Abstract
Immune rejection poses a major challenge in organ transplantation, with tissue-resident memory T (TRM) cells playing a critical role in graft rejection. This study investigated the impact of CCR8 on TRM cells using single-cell RNA sequencing (scRNA-seq), flow cytometry, and immunohistochemistry. The results show that CCR8 expression was upregulated on CD8⁺ TRM cells after transplantation, peaking on day 7. Blocking or knocking out CCR8, as well as neutralizing CCL1 and CCL8, significantly reduced CD8⁺ TRM cell accumulation in the graft and their cytokine production. These treatments prolonged graft survival, alleviated rejection severity, and impaired the ability of CD8⁺ TRM cells to produce GZMB, IFN-γ, and IL-2. Single-cell analysis of skin transplantation revealed that loss of CCR8 disrupted macrophage–T cell interactions, particularly CD8⁺ TRM–macrophage crosstalk, while enhancing CD40, PD-L1, and NRG signaling. These findings suggest that targeting CCR8 to limit the accumulation and function of CD8⁺ TRM cells may be an effective strategy to alleviate transplant rejection.
Similar content being viewed by others
Introduction
Organ transplantation is a life-saving intervention for end-stage organ failure1. However, the long-term survival of grafts is limited by immune-mediated rejection. Therefore, understanding the mechanisms of rejection and inducing immune tolerance is crucial for improving transplant outcomes. Allogeneic transplantation immunity is a multifaceted process, resulting from interactions between various immune cells, including lymphocytes, macrophages, monocytes, and dendritic cells (DC), with T cells being the central mediator of allogeneic transplant rejection. T cells recognize polymorphisms in donor minor histocompatibility and major histocompatibility complex proteins2,3. Despite significant progress in organ transplantation due to the application of immunosuppressive agents, long-term outcomes remain disappointing4. Therefore, inducing immune tolerance to grafts remains a challenging issue.
Tissue-resident memory T (TRM) cells are a specialized subset of immune cells found in non-lymphoid tissues. Unlike circulating memory T cells, TRM possess unique homing and functional characteristics. They are defined by the expression of activation, retention, and adhesion markers, the downregulation of cell migration molecules such as S1PR1, CCR7, and CD62L to prevent egress, and the promotion of active retention through receptors and integrins (including CD69, CD49a, and CD103) binding to extracellular matrix components5,6,7,8. TRM can also produce effector cytokines, such as GZMB, IFN-γ, and local proliferation9,10,11. They play a critical role in adaptive immune responses, providing local and long-term protection through continuous tissue surveillance in non-lymphoid tissues6,12,13,14, and are crucial in transplant rejection of solid organs15,16,17. TRM influence long-term survival in heart18, kidney19, liver9, and lung transplants20, leading to the occurrence of rejection. Thus, targeting and reducing CD8+ TRM in the graft to alleviate transplant rejection is an attractive strategy.
Recent studies have greatly enhanced our understanding of chemokine-mediated leukocyte recruitment and responses, including differentiation, activation21, apoptosis22, development23, and angiogenesis24. The chemokine receptor CCR8 is considered a key chemokine receptor for TRM cells. CCR8 is expressed in about 50% of TRM cells, but rarely in central memory T cells (TCM)25. In mice, the known chemokine ligands for CCR8 are CCL1 and CCL826,27, while in humans, the ligands are CCL1 and CCL1828. In pulmonary fibrosis, blocking CCL18-CCR8 signaling can inhibit the accumulation of CD8+ TRM cells and alleviate fibrosis29. However, it is not yet known whether blocking CCR8 in organ transplants can reduce TRM cells in the graft and thereby reduce transplant rejection.
Here, we investigate the role of CCR8 in CD8⁺ TRM cells during transplantation. By integrating scRNA-seq data from mouse skin grafts, we identify CCR8 as a key molecule highly expressed in recipient-derived CD8⁺ TRM cells. Our findings suggest that CCR8 plays a critical role in regulating CD8⁺ TRM cell function in transplantation.
Methods
Ethics statement
The studies were reviewed and approved by the ethics committee of the Qingdao University (No.20241205C5710020251205158). The housing and experimental procedures of the mice were conducted in accordance with the Principles of Laboratory Animal Care and the ARRIVE guidelines.
Mice
C57BL/6J (CD45.1+ or CD45.2+), BALB/c, and CCR8 knockout (CCR8KO) mice were purchased from GemPharmatech. All mice were housed in the SPF-grade animal facility at Qingdao University. Male mice aged 7–8 weeks were used in all experiments.
Skin transplantation and histology
Full-thickness dorsal skin (~ 1 cm × 1 cm) was harvested from BALB/c donor mice. Hair was removed from the backs of recipient mice, and a square piece of skin (~ 1.2 cm × 1.2 cm) was excised to create the graft bed. The grafts were secured to the recipient bed using at least eight interrupted 7 − 0 nylon sutures. On the day of transplantation, the graft site was covered with an elastic bandage, which was removed on day 6 post-transplantation. Sutures were removed on day 7. Two personnel monitored the grafts daily for signs of rejection. Grafts were considered rejected when more than 80% of the graft area became necrotic.
Anti-CCL1 and Anti-CCL8 antibody treatments
Mice were treated with anti-CCL1 monoclonal antibody (R&D; MAB845; 50 µg per intraperitoneal injection) and anti-CCL8 monoclonal antibody (R&D; MAB790; 50 µg per intraperitoneal injection) on days 0, 2, 4, and 6 post-transplantation. Mice were euthanized for tissue collection on day 7.
Measurement of cytokine levels
Blood was drawn from the choroid plexus of the posterior eyeball of mice and collected in sterile tubes. The serum was stored at -80 °C. All serum samples were analyzed within 1–2 months after collection. According to the manufacturer’s instructions, a panel of 31 mouse cytokines (LXR-MultiDTM-31, LabEx Co., Shanghai, China) was used to determine serum cytokine levels. The Luminex X-200 system (Luminex) and Milliplex Analyst software (version 5.1) were employed for the detection and data analysis of 36 mouse cytokines30.
Tissue processing
Mice were anesthetized with 1% pentobarbital sodium (60 mg/kg, intraperitoneal injection) prior to the procedures, and blood was collected via enucleation. Blood samples were anticoagulated with EDTA and centrifuged at 3000 rpm for 20 min. Plasma was collected, and the remaining blood was resuspended to its original volume using whole blood and tissue dilution buffer. Red blood cells were lysed with red blood cell lysis buffer for 2 min, neutralized with PBS, and centrifuged at 1500 × g for 5 min. The supernatant was discarded, and the pellet was retained.
Skin-draining lymph nodes (axillary, brachial, and inguinal) and spleens were ground and filtered through a 70 μm cell strainer, followed by a 40 μm cell strainer and centrifugation. The spleen underwent one round of red blood cell lysis as described above.
Transplanted skin was excised, washed three times in pre-cooled PBS, and then cut into ~ 2 mm fragments. These were digested in a mixture containing 1 mg/mL Collagenase IV (Glpbio, GC19591), 3.5 mg/mL Collagenase II (Glpbio, GC19588), 0.04 mg/mL DNase I (Glpbio, GC19804), and 0.002 mg/mL hyaluronidase (Glpbio, GC36265). Samples were incubated on a shaker at 37 °C and 250 rpm for 60 min, then filtered through 70 μm and 40 μm cell strainers and centrifuged.
Flow cytometry staining
Single-cell suspensions were washed with PBS and blocked with 5% bovine serum albumin (BSA) for 15 min. Viability was assessed using a fixable live/dead dye before surface staining. Cells were stained with fluorochrome-conjugated antibodies at 4 °C for 30 min.
For intracellular cytokine staining, cells were stimulated with phorbol 12-myristate 13-acetate (PMA), ionomycin, and brefeldin A (BioLegend, 423304) for 5–6 h at 37 °C in a humidified 5% CO₂ incubator using DMEM/F-12 medium (ViyaCell, C3130-0500) supplemented with 10% fetal bovine serum (FBS). Surface staining was performed at room temperature for 30 min. Intracellular staining was conducted using the Intracellular Fixation & Permeabilization Buffer Set (eBioscience) according to the manufacturer’s instructions.
Data acquisition was performed on a CytoFLEX S flow cytometer (Beckman Coulter) and URIT Flow Cytometer, BF-730 B5R3V6, URIT Medical Electronic Co.,Ltd.,Guilin China, and results were analyzed using FlowJo software (version 10.8.1).
The following fluorochrome-conjugated antibodies (all from BioLegend) were used: CD3-PerCP/Cyanine5.5 (17A2), CD3-FITC (17A2), CD4-FITC (RM4-5), CD8-PE/Cyanine7 (53 − 6.7), CD8-PC7 (53 − 6.7), CD25-APC (PC61), CD69-PE (H1.2F3), CD103-APC (2E7), PD-1-ECD (29 F.1A12), PD-1-PC7 (29 F.1A12), CTLA-4-BV610 (UC10-4B9), TIGIT-PB450 (1G9), FOXP3-PE (150D), CCR8-APC-A750 (SA214G2), Granzyme B-PB450 (QA18A28), IL-2-BV610 (JES6-5H4), IFN-γ-BV660 (XMG1.2), CD11c-APC (N418), CD11b-PB450 (M1/70), F4/80-BV610 (BM8), NK1.1-PE (PK136), CD45-PC5.5 (30-F11), CD45.1-PB450 (A20), CD45.2-PerCP (104), CD19-PC7 (6D5).
Tissue histology and Immunofluorescence
The skin grafts were harvested at the indicated time points. Sections (4 to 6 mm) were fixed in 4% paraformaldehyde and stained with hematoxylin and eosin (H&E) and immunohistochemistry (IHC) using antibodies against CD3, CD4, CD8, CD69, CD103, FOXP3 and CCR8. A blinded pathologist performed the analysis of H&E-stained sections based on the following criteria: 1 point-folliculitis->25% of all hair follicles have moderate or severe lymphocytic infiltrates; 1 point-dermal inflammation-dense aggregates of lymphocytes in a band-like distribution or multiple patchy distributions; 1 point-epidermal infiltration-basilar and epidermal damage or necrosis for a total of 3 points. Some slides were analyzed and scored for the degree of CD4/CD8 infiltration according to the following criteria: 1 point-scattered, increased dermal positive staining; 2 points-mild, scattered, increased dermal staining with focal infiltration into the dermis and hair follicles; 3 points-marked, scattered positive staining, and dermal and hair follicle infiltration31.
Multiplex Immunofluorescence staining
Tissue sections were first deparaffinized and rehydrated. Antigen retrieval was then performed by heating the slides in Tris-EDTA buffer at pH 6.0 or pH 9.0. Endogenous peroxidase activity was quenched with 3% hydrogen peroxide (H₂O₂), and nonspecific binding was blocked using 3% bovine serum albumin (BSA). The slides were subsequently incubated with primary antibodies against CD4, CD8A, CD69, CD103, CCR8, and FOXP3. Signal detection was carried out using a TSA kit (Nanjing Freethinking Biotechnology Co., Ltd., China). Nuclei were counterstained with DAPI. Finally, the stained slides were scanned using a Pannoramic MIDI scanner (3DHISTECH, Hungary), and image analysis was performed using the HALO 2.0 Area Quantification algorithm (Indica Labs, Corrales, NM) at Nanjing Freethinking Biotechnology Co., Ltd. (China)9.
Library preparation and sequencing
Single-cell RNA sequencing (scRNA-seq) libraries were prepared using the DNBelab C Series High-throughput Single-cell System (BGI-research). Single-cell suspensions were converted into barcoded libraries through a workflow involving droplet encapsulation, emulsion breaking, mRNA capture on beads, reverse transcription, and subsequent cDNA amplification and purification. The amplified cDNA was then fragmented to a target size of 250–400 base pairs. Indexed libraries were constructed according to the manufacturer’s protocol. Library quality was assessed using the Qubit ssDNA Assay Kit (Thermo Fisher Scientific) and the Agilent Bioanalyzer 2100 system. Sequencing was performed on the DIPSEQ T7 platform (China National GeneBank) using paired-end mode. The sequencing read structure included a 30 bp Read 1 (comprising two 10-bp cell barcodes and a 10-bp UMI), a 100 bp Read 2 for transcriptome profiling, and an additional 10-bp read for sample indexing9.
Single-cell RNA sequencing data pre-processing
Raw sequencing data were processed using an open-source pipeline provided by MGI (https://github.com/MGI-tech-bioinformatics/DNBelab_C_Series_HT_scRNA-analysis-software). The workflow included sample demultiplexing, barcode parsing, and quantification of 3′ unique molecular identifiers (UMIs) per cell, all performed using default pipeline settings. Reads were aligned to the Mus musculus reference genome (GRCm38). Cell quality was assessed based on UMI count distribution, applying the barcodeRanks function from the DropletUtils package to distinguish valid cells from ambient RNA or bead-associated noise. Low-quality barcodes falling below the UMI threshold were excluded. Gene expression quantification was then performed using the PISA framework, resulting in gene-by-cell count matrices for downstream analysis.
Single-cell gene expression analysis and subcluster identification
Single-cell RNA sequencing data were processed using the Seurat R package (version 4.3.0)32 according to our previous studies33,34,35. Initial steps included importing the raw gene expression matrix and implementing strict quality control criteria. Cells expressing fewer than 501 genes or containing over 25% mitochondrial transcripts were excluded to eliminate low-quality profiles. Remaining high-quality cells were subsequently normalized and scaled using Seurat’s default workflow.
To capture biological heterogeneity, highly variable genes were identified via the FindVariableFeatures function, serving as the basis for principal component analysis (PCA) on the scaled dataset. Dimensionality reduction and clustering were then performed using FindNeighbors (based on the top 10 principal components) and FindClusters (with a resolution of 0.5). Finally, uniform manifold approximation and projection (UMAP) was applied to visualize the cellular landscape and delineate transcriptionally distinct subclusters.
Cell type annotation
To characterize distinct cell populations, we identified cluster-specific gene expression patterns using the FindMarkers and FindAllMarkers functions. Cell type assignment was conducted based on canonical marker genes corresponding to established biological identities. Marker selection was informed by the CellMarker database (http://bio-bigdata.hrbmu.edu.cn/CellMarker/) and validated through reference to peer-reviewed literature. For enhanced accuracy, we additionally utilized the SingleR package (version 2.0.0)36, which aligns individual cell transcriptomes with annotated reference datasets to refine cell type classification.
Functional enrichment analysis
Following cell type annotation, functional enrichment analysis was performed to explore the biological roles and signaling pathways associated with differentially expressed genes across cell clusters. Gene Ontology (GO) terms and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways were evaluated using the clusterProfiler package (version 3.17.0)37,38,39,40 in conjunction with org.Hs.eg.db (version 3.11.4). A significance threshold of p < 0.05 was applied to both analyses to ensure statistical robustness. The top enriched GO and KEGG terms for each cell type were selected and visualized using bar plots or dot plots, providing an intuitive overview of key biological processes and molecular functions characterizing each cellular subset.
Pseudotime analysis
Trajectory analysis was performed using the Monocle package (version 2.28.0) to trace the developmental progression of cells along a pseudo-temporal continuum. The analysis included various T cell subsets, such as TRM and Teff cells. For each cell group, parameters were defined as follows: a lower detection limit of 0.5, a minimum expression threshold of 0.1, and expression in at least 10 cells. Cell trajectories were visualized using the plot_cell_trajectory function, which mapped potential developmental pathways based on pseudotime, Seurat cluster assignments, and associated metadata. This approach provided a dynamic depiction of cellular differentiation and maturation processes.
Cell-cell communication analysis
To elucidate intercellular signaling dynamics, we applied the CellChat package (version 1.6.1)41, which infers cell–cell communication based on curated ligand–receptor interactions, including those from the KEGG signaling pathway database and recent experimental datasets. The analytical workflow began with the identification of differentially expressed signaling genes, enabling the detection of key interaction candidates. Subsequently, average expression levels of ligand–receptor pairs were computed across cell types to profile communication patterns. Finally, the probability of cell–cell interactions was inferred to reconstruct the global intercellular communication network, offering insights into the regulatory crosstalk among distinct cellular populations.
Statistical analysis
All statistical analyses and data visualizations were performed using R software (version 4.3.2) and GraphPad Prism 8. The data are presented as mean ± standard error of the mean (SEM). Statistical significance between two groups was assessed using an unpaired Student’s t test. Statistical differences among multiple groups were evaluated using one-way analysis of variance (ANOVA). Statistical significance was considered at P < 0.05. This threshold was applied to ensure robust and interpretable comparisons within the designated confidence level.
Results
TRM cells are enriched in rejecting allografts and secondary lymphoid organs
To assess the role of TRM cells in transplant rejection, we established a murine skin graft model using either syngeneic C57BL/6 (B6) or allogeneic BALB/c donors (Fig. 1A). Tail skin was grafted onto B6 recipients (n = 10 per group), and tissue samples, including skin, spleen, blood, lymph nodes, liver and kidney, were collected on day 7 post-transplantation. Graft survival was significantly reduced in the allogeneic group compared to syngeneic controls, as shown by survival curves and statistical analysis (Fig. 1B). Macroscopic observation showed visible rejection in BALB/c→B6 grafts (median survival times [MST] 16 days posttransplant; P < 0.0001) beginning around day 9 (Fig. 1C). Histological examination revealed dense mononuclear infiltration in allogeneic grafts, while liver and kidney tissues remained histologically unremarkable (Fig. 1D, Fig. S1G). Immunohistochemical staining confirmed increased CD3⁺, CD4⁺, and CD8⁺ T cell infiltration in allografts (Fig. 1D).
TRM cells are enriched in rejecting allografts and secondary lymphoid organs. (A) The skin graft from BALB/c mice was transplanted to the back of C57 mice, and on day seven, the graft, spleen, peripheral blood, and draining lymph nodes were harvested. (B) The graft survival time after syngeneic(n = 6) and allogeneic(n = 6) skin transplantation. (C) Photos of C57 and BALB/c skin grafts in C57 recipient mice. (D) Hematoxylin and Eosin (HE), CD3, CD4, and CD8 immunohistochemistry of syngeneic and allogeneic grafts. (E) Flow cytometric analysis of changes in CD8+TRM cells in syngeneic and allogeneic grafts. (F) Immunohistochemistry of CD69 and CD103 in syngeneic and allogeneic grafts. Changes in immune checkpoint molecules PD-1 (G), TIGIT (H), and CTLA-4 (I) on CD8⁺ TRM cells in the grafts. Statistical analysis was performed using the t-test to compare the differences between groups. Bars represent mean ± SD from 10 independent experiments. Statistical significance is indicated by *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001; “ns” denotes no significant difference.
Flow cytometric analysis demonstrated a significant increase in the proportion of CD69⁺CD103⁻ TRM (P < 0.0001) within CD8⁺ T cells in allogeneic grafts, whereas the frequencies of CD69⁺CD103⁺ TRM (P = 0.51) within CD8⁺ T cells and both CD69⁺CD103⁺ (P = 0.09) and CD69⁺CD103⁻ TRM (P = 0.7) within CD4⁺ T cells remained unchanged. In the draining lymph nodes, both CD69⁺CD103⁺ (P = 0.016) and CD69⁺CD103⁻ TRM (P < 0.0001) subsets within CD8⁺ T cells were significantly increased, while in the spleen, the alterations in CD8⁺ TRM mirrored those observed in the grafts (Fig. 1E, Fig. S1I, H). Immunohistochemistry also showed elevated expression of TRM-associated markers CD69 (P = 0.0025) and CD103 (P = 0.0243) in allogeneic skin (Fig. 1F).
To assess functional activation, we measured expression of immune checkpoints on CD8⁺ TRM cells. Cells from allogeneic recipients expressed significantly higher levels of PD-1 (P < 0.0001), TIGIT (P < 0.0001), and CTLA-4 (P < 0.0001) in the skin, lymph nodes, and spleen (Fig. 1G-I, Fig. S1J-L), indicating activation or potential exhaustion.
Treg cells, as immune suppressive cells, can inhibit the activation and proliferation of CD8+ T cells, playing a crucial role in transplant immune rejection42. Additionally, Tregs (P < 0.0001) were expanded in lymph nodes, spleen, and peripheral blood of allograft recipients (Fig. S1B, C). Notably, TIGIT (P < 0.0001) expression on Tregs was significantly increased, while PD-1(P < 0.0001) expression was reduced (Fig. S1D-F), suggesting a functional shift in Tregs under allogeneic challenge.
Single-cell RNA sequencing reveals immune cell subpopulations and signaling differences between syngeneic and allogeneic grafts
To further investigate the immune cell heterogeneity between syngeneic and allogeneic grafts, we performed an analysis using publicly available scRNA-seq datasets43. Unsupervised clustering followed by t-SNE visualization identified 14 distinct immune cell clusters (Fig. 2A). Based on the expression levels of characteristic genes, these clusters were annotated as major immune cell types, including T cells, B cells, macrophages, granulocytes, and natural killer (NK) cells (Fig. 2B). The expression of marker genes in each immune cell type was visualized, confirming the accuracy of the cell type classification (Fig. 2C).
Single-cell RNA sequencing reveals immune cell subpopulations and signaling differences between syngeneic and allogeneic grafts. (A) t-SNE plot showing the clustering of immune cell populations based on single-cell RNA sequencing data. Each cluster is represented by a different color, with cell populations identified by their respective markers. (B) t-SNE plot highlighting the main immune cell subsets: B cells, T cells, NK cells, macrophages, and granulocytes. (C) Dot plot showing the expression of key genes in different immune cell populations. The size of the dots represents the percentage of cells expressing the gene, and the color indicates the level of gene expression. (D) t-SNE plot depicting the distribution of T cell subsets including CD4⁺ and CD8⁺ T cells, and their various functional states. (E) t-SNE plot detailing the different CD4⁺ and CD8⁺ T cell subsets, including CD4⁺ TRM cells, CD8⁺ cytotoxic cells, and CD8⁺ TRM cells. (F) Dot plot showing the expression of key transcription factors and surface markers across different T cell populations, highlighting the differential expression in subsets like CD8⁺ TRM and other T cell types. (G) Heatmap illustrating the interaction strength between Ccr7 and Ccr8 in different T cell subsets, with higher values indicating stronger interactions. (H) Bar chart comparing the interaction strength of immune receptors between syngeneic and allogeneic conditions. (I) Sankey plot illustrating the relative information flow between key genes, comparing the distribution of expression in syngeneic and allogeneic conditions.
Further subclustering of the T cell population revealed several subsets, including CD4⁺ conventional T cells (Tconv), Treg, CD4⁺ PD-1⁺ T cells, CD8⁺ T cells, and CD8⁺ TRM (Fig. 2D, E). The specific expression of key marker genes (CD4, CD8a, Foxp3, Pdcd1) in these subsets further validated their classification accuracy (Fig. 2F). Heatmap analysis demonstrated differential expression of chemokine receptors among the T cell subsets, with circulating T cells showing high expression of Ccr7, whereas TRM-like cells exhibited significantly higher expression of Ccr8 (Fig. 2G). These findings suggest that Ccr8 may serve as a key marker for identifying TRM cells within grafts.
Immunohistochemical staining validated the potential contribution of TRM cells to graft rejection in skin transplants. In the allogeneic group, the number of CCR8⁺ cells within the graft were significantly increased, particularly in the inflammatory regions of the transplant area (Fig. S2B). This observation suggests that CCR8⁺ TRM cells may play a critical role in the immune rejection of allogeneic skin grafts.
To assess differences in immune cell interactions between the syngeneic and allogeneic groups, we performed cell-cell interaction analysis using a cell communication tool. The total cell interaction intensity in the allogeneic group was significantly higher than in the syngeneic group (Fig. 2H), indicating more active immune cell communication in the context of allogeneic transplantation. Notably, the interactions between TRM cells and other T cells (CD8⁺ T cells), macrophages, and NK cells were particularly pronounced (Fig. S2C).
Further analysis of relative signaling flow across various pathways revealed heightened signaling activity in multiple immune-regulatory pathways in the allogeneic group, suggesting stronger immune signaling in the allogeneic immune context. Additionally, the analysis of ligand-receptor signaling pathways indicated significantly enhanced relative information flow in the allogeneic group, particularly in pathways involved in chemotaxis, costimulation, and T cell retention (Fig. 2I). These ligand-receptor interactions are likely crucial for the rejection response mediated by CCR8⁺ TRM cells.
In conclusion, single-cell transcriptomic analysis revealed highly activated immune cell populations with functional specialization in allogeneic grafts. The enrichment of CCR8⁺ TRM cells and enhanced intercellular signaling could play a pivotal role in mediating rejection responses.
Recipient-derived CD8⁺ TRM cells accumulate in allografts and express CCR8
To delineate the origin and dynamics of CD8⁺ TRM cells following transplantation, we employed an allogenic skin graft model using CD45.2⁺ BALB/c mice as donors and CD45.1⁺ C57BL/6 mice as recipients. Skin grafts, dLNs, spleens, and peripheral blood were collected on days 3, 5, 7, and 9 post-transplantation for flow cytometric analysis (Fig. 3A, B).
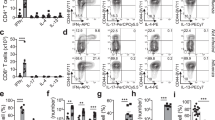
Recipient-derived CD8⁺ TRM cells accumulate in allografts and express CCR8. (A) BALB/c mice (CD45.2) serve as skin donors for C57BL/6 mice (CD45.1) recipients. Grafts, lymph nodes, and spleen were collected on days 3, 5, 7, and 9 post-transplant. (B) Representative flow cytometry plots showing the distribution of CD45.1 and CD45.2 cells in grafts on days 3, 5, 7, and 9 post-transplant, gated on live cells. (C) Quantification of the percentage of CD45.1 and CD45.2 (D) cells in the skin, lymph nodes, spleen, and blood at the indicated time points. (E) Quantification of CD8⁺ TRM cell percentages (CD45.1⁺CD8⁺CD69⁺CD103⁺) in skin over time post-transplant. (F) Quantification of CCR8 expression on CD45.1⁺CD8⁺CD69⁺CD103⁺ cells in the skin at different time points post-transplant. Statistical analysis was performed using the t-test to compare the differences between groups. Bars represent mean ± SD from 8 independent experiments. Statistical significance is indicated by *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001; “ns” denotes no significant difference.
Recipient-derived immune cells progressively infiltrated the grafted skin, peaking at day 7 before declining. In contrast, donor-derived immune cells remained scarce in the skin throughout. In dLNs, spleen, and peripheral blood, recipient-derived cells predominated across all time points, whereas donor-derived cells transiently decreased and reached a minimum at day 5 (Fig. 3C, D, Fig. S3B).
Analysis of T cell subsets revealed that donor-derived CD4⁺ and CD8⁺ T cells were virtually absent in the skin, while recipient-derived counterparts increased steadily. In secondary lymphoid organs and blood, recipient-derived CD4⁺ and CD8⁺ T cells remained dominant, with minimal contribution from donor cells (Fig. S3C, D).
Focusing on CD8⁺ TRM cells, we observed a marked expansion of recipient-derived CD8⁺ TRM (P = 0.004) in the graft, peaking on day 7. Donor-derived CD8⁺ TRM cells also transiently accumulated in dLNs and spleen, reaching peak levels at day 7, but remained a minority compared to recipient-derived populations (Fig. 3E, Fig. S3E).
Notably, CCR8 expression was nearly undetectable on donor-derived CD8⁺ TRM cells, but was progressively upregulated on recipient-derived CD8⁺ TRM cells, with maximal expression observed at day 7 (P = 0.0006) (Fig. 3F, Fig. S3F). These findings suggest that recipient-derived CD8⁺ TRM cells are preferentially recruited to the graft via CCR8-dependent mechanisms and may contribute to local alloimmune responses.
CCR8 inhibition prolongs graft survival and attenuates CD8⁺ TRM activation
To evaluate the impact of CCR8 blockade on transplant rejection, mice receiving allogeneic skin grafts were treated intraperitoneally with the CCR8 inhibitor R243 (1 mg/kg) on days 0, 2, 4, and 6 post-transplantation. Graft tissues, dLNs, spleens, and peripheral blood were collected on day 7, and graft necrosis was monitored daily until complete graft detachment (Fig. 4A). R243 treatment (MST 12.5 vs. 23.5 days posttransplant, P < 0.0001) significantly prolonged graft survival compared to vehicle-treated controls (Fig. 4B, C).
CCR8 inhibition prolongs graft survival and attenuates CD8⁺ TRM activation. (A) Mice were treated with R243 (1 mg/kg) on days 0, 2, 4, and 6 post-transplantation. Grafts and lymph nodes, spleen, and blood were harvested at days 7, 9, 11, and 13 post-transplant. (B) Photos of skin grafts from control and R243-treated mice on days 7, 9, 11, 13, 15, 17, 19, 21, 23, and 25 post-transplant. (C) Graft survival rates in control and R243-treated mice. (D) Histological examination of grafts using H&E staining, showing reduced immune infiltration in R243-treated mice. (E) Representative H&E-stained sections of skin grafts from control and R243-treated mice, illustrating immune cell infiltration and tissue damage. (F) Quantification of CD4⁺ and CD8⁺ T cell infiltration in grafts using immunohistochemistry for CD4 and CD8 markers. (G) Immunohistochemistry of CD69 and CD103 in syngeneic and allogeneic grafts. (H) Flow cytometry analysis showing the percentage of immune cell subsets (T cells, B cells, NK cells, DCs, and macrophages) in skin grafts from control and R243-treated mice. (I) Flow cytometry plots depicting CD69 and CD103 expression on CD4⁺ and CD8⁺ T cells in skin grafts from control and R243-treated mice. (J) Immunofluorescence staining of skin grafts for DAPI (blue), CD4 (cyan), CD8 (white), CD69 (orange), CCR8 (red), FOXP3 (green), and CD103 (yellow) in control and R243-treated mice. (K) Flow cytometry analysis of immune checkpoint markers (PD-1, TIGIT, and CTLA-4) on CD8⁺TRM cells in skin grafts from control and R243-treated mice. (L) Flow cytometry analysis of cytokine production (GZMB, IFN-γ, TNF-α, and IL-2) by CD8⁺TRM cells in skin grafts from control and R243-treated mice. Statistical analysis was performed using the t-test to compare the differences between groups. Bars represent mean ± SD from 10 independent experiments. Statistical significance is indicated by *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001; “ns” denotes no significant difference.
Immunohistochemical analysis showed reduced infiltration of both CD4⁺ and CD8⁺ T cells in grafts from R243-treated mice. Histological examination of skin grafts revealed that R243-treated mice exhibited focal dermal-epidermal and periadnexal inflammation with limited stromal infiltration, whereas control mice developed diffuse inflammatory infiltrates, tissue injury, and dermal thickening (Fig. 4D-F). Multiplex immunofluorescence and immunohistochemistry also showed that the expression of TRM-related markers CD69 and CD103 in the skin of the R243 treatment group was significantly reduced (Fig. 4G, J). Flow cytometry further confirmed a significant reduction in T cells (P = 0.0001) within graft tissues—the most abundant and most substantially decreased population—while NK cells (P = 0.0003) were significantly increased. B cells (P = 0.0125) showed a trend towards elevation but without statistical significance, and no significant changes were observed in DCs (P = 0.7008) or macrophages (P = 0.9513). In dLNs, spleen, and peripheral blood, T cells were consistently reduced following R243 treatment. NK cells were significantly increased in dLNs, while B cells showed a non-significant downward trend; DCs and macrophages remained unchanged. In the spleen, DCs were significantly elevated, but no changes were detected in B cells, NK cells, or macrophages. In peripheral blood, NK cells were reduced, while the proportions of B cells, DCs, and macrophages were unaffected (Fig. 4H, Fig. S4B-D).
To determine whether R243 influenced TRM populations, CD8⁺CD69⁺CD103+ T cells and CD8⁺CD69⁺CD103− T cells were analyzed across tissues. R243 treatment led to a significant decrease in both CD8⁺CD69⁺CD103+ T cells (P < 0.0001) and CD8⁺CD69⁺CD103− T cells (P < 0.0001) in grafts, dLNs and spleen (Fig. 4I, S4E, F). Notably, the reduction of CD69⁺CD103⁻ CD8⁺ TRM cells in the graft was particularly pronounced. In parallel, expression of immune checkpoint molecules PD-1 (P < 0.0001), CTLA-4 (P < 0.0001), and TIGIT (P < 0.0001) on CD8⁺CD69⁺CD103+ T cells and CD8⁺CD69⁺CD103− T cells was significantly reduced in grafts, dLNs, and spleen of R243-treated mice compared to controls (Fig. 4K, Fig. S4G, H).
To assess TRM effector function, intracellular staining revealed that both CD8⁺CD69⁺CD103⁺ T cells and CD8⁺CD69⁺CD103- T cells in the grafts of R243-treated mice produced significantly less GZMB (P < 0.0001), IFN-γ (P < 0.0001), and IL-2 (P < 0.0001) compared to the control group. However, in the spleen and dLNs, there were no significant changes in the production of GZMB and IFN-γ by CD8⁺CD69⁺CD103⁺ T cells and CD8⁺CD69⁺CD103- T cells. In the dLNs, IL-2 production by CD8⁺CD69⁺CD103⁺ T cells was significantly increased, while IL-2 production by CD8⁺CD69⁺CD103- T cells showed no significant change. In the spleen, IL-2 production by CD8⁺CD69⁺CD103⁺ T cells showed no significant change, while IL-2 production by CD8⁺CD69⁺CD103- T cells was significantly reduced (Fig. 4L, Fig. S4I, J). Meanwhile, in the R243-treated mice, Treg cells were reduced in the grafts, dLNs, spleen, and peripheral blood. The expression of immune checkpoint molecules (TIGIT and PD-1) on Treg cells was also decreased, indicating that although Treg cells were reduced, the exhausted Treg cells were also diminished (Fig. S4K, L). These findings suggest that CCR8 inhibition selectively dampens CD8⁺ TRM activation and effector function within the graft microenvironment, potentially contributing to prolonged graft survival.
CCR8 knockout impairs TRM infiltration and reduces inflammatory response in skin graft rejection
To further investigate the role of CCR8 in allogeneic transplantation rejection, we generated CCR8 knockout (CCR8−/−) mice. Skin grafts from MHC completely mismatched BALB/c mice were transplanted onto wild-type (WT) or CCR8−/− recipients, as shown in Fig. 5A. Histological analysis of the grafts from CCR8−/− mice revealed significantly reduced tissue damage and decreased infiltration of inflammatory cells compared to the WT group (Fig. 5B, C). Immunohistochemical analysis of CD4+ and CD8+ T cell infiltration in the grafts further revealed that, compared to the control group, the severity of immune infiltration in the CCR8−/− grafts was markedly reduced. Additionally, immunohistochemical analysis of CD69 and CD103 expression showed a significant decrease in both markers in the grafts of CCR8−/− mice (Fig. 5D), suggesting a substantial reduction in CD4+ TRM and CD8+ TRM cells in the grafts.
CCR8 knockout impairs tissue-resident memory T cell infiltration and reduces inflammatory response in skin graft rejection. (A) BALB/c mice served as skin donors, and C57BL/6 CCR8KO or WT mice were used as recipients. Grafts, lymph nodes, spleen, and blood were collected on day 7 post-transplantation for histological examination, flow cytometry, and single-cell RNA sequencing. (B) Severity of immune infiltration in skin grafts assessed by H&E staining in WT and CCR8KO mice. (C) Quantification of CD4⁺ and CD8⁺ T cell infiltration in skin grafts by immunohistochemistry, comparing WT and CCR8KO mice. (D) Representative IHC images showing CD4, CD8, CD69, and CD103 expression in skin grafts from WT and CCR8KO mice. (E) Cytokine levels in serum from WT and CCR8KO mice. (F) Flow cytometry analysis of CD8+TRM cells in draining lymph nodes and spleen of WT and CCR8KO mice. (G) Flow cytometry analysis of immune checkpoint markers (PD-1, TIGIT, and CTLA-4) on CD8⁺TRM cells in dLNs and spleen of WT and CCR8KO mice. Statistical analysis was performed using the t-test to compare the differences between groups. Bars represent mean ± SD from 8–10 independent experiments. Statistical significance is indicated by *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001; “ns” denotes no significant difference.
To further explore how CCR8 regulates the activation and function of CD8+ TRM cells, we analyzed the serum cytokine profiles in skin-transplanted mice. In the CCR8−/− group, levels of TNF-α (P = 0.021), IFN-γ (P = 0.022), IL-1β (P = 0.021), GM-CSF (P = 0.004), and IL-2 (P = 0.041) were decreased, while levels of IL-10 (P = 0.025), MCP-1 (P = 0.036), and MCP-5 (P = 0.004) were elevated (Fig. 5E), indicating a reduction in inflammation. Meanwhile, several chemokines, including CXCL1, CXCL5, CXCL11, CXCL12, CXCL13, CXCL16, CCL3, CCL4, CCL5, CCL11, CCL17, CCL19, CCL20, CCL22, CCL24, CCL27, and CX3CL1, showed no significant change (Fig. S5D). Moreover, we examined the changes in lymphocyte subset distribution in the dLNs, spleen, and peripheral blood. Compared with the control group, in the dLNs, the proportion of T cells (P < 0.0001) was significantly reduced, whereas NK cells (P < 0.0001), DCs (P = 0.0001), and macrophages (P = 0.0003) increased, with no significant change in B cells (P = 0.5865); in the spleen, the proportions of T cells (P < 0.0001), DCs (P = 0.0001), and macrophages (P < 0.0001) decreased, with T cells showing the most significant reduction. B cells (P = 0.0956) showed a trend toward increase, but this did not reach statistical significance, and NK cells (P = 0.6498) did not exhibit significant changes; in peripheral blood, the proportion of T cells (P = 0.0001) was significantly lower, with no significant changes in NK cells (P = 0.5558) or macrophages (P = 0.9476), while B cells (P = 0.0873) showed a trend toward decrease and DCs (P = 0.1177) a trend toward increase, but neither reached statistical significance (Fig. S5A-C).
We also performed flow cytometry to assess the proportion of CD8⁺CD69⁺CD103+ T cells and CD8⁺CD69⁺CD103− T cells in the dLNs and spleen. Compared to the WT group, CCR8−/− mice exhibited significantly reduced proportions of CD8⁺CD69⁺CD103+ T cells and CD8⁺CD69⁺CD103− T cells in all tissues, suggesting a substantial reduction of TRM following CCR8 knockout (Fig. 5F). Finally, we assessed the expression of immune checkpoints, including PD-1, CTLA-4, and TIGIT, on CD8⁺CD69⁺CD103+ T cells and CD8⁺CD69⁺CD103− T cells in the dLNs and spleen. In both the spleen and dLNs, the expression of PD-1, CTLA-4, and TIGIT was significantly decreased on CD8⁺CD69⁺CD103+ T cells and CD8⁺CD69⁺CD103− T cells (Fig. 5G), indicating reduced activation of CD8+TRM cells.
CCR8 deficiency restructures the skin immune landscape and reduces CD8⁺ TRM cell proportions
To investigate the role of CCR8 in shaping the skin immune microenvironment, we performed scRNA-seq on skin cells from three groups of mice, including wild-type controls (n = 3), anti-CCR8 antibody-treated mice (Anti_Ccr8, n = 3), and CCR8 knockout mice (KO_Ccr8, n = 4), with the total of 109, 212 high quality cells. Unsupervised clustering of all cells identified 21 major cell clusters (Fig. 6A), with broadly similar cellular compositions observed across the different treatment groups (Fig. 6B). Based on canonical marker gene expression patterns, these clusters were annotated into seven major immune cell lineages: T cells, NK cells, neuron-like cells, myeloid cells, granulocytes, fibroblasts, and B cells (Fig. 6C, D).
CCR8 deficiency restructures the skin immune landscape and reduces CD8⁺ TRM cell proportions. (A) UMAP plot showing the clustering of immune cell populations based on scRNA-seq data. Each cluster is represented by different colors, with each number representing a distinct cluster of cells. (B) UMAP plot depicting immune cell distribution in control, Anti_Ccr8, and KO_Ccr8 groups. (C) UMAP plot with distinct immune cell types highlighted, including B cells, fibroblasts, granulocytes, myeloid cells, neurons-like cells, NK cells, and T cells. (D) Dot plot showing the expression of specific markers for various immune cell types. The size of the dots indicates the percentage of cells expressing each marker, and the color represents the average expression level. (E) Bar graph showing the distribution of immune cell types in the Anti_Ccr8, Control, and KO_Ccr8 groups. (F) UMAP plot showing detailed subsets of CD8⁺ T cells, such as CD8⁺ T effector, CD8⁺ TRM, CD8⁺ TEX, and CD8⁺ proliferative cells, as well as CD4⁺ T cells and their subsets. (G) Bar graph showing the distribution of CD4⁺ and CD8⁺ T cell subsets in the Anti_Ccr8, Control, and KO_Ccr8 groups. (H-I) Pseudotime analysis with identified CD8+ TRM, and Teff cells.
Further comparative analysis revealed a significant reduction in the T cell population in the KO_Ccr8 (Fig. 6E). Reclustering of the T cell compartment identified 11 subpopulations, including Tregs, CD4⁺ T cells, proliferative CD8⁺ T cells, effector CD8⁺ T cells, exhausted CD8⁺ T cells (TEX), CD8⁺ TRM, double-negative T cells (DNT), and double-positive T cells (DPT) (Fig. 6F, Fig. S6A). The specific expression of key marker genes (CD4, CD8a, Foxp3, Pdcd1) in these subsets further validated their classification accuracy (Fig. S6B, C). Notably, the proportion of CD8⁺ TRM cells was markedly reduced in the KO_Ccr8 group compared to both the control and anti-CCR8-treated groups (Fig. 6G), suggesting that CCR8 plays a critical regulatory role in the development or maintenance of CD8⁺ TRM cells. Pseudotime trajectory analysis reveals that CD8+ Teff cells are in the mid-stage of differentiation, while CD8+ TRM cells are in the late stage with stable functionality, suggesting the transition of CD8+ Teff cells into functionally mature CD8+ TRM cells. CD8+ TRM cells play a critical role in transplantation (Fig. 6H, I).
These results indicate that CCR8 is an essential factor in maintaining immune homeostasis, particularly in supporting the presence of local protective CD8⁺ TRM.
CCR8 deficiency alters immune cell interactions and gene expression in graft microenvironment
To investigate the impact of CCR8 deletion on the skin immune microenvironment, we performed unsupervised clustering analysis on Myeloid cells from three groups of mice. This analysis revealed 14 distinct Myeloid cell subclusters (Fig. 7A, B), and the different treatment groups showed variations in cell composition, particularly within macrophages and monocyte populations, mainly including Apoe+ Macro, Ccl8+ macro, Arg1+ Macro, Spp1+ Macro, Cst3+ DC, Ly6c2+ Mono, among others. We observed an increase in Ccl8+ Macro in the CCR8_KO group, which may indicate reactive upregulation. GO analysis suggests that this population of cells primarily mediates lymphocyte migration and T cell proliferation (Fig. 7C, Fig. S7). Figure 7D displays the expression patterns of key immune marker genes across different clusters (Table S1).
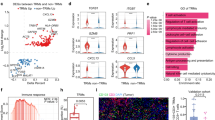
CCR8 deficiency alters immune cell interactions and gene expression in skin microenvironment. (A) UMAP plot showing the clustering of immune cell populations based on scRNA-seq data. Each cluster is represented by a different color, with each number corresponding to a distinct cell identity. (B) UMAP plot depicting the immune cell distribution in control, Anti_Ccr8, and KO_Ccr8 groups. (C) Bar graph showing the distribution of different immune cell types across the Anti_Ccr8, Control, and KO_Ccr8 groups. (D) Dot plot showing the expression of specific markers in various immune cell populations, where the size of the dots represents the percentage of cells expressing the marker and the color indicates the average expression level. (E) Bar graphs showing the number of interactions and interaction strength between immune cell populations in the control, Anti_Ccr8, and KO_Ccr8 groups. (F) Network plots showing the interactions between immune cells in the control, Anti_Ccr8, and KO_Ccr8 groups, with nodes representing immune cell populations and edges representing interactions. (G) Bar chart showing the relative information flow between immune cells in the control, Anti_Ccr8, and KO_Ccr8 groups. (H) Interaction matrix showing the p-value and the strength of communication between different immune cell populations in response to CCR8 inhibition. Each row and column represent different immune cell subsets, with the color intensity indicating the level of communication.
We used functional enrichment Analysis to assess functional differences in gene expression between the groups (Table S2, 3). The bar plots in Figure S7 (C0 to C13) show that in the CCR8 knockout group, many immune-related pathways, such as wound healing (C0), inflammatory response regulation (C1), and skin development (C11), were significantly altered. In particular, genes associated with T cell proliferation (C9), inflammatory response (C8), and cell migration (C7) were significantly upregulated in the KO_Ccr8 group, while pathways related to collagen formation (C7) and antigen processing (C2) were downregulated. These results suggest that CCR8 is essential for maintaining immune cell interactions and supporting normal immune function, especially in regulating T cell activation and proliferation, macrophage interactions, and skin-specific immune responses.
We further examined cellular interaction analysis between Myeloid and T cell (Table S4). As shown in Fig. 7E, the KO_Ccr8 exhibited significantly reduced numbers (left plot) and intensities (right plot) of immune cell interactions compared to the control and anti-CCR8 groups, suggesting that CCR8 plays a crucial role in promoting skin immune cell interactions. Furthermore, Fig. 7F illustrates the interaction network between key immune cell types. This figure shows that in the CCR8 knockout group, the interaction between CD8⁺ TRM cells and macrophages is markedly reduced, contrasting with the control and anti-CCR8 groups, indicating a disruption in immune cell cooperation.
The analysis of ligand-receptor signaling pathways revealed significant changes in immune molecular signaling in the KO_Ccr8 group compared to the control group. Specifically, the information flow for CD40, PD-L1, and NRG was markedly increased in the KO_Ccr8 group, indicating a substantial upregulation of these immune markers in the absence of Ccr8. This suggests that Ccr8 plays a critical role in regulating these immune molecules, potentially modulating immune responses to maintain immune homeostasis (Fig. 7G). These findings demonstrate that the loss or inhibition of Ccr8 significantly impacts the regulation of immune molecules, with a greater magnitude of effect observed in the KO_Ccr8 group compared to the Anti_Ccr8 group. These results provide valuable insights into the role of Ccr8 in immune signaling and its potential implications for immune-based therapies. The intercellular communication network elucidates the interactions between macrophage chemokines (Cxcl4, Cxcl16, Cxcl10, Ccl8) and T cell subset (CD4+Treg, CD4+T, CD8+Teff, CD8+TEX, CD8+TRM) chemokine receptors (Cxcr3, Ccr2, Ccr5) across the three groups (Fig. 7H). It indicates that the interaction between macrophages and CD8+ TRM cells is reduced after CCR8 inhibition or knockout.
Blockade of CCL8 or CCL1 prolongs graft survival by reducing TRM cell infiltration and activation
In mice, CCR8 functions as the receptor for both CCL8 and CCL1. Given the chemotactic nature of CCR8⁺ TRM cells, we hypothesized that their migration toward the graft may be mediated by CCL8 and CCL1, and that blocking either chemokine could inhibit this process. To test this, we administered monoclonal antibodies against CCL8 or CCL1 to transplant recipients and compared outcomes with control-treated animals. The treatment regimen is illustrated in Fig. 8A.
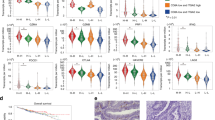
Blockade of CCL8 or CCL1 prolongs graft survival by reducing TRM cell infiltration and activation. (A) BALB/c mice received skin grafts from C57BL/6 mice and were treated with 50 µg of Anti-CCL1 mAb or Anti-CCL8 mAb on Days 0, 2, 4, and 6. Mice were sacrificed on Day 7, and tissues from skin, lymph nodes, spleen, and blood were analyzed. (B) Photos of skin grafts on Days 7, 9, 13, 15, 17, and 21 after transplantation in mice treated with either Anti-CCL1 or Anti-CCL8 mAb. (C) Graft survival curves and graft survival time for control, Anti-CCL1 mAb, and Anti-CCL8 mAb groups. (D) Representative histological images showing CD4, CD8, CD69 and CD103 staining of skin grafts at Day 7 post-transplantation in Anti-CCL1 and Anti-CCL8 mAb-treated mice. (E–K) Changes in CD8⁺ TRM cells in the graft, their expression of immune checkpoints (PD-1, TIGIT, and CTLA-4), and their ability to produce cytokines (GZMB, IFN-γ, and IL-2). Statistical analysis was performed using the t-test to compare the differences between groups. Bars represent mean ± SD from 6 independent experiments. Statistical significance is indicated by *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001; “ns” denotes no significant difference.
Starting on day 7 post-transplantation, graft necrosis areas were monitored daily until complete rejection. Graft survival was significantly prolonged in both anti-CCL8 (MST 13.6 vs. 21 days posttransplant; P = 0.0011) and anti-CCL1 (MST 13.6 vs. 20.67 days posttransplant; P = 0.0011) treatment groups compared to controls, with no significant difference between the two treatment arms (Fig. 8B, C). H&E staining revealed markedly reduced inflammatory cell infiltration and attenuated tissue damage in the treated groups (Fig. 8D).
Immunohistochemical analysis showed decreased infiltration of CD4⁺ and CD8⁺ T cells in the grafts, accompanied by significantly reduced expression of CD69 and CD103, indicating a reduction in both CD4⁺ and CD8⁺ TRM populations (Fig. 8D). Flow cytometry confirmed a significant decrease in graft-infiltrating CD4⁺ and CD8⁺ T cells in both treatment groups compared to controls, with a more pronounced reduction in the anti-CCL1 group. Similar reductions were observed in the spleen and peripheral blood. Interestingly, in dLNs, CD4⁺ T cells were significantly reduced in both treatment groups. CD8⁺ T cells were significantly decreased only in the anti-CCL1 group, with no significant change in the anti-CCL8 group (Fig. S8A, B).
Further flow cytometric analysis of TRM subsets demonstrated significant reductions in CD69⁺CD103⁻ CD8⁺ T (P = 0.01) cells within grafts in both treatment groups. Although CD69⁺CD103⁺ CD8⁺ T cells (P = 0.39) exhibited a downward trend in the anti-CCL1 group, the difference did not reach statistical significance. In dLNs, CD69⁺CD103⁺ CD8⁺ T cells (P = 0.014 and P = 0.0073) were significantly reduced in both treatment groups. CD69⁺CD103⁻ CD8⁺ T cells (P = 0.085) also showed a decreasing trend, but this was not statistically significant in the anti-CCL8 group. In the spleen, both CD8⁺CD69⁺CD103+ T cells (P < 0.0001) and CD8⁺CD69⁺CD103− T cells (P = 0.0051 and P = 0.02) subsets were significantly diminished in both treatment groups (Fig. 8E, Fig. S8C).
Intragraft CD8⁺ TRM cells were markedly decreased following treatment. Correspondingly, the expression of immune checkpoint molecules PD-1, TIGIT, and CTLA-4 on CD8⁺ TRM cells was significantly reduced. In dLNs, PD-1 expression remained unchanged, while TIGIT and CTLA-4 were significantly downregulated. In the spleen, PD-1 expression on CD8⁺ TRM cells was unchanged in the anti-CCL1 group but significantly reduced in the anti-CCL8 group. Both treatment groups exhibited significant reductions in TIGIT and CTLA-4 expression (Fig. 8F-H, Fig. S8D-F).
Functionally, CD8⁺ TRM cells in the grafts of both treatment groups showed reduced production of GZMB (P = 0.028 and P = 0.003), IFN-γ (P = 0.017 and P = 0.006), and IL-2 (P = 0.007 and P = 0.014). Interestingly, in dLNs and the spleen, GZMB production by CD8⁺ TRM cells was increased, with the anti-CCL1 group showing the most pronounced enhancement. No significant changes were observed in IFN-γ or IL-2 production in these compartments (Fig. 8I-K, Fig. S8G-I).
In parallel, we assessed the effects on Treg cells. The results showed a significant reduction of Treg cells in the grafts of the anti-CCL1 and anti-CCL8 treatment groups, while no notable changes were observed in the spleen, dLNs, or blood (Fig. S8J).
In conclusion, after neutralizing CCR8 ligands, we found no significant changes in total CD8⁺ T cells within the grafts, whereas CD8⁺ TRM cells were significantly reduced, along with suppression of their activation and function. This indicates that CCR8 is a key molecule for the activation and functional regulation of CD8⁺ TRM cells.
Discussion
In our previous research, single-cell sequencing analysis of mouse liver transplantation revealed that CD8⁺ TRM cells play a major role in acute transplant rejection9,44. Based on this, the present study first established both syngeneic and allogeneic skin transplantation tolerance and rejection models. In the rejection group mice, there was a significant increase in CD8⁺ TRM cells in the grafts, dLNs and spleen, consistent with previous findings18,45. Meanwhile, Treg cells were significantly elevated during the acute rejection phase (Fig. 1, Fig. S1). Previous studies have shown that a decrease in Treg cell levels is characteristic of transplants that do not undergo rejection46,47. Subsequently, by analyzing single-cell data, we found that compared to syngeneic grafts, CCR8 expression in CD8⁺ TRM cells was significantly increased in allogeneic grafts, and interactions between CD8⁺ TRM cells and other immune cells were also enhanced (Fig. 2). More interestingly, consistent with previous studies, immune cells in the graft were almost completely replaced by recipient cells5,48. Our study further discovered that nearly all CD8⁺ TRM cells in the graft originated from the recipient, and the expression of CCR8 on recipient-derived CD8⁺ TRM cells gradually increased over time (Fig. 3). Blocking CCR8 (Fig. 4), knocking out CCR8 (Fig. 5), and intervening with the anti-CCR8 ligands CCL1 and CCL8 (Fig. 8) all significantly prolonged the survival of allogeneic grafts and reduced the infiltration and activation of CD8⁺ TRM cells. In conclusion, our findings indicate that CCR8 plays an important role in alleviating the allogeneic antigen response in skin transplantation recipients, and also highlight the potential of targeting TRM cells as a therapeutic intervention strategy.
CCR8, as a chemokine receptor, is expressed on various immune cells29,49. It is notable that CCR8 is not expressed on homeostatic T cells. Instead, upon stimulation, the CCR8 stored intracellularly is rapidly transported to the cell surface50. We observed that the most significant changes in immune cell subsets (including T cells, B cells, NK cells, macrophages, and DCs) after CCR8 blockade occurred in T cells. Earlier work identified an early inflammatory amplification pathway in which donor kidney–resident macrophages promote recipient immune-cell infiltration through the CCL8–CCR8 axis51. Building on this, our study centers on CD8⁺ TRM cells, a key effector population in transplant rejection. It has been reported that CCR8 is a marker for TRM cells in human skin52. In our study, we found that CCR8 is a key regulator of TRM cell migration and retention in the graft. In recipient-derived CD8⁺ TRM cells in allogeneic transplants, the expression of CCR8 was significantly upregulated in the inflammatory regions. This suggests that CCR8 mediates the recruitment of TRM cells to the transplant site, thereby promoting immune rejection. Inhibition of CCR8 using the R243 inhibitor significantly prolonged graft survival and reduced TRM cell infiltration. Meanwhile, using CCR8 knockout mice also yielded the same results, further supporting the importance of CCR8 in transplant rejection. These results are consistent with previous studies, indicating that CCR8 is crucial for the migration of TRM cells and their retention in tissues29.
The ability to regulate the dynamics of TRM cells offers an exciting therapeutic opportunity to improve transplant outcomes. In lung transplantation, reducing the infiltration of TRM cells in the graft has been shown to reduce graft dysfunction53. Based on these findings, we made the novel observation that, in a skin graft model, targeting CCR8 prolongs graft survival by reducing the accumulation of TRM cells within the graft. We further found that blocking CCR8 reduced the expression of immune checkpoint molecules on TRM cells and diminished their ability to produce cytokines such as GZMB, IFN-γ, and IL-2, suggesting that CCR8 not only regulates TRM cell migration but may also affect their functional status. The study shows that CD8+ TRM cells have a high ability to secrete IL-2, which can promote the long-term survival of CD8+ TRM cells54,55. Furthermore, targeting CCR8 effectively reduces the accumulation of CD8+ TRM cells within the graft, further supporting the critical role of CCR8 in regulating immune responses. Therefore, targeting CCR8 effectively reduces the accumulation of CD8⁺ TRM cells within the graft, suppresses their function, and prolongs graft survival, providing an effective strategy to mitigate transplant rejection. Studies have shown that Treg cells exert immunosuppressive effects in transplant rejection by inhibiting effector T-cell activation, regulating dendritic cell function, and modulating immune responses through immune checkpoint molecules56,57,58,59. Treg cells play a role in promoting immune tolerance, while CD8 + TRM cells play a major role in transplant rejection. Interestingly, despite the simultaneous reduction of both Treg cells and CD8 + TRM cells, transplant rejection was not exacerbated; instead, graft survival was promoted. This suggests that in transplant immune responses, the pro-rejection effect of CD8 + TRM cells is significantly stronger than the immune tolerance-promoting effect of Treg cells. These findings open new avenues for exploring the potential of CCR8 antagonists as adjunctive therapy in transplantation to promote graft survival and alleviate rejection.
Previous studies have shown that CD8⁺ TRM cells increase after liver transplantation, but their donor versus recipient origin has not been defined5,18. Here, using CD45.1/CD45.2 congenic mice in a skin-graft model, we tracked the dynamics of donor- and recipient-derived T_RM and found that post-transplant intragraft CD8⁺ TRM are predominantly recipient-derived. Mechanistically, recipient perilesional CD8⁺ TRM likely undergo antigen-driven proliferation, upregulate CCR8, and migrate into the graft via chemotaxis to mediate rejection. Consistent with this, CCR8 blockade effectively reduced intragraft CD8⁺ TRM accumulation, suggesting a therapeutic strategy to promote long-term graft survival.
Our study provides valuable insights into the role of TRM cells in transplant rejection. Although TRM cells have been identified as key players in rejection responses, the mechanisms regulating their migration and retention in allogeneic transplantation have not been fully explored. We used single-cell RNA sequencing to provide insights into the cell composition of the graft after CCR8 blockade, as well as the role of CCR8 in mediating TRM cell accumulation. By identifying interactions between TRM cells and other immune cell populations (such as T cells, macrophages, and NK cells), our research offers a more detailed understanding of the immune processes driving transplant rejection.
Although our findings are encouraging, there are several limitations to consider. Although the murine skin transplantation model is informative, it cannot replace vascularized solid-organ transplantation, and the long-term effects of CCR8 inhibition on immune function and transplant tolerance still require further investigation. Additionally, further studies are needed to investigate the interactions between TRM cells and other immune cell populations, including regulatory T cells and DCs, to gain a comprehensive understanding of the mechanisms underlying transplant rejection.
Our data indicate that CCR8 is a key regulator of the accumulation and activation of CD8⁺ TRM cells in allogeneic grafts, linking this pathway to stronger rejection responses. These results suggest that CCR8 modulation may represent a translationally promising strategy to improve graft survival by limiting pathogenic TRM-mediated local inflammation. In principle, targeting CCR8 could serve as a complement to current immunosuppressive regimens by acting on tissue-resident effector populations that are not fully controlled by conventional systemic immunosuppression. From a clinical perspective, CCR8 inhibition might be applied as an induction-phase adjunct to prevent the early seeding and expansion of TRM cells within the graft. However, multiple challenges must be addressed before clinical translation. First, CCR8 is not exclusively expressed on CD8⁺ TRM cells; it may also be present on other immune subsets depending on tissue context and inflammatory cues. Thus, CCR8 blockade could inadvertently disrupt protective immunity or regulatory networks. Therefore, defining the expression landscape of CCR8 across different immune cell types in transplantation and clarifying its downstream signaling features will be essential for determining whether antagonism or agonism should be prioritized clinically. Second, TRM cells play important roles in local antiviral and antibacterial defense, and long-term CCR8 inhibition may compromise tissue immunity or impair vaccine-elicited peripheral memory, particularly in the setting of concomitant standard immunosuppression. Third, the optimal timing, dosage, and tissue penetration of CCR8-targeted therapy remain unclear, and excessive suppression of local immune surveillance may increase the risk of malignancy in the graft or surrounding tissues.
In conclusion, our study provides strong evidence that CCR8 plays a key role in the accumulation and activation of CD8⁺ TRM cells in allogeneic skin grafts, thereby promoting transplant rejection. Targeting CCR8 may represent a novel therapeutic approach to extend graft survival and reduce rejection by modulating TRM cell responses. CCR8 targeting significantly reduced TRM functional activation in this study, which may reduce the graft’s reliance on conventional immunosuppressants and also allow the development of tissue-specific delivery systems to further minimize systemic side effects. These findings contribute to a deeper understanding of tissue-resident immune mechanisms in transplantation and lay the foundation for future therapeutic strategies aimed at improving transplant outcomes.
Data availability
The datasets generated during the current study are available in the NGDC Genome Sequence Archive repository with the dataset identifier CRA029911 (https://ngdc.cncb.ac.cn/gsub/submit/gsa/subCRA048623). All processed data are available from the corresponding author on reasonable request.
References
Short, S., Lewik, G. & Issa, F. An immune atlas of T cells in transplant rejection: pathways and therapeutic opportunities. Transplantation 107, 2341–2352. https://doi.org/10.1097/tp.0000000000004572 (2023).
Marino, J., Paster, J. & Benichou, G. Allorecognition by T lymphocytes and allograft rejection. Front. Immunol. 7 https://doi.org/10.3389/fimmu.2016.00582 (2016).
Liu, Y. et al. Donor MHC-specific thymus vaccination allows for immunocompatible allotransplantation. Cell Res. 35, 132–144. https://doi.org/10.1038/s41422-024-01049-5 (2025).
Ohm, B. et al. Chimerism and immunological tolerance in solid organ transplantation. Semin. Immunopathol. 47 https://doi.org/10.1007/s00281-025-01052-x (2025).
Prosser, A. et al. Dynamic changes to tissue-resident immunity after MHC-matched and MHC-mismatched solid organ transplantation. Cell. Rep. 35, 109141. https://doi.org/10.1016/j.celrep.2021.109141 (2021).
Masopust, D. & Soerens, A. G. Tissue-resident T cells and other resident leukocytes. Annu. Rev. Immunol. 37, 521–546. https://doi.org/10.1146/annurev-immunol-042617-053214 (2019).
Iijima, N. & Iwasaki, A. Tissue instruction for migration and retention of TRM cells. Trends Immunol. 36, 556–564. https://doi.org/10.1016/j.it.2015.07.002 (2015).
Matloubian, M. et al. Lymphocyte egress from thymus and peripheral lymphoid organs is dependent on S1P receptor 1. Nature 427, 355–360. https://doi.org/10.1038/nature02284 (2004).
Li, X. et al. Single cell RNA-sequencing delineates CD8(+) tissue resident memory T cells maintaining rejection in liver transplantation. Theranostics 14, 4844–4860. https://doi.org/10.7150/thno.96928 (2024).
Wakim, L. M. et al. The molecular signature of tissue resident memory CD8 T cells isolated from the brain. J. Immunol. 189, 3462–3471. https://doi.org/10.4049/jimmunol.1201305 (2012).
Mackay, L. K. et al. The developmental pathway for CD103(+)CD8 + tissue-resident memory T cells of skin. Nat. Immunol. 14, 1294–1301. https://doi.org/10.1038/ni.2744 (2013).
Mueller, S. N. & Mackay, L. K. Tissue-resident memory T cells: local specialists in immune defence. Nat. Rev. Immunol. 16, 79–89. https://doi.org/10.1038/nri.2015.3 (2016).
Bergsbaken, T. & Bevan, M. J. Proinflammatory microenvironments within the intestine regulate the differentiation of tissue-resident CD8⁺ T cells responding to infection. Nat. Immunol. 16, 406–414. https://doi.org/10.1038/ni.3108 (2015).
Sasson, S. C., Gordon, C. L., Christo, S. N., Klenerman, P. & Mackay, L. K. Local heroes or villains: tissue-resident memory T cells in human health and disease. Cell Mol. Immunol. 17, 113–122. https://doi.org/10.1038/s41423-019-0359-1 (2020).
Fu, J. & Sykes, M. Emerging concepts of tissue-resident memory T cells in transplantation. Transplantation 106, 1132–1142. https://doi.org/10.1097/tp.0000000000004000 (2022).
Poon, M. M. L. et al. Tissue adaptation and clonal segregation of human memory T cells in barrier sites. Nat. Immunol. 24, 309–319. https://doi.org/10.1038/s41590-022-01395-9 (2023).
Alexander, K. L. & Ford, M. L. The entangled world of memory T cells and implications in transplantation. Transplantation 108, 137–147. https://doi.org/10.1097/tp.0000000000004647 (2024).
Tian, Q. et al. Skin and heart allograft rejection solely by long-lived alloreactive T(RM) cells in skin of severe combined immunodeficient mice. Sci. Adv. 8, eabk0270. https://doi.org/10.1126/sciadv.abk0270 (2022).
Asada, N., Ginsberg, P., Gagliani, N., Mittrucker, H. W. & Panzer, U. Tissue-resident memory T cells in the kidney. Semin Immunopathol. 44, 801–811. https://doi.org/10.1007/s00281-022-00927-7 (2022).
Snyder, M. E. et al. Generation and persistence of human tissue-resident memory T cells in lung transplantation. Sci. Immunol. 4 https://doi.org/10.1126/sciimmunol.aav5581 (2019).
Takahama, Y. Journey through the thymus: stromal guides for T-cell development and selection. Nat. Rev. Immunol. 6, 127–135. https://doi.org/10.1038/nri1781 (2006).
Murooka, T. T. et al. CCL5-CCR5-mediated apoptosis in T cells: requirement for glycosaminoglycan binding and CCL5 aggregation. J. Biol. Chem. 281, 25184–25194. https://doi.org/10.1074/jbc.M603912200 (2006).
Zou, Y. R., Kottmann, A. H., Kuroda, M., Taniuchi, I. & Littman, D. R. Function of the chemokine receptor CXCR4 in haematopoiesis and in cerebellar development. Nature 393, 595–599. https://doi.org/10.1038/31269 (1998).
Strieter, R. M. et al. The functional role of the ELR motif in CXC chemokine-mediated angiogenesis. J. Biol. Chem. 270, 27348–27357. https://doi.org/10.1074/jbc.270.45.27348 (1995).
Liu, G., Wang, Z. & Li, S. Heterogeneity and plasticity of tissue-resident memory T cells in skin diseases and homeostasis: a review. Front. Immunol. 15, 1378359. https://doi.org/10.3389/fimmu.2024.1378359 (2024).
Islam, S. A. et al. Mouse CCL8, a CCR8 agonist, promotes atopic dermatitis by recruiting IL-5 + T(H)2 cells. Nat. Immunol. 12, 167–177. https://doi.org/10.1038/ni.1984 (2011).
Jiang, S. et al. Unveiling the structural mechanisms of nonpeptide ligand recognition and activation in human chemokine receptor CCR8. Sci. Adv. 10, eadj7500. https://doi.org/10.1126/sciadv.adj7500 (2024).
Moser, B. Chemokine Receptor-Targeted therapies: special case for CCR8. Cancers 14 https://doi.org/10.3390/cancers14030511 (2022).
Feng, X. et al. CD8(+) tissue-resident memory T cells are essential in bleomycin-induced pulmonary fibrosis. Am. J. Physiol. Cell Physiol. 327, C1178–c1191. https://doi.org/10.1152/ajpcell.00368.2024 (2024).
Hou, Y. et al. Optimizing stem cell infusion timing in the prevention of acute graft-versus-host disease. Cell 188, 3030–3044e3017. https://doi.org/10.1016/j.cell.2025.03.022 (2025).
Wang, P. et al. Oral alloantigen exposure promotes donor-specific tolerance in a mouse model of minor-mismatched skin transplantation. Am. J. Transpl. 22, 2348–2359. https://doi.org/10.1111/ajt.17107 (2022).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell. 184, 3573–3587.e3529. https://doi.org/10.1016/j.cell.2021.04.048 (2021).
Zhou, L. X. et al. Myeloid-derived suppressor cells-induced exhaustion of CD8 + T-cell participates in rejection after liver transplantation. Cell. Death Dis. 15, 507. https://doi.org/10.1038/s41419-024-06834-z (2024).
Jiang, P. et al. PTX3 exacerbates hepatocyte pyroptosis in hepatic ischemia-reperfusion injury by promoting macrophage M1 polarization. Int. Immunopharmacol. 143, 113604. https://doi.org/10.1016/j.intimp.2024.113604 (2024).
Li, X. et al. Integrated single cell analysis reveals an atlas of tumor associated macrophages in hepatocellular carcinoma. Inflammation 47, 2077–2093. https://doi.org/10.1007/s10753-024-02026-1 (2024).
Aran, D. et al. Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat. Immunol. 20, 163–172. https://doi.org/10.1038/s41590-018-0276-y (2019).
Wu, T. et al. ClusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innov. (Camb). 2, 100141. https://doi.org/10.1016/j.xinn.2021.100141 (2021).
Kanehisa, M., Furumichi, M., Sato, Y., Matsuura, Y. & Ishiguro-Watanabe, M. KEGG: biological systems database as a model of the real world. Nucleic Acids Res. 53, D672–d677. https://doi.org/10.1093/nar/gkae909 (2025).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Science: Publication Protein Soc. 28, 1947–1951. https://doi.org/10.1002/pro.3715 (2019).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30. https://doi.org/10.1093/nar/28.1.27 (2000).
Jin, S. et al. Inference and analysis of cell-cell communication using cellchat. Nat. Commun. 12, 1088. https://doi.org/10.1038/s41467-021-21246-9 (2021).
Gauthier, J. M., Harrison, M. S., Krupnick, A. S., Gelman, A. E. & Kreisel, D. The emerging role of regulatory T cells following lung transplantation. Immunol. Rev. 292, 194–208. https://doi.org/10.1111/imr.12801 (2019).
Lee, H. et al. Single-cell RNA sequencing revealed the role of the Th17 pathway in the development of anti- human leukocyte antigen antibodies in a highly sensitized mouse model. Kidney Res. Clin. Pract. https://doi.org/10.23876/j.krcp.23.317 (2024).
Li, X. et al. Landscape of immune cells heterogeneity in liver transplantation by single-cell RNA sequencing analysis. Front. Immunol. 13 https://doi.org/10.3389/fimmu.2022.890019 (2022).
Abou-Daya, K. I. et al. Resident memory T cells form during persistent antigen exposure leading to allograft rejection. Sci. Immunol. 6 https://doi.org/10.1126/sciimmunol.abc8122 (2021).
Rubino, V. et al. T(R3-56) and Treg regulatory T cell subsets as potential indicators of graft tolerance control in kidney transplant recipients. Int. J. Mol. Sci. 25 https://doi.org/10.3390/ijms251910610 (2024).
Li, X. et al. Impact of exosomes derived from adipose stem cells on lymphocyte proliferation and phenotype in mouse skin grafts. Extracell. Vesicles Circ. Nucl. Acids. 6, 141–157. https://doi.org/10.20517/evcna.2024.52 (2025).
Huang, H. et al. The time-dependent shift in the hepatic graft and recipient macrophage pool following liver transplantation. Cell Mol. Immunol. 17, 412–414. https://doi.org/10.1038/s41423-019-0253-x (2020).
Wen, Y., Xia, Y., Yang, X., Li, H. & Gao, Q. CCR8: a promising therapeutic target against tumor-infiltrating regulatory T cells. Trends Immunol. 46, 153–165. https://doi.org/10.1016/j.it.2025.01.001 (2025).
Gombert, M. et al. CCL1-CCR8 interactions: an axis mediating the recruitment of T cells and Langerhans-type dendritic cells to sites of atopic skin inflammation. J. Immunol.. 174, 5082–5091. https://doi.org/10.4049/jimmunol.174.8.5082 (2005).
Dangi, A. et al. Blocking CCL8-CCR8-mediated early allograft inflammation improves kidney transplant function. J. Am. Soc. Nephrology: JASN. 33, 1876–1890. https://doi.org/10.1681/asn.2022020139 (2022).
Zaid, A. et al. Chemokine receptor-dependent control of skin tissue-resident memory T cell formation. J. Immunol. (Baltimore Md. : 1950). 199, 2451–2459. https://doi.org/10.4049/jimmunol.1700571 (2017).
Moghbeli, K. et al. NKG2D Blockade impairs tissue-resident memory T cell accumulation and reduces chronic lung allograft dysfunction. JCI Insight. 10 https://doi.org/10.1172/jci.insight.184048 (2025).
Pallett, L. J. et al. IL-2(high) tissue-resident T cells in the human liver: sentinels for hepatotropic infection. J. Exp. Med. 214, 1567–1580. https://doi.org/10.1084/jem.20162115 (2017).
Feau, S., Arens, R., Togher, S. & Schoenberger, S. P. Autocrine IL-2 is required for secondary population expansion of CD8(+) memory T cells. Nat. Immunol. 12, 908–913. https://doi.org/10.1038/ni.2079 (2011).
Bradley, J. A. Transplant tolerance by Treg therapy. Am. J. Transplantation: Official J. Am. Soc. Transplantation Am. Soc. Transpl. Surg. 14, 5–6. https://doi.org/10.1111/ajt.12510 (2014).
Bernaldo-de-Quirós, E. et al. First-in-human therapy with Treg produced from thymic tissue (thyTreg) in a heart transplant infant. J. Exp. Med. 220 https://doi.org/10.1084/jem.20231045 (2023).
Tang, Q. & Vincenti, F. Transplant trials with tregs: perils and promises. J. Clin. Investig. 127, 2505–2512. https://doi.org/10.1172/jci90598 (2017).
Juneja, T., Kazmi, M., Mellace, M. & Saidi, R. F. Utilization of Treg cells in solid organ transplantation. Front. Immunol. 13, 746889. https://doi.org/10.3389/fimmu.2022.746889 (2022).
Acknowledgements
The authors would like to thank Organ Transplantation Center and the Key Laboratory of Organ Transplantation of Affiliated Hospital of Qingdao University and Organ Transplant Center of Fujian Medical University Union Hospital for their technical support.
Funding
This work was supported by the National Natural Science Foundation of China (No. 82370666 and No. 82470686), the Science Foundation of Shandong Province (No. ZR2022MH292), the Science Foundation of Qingdao Science and Technology Bureau (No. 24-1-8-smjk-1-nsh), Fujian provincial health technology project (No. 2023CXA014), Science Foundation of Fujian Medical University Union Hospital (No. 2020XH010 and 2020XH011), Henan Province Science and Technology Key Project (No. 242102310072), and Shanxi Province Key Laboratory of Digestive Diseases and Organ Transplantation Open Funded Research Project (No. DDOT2024002).
Author information
Authors and Affiliations
Contributions
JZC, XQL, and XTW contributed to the research design. XTW, XQL, SPL, HLC, XZ, ZYJ, RDD, FW, DHT HLand MYS performed basic experiments. XTW, XQL, SPL, HLC, KZ, BZ, BW and DHT contributed to the data management and statistical analyses. JZC, XQL, and XTW wrote the manuscript. All authors contributed to the article and approved the submitted version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, X., Li, X., Cai, H. et al. Single-cell analysis indicating CCR8 modulates CD8+ tissue-resident memory T cells to attenuates rejection in transplantation. Sci Rep 16, 2936 (2026). https://doi.org/10.1038/s41598-025-32785-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-32785-2