in diabetic mice. Front. Genet. 15, 1387688 (2024).
Li, L. et al. Single-nucleus transcriptomic profiling of the diaphragm during mechanical ventilation. Sci. Rep. 14, 1–17 (2024).
Morris, I. S., Bassi, T., Oosthuysen, C. & Goligher, E. C. Phrenic nerve stimulation for acute respiratory failure. Respir. Care 68, 1736–1747 (2023).
Yang, M. et al. Phrenic nerve stimulation protects against mechanical ventilation-induced diaphragm dysfunction in rats. Muscle Nerve 48, 958–962 (2013).
Fernandez, K. C. et al. Transvenous stimulation yields exposure-dependent protection from ventilator-induced diaphragm atrophy. J. Appl. Physiol. 135, 15–25 (2023).
Iranpour, Y. & Zandifar, A. Diaphragm muscle parameters as a predictive tool for weaning critically ill patients from mechanical ventilation: a systematic review and meta-analysis study. Eur. J. Transl. Myol. 34, 12642 (2024).
Martin, A. D. et al. Effect of intermittent phrenic nerve stimulation during cardiothoracic surgery on mitochondrial respiration in the human diaphragm. Crit. Care Med. 42, e152 (2014).
Tang, H. B. & Shrager, J. B. The signaling network resulting in ventilator-induced diaphragm dysfunction. Am. J. Respir. Cell Mol. Biol. 59, 417–427 (2018).
Das, A., Shyamal, S., Sinha, T., Mishra, S. S. & Panda, A. C. Identification of potential circRNA-microRNA-mRNA regulatory network in skeletal muscle. Front. Mol. Biosci. 8, 762185 (2021).
Wei, L. et al. Identification of key pathways and RNAs associated with skeletal muscle atrophy after spinal cord injury. J. Musculoskel. Neuron 21, 550–559 (2021).
Wang, W. L. et al. LncRNA Xist acts as a miR-486–5p sponge to modulate myoblast proliferation by recruiting Stk4. J. Muscle Res. Cell M 1, 1–14 (2025).
Liu, D. S. et al. The novel regulatory role of lncRNA-miRNA-mRNA axis in amyotrophic lateral sclerosis: an integrated bioinformatics analysis. Comput. Math. Method M 20, 5526179 (2021).
Chen, R. et al. CircTmeff1 promotes muscle atrophy by interacting with TDP-43 and encoding a novel TMEFF1–339aa protein. Adv. Sci. 10, 2206732 (2023).
Moroz, N. et al. Oxidants regulated diaphragm proteolysis during mechanical ventilation in rats. Anesthesiology 131, 605–618 (2019).
Zhou, Z. X. et al. Circular RNA cVIM promotes hepatic stellate cell activation in liver fibrosis via miR-122–5p/miR-9–5p-mediated TGF-β signaling cascade. Commun. Biol. 7, 113 (2024).
Liu, R. N. et al. Transcriptome profiling of the diaphragm in a controlled mechanical ventilation model reveals key genes involved in ventilator-induced diaphragmatic dysfunction. BMC Genomics 22, 472 (2021).
Zhang, D., Hao, W. Y., Li, X. J., Han, P. Y. & Niu, Q. Aldh1a1 and Scl25a30 in diaphragmatic dysfunction. Exp. Biol. Med. 247, 1013–1029 (2022).
Nakai, H. Electrical stimulation mitigates muscle degradation shift in gene expressions during 12-h mechanical ventilation. Sci. Rep. 13, 20136 (2023).
Langmead, B. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Lee, E. K. et al. The involvement of serum exosomal miR-500-3p and miR-770-3p in aging: modulation by calorie restriction. Oncotarget 9, 5578–5587 (2018).
Donate, P. B. et al. T cell post-transcriptional miRNA-mRNA interaction networks identify targets associated with susceptibility/resistance to collagen-induced arthritis. PLoS ONE 8, e54803 (2013).
Liu, H. L. et al. Identification of the microRNA expression profile in the regenerative neonatal mouse heart by deep sequencing. Cell Biochem. Biophys. 70, 635–642 (2014).
Song, Y., Shang, D. T., Cheng, H. H. & Zhou, R. J. The small GTPase RAB37 functions as an organizer for autophagosome biogenesis. Autophagy 14, 727–729 (2018).
Wang, Z. X. et al. Knockout of GGPPS1 restrains rab37-mediated autophagy in response to ventilator-induced lung injury. Hum. Cell 35, 871–884 (2022).
Duan, L. J. et al. miR-133 modulates TGF-β1-induced bladder smooth muscle cell hypertrophic and fibrotic response: implication for a role of microRNA in bladder wall remodeling caused by bladder outlet obstruction. Cell. Signal. 27, 215–227 (2015).
Matkovich, S. J. et al. MicroRNA-133a protects against myocardial fibrosis and modulates electrical repolarization without affecting hypertrophy in pressure-overloaded adult hearts. Circ. Res. 106, 166-U340 (2010).
Roderburg, C. et al. miR-133a mediates TGF-β-dependent derepression of collagen synthesis in hepatic stellate cells during liver fibrosis. J. Hepatol. 58, 736–742 (2013).
Wang, Y. et al. Differentially expressed microRNAs in bone marrow mesenchymal stem cell-derived microvesicles in young and older rats and their effect on tumor growth factor-β1-mediated epithelial-mesenchymal transition in HK2 cells. Stem Cell Res. Therapy 6, 185 (2015).
Zhang, X., Zhang, W., Chen, X. & Cai, Y. Prevention of bleomycin-induced pulmonary inflammation and fibrosis in mice by bilobalide. Evid. Based Complement. Alternat. Med. 23, 1973163 (2023).
Zhang, X., Wu, H., Hong, X., Xiao, Y. & Zeng, X. Circular RNA: from non-coding regulators to functional protein encoders. Pharm. Sci. Adv. 3, 100085 (2025).
Peng, L., Yang, C., Chen, Y. F. & Liu, W. Predicting CircRNA-disease associations via feature convolution learning with heterogeneous graph attention network. IEEE J. Biomed. Health Inform. 27, 3072–3082 (2023).
Ying, K., Chen, J., Fu, Z. H. & Ren, B. FAS-mediated circRNA-miRNA-mRNA crosstalk network regulates immune cell infiltration in cerebral infarction. J. Mol. Neurosci. 73, 117–128 (2023).
Liang, G. H. et al. Autophagy-associated circRNA circCDYL augments autophagy and promotes breast cancer progression. Mol. Cancer 19, 65 (2020).
Acknowledgements
We thank OE Biotech Co., Ltd. for providing the small RNA sequencing and whole RNA sequencing. We thank Figdraw (www.figdraw.com) for assistance in the pattern drawing.
Funding
This work was supported by Fundamental Research Program of Shanxi Province (202303021221245).
Author information
Authors and Affiliations
Contributions
Dong Zhang, Lei Li, Fangyuan Li and Min Wang performed the experiments. Xuejiao Duan interpreted the data. Mingming Zhang, Dong Zhang wrote the manuscript and Mingming Zhang prepared the figures. Wenyan Hao designed the pulse generator. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics declarations
All animal care and experimental procedures associated with the research were approved by the Institutional Animal Care and Use Committees of Changzhi Medical College, and all experiments were performed in accordance with ARRIVE guidelines.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, M., Li, F., Wang, M. et al. Identification of competing endogenous RNA networks involved in phrenic nerve stimulation preventing mechanical ventilation induced diaphragm dysfunction. Sci Rep 16, 2065 (2026). https://doi.org/10.1038/s41598-025-31787-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-31787-4