Abstract
Bromodomain adjacent to zinc finger domain 2B (BAZ2B) is involved in chromatin remodeling, non-coding RNA regulation, and aging. However, its role in cancer remains unclear. Using multiple databases (TCGA, GEO, GTEx, TIMER2.0, cBioPortal), we found that BAZ2B is downregulated in many cancers, including breast cancer. Low BAZ2B correlates with poor prognosis, immune infiltration, and diagnostic potential, as shown by ROC and Kaplan–Meier analyses. Functionally, BAZ2B knockdown promoted breast cancer cell proliferation, colony formation, migration, and invasion, but suppressed apoptosis. Western blotting showed that BAZ2B depletion decreased Bax and p53, but increased Bcl-2. BAZ2B deficiency increased lactate production and upregulated glycolytic enzymes (LDHA, HK1, HK2, PKM2, PFK1). This suggests BAZ2B suppresses tumors by inhibiting glycolysis. Multi-omics integration further uncovered BAZ2B loss-induced co-activation of pro-inflammatory factor release and mTOR signaling, concurrent with suppression of apoptotic pathways and dysregulation of DNA repair machinery. Clinical validation using tissue microarrays confirmed reduced BAZ2B expression in tumors versus adjacent tissues, consistent with CPTAC proteomic data from UALCAN and immunohistochemical evidence from HPA. Collectively, BAZ2B constrains breast cancer progression by orchestrating proliferation, apoptosis, EMT, and glycolytic metabolism, positioning it as a promising therapeutic target and multi-cancer biomarker.
Similar content being viewed by others
Introduction
Breast cancer is the most common cancer in women worldwide1. Recently, breast cancer surpassed lung cancer as the most frequently diagnosed cancer2. Cancer progression burdens societal productivity and economies3. Early screening, diagnosis, and treatment can reduce these losses. Preventing cancer through etiology-based strategies may lower incidence and mortality. Rational healthcare systems can also ease economic burdens4,5. The WHO Global Cancer Report estimates that investments in cancer services between 2020 and 2030 could prevent approximately 7.3 million cancer-related deaths, expanding health benefits for individuals and families6. Notably, the U.S. "Cancer Moonshot Initiative" relaunched in February 2022 aims to accelerate cancer research, emphasizing a 50% improvement in treatment efficacy alongside mortality reduction. The WHO7 further advocates a 25% relative reduction in mortality from cardiovascular diseases, cancers, diabetes, and chronic respiratory diseases by 2025. Breast cancer prognosis is dynamically regulated by multifactorial interactions, where variables such as age, genetic predisposition, and lifestyle influence early diagnosis and therapeutic windows through complex mechanisms8. Molecularly, tumor heterogeneity drives subtype-specific dysregulation of key signaling pathways like PI3K/AKT/mTOR9, Despite advances in targeted therapies (e.g., Alpelisib), stromal cell-mediated paracrine signaling within the tumor microenvironment often induces adaptive resistance10. Concurrently, glutamine-dependent metabolic reprogramming11 and enhanced mitochondrial oxidative phosphorylation not only sustain cancer stemness but also promote immune evasion. However, metabolism-targeted therapies face challenges due to toxicity and inconsistent efficacy12. These biological complexities underscore the urgency of advancing fundamental research and translational applications in breast oncology.
BAZ2B (also designated WALp4), a 240-kDa protein within the bromodomain-containing family, functions in chromatin remodeling13 and non-coding RNA regulation. This multidomain histone-binding protein harbors two evolutionarily conserved reader modules—a plant homeodomain (PHD) and a bromodomain (BRD)—connected by disordered linkers. The BRD, a validated therapeutic target, has spurred development of small-molecule inhibitors14,15,16,17,18,19,20. BAZ2B additionally regulates hematopoietic development and pluripotent stem cell reprogramming21. While prior studies associate BAZ2B with cancer risk and cellular differentiation22, pan-cancer evidence linking BAZ2B to diverse tumor types remains lacking.
In this study, we conducted the first comprehensive analysis of BAZ2B expression profiles and survival correlations across cancers using TCGA and GEO datasets. Multidimensional parameters—including gene expression, DNA methylation, mutational landscape, and immune infiltration—were integrated to elucidate BAZ2B’s role in oncogenesis. Functional validation in breast cancer models confirmed its regulatory effects on malignant progression. Our findings systematically delineate BAZ2B’s pan-cancer significance and establish its oncogenic relevance in breast cancer.
Materials and methods
Gene expression analysis
The expression levels of BAZ2B between cancer tissues and adjacent normal tissues were investigated using TIMER2.023 (http://timer.cistrome.org/). We analyzed unpaired samples from GTEx and TCGA using R (v3.6.3) and visualized them with "ggplot224. Gene expression levels were displayed on a log2(TPM + 1) scale, where TPM (Transcripts Per Kilobase Million) represents transcripts per million.
Differences in BAZ2B protein expression across six cancer types—breast cancer (BRCA), glioblastoma multiforme (GBM), head and neck squamous cell carcinoma (HNSC), liver cancer (LIHC), pancreatic cancer (PAAD), and uterine corpus endometrial carcinoma (UCEC)—were compared using the CPTAC (Clinical Proteomic Tumor Analysis Consortium) analysis module in the UALCAN database25 (https://ualcan.path.uab.edu/). The expression patterns of BAZ2B (NP_001276904.1) across protein stages and its phosphorylation sites (T1247, S465, T542, T547, S552) were further analyzed to investigate their differences between primary tumor tissues and normal tissues.
Additionally, immunohistochemical staining intensities of BAZ2B in tissues including breast, pharynx, liver, uterus, pancreas, and cerebral cortex were evaluated using the Human Protein Atlas (HPA) database (https://www.proteinatlas.org/).
Survival prognostic and diagnostic analyses
The association between BAZ2B expression and overall survival (OS) was evaluated using the Kaplan–Meier Plotter database (https://kmplot.com/analysis/). Patients surviving beyond the predefined follow-up threshold were excluded, with the optimal cutoff value automatically selected and validated via the log-rank test. A statistically significant threshold was defined as a P-value < 0.05. RNA sequencing (RNA-seq) data from TCGA (The Cancer Genome Atlas) and GTEx (Genotype-Tissue Expression) were analyzed and visualized using the R packages “pROC” and “ggplot2”. The area under the receiver operating characteristic (ROC) curve (AUC > 0.5) was calculated to assess diagnostic and prognostic potential.
DNA methylation and mutation analysis
RNA sequencing (RNA-seq) and Methylation450 data from TCGA (The Cancer Genome Atlas) were analyzed and visualized using R (version 3.6.3) and the “ggplot2” package. Mutation profiles of BAZ2B across 32 cancer types—including gene alteration frequency, mutation types, and mutation sites—were retrieved through the “Quick Search Beta!” option in cBioPortal (https://www.cbioportal.org/) by querying “BAZ2B”. Specific mutation sites and three-dimensional protein structural details were examined in the “Mutations” module. Clinical outcomes of BAZ2B mutations in BRCA (breast cancer), including overall survival (OS), disease-specific survival (DSS), progression-free survival (PFS), and disease-free survival (DFS), were further analyzed using the “Comparison/Survival” module.
Immune infiltration analysis
The relationship between BAZ2B gene expression and immune infiltration was investigated using the “Immune-Gene” module of the TIMER 2.0 database (http://timer.cistrome.org/), which analyzed tumor expression profiles from TCGA (The Cancer Genome Atlas).
Enrichment analysis
We built a PPI network for BAZ2B using the STRING database. The search parameters included entering “BAZ2B” in the “protein name” module, selecting “Homo sapiens” under “Organism”, and configuring the following settings: network type (“full STRING network”), edge meaning (“types of interaction evidence”), active interaction sources (“Experiments, Co-expression, Neighborhood, Co-occurrence, Textmining”), minimum interaction score [“low confidence (0.150)”], and maximum interactors (“no more than 10 interactors in the first shell, 30 in the second shell”).
To predict BAZ2B-binding proteins, the “Similar Gene Analysis” module in GEPIA2.0 (http://gepia.cancer-pku.cn/index.html) was utilized to identify the top 100 genes co-expressed with BAZ2B in TCGA tumor and normal tissues. Pearson correlation analysis was subsequently performed for the top six BAZ2B-associated genes. RNA-seq data from TCGA projects (BLCA [bladder urothelial carcinoma], BRCA [breast cancer], HNSC [head and neck squamous cell carcinoma], LUAD [lung adenocarcinoma], and UCEC [uterine corpus endometrial carcinoma]) were analyzed to evaluate co-expression patterns among BAZ2B, ARID4A, PIKFYVE, CHD9, KMT2A, CARF, and NR2C2. Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway26,27,28 analyses were visualized using the R packages “ggplot2” and “clusterProfiler”.
Cell culture and transfection
The human normal mammary epithelial cell line MCF-10A and breast cancer cell lines MDA-MB-453, HCC38, MDA-MB-231, MCF7, and SK-BR-3 (authenticated by short tandem repeat [STR] profiling) were provided by Saier Biotechnology Laboratory (Tianjin, China). Cells were cultured in RPMI-1640 or DMEM (high-glucose) medium (GIBCO BRL, USA) supplemented with 10% fetal bovine serum (FBS; GIBCO) at 37 °C under a humidified atmosphere of 5% CO2. Transfection of siRNA (purchased from Saier Biotechnology Co., Ltd., Tianjin, China) into the aforementioned cell lines was performed using Lipofectamine 2000 Reagent (Invitrogen, USA). The siRNA sequences were as follows: 5′-CAGTGATGTTTGGCCAAGATCGTTA-3′ (si-BAZ2B) and 5′-UUCUCCGAACGUGUCACGUTT-3′ (siRNA-NC).
Reverse transcription-quantitative (RT-q)PCR
Total RNA was extracted from cells using TRIzol Reagent (Invitrogen, USA). Reverse transcription of total RNA was performed using a Reverse Transcription Kit (Takara, Japan). Quantitative real-time PCR (qRT-PCR) was carried out on an iQ5 Real-Time PCR System (Applied Biosystems, USA) with SYBR Premix Ex Taq (Takara, Japan). The thermal cycling conditions were as follows: initial denaturation at 94 °C for 4 min, followed by 40 cycles of denaturation at 94 °C for 30 s, annealing at 58 °C for 30 s, and extension at 72 °C for 30 s. Relative expression of BAZ2B was calculated using the 2−ΔΔCt method and normalized to β-actin. The primer sequences used for qRT-PCR are listed in the following Table 1:
CCK8
Cells transfected with siRNA were seeded into 96-well plates at a density of 1 × 103 cells/well in 100 μL medium. At 24 h, 48 h, and 72 h time points, 10 μL of CCK8 solution was added to each well. After further incubation for 2 h, absorbance (OD) values were measured at a wavelength of 450 nm using a microplate reader.
Western blotting
Cells were collected for protein extraction, and protein concentrations were quantified using a BCA Protein Assay Kit (Solarbio, Beijing, China). Proteins were separated by sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) and transferred onto polyvinylidene fluoride (PVDF) membranes (Millipore, USA). The membranes were blocked in 5% non-fat milk for 2 h at room temperature with gentle shaking. Subsequently, membranes were cut into strips according to the molecular weights of target proteins and incubated overnight at 4 °C with gentle shaking in blocking buffer containing the following primary antibodies: GAPDH, Bax, Bcl-2, p53, E-cadherin, N-cadherin, vimentin, and snail. Membranes were then incubated with corresponding horseradish peroxidase (HRP)-conjugated secondary antibodies for 1 h at room temperature. Protein bands were visualized using an enhanced chemiluminescence (ECL) substrate (PerkinElmer, USA).
Migration and invasion assays
Invasion and migration assays were performed using Transwell chambers with or without Matrigel coating. Cells were digested, resuspended in serum-free medium, and seeded into the upper chamber, while the lower compartment of the 24-well plate was filled with complete medium containing 10% fetal bovine serum (FBS). The chambers were incubated at 37 °C for 24 h. After incubation, non-invading/migrating cells on the upper surface of the membrane were removed with a cotton swab, and residual cells were gently washed with phosphate-buffered saline (PBS). Cells adhering to the lower surface were fixed with a methanol-glacial acetic acid mixture (3:1 ratio) for 30 min and stained with crystal violet staining solution (Solarbio, G1063) for 15 min. The chambers were rinsed thoroughly, mounted on glass slides, and visualized under a microscope (OLYMPUS, Japan). Cells in three randomly selected fields per chamber were imaged and quantified.
Colony formation assay
Suspended cells (500 cells/well) were seeded into 12-well plates and cultured in 2 mL complete medium at 37 °C with 5% CO2 for 1–2 weeks. When single-cell-derived colonies exceeded 50 cells per clone, the medium was aspirated, and cells were rinsed once with PBS. Cells were fixed with 500 μL of 4% paraformaldehyde at 4 °C for 30 min, followed by PBS washing. Colonies were stained with 500 μL of 0.1% crystal violet for 10 min and rinsed with PBS. Plates were air-dried, photographed, and colonies containing > 50 cells were counted microscopically.
Apoptosis analysis by flow cytometry
Transfected SK-BR-3 breast cancer cells were digested with 0.25% trypsin, resuspended (5 × 104–1 × 105 cells), and centrifuged at 800 × g for 10 min. The pellet was gently resuspended in 195 μL Annexin V-FITC binding buffer (Beyotime, China), mixed with 10 μL propidium iodide (PI), and incubated in the dark at room temperature for 10–20 min. Samples were kept on ice and analyzed immediately using a flow cytometer.
Lactate assay
Cells were collected 48 h post-transfection, lysed by sonication, and centrifuged at 2500 rpm for 10 min. Supernatants were assayed as follows: blank controls, standards, and test samples were prepared. Enzyme working solution and chromogenic substrate were added sequentially, vortexed, and incubated at 37 °C for 10 min. Reactions were terminated, and absorbance (A) was measured at 530 nm using a 1 cm light-path cuvette with distilled water as zero reference.
Immunohistochemistry
This study was approved by the Ethics Committees of North China University of Science and Technology, and Shanghai YEPCOME biotechnology Co.,Ltd. All participants provided informed consent. Paraffin-embedded sections were deparaffinized in xylene, dehydrated in ethanol, and washed with PBS. Endogenous peroxidase activity was blocked with 3% H2O2 (25 min, room temperature). Antigen retrieval was performed by high-pressure heating. Sections were blocked, incubated with primary anti-BAZ2B antibody (1:200 dilution, HPA019819, ATLAS) at 4 °C overnight, washed, and treated with secondary antibody (30 min, room temperature). Signal development utilized 3,3′-diaminobenzidine (DAB), followed by hematoxylin counterstaining.
Transcriptome sequencing analysis
RNA extraction was performed using Trizol reagent (Invitrogen, USA) at a density of 5 × 10⁶ cells/mL. Transcriptome sequencing was conducted by Zhongke Xinshengming Biotechnology Co., Ltd. (China) following established protocols29,30,31.
Tandem mass tag(TMT)-labeled quantitative proteomics
Cells (1 × 107–10 × 107) were lysed in RIPA buffer (Beyotime, China) and subjected to 4D label-free quantitative proteomics by Zhongke Xinshengming Biotechnology Co., Ltd.32,33,34,35.
Statistical analysis
Data were analyzed using GraphPad Prism 8.2.1. Student’s t-test or one-way ANOVA was applied, with results expressed as mean ± SD. A. P < 0.05 was considered statistically significant.
Ethics approval
This study was performed in line with the principles of the Declaration of Helsinki. All studies performed with human tissue specimens were approved by the Ethics Committee of North China University of Science and Technology, and Shanghai YEPCOME biotechnology Co.,Ltd.
Consent publication
All patients provided their informed consent.
Results
BAZ2B expression in human pan-cancer
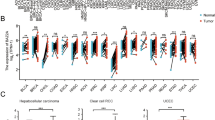
We analyzed TCGA and GEO databases to compare BAZ2B expression between tumors and normal tissues. BAZ2B mRNA was significantly lower in multiple cancers, including bladder cancer (BLCA), breast cancer (BRCA), chromophobe renal cell carcinoma (KICH), clear cell renal carcinoma (KIRC), lung squamous cell carcinoma (LUSC), prostate adenocarcinoma (PRAD), thyroid carcinoma (THCA), and uterine corpus endometrial carcinoma (UCEC), compared to normal tissues (Fig. 1A). As shown in Supplementary Fig. 1A, BAZ2B expression was markedly lower in tumor tissues of BLCA, BRCA, KICH, LUSC, and UCEC compared to paired adjacent normal tissues. For cancers lacking matched normal tissues, GTEx dataset-derived normal tissues were used as controls, further confirming BAZ2B downregulation in BLCA, BRCA, KICH, lung adenocarcinoma (LUAD), LUSC, PRAD, THCA, and UCEC (Supplementary Fig. 1B).
Expression of BAZ2B gene across various cancer types. (A) We analyzed the mRNA expression levels of BAZ2B in tumor and adjacent normal tissues using the TIMER database (http://timer.cistrome.org/). *: P < 0.05; **: P < 0.01; ***: P < 0.001. A P-value < 0.05 was considered statistically significant.
CPTAC analysis demonstrated that BAZ2B protein levels were downregulated in BRCA, head and neck squamous cell carcinoma (HNSC), and liver hepatocellular carcinoma (LIHC), but upregulated in UCEC, pancreatic adenocarcinoma (PAAD), and glioblastoma (GBM) compared to normal tissues (Fig. 2A–F). Human Protein Atlas IHC staining corroborated low BAZ2B expression in breast, throat, and liver tumors, but high expression in uterine, pancreatic, and neural tumors (Fig. 2A–F). CPTAC data further revealed a significant negative correlation between BAZ2B expression and pathological staging in BRCA (P < 0.05), but positive correlations in PAAD and UCEC (Supplementary Fig. 2A–C). BAZ2B phosphorylation levels also correlated with BRCA and HNSC progression (Supplementary Fig. 2D–E).
Expression of BAZ2B protein in different tumors. (A–F) BAZ2B protein expression in BRCA, HNSC, PAAD, GBM, LIHC, and UCEC was analyzed based on CPTAC data, along with immunohistochemical (IHC) analysis of BAZ2B protein in tumor versus normal tissues from the HPA database. (G–I) The relationship between BAZ2B expression levels and pathological stages was explored using the UALCAN database. (J–K) Differences in BAZ2B protein phosphorylation levels between tumor and normal tissues in BRCA and HNSC were analyzed using the CPTAC module. *: P < 0.05; **: P < 0.01; ***: P < 0.001.
Prognostic and diagnostic value of BAZ2B
Kaplan–Meier analysis via the Kaplan–Meier Plotter database indicated that low BAZ2B expression was associated with shorter overall survival (OS) in BLCA, BRCA, HNSC, LUAD, and THCA patients (Fig. 3A). Receiver operating characteristic (ROC) curve analysis demonstrated diagnostic accuracy for BAZ2B in BLCA (AUC = 0.847), BRCA (0.741), HNSC (0.585), LUAD (0.800), THCA (0.650), and UCEC (0.932) (Fig. 3B). These findings suggest BAZ2B as a potential prognostic biomarker across cancers.
Prognostic and diagnostic analysis of BAZ2B expression in cancers. (A) Overall survival (OS) analysis of the BAZ2B gene in TCGA datasets (https://kmplot.com/analysis/). (B) Receiver operating characteristic (ROC) analysis of BAZ2B gene expression in the TCGA database.
Epigenetic regulation and genetic alterations
Promoter DNA methylation is a pivotal epigenetic mechanism implicated in carcinogenesis, often associated with tumor metastasis and unfavorable prognosis36,37,38. Our analysis first interrogated the methylation landscape of BAZ2B, revealing significant hypermethylation at its promoter region across multiple cancer types, including BLCA, BRCA, HNSC, LUAD, and UCEC, compared to normal tissues (Fig. 4A). This epigenetic silencing was functionally corroborated by a significant negative correlation between BAZ2B expression and its promoter methylation levels, as quantified using two independent probes (cg19903071 and cg04706995) on TCGA RNA-seq and Methylation450 datasets (Fig. 4B).
Promoter methylation and mutational characteristics of the BAZ2B gene in tumor tissues. (A) Methylation levels of BAZ2B in normal and primary tumor tissues were analyzed using UALCAN. (B) DNA methylation of BAZ2B in BLCA, BRCA, HNSC, LUAD, and UCEC. (C,D) Mutation frequency (C) and mutation sites (D) of the BAZ2B gene. (E) Mutation sites, types, and corresponding case numbers in TCGA. (F) Potential correlations between mutational status and OS, DSS, DFS, and PFS in BRCA were analyzed using the cBioPortal database.
Beyond epigenetic regulation, we comprehensively characterized the genetic alteration profile of BAZ2B. Analysis of TCGA cohorts unveiled a spectrum of aberrations—including mutations, structural variants, amplifications, and deletions—with mutations being the most prevalent (Fig. 4C). Notably, endometrial carcinoma (UCEC) exhibited the highest alteration frequency (> 12%), predominantly driven by mutations. Profiling of specific mutation sites identified truncating mutations as the major type, with a recurrent K2160Sfs*8 frameshift mutation observed in single cases of UCEC, BRCA, and COAD (Supplementary Fig. 3A). To assess the clinical impact of these genetic changes, we evaluated survival outcomes. In breast cancer, patients harboring BAZ2B alterations demonstrated significantly poorer overall survival (OS, P = 0.0345) and disease-specific survival (DSS, P = 8.732e−3) compared to those without alterations. However, no significant association was found with disease-free survival (DFS, P = 0.200) or progression-free survival (PFS, P = 0.0590) (Fig. 4D).
Association of BAZ2B gene expression and immune infiltration
Next, we employed algorithms including TIMER, XCELL, EPIC, and TIDE to evaluate the potential correlation between BAZ2B expression and immune cell infiltration across various cancer types. The results revealed that BAZ2B expression in most cancers (e.g., BLCA, BRCA, HNSC, LUAD, and UCEC) showed significant associations with CD8 + T cells, NK cells, dendritic cells (DCs), and regulatory T cells (Tregs) (Fig. 5A). TIMER-generated scatterplots further demonstrated that BAZ2B expression exhibited significant positive correlations with infiltration levels of CD8 + T cells, dendritic cells (DCs), and NK cells, while displaying significant negative correlations with regulatory T cells (Tregs) and mast cells (Supplementary Fig. 4A, Fig. 5B–E).
Correlation analysis between BAZ2B expression and tumor-infiltrating immune cells. (A) Correlation of BAZ2B expression with NK cell and CD8 + T cell infiltration using multiple algorithms from the TIMER2.0 database. (B–E) Associations between BAZ2B expression and immune cell infiltration in BRCA, HNSC, LUAD, and UCEC.
Enrichment analysis of proteins and genes associated with BAZ2B
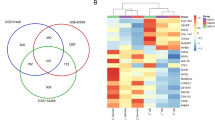
Next, we identified proteins interacting with BAZ2B using the STRING database, which revealed 40 interacting proteins. The protein–protein interaction (PPI) network is illustrated in Fig. 6A. Additionally, the top 100 co-expressed genes of BAZ2B across pan-cancer datasets were analyzed via GEPIA2.0, among which ARID4A, PIKFYVE, CHD9, KMT2A, CARF, and NR2C2 exhibited correlation coefficients exceeding 0.8 with BAZ2B in most cancer types (Fig. 6B). TCGA RNA-seq data further confirmed the co-expression of BAZ2B with ARID4A, PIKFYVE, CHD9, KMT2A, CARF, and NR2C2 in BLCA, BRCA, HNSC, LUAD, and UCEC (Fig. 6C). Concurrently, Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses demonstrated that BAZ2B-associated co-expressed genes in breast cancer were linked to aberrant transcriptional regulation and spinocerebellar ataxia (Supplementary Fig. 5A). Extended pathway relationships are depicted in Supplementary Fig. 5B.
Enrichment analysis of BAZ2B-related genes. (A) Protein interaction network of BAZ2B obtained from the STRING database (https://string-db.org/; http://gepia.cancer-pku.cn/index.html). (B) GEPIA2.0 revealed positive correlations between BAZ2B and six genes (GCC2, STAM2, CARF, ARID4A, BMPR2, and MBD5). (C) Co-expression analysis of BAZ2B and related genes in BRCA. (D,E) KEGG (Kyoto Encyclopedia of Genes and Genomes) and GO (Gene Ontology) enrichment analyses of BAZ2B and its related genes. *: P < 0.05; **: P < 0.01; ***: P < 0.001. A P-value < 0.05 was considered statistically significant.
BAZ2B is lowly expressed in breast cancer and promotes the malignant behavior of breast cancer cells
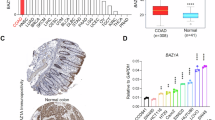
We focused on breast cancer (BRCA) cells to study BAZ2B. First, we measured BAZ2B mRNA expression levels in five breast cancer cell lines (MDA-MB-453, HCC38、, DA-MB-231, MCF7 and SK-BR-3) using RT-qPCR, identifying SK-BR-3 cells as having the highest BAZ2B expression (Fig. 7A), which were therefore selected for further experiments. We designed siRNAs targeting BAZ2B; qRT-PCR results confirmed that the interfering RNA si-BAZ2B-2 exhibited the most effective knockdown (Fig. 7B) and was consequently used in subsequent assays. Immunohistochemical (IHC) analysis of 80 paired breast cancer tissues and adjacent non-tumor tissues revealed significantly lower BAZ2B expression in 72 cases (90%) of cancerous tissues compared to their adjacent counterparts (Fig. 7C). Following transfection of si-BAZ2B-2 into SK-BR-3 cells (with NC-siRNA as a negative control), CCK-8 cell proliferation assays and colony formation assays demonstrated that BAZ2B downregulation significantly promoted cell proliferation and clonogenic capacity (Fig. 7D–E). Transwell assays further confirmed that BAZ2B knockdown enhanced cell migration and invasion capabilities (Fig. 7F–G). Flow cytometry analysis revealed that BAZ2B positively regulates apoptosis (Fig. 7H), and Western blot detection showed that BAZ2B knockdown significantly decreased levels of the pro-apoptotic proteins Bax and p53 while markedly increasing expression of the anti-apoptotic protein Bcl-2 (Fig. 7I). Furthermore, results from detecting epithelial-mesenchymal transition (EMT)-related molecule levels provided additional support for BAZ2B's role in inhibiting cancer cell migration (Fig. 7J). Metabolic studies revealed that BAZ2B downregulation led to a significant increase in cellular lactate levels (Fig. 7K), accompanied by a marked upregulation in the mRNA expression of key glycolytic enzymes (LDHA, HK1, HK2, PKM2 and PFK1), indicating that BAZ2B knockdown significantly promotes the glycolytic pathway (Fig. 7L).
BAZ2B exhibits downregulated expression in breast cancer (BRCA) and correlates with malignant progression of the disease. (A) qRT-PCR analysis of BAZ2B mRNA expression in five breast cancer (BRCA) cell lines. (B) Validation of siRNA efficiency via qRT-PCR in SK-BR-3 cells. (C) Immunohistochemical detection of BAZ2B expression in cancerous and adjacent tissues. (D–G) Effects of BAZ2B downregulation on the proliferation (D), colony formation (E), migration (F), and invasion (G) of SK-BR-3 breast cancer cells (experimental details are provided in the Materials and Methods section). (H) Flow cytometry analysis of apoptosis in SK-BR-3 cells induced by BAZ2B knockdown (experimental details are provided in the Materials and Methods section). (I,J) Western blot analysis of the effects of BAZ2B on apoptosis- and EMT-associated molecules in SK-BR-3 cells transfected with siRNA (total protein was extracted post-transfection). (K) Measurement of lactate production following BAZ2B downregulation. (L) Real-time quantitative PCR determination of mRNA expression levels of the knocked-down BAZ2B gene and key glycolytic enzymes (HK1, HK2, LDHA, PKM2, PFK1).
Multiomics analysis of BAZ2B
To further validate the relationship between BAZ2B and cancer, SK-BR-3 breast cancer cells transfected with BAZ2B-siRNA or NC-siRNA were subjected to transcriptomic and proteomic analyses.
Transcriptomic sequencing identified 1,095 differentially expressed genes (DEGs). The top 30 DEGs (19 upregulated, 11 downregulated), ranked by p-value, included genes associated with inflammatory responses and cell cycle regulation. Significantly upregulated genes comprised SAA1/SAA2 (acute-phase response), WFDC2 (antimicrobial peptide), and LCN2, while downregulated genes included cell division-related NUF2 and ANLN (Fig. 8A).

Kanehisa Laboratories. (D–F) Proteomic analysis: Heatmap of differentially expressed proteins (D), KEGG pathway dynamics (E,F). (G) Venn diagram illustrating 68 overlapping DEGs between transcriptomic and proteomic datasets. (H) Sankey diagram visualizing regulatory associations between multi-omics data in key pathways (e.g., PI3K-AKT signaling, RNA transport), constructing a multi-dimensional framework of BAZ2B’s transcriptional-translational coupling in cellular functions.
Multi-omics analysis of BAZ2B. (A) Heatmap of differentially expressed genes (DEGs) showing transcriptomic profiles. (B,C) Significantly enriched upregulated and downregulated KEGG pathways (reproduced from the KEGG pathway database, including but not limited to: https://www.kegg.jp/entry/map04657, https://www.kegg.jp/entry/map04150, https://www.kegg.jp/entry/map04210, https://www.kegg.jp/entry/map04110, https://www.kegg.jp/entry/map04151, https://www.kegg.jp/entry/map03410). (C)
KEGG pathway enrichment analysis revealed coordinated activation of pro-tumorigenic pathways. The upregulated pathways were proliferation drivers, primarily ribosome biogenesis and the mTOR signaling pathway. Inflammatory remodeling: Aberrant activation of IL-17 signaling and arachidonic acid metabolism. Metabolic adaptation: Reprogramming of glutathione and tyrosine metabolism, indicative of enhanced oxidative stress resistance (Fig. 8B). Downregulated pathways exhibited three-tiered functional impairment: Genomic instability: Suppression of DNA repair mechanisms (mismatch repair, base excision repair, nucleotide excision repair), suggesting compromised DNA damage response. Tumor suppressor inactivation: Reduced activity of the Hippo signaling pathway (multiple species) and cellular senescence pathways. Apoptosis inhibition: Impaired clearance of aberrant cells via apoptosis pathway suppression (Fig. 8C).
Proteomic analysis identified 28 differentially expressed proteins (DEPs; > twofold change, Fig. 8D). Metabolic regulators HSD17B7 and FN1 were significantly upregulated, while lipid metabolism enzymes PGM5 and BASP1 were downregulated. KEGG pathway analysis highlighted multidimensional regulatory networks: Upregulation of steroid hormone biosynthesis and ovarian steroidogenesis pathways, coupled with HSD17B7 overexpression, suggested estrogen-dependent growth potentiation. Suppression of glycerolipid metabolism and PPAR signaling indicated lipid metabolic imbalance. Synergistic activation of PI3K-Akt and ECM-receptor interaction pathways drove cell cycle progression (PKMYT1↑) and EMT (FN1↑). IL-17 signaling and complement/coagulation cascades were inversely regulated, reshaping an immunosuppressive microenvironment. Inhibition of drug-metabolizing enzymes (CYP450) implied potential metabolic drug resistance (Fig. 8E–F).
Integrated multi-omics analysis uncovered a BAZ2B deficiency-induced cross-omics regulatory network. Among 960 DEGs (transcriptome) and 1,565 DEPs (proteome), 87 overlapping molecules were identified, with 52 showing concordant expression (Fig. 8G). KEGG enrichment analysis showed persistent activation of IL-17 signaling (fold enrichment = 13.33), with key genes including S100A7/A8 and LCN2. Dysregulated oxidative phosphorylation (NDUFA11, ATP5F1D↑; fold enrichment = 6.88) alongside suppression of butyrate metabolism (ACAT2↓; Fig. 8H). ACAT2 deficiency correlated with reduced H3K9ac levels, suggesting altered chromatin accessibility. This cascade illustrates BAZ2B’s tumor-suppressive role through coordinated regulation of inflammatory responses, metabolic homeostasis, and epigenetic remodeling.
Discussion
Cancer is fundamentally a genetic disease caused by mutations in genes controlling cellular functions, particularly those regulating cell growth and division39. In early cancer detection, large numbers of oncogenes were identified through functional assays of tumor genetic material, and a subset of tumor suppressor genes was discovered via loss of heterozygosity (LOH)40. Pan-cancer analysis refers to the examination of genomic expression, prognosis, and mutations across different tumors using multiple databases, providing new clues for cancer prevention and diagnostic biomarkers41. We performed a comprehensive analysis of the BAZ2B gene across 33 distinct tumor types. We found that BAZ2B mRNA is downregulated in many cancers and correlates with pathological stage. This suggests BAZ2B may function as a tumor suppressor, exerting inhibitory effects on malignant progression. Interestingly, while our pan-cancer analysis revealed consistent downregulation of BAZ2B in multiple malignancies including LUAD, a previous study by Li et al. reported upregulation of BAZ2B in LUAD42. This discrepancy may stem from differences in data processing, normal tissue selection, or statistical thresholds. Nevertheless, our functional assays in breast cancer models strongly support a tumor-suppressive role for BAZ2B, which is consistent with its downregulation in the majority of cancers we analyzed. Further investigations are warranted to elucidate the context-dependent roles of BAZ2B across different tumor types.
BAZ2B knockdown increased lactate and promoted glycolysis, suggesting it drives the Warburg effect43. Lactate accumulation acidifies the tumor microenvironment44, activating proteases such as MMPs to degrade the extracellular matrix, thereby metabolically explaining the enhanced migratory/invasive capacities observed upon BAZ2B knockdown; Upregulation of key glycolytic enzymes (e.g., HK2, PKM2) not only accelerates glucose breakdown for energy production but also provides accumulated intermediate metabolites as precursors for nucleic acid/lipid synthesis, thus supporting the BAZ2B deficiency-induced proliferation and clonogenic phenotypes. Notably, the increased expression of LDHA—a target gene of the hypoxia-inducible factor HIF-1α—suggests that BAZ2B likely regulates glycolysis through the HIF-1α pathway. This observation forms a mechanistic loop with the multi-omics analysis indicating mTOR pathway activation (known to stabilize HIF-1α). In conclusion, BAZ2B functions as a “metabolic checkpoint” by suppressing the expression of rate-limiting glycolytic enzymes. Its inactivation reprograms tumor cell metabolism, thereby driving malignant progression.
While our study provides compelling evidence for the tumor-suppressive role of BAZ2B through loss-of-function approaches, we acknowledge that overexpression rescue experiments would further strengthen these findings. Unfortunately, due to resource constraints, such experiments were not feasible in the current study. Nonetheless, based on the consistent phenotypic changes observed upon BAZ2B knockdown—including enhanced proliferation, migration, invasion, glycolytic flux, and apoptosis evasion—we hypothesize that BAZ2B re-expression would likely reverse these malignant phenotypes. Future studies employing inducible expression systems or stable transgenic models in BAZ2B-low backgrounds could definitively test this hypothesis and provide mechanistic insights into its reversible tumor-suppressive functions. Such approaches would also help elucidate whether BAZ2B restoration could synergize with existing therapies, particularly in metabolic or immune-contextualized treatment regimens.
Conclusion
This study demonstrates that the expression level of BAZ2B correlates with the prognosis of breast cancer patients and suggests its potential as a diagnostic marker. In breast cancer cells, BAZ2B exerts a tumor-suppressive role: its downregulation significantly reduced apoptosis while promoting cell proliferation, migration, invasion, and glycolysis, manifested by increased lactate production and upregulation of key glycolytic enzymes. Collectively, these findings indicate that BAZ2B represents a promising therapeutic intervention target in breast cancer.
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information files. For additional details, data are available from the corresponding author upon reasonable request. The mass spectrometry proteomics data has been uploaded to the Integrated Proteome Resources center (http://proteomecentral.proteomexchange.org/cgi/GetDataset?ID=PCD054319) with the dataset identifier PXD054319. The raw RNA-seq data has been deposited in NCBI’s Sequence Read Archive (https://submit.ncbi.nlm.nih.gov/subs/sra/SUB15363302/overview) with the accession number is PRJNA1271767. Other data are available from the corresponding author.
References
Ensenyat-Mendez, M. et al. Current triple-negative breast cancer subtypes: Dissecting the most aggressive form of breast cancer. Front. Oncol. 11, 681476 (2021).
Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249 (2021).
Kocarnik, J. M. et al. Cancer incidence, mortality, years of life lost, years lived with disability, and disability-adjusted life years for 29 cancer groups from 2010 to 2019: A systematic analysis for the global burden of disease study 2019. JAMA Oncol. 8, 420–444 (2022).
de Martel, C., Georges, D., Bray, F., Ferlay, J. & Clifford, G. M. Global burden of cancer attributable to infections in 2018: A worldwide incidence analysis. Lancet Glob. Health 8, e180–e190 (2020).
Sung, H., Siegel, R. L., Rosenberg, P. S. & Jemal, A. Emerging cancer trends among young adults in the USA: Analysis of a population-based cancer registry. The Lancet. Public health 4, e137–e147 (2019).
Chen, S. et al. Estimates and projections of the global economic cost of 29 cancers in 204 countries and territories from 2020 to 2050. JAMA Oncol. 9, 465–472 (2023).
Schüz, J. et al. European Code against Cancer 4th Edition: 12 ways to reduce your cancer risk. Cancer Epidemiol. 39(Suppl 1), S1-10 (2015).
Zhou, M. et al. Mendelian randomization analysis revealed potential metabolic causal factors for breast cancer. Sci. Rep. 13, 14290 (2023).
Miricescu, D. et al. PI3K/AKT/mTOR signaling pathway in breast cancer: From molecular landscape to clinical aspects. Int. J. Mol. Sci. 22, 173 (2020).
André, F. et al. Alpelisib for PIK3CA-mutated, hormone receptor-positive advanced breast cancer. N. Engl. J. Med. 380, 1929–1940 (2019).
Li, T., Copeland, C. & Le, A. Glutamine metabolism in cancer. Adv. Exp. Med. Biol. 1311, 17–38 (2021).
Stine, Z. E., Schug, Z. T., Salvino, J. M. & Dang, C. V. Targeting cancer metabolism in the era of precision oncology. Nat. Rev. Drug Discovery 21, 141–162 (2022).
Feng, Y. et al. Crystal structure of the BAZ2B TAM domain. Heliyon 8, e09873 (2022).
Amato, A., Lucas, X., Bortoluzzi, A., Wright, D. & Ciulli, A. Targeting ligandable pockets on plant homeodomain (PHD) zinc finger domains by a fragment-based approach. ACS Chem. Biol. 13, 915–921 (2018).
Chen, P. et al. Discovery and characterization of GSK2801, a selective chemical probe for the bromodomains BAZ2A and BAZ2B. J. Med. Chem. 59, 1410–1424 (2016).
Dalle Vedove, A. et al. Structural analysis of small-molecule binding to the BAZ2A and BAZ2B Bromodomains. ChemMedChem 13, 1479–1487 (2018).
Drouin, L. et al. Structure enabled design of BAZ2-ICR, A chemical probe targeting the bromodomains of BAZ2A and BAZ2B. J. Med. Chem. 58, 2553–2559 (2015).
Ferguson, F. M. et al. Binding hotspots of BAZ2B bromodomain: Histone interaction revealed by solution NMR driven docking. Biochemistry 53, 6706–6716 (2014).
Marchand, J. R., Lolli, G. & Caflisch, A. Derivatives of 3-Amino-2-methylpyridine as BAZ2B bromodomain ligands. in silico discovery and in crystallo validation. J. Med. Chem. 59, 9919–9927 (2016).
Unzue, A. et al. The “Gatekeeper” residue influences the mode of binding of acetyl indoles to bromodomains. J. Med. Chem. 59, 3087–3097 (2016).
Arumugam, K. et al. The master regulator protein BAZ2B can reprogram human hematopoietic lineage-committed progenitors into a multipotent state. Cell Rep. 33, 108474 (2020).
Bingen, J. M., Clark, L. V., Band, M. R., Munzir, I. & Carrithers, M. D. Differential DNA methylation associated with multiple sclerosis and disease modifying treatments in an underrepresented minority population. Front Genet. 13, 1058817 (2022).
Li, T. et al. TIMER2.0 for analysis of tumor-infiltrating immune cells. Nucleic Acids Res. 48, W509-w514 (2020).
Blum, A., Wang, P. & Zenklusen, J. C. SnapShot: TCGA-analyzed tumors. Cell 173, 530 (2018).
Chandrashekar, D. S. et al. UALCAN: An update to the integrated cancer data analysis platform. Neoplasia 25, 18–27 (2022).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Prot. Sci. Publ. Prot. Soc. 28, 1947–1951 (2019).
Kanehisa, M., Furumichi, M., Sato, Y., Matsuura, Y. & Ishiguro-Watanabe, M. KEGG: Biological systems database as a model of the real world. Nucleic Acids Res. 53, D672-d677 (2025).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Anders, S., Pyl, P. T. & Huber, W. HTSeq–a Python framework to work with high-throughput sequencing data. Bioinformatics (Oxford, England) 31, 166–169 (2015).
Gu, Z., Eils, R. & Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics (Oxford, England) 32, 2847–2849 (2016).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics (Oxford, England) 26, 139–140 (2010).
Cox, J. et al. Accurate proteome-wide label-free quantification by delayed normalization and maximal peptide ratio extraction, termed MaxLFQ. Mol. Cel. Proteomics : MCP 13, 2513–2526 (2014).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Cox, J. et al. Andromeda: A peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 10, 1794–1805 (2011).
Wiśniewski, J. R., Zougman, A., Nagaraj, N. & Mann, M. Universal sample preparation method for proteome analysis. Nat. Methods 6, 359–362 (2009).
Yamashita, K. et al. Epigenetic biomarkers of promoter DNA methylation in the new era of cancer treatment. Cancer Sci. 109, 3695–3706 (2018).
Chen, Y. C. et al. Increased S100A15 expression and decreased DNA methylation of its gene promoter are involved in high metastasis potential and poor outcome of lung adenocarcinoma. Oncotarget 8, 45710–45724 (2017).
Sengupta, D. et al. DNA methylation and not H3K4 trimethylation dictates the expression status of miR-152 gene which inhibits migration of breast cancer cells via DNMT1/CDH1 loop. Exp. Cell Res. 346, 176–187 (2016).
Vogelstein, B. & Kinzler, K. W. Cancer genes and the pathways they control. Nat. Med. 10, 789–799 (2004).
Baker, S. J. et al. Chromosome 17 deletions and p53 gene mutations in colorectal carcinomas. Science 244, 217–221 (1989).
Lv, Z. et al. Zic family member 2 (ZIC2): A potential diagnostic and prognostic biomarker for pan-cancer. Front Mol. Biosci. 8, 631067 (2021).
Li, Y. et al. The emerging role of ISWI chromatin remodeling complexes in cancer. J. Exp. Clin. Cancer Res. 40, 346 (2021).
Vander Heiden, M. G., Cantley, L. C. & Thompson, C. B. Understanding the Warburg effect: The metabolic requirements of cell proliferation. Science 324, 1029–1033 (2009).
Estrella, V. et al. Acidity generated by the tumor microenvironment drives local invasion. Can. Res. 73, 1524–1535 (2013).
Acknowledgements
We would like to extend our appreciation to Tianjin Saier Biology Technology Co., Ltd. and Dr. Guo Liming for their valuable technical support throughout this study.
Funding
This study was supported by the Medical Science Research Project of Hebei Provincial Health Commission (Grant No. 20250357) and the National Natural Science Foundation of China (Grant No. 82173065), Hebei Provincial Central Government Guiding Fund for Local Science and Technology Development Program (246Z7740G).
Author information
Authors and Affiliations
Contributions
Writing—review & editing, J.G and S.W.; Writing—original draft. J.W and B.Z.; Methodology, J.Z. and G.Q.; Investigation, Z.S. and R.G.; Software, Formal analysis, M.Z. and L.H.; Supervision, L.Z. and Y.L.; Supervision, Resources, Funding acquisition, Y.L. and Y.Z.; Conceptualization, P.D. and Y.Z. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Guo, J., Wang, S., Liu, Y. et al. Pan-cancer and multi-omics analyses reveal the diagnostic and prognostic value of BAZ2B in cancer. Sci Rep 16, 3578 (2026). https://doi.org/10.1038/s41598-025-33622-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-33622-2