Abstract
Lung adenocarcinoma (LUAD) is the most common subtype of lung cancer worldwide, in which fatty acid metabolism plays a key role in tumor growth, metastasis, and immune evasion. PPIA has been confirmed to be closely related to tumor metabolism and immune regulation, but its specific mechanism of action is still unclear. Multi-omics data from TCGA and GEO, combined with single-cell RNA sequencing (scRNA-seq), were used to characterize metabolic heterogeneity and the immune microenvironment in lung adenocarcinoma (LUAD). Fatty acid metabolism–related genes were identified via WGCNA and univariate Cox regression, and a prognostic risk model was constructed. Virtual knockout, cell function assays, and metabolic profiling were performed to investigate the role of PPIA in metabolic reprogramming and tumor progression. In vitro, PPIA was silenced using siRNA or co-overexpressed with pCMV-C-Myc in A549 and H1975 cells. Functional validation included qRT-PCR and Western blotting, as well as CCK-8, colony formation, wound healing, and Transwell assays to assess proliferation, migration, and invasion. Metabolic assays measured glutamine uptake, α-ketoglutarate production, free fatty acid levels, and the GSH/GSSG ratio, elucidating PPIA’s regulatory effects on metabolism and LUAD progression. Integrative multi-omics analysis identified a fatty acid metabolism–related gene module associated with poor prognosis in LUAD. A four-gene risk model (ERCC1, KYNU, AKR1A1, and PPIA) demonstrated strong predictive power for overall survival across multiple datasets. Single-cell analysis revealed that PPIA was highly expressed in malignant, metabolically active cell populations and strongly correlated with fatty acid and glutamine metabolism. Functional assays confirmed that PPIA silencing inhibited LUAD cell proliferation, migration, and EMT, while reducing key metabolites including glutamine uptake, α-ketoglutarate, free fatty acids, and the GSH/GSSG ratio. Mechanistically, PPIA depletion downregulated c-Myc expression, whereas c-Myc overexpression partially reversed these effects, suggesting that PPIA drives LUAD progression through the c-Myc–mediated fatty acid–glutamine metabolic axis. This study reveals that PPIA promotes the malignant progression of LUAD and affects the immune microenvironment by regulating the c-Myc-mediated fatty acid-glutamine metabolism network remodeling. A prognosis model based on fatty acid metabolism can serve as an effective tool for assessing the prognosis of LUAD patients. Due to incomplete clinical information in some datasets, comprehensive subgroup analyses could not be performed.
Similar content being viewed by others
Introduction
Lung adenocarcinoma (LUAD), the predominant histological subtype of non-small cell lung cancer (NSCLC), accounts for 40 ~ 50% of global lung cancer cases and exhibits rising incidence and mortality trends1. Compounding this challenge, lung adenocarcinoma (LUAD) demonstrates highly aggressive metastatic behavior and is linked to unfavorable clinical outcomes due to its intricate molecular heterogeneity and frequent late-stage diagnosis2.
In recent years, metabolic reprogramming within the tumor microenvironment (TME) has been increasingly recognized as a key driver of cancer progression. Faced with nutrient deprivation and hypoxic conditions, cancer cells reprogram their metabolism to rapidly acquire energy and biosynthetic precursors necessary for sustaining malignant proliferation3. This metabolic shift not only directly influences the metabolic state and functional differentiation of immune cells4, but also contributes to the formation of an immunosuppressive microenvironment by modulating tumor-derived metabolites such as lactate and reactive oxygen species, thereby facilitating immune evasion5. Accumulating evidence indicates that aberrant glucose, amino acid, and lipid metabolism are closely associated with tumor aggressiveness and therapeutic resistance6,7,8. Among these, lipid metabolism—particularly fatty acid metabolism—has emerged as a critical pathway regulating tumor growth, metastasis, and immune responses9. Tumor cells enhance fatty acid synthesis by upregulating key enzymes such as fatty acid synthase (FASN), supporting membrane integrity and energy storage. Concurrently, immune cells undergo metabolic adaptations, including increased fatty acid oxidation, to enable functional differentiation, collectively shaping tumor immunogenicity and response to therapy10.
Notably, glutathione metabolism, an essential antioxidant and metabolic regulatory pathway, exhibits significant crosstalk with fatty acid metabolism. Glutathione (GSH) not only scavenges reactive oxygen species (ROS) to protect tumor cells from oxidative stress, but also enhances their metabolic flexibility by regulating the expression of fatty acid metabolism-related genes, such as FASN and SCD111. Furthermore, α-ketoglutarate (α-KG), a central intermediate of the tricarboxylic acid (TCA) cycle, can promote tumor progression through its influence on epigenetic modifications, such as DNA demethylation, when its metabolism is dysregulated, and contribute to the aberrant activation of the metabolic-immune axis12. As a key regulator of cellular metabolism and proliferation, c-Myc drives tumor cells into a hypermetabolic state by activating fatty acid biosynthesis pathways and suppressing oxidative phosphorylation, while simultaneously impairing anti-tumor immune function through metabolite deprivation13.
Peptidylprolyl isomerase A (PPIA), a member of the peptidylprolyl isomerase (PPIase) family, is a highly conserved protein-folding chaperone14,15,16. With a molecular weight of approximately 18 kDa, PPIA is predominantly cytoplasmic and catalyzes the cis-trans isomerization of proline residues, thereby facilitating protein folding, assembly, and stability17,18,19. Its catalytic activity relies on a hydrophobic pocket at the active site, which binds specific peptide substrates and accelerates protein conformational changes20,21. Recent studies have shown that loss of PPIA suppresses the expression of SLC1A5, a key enzyme in glutamine metabolism, disrupts NRF2-mediated metabolic adaptation, and inhibits tumor growth22.
In this study, we focused on lung adenocarcinoma (LUAD), integrating multi-omics datasets with metabolomic profiling to systematically elucidate the role of metabolic dysregulation in tumor progression and immune evasion. By constructing a prognostic model based on fatty acid metabolism-associated genes, we identified PPIA as a key molecule and explored its potential role in modulating tumor behavior and therapeutic response via the metabolic-immune network. These findings not only provide new insights for molecular subtyping and prognostic assessment in LUAD but also offer a rationale for the clinical translation of metabolic-targeted therapies in combination with immune checkpoint inhibitors.
Materials and methods
Data collection
This study obtained comprehensive multi-omics data for lung adenocarcinoma (LUAD) from the TCGA (The Cancer Genome Atlas) and GEO (Gene Expression Omnibus) databases, which included gene expression profiles, single-cell sequencing data, and detailed clinical and pathological information. Specifically, the TCGA database provides RNA sequencing (RNA-seq) data from 480 LUAD tumor samples and 53 paired normal lung tissue samples. The GEO database includes two bulk transcriptome datasets (GSE11969, n = 54; GSE68465, n = 63) and a single-cell RNA sequencing dataset (GSE150938, n = 4,081 cells). To systematically analyze genes involved in fatty acid metabolism, this study obtained three authoritative gene sets from the Molecular Signatures Database v7.2 (MSigDB): the KEGG fatty acid metabolism pathway, the Reactome fatty acid metabolism gene set, and the Hallmark fatty acid metabolism gene set, ultimately identifying 309 genes closely associated with fatty acid metabolism.
Single-cell RNA sequencing data processing and cell annotation
The processing of single-cell RNA sequencing (scRNA-seq) data and cell annotation was performed according to established protocols. The “Seurat” package was utilized for normalization and quality control of the GSE150938 single-cell data. Specifically, genes expressed in fewer than three cells, cells with fewer than 200 or more than 10,000 genes, and cells with mitochondrial gene expression exceeding 20% were excluded, ensuring the retention of high-quality cell populations. After rigorous filtering, 39,550 cells were retained for further in-depth analysis.
To correct for technical biases, linear regression models and log normalization techniques were applied. The “FindVariableFeatures” function was used to identify the 2,000 most variable genes, which helped capture the primary sources of intercellular heterogeneity. To address potential batch effects, the “RunHarmony” algorithm was employed to integrate and correct the data, effectively eliminating the influence of experimental batches on subsequent analyses. Principal component analysis (PCA) was conducted to reduce the dimensionality of the data, and the relevant components for further analysis were identified. Subsequently, t-distributed stochastic neighbor embedding (t-SNE) was performed for nonlinear dimensionality reduction of the first 20 principal components, revealing distinct clustering patterns among the cell populations. Using the “FindNeighbors” and “FindClusters” functions (with a resolution setting of 0.8), 16 distinct cell clusters were identified and visualized, reflecting the heterogeneity of the LUAD microenvironment at the single-cell level. Differentially expressed genes (DEGs) within each cluster were identified using the “FindAllMarkers” function in “Seurat,” with criteria of min.pct > 0.25, log2(foldchange) > 0.25, and two-sided testing (only.pos = TRUE) to ensure the reliability of the results. Finally, the “SingleR” package was used for accurate cell cluster annotation, revealing the biological characteristics and functions of different cell types, which provided important insights into the LUAD microenvironment.
AUCell
The AUCell R package was used to quantify the activity levels of specific gene sets within individual cells, providing a quantitative tool for assessing the activity of metabolic pathways at the cellular level. This method ranks the genes according to their expression levels based on the area under the curve (AUC) value for model genes, where the AUC value reflects the proportion of cells with high expression of the specific gene sets. Specifically, gene set activity is calculated by evaluating the relative expression levels of each gene within the cell, which is then used to derive an overall activity score for the gene set. A higher score indicates greater activity of the gene set within the cell.
To accurately identify active cell populations, this study used the “AUCell explore Thresholds” function to determine the optimal threshold for gene set activity. This function automatically calculates the best threshold based on the distribution of AUC values, ensuring the objectivity and accuracy of active cell identification. Cells with higher AUC values demonstrate more active expression of gene sets, indicating that specific metabolic pathways are more active in these cells. To visually display the spatial distribution of gene set activity within the cells, this study utilized the “ggplot2” R package to map the AUC scores to the two-dimensional space obtained through t-SNE dimensionality reduction. The resulting density plots and heatmaps effectively visualized active cell clusters. The process involved associating the AUC score of each cell with its two-dimensional coordinates after t-SNE reduction, followed by generating a scatter plot with a color gradient representing the level of gene set activity.
Single-Sample gene set enrichment analysis (ssGSEA) and weighted gene Co-expression network analysis (WGCNA)
In this study, the single-sample gene set enrichment analysis (ssGSEA) method was used to systematically calculate the enrichment score for specific gene sets in each sample. Specifically, to evaluate the overall activity of fatty acid metabolism in LUAD, ssGSEA was applied to the TCGA-LUAD dataset to compute the fatty acid metabolism enrichment score for each sample. This score reflects the combined expression levels of fatty acid metabolism-related genes in the sample, providing a quantitative metric for the tumor’s metabolic characteristics. Weighted Gene Co-expression Network Analysis (WGCNA) was used to construct a scale-free co-expression network, revealing the systematic co-expression relationships between genes. During the analysis, an appropriate soft threshold (soft-thresholding power = 5) was selected to optimize network construction, ensuring that the network’s topological structure approximated a scale-free distribution, which is crucial for the network’s modularity analysis reliability. An adjacency matrix was then constructed based on the Pearson correlation coefficient, which was further transformed into a Topological Overlap Matrix (TOM), with the corresponding dissimilarity matrix (1-TOM) calculated. In this network, nodes represent individual genes, and edge weights indicate the strength of the co-expression relationships between genes. To effectively identify gene co-expression modules, hierarchical clustering was performed on the TOM matrix, with a minimum module size set to 30 genes. Similar modules were merged using a cutting height parameter (abline = 0.25). As a result, 15 co-expression modules were identified. The gray module included genes that could not be classified into other modules and was treated separately for further analysis. By calculating the dissimilarity of module characteristic genes and performing hierarchical clustering, a comprehensive gene co-expression network was successfully constructed, providing a methodological foundation for identifying core modules significantly related to fatty acid metabolism. The study found that one of these modules showed a significant correlation with fatty acid metabolism pathways. This module was selected for further functional validation and mechanistic exploration, offering a new perspective for understanding the role of fatty acid metabolism in LUAD.
Construction and validation of the risk score model
In this study, based on the results of WGCNA, differentially expressed genes significantly associated with fatty acid metabolism were selected as candidate prognostic factors. First, univariate Cox proportional hazards regression analysis (P < 0.05) was employed to identify genes that were significantly associated with patient prognosis from the candidate genes. Subsequently, Lasso regression was performed to further refine the model and reduce the risk of overfitting. Finally, a prognostic risk score model for LUAD was constructed using a multivariate Cox proportional hazards regression model. The TCGA-LUAD dataset was used as the training set for model development, while the GEO datasets (GSE11969 and GSE68465) were used as independent validation sets to evaluate model performance. Patients were classified into high-risk and low-risk groups based on the median risk score. Kaplan-Meier survival analysis was conducted to compare the overall survival between the two groups, and the model’s predictive performance was assessed using the receiver operating characteristic (ROC) curve. To further validate the model’s generalizability, the same analysis process was repeated in two independent GEO datasets, yielding consistent prognostic discrimination ability and confirming that the risk score model can serve as an effective tool for prognostic evaluation in LUAD patients.
Model evaluation
To enhance the clinical applicability of the prognostic model, this study integrated clinical pathological parameters (such as age, sex, and tumor stage) with the risk score to construct a comprehensive prognostic model. Visualization was performed using the “regplot” R package, and calibration curves were employed to assess the consistency between predicted probabilities and observed outcomes. ROC curve analysis was used to evaluate the model’s predictive performance at different time points. Additionally, decision curve analysis (DCA) was conducted to assess the net clinical benefit of the nomogram in decision-making.
scRNA-seq metabolic analysis and virtual knockout
In the scRNA-seq data, patients were categorized into high-expression and low-expression groups based on the median expression level of PPIA. The “scMetabolism” package was used to identify and analyze the metabolic pathway enrichment in the high-expression and low-expression groups of PPIA within single cells, further exploring the metabolic levels across different subpopulations. The scMetabolism package utilizes known metabolic pathway databases, such as KEGG and Reactome, to compute scores for various metabolic pathways using methods like ssGSEA and AUCell.
This study employed the scTenifoldKnk package for virtual gene knockout functional prediction23. This tool constructs a gene regulatory network and simulates network alterations following PPIA gene knockout, identifying affected genes and inferring the potential functions of target genes. The analysis includes three core modules: network construction, virtual knockout, and manifold alignment. The aim of the virtual knockout is to identify genes whose regulation is altered, which are then used to infer the functional roles of target genes within cells.
Biological function of PPIA in RNA-seq
The TIMER database was queried for the expression of PPIA across various cancer types. Based on the expression levels of PPIA, patients were divided into high-expression and low-expression groups, and survival differences were analyzed. Finally, the correlation between PPIA gene expression and enrichment scores was examined.
Cell culture
The normal human lung epithelial BEAS-2B cells and human lung adenocarcinoma cell lines A549 and H1975 used in this study were all purchased from the Cell Resource Center of Shanghai Institutes for Biological Sciences. The cell culture conditions were as follows: BEAS-2B cells were cultured in F12K medium, while A549 and H1975 cells were cultured in RPMI-1640 medium (both products from Gibco BRL, USA), with the addition of 10% fetal bovine serum (FBS) and 1% penicillin-streptomycin solution (Gibco, Invitrogen, USA). All cells were maintained under standard cell culture conditions: 5% CO₂, 95% relative humidity, and a constant temperature of 37 °C.
Cell transfection
To investigate the functional role of PPIA, small interfering RNA (siRNA) technology was used to specifically knock down its expression. The siRNA constructs were purchased from Siwega Biotechnology Co (Wuhan, China), and the pCMV-C-Myc plasmid was obtained from Beyotime Biotechnology (Shanghai, China). The specific sequences are listed in Supplementary Table S1.
Transfection was performed in 6-well plates, with a cell density controlled to 50% confluence. Lipofectamine 3000 (Invitrogen, USA) was used as the transfection reagent, following the manufacturer’s instructions. After transfection, the cells were cultured for 48 h and then subjected to subsequent experimental analysis.
RNA extraction and qRT-PCR
Total RNA from the cell lines was extracted using TRIzol reagent (R1000, Solarbio) according to the standard procedure, and cDNA synthesis was performed using PrimeScript™ RT Master Mix (R232-01, Vazyme). The real-time quantitative PCR (qRT-PCR) reaction mixture used SYBR Green Pro Taq HS Premix (AG11701, Agbio), with GAPDH as the internal control gene to normalize the mRNA expression levels. The relative gene expression levels were calculated using the 2-ΔΔCT method, and the primers used in the experiment were synthesized by Shanghai Shenggong Bioengineering Co., Ltd. (full sequences are shown in Supplementary Table S1).
Cell proliferation assay (CCK-8)
The cell proliferation ability was assessed using the CCK-8 kit (UE, Suzhou, China) according to the manufacturer’s instructions. 1000 cells/well were seeded in a 96-well plate and cultured under the specified experimental conditions. Every 24 h, 10 µL of CCK-8 reagent was added to the serum-free medium, and after 2 h of incubation in the dark, absorbance was measured at 450 nm using a microplate reader. Cell proliferation was monitored over 5 days to plot the growth curve.
Colony formation assay
1000 cells/well were seeded in a 6-well plate, and the experiment was terminated after 14 days of culture. The medium was discarded, and cells were washed twice with PBS, fixed with 4% paraformaldehyde for 15 min, and stained with 0.1% crystal violet solution at room temperature for 30 min. After slow washing with running water, the cells were air-dried. Colonies consisting of more than 50 cells were counted under a microscope, and the colony formation rate was calculated.
Wound healing assay
After 48 h of transfection, when the cells had fused to 95%, a 200 µL sterile pipette tip was used to create a scratch in the monolayer of cells in the 6-well plate. The plate was then gently washed three times with PBS to remove floating cells. Serum-free medium was added for further culture, and images were captured under a microscope at 0 and 48 h. ImageJ software was used to measure the change in scratch width, and the relative migration rate was calculated as: migration rate (%) = (0 h scratch width − 48 h scratch width)/0 h scratch width × 100%.
Transwell migration and invasion assay
Cells were incubated for 24 h in starvation in serum-free basal medium to synchronize their growth condition. Following the conclusion of starvation cultures, cells were enzymatically digested with trypsin and subsequently harvested. Following the removal of the original medium, the cells were resuspended in basal medium. For migration tests, Transwell chambers were positioned on 24-well plates, and 2 × 10^4 cells were inoculated into the top compartment, while complete media with 10% FBS was introduced into the lower chamber. In the invasion assay, matrix glue was initially applied to the upper chamber of the Transwell chamber prior to cell seeding, followed by the identical process utilized in the migration assay. Following a 24-hour culture period, the Transwell chamber was extracted, and the cells were preserved with 4% paraformaldehyde. Following cell fixation, staining was conducted utilizing a 0.1% crystal violet solution. Upon completion of the staining, the residual non-migratory or non-invasive cells in the upper chamber were meticulously scraped away, and the Transwell chamber was positioned beneath an inverted microscope for inspection and photographic documentation.
Immunohistochemical validation
Immunohistochemical staining slides for PPIA in lung cancer and normal lung tissues were obtained from The Human Protein Atlas (HPA) database, and staining intensity and positive cell proportions were analyzed using ImageScope software. The H-score differences between tumor and normal tissues were compared to assess the expression characteristics of PPIA in clinical samples.
Assessment of fatty acid-glutamine network
The transfected A549 and H1975 cells were grown in culture medium for 24 h. According to the manufacturer’s instructions, the Glutamine Assay Kit (Colorimetric) (ab197011), α-Ketoglutarate (alpha KG) Detection Kit (ab83431), GSH/GSSG Ratio Detection Kit (Fluorescent - Green) (ab138881), and Free Fatty Acid Assay Kit - Quantification (ab65341) were used to detect glutamine uptake, α-KG production, GSH/GSSG ratio, and free fatty acid content in lung adenocarcinoma cells.
Western blot analysis
Cells in six-well plates were lysed using RIPA lysis buffer supplemented with protease and phosphatase inhibitors to extract total protein. The protein concentration was quantified with the BCA technique, and the samples were standardized to an equivalent concentration. Identical quantities of protein samples were subjected to SDS-PAGE gel electrophoresis and then transferred to PVDF membranes. The membranes were blocked with 5% BSA in TBST buffer at room temperature for one hour after the transfer. The primary antibody was administered, and the membranes were incubated at 4 °C overnight. Subsequent to the removal of the primary antibody, the membranes were subjected to three washes with TBST buffer, each lasting 5 min. Following this, the HRP-conjugated secondary antibody was introduced, and the membranes were incubated at ambient temperature for 1 h. The membranes were created using ECL, and the bands were documented while the expression levels of the target proteins were assessed by an image analysis system. The antibodies used in this section are as follows: PPIA (Cyclophilin A Polyclonal antibody, 1:1000, Proteintech, Cat No. 10720-1-AP), N-cadherin (N-cadherin Polyclonal antibody, 1:5000, Proteintech, Cat No: 22018-1-AP), E-cadherin (E-cadherin Polyclonal antibody, 1:5000, Proteintech, Cat No: 20874-1-AP), Vimentin (Vimentin Polyclonal antibody, 1:20000, Proteintech, Cat No: 10366-1-AP), c-MYC (c-MYC Polyclonal antibody, 1:12000, Proteintech, Cat No. 10828-1-AP), GAPDH (GAPDH Monoclonal antibody, 1:20000, Proteintech, Cat No: 60004-1-Ig), Rabbit secondary antibody (HRP-conjugated Goat Anti-Rabbit IgG(H + L), 1:7500, Proteintech, Cat No: SA00001-2), Mouse secondary antibody (HRP-conjugated Goat Anti-Mouse IgG(H + L), 1:7500, Proteintech, Cat No: SA00001-1).
Statistical analysis
All data were processed and visualized using R (v4.2.3) and GraphPad Prism (v9.4.0). Survival analysis was conducted using the Kaplan-Meier method and log-rank test (*P < 0.05, **P < 0.01, ***P < 0.001), and the predictive efficacy of the risk score was evaluated using time-dependent ROC curves (AUC > 0.5 is considered effective). After testing the normality of quantitative data, inter-group comparisons were conducted using independent samples t-test or one-way ANOVA, while non-normally distributed data were analyzed using the Wilcoxon rank-sum test or Kruskal-Wallis test. Bivariate correlation analysis uses the Spearman correlation coefficient (ρ).
Two-tailed tests were performed for all statistical analyses, and differences with P values less than 0.05 were considered statistically significant.
Results
Single-cell RNA sequencing data processing
Strict quality control was applied to the single-cell RNA sequencing (scRNA-seq) data in this study, resulting in the exclusion of low-quality cells and the retention of 39,550 high-quality cells for downstream analysis. Quality control criteria included the detection of at least three genes per cell, mitochondrial gene expression below 20%, and each gene being expressed in at least three cells. The 12 analyzed samples showed uniform cell distribution without significant batch effects (Fig. 1A), indicating suitability for further analysis. Dimensionality reduction was performed using t-distributed stochastic neighbor embedding (t-SNE). It categorized these cells into 16 distinct clusters (Fig. 1B). Based on canonical cell marker expression, eight major cell types were identified, including T cells, B cells, endothelial cells, epithelial cells, NK cells, monocytes, macrophages, and smooth muscle cells (Fig. 1C). Fatty acid metabolism activity at the single-cell level was evaluated using the “AUCell” R package, which calculated gene set enrichment scores (AUC values) for each cell. Macrophages, monocytes, and epithelial cells showed higher fatty acid metabolic activity (Fig. 1D). A total of 2,445 marker genes from these three cell types were extracted as fatty acid metabolism-related genes for subsequent analysis.
Annotation of single-cell data and fatty acid metablism activity. (A) The t-SNE plot displayed all cells from the 12 LUAD samples. (B) The t-SNE plot categorized these cells into 16 distinct clusters. (C) The t-SNE map indicated that LUAD samples can be annotated as 8 cell types in the tumor microenvironment. (D) AUCell score of model genes in each cell.
Identification of fatty acid metabolism-related genes and construction of a prognostic risk model
A prognostic risk scoring model for LUAD was constructed based on the TCGA-LUAD cohort. Fatty acid metabolism scores were first calculated for each sample using single-sample Gene Set Enrichment Analysis (ssGSEA). Weighted gene co-expression network analysis (WGCNA) identified gene modules significantly associated with fatty acid metabolism, particularly the turquoise (r = 0.53) and brown (r = 0.39) modules (Fig. 2A and F). By integrating the 2,445 fatty acid metabolism-related genes identified from scRNA-seq data, 70 overlapping genes were screened. Subsequently, univariate Cox regression analysis, LASSO regression for dimensionality reduction, and multivariate Cox regression analysis were performed (Fig. 2G and J). Four key genes—ERCC1, KYNU, AKR1A1, and PPIA—were ultimately selected to construct the prognostic risk model: Risk score = (0.4405 × ERCC1) + (0.1583 × KYNU) + (− 0.8604 × AKR1A1) + (0.4176 × PPIA). High expression of ERCC1 and KYNU, low expression of AKR1A1, and high expression of PPIA were all associated with poor prognosis. The model exhibited strong predictive performance in both the training cohort and two independent GEO validation datasets (AUC > 0.65), and high-risk patients showed significantly reduced overall survival.
Construction and validation of the prognostic model associated with fatty acid metabolism.(A) Clustering dendrogram of whole-genome genes from colorectal adenocarcinoma samples. (B) Heatmap showing the correlation between module eigengenes and fatty acid metabolism. (C, D) Relationship between soft-thresholding power and average connectivity; the scale-free topology model fit index (R²) is displayed on the y-axis. (E, F) Scatter plots of gene significance (GS) versus module membership (MM) in the turquoise module (E) and the brown module (F). (G–J) Prognostic genes were further identified using univariate Cox regression, LASSO regression, and multivariate Cox regression analyses.
Survival analysis and model evaluation
Patients in the TCGA-LUAD cohort were stratified into high-risk and low-risk groups based on the median risk score. Kaplan-Meier survival analysis demonstrated a significant difference in overall survival between the two groups (P < 0.05, Fig. 3A and B), with high-risk patients showing markedly poorer prognosis. This finding was validated in two independent GEO datasets (GSE11969 and GSE68465), which also exhibited significant survival differences (P < 0.05, Fig. 3C and D), confirming the model’s reproducibility and robustness. The predictive performance of the model was further assessed using receiver operating characteristic (ROC) curves. The AUC values in the TCGA dataset and the two GEO validation sets were all above 0.6 (Fig. 3E and H), indicating favorable predictive stability and clinical applicability across datasets. Collectively, these results support the utility of this risk model as an effective tool for prognostic evaluation in LUAD patients.
Performance evaluation of the risk prediction model. (A-D) Kaplan–Meier survival analysis of the prognostic signature across the training set, validation set, and two independent GEO datasets. (E-H) ROC curves demonstrating the predictive performance of the model in the training set, validation set, and two independent GEO datasets.
Construction of the prognostic nomogram
To further enhance the clinical utility of the prognostic model, we integrated clinicopathological parameters—including patient age and tumor stage—with the risk score to construct a personalized prognostic nomogram for patients in the TCGA-LUAD cohort (Fig. 4A). The nomogram was developed using multivariate Cox regression analysis, which determined the weighting coefficients for each variable. Among these, the risk score emerged as the most dominant predictive factor, while clinical features such as age and tumor stage were incorporated to refine survival risk quantification. The calibration curve (Fig. 4B) showed excellent agreement between predicted survival probabilities and actual outcomes, validating the model’s predictive accuracy. To comprehensively assess the model’s performance, time-dependent ROC curve analysis was conducted to evaluate 1-, 3-, 5-, and 10-year survival predictions, with corresponding AUC values of 0.78, 0.74, 0.75, and 0.78 (Figs. 4C–F), indicating the nomogram’s robust predictive capability across multiple time points.
Development of an integrated prognostic nomogram. (A) A prognostic nomogram was developed by integrating clinical parameters with the risk score. (B) Calibration plots for 1-, 3-, 5-, and 10-year overall survival. (C–F) Time-dependent ROC curves for 1-, 3-, 5-, and 10-year survival illustrating the AUC values for clinical variables, the risk score, and the nomogram.*P < 0.05, **P < 0.01, ***P < 0.001.
The expression and biological function of PPIA in LUAD
An in-depth analysis of the PPIA gene revealed the highest hazard ratio (HR) among the studied genes. Using the TIMER database, we evaluated PPIA expression across different cancer types and adjacent normal tissues. Results showed significant overexpression of PPIA in tumor tissues of most cancers (Fig. 5A). Additionally, using the GEPIA database, TCGA-LUAD patients were categorized into high and low PPIA expression groups, with patients in the high expression group exhibiting poorer prognosis (Fig. 5B). To explore the underlying mechanisms of this observation, we analyzed the correlation between PPIA expression and hallmark gene sets (Fig. 5C). Notably, PPIA expression was positively correlated with pathways such as Myc_Target_v1, Myc_Target_v2, glycolysis, oxidative phosphorylation, and TORC1 signaling (all correlations > 0.5).
The expression of PPIA is an important factor affecting the metabolism of lung adenocarcinoma cells
Lung adenocarcinoma originates primarily from epithelial cells, so we focused on analyzing the epithelial cell subpopulation. KEGG enrichment analysis of differentially regulated genes after virtual knockout of the PPIA gene revealed significant enrichment in metabolic pathways, including Glutathione metabolism and Metabolism of xenobiotics by cytochrome P450 (Fig. 5D and E). Cells were divided into high and low PPIA expression groups based on the median expression of PPIA, and the differences in expression levels were visualized (Fig. 5F). Metabolic analysis showed significant differences in various metabolic pathways between the high and low PPIA expression groups (Fig. 6).
The role of PPIA in cancer. (A) Boxplot showing PPIA expression across pan-cancer datasets. (B) Survival analysis indicating the prognostic impact of PPIA expression. (C) Correlation between PPIA expression and 50 hallmark signaling pathways. (D, E) KEGG and GO enrichment analyses of differentially regulated genes after in silico PPIA knockout. (F) Differential expression of PPIA in lung adenocarcinoma epithelial cells.*P < 0.05, **P < 0.01, ***P < 0.001.
PPIA shows significantly high expression in lung adenocarcinoma
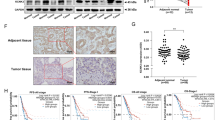
To compare PPIA expression between normal lung cells and LUAD cells, we used the BEAS-2B cell line and two LUAD cell lines, A549 and H1975. qRT-PCR results showed that PPIA expression was significantly elevated in LUAD cell lines (Fig. 7A). PPIA expression was efficiently silenced by siRNA in A549 and NCI-H1975 cells. Four experimental groups were established (si-NC, si-NC + pCMV-C-Myc, si-PPIA, and si-PPIA + pCMV-C-Myc). Both qRT-PCR and Western blot analyses confirmed a marked decrease in PPIA mRNA and protein expression following siRNA transfection, while pCMV-C-Myc co-transfection partially restored PPIA expression (Fig. 7B and D). To assess PPIA expression in patient samples, we analyzed immunohistochemical pathological slides from the HPA database, revealing significantly higher expression in lung adenocarcinoma tissue compared to normal lung tissue (Fig. 7E).
(A) Relative expression level of PPIA. (B) Evaluation of PPIA knockdown efficiency in A549 and NCI-H1975 cells with four experimental groups (si-NC, si-NC + pCMV-C-Myc, si-PPIA, and si-PPIA + pCMV-C-Myc). (C) Western blot analysis and quantification of PPIA protein levels in the same groups. (D) Quantification of PPIA protein levels based on Western blot results in A549 and NCI-H1975 cells. (E) Immunohistochemistry of PPIA in tumor and adjacent normal tissues.
Data are presented as mean ± standard deviation from three independent replicates. Statistical significance was indicated as follows: *P < 0.05, **P < 0.01, ***P < 0.001.
Interfering with PPIA can inhibit the proliferation and metastasis of lung adenocarcinoma cells
In LUAD cell lines A549 and NCI-H1975, CCK-8 assays revealed that PPIA knockdown markedly reduced cell proliferation, whereas co-transfection with pCMV-C-Myc partially reversed this inhibitory effect (Fig. 8A). Consistently, colony formation assays showed that PPIA silencing significantly suppressed colony formation, with both the number and plating efficiency of colonies notably decreased in the si-PPIA group compared with the si-NC group, while Myc overexpression mitigated this reduction (Fig. 8B). In the scratch wound healing assay, the wound closure rate was markedly slower in the si-PPIA group than in the control, and pCMV-C-Myc co-transfection partially restored migratory capacity (Fig. 9A). Similarly, Transwell migration and invasion assays demonstrated that PPIA knockdown significantly inhibited the migratory and invasive abilities of LUAD cells, which were partially rescued by Myc overexpression (Fig. 9B).
(A) CCK-8 assay showed that PPIA knockdown markedly reduced cell viability, while Myc overexpression partially rescued the inhibitory effect. (B) Colony formation assay demonstrated that the number of colonies was significantly decreased in PPIA-silenced cells compared with the control groups, and Myc overexpression attenuated this reduction.
(A) Scratch-wound healing assay revealed that PPIA knockdown significantly delayed wound closure compared with the control groups, whereas Myc overexpression partially reversed this effect. (B) Transwell migration and invasion assays demonstrated that downregulation of PPIA markedly suppressed the migratory and invasive abilities of LUAD cells, while Myc overexpression alleviated these inhibitory effects. all experiments were replicated using two LUAD cell lines, A549 and H1975.Data are presented as mean ± standard deviation from three independent replicates. Statistical significance was indicated as follows: *P < 0.05, **P < 0.01, ***P < 0.001.
Interfering with PPIA can reshape the fatty acid-glutamine network and inhibit EMT through the c-Myc pathway
Western blot analysis was used to verify the protein expression of EMT (epithelial-mesenchymal transition) markers in the si-PPIA and si-NC groups. After PPIA interference, N-cadherin and Vimentin expression significantly decreased, while E-cadherin expression increased (Fig. 10 A and B).
To explore the impact of PPIA on fatty acid and glutamine metabolism reprogramming, we assessed relevant metabolic markers. PPIA interference significantly reduced the levels of glutamine, α-ketoglutarate, GSH/GSSG, and free fatty acids. c-Myc is a key factor in the glutamine metabolic pathway, directly influencing the transport of glutamine into cells (Fig. 10 C F). Western blot analysis revealed that c-Myc expression was significantly downregulated in LUAD cells following PPIA knockdown, whereas co-transfection with pCMV-C-Myc partially restored c-Myc levels (Fig. 10G). Consistently, densitometric quantification and RT-qPCR confirmed this trend in both A549 and NCI-H1975 cells (Fig. 10H J), suggesting that c-Myc overexpression may partially reverse the molecular and cellular effects induced by PPIA depletion. These findings indicate that PPIA knockdown reduces c-Myc expression, which may contribute to the observed suppression of glutamine and fatty acid metabolism in LUAD cells.
(A) Western Blot of EMT markers after interfering with PPIA. (B) Statistical plot of EMT markers in A549 cells and NCI-H1975 cells after interfering with PPIA. (C) Quantification of relative glutamine uptake following PPIA knockdown in A549 and NCI-H1975 cells. (D) Quantification of relative Alpha Ketoglutarate production following PPIA knockdown in A549 and NCI-H1975 cells. (E) Quantification of GSH/GSSG ration following PPIA knockdown in A549 and NCI-H1975 cells. (F) Quantification of FFA following PPIA knockdown in A549 and NCI-H1975 cells. (G) Western blot analysis of c-Myc protein expression in A549 and NCI-H1975 cells following PPIA knockdown and/or Myc overexpression. (H, J) RT-qPCR analysis of c-Myc mRNA expression in A549 and NCI-H1975 cells after PPIA knockdown and/or Myc overexpression. (I) Quantification of c-Myc protein levels based on Western blot results in A549 and NCI-H1975 cells.
Discussion
Despite the significant heterogeneity of the tumor microenvironment (TME) across various cancer types, all tumor cells share the common feature of acquiring nutrients through metabolic reprogramming to support their rapid proliferation. To maintain mitochondrial function and the normal operation of the TCA cycle, tumor cells have developed a unique glutamine replenishment pathway26. As the most abundant free amino acid in the blood, glutamine is extensively taken up by tumor cells, primarily through the ASCT2 and SN2 transporters. It is then converted into glutamate by glutaminase (GLS/GLS2), which is further transformed into α-ketoglutarate to replenish TCA cycle intermediates27. This metabolic pattern makes tumor cells highly dependent on glutamine, resulting in what is known as “glutamine addiction,” where glutamine deprivation leads to tumor cell growth arrest or even cell death28. Recent single-cell RNA sequencing studies have revealed significant metabolic heterogeneity among different tumor subgroups, with some particularly dependent on glutamine metabolism to maintain their stemness and invasiveness. This provides a theoretical foundation for precision therapies targeting glutamine metabolism.
Using WGCNA and AUCell algorithms, we identified genes strongly associated with fatty acid metabolism and selected four key genes—ERCC1, KYNU, AKR1A1, and PPIA—to construct a risk score model. The fatty acid metabolism-based risk score model demonstrated strong predictive performance across multiple independent datasets and was significantly associated with various clinical parameters. Kaplan-Meier survival curves showed that patients with high-risk scores had significantly poorer prognoses than those with low-risk scores. The model’s predictive efficacy for patient survival was evaluated using ROC curves and prognostic nomograms, demonstrating strong predictive power.
Among the four genes included in the model, PPIA attracted particular attention due to its strong association with lipid metabolic processes and patient prognosis. This study is the first to systematically reveal the key regulatory role of PPIA in lung adenocarcinoma (LUAD), demonstrating that PPIA not only plays a critical role in tumor microenvironment remodeling but also promotes tumor progression by influencing the fatty acid-glutathione metabolic network and EMT processes. Using single-cell RNA sequencing (scRNA-seq) and RNA-seq analysis, we found that PPIA expression was significantly upregulated in macrophages, monocytes, and epithelial cells, and was closely associated with fatty acid metabolic activity. Differential expression analysis and Kaplan-Meier survival curves further confirmed that high PPIA expression was associated with shorter survival in LUAD patients, suggesting that it could serve as a potential independent prognostic marker. Single-gene GSVA analysis revealed that high PPIA expression was linked to enrichment in the Myc_Target_v1, Myc_Target_v2, glycolysis, oxidative phosphorylation, and mTORC1 signaling pathways. In silico knockout analysis showed that PPIA knockout significantly enriched genes involved in glutathione metabolism in LUAD cells. These findings suggest that PPIA may be closely linked to energy metabolism pathways associated with LUAD growth, and interfering with its expression may affect glutathione-related metabolic pathways.
Based on the results of bioinformatics analyses, we performed a series of in vitro experiments to validate the functional role of PPIA in LUAD. qRT-PCR, Western blot, and immunohistochemistry consistently confirmed that PPIA was markedly overexpressed in LUAD tissues, and its expression level was significantly associated with patient prognosis. CCK-8 and colony formation assays demonstrated that PPIA knockdown significantly suppressed the proliferative capacity of LUAD cells. Wound healing and Transwell migration/invasion assays showed that silencing PPIA markedly impaired the migratory and invasive potential of LUAD cells. Western blot analysis further revealed that PPIA knockdown led to increased expression of E-cadherin and reduced levels of N-cadherin and Vimentin, suggesting that inhibition of PPIA suppresses LUAD progression by attenuating the epithelial-mesenchymal transition (EMT) process. To further elucidate the metabolic consequences of PPIA knockdown and validate the bioinformatics predictions, we assessed c-Myc expression and key metabolic intermediates in LUAD cells. Both mRNA and protein levels of c-Myc were significantly reduced upon PPIA knockdown. In addition, intracellular levels of glutamine, α-ketoglutarate (α-KG), and the GSH/GSSG ratio were all significantly decreased. Notably, PPIA knockdown also led to a marked reduction in free fatty acid levels, indicating disrupted lipid metabolism.
High PPIA expression is strongly associated with fatty acid metabolism in LUAD, and its knockdown substantially disrupts fatty acid metabolic processes in tumor cells. As a central regulator of cellular metabolism, c-Myc facilitates glutamine uptake by upregulating the amino acid transporter SLC1A529,30. Knockdown of PPIA results in downregulation of c-Myc expression, which leads to reduced glutamine uptake and subsequent depletion of downstream metabolites such as α-ketoglutarate (α-KG) and glutathione (GSH). However, the underlying mechanism by which PPIA regulates c-Myc expression (e.g., through transcriptional regulation, modulation of protein stability, or other indirect pathways) remains unclear and requires further investigation.
Notably, the decreased GSH/GSSG ratio compromises the cellular antioxidant defense against reactive oxygen species (ROS), thereby increasing susceptibility to oxidative stress and triggering programmed cell death31,32. The reduction in glutamine metabolism following PPIA knockdown may also impair both the tricarboxylic acid (TCA) cycle and fatty acid metabolism. As a critical intermediate in energy production, α-KG depletion disrupts these pathways and hinders metabolic homeostasis33. Recent studies have shown that miRNA-mediated regulation of cell-cycle and cytoskeletal pathways profoundly affects LUAD prognosis34, underscoring a broader network where non-coding RNAs cooperate with metabolic drivers such as PPIA and c-Myc to shape tumor progression and therapeutic response. Collectively, these findings suggest that PPIA holds potential as a therapeutic target in LUAD, with preliminary evidence indicating that targeting PPIA may exert anti-tumor effects by simultaneously interfering with glutamine metabolism, lipid utilization, and redox homeostasis—though further preclinical validation is necessary to confirm its translational value.
In summary, this study constructed and validated a prognostic model based on fatty acid metabolism-related genes, which exhibited strong predictive accuracy for LUAD prognosis. Among the identified genes, PPIA emerged as a potential high-risk biomarker, and in vitro experiments demonstrated that PPIA interference suppressed LUAD cell proliferation, migration, and invasion. Mechanistically, PPIA knockdown simultaneously decreased c-Myc expression and disrupted glutamine metabolism(including fatty acid, α-ketoglutarate and glutathione), positioning PPIA as a promising therapeutic target that could simultaneously impact fatty acid and glutamine metabolism in LUAD.
However, this study has several limitations that should be acknowledged. First, all functional experiments were conducted in vitro, and the lack of in vivo validation limits the generalizability of our findings, particularly given the immunological and systemic complexities of the tumor microenvironment in living organisms. Future work will include in vivo models such as xenograft assays and immune checkpoint inhibitor (ICI) treatment models to validate the role of PPIA in LUAD progression and its biological relevance. Second, while we propose PPIA as a potential therapeutic target, our current data are insufficient to fully support its translational readiness, and the discussion of its therapeutic potential should be interpreted with caution until additional preclinical data (e.g., in vivo efficacy of pharmacological inhibition) are obtained. Third, we did not perform subgroup analyses to evaluate the prognostic model’s performance across key clinical subgroups such as age, sex, tumor stage, or tumor mutational burden (TMB) level, which may limit our understanding of the model’s applicability in diverse patient populations. If feasible, future studies should incorporate these subgroup analyses to enhance the clinical relevance of the model. Fourth, as highlighted earlier, the mechanism underlying the regulatory interaction between PPIA and c-Myc remains speculative. To address this, future experiments could include targeted assays such as promoter activity analysis, ChIP-qPCR, or co-immunoprecipitation to clarify how PPIA modulates c-Myc expression.
Conclusion
The prognostic model constructed based on fatty acid metabolism-related genes demonstrates strong predictive accuracy. Among these, PPIA, identified as a high-risk prognostic factor, appears to promote the aggressive progression of LUAD by regulating c-Myc protein levels and driving the reprogramming of the fatty acid–glutamine metabolic network. Further in vivo and mechanistic studies are warranted to validate the therapeutic potential of PPIA and clarify its precise role in LUAD pathogenesis.
Data availability
To access the data supporting this study’s findings, please contact the corresponding author with a reasonable request.
References
Takamori, S., Komiya, T. & Powell, E. Survival benefit from immunocheckpoint inhibitors in stage IV non-small cell lung cancer patients with brain metastases: A National cancer database propensity‐matched analysis[J]. Cancer Med. 10 (3), 923–932. https://doi.org/10.1002/cam4.3675 (2021).
Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 Countries[J]. Cancer J. Clin. 71 (3), 209–249. https://doi.org/10.3322/caac.21660 (2021).
Pavlova, N. N. & Thompson, C. B. The emerging hallmarks of cancer Metabolism[J]. Cell Metabol. 23 (1), 27–47. https://doi.org/10.1016/j.cmet.2015.12.006 (2016).
Vander Heiden, M. G., Cantley, L. C. & Thompson, C. B. Understanding the Warburg effect: the metabolic requirements of cell Proliferation[J]. Science 324 (5930), 1029–1033. https://doi.org/10.1126/science.1160809 (2009).
Carracedo, A., Cantley, L. C. & Pandolfi, P. P. Cancer metabolism: fatty acid oxidation in the limelight[J]. Nat. Rev. Cancer. 13 (4), 227–232. https://doi.org/10.1038/nrc3483 (2013).
Zhao, Y., Butler, E. B. & Tan, M. Targeting cellular metabolism to improve cancer therapeutics[J]. Cell Death Dis. 4 (3), e532–e532. https://doi.org/10.1038/cddis.2013.60 (2013).
Vander Heiden, M. G. Targeting cancer metabolism: a therapeutic window opens[J]. Nat. Rev. Drug Discovery. 10 (9), 671–684. https://doi.org/10.1038/nrd3504 (2011).
Menendez, J. A. Fine-tuning the lipogenic/lipolytic balance to optimize the metabolic requirements of cancer cell growth: Molecular mechanisms and therapeutic perspectives[J]. Biochimica et Biophysica Acta (BBA) - Molecular and Cell Biology of Lipids 1801(3), 381–391. https://doi.org/10.1016/j.bbalip.2009.09.005 (2010).
Chen, X., Chen, S. & Yu, D. Metabolic reprogramming of chemoresistant cancer cells and the potential significance of metabolic regulation in the reversal of cancer Chemoresistance[J]. Metabolites 10 (7), 289. https://doi.org/10.3390/metabo10070289 (2020).
Corazao-Rozas, P. et al. Mitochondrial oxidative phosphorylation controls cancer cell’s life and death decisions upon exposure to MAPK inhibitors[J]. Oncotarget 7 (26), 39473–39485. https://doi.org/10.18632/oncotarget.7790 (2016).
Luo, Y. et al. Fatty acid metabolism and cancer Immunotherapy[J]. Curr. Oncol. Rep. 24 (5), 659–670. https://doi.org/10.1007/s11912-022-01223-1 (2022).
Li, Y., Wan, Y. Y., Zhu, B. & Pan, B. Immune Cell Metabolism in Tumor Microenvironment[M]//Li F. Immune Metabolism in Health and Tumor: 1011, 163–196. (Dordrecht: Springer Netherlands). (2017). https://doi.org/10.1007/978-94-024-1170-6_5.
Kopecka, J. et al. Phospholipids and cholesterol: inducers of cancer multidrug resistance and therapeutic targets[J]. Drug Resist. Updates. 49, 100670. https://doi.org/10.1016/j.drup.2019.100670 (2020).
Zhang, L. et al. PPIA is a novel adipogenic factor implicated in obesity[J]. Obesity 23 (10), 2093–2100. https://doi.org/10.1002/oby.21208 (2015).
Han, J. M., Jung, H. J. & Cyclophilin A/CD147 interaction: A promising target for anticancer Therapy[J]. Int. J. Mol. Sci. 23 (16), 9341. https://doi.org/10.3390/ijms23169341 (2022).
Palacín, M. et al. A search for cyclophilin‐A gene (PPIA) variation and its contribution to the risk of atherosclerosis and myocardial infarction[J]. Int. J. Immunogenet. 35 (2), 159–164. https://doi.org/10.1111/j.1744-313X.2008.00755.x (2008).
Liu, J., Chen, C. M. & Walsh, C. T. Human and Escherichia coli cyclophilins: sensitivity to Inhibition by the immunosuppressant cyclosporin A correlates with a specific Tryptophan residue[J]. Biochemistry 30 (9), 2306–2310. https://doi.org/10.1021/bi00223a003 (1991).
Davis, T. L. et al. Structural and biochemical characterization of the human Cyclophilin family of Peptidyl-Prolyl Isomerases[J]. PLoS Biol. 8 (7), e1000439. https://doi.org/10.1371/journal.pbio.1000439 (2010).
Song, F. et al. Cyclophilin A (CyPA) induces chemotaxis independent of its Peptidylprolyl Cis-Trans isomerase Activity[J]. J. Biol. Chem. 286 (10), 8197–8203. https://doi.org/10.1074/jbc.C110.181347 (2011).
Foster, T. L. et al. Cyclophilin A interacts with domain II of hepatitis C virus NS5A and stimulates RNA binding in an Isomerase-Dependent Manner[J]. J. Virol. 85 (14), 7460–7464. https://doi.org/10.1128/JVI.00393-11 (2011).
Lauranzano, E. et al. Peptidylprolyl isomerase A governs TARDBP function and assembly in heterogeneous nuclear ribonucleoprotein complexes[J]. Brain 138 (4), 974–991. https://doi.org/10.1093/brain/awv005 (2015).
Lu, W. et al. PPIA dictates NRF2 stability to promote lung cancer progression[J]. Nat. Commun. 15 (1), 4703. https://doi.org/10.1038/s41467-024-48364-4 (2024).
Osorio, D. et al. ScTenifoldKnk: an efficient virtual knockout tool for gene function predictions via single-cell gene regulatory network perturbation[J]. Patterns 3 (3), 100434. https://doi.org/10.1016/j.patter.2022.100434 (2022).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes[J]. Nucleic Acids Res. 28 (1), 27–30. https://doi.org/10.1093/nar/28.1.27 (2000).
Kanehisa, M. et al. KEGG as a reference resource for gene and protein annotation[J]. Nucleic Acids Res. 44 (D1), D457–D462. https://doi.org/10.1093/nar/gkv1070 (2016).
Jin, L., Alesi, G. N. & Kang, S. Glutaminolysis as a target for cancer therapy[J]. Oncogene 35 (28), 3619–3625. https://doi.org/10.1038/onc.2015.447 (2016).
Durán, R. V. & Hall, M. N. Glutaminolysis feeds mTORC1[J]. Cell. Cycle. 11 (22), 4107–4108. https://doi.org/10.4161/cc.22632 (2012).
Yang, L., Venneti, S., Nagrath, D. & Glutaminolysis A hallmark of cancer Metabolism[J]. Annu. Rev. Biomed. Eng. 19 (1), 163–194. https://doi.org/10.1146/annurev-bioeng-071516-044546 (2017).
Li, X. et al. Glutamine addiction in tumor cell: oncogene regulation and clinical treatment[J]. Cell. Communication Signal. 22 (1), 12. https://doi.org/10.1186/s12964-023-01449-x (2024).
Hu, X., Jin, H. & Zhu, L. Effect of glutamine metabolism on chemoresistance and its mechanism in tumors[J]. J. Zhejiang Univ. (Medical Sciences). 50 (1), 32–40. https://doi.org/10.3724/zdxbyxb-2021-0040 (2021).
Georgiou-Siafis, S. K. & Tsiftsoglou, A. S. The Key Role of GSH in Keeping the Redox Balance in Mammalian Cells: Mechanisms and Significance of GSH in Detoxification via Formation of Conjugates[J]. Antioxidants, 12(11), 1953. (2023). https://doi.org/10.3390/antiox12111953
Xiong, Y. et al. Engineering nanomedicine for glutathione depletion-augmented cancer therapy[J]. Chem. Soc. Rev. 50 (10), 6013–6041. https://doi.org/10.1039/D0CS00718H (2021).
Yuan, Y. et al. Exercise-induced α‐ketoglutaric acid stimulates muscle hypertrophy and fat loss through OXGR1‐dependent adrenal activation[J]. EMBO J. 40 (14), e108434. https://doi.org/10.15252/embj.2021108434 (2021).
Sahin, Y. et al. Investigating miR-30a’s tumor suppressor role and its targets (MYBL2, TWF1, GALNT7, PFN2) as prognostic biomarkers in lung adenocarcinoma. Biochem. Biophys. Rep. 43, 102122. https://doi.org/10.1016/j.bbrep.2025.102122 (2025).
Funding
This work was financially supported by the Guangxi Natural Science Foundation (Grant No. 2025GXNSFHA069094); The Yunnan Key Laboratory of Laboratory Medicine & Yunnan Province Clinical Research Center for Laboratory Medicine (Grant No. 2017DG0005-2024-02); The Zhejiang Key Laboratory of Diagnosis & Treatment Technology on Thoracic Oncology (Lung and Esophagus) (Grant No. 2022K001); The Xiansheng Clinical Research Special Fund of the China International Medical Exchange Foundation (Grant No. Z-2014-06-2102); The Open Project Program of Chongqing Key Laboratory of Development and Utilization of Genuine Medicinal Materials in the Three Gorges Reservoir Area (Grant No. KFKT2022012); The Fujian Provincial Key Laboratory of Innovative Drug Target Research (Grant No. FJ-YW-2025KF06); and The Guangxi Undergraduate Innovation and Entrepreneurship Training Program (Grant No. S202510599070).
Author information
Authors and Affiliations
Contributions
GY: Conceptualization, Data curation, Formal analysis, Investigation, Software, Writing-original draft, Writing-review & editing; WW: Conceptualization, Data curation, Visualization, Writing-original draft; BJ: Conceptualization, Formal analysis, Software, Writing-original draft; XZ: Data curation, Investigation, Software; LJ: Data curation, Investigation, Formal analysis, Visualization, Writing-original draft; LT: Methodology, Software, Writing-original draft; XX: Formal analysis, Writing-original draft; YL: Investigation, Writing-original draft; QX: Methodology, Writing-original draft; LJ: Conceptualization, Supervision, Methodology, Resources, Funding acquisition, Formal analysis, Data curation, Writing–original draft, Writing-review & editing, Organizing and training this research team.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Consent for publication
All listed authors have actively participated in the study and have read and approved the submitted manuscript.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Gan, Y., Wei, W., Jiang, B. et al. PPIA regulates fatty acid and glutamine metabolism in lung adenocarcinoma based on multiomics prognostic model and experiment validation. Sci Rep 16, 4154 (2026). https://doi.org/10.1038/s41598-025-34313-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-34313-8