Abstract
Colorectal cancer (CRC) is the second most frequently diagnosed cancer worldwide and represents a major challenge for public health. Despite advances in molecular profiling, important gaps remain in our understanding of tumorigenesis and the regulatory mechanisms underlying CRC progression. The most widely adopted classification system is the Consensus Molecular Subtypes (CMS), which stratifies CRC into four biologically distinct subtypes. We investigated the role of A-to-I RNA editing across CMS subtypes in a cohort of 100 CRC patients at various disease stages. Bulk RNA-seq data were analyzed using REDItools to detect editing events, focusing on both recoding sites and edits within repetitive elements, such as Alu sequences. Furthermore, expression levels of the ADAR enzyme family were assessed, and deconvolution analyses were performed on single-cell RNA-seq data from an independent cohort of stage II CRC patients to characterize editing activity within the tumor microenvironment (TME). Competitive endogenous RNA (ceRNA) networks, specific to each CMS subtype, were constructed based on editing events in repetitive elements. A multivariate Cox proportional hazards model was applied to evaluate associations with overall survival (OS). We observed statistically significant differences in ADARB1 expression across CMS subtypes. Single-cell RNA-seq data revealed subtype-specific distribution patterns of ADAR enzymes within the TME. Analysis of editing events showed subtype-specific signatures in both known cancer-related genes (e.g., COPA, CADPS, IGFBP7) and novel candidates (ZNF552, RALGPS1). Editing in repetitive elements informed the construction of distinct ceRNA networks for each CMS subtype, suggesting different post-transcriptional regulatory mechanisms. Survival analysis identified three variables significantly associated with OS, independent of CMS classification and clinical stage: ADARB1 expression, and editing events in NOP14-AS1 (chr4:2960236; p = 0.036; HR = 0.0069), previously linked to 5-FU sensitivity, and ST7-AS2 (chr4:117120557). This study underscores the biological relevance of RNA editing in CRC, highlighting its impact on chemoresistance, the tumor microenvironment, and subtype-specific gene regulation. Our findings suggest that RNA editing represents a critical post-transcriptional regulatory layer in CRC and holds potential as a biomarker and therapeutic target.
Similar content being viewed by others
Introduction
Colorectal cancer (CRC) is the second most frequently diagnosed cancer worldwide1 and represents a major challenge for public health. In recent years, significant progress has been made in both CRC therapy and screening programs. However, important gaps remain in our understanding of tumorigenesis and the molecular characteristics of this disease. Several molecular classification models have been proposed, such as CRIS2 (Colon Cancer Intrinsic Subtype), but the most widely adopted in the literature is the Consensus Molecular Subtypes (CMS) model.
The CMS classification was introduced in 20153 with the aim of improving the biological understanding of CRC and guiding clinical treatment decisions. This model identifies four distinct molecular subtypes:
-
CMS1 (“immune subtype”): characterized by high immune infiltration,
-
CMS2 (“canonical subtype”): marked by alterations in key CRC pathways such as WNT and MYC,
-
CMS3 (“metabolic subtype”): defined by widespread dysregulation of metabolic pathways,
-
CMS4 (“mesenchymal subtype”): associated with stromal infiltration and the poorest overall survival.
In 2022, the CMS model was further refined with the introduction of the integrative CMS (iCMS) classification4. This model highlighted that each CMS subtype can be further stratified into two groups, iCMS2 and iCMS3, providing an even deeper biological characterization of CRC tumors.
This work was designed to enhance the understanding of the tumor microenvironment (TME) that characterizes each CMS subtype. To achieve this, we not only characterized the TME composition of each CMS, but also investigated whether RNA modifications could drive the enrichment or depletion of specific cellular populations.
RNA modifications have emerged as important regulators of tumorigenesis, influencing protein structure, stability, localization, and RNA interactions with other molecules5. Several types of RNA modifications are now well characterized, including N6-methyladenosine (m6A), N1-methyladenosine (m1A), 5-methylcytosine (m5C), N7-methylguanosine (m7G), pseudouridine (Ψ), adenosine-to-inosine (A-to-I) editing, and APOBEC-mediated cytidine-to-uridine (C-to-U) editing.
In this study, we focused specifically on A-to-I RNA editing6, a post-transcriptional modification catalyzed by the ADAR (adenosine deaminase acting on RNA) family of enzymes. These proteins typically contain two or three RNA-binding domains at the N-terminus and a deaminase domain at the C-terminus. In mammals, three ADAR proteins have been identified. ADAR1 is expressed in two isoforms: ADARp110, a 110-kDa protein that is constitutively expressed and localized in the nucleus, and ADARp150, a 150-kDa isoform induced by interferon that can be found both in the nucleus and cytoplasm. ADAR2, also known as ADARB1, is predominantly expressed in the central nervous system7, whereas ADAR3, or ADARB2, lacks catalytic activity, and is thought to act as a competitive inhibitor of the other ADAR enzymes8.
For a long time, the study of A-to-I editing was mainly restricted to brain cancers. However, in recent years, the development of new computational tools, such as REDItool and JACUSA, has greatly simplified the identification of editing events in RNA, allowing researchers to expand the investigation to other tumor types. Several studies have now reported genes affected by RNA editing in different cancers9. Among these, AZIN1 is one of the most extensively studied. Editing of AZIN1 has been associated with various malignant tumors, including hepatocellular carcinoma (HCC), esophageal squamous cell carcinoma (ESCC), and CRC. The editing event leads to a serine-to-glycine substitution at position 367 (Ser367Gly) of the AZIN1 protein, enhancing its affinity for key regulators such as ornithine decarboxylase and cyclin D1, and protecting them from proteasomal degradation. This mechanism promotes the maintenance of cancer cell stemness and tumor progression10.
Despite these findings, to the best of our knowledge, no studies have specifically investigated the relationship between RNA editing and CMS profiles in CRC. Most of the available literature has focused on other types of RNA modifications11. A better understanding of RNA editing in the context of CMS classification could not only help identify new prognostic biomarkers but also provide insight into molecular features such as the enrichment of alternative splicing isoforms observed in subtypes like CMS212. It is now well recognized that RNA editing can influence splicing, further highlighting its potential relevance in shaping CMS-specific tumor biology.
Materials and methods
100 colon cancer patients were retrospectively enrolled at the IRCCS Istituto Tumori “Giovanni Paolo II”. In detail, Formalin-Fixed Paraffine-Embedded (FFPE) samples were collected with the support of Pathology Department performing quality check and selection of the most representative block for each case.
The Institutional Ethics Committee “Gabriella Serio” of the IRCCS Istituto Tumori “Giovanni Paolo II” approved the study. Patients signed an informed consent. Given the retrospective nature of the study, a Data Protection Impact Assessment document was redacted by the Institutional Data Protection Officer in compliance with Article 36 of the EU General Data Protection Regulation (Regulation (EU) 2016/679). The document was evaluated and approved by the Institutional Ethics Committee “Gabriella Serio” (Prot n. 780/CE). The study is compliant with the Declaration of Helsinki.
DNA and RNA extraction
Nucleic acid extraction was performed using the MagMAX™ FFPE DNA/RNA Ultra Kit (ThermoFisher Scientific) on the KingFisher™ Duo Prime System (ThermoFisher Scientific) following the manufacturer’s instructions. Briefly, 6 μm-thick tissue sections were deparaffinized and subjected to proteinase K digestion at 56 °C overnight to ensure effective nucleic acid release. Nucleic acids were captured using paramagnetic beads and washed with buffers provided in the kit to remove contaminants. The extracted DNA and RNA were eluted in 50 µL of the elution buffer provided and quantified using Qubit™ Fulorometric Quantification (ThermoFisher Scientific) to assess concentration. Extracted nucleic acids were stored at −20 °C and − 80 °C, respectively, until further processing.
Bulk RNAseq
RNA-seq sequencing experiments were performed by Genomix4life S.R.L. (Baronissi, Salerno, Italy). RNA concentration in each sample was assayed with a ND-1000 spectrophotometer (NanoDrop) and its quality assessed with the TapeStation 4200 (Agilent Technologies). Indexed libraries were prepared from 800 ng/ea purified RNA with TruSeq Stranded Total RNA Library Prep Gold (Illumina) according to the manufacturer’s instructions. Libraries were quantified using the TapeStation 4200 (Agilent Technologies) and Qubit fluorometer (Invitrogen Co.), then pooled such that each index-tagged sample was present in equimolar amounts, with final concentration of the pooled samples of 1.5 nM. The pooled samples were subject to cluster generation and sequencing using an Illumina NovaSeq 6000 System (Illumina) in a 2 × 100 paired-end format. The raw sequence files generated (.fastq files) underwent quality control analysis using FastQC.
Once the quality of all samples was confirmed to be adequate, the reference genome (hg38, downloaded from UCSC) was indexed using STAR13 (v. 2.7.11b), and alignment was performed. Following alignment, BAM files were quantified using RSEM14, generating two types of quantification for each sample: one at the gene level and one at the isoform level.
CMS and iCMS classifications
For the classification of samples into CMS and iCMS profiles, the CMScaller (v. 0.99.2) package was used. This package requires input data normalized in TPM. For CMS classification, the template provided by the package was used. For iCMS classification, the adjusted function was applied, followed by the ntp function. For ntp, it is necessary to provide a template; in this case, the template downloaded from the supplementary material of Joanito et al.4.
Bioinformatics analysis
DGE and DTU analysis
Gene- and isoform-level count matrices were imported into RStudio (v. 4.3.2) using the tximport (v. 3.21) package. For the gene-level matrix, the parameters txIn and txOut were set to “FALSE”, whereas for the isoform-level matrix, both parameters were set to “TRUE”. Data normalization was performed using DESeq215 (release 3.21). The variance-stabilized transformation (VST) results were used to perform ANOVA and Tukey’s post hoc test. P-values were adjusted for multiple testing using Benjamini-Hochberg (BH) correction. Statistical analyses were carried out using the rstatix package (v0.7.2). All figures were generated in R using the ggpubr and ggplot2 packages (v.s 0.6.0 and 3.5.2 respectively).
Cell type Deconvolution and digital expression assessment
In this project, three different deconvolution methods were applied: CIBERSORTx16, MuSiC17 (v. 0.3) and BayesPrism18 (v. 2.2).
Pre-processing single cell data
To estimate cell-type proportions from bulk RNA-seq data, a signature matrix was first constructed using single-cell RNA-seq data from an independent cohort of nine stage II colon cancer tissue samples. The Cell Ranger output was imported into RStudio and converted into a Seurat object.
In detail, Single-cell RNA-seq count matrices were imported into Seurat v5 using the CreateSeuratObject function (min.cells = 5, min.features = 200). For each sample, the percentage of mitochondrial transcripts was computed with PercentageFeatureSet, and quality-control filtering was applied to retain cells with more than 100 and fewer than 7,000 detected genes and with less than 15% mitochondrial RNA content. After filtering, the objects were normalized independently with Seurat’s log-normalization method (NormalizeData). To mitigate batch effects across the nine samples and harmonize the expression profiles, we applied Seurat’s anchor-based integration workflow. Integration anchors were identified using FindIntegrationAnchors (dims = 1:30, k.filter = 200), and the datasets were combined into a single integrated object using IntegrateData with the same dimensional parameters. This integrated Seurat object was used for all downstream analyses.
After normalization, clustering was performed using the FindClusters() function with a resolution of 0.6, identifying 11 distinct clusters, theses clusters were manually annotated using the FindAllMarkers() function. UMAP and dot plot of markers genes are displayed, respectively in Supplementary Fig. 1 A and Supplementary Fig. 1B.
Signature matrix constructions
For CIBERSORTx, the signature matrix was built by selecting samples with the best cell distribution (samples: S01, S02, and S05). CPM normalization was applied, and the matrix was exported with 20 replicates per cell type. For the bulk RNA-seq data, TPM normalization was applied. Deconvolution was performed using Docker Desktop (v. 4.38), running the CellFraction function with default parameters and enabling batch correction in S-mode.
For MuSiC, the MuSiC R package was used. The main difference in this method was the data format: both the single-cell and bulk RNA-seq data had to be provided as raw counts.
BayesPrism was run with default parameters, using the same data normalization as for MuSiC (raw counts).
Cell-fraction Estimation
To ensure comparability across methods, data were scaled between 0 and 1. Subsequently, pairwise contrasts were built for each CMS subtype against all other profiles. Statistical testing was performed using the Wilcoxon test, and only results that were consistent across all three methods—both in terms of statistical significance (< 0.05 p-value) and direction of enrichment or depletion— were retained.
Gene expression analysis in deconvolved cell populations
Gene expression in the different deconvolved cell populations was assessed. This type of analysis is available only with CIBERSORTx, where GroupMode was run using the signature matrix and bulk data for each CMS profile. GroupMode was executed twice: once using only the samples corresponding to each CMS subtype and once using the samples from all other CMS profiles. Differential gene expression (DGE) analysis was then performed using scripts presented in this review19.
In the case of BayesPrism, the output matrix “Z” (containing estimated expression values per cell type) was used. Following the official GitHub recommendations, normalization was performed using DESeq2, and pairwise contrasts were built for all CMS subtypes.
Editing detection site
To detect RNA editing sites in bulk samples, REDItools20,21 (v. 1.3) was used with the REDItoolDnaRNA.py script. The first step involved indexing the BAM files and the reference genome. Subsequently, the REDItoolRNARNA.py function was applied without DNA sequencing data. This process generated output tables containing information about editing sites in the samples. The output tables were filtered using the dbSNP file and annotated using the RepeatMasker track (downloaded from UCSC), which was used to categorize the editing site into “Repeated” elements (Rep site) and “Unique” elements (NoRep site).
Editing statistical analyses
Editing Statistical analyses were performed to assess global editing differences among CMS profiles. Initially, the GetOverall.py script from REDItools was applied to compute global editing rates per sample. Subsequently, the GetREI.py script was run separately on Rep and NoRep files to quantify editing events within repetitive and unique elements, respectively. To specifically investigate editing within Alu elements, the RNAEditingIndexer tool22 was used. A custom RepeatMasker annotation containing only Alu elements (in BED format) was generated and applied to all BAM files.
Differential gene editing site analysis
To identify genes with editing sites differentially edited across CMS profiles, Rep and NoRep files were imported and merged by sample in RStudio. The resulting data frame was filtered to retain only sites with: a coverage > 10 reads, and an editing frequency > 0.1.
Kruskal-Wallis tests were performed at each editing site, followed by Dunn’s post hoc tests.
Editing sites with an adjusted p-value (p-adj < 0.01), corrected using the Benjamini-Hochberg (BH) method, were considered statistically significant.
CeRNA network construction for CMS subtypes
To investigate editing sites located within repetitive elements, we built a competing endogenous RNA (ceRNA) network specific to each CMS subtype. In this model, long non-coding RNAs (lncRNAs) act as molecular sponges for miRNAs, preventing them from binding and repressing their mRNA targets, thus allowing mRNA expression.
First, a correlation matrix was generated between mRNA and lncRNA expression using the variance-stabilized (VST) gene-level matrix. Correlations were filtered by p-value (< 0.05) and r > |0.7|.
The interaction file was constructed using two databases containing experimentally validated interactions. From lncRNA-miRNA interactions lncPedia23 only interactions with experimental validation and showing a negative regulatory effect were retained.
For miRNA–mRNA interactions, data were retrieved from TarBase24; this dataset was filtered to include only direct interactions, regardless of whether the effect was positive or negative.
Next, Gene Set Enrichment Analysis (GSEA) was performed separately for each CMS subtype using gene sets from the H1 and C2 collections (GMT format). Pathways were considered statistically significant if they exhibited a nominal p-value < 0.05 and a q-value < 0.25. Genes enriched for each CMS subtype were then used to filter the preliminary ceRNA networks, generating CMS-specific networks.
Subsequently, significant Rep-editing sites were analyzed. Only genome positions falling within annotated lncRNAs (according to a GTF file v.40) were retained. These lncRNAs were used to further filter the CMS-specific ceRNA networks, producing the final networks for each CMS subtype.
Visualization and exploration of the network
Finally, the networks were analyzed using Cytoscape25 (v. 3.10.3). The MCODE (v. 2.0.0) plugin was used to identify densely connected modules, and CytoHubba (v. 0.1) was used to rank interactions, selecting the top five hubs for each network. Functional enrichment of hub genes was performed using WebGestalt26.
Survival analysis
Univariate and multivariate Cox proportional hazards regression analyses, as well as Kaplan–Meier survival analyses, were performed using the “survival” R package (v. 3.7).
Clinicopathological variables were selected based on univariate Cox models with p-value filtering (p < 0.05), while numerical variables were selected using various feature selection strategies, including p-value filtering, correlation-based filtering, forward selection, and LASSO regression, implemented via the “glmnet” R package (v. 4.1).
The performance of each multivariate model was evaluated using 5-fold cross-validation with the “survcomp” R package (v. 1.54.0), employing the concordance index (C-index) as the evaluation metric. Among the three approaches tested, the forward selection model achieved the highest C-index, indicating the best predictive performance.
This best-performing model was then selected based on the evaluation criteria. A risk score was subsequently calculated for each sample using the formula derived from the final multivariate Cox model.
Optimal cutpoints for stratifying the risk score were identified using the “surv_cutpoint” function from the “survminer” R package.
Results
ADAR family gene expression across CMS
The clinical characteristics of the patient cohort included in the study are summarized in Table 1. The median age was 74 years (Q1: 65, Q3: 79), with a balanced distribution between males and females. Most patients were diagnosed with stage III or IV colon cancer.
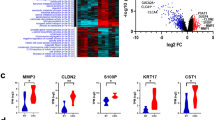
Violin plot representing the expression levels of editing-related genes across CMS subtypes. ANOVA followed by Tukey’s post-hoc test was performed for pairwise comparisons (ns = not significant, p ≤ 0.05 (*), p ≤ 0.01 (**), p ≤ 0.001 (***), p ≤ 0.0001 (****). (A) ADARB1 expression across CMS subtypes(p < 0.0001). (B) ADARB2 expression across CMS subtypes (p = 0.057). (C) ADAR expression across CMS subtypes (p = 0.063).
The expression of enzymes belonging to the ADAR family across CMS subtypes was explored. ADARB1 emerged as the only ADAR gene showing significant expression differences among CMS subtypes (Fig. 1A). ADARB2 showed a statistical trend with higher expression in CMS2 then in CMS4, whilst no statistical significant difference was observed for ADAR (Fig. 1B-C).
As illustrated in Fig. 1A, CMS4 exhibited the highest expression levels of ADARB1, followed by CMS1. In contrast, CMS2 and CMS3 showed the lowest levels of expression. These findings suggest that the elevated expression of ADARB1 in CMS1 and CMS4 may be associated with the TME characteristics specific to the features of these subtypes—namely, immune enrichment in CMS1 and stromal enrichment in CMS4.
Mapping Cell-Type composition in colon cancer CMS landscapes
To investigate this possibility, bulk RNA-seq data were deconvoluted using a single-cell RNA-seq dataset derived from colon cancer patients to estimate the cellular composition of the TME and identifying potential associations between ADAR family gene expression and specific immune or stromal cell populations. Three deconvolution algorithms were employed: CIBERSORTx, MuSiC, and BayesPrism. Results from each method are presented in Supplementary Fig. 2–13.
Cell fraction estimates from each method were used to identify differentially enriched cell populations across CMS subtypes. Subsequently, immune and stromal populations showing significant associations with the expression of ADAR genes were examined.
Statistical comparison of cell type proportions in CMS1 versus others; Cell populations consistently identified across all three methods (BayesPrism, CIBERSORTx, and MuSiC) as significantly enriched or depleted in CMS1 compared to other CMS subtypes.
In CMS1 (Fig. 2), an enrichment of monocytes was observed; this population also showed elevated expression of ADARB1 and ADAR (Supplementary Fig. 14).
A-B Statistical comparison of cell type proportions in CMS2 versus others;. Cell populations consistently identified across all three methods (BayesPrism, CIBERSORTx, and MuSiC) as significantly enriched or depleted in CMS2 compared to other CMS subtypes.
In CMS2, epithelial cells were enriched, while fibroblast were depleted (Fig. 3A-B). Epithelial cells in this subtype showed increased expression of ADARB2, which was also observed in the remaining fibroblast population (Supplementary Fig. 15).
A-B. Statistical comparison of cell type proportions in CMS3 versus others; Cell populations consistently identified across all three methods (BayesPrism, CIBERSORTx, and MuSiC) as significantly enriched or depleted in CMS3 compared to other CMS subtypes.
In CMS3, an enrichment of CALB1 + epithelial cells and a reduction in smooth muscle cells were detected (Fig. 4A-B). CALB1 + epithelial cells exhibited reduced ADARB2 expression, whereas smooth muscle cells displayed elevated ADARB1 levels (Supplementary Fig. 16).
A-B-C-D.Statistical comparison of cell type proportions in CMS4 versus others; Cell populations consistently identified across all three methods (BayesPrism, CIBERSORTx, and MuSiC) as significantly enriched or depleted in CMS4 compared to other CMS subtypes.
CMS4 showed enrichment in endothelial cells, fibroblasts, mast cells, and macrophages (Fig. 5A-B-C-D). Among these, mast cells and fibroblasts were enriched in ADARB1, while macrophages showed increased expression of ADAR (Supplementary Fig. 17).
Notably, fibroblasts in CMS2 and CMS4 exhibited contrasting expression patterns: in CMS4 fibroblasts, ADARB1 and ADAR were upregulated (Supplementary Fig. 17), whereas in CMS2 fibroblasts, only ADARB2 expression was increased (Supplementary Fig. 15).
This differential pattern may suggest a role for ADARB1 and ADAR in promoting fibroblast enrichment in CMS4.
RNA editing across CMS
The first step of the analysis involved assessing global RNA editing activity across the samples. No statistically significant differences were observed among the CMS subtypes (Supplementary Fig. 18 A).
Most editing events occur in Alu inverted repeats, while some are found in non-repetitive regions and can lead to amino acid substitutions. To globally quantify editing in colon cancer, we used three metrics: GetREI for unique regions (Supplemetary Fig. 18B), GetNoREI for repetitive elements (Supplementary Fig. 18 C) and the RNAEditingIndex for Alu elements (Supplementary Fig. 18D-I). None of the analyses showed differences across the CMS subtype.
We then evaluated editing at the gene level (GetREI) and lncRNA level (GetNoREI) to identify edited transcripts. This editing is mainly mediated by ADARB1, which is enriched in CMS4 and CMS1. Gene editing was further explored through differential transcript usage (DTU). Significant recoding events are reported in Table 2, and edited lncRNA in Supplementary Table 1.
Differential Editing Events by CMS.
Among the significant recoding events identified in association with CMS profiles were editing sites in COPA27 and IGFBP728, which have been previously implicated in tumorigenesis. As shown in Supplementary Fig. 19, COPA editing event is enriched in the CMS4 subtype, followed by CMS1, compared to other CMS profiles.
Notably, DTU analysis revealed significant isoform switching in COPA, with enriched isoforms in CMS4.
The editing event on IGFBP7 was found to be significantly enriched in the contrasts CMS4 vs. CMS3 and CMS4 vs. CMS2, as well as in CMS1. The subtype with the lowest enrichment of this editing event was CMS3 (Supplementary Fig. 20 A).
In our analysis, the isoform IGFBP7-202 (Supplementary Fig. 20B) emerged as significantly enriched in CMS4 according to DTU analysis.
CADPS showed increased RNA editing in the CMS1 (Supplementary Fig. 21 A) subtype compared to other profiles. This gene encodes a protein involved in the exocytosis of neurotransmitter-containing vesicles. However, due to the high number of transcript isoforms, that reached statistical significance in DTU analysis (Supplementary Fig. 21B-L), interpreting the functional implications of this editing event remains complex and requires further investigation.
In contrast, other editing sites, such as those in ZNF552, and RALGPS1, have not yet been reported in the context of colorectal or other cancers.
Editing in ZNF552 (chr19:57813360) (Supplementary Fig. 21 A) was significantly different in CMS3 vs. CMS1/CMS4 and CMS2 vs. CMS1 comparisons. The DTU data shows an enrichment of two isoform (Supplementary Fig. 22 C) in the same CMS subtype.
Finally, RALGPS1 displayed two distinct editing events: one (chr9:127183878) significantly enriched in CMS4 vs. CMS2/CMS3 (Supplementary Fig. 23 A), and another (chr9:127183894) enriched in CMS2 vs. CMS4 (Supplementary Fig. 23B). This gene encodes a guanine nucleotide exchange factor (GEF) that activates members of the Ral family.
CeRNA editing network
To investigate the potential regulatory roles of editated lncRNA (Supplementary Table 1) and their contribution to the molecular features of each CMS subtype, we systematically reconstructed a ceRNA network for each group, allowing us to explore how these transcripts might modulate gene expression throught interactions with miRNA and their downstream targets, thereby shedding light on the distictive molecular characteristics of each subtype. In these networks, it was hypothesized that lncRNAs may function as sponges for miRNAs, thereby preventing miRNAs from binding to their mRNA targets and leading to increased expression of those target genes. In Supplementary Fig. 24, significantly enriched gene sets in CMS1 are shown. They were used to filter the overall ceRNA network obtaining CMS1-related subnetwork.
CMS1 ceRNA network: Competing endogenous RNA network specific to the CMS1 subtype. The network highlights the top 10 hub interactions identified using the CytoHubba.
In the CMS1 ceRNA network, only one lncRNA, RC3H1-IT1, was identified (Fig. 6). This lncRNA is hypo-edited in two genomic positions shown in Fig. 6, in comparison CMS3 to CMS1. The corresponding mRNA target is TNFRSF1A which encodes for the tumor necrosis factor receptor, having a well-established role in inflammatory processes. Significantly enriched gene sets and the edited lncRNAs are shown in Supplementary Fig. 24 and Supplementary Table 2, respectively.
CMS2 ceRNA network: Competing endogenous RNA network specific to the CMS2 subtype. The network highlights the top 10 hub interactions identified using the CytoHubba.
The ceRNA network of CMS2 (Fig. 7) was characterized by the involvement of the genes MED14, OGT, and RBM39 (significantly enriched gene sets are displayed in Supplementary Fig. 25).
The network also included the lncRNAs ANKRD10-IT1 and LINC01473. ANKRD10-IT1 was found to be edited at the genomic position chr13:110899024, with editing levels enriched in CMS4 compared to CMS2. In contrast, LINC01473 displayed four distinct editing sites, listed in Supplementary Table 3, all of which were hyper-edited in CMS2 relative to CMS4.
CMS3 ceRNA network:. A. WebGestalt enrichment results (unweighted analysis), B-C. the top 20 hub interactions identified by CytoHubba.
The ceRNA network identified in CMS3 was markedly extensive and was therefore stratified based on functional annotations. Functional enrichment analysis was performed using the WebGestalt tool (Fig. 8A), applied to the mRNA components of the CMS3-specific ceRNA network.
As shown in Fig. 8B–C, pathway enrichment results enabled the manual subdivision of the network into functionally distinct modules. Among the most enriched genes, SPHK2 appeared across multiple pathways, including sphingolipid metabolism and Rho signaling. Based on these results, the network was refined to highlight genes involved in these two pathways (Fig. 8B).
Additional modules were identified based on the enrichment of other genes such as PGAM5 and SCAF1.
-
PGAM5 was included in the network due to its association with mitochondrial-related pathways (Fig. 8C);
-
SCAF1, also known as COX7A2L, was linked to pathways related to oxidative phosphorylation and mitochondrial supercomplex assembly.
The lncRNAs involved in the CMS3-specific ceRNA network are listed in Supplementary Table 4. Interestingly, a large number of these lncRNAs showed enrichment in CMS3 compared to CMS4.
CMS4 ceRNA network: network specific to the CMS4 subtype. (A) WebGestalt enrichment results (unweighted analysis), (B) WebGestalt results (weighted analysis), (C) top 10 hub interactions identified by CytoHubba.
The same analytical approach was applied to the CMS4 ceRNA network. Functional enrichment analysis revealed a significant association with pathways involving fibroblasts and smooth muscle cells, as shown in Fig. 9A-B. Given the fibroblast enrichment identified through deconvolution methods, the associated gene lists were further explored. The genes contributing to these enriched pathways were merged into a unified list and used to manually curate the ceRNA network, resulting in a refined version illustrated in Fig. 9C. The lncRNAs that are involved in this network are listed in Supplementary Table 5.
Survival analysis and prognostic model definition
To investigate the prognostic relevance of ADAR family gene expression and RNA editing events, survival analyses were performed on colon cancer patient cohorts. Overall survival (OS) was estimated using three Cox proportional hazards models (Univariant Model Cox-ph is represented in Supplementary Table 6), each incorporating a different feature selection strategy: correlation-based selection (Supplementary Fig. 26–27), forward selection, and LASSO regression (Supplementary Fig. 28–29). Among the three approaches, the forward selection model demonstrated the highest C-index, indicating superior predictive performance (Fig. 10A).
(A) Overall Survival analysis, presenting the C-index evaluated using 5-fold cross-validation for three different feature selection methods: Correlation, Forward Selection, and Lasso Selection. (B) multivariate Cox proportional hazards model based on features selected by Forward Selection. (C) Kaplan–Meier survival curves stratified by risk score derived from the multivariate Cox model.
Notably, this model identified (Fig. 10B) three variables as significant independent predictors of OS: ADARB1 expression, and RNA editing events in the lncRNAs ST7-AS2 and NOP14-AS1. ADARB1 showed a hazard ratio (HR) of 4.3352 (p < 0.001), suggesting a central role for ADARB1 in influencing patient survival.
In addition, two RNA editing sites were significantly associated with patient prognosis. The first, located in ST7-AS2 (chr4:117120557), had a p-value of 0.026 and an HR of 6.6990. The second editing site, located in NOP14-AS1 (chr4:2960236), was associated with improved survival (p = 0.036; HR = 0.0069).
Based on the coefficients of the selected multivariate Cox model, a risk score was computed for each patient (Fig. 10C), enabling stratification into high- and low-risk groups. The optimal cut-off value (RiskScore cutpoint = 1.0006) for this dichotomization was derived from the distribution of the risk scores. Kaplan–Meier survival curve showed statistically significant difference in the two groups (p-value ≤ 0.0001). In detail, high risk patients had a median survival of 870 days (95% CI 537–1325), whereas the low-risk group did not reach the median OS within the available follow-up period.
Discussion
The principal aim of this study was to investigate the impact of A-to-I RNA editing across the CMS subtypes in CRC. One of the key findings that motivated the exploration of this RNA modification was the specific enrichment of the ADARB1 gene in CMS4, followed by CMS1. These results suggest that RNA editing might contribute to the enrichment of immune cells in CMS1 and stromal cells in CMS4. Based on this observation, we examined the expression of ADAR enzymes in the TME using RNA-seq-based deconvolution methods.
A particularly promising result was the contrasting fibroblast abundance between CMS4 and CMS2. CMS4 displayed an increased fibroblast population, while CMS2 showed a notable reduction. Interestingly, fibroblasts in CMS4 exhibited elevated ADARB1 expression, whereas CMS2 showed increased levels of ADARB2. Given that ADARB2 is known to act as a competitive endogenous inhibitor at editing sites29, its higher expression in CMS2 may lead to a reduction in editing activity, potentially contributing to the decreased fibroblast levels observed. In their study30, the authors also highlighted that cancer cells can promote the expression of ADAR1 in fibroblasts, and that edited AZIN1 plays a role in enhancing invasion and migration. These findings together suggest a possible link between RNA editing dynamics, particularly the ADAR-mediated editing of key transcripts, and fibroblast behavior across CMS subtypes.
We further analyzed gene-level editing events using REDItools on bulk RNA-seq data. This analysis identified several genes with subtype-specific editing patterns. Some of these editing sites, such as those in COPA, IGFBP7, and CADPS, are already well-documented in cancer biology. In particular COPA encodes a protein involved in retrograde transport between the trans-Golgi network and the endoplasmic reticulum. The editing event occurs at chr1:160332454, resulting in an amino acid substitution at position 164, from isoleucine (I) to valine (V) (Ile164Val). This functional recoding event has been previously described in the literature27; in particular, the edited COPAI164V isoform has been reported to promote endoplasmic reticulum stress, thereby facilitating metastatic progression in CRC through upregulation of MALAT1, MET, and ZEB1. IGFBP7 encodes an insulin-like growth factor-binding protein with lower affinity for IGFs compared to other members of the same family. It is widely recognized as a tumor suppressor in several cancer types28. The identified editing site at chr4:57110068 results in an amino acid substitution from lysine (K) to glycine (G) at position 95 (Lys95Gly). This recoding event has been previously reported28, and some studies suggested that it may alter the protein’s affinity for IGF1 and IGF2, potentially increasing its binding capacity compared to the unedited form. While the editing site on CADPS wasn’t related with tumorigenesis but in this work the authors showed the effect of editing of CADPS in synaptic transmission31.
Additionally, novel editing events were found in genes not previously associated with editing, such as ZNF552 and RALGPS1. ZNF552 encodes a zinc finger protein involved in transcriptional regulation by RNA polymerase. Although the current data do not allow for detailed functional analysis, zinc finger proteins have been broadly implicated in cancer-related processes. For example, ZNF217, another member of this family, is known to promote metastasis and cell proliferation when overexpressed32. Other processes, such as inflammation, are also influenced by dysregulated zinc finger proteins33. These observations suggested that RNA editing in ZNF552 could potentially affect similar oncogenic pathways. RALGPS shows two different editing (chr9:127183878) and (chr9:127183894). This gene encodes a guanine nucleotide exchange factor (GEF) that activates members of the Ral family and it is involved cytoskeletal dynamics and vesicle trafficking.
The analysis of differential transcript usage (DTU) for the edited gene did not allow us to clearly establish a relationship between the editing site and its potential role in promoting alternative splicing. Investigating such associations would require different types of data, as bulk RNA-seq alone is insufficient to resolve these events. Moreover, the large number of isoforms further complicated data interpretation.
Only IGFBP7 showed a clear result, with a single isoform, IGFBP7-202, significantly enriched in a specific CMS subtype. Notably, this enrichment matched the same subtype in which the editing site was enriched—CMS4. Interestingly, these editing events were enriched in CMS4 and CMS1, the same subtypes in which we also observed ADARB1 upregulation.
To better understand the complex regulatory landscape of each CMS subtype, we also constructed ceRNA networks. These networks revealed subtype-specific interactions.
CMS1 ceRNA network shows the presence of the lncRNA RC3H1-IT1 and the mRNA TNFRSF1A. A previous study34 employing a similar ceRNA network approach in the context of rheumatoid arthritis, a disease typically characterized by autoimmune etiology, also identified RC3H1-IT1 as a central node. Based on this evidence, the ceRNA interaction involving RC3H1-IT1 and TNF can be interpreted as strongly associated with the CMS1 subtype, which is known to be enriched in immune-related signatures.
CMS2 network includes: MED14, OGT and RBM39 mRNAs. RBM39 is a well-characterized splicing regulator. In a previous study35, the axis composed of MORC2–RBM39–CDK5RAP2 was shown to modulate alternative splicing of CDK5RAP2, promoting the CDK5RAP2-S isoform, which is implicated in epithelial-to-mesenchymal transition (EMT) and metastasis in in vitro colon cancer models. MED14 is a component of the Mediator complex, originally identified as a co-regulator of RNA polymerase II36. This multiprotein complex functions as a molecular bridge between transcription factors and gene promoters, playing a central role in transcriptional regulation. OGT encodes O-linked N-acetylglucosamine transferase, an enzyme responsible for the O-GlcNAcylation of serine and threonine residues on target proteins. Elevated OGT expression has been reported as a negative prognostic marker in CRC.
The network also included the lncRNAs ANKRD10-IT1 and LINC01473. ANKRD10-IT1 was found to be edited at the genomic position chr13:110899024, with editing levels enriched in CMS4 compared to CMS2. In contrast, LINC01473 displayed four distinct editing sites, all of which were hyper-edited in CMS2 relative to CMS4.
While no studies to date have linked LINC01473 to colorectal or other cancers, ANKRD10-IT1 has been previously implicated in CRC. In one study37, a non-parametric classification of CRC revealed three novel subtypes, one of which—subtype 3—was strongly associated with m6A RNA modification. Among the 21 lncRNAs involved in this subtype, four were linked to poor prognosis, including ANKRD10-IT1. Additionally, gene set enrichment analysis of this subtype showed a significant enrichment in spliceosome-related pathways.
This evidence supports a hypothesis that the CMS2-associated ceRNA network may be functionally related to RNA splicing and epitranscriptomic modifications. Supporting this, a proteomic-based study of CMS classification further divided CMS2 into two proteomic subgroups (HCL1 and HCL2)12. HCL2 was characterized by splicing regulation and enhanced transcriptional activity, reinforcing the potential relevance of this ceRNA network in defining CMS2 subtype biology.
The CMS3 ceRNA network was divided into two subclusters, one of which showed enrichment of SPHK2, a key gene in sphingolipid metabolism. SPHK2 is known to be upregulated in CC, and its silencing via siRNA reduces cell proliferation and migration in vitro. Treatment with FTY72038, a sphingolipid signaling modulator, has also been shown to inhibit RhoA activity and downregulate metastasis-related proteins. Based on these findings and the enrichment of Rho pathway components in the WebGestalt analysis, the ceRNA network was manually refined to highlight genes involved in sphingolipid metabolism and Rho signaling.
Two other genes of interest in this subcluster were PGAM5 and SCAF1. PGAM5 plays a key role in mitochondrial homeostasis. A previous study39 has shown that mitochondrial ROS can trigger its cleavage, leading to YAP activation, a known tumorigenic factor. SCAF1 (also known as COX7A2L) is involved in assembling mitochondrial respiratory supercomplexes, particularly by stabilizing the interaction between Complex III (CIII₂) and Complex IV (CIV). This improves electron transport efficiency and reduces ROS production, supporting effective oxidative phosphorylation. SCAF1 enables formation of the CS-respirasome (CI + CIII₂ + 2×CIV), which provides a kinetic advantage under low oxygen conditions40,41.
Altered expression of PGAM5 and SCAF1, both linked to mitochondrial function and homeostasis, may underlie the significant metabolic reprogramming observed in the CMS3 subtype. Supporting this hypothesis is the simultaneous involvement of sphingolipid metabolism pathways within the same network3,12.
At least the CMS4 network shows a marker genes of Fibroblast population and Myofibroblast in WebGestalt results; moreover a recent study proposed a novel annotation of fibroblast subtypes in CRC. In this work42, a specific population defined as RGS5⁺ACTA2^hi fibroblasts was identified, which is characterized by the expression of MYH11 and MYL9—genes that were also present in the CMS4 ceRNA network. These findings support the hypothesis that the CMS4-associated network may play a role in the expansion and activity of fibroblast populations, a feature consistently observed in CMS4, both in the literature and in the cellular composition inferred by deconvolution analysis in the present study.
We also evaluated the prognostic impact of RNA editing and ADAR expression through a multivariate Cox proportional hazards model. Three independent prognostic variables were identified: the expression of ADARB1, and two editing sites—NOP14-AS1 (chr4:2960236), which was associated with improved survival (p = 0.036; HR = 0.0069) and has previously been linked to increased sensitivity to 5-FU. This effect was demonstrated in an in vitro study, that showed that this lncRNA interacts with miR-30a-5p and regulates glucose metabolism. It is plausible that editing at this site disrupts the miRNA interaction, potentially reducing chemoresistance and enhancing treatment response43. Moreover ST7-AS2 (chr4:117120557) was not already described in the tumorigenesis. Such a prognostic model, further validated, could be useful in the clinical practice to better stratify patients.
Conclusion
In conclusion, this study not only dissects the distribution and role of ADAR-mediated RNA editing across CMS subtypes but also highlights its multifaceted biological significance—from its role in chemoresistance (e.g., NOP14-AS1), to its contribution to the regulation of the tumor microenvironment and subtype-specific transcriptional programs. Our findings suggest that RNA editing, particularly, represents a promising layer of post-transcriptional regulation in CRC.
Data availability
Bulk RNAseq raw data were deposited in SRA repository under the accession number PRJNA1304982 (https:/www.ncbi.nlm.nih.gov/bioproject/PRJNA1304982). Single cell RNAseq data were deposited in SRA repository under the accession number (https://www.ncbi.nlm.nih.gov/bioproject/PRJNA1369595).
References
Morgan, E. et al. Global burden of colorectal cancer in 2020 and 2040: incidence and mortality estimates from GLOBOCAN. Gut 72 (2), 338–344 (2023).
Isella, C. et al. Selective analysis of cancer-cell intrinsic transcriptional traits defines novel clinically relevant subtypes of colorectal cancer. Nat. Commun. 8, Article number: 15107 (2017)
Guinney, J. et al. The consensus molecular subtypes of colorectal cancer. Nat. Med. 21 (11), 1350–1356 (2015).
Joanito, I. et al. Single-cell and bulk transcriptome sequencing identifies two epithelial tumor cell States and refines the consensus molecular classification of colorectal cancer. Nat. Genet. 54 (7), 963–975 (2022).
Barbieri, I. & Kouzarides, T. Role of RNA modifications in cancer. Nat. Reviews Cancer Nat. Res. 20, 303–322 (2020).
Wang, H. et al. RNA Editing in Cancer: from Evaluating the Editing Level To Exploring the Editing Effects 10 (Frontiers Media S.A., 2021). Frontiers in Oncology.
Savva, Y. A., Rieder, L. E. & Reenan, R. A. The ADAR protein family. Genome biology. 13, 252. (2012).
Chen, C. X. et al. A third member of the RNA-specific adenosine deaminase gene family, ADAR3, contains both single-and double-stranded RNA binding domains. (2000).
Zhang, Y. et al. Advances in A-to-I RNA Editing in Cancer 23 (BioMed Central Ltd, 2024).
Shigeyasu, K. et al. AZIN1 RNA editing confers cancer stemness and enhances oncogenic potential in colorectal cancer. JCI Insight. 2018 Jun 21;3(12):e99976
Chen, H. et al. Cross-talk of four types of RNA modification writers defines tumor microenvironment and Pharmacogenomic landscape in colorectal cancer. Mol Cancer. 2021 Feb 8;20(1):29. doi: 10.1186/s12943-021-01322-w
Feliu, J. et al. Functional proteomics of colon cancer consensus molecular subtypes. Br. J. Cancer. 130 (10), 1670–1678 (2024).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29 (1), 15–21 (2013).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics. 2011 Aug 4;12:323. doi: 10.1186/1471-2105-12-323
Love, M. I., Huber, W. & Anders, S. Moderated Estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15(12):550. doi: 10.1186/s13059-014-0550-8.
Newman, A. M. et al. Determining cell type abundance and expression from bulk tissues with digital cytometry. Nat. Biotechnol. 37 (7), 773–782 (2019).
Wang, X., Park, J., Susztak, K., Zhang, N. R. & Li, M. Bulk tissue cell type Deconvolution with multi-subject single-cell expression reference. Nat Commun. 2019 Jan 22;10(1):380. doi: 10.1038/s41467-018-08023-x
Chu, T., Wang, Z., Pe’er, D. & Danko, C. G. Cell type and gene expression Deconvolution with bayesprism enables bayesian integrative analysis across bulk and single-cell RNA sequencing in oncology. Nat. Cancer. 3 (4), 505–517 (2022).
Steen, C. B., Liu, C. L., Alizadeh, A. A. & Newman, A. M. Profiling cell type abundance and expression in bulk tissues with CIBERSORTx. Methods Mol Biol. 2020;2117:135-157.
Picardi, E., Pesole, G. & REDItools High-throughput RNA editing detection made easy. Bioinformatics 29 (14), 1813–1814 (2013).
Lo Giudice, C., Tangaro, M. A., Pesole, G. & Picardi, E. Investigating RNA editing in deep transcriptome datasets with reditools and rediportal. Nat. Protoc. 15 (3), 1098–1131 (2020).
Roth, S. H., Levanon, E. Y. & Eisenberg, E. Genome-wide quantification of ADAR adenosine-to-inosine RNA editing activity. Nat. Methods. 16 (11), 1131–1138 (2019).
Volders, P. J. et al. Lncipedia 5: towards a reference set of human long non-coding Rnas. Nucleic Acids Res. 47 (D1), D135–D139 (2019).
Dong, Y. et al. Autophagy-related lncRNAs and Exosomal lncRNAs in Colorectal Cancer: Focusing on lncRNA-targeted Strategies Vol. 24 (BioMed Central Ltd, 2024). Cancer Cell International.
Otasek, D., Morris, J. H., Bouças, J., Pico, A. R. & Demchak, B. Cytoscape automation: empowering workflow-based network analysis. Genome Biol. 2019 Sep 2;20(1):185
Elizarraras, J. M. et al. WebGestalt 2024: faster gene set analysis and new support for metabolomics and multi-omics. Nucleic Acids Res. 52 (W1), W415–W421 (2024).
Wang, S. et al. qi, COPA A-to-I RNA editing hijacks endoplasmic reticulum stress to promote metastasis in colorectal cancer. Cancer Lett. ;553:215995. (2023).
Godfried Sie, C., Hesler, S., Maas, S. & Kuchka, M. IGFBP7’s susceptibility to proteolysis is altered by A-to-I RNA editing of its transcript. FEBS Lett. 586 (16), 2313–2317 (2012).
Raghava Kurup, R. et al. RNA binding by ADAR3 inhibits adenosine-to-inosine editing and promotes expression of immune response protein MAVS. J Biol Chem. 2022 Sep;298(9):102267.
Takeda, S. et al. Activation of AZIN1 RNA editing is a novel mechanism that promotes invasive potential of cancer-associated fibroblasts in colorectal cancer. Cancer Lett. 444, 127–135 (2019).
Shumate, K. M., Tas, S. T., Kavalali, E. T. & Emeson, R. B. RNA editing-mediated regulation of calcium-dependent activator protein for secretion (CAPS1) localization and its impact on synaptic transmission. J. Neurochem. 158 (2), 182–196 (2021).
Zhang, Z. C., Zheng, L. Q., Pan, L. J., Guo, J. X. & Yang, G. S. ZNF217 is overexpressed and enhances cell migration and invasion in colorectal carcinoma. Asian Pac. J. Cancer Prev. 16 (6), 2459–2463 (2015).
Liu, S., Sima, X., Liu, X. & Chen, H. Zinc Finger Proteins: Functions and Mechanisms in Colon Cancer 14 (Cancers. MDPI, 2022).
Huang, Z. & Kuang, N. Construction of a CeRNA network related to rheumatoid arthritis. Genes (Basel). 2022 Apr 6;13(4):647.
He, Y. et al. MORC2 regulates RBM39-mediated CDK5RAP2 alternative splicing to promote EMT and metastasis in colon cancer. Cell Death Dis. 2024 Jul 24;15(7):530.
Jaeger, M. G. et al. Selective mediator dependence of cell-type-specifying transcription. Nat. Genet. 52 (7), 719–727 (2020).
Tan, X. et al. Integrative analysis reveals potentially functional N6-Methylandenosine-Related long noncoding RNAs in colon adenocarcinoma. Front Genet. 2021 Sep 17;12:739344
Jafari, N. et al. De Novo fatty acid synthesis-driven sphingolipid metabolism promotes metastatic potential of colorectal cancer. Mol. Cancer Res. 17 (1), 140–152 (2019).
Wang, S. et al. ROS-induced cytosolic release of mitochondrial PGAM5 promotes colorectal cancer progression by interacting with MST3. Nat Commun. 2025 Feb 6;16(1):1406
García-Poyatos, C. et al. Scaf1 promotes respiratory supercomplexes and metabolic efficiency in zebrafish. EMBO Rep. 2020 Jul 3;21(7):e50287.
Benegiamo, G. et al. COX7A2L genetic variants determine cardiorespiratory fitness in mice and human. Nat. Metab. 4 (10), 1336–1351 (2022).
Li, S. et al. An integrated map of fibroblastic populations in human colon mucosa and cancer tissues. Commun Biol. 2022 Dec 3;5(1):1326.
Lu, Y. N. Blocking LncRNA NOP14-AS1 overcomes 5-Fu resistance of colon cancer cells by modulating miR-30a-5p-LDHA-glucose metabolism pathway. Discov Oncol. 2025 Apr 3;16(1):458.
Acknowledgements
All the patients.
Funding
“Tecnopolo per la Medicina di Precisione”, CUP: B84I18000540002.
SingleCell RNAseq dataset: Ricerca Corrente 2025-Italian Ministry of Health.
Author information
Authors and Affiliations
Contributions
DM, SDS: conducted data interpretation and data analysis. DM, DT: contributed to the writing of the manuscript. EM, FAZ, GC: contributed to the organization and processing of histology data. FB, SD, TG: contributed to the collection of clinical data. DT, RP, AL, RL: contributed to the generation of all the molecular data. EP, ST, SDS conceived the study and supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
The Institutional Ethics Committee “Gabriella Serio” of the IRCCS Istituto Tumori “Giovanni Paolo II” approved the study Given the retrospective nature of the study, a Data Protection Impact Assessment document was redacted by the institutional Data Protection Officer in compliance with Article 36 of the EU General Data Protection Regulation (Regulation (EU) 2016/679). The document was evaluated and approved by the Institutional Ethics Committee “Gabriella Serio” (Prot n. 780/CE). The study is compliant with the Declaration of Helsinki.
The authors affiliated to the IRCCS Istituto Tumori “Giovanni Paolo II”, Bari, are responsible for the views expressed in this article, which do not necessarily represent the Institute.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Monaco, D., Traversa, D., Mattioli, E. et al. Biological and prognostic relevance of A-to-I RNA editing across consensus molecular subtypes of colon cancer. Sci Rep 16, 4018 (2026). https://doi.org/10.1038/s41598-025-34043-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-34043-x