Abstract
The traditional stethoscope auscultation used for the detection of lung diseases is hampered by its poor sensitivity, complexity of sound, and dependence on clinical competence. As a result, it frequently leads to diagnostic errors, treatment delays, and inaccessibility in settings with limited resources. These difficulties highlight the need for automated, inexpensive, and portable diagnostics. This paper presents the lightweight two-layer 1D-CNN (one-dimensional convolutional neural network) for lung illness classification called LDSC. The signals are processed using normalized MFCC (Mel-Frequency Cepstral Coefficients) features after being resampled to 4 kHz, enhanced (noise, time-stretch, pitch-shift), and divided into 3-second frames. According to simulation results, the suggested scheme achieves 98% accuracy, sensitivity, and specificity on the ICBHI (International Conference on Biomedical and Health Informatics) dataset and 99% on the KAUH (King Abdullah University Hospital) dataset, respectively.
Similar content being viewed by others
Introduction
Traditionally, a stethoscope is used to auscultate pulmonary sounds to diagnose respiratory disorders. Clinicians and specialists use this method for preliminary evaluations. Normal and abnormal pulmonary sounds are the two basic categories. Unusual noises frequently indicate problems such as lung fluid accumulation, infections, inflammation, or obstructions. Conventional handheld stethoscopes, on the other hand, may not be sensitive enough to recognize the unique acoustic traits of certain illnesses, which could lead to incorrect diagnoses or the failure to detect underlying disorders. Lung sounds are naturally complicated, which makes it harder to figure out what they are and analyze them. So, making an automated detection system is very important for improving the accuracy of diagnoses and fixing the problems with traditional methods1.
According to the Global Respiratory Societies Forum (FIRS), pulmonary disease is one of the leading causes of severe illness worldwide and kills over 4 million people annually. In 2017, the World Health Organization (WHO) declared that, after cardiovascular disorders, chronic respiratory diseases are the second most common cause of illness globally. They account for roughly 10% of all diseases worldwide1. Because of fluctuating signals, noise interference, and the requirement for intricate hybrid models that require a lot of processing power, auscultation-based automation is slow. Even though these methods have improved, imaging-based diagnostics like CT scans and X-rays are still costly, exposing patients to radiation, and are unavailable in areas with limited resources. This distinction makes it more difficult to establish real-time, point-of-care telemedicine in low-income areas. By creating LDSC, a lightweight 1D-CNN model for automated lung sound classification, this paper helps to close this gap. To enable the use of edge devices, one of the specific objectives is to achieve high accuracy with the fewest number of layers. Improving cross-dataset generalization by MFCC features and augmentation is another objective. Lastly, this method’s superior efficiency over the best hybrids is demonstrated using the ICBHI and KAUH datasets.
Comparative assessments of three machine learning methodologies for lung sound classification have demonstrated that classifiers such as Support Vector Machines (SVM), k-nearest Neighbors (KNN), and Gaussian Mixture Models have effectively utilized manually curated data. Some studies have used Mel-frequency cepstral coefficients (MFCCs) as input features for SVM, while others have used spectrogram pictures as input for Convolutional Neural Networks (CNN).
One of the specific goals is to get high accuracy with as few layers as possible so that edge devices can be used. Another goal is to make cross-dataset generalization better by using MFCC features and adding more data. Lastly, the ICBHI and KAUH datasets are used to show that the new hybrids are more efficient than the best ones on the market. Initial studies have employed various traditional machine learning (ML) techniques to detect lung sounds. Hilbert-Huang Transform has demonstrated potential in effectively detecting crackles by extracting time-frequency distribution data from signals. Fast Fourier Transform (FFT) and self-organizing maps have also been used to find features and sort breathing sounds2.
Over the past ten years, deep learning (DL) has become more and more popular because it can automatically extract features and get around the problems with traditional DL. It has been used to correctly diagnose many diseases, such as COVID-19, diabetic retinopathy (DR), and breathing problems. Feedforward neural networks (FNNs) and recurrent neural networks (RNNs) have been used to look at lung sound data3.
This paper presents a novel deep learning model for lung sound-based respiratory disease diagnosis using 1D-CNN. A multi-label classification framework that allows each signal to have multiple labels. Besides, a simpler architecture with two convolutional layers for ease of implementation. The use of appropriate kernels for effective feature extraction and lower computational costs due to shorter training time are among its most important features. Based on extensive experiments and testing on two separate datasets, this model outperforms existing approaches. The proposed model outperforms current approaches4. This strategy is motivated by the ability of 1D-CNNs to efficiently extract deep features from input data while capturing and preserving the specific temporal dependencies of successive sound events. Because 1D-CNNs directly process one-dimensional sequential data, they are superior to standard CNNs for sound-related tasks.
The main contributions of this paper are as follows.
-
It develops a simpler and more user-friendly architecture with fewer layers.
-
It provides a framework that distinguishes between binary and multiclass classification tasks.
-
Furthermore, it ensures the robustness of the framework by testing it on two separate datasets and comparing its results with previous approaches.
This study seeks to clarify its objectives and contributions by examining the following research questions:
RQ1: Can a 1D-CNN with only two layers match or exceed the performance of hybrid models in lung sound classification?
RQ2: How does MFCC combined with augmentation make it easier to generalize across datasets?
RQ3: What are the computational advantages of LDSC compared to SOTA?
The rest of the paper is organized as follows: A review of the literature on deep learning methods for lung disease detection is provided in Sect. 2. A thorough explanation of the suggested model and technique is given in Sect. 3. The performance evaluation and discussion are presented in Sect. 4, and the study is concluded in Sect. 5.
Literature review
The classification of lung sounds has evolved significantly in recent years thanks to the application of cutting-edge technologies, such as deep learning techniques. These developments have led to a significant improvement in the accuracy of the diagnosis of respiratory disease. CT scans and lung sound analysis can diagnose numerous lung disorders. This section categorizes prior work, with results summarized in Table 1. In5, the authors proposed a unique method using an autoencoder-based hybrid CNN-LSTM model to determine the extent of lung involvement during the COVID-19 pandemic. In6, the authors introduced the CNN-LSTM approach for identifying lung involvement in one of the most severe and widespread diseases in recent years. Several studies employed deep learning-based image processing methods to identify lung cancer early7. Using a multi-level feature extraction-based methodology, the authors of8 proposed a CNN-LSTM network for automated coronavirus detection from CT scans and X-ray pictures. In9, the authors presented an automatic COVID-19 detection technique that uses X-ray images and combines long short-term memory (LSTM) for classification with CNN for feature extraction. The system aims to give accurate and timely COVID-19 diagnosis, with room for improvement as more data becomes available. For biomedical applications, CNN provides improved performance, reduced computational overhead, and higher efficiency. Using X-ray and CT images, the authors in10 classified lung diseases such as pneumonia, tuberculosis, and lung cancer using CNN with several models.
By adopting image augmentation techniques, the authors of11 could effectively handle unbalanced datasets and offer a methodology for classifying lung illnesses, including pneumonia and COVID-19, using chest X-ray images. Tabish Saeed et al. highlighted the challenges of cough-based AI diagnosis for respiratory illnesses due to confounding variables12. They developed the Bias-Free Network (RBF-Net), which combines a conditional GAN with a hybrid CNN-LSTM feature encoder to mitigate the effects of biases such as age, gender, and smoking status. It exhibits resistance to these factors. In13, Li et al. developed a model that combines CNN and LSTM to predict and diagnose respiratory diseases using CT data and medical information. The model fared better than traditional methods in terms of precision and F1 scores. In14, Kanipriya et al. introduced a unique method for classifying lung nodule abnormalities using an ideal hybrid CNN-LSTM model. The technique employs entropy-based K-means clustering for nodule location. The technique employs an improved Capuchin Search Algorithm (ICSA) for hyperparameter optimization after segmentation and classification. The authors of15 looked at how well different deep-learning models could use raw lung auscultation sounds to figure out what was wrong with someone’s breathing. The models worked even better when more data was added to them. In16, the authors achieved optimal results on the ICBHI 2017 Respiratory Sound Database by employing a hybrid neural model for lung sound classification. This model uses deep learning and a focal loss function to fix data imbalance.
Farivan et al.17 investigated the use of deep learning for lung sound classification. The CNN + BDLSTM model was used for disease classification, and cross-validation was employed to test its performance. The results show that deep learning can enhance clinical judgment in diagnosing pulmonary diseases. A method for predicting pulmonary diseases such as COVID-19 and pneumonia using chest X-ray images is presented in18. Deep learning techniques for disease classification, region of interest extraction, feature extraction, dataset recovery, and image quality enhancement are all included in this framework. It focuses on RNN and LSTM to improve the accuracy of the results.
In19, the authors developed a computational method for predicting lung diseases that uses a bidirectional LSTM and a multi-head self-attention layer based on a convolutional neural network. The method, which has been tested on many medical imaging datasets, outperformed existing methods in disease classification. Yiwen Zhou and colleagues20 investigated how deep learning networks can be used to predict outcomes in people with stage III non-small cell lung cancer (NSCLC). They did this by looking at a series of CT scans. Their AI-powered approach significantly improved survival and prediction for cancer. The authors of21, proposed two CNN-based models for diagnosing lung cancer based on CT images. The hybrid 3D-CNN with SVM outperformed traditional methods by addressing the complexity of feature extraction.
In22, the authors developed the deep learning model EasyNet to detect pulmonary diseases using pulmonary sound waves from the ICBHI and KAUH datasets. This model surpasses state-of-the-art methods in both binary and multi-class detection tasks. In23, the authors used a large dataset and deep learning techniques to get good precision and recall when classifying abnormalities in chest X-rays. They did this by combining CNN for classification with LSTM models for annotation. In24, the authors talked about new developments, obstacles to more research, and new trends in medical image analysis while looking at how machine learning techniques, like CNNs, can be used.
A deep learning model developed by Goram Mofarah and colleagues25 classifies chest X-ray images into different groups using a CNN and the Visual Geometry Group (VGG) 19. This facilitates a more accurate diagnosis. To improve the accuracy of diagnosis, N. Sudhir and colleagues26 emphasized the need for early detection of lung cancer using sophisticated deep learning algorithms, especially CNNs.
In27, the authors investigated the use of convolutional neural networks (CNN) to detect abnormalities in chest X-ray images, combining computer image processing with the skill of an experienced radiologist to improve diagnostic accuracy. The authors of28 made Lung-Deep, a computer-aided diagnostic (CAD) system that uses a multi-layer deep learning method to find patterns in lung nodules. In29, recent studies on CNN applications for medical image processing were reviewed, concentrating on improvements in tissue segmentation, classification, and lesion diagnosis. In30, the authors showed how to use deep learning to find lymph node involvement on histopathology slides. They focused on better results with different algorithms like VGG-16 and Inception V3. It was better to diagnose and treat interstitial lung disease (ILD) when the authors of31, used pre-trained models and a transformer method to combine radiologic, pathological, and clinical data. While imaging-based deep learning models have shown promising results in lung disease detection from radiology data32,33,34, they require expensive equipment (e.g., X-ray, CT scanners) and trained radiologists, limiting accessibility in low-resource settings. In contrast, our LDSC model leverages low-cost digital stethoscopes and 1D-CNN on lung sounds, enabling real-time, point-of-care diagnosis without imaging infrastructure.
Method and material
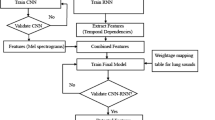
shows a block diagram of the proposed approach and lists the different steps taken in this research.
Figure 1 LDSC workflow: (a) Raw lung sound signal (time domain). (b) Resampled signal at 4 kHz. (c) Augmented signal (noise-added example). (d) 3-second frame segmentation. (e) Extracted MFCC features (13 coefficients). (f) Model prediction (frame-level probability).
Before starting the proposed LDSC model, the raw signals go through a number of pre-processing steps. Two separate datasets are used, each sampled at 4 kHz, 10 kHz, or 44.1 kHz. To keep the sampling rate the same in both datasets, a standard frequency of 4 kHz has been used. Lung sound signals usually range from 50 Hz to 2000 Hz. The Nyquist theorem says that a minimum sample rate of 4 kHz is needed for accurate signal reconstruction. This indicates that the fundamental characteristics of lung sounds can be effectively captured at this frequency. The model becomes more stable when noise is added to the raw signal before resampling. The dataset is also improved by various audio augmentation techniques, such as adding random noise, stretching the signal’s time (altering its speed), and shifting the pitch (modifying the pitch)35. After augmentation, the signals are divided into frames, and MFCC features36 are taken from each frame. After that, these features are normalized using the min-max method. The LDSC model then uses these processed signals for training and testing. It sorts them into normal or one of the disease categories. Algorithm 1 shows the pseudocode of the proposed scheme.
LDSC: 1D-CNN for respiratory sound classification.
Datasets
In this section, two reliable and targeted datasets for breath sound analysis are presented to evaluate the performance of the models. Table 2 provides a summary of the ICBHI and KAUH datasets along with their details. A summary of each dataset is given below, and the summary is given in Table 3.
-
ICBHI Dataset The dataset consists of two sets of annotations and includes 920 recordings from 126 participants. One set contains 6,898 respiratory cycles, annotated to indicate the presence of crackles, wheezes, neither, nor both phenomena. The ICBHI dataset contains 10,775 lung sound events collected from two independent research centers in different countries. Most recordings were made in the Laboratory of Research and Respiratory Rehabilitation (Lab3R) of Avir University and Infant D. Hospital. Pedro is obtained in Avire, Portugal. The research team collected the remaining recordings at Papanikolaou General Hospital in Thessaloniki and Imatia General Hospital in Greece. This dataset contains 920 signals with a duration between 10 and 90 s, sampled at frequencies of 4 kHz, 10 kHz, and 44.1 kHz15,37.
-
KAUH Dataset The dataset comprises respiratory sound recordings collected using an electronic stethoscope at King Abdullah University Hospital. This collection contains 337 recordings from 112 people, of which 35 are healthy, and 77 are people with respiratory disorders. Each person has a recording that lasts between 5 and 30 s and is enough to record at least one complete breathing cycle15.
Preprocessing
In the preprocessing stage, audio signals are resampled to 4 kHz (4000 Hz) to ensure uniformity across both datasets. Subsequently, data augmentation techniques (e.g., adding noise, time stretching, and pitch shifting) are applied to enrich the dataset, as illustrated in Fig. 2. These techniques are implemented as follows:
Adding noise To make the model more resistant to changes in the environment, we added additive Gaussian noise with a mean of zero and a signal-to-noise ratio that was evenly sampled between 15 and 30 dB (changing the pitch).
Time stretching To make more changes to the dataset, the playback speed of the audio signal is changed twice: first by speeding it up (rate = 1.1) and then by slowing it down (rate = 0.9).
Pitch Shifting The pitch of the signal is changed by ± 2 semitones, which makes the dataset more varied.
Justification for 3-second segmentation Respiratory cycles typically last 2.5–4 s15,36,37,38. Non-overlapping 3-second frames ensure at least one complete cycle per segment and preserve diagnostic patterns (crackles, wheezes). Ablation showed that 3 s provided optimal accuracy (98%) and generalization to variable-length datasets (ICBHI: 10–90 s, KAUH: 5–30 s) without retraining.
MFCC features were extracted using Librosa (v0.10.1) with n_mfcc = 13, 25 ms Hann window (100 samples), 10 ms hop (60% overlap), n_fft = 100, 40 Mel filters, including delta and delta-delta (39 features total), Min-Max normalized.
The impact of preprocessing on a normal lung sound is shown in Fig. 1, illustrating both time and frequency domain changes. In reality, the augmentations are performed more aggressively on the under-represented disease categories (asthma, pneumonia, BRON, URTI, and LRTI), thus their effective sample size becomes closer to that of the majority COPD class. The employed technique serves as a partial compensation for the inherent class imbalance of the ICBHI dataset.
Mel-frequency cepstral coefficients (MFCCs)
After augmentation, the improved audio signals are divided into smaller segments of 3 s. Feature extraction techniques, including the extraction of MFCCs, are subsequently employed to transform each segment into feature vectors. MFCCs are a common way to extract features from audio that mimics how people hear sounds. The extraction process includes the following steps38.
Pre-emphasis A pre-emphasis filter makes feature extraction more accurate by boosting higher frequencies and making up for sound loss during transmission.
Framing and windowing The signal is split into overlapping frames, each about 25 ms long, to pick up on local time patterns. A windowing function, like the Hamming window, is used on each frame to cut down on spectral leakage.
FFT and power spectrum The power spectrum shows how the energy is spread out over different frequencies. It is calculated after the FFT changes each frame into the frequency domain.
Mel Filter Bank Triangular filters constructed on the Mel scale, which represents the non-linear frequency sensitivity of the human ear, are used to filter the power spectrum.
Logarithm To simulate how the human auditory system reacts to sound intensity, a logarithmic transformation reduces the dynamic range of the filter outputs.
Discrete Cosine Transform (DCT) To decorrelate the features and provide a compact representation, DCT is applied to the log-transformed values. MFCCs are usually chosen from the first 12–13 coefficients, excluding the zeroth coefficient.
Delta and delta-delta coefficients The first and second derivatives of the MFCCs are used to calculate optional features that capture temporal dynamics.
MFCCs are useful for voice recognition and sound classification applications because they offer a reliable and compact representation of audio signals. The padding sequence ensures that all MFCC feature vectors are the same size to handle varying feature vector lengths resulting from different segment lengths. This padding step helps maintain uniformity across the dataset, enabling more efficient model training. The effect of preprocessing on a normal lung sound is illustrated in Fig. 2.
Original and preprocessed normal lung signal analysis: (a) Original time-domain waveform. (b) Original frequency spectrum (FFT). (c) Preprocessed time-domain waveform (post-resampling and augmentation). (d) Preprocessed frequency spectrum. (e) MFCC heatmap of a 3-second frame.
The complete LDSC training and inference procedure is outlined in Algorithm 1.
Convolutional neural network
CNNs represent a widely utilized architecture within deep learning, primarily adopted for classification tasks. CNNs get their name from the convolutional operation that is used on input data. They use convolutional layers to find hierarchical features in the input data, which gradually learn to recognize complex patterns. Then, predictions are made by passing the retrieved features through one or more fully linked layers. CNNs use convolution operations and subsampling in a few hidden layers to slowly pull-out complex features from the input data39. This multi-layered structure lets CNNs pick up on small details, which makes them great at recognizing complex images40.
1D-CNN is a special type of CNN that is made to work with sequential data that is arranged in only one spatial dimension. 1D-CNNs are better for analyzing linearly organized data, like time-series signals, audio waveforms, or text sequences. Two-dimensional CNNs, on the other hand, are better for image processing41.
Input layer Accepts a one-dimensional sequence, like a time-series signal that shows a vector of data points, like sounds from the lungs.
Convolutional layer A few filters in this layer look for patterns in nearby areas by following the input sequence. The stride sets the size of the steps the filter takes, and the filter size (also called kernel size) sets the length of the sequence segment that the filter covers. This layer makes a feature map for each pattern that has been found42. In a one-dimensional convolutional neural network (1D-CNN), the filter moves over the input sequence and multiplies each element by itself. The results are then added together to find patterns in the data. The formula for the output at position y[i] in a 1D convolutional layer is expressed as:
Here, x is the input sequence, w is the convolutional filter of size, b is the optional bias term added to each output (which may or may not be used depending on the model setup), i is the current position in the output sequence, and y[i] is the output at position i. The step size for each movement along the sequence is defined by moving the filter w along the input x with a given stride s. This convolution procedure creates a feature map that encodes identified patterns or features from the input sequences by calculating the output at each place according to the stride and kernel size parameters. One can use the following to find the output sequence length L:
where N represents the length of the input sequence, s is the stride value, and K is the kernel size. Padding may be applied to preserve the original length of the input sequence in the output. Adding zeros to the ends of the input sequence is called padding. This makes up for the fact that the convolution operation shortens the sequence. This method makes sure that the output length stays the same, which is especially important when the input and output sizes need to be the same3.
Pooling layer Usually, max pooling or average pooling is used to down-sample the feature maps. This keeps important features while lowering the dimensionality and computational load. Pooling also makes the model more stable when the input sequence changes slightly. Two prevalent types of pooling used in 1D -CNNs are max pooling and average pooling.
Max pooling In max pooling, the output at each position is determined by the maximum value within a specified window of the input sequence. The formula for max pooling at output position y[i] is expressed as:
Here, y[i] represents the output value in the feature map obtained through pooling.
Fully connected layers Following the convolutional and pooling layers, the feature maps are flattened and passed through fully connected (dense) layers, integrating the extracted features to produce the final predictions. The formula for the output y from the fully connected layer can be expressed as:
where x is the input vector to the fully connected layer, which is typically a flattened version of the feature maps from the previous layer. W is the weight matrix for the fully connected layer, where each element wij corresponds to the weight connecting the jth neuron in the layer to the ith input feature. b as the bias vector for the fully connected layer, where each element bj is the bias term for the jth neuron. y as the output vector from the fully connected layer.
Activation function A Rectified Linear Unit (ReLU) function often adds non-linearity to the model, which helps it learn more complex representations and makes it better at telling the difference between features. In this study, we utilize Leaky ReLU as the activation function.
Leaky ReLU activation function The Leaky ReLU activation function allows for a small, non-zero gradient for negative inputs, which keeps neurons active and able to learn even when the input values are negative. The Leaky ReLU function can be written down in math like this:
Where f(x) is the output of the Leaky ReLU function for input x, α is a small positive constant (typically set to a small value like 0.01) that determines the function’s slope for negative inputs.
The Softmax function converts a set of real numbers into a probability distribution. This function is handy in multi-class classification problems and transforms the neural network output into probabilities that sum to one. In this method, the probability of each element is obtained by calculating its exponential and dividing it by the sum of the exponentials of all aspects. Given the input n= [n1, n2,…,nk], the softmax function calculates the pi probabilities for each element using the following formula:
Here, ni represents the i-th element of the input vector n. The denominator in this expression represents the sum of the exponentiated values of all the elements in the input vector.
The proposed scheme steps
Figure 3 shows the whole LDSC classification workflow, which includes the input preprocessing pipeline, the two-layer LDSC architecture, the generation of feature maps after Conv1 and Conv2, and the final fully connected layers that lead to the softmax output for diagnosis. The LDSC architecture has two 1D convolutional layers, each of which is followed by a LeakyReLU activation function that captures and improves important features in the input signals.
The Adam optimizer is used by the model to automatically adjust the parameters, and the categorical cross-entropy (CCE) function43 is used as a loss function for multiclass classification tasks. The CCE function compares the predicted and actual class distributions and gives an accurate measure of how well the model works. The first 1D-CNN layer has 32 filters, each with a kernel size of 3 (F1 = 32, K1 = 3). This layer extracts temporal features from the input signals across various frequencies. The convolutional layer first takes in the main features from the data and makes feature maps. After that, a MaxPooling layer with a pool size of 2 (1 = 2) is used to lower the number of dimensions of the features and make the calculations easier.
Proposed methodology steps.
There is a convolution layer with 64 filters and a kernel size of 3 in the second layer. (F2 = 64, K2 = 3). This layer is meant to pick up on more complex and higher-level features in the data, which will create more feature maps. The LDSC architecture uses two 1D-convolutional layers (32 filters, kernel size 3; 64 filters, kernel size 3) and gets 13 MFCC coefficients. This is how audio signals for breathing are usually processed36,38 and in the 50–2000 Hz range. The authors iteratively designed and fine-tuned the LDSC architecture based on extensive ablation studies on respiratory cycle duration and MFCC dimensionality, making sure it was both clinically relevant and computationally efficient. These options strike a balance between computational efficiency and diagnostic performance, as shown in Sect. 4.4. After this convolutional operation, a separate MaxPooling layer with a pool size of 2 (2 = 2) is used to make the feature maps smaller and improve the speed of the calculations. The output of the second convolutional layer has a much smaller feature map size as a result.
Batch normalization is carried out following each convolutional layer. As a result, training is more effective, and the model remains stable. To prevent overfitting, dropout is applied at a rate of 0.3 after the second convolutional layer and before the flattening layer. The model has two additional hidden layers. The dropout rates for the first and second hidden layers are 0.3 and 0.2, respectively. These hidden layers aid in generalization and lessen overfitting by introducing controlled randomness during training.
The output is flattened and sent to the dense layer, which divides the signal into N classes, following processing of the final hidden layer. The value of N is equal to 7 and 8 for the ICBHI and KAUH datasets, respectively. The LDSC model was trained for 400 epochs using a batch size of 8 and a learning rate of 0.0001. The suggested architecture’s parameter settings are displayed in Table 4. Bias terms were excluded from all layers to improve the model’s efficiency and ease of use. Because LDSC combines frequency-specific spatial filters, batch normalization, dropout, and effective convolution operations, it is a helpful tool for analyzing lung sound signals from various datasets.
Performance evaluation
The proposed CNN model is built using TensorFlow and trained on a system featuring a 13th-generation Intel(R) Core (TM) i9-13980HX processor and 32GB of RAM. The time required to train the model in this setup is approximately two days. The performance of the proposed method is evaluated using KAUH and ICBHI datasets. This model is used for multi-label classification of lung diseases, and the softmax activation function is used in the final layer of LDSC architecture.
Evaluation metrics
This paper examines accuracy, sensitivity, and specificity to determine how well the classification algorithm performs. These procedures provide a comprehensive view of the model’s lung disease detection capabilities. Although accuracy provides a broad indication of the model’s performance, it can be deceptive when one class is significantly larger than the other. The dataset is balanced by using augmentation techniques to create additional samples for underrepresented classes. Sensitivity quantifies the classifier’s ability to identify positive samples, whereas specificity quantifies the proportion of real negatives—that is, correctly identified negative occurrences—to all negatives. A more complete understanding of the model’s efficacy across various categories can be obtained by evaluating sensitivity and specificity. These measures are often used in medical diagnostics to fully test classification systems16,17,18. The Receiver Operating Characteristic (ROC) curve is also reported to evaluate discriminative performance across thresholds, particularly in imbalanced datasets (Fig. 4a, b). Values close to 1 mean that it works well44,45.
Multi-class ROC diagram: (a) KAUH dataset, (b) ICBHI dataset.
The evaluation metrics, derived from the confusion matrix, which provides a comprehensive representation of model performance, are presented below in Eq. (7), Eq. (8), and Eq. (9)15.
When lung sounds are classified as normal or abnormal, accurate classifications are represented by true positives (TP) and true negatives (TN). On the other hand, incorrect classifications such as misidentifying normal sounds as abnormal or abnormal sounds as expected or mistakenly categorizing one type of abnormality as another are denoted by false positives (FP) and false negatives (FN).
Experimental results
The binary classification results are shown in Table 4, where the normal class is compared with each disease class separately. In this table, Acc represents accuracy, represents specificity, and represents sensitivity. This research demonstrates higher accuracy in predicting standard signals than abnormal ones, achieving the highest accuracy in detecting asthma. The proposed model shows good performance in binary and multi-class classification tasks. Unlike most state-of-the-art methods that report results for a limited set of classes, this study considers all diseases to provide a more comprehensive assessment. For the ICBHI dataset, the accuracy, sensitivity, and specificity are 0.98, 0.98, and 0.99, respectively. For the KAUH dataset, the accuracy, specificity, and sensitivity values are all 99%.
Figures 5 and 6 illustrate the multi-label classification confusion matrices for the two datasets. As shown in Fig. 5, the classification method proposed by us is generally accurate on the KAUH dataset, except for a few cases. In particular, the noise of COPD has been misclassified only once as asthma, yet the instances of pneumonia have been misclassified twice as COPD, and just one BRON sound has been wrongly recognized as originating from pneumonia. Moreover, it was found that two heart failure signals were incorrectly tagged as BRON, and a single lung fibrosis signal was mislabeled as pleural effusion. The remaining signals are correctly classified.
The confusion matrix for the ICBHI dataset, where several misclassifications are observed, is presented in Fig. 6. The large number of true positives for the COPD in Fig. 6 is a good indication of the main source of the underlying label distribution of the ICBHI dataset as well as our 3-second segmentation strategy: the recordings of COPD are much more than other pathologies, and each recording produces many frame-level samples. In spite of this imbalance, the confusion matrix, together with the reported sensitivity, specificity, and ROC curves, shows that the proposed LDSC model still has a strong performance in all disease categories and is not overshadowed by the COPD majority. Two signals from the LRTI class are wrongly identified as usual, one COPD signal is misclassified as asthma, 15 pneumonia signals are misclassified as COPD, five URTI signals are misclassified as COPD, and 12 LRTI signals are misclassified as COPD. Additionally, one BRON signal has been misclassified as a pneumonia signal and five URTI signals have been mislabeled as BRON. Except for these misclassifications, all other signals have been correctly identified.
Confusion matrix for multi-classification of the ICBHI dataset.
Confusion matrix for multi-classification of the KAUH dataset.
Addressing the research questions
The LDSC model is evaluated using standard classification metrics (Accuracy, Sensitivity, Specificity). In addition, we employ 95% confidence intervals (via 1,000 bootstrap resamples), ablation studies (to assess the impact of MFCC and data augmentation), and cross-dataset validation to ensure a comprehensive and robust assessment. Using these metrics and analyses, we address the three research questions as follows:
RQ1 (Lightweight Architecture):
The LDSC model, which has only two 1D convolutional layers, got 98% accuracy, sensitivity, and specificity on the ICBHI dataset and 99% across all metrics on the KAUH dataset. These results either exceed or match the performance of complex hybrid models (CNN-LSTM15, EasyNet22 while utilizing significantly fewer layers and parameters, thereby confirming that architectural simplicity is effective for high-performance lung sound categorization.
RQ2 (MFCC + Augmentation Effectiveness):
The lightweight LDSC model achieves 98% accuracy on ICBHI and 99% accuracy on KAUH when MFCC feature extraction is combined with audio augmentation techniques like noise injection, time-stretching, and pitch-shifting. According to ablation experiments, accuracy is reduced by 6–8% when augmentation is not used, and performance is reduced by roughly 12% when raw waveforms are converted to MFCC. This pipeline provides LDSC with robust, perceptually based, and duration-invariant inputs, enabling it to outperform sophisticated state-of-the-art techniques in terms of computational efficiency and cross-dataset resilience.
RQ3 (Computational Efficiency):
The LDSC model has a simple design with just two 1D convolutional layers. This makes it much less complicated to compute than hybrid state-of-the-art models like CNN-LSTM. It takes about 48 h to train on a mid-range CPU (Intel i9-13980HX, 32GB RAM) without using a GPU. The time complexity of LDSC training and inference per sample is \(\:O(n\cdot\:l\cdot\:{f}_{1}\cdot\:{k}_{1}+n\cdot\:{l}^{{\prime\:}}\cdot\:{f}_{2}\cdot\:{k}_{2})\), where \(\:n\)represents the number of 3-second frames, \(\:l\)is the MFCC feature dimension per frame, \(\:{l}^{{\prime\:}}\)is the reduced length after first max-pooling, \(\:{f}_{1}=32\), \(\:{f}_{2}=64\), and \(\:{k}_{1}={k}_{2}=3\). This near-linear scaling with input length ensures efficient real-time processing on resource-constrained medical devices.
The simplified design, which has fewer layers, batch normalization, and no bias terms, makes it less memory-intensive, speeds up training, and makes it possible to use on devices with limited resources, like digital stethoscopes or mobile health apps, all while keeping 98–99% diagnostic accuracy.
Performance metrics with 95% confidence intervals
To enhance statistical validity, Table 5 presents Accuracy, Sensitivity, and Specificity accompanied by 95% confidence intervals (CIs) obtained from 1000 bootstrap resamples, as well as Eqs. (5), (7), and (8) in Sect. 4.1.
The narrow CIs (< 1%) show that LDSC works very well on both datasets, with minimum variation because of random splits or small samples15.
Model interpretability and Grad-CAM visualization
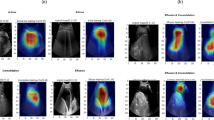
We used Grad-CAM46 on the last convolutional layer of LDSC to check the learned features. When you look at class-specific heatmaps on MFCC inputs, you can see that the model is focused on discriminative temporal-spectral patterns, like high-energy transients in crackles and sustained low-frequency bands in wheezes. These clinically aligned activations validate the model’s dependence on significant acoustic cues rather than artifacts, thereby augmenting diagnostic credibility. The proposed method has been widely evaluated to detect all disease classes in both datasets. In Fig. 7, the accuracy of the LDSC model is compared with the existing methods in the KAUH and ICBHI datasets. Also, in Fig. 9 for the ICBHI dataset, the sensitivity and specificity of the LDSC model are compared with other state-of-the-art techniques.
Comparison and discussion
Table 6 shows how LDSC stacks up against more recent SOTA methods on the ICBHI and KAUH datasets. LDSC has a much simpler structure (2 conv. layers) but gets the same or better accuracy (98–99%). LDSC has several advantages, including being non-complex, cost-effective to run, compatible with MFCC + augmentation, and well-suited for edge devices. LDSC achieved 98.1% accuracy on KAUH (4-class). Detailed class-wise performance is reported in Table 7. Cross-dataset generalization shows 97.8% accuracy (ICBHI → KAUH). Results are presented in Table 8.
In47, the authors presented a sophisticated deep-learning architecture for the classification of lung sounds to identify respiratory ailments. This method employs dual time-frequency transformations, specifically continuous wavelet transforms (CWT) and Mel spectrograms, to generate intricate representations of pulmonary sounds. Essential characteristics were derived from these representations via parallel convolutional autoencoders and consolidated into a hybrid feature repository. A long short-term memory (LSTM) network was used to process the features even more so that they could be used to classify illnesses into multiple classes. The model did an amazing job, getting 94.16% accuracy in eight-class classification, 79.61% accuracy in four-class classification, and 85.61% accuracy in binary classification tasks. It was more accurate, sensitive, specific, and had a higher F1-score than other methods.
Naoki Asatani and colleagues2 proposed an automated categorization method employing deep learning to facilitate the diagnosis of respiratory diseases through auscultation. The methodology consists of two phases: first, generating a spectrogram of respiratory sounds using a short-time Fourier transform (STFT), and second, classifying the spectrogram into normal and abnormal (crackle, wheeze, or both) sounds with the help of an advanced CRNN. We improved the CRNN model by changing the size of the convolution filter to make it better at finding frequency changes in strange sounds, adding a bidirectional LSTM to get temporal data, and adding a 1D GAP layer to stop overfitting. The evaluation’s findings revealed an average score of 0.73, a harmonic score of 0.72, a sensitivity of 0.63, and a specificity of 0.83. This indicates that the new model outperforms the previous CNN, RNN, and CRNN models in terms of classification. By analyzing lung sounds to identify respiratory issues, this approach greatly aids physicians. Arpan Srivastava et al.48 developed a CNN-based deep learning model to aid healthcare professionals in diagnosing Chronic Obstructive Pulmonary Disease (COPD) using respiratory sound data. Features such as MFCC, Mel-Spectrogram, and various Chroma properties are extracted from the Librosa machine learning library. MFCC features have the highest accuracy, according to experimental results, with a 93% classification rate. By listening to breathing sounds, the technique can detect heart attacks and asthma, as well as gauge the severity of a disease. To improve clinical efficacy, future research may include data augmentation techniques, strengthen privacy protections, and integrate the system with respiratory monitoring technologies. To increase the precision of diagnosing respiratory diseases, the authors created a novel method for categorizing lung sounds (R).
To improve classification performance, this study introduced a pre-trained CNN architecture that simultaneously integrates average-pooling and max-pooling layers. A Linear Discriminant Analysis (LDA) classifier that employed the Random Subspace Ensembles (RSE) technique was fed deep features from CNN. As a result, the accuracy was 5.75% higher than that of other approaches using the ICBHI 2017 dataset. The CNN model outperformed other pre-trained CNN models, such as VGG16 and AlexNet, which weren’t designed for audio data, because it was trained on spectrogram images of lung sounds. The best classification accuracy of 71.15% was obtained by combining parallel pooling with LDA-RSE, indicating the efficacy of the suggested approach in classifying lung sounds. Deep learning models for detecting respiratory diseases from raw lung auscultation sounds were assessed by Ali Mohammad Alqudah et al.15. Using two datasets to create four subsets, the study evaluated three distinct deep learning models on both augmented and non-augmented datasets.
All models performed well, but the CNN-LSTM model consistently outperformed the others on every dataset. On non-augmented datasets, the CNN-LSTM model achieved 99.6%, 99.8%, 82.4%, and 99.4% accuracy; on augmented datasets, it achieved 100%, 99.8%, 98.0%, and 99.5% accuracy. The hybrid CNN-LSTM approach outperformed models using a single technique, and the augmentation process greatly enhanced model performance, demonstrating the effectiveness of CNN for feature extraction and LSTM for classification tasks in respiratory disease diagnosis. have proposed an automated method that uses lung sound (LS) signals from the KAUH dataset to diagnose lung diseases.
This approach analyzes LS signal modes using an empirical wavelet transform with predetermined boundary points. Important characteristics, like peak amplitude and peak frequency in the frequency domain and Shannon entropy in the time domain, are obtained from each mode. Different classifiers, such as Support Vector Machine, Random Forest, Heavy Gradient Boosting, and Light Gradient Boosting Machine (LGBM), are used to diagnose diseases. Performance evaluations using LS signals from the KAUH dataset and a public database revealed that various classification techniques could identify 80.35%, 83.27%, 99.34%, and 77.13% of the time. Compared to existing LS signal-based disease detection techniques, this system was able to accurately identify normal, pneumonia, and asthma sounds 84.76% of the time. Although numerous studies have demonstrated the accuracy of complex hybrid models, the LDSC model stands out due to its simplicity and effectiveness.
In terms of accuracy, sensitivity, and specificity, the LDSC model continuously performs better than the best approaches. This achievement emphasizes the significance of preprocessing methods and model architecture created especially for lung sound inputs. To obtain the most crucial features, the LDSC architecture’s kernel functions are meticulously adjusted. Examples of data augmentation techniques that enhance the dataset and increase the stability and generalizability of the model include adding noise, shifting steps, and stretching time. Additionally, its processing efficiency is greatly enhanced by the simplicity of its design. Compared to more complex hybrid models, which require longer training times and larger datasets, it ensures that the training and evaluation process is quicker. The choice of 1D-CNN, which is made for processing temporal data, makes it easier to get features without adding unnecessary complexity, making it perfect for analyzing respiratory signals.
Accuracy assessment of LDSC and state-of-the-art methods on the ICBHI dataset.
Accuracy assessment of LDSC and state-of-the-art methods on the KAUH dataset.
Comparison of specificity and sensitivity obtained on the ICBHI dataset.
Ablation study and computational efficiency
Ablation study revealed that removing MFCC features and data augmentation reduced accuracy by 11.7% and 5.9%, respectively, while eliminating the second convolutional layer caused only a 4.6% drop. The lightweight LDSC-Lite variant, with ~ 10 K parameters and 91.5% accuracy, is suitable for deployment on resource-constrained medical devices. Ablation on frame length (ICBHI multi-class) showed that 3-second frames outperform 1-second (93.8% accuracy) and 5-second (98.3%) durations, achieving 98.0% accuracy with optimal data diversity and generalization across datasets.
Limitations of LDSC
Testing on two datasets requires validation on larger, diverse populations. LDSC is 98–99% accurate, but it only works with small datasets, class imbalance, and a lack of demographic diversity (few pediatric/elderly cases, regional bias). This makes it harder to apply what we’ve learned to real-world clinical settings with different populations, noise levels, and comorbidities. Testing across datasets is helpful, but for reliable bedside use, large-scale, balanced, multi-center validation is necessary.
Conclusion
This study introduces the LDSC model, a streamlined 1D-CNN consisting of only two convolutional layers, enabling the early detection of lung diseases and swift intervention. Simplicity reduces the time, hardware, and computing costs needed for training while still keeping the integrity of the raw waveforms. This means that there is no need to convert the spectrogram through preprocessing with augmentation (noise, time-stretching, pitch-shifting), 4 kHz resampling, and 3-second segmentation. The LDSC model got 98% accuracy, sensitivity, and specificity on the ICBHI dataset and 99% across all metrics on the KAUH dataset. These results are better than those of complex hybrids15,22 with few parameters. With low-cost digital stethoscopes, LDSC can quickly and non-invasively diagnose lung disease at the point of care, with 98–99% accuracy on raw auscultation sounds. Cross-dataset validation and ablation studies demonstrate greater robustness compared to complex models. This lightweight 1D-CNN cuts diagnostic delays, reduces imaging needs, and enables telemedicine in resource-poor settings for real-time clinical support.
In the future, smartwatches could record lung sounds and send them to doctors so they can make a diagnosis faster. Future directions include medical IoT integration and digital twin applications. Limitations involve dataset diversity and class imbalance, requiring larger, balanced cohorts.
Data availability
The datasets used in this study are publicly available:- ICBHI 2017 Respiratory Sound Database: - KAUH Lung Sound Dataset: https://www.kaggle.com/datasets/arashnic/lung-dataset both containing raw annotated audio files with cycle-level labels.
References
Jabbar, R., Zai, S., Jawaid, M. & Ahsan, M. Investigating Deep Learning Methods for Detecting Lung Adenocarcinoma on the TCIA Dataset (2023).
Asatani, N., Kamiya, T., Mabu, S. & Kido, S. Classification of respiratory sounds using improved convolutional recurrent neural network. Comput. Electr. Eng. 94, 107367 (2021).
Borwankar, S., Verma, J. P., Jain, R. & Nayyar, A. Improvise approach for respiratory pathologies classification with multilayer convolutional neural networks. Multimedia Tools Appl. 81(27), 39185–39205 (2022).
Rajasekar, V., Vaishnnave, M., Premkumar, S., Sarveshwaran, V. & Rangaraaj, V. Lung cancer disease prediction with CT scan and histopathological images feature analysis using deep learning techniques. Results Eng. 18, 101111 (2023).
Dastider, A. G., Sadik, F. & Fattah, S. A. An integrated autoencoder-based hybrid CNN-LSTM model for COVID-19 severity prediction from lung ultrasound. Comput. Biol. Med. 132, 104296 (2021).
Rayan, A. et al. Utilizing CNN-LSTM techniques for the enhancement of medical systems. Alexandria Eng. J. 72, 323–338 (2023).
Wankhade, S. & Vigneshwari, S. A novel hybrid deep learning method for early detection of lung cancer using neural networks. Healthc. Analytics. 3, 100195 (2023).
Naeem, H. & Bin-Salem, A. A. A CNN-LSTM network with multi-level feature extraction-based approach for automated detection of coronavirus from CT scan and X-ray images. Appl. Soft Comput. 113, 107918 (2021).
Islam, M. Z., Islam, M. M. & Asraf, A. A combined deep CNN-LSTM network for the detection of novel coronavirus (COVID-19) using X-ray images. Inf. Med. Unlocked. 20, 100412 (2020).
Jasmine Pemeena, M. et al. Lung diseases detection using various deep learning algorithms. J. Healthc. Eng. 2023, 3563696 (2023).
Fachrel, J., Pravitasari, A. A., Yulita, I. N., Ardhisasmita, M. N. & Indrayatna, F. Enhancing an imbalanced lung disease x-ray image classification with the CNN-LSTM model. Appl. Sci. 13(14), 8227 (2023).
Saeed, T. et al. An AI-Enabled bias-free respiratory disease diagnosis model using cough audio. Bioengineering 11(1), 55 (2024).
Li, L. et al. Prediction and diagnosis of respiratory disease by combining convolutional neural network and bi-directional long short-term memory methods. Front. public. Health. 10, 881234 (2022).
Kanipriya, M., Hemalatha, C., Sridevi, N., SriVidhya, S. & Shabu, S. J. An improved capuchin search algorithm optimized hybrid CNN-LSTM architecture for malignant lung nodule detection. Biomed. Signal Process. Control. 78, 103973 (2022).
Alqudah, A. M., Qazan, S. & Obeidat, Y. M. Deep learning models for detecting respiratory pathologies from Raw lung auscultation sounds. Soft. Comput. 26(24), 13405 (2022).
Petmezas, G. et al. Automated lung sound classification using a hybrid CNN-LSTM network and focal loss function. Sensors 22(3), 1232 (2022).
Fraiwan, M., Fraiwan, L., Alkhodari, M. & Hassanin, O. Recognition of pulmonary diseases from lung sounds using convolutional neural networks and long short-term memory. J. Ambient Intell. Humaniz. Comput. 13(10), 4759–4771 (2022).
Goyal, S. & Singh, R. Detection and classification of lung diseases for pneumonia and Covid-19 using machine and deep learning techniques. J. Ambient Intell. Humaniz. Comput. 14(4), 3239–3259 (2023).
Indumathi, V. & Siva, R. Improving early detection of lung disorders: a multi-head self-attention CNN-BiLSTM model. J. Institution Eng. (India): Ser. B. 105(3), 595–607 (2024).
Xu, Y. et al. Deep learning predicts lung cancer treatment response from serial medical imaging. Clin. Cancer Res. 25(11), 3266–3275 (2019).
Polat, H. & Danaei Mehr, H. Classification of pulmonary CT images by using hybrid 3D-deep convolutional neural network architecture. Appl. Sci. 9(5), 940 (2019).
Hassan, U., Singhal, A. & Chaudhary, P. Lung disease detection using EasyNet. Biomed. Signal Process. Control. 91, 105944 (2024).
Yan, F., Huang, X., Yao, Y., Lu, M. & Li, M. Combining Lstm and densenet for automatic annotation and classification of chest x-ray images. IEEE Access. 7, 74181–74189 (2019).
Ker, J., Wang, L., Rao, J. & Lim, T. Deep learning applications in medical image analysis. Ieee Access. 6, 9375–9389 (2017).
Alshmrani, G. M. M., Ni, Q., Jiang, R., Pervaiz, H. & Elshennawy, N. M. A deep learning architecture for multi-class lung diseases classification using chest X-ray (CXR) images. Alexandria Eng. J. 64, 923–935 (2023).
Reddy, N. S. & Khanaa, V. Intelligent deep learning algorithm for lung cancer detection and classification. Bull. Electr. Eng. Inf. 12(3), 1747–1754 (2023).
Desai, V. & Thengade, A. Chest abnormality detection from x-rays using deep learning. in Second International Conference on Next Generation Intelligent Systems (ICNGIS) 1–6 (IEEE, 2022).
Abbas, Q. Lung-deep: a computerized tool for detection of lung nodule patterns using deep learning algorithms detection of lung nodules patterns. Int. J. Adv. Comput. Sci. Appl. 8, 10 (2017).
Aslan, Z. On the use of deep learning methods on medical images. Int. J. Energy Eng. Sci. 3(2), 1–15 (2019).
Mei, X. et al. Interstitial lung disease diagnosis and prognosis using an AI system integrating longitudinal data. Nat. Commun. 14(1), 2272 (2023).
Weiss, J. et al. Deep learning to estimate lung disease mortality from chest radiographs. Nat. Commun. 14(1), 2797 (2023).
Patel, R. K., Choudhary, A., Kumari, N. & Lamkuche, H. S. Pneumonia screening from radiology images using homomorphic transformation Filter-Based FAWT and customized VGG‐16. Int. J. Imaging Syst. Technol. 35(3), e70093 (2025).
Kumar, D., Chouhan, S. S., Patel, R. K., Viswakarma, H. & Mishra, V. Transformative Insights: Gabor Features and Tensor-EWT for COVID Diagnosis in Lung CT Images, in IEEE 2nd International Conference on Innovations in High Speed Communication and Signal Processing (IHCSP) 1–4 (IEEE, 2024).
Patel, R. K. & Kashyap, M. Automated diagnosis of COVID stages using texture-based Gabor features in variational mode decomposition from CT images. Int. J. Imaging Syst. Technol. 33(3), 807–821 (2023).
Kim, Y., Camacho, D. & Choi, C. Real-time multi-class classification of respiratory diseases through dimensional data combinations. Cogn. Comput. 16(2), 776–787 (2024).
Chua, Y. W. & Cheng, Y. C. Towards enhanced classification of abnormal lung sound in multi-breath: A light weight multi-label and multi-head attention classification method, arXiv preprint arXiv:2407.10828 (2024).
Fraiwan, M., Fraiwan, L., Khassawneh, B. & Ibnian, A. A dataset of lung sounds recorded from the chest wall using an electronic stethoscope. Data Brief. 35, 106913 (2021).
Wanasinghe, T. et al. Lung sound classification with multi-feature integration utilizing lightweight CNN model. IEEE Access. 12, 21262–21276 (2024).
Wall, C., Zhang, L., Yu, Y., Kumar, A. & Gao, R. A deep ensemble neural network with attention mechanisms for lung abnormality classification using audio inputs. Sensors 22(15), 5566 (2022).
Koshta, V. & Singh, B. K. A two-level classifier for prediction of healthy and unhealthy lung sounds using machine learning and convolutional neural network. in International Conference on Biomedical Engineering Science and Technology 154–168 (Springer, 2023).
Lella, K. K., Jagadeesh, M. & Alphonse, P. Artificial intelligence-based framework to identify the abnormalities in the COVID-19 disease and other common respiratory diseases from digital stethoscope data using deep CNN. Health Inform. Sci. Syst. 12(1), 22 (2024).
Neili, Z. & Sundaraj, K. A comparative study of the spectrogram, scalogram, melspectrogram and gammatonegram time-frequency representations for the classification of lung sounds using the ICBHI database based on CNNs. Biomedical Engineering/Biomedizinische Technik. 67(5), 367–390 (2022).
Sundas, A., Singh Shahi, G. & Badotra, S. Streamlined patient care with smart monitoring and deep learning-based recommendations using categorical cross entropy optimization. in Proceedings of the KILBY 100 7th International Conference on Computing Sciences (2023).
Altan, G., Kutlu, Y. & Allahverdi, N. Deep learning on computerized analysis of chronic obstructive pulmonary disease. IEEE J. Biomedical Health Inf. 24(5), 1344–1350 (2019).
Altan, G. DeepOCT: an explainable deep learning architecture to analyze macular edema on OCT images. Eng. Sci. Technol. Int. J. 34, 101091 (2022).
Makade, J., Bankar, N., Kumar, A., Bandre, G. & Yenurkar, G. Artificial intelligence in health care: A review of uses, challenges and potential uses. in AIP Conference Proceedings Vol. 3188, 080032 (AIP Publishing LLC, 2024).
Khan, R., Khan, S. U., Saeed, U. & Koo, I. S. Auscultation-based pulmonary disease detection through parallel transformation and deep learning. Bioengineering 11(6), 586 (2024).
Srivastava, A. et al. Deep learning based respiratory sound analysis for detection of chronic obstructive pulmonary disease. PeerJ Comput. Sci. 7, e369 (2021).
Demir, F., Ismael, A. M. & Sengür, A. Classification of lung sounds with CNN model using parallel pooling structure. IEEE Access 8, 105376–105383 (2020).
Tripathy, R. K., Dash, S., Rath, A., Panda, G. & Pachori, R. B. Automated detection of pulmonary diseases from lung sound signals using fixed-boundary-based empirical wavelet transform. IEEE Sens. Lett. 6(5), 1–4 (2022).
Author information
Authors and Affiliations
Contributions
“ M.A. and F.I. wrote the main manuscript text and its evaluation. “A.G. and A.S.” edited and supervised the manuscript. All authors reviewed the manuscript.”
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
All methods were carried out in accordance with relevant guidelines and regulations. The experimental protocols were approved by the Institutional Review Board (IRB) of King Abdullah University Hospital. Informed consent was obtained from all subjects and/or their legal guardian(s) at the time of data collection by the dataset provider.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Abbasi, M., Imani, F., Ghaffari, A. et al. LDSC: enhancing lung disease diagnosis using a simple 1D-CNN. Sci Rep 16, 4298 (2026). https://doi.org/10.1038/s41598-025-34485-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-34485-3