Abstract
The diagnosis of leukemia is based on the simultaneous analysis of morphological patterns of hematological images and the presence of clinical indicators in written reports. Majority of machine learning models are unimodal and centralized. They are not able to integrate information with the institutions or give clinically useful explanations. This paper suggests a federated multimodal architecture that integrates Vision Transformers (ViT) and ClinicalBERT to encode images and classify texts to conduct joint leukemia diagnosis and staging in decentralized medical devices, respectively. Both modalities are synthesised into a single semantic space to form a cross-modal fusion layer, and binary diagnosis and multiclass staging are facilitated by dual output heads. The framework uses federated learning protocol which maintains the privacy of data by the fact that the local data does not move out of institutional boundaries. To improve the level of transparency, SHAP-based explanations are provided on each prediction, where both visual regions and clinical tokens are considered important. The results of the experiments indicate that the suggested system is more accurate and has a higher F1-score than unimodal and centralized baselines and also has interpretable and patient-specific explanation, which is consistent with clinical expectations. The architecture is robust in the non-IID data distributions and is scaled through simulated healthcare networks, which makes it appropriate to deploy to actual health care in diagnostic oncology.
Similar content being viewed by others
Introduction
Leukemia is also a rather complicated type of cancer in terms of diagnosis, where the identification of abnormal hematological patterns and the thorough analysis of the clinical record and laboratory reports is needed1. The conventional process of diagnostic activities is based on the human eye analysis of blood smears and the semantic meaning of patient records, such as the number of white blood cells, the percentage of blast, and the early symptom pattern2. Morphological evidence combined with structured and unstructured clinical data is necessary to draw accurate diagnoses and stage3. Nevertheless, in reality, these data modalities tend to be handled separately, which restricts the possibility of deriving composite information that reflects the cognitive problem-solving of the hematologists4.
The modern developments in the field of machine learning have enabled the ability to automatize the elements of the diagnostic process with the help of image-based convolutional model or text-based transformers5. However, such methods are rather independent, either revealing leukemic manifestations in smear images or disease indicators in free-text reports without developing a comprehensive interpretation between modalities6. More to the point, these models are usually trained on centralized data, which not only endangers the privacy of patients7 but also does not allow reflecting the institutional heterogeneity of data collection and diagnostic standards8. It is clear that there is a requirement of smart systems that uphold privacy, clinically challenging capture, and provide transparent support in decision systems9.
Besides the technical disconnectivity, there is an increasing interest in the area of the interpretability of AI-based systems of medical decision-making10. Clinicians do not have faith in black-box models that are unable to explain their findings by visible cell abnormalities or pattern of written symptoms11. Although predictive performance should be high, it should not be good enough to be deployed in high-risk environments like oncology12. They should not only be correct, but also provide explanations, privacy-conformant, and adaptable to decentralized healthcare ecosystems13. The research study addresses the absence of a privacy-sensitive and explainable multimodal system with the capability to diagnose leukemia and simultaneously determine its clinical stage based on the information of various healthcare facilities14. Current solutions do not adequately support image-text fusion, ensure local data ownership, or offer necessary post hoc interpretability for clinician trust15.
Several research efforts have attempted to address isolated facets of this challenge16. Image-based models17 using convolutional neural networks and more recently ViTs have shown encouraging results in detecting leukemic cells based on visual cues such as nuclear size and blast morphology18. Similarly, textual encoders like BERT and its clinical variants have been employed to parse physician notes and laboratory findings to infer disease presence19. In some cases, fusion models have been proposed to concatenate image and text embeddings into a unified vector space. However, these solutions often assume data centralization and overlook the practical constraints of hospital data silos. Additionally, fusion is typically shallow, and interpretability mechanisms are often either absent or too general to provide patient-level explanations20. A parallel line of work in federated learning offers promising strategies for decentralized model training across multiple clients without exchanging raw data21. While these methods address the privacy concern, most federated learning models in healthcare are unimodal and do not explore the rich interaction between vision and language inputs. Furthermore, existing explainable AI techniques in federated settings are mostly limited to global feature importance and do not provide instance-specific interpretations aligned with clinical reasoning22. Thus, a comprehensive solution that integrates multimodality, federated computation, and explainability remains an open and underexplored area23.
This research proposes a federated multimodal learning architecture for leukemia detection and staging that integrates vision transformers for image encoding and ClinicalBERT for textual understanding. The model incorporates a cross-modal fusion layer to synthesize both modalities and is trained using a privacy-preserving federated protocol across multiple simulated healthcare clients. To enhance interpretability, SHAP-based local explanations are computed for both image patches and text tokens, providing clinicians with actionable insight into each prediction. Unlike existing models, this framework operates under realistic non-IID conditions, offers dual-head classification for stage prediction, and maintains explainability at the individual patient level. The aim of this study is to develop and evaluate a privacy-preserving, interpretable, and multimodal deep learning framework for automated leukemia diagnosis and staging under federated conditions. The proposed architecture represents a innovative contribution that again fits the current privacy-sensitive clinical AI requirements to the best of our knowledge, as none of the previous works combines ViT- and ClinicalBERT-based multimodal fusion and SHAP-assisted interpretability in a federated environment.
The importance of this study is that it can help bridge an important gap between high-performing AI systems and clinically deployable solutions for hematological oncology. By bringing together the benefits of decentralized computation with modality-aware interpretability, the proposed framework accounts for practical issues in hospital deployment including data governance, diagnostic transparency and regional bias. Furthermore, it will help further the new research field of explainable federated learning, which can be used to provide a blueprint of secure and interpretable AI in other clinical areas. The framework can be extended to other related diagnostic areas where multimodal evidence and justification at the patient-level are crucial such as cardiology, pathology, and radiology. Main contributions of this research are:
-
Federated multimodal learning architecture: Integrates Vision Transformers (ViT) for image encoding and ClinicalBERT for processing clinical text, ensuring privacy and data decentralization.
-
Cross-modal fusion mechanism: Combines image and text modalities to form a unified representation for both leukemia detection and staging.
-
SHAP-based interpretability: Provides patient-specific explanations for both visual and textual inputs, enhancing transparency and trust in AI predictions.
-
Real-world applicability: Demonstrates robust performance under non-IID data distributions and scalability across simulated healthcare networks.
The remaining part of this paper is laid out as follows. The literature review is given in detail in Sect. 2, the proposed methodology is outlined in Sect. 3, the experimental setup and results are discussed in Sect. 4 and finally the paper is concluded with the future research directions.
Literature review
Federated learning (FL) has emerged as a viable approach to decentralized model training in healthcare, allowing institutions to collaboratively develop machine learning models without sharing sensitive patient data24. FL has already been successfully used in the medical imaging field, e.g., in tumor segmentation, diabetic retinopathy classification, and COVID-19 detection, and its frameworks are able to support non-IID data distributions and institution heterogeneity25. Parekh et al.26 proved that federated learning is practicable in cross-domain scenarios by modeling cross-domain training simulation in hospitals with different data distributions. Despite these developments, the vast majority of FL applications in medicine have been unimodal and tend to analyze image data only. Minimal research has been done on how to unify heterogeneous modalities e.g. clinical text and visual diagnostics into a single framework. Peng et al.27 also tried to solve the modality heterogeneity in computational pathology, on a multi-modal federated architecture, but their system was limited to structured imaging and tabular data. The combination of complex unstructured clinical narratives, e.g. physician notes or diagnostic summaries, is also under-researched in the federated setting.
Multimodal learning Multimodal learning in particular has been an expanding field of computational medicine motivated by the finding that using multiple modalities of data improves diagnostic accuracy and strength28. Combining radiological scans and patient records, especially in oncology, has been demonstrated to be successful in increasing both predictive power and interpretability. Research articles such as Thrasher et al.29 review multimodal learning in the healthcare field extensively, and the authors acknowledge the advantages of using electronic health records in conjunction with imaging data. Nevertheless, these methods usually presuppose centralized access to multimodal inputs, which is infeasible in practice in the real clinical system limited by privacy legislations and facility fragmentation30. Even in the context of fusion architecture, they often use naive concatenation of embeddings or even use joint encoders that are not easy to interpret and are computationally inefficient31. Moreover, not many studies provide posthoc justification of predictions in modality specific like that which is important in making transparent decisions in medicine32.
The emergence of explainable artificial intelligence (XAI) has sought to bridge the gap between black-box models and clinician trust. A SHapley Additive explanation (SHAP)33 has become a widely adopted method for attributing prediction importance to input features by leveraging cooperative game theory. SHAP has been applied to various medical domains such as breast cancer risk prediction34 and Parkinson’s disease diagnosis, where model transparency is vital for adoption. However, most XAI implementations in healthcare are confined to tabular or image data, and few extend to text, let al.one to simultaneous multimodal interpretation. Moreover, SHAP’s application within federated environments poses both computational and architectural challenges, particularly when trying to ensure consistency and privacy during explanation generation35. There is currently no established methodology for aligning SHAP interpretations across modalities within decentralized, privacy-sensitive settings36.
In the context of leukemia detection, deep learning has been applied with considerable success using high-resolution microscopy images37. CNNs have long been the dominant architecture for image-based diagnosis, but more recent studies have introduced ViTs due to their superior performance in capturing global visual dependencies. Cho et al.38 evaluated ViT models for classifying acute lymphoblastic leukemia (ALL) and found them to outperform traditional CNNs in capturing nuclear irregularities and blast distributions. Nevertheless, these models were trained on centralized datasets and did not incorporate associated clinical information that typically accompanies diagnostic workflows38. As a result, their practical value in real-world settings is limited, especially in complex cases where morphological evidence alone is inconclusive39. Furthermore, these image-only models provide limited interpretability beyond pixel-level heatmaps, which are insufficient for comprehensive diagnostic explanations.
The most recent developments in computational pathology have seen the introduction of highly effective representation learning models and diagnostic models which are vastly superior to prior models in making predictions on complex histopathological problems. Quan et al. (2024) also illustrated how Global Contrast-Masked Autoencoders can be trained to learn very discriminative and robust features of pathology using a self-supervised approach that is morphologically sensitive and that their Dual-Channel Prototype Network can also solve the few-shot classification problem by inferring scarce pathology samples through prototype-based reasoning40. Likewise, Wang et al. suggested a pyramid-based self-supervised strategy to grasp multi-scale pathologic signs, which, in turn, allows defining the tissue heterogeneity more properly41. In addition to representation learning, Nan et al. proposed a deep learning system that can measure visual patterns of pathologists on a whole-slide image, which is an indication of the possibility of AI systems imitating and formalizing expert interpretive behavior42. Other contributions are the use of capsule networks in medical imaging, like DenseCapsNet to detect COVID-19 in chest X-rays that showed better spatial relationship modeling in tasks of diagnosing diseases43. Reinforcement learning has become another useful paradigm as demonstrated by Zheng et al. who have designed an efficient melanoma diagnosis strategy using deep RL through the learning of optimal region-of-interest selection policies44. Together, these studies have highlighted the fast advancement of pathology-driven deep learning, with the essence of strong feature learning, effective data use, and clinically consistent diagnostic logic, which are highly encouraging and supportive of the multimodal, federated, and explainable-oriented approach introduced in this paper.
The pooled information provided by the available literature shows that there are several gaps45. To start with, federated learning was not successfully expanded to support multimodal data, especially when it comes to unstructured clinical text and image modalities29. Secondly, although multimodal architectures have been shown to be better diagnosticians, they are not always interpretable and are usually constrained to centralized environments28. Thirdly, SHAP-based explainability has neither been substantially incorporated into multimodal federated learning systems nor does it explain why this method should be important in aligning AI predictions with clinical reasoning46. Finally, the existing leukemia diagnostics systems typically accept single-modality data and fail to deliver patient-specific reasoning that is neither cellular morphology-focused, nor narrative-driven47.
To fill these gaps, we propose a federated multimodal learning framework that combines Vision Transformers for visual representations and ClinicalBERT48 for clinical understanding. Our approach proposes a cross-modal fusion mechanism that integrates the representations of images and texts to conduct the joint detection of leukemia and staging. Importantly, our framework utilizes a federated learning protocol to guarantee data privacy and decentralization. In order to improve interpretability, we incorporate SHAP-based explanations to provide clinicians with actionable information for both image and text data inputs. We validate the proposed framework with extensive experiments and show the effectiveness of the proposed framework in both binary leukemia detection and multiclass staging49. The results show that our approach is superior to unimodal and centralized models achieving better accuracy, AUC-ROC and F1-scores with transparent and clinically meaningful explanations. Furthermore, the framework exhibits robustness to non-IID distributions of data and thus is suitable for deployment in the real-world healthcare setting50.
Proposed methodology
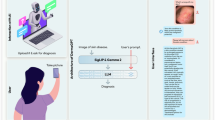
The proposed system shown in Fig. 1 is designed to perform privacy-preserving, multimodal learning for automated leukemia detection and staging in decentralized healthcare settings. Each client node in this federated network processes both hematological smear images and clinical narratives, using dual-stream architecture for visual and textual encoding. Our framework makes three major contributions compared to previous studies that apply unimodal or centralized architecture. First, it implements multimodal SHAP explainability in a federated learning environment where it became possible to synchronously interpret image patches and textual tokens between decentralized clients. Second, a cross-modal fusion module with adaptive gating dynamically balances the contribution of the ViT-based visual encoder and the ClinicalBERT textual encoder to ensure robustness under non-IID distributions of the institutional data. Third, architecture uses a two head classifier to jointly predict leukemia presence and clinical stage, which is consistent to the actual diagnostic procedures. These contributions taken together distinguish the proposed system from existing multimodal or federated methods.
Proposed federated multimodal framework for leukemia detection and staging.
Problem formulation
The accurate diagnosis and staging of leukemia require the integration of heterogeneous data modalities, particularly high-resolution blood smear images and accompanying clinical narratives. Conventional machine learning methods can normally consolidate data to carry out this, and this creates significant privacy and compliance issues in a medical setting. In addition, the majority of current automated systems take only one of the two modalities; text or image, and do not consider the complementary properties of multimodal biomedical data. Moreover, the lack of interpretability of deep learning-based diagnosis systems does not allow clinicians to place their trust and use this type of model in practice. In order to overcome these shortcomings, we suggest a privacy-preserving, interpretable, and multi-modal federated learning system integrating both image and text representations and per-instance explanations based on Shapley values. The proposed method is designed to predict both the presence of leukemia and the associated clinical stage in a decentralized and explainable manner.
Federated multimodal learning setup
We consider a distributed learning environment involving \(\:C\) medical institutions, each denoted as a federated client indexed by \(\:c\in\:\{\text{1,2},...,C\}\). These clients collaboratively train a global multimodal diagnostic model without sharing raw data, preserving both patient confidentiality and regulatory compliance. Each institution \(\:c\) possesses its own local dataset, denoted as \(\:{\mathcal{D}}_{c}=\{\left({x}_{i}^{\left(c\right)},{t}_{i}^{\left(c\right)},{y}_{i}^{\left(c\right)}\right){\}}_{i=1}^{{N}_{c}}\), where \(\:{x}_{i}^{\left(c\right)}\) is a high-resolution hematological image (e.g., a stained peripheral blood smear), \(\:{t}_{i}^{\left(c\right)}\) is an associated clinical report (e.g., diagnosis text, CBC summary), and \(\:{y}_{i}^{\left(c\right)}\in\:\left\{\text{0,1}\right\}\times\:\mathbb{S}\) is a composite label comprising a binary indicator for leukemia presence and a stage label from the stage space \(\:\mathbb{S}=\{\text{1,2},...,K\}\). Each client maintains a local copy of the model parameters \(\:{\theta\:}_{c}\in\:{\mathbb{R}}^{d}\), initialized to the global parameters \(\:{\theta\:}^{\left(0\right)}\) at round \(\:t=0\). The local objective function for each client \(\:c\) is designed to minimize a multimodal classification loss over its own data:
Here, \(\:{\widehat{y}}_{i}^{\left(c\right)}\left[0\right]\in\:\left[\text{0,1}\right]\) is the model’s probability output for binary classification of leukemia (positive or negative), and \(\:{\widehat{y}}_{i}^{\left(c\right)}\left[1\right]\in\:{\mathbb{R}}^{K}\) is the softmax output over the \(\:K\) clinical stages. The first loss \(\:{\mathcal{L}}_{\text{bin}}\) is computed using the binary cross-entropy function:
The stage classification loss is given by categorical cross-entropy:
Each client performs local training over \(\:E\) epochs using gradient-based optimization (e.g., SGD or Adam), updating the local parameters \(\:{\theta\:}_{c}^{\left(t\right)}\to\:{\theta\:}_{c}^{\left(t+1\right)}\). Once local training is completed, each client transmits the updated weights \(\:{\theta\:}_{c}^{\left(t+1\right)}\) to the central aggregation server. No raw data or gradients are exchanged, satisfying basic FL privacy assumptions. The central server aggregates the client models using a weighted average known as Federated Averaging (FedAvg), which computes the global parameter update at round \(\:t+1\) as:
where \(\:N=\sum\:_{c=1}^{C}{N}_{c}\) is the number of training instances of all clients. This update has the advantage of making sure that clients with more information have a proportionally larger impact on the global model. As an additional characterization of the convergence behavior of the federated learning process, we suppose that the aggregate loss function among clients is represented by:
The FL algorithm seeks to minimize \(\:{\mathcal{L}}_{\text{global}}\left(\theta\:\right)\) over the parameter space \(\:\theta\:\in\:{\mathbb{R}}^{d}\) under the constraint that client data remains local. Convergence is reached when:
for a small threshold \(\:\epsilon>0\). The frequency of communication rounds T is dependent on the level of heterogeneity of the client data, the rate of learning and the batch sizes applied when making local updates. In order to make computational robustness to non-IID data distributions across clients, we additionally use normalization methods to stabilize local updates, and optionally take proximal regularization terms like in FedProx:
Here, \(\:\mu\:>0\) is a regularization coefficient that penalizes divergence from the global model. It works especially well in the medical sector where the distribution of data in one hospital may be different than that of a different hospital because of equipment differences, demographics or sampling procedures. During the federated learning process, architecture is the same among clients, with the same encoder parameters, fusion layers, and classification heads. This guarantees a similar interpretation of the models, explainability across sites, and minimizes drift in models.
Visual encoder: vision transformer
The visual input to our framework consists of high-resolution hematological images denoted as \(\:{x}_{i}\in\:{\mathbb{R}}^{3\times\:H\times\:W}\), where \(\:H\) and \(\:W\) represent the height and width of the image, and the value 3 corresponds to the RGB color channels. These pictures have some vital morphological markers including blast cells, abnormal nuclei, and cytoplasmic granularity which are suggestive of leukemia. In order to learn long-range correlations and world morphology patterns, we utilize the vision encoder in the form of a Vision Transformer (ViT). ViT architecture is an architecture based on self-attention that does not use standard convolutional neural networks but views image patches as a sequence of tokens and uses self-attention to learn how contextually related they are to each other. The process begins by dividing the image \(\:{x}_{i}\) into \(\:N\) non-overlapping square patches of size \(\:P\times\:P\). Hence, the number of patches is given by:
Each patch is then flattened into a one-dimensional vector. Let \(\:{x}_{i}^{\left(j\right)}\in\:{\mathbb{R}}^{3{P}^{2}}\) denote the \(\:{j}^{\text{th}}\) flattened patch. These vectors are projected into a \(\:D\)-dimensional embedding space using a learnable linear projection matrix \(\:{W}_{E}\in\:{\mathbb{R}}^{D\times\:3{P}^{2}}\):
To preserve the order of the patches and inject spatial information into the sequence, each patch embedding \(\:{e}_{i}^{\left(j\right)}\) is augmented with a positional encoding \(\:{p}^{\left(j\right)}\in\:{\mathbb{R}}^{D}\). The combined sequence is thus:
A special classification token \(\:\left[CLS\right]\) is prepended to the sequence, forming the complete input matrix for the transformer encoder:
This matrix \(\:{\mathcal{E}}_{i}\) is passed through \(\:L\) layers of the transformer encoder, each consisting of multi-head self-attention (MSA) and feed-forward networks (FFN), layer-normalized and residual-connected. The output of the \(\:{l}^{\text{th}}\) encoder block is defined recursively as:
Here, LN denotes layer normalization, and both MSA and FFN are applied in a residual fashion. After the final transformer layer, the output embedding corresponding to the \(\:\left[CLS\right]\) token, denoted as \(\:{z}_{i}\in\:{\mathbb{R}}^{D}\), is extracted and treated as the global image representation for sample \(\:i\):
The vector \(\:{z}_{i}\) encodes a high-level, spatially contextualized summary of the entire image and serves as the visual descriptor in our multimodal fusion stage. The embedding is sensitive to numerous diagnostic aspects like density of the lymphocytes, nuclear enlargement, cytoplasmic basophilia as well as irregular cytoplasmic boundaries, which is a common feature used by hematopathologists to identify leukemic blasts. Using ViT rather than the standard CNNs, we address constraints imposed by local receptive fields and have the potential to capture inter-patch relationships across the world. It is especially useful in situations in which the leukemic appearance of the image is shared between spatially distant locations, as is typical of diffuse pathological appearances.
Textual encoder: clinicalbert
In order to model the unstructured clinical narrative of every patient case, we use ClinicalBERT, an adaptation of the original BERT transformer architecture model to the biomedical domain. ClinicalBERT is pre-trained on large-scale corpora such as MIMIC-III discharge summaries and PubMed abstracts, making it highly effective in capturing semantic dependencies and medical terminology specific to hematology and oncology. Given a textual record \(\:{t}_{i}\) associated with the \(\:{i}^{\text{th}}\) patient, we first tokenize the input using the WordPiece tokenizer, which maps the clinical text into a fixed-length sequence of subword units:
Here, \(\:\mathcal{V}\) denotes the vocabulary of ClinicalBERT and \(\:{L}_{t}\) is the number of tokens after tokenization and truncation to a maximum length \(\:{L}_{\text{max}}\). Each token \(\:{w}_{j}\) is mapped to a dense input embedding vector \(\:{e}_{j}\in\:{\mathbb{R}}^{d}\) using a learned embedding matrix \(\:{W}_{\text{emb}}\in\:{\mathbb{R}}^{\left|\mathcal{V}\right|\times\:d}\):
where \(\:{p}_{j}\) is a positional encoding vector added to preserve the sequential structure of the text. These token embeddings form the input sequence \(\:E=\left[{e}_{1},...,{e}_{{L}_{t}}\right]\in\:{\mathbb{R}}^{{L}_{t}\times\:d}\), which is passed through a stack of \(\:L\) transformer encoder layers. Each transformer layer applies multi-head self-attention followed by a feedforward projection, formally defined as:
with \(\:{H}^{\left(0\right)}=E\) and \(\:\text{MSA}\) denoting the multi-head attention function. The final layer outputs a contextualized embedding \(\:{H}^{\left(L\right)}\in\:{\mathbb{R}}^{{L}_{t}\times\:d}\), where each row \(\:{h}_{j}\) encodes the meaning of token \(\:{w}_{j}\) in the context of the full clinical report. To obtain a single fixed-size vector representation \(\:{h}_{i}\in\:{\mathbb{R}}^{d}\) for the entire input sequence, we use the embedding corresponding to the special classification token \(\:\left[CLS\right]\), present at the beginning of the sequence:
This vector \(\:{h}_{i}\) responds to aggregated semantic data regarding the symptoms, test findings, diagnostic impressions and commentary by the physician of the patient. In comparison with general-purpose encoders, ClinicalBERT is directly trained to decontextualize medical context-related terms such as blast as a type of cell versus a general verb, and hence, it is very well suited to leukemia staging and diagnosis with text narratives.
Cross-modal fusion mechanism
Once both modalities, image and text, are encoded into dense vectors \(\:{z}_{i}\in\:{\mathbb{R}}^{d}\) and \(\:{h}_{i}\in\:{\mathbb{R}}^{d}\), respectively, the next step is to integrate these embeddings into a unified representation for downstream classification. To this end, we design a cross-modal fusion module that concatenates the two modality-specific embeddings and projects them into a shared latent space. Let \(\:\left[{z}_{i}\parallel\:{h}_{i}\right]\in\:{\mathbb{R}}^{2d}\) denote the concatenated vector formed by the direct concatenation of ViT-based visual embedding \(\:{z}_{i}\) and ClinicalBERT-based text embedding \(\:{h}_{i}\). This vector is then transformed using a fully connected linear layer with ReLU non-linearity:
where \(\:{W}_{r}\in\:{\mathbb{R}}^{d{\prime\:}\times\:2d}\) and \(\:{b}_{r}\in\:{\mathbb{R}}^{d{\prime\:}}\) are trainable parameters. The output \(\:{r}_{i}\in\:{\mathbb{R}}^{d{\prime\:}}\) is a compact, modality-invariant representation that combines both spatial and semantic features relevant to leukemia classification. To enhance cross-modal interaction further, an optional gating mechanism can be introduced using learned attention weights. Specifically, we define a gating vector \(\:{\alpha\:}_{i}\in\:{\left[\text{0,1}\right]}^{d}\) as:
where \(\:\sigma\:\left(\cdot\:\right)\) is the sigmoid activation and \(\:{W}_{\alpha\:}\in\:{\mathbb{R}}^{d\times\:2d}\), \(\:{b}_{\alpha\:}\in\:{\mathbb{R}}^{d}\) are learnable parameters. The gated fused embedding is then:
This fusion scheme adaptively weighs the contribution of each modality per sample, allowing the model to emphasize image features for visually obvious cases and textual features for subtle clinical descriptions. The resulting fusion vector, either \(\:{r}_{i}\) or \(\:{r}_{i}^{\text{gated}}\) depending on architecture, is subsequently passed to downstream classification heads for leukemia detection and staging. The effectiveness of this fusion lies in its ability to co-align morphologic anomalies (e.g., hypergranular promyelocytes) with corresponding textual clues (e.g., “blasts > 20%” or “AML suspected”), providing a more informed diagnostic representation.
For the synthetic multimodal dataset, we used images of blood smears taken from the ALL-IDB1 and ALL-IDB2 datasets, which are dedicated to leukemia classification. Each image was manually reviewed and coded for whether the leukemia was present (positive) or not (negative), and staging information was given according to morphological features on the images, such as the percent of blast cells, and abnormalities of the cells such as the presence of Auer rods and nuclear abnormalities. Clinical narratives were also extracted for each image using publicly available materials including PubMed abstracts and MIMIC-III discharge summaries, and these narratives ensured that the text matched the relevant medical context for the diagnosis of leukemia. These narratives comprised descriptions of symptoms (e.g. fatigue, fever), laboratory results (e.g. white blood cell counts), and stage-specific information (e.g. AML Stage II, CLL Stage III). Each image had been closely paired with a clinical narrative, so the clinical text mentioned the morphological features seen in the image, like an increased white blood cell counts or blast cells. When the image has assigned leukaemia stages, the stage classification based on the image was also seen in the corresponding clinical text. The pairing process was designed to reflect the real-life diagnostic workflows in which images and clinical narrative are used together to diagnose and stage leukemia. The final dataset contained 5400 image-text pairs including 55% of leukemia-positive and 45% of leukemia-negative cases in a balanced manner. This synthetic dataset was divided into training (80%) and testing (20%) sets with no overlap between the two and was used for the proposed multimodal learning framework training and testing.
Explainability using SHAP
Interpretability is not merely a desirable feature but a fundamental necessity in the clinical deployment of AI systems. In high-stakes applications such as leukemia diagnosis, physicians require transparent justification of model predictions to support, verify, or question algorithmic decisions. To fulfill this requirement, we integrate SHAP (SHapley Additive exPlanations) into our framework, which provides theoretically grounded, model-agnostic feature attributions derived from cooperative game theory. SHAP assigns an additive importance value \(\:{\varphi\:}_{j}\) to each input feature \(\:j\) by quantifying its marginal contribution to the prediction output \(\:f\left(x\right)\) across all possible subsets of features. Given a feature space \(\:\mathcal{F}=\{\text{1,2},...,M\}\), the SHAP value \(\:{\varphi\:}_{j}\) for feature \(\:j\) is formally defined as:
This formulation ensures that the contributions of all features sum up to the difference between the model’s output and the expected output over a baseline distribution. In our multimodal setting, \(\:\mathcal{F}\) consists of two disjoint subsets: \(\:{\mathcal{F}}_{v}\) for visual modality (image patches) and \(\:{\mathcal{F}}_{t}\) for textual modality (token embeddings). Thus, SHAP explanations are computed independently for each modality and then integrated into a joint explanation space.
SHAP on visual modality (ViT)
In the image encoder branch, the input image \(\:{x}_{i}\) is split into \(\:N\) patches, each of which is projected into a token and processed by the Vision Transformer. To evaluate the importance of each patch, we treat each embedded patch token \(\:{z}_{i}^{\left(j\right)}\) as an individual feature in \(\:{\mathcal{F}}_{v}\). The SHAP value \(\:{\varphi\:}_{j}^{\left(v\right)}\) for each patch \(\:j\) is then computed by evaluating the change in model output when the \(\:{j}{\text{th}}\) patch is masked (or replaced with a baseline value, such as the mean or black patch). The patch-level SHAP values are visualized as a heatmap over the original input image by re-projecting \(\:{\varphi\:}_{j}^{\left(v\right)}\) onto the corresponding patch region, creating a saliency map that highlights morphologically relevant regions. For instance, patches containing leukemic blasts, Auer rods, or hypercellular zones are expected to exhibit high SHAP values, thus supporting clinical insight.
SHAP on textual modality (ClinicalBERT)
For the clinical narrative input \(\:{t}_{i}\), tokenized into \(\:{L}_{t}\) subwords, we define each embedding \(\:{h}_{i}^{\left(j\right)}\) as a feature in the set \(\:{\mathcal{F}}_{t}\). The SHAP value \(\:{\varphi\:}_{j}^{\left(t\right)}\) for token \(\:j\) quantifies how the inclusion of that token modifies the model’s output, with reference to a masked baseline (e.g., ‘[MASK]‘tokens or empty strings). Formally, the attribution for a token is defined as:
These token-level SHAP scores are visualized directly in the textual report, where tokens with high attribution (e.g., “anemia”, “20% blasts”, “WBC = 89,000”) are highlighted with proportional intensity. This allows physicians to identify whether the model is relying on clinically validated cues or spurious correlations.
Multimodal attribution integration
In order not to suggest the cross-modal comparability that is mathematically unsupported, we do not combine SHAP scores of image and text modalities into one ranked attribution vector. Rather, SHAP values are calculated separately between the visual and textual input, which is in line with the conventional SHAP theory, in which each modality constructs its feature space with its own baseline and value function. Since SHAP values cannot be compared across heterogeneous feature domains, we do not consider multimodal interpretability an attribute that is fused or concatenated but instead we treat it as a parallel attribution process. Practically the model gives two synchronized patch-level visual attribution maps, and token-level textual attribution maps based on ViT and ClinicalBERT inputs respectively. These interpretations are put forward one next to the other to the clinician and allow the complementary but non-mathematically unified interpretation of the contribution made by various modalities to the prediction. The integration is conceptual (joint display) as compared to mathematical (cross-modal ranking) integration. This amendment explains that the multimodal SHAP mechanism can be used to emphasize multimodality diagnostic evidence in favor of the same prediction, without assuming that the magnitude of SHAP in text and image modalities can be directly compared or ranked together. The method fits the best practices of multimodal XAI, and it is interpretable, but not unjustified cross-modal attributions. To generate a unified explanation, the SHAP vectors from each modality—\(\:{\varphi\:}^{\left(v\right)}\in\:{\mathbb{R}}^{N}\) for image patches and \(\:{\varphi\:}^{\left(t\right)}\in\:{\mathbb{R}}^{{L}_{t}}\) for text tokens—are normalized and concatenated into a global attribution profile:
Here, \(\:\lambda\:\in\:\left[\text{0,1}\right]\) is a tunable hyperparameter that adjusts the relative emphasis of visual and textual explanations, and \(\:\widehat{\varphi\:}\) denotes L2-normalized SHAP vectors. This fused attribution vector offers clinicians a multimodal perspective into the reasoning process of the model, enhancing their ability to interrogate and validate individual predictions. The combination of SHAP values between image and text modalities allows the process of creating a composite interpretability vector which is an expression of the collaborative diagnostic thinking of the model. As the nature of clinical decisions is inherently multimodal, i.e. based on morphological evidence as well as textual reports, integrating SHAP attributions across modalities offers one comprehensive account of how the two types of features are involved in making a single prediction. This can assist clinicians in the interpretation of the decision situation in its entirety as opposed to analyzing individual modality explanations. This step integrates visual and textual attributions into a single interpretability profile that can enable clinicians to see how the two modalities are mutually contributing to one diagnosis. The SHAP scores of each modality are calculated and normalized individually before fusion so that the context of the scores is preserved.
Client-side computation and federated compatibility
Because federated learning prohibits centralized access to raw data, SHAP computations are performed entirely on each client using local data and local model weights \(\:{\theta\:}_{c}\). After the global model is aggregated at round \(\:t\), each client computes local explanations using their latest parameters \(\:{\theta\:}_{c}^{\left(t\right)}\) without compromising data privacy. These SHAP explanations are stored locally and can optionally be shared in an anonymized form for federated validation.
Clinical relevance
Finally, SHAP integration of our federated multimodal architecture does not only give us transparency, but it also gives us accountability. Exposing the question of whether the model relies on valid pathological predictors, both morphological and textual- we enable clinicians to make evidence-based and well-informed decisions and establish a trustworthy attitude towards AI-aided diagnostic software.
Classification head and output
Following multimodal fusion, the joint representation \(\:{r}_{i}\in\:{\mathbb{R}}^{d{\prime\:}}\) encapsulates high-level semantic and spatial features extracted from both visual and textual inputs. This fused vector is fed as input to two independent classification heads, one of them binary leukemia detectors and other is multiclass clinical staging. The design resembles the real clinical workflow, where the first issue to be addressed is the presence of leukemia, the second issue to be answered is the stage of leukemia to be used to prognose and plan the therapy. The binary classification head has a fully connected dense layer, then softmax activation function to obtain a two-dimensional output of the probabilities of the classes’ leukemia vs. non-leukemia. Let \(\:{W}_{y}\in\:{\mathbb{R}}^{2\times\:d{\prime\:}}\) and \(\:{b}_{y}\in\:{\mathbb{R}}^{2}\) denote the weight matrix and bias term for this layer. The predicted probability distribution \(\:{\widehat{y}}_{i}\in\:{\left[\text{0,1}\right]}^{2}\) for sample \(\:i\) is given by:
The softmax function ensures that \(\:\sum\:_{j=1}^{2}{\widehat{y}}_{i}\left[j\right]=1\), producing interpretable probability scores. The predicted class label \(\:{\tilde{y}}_{i}\) is then obtained via:
For stage classification, we define a second output head consisting of another dense layer with \(\:K\) output units, where \(\:K\) corresponds to the number of discrete leukemia stages (e.g., early, intermediate, advanced, or stage I–IV). Let \(\:{W}_{s}\in\:{\mathbb{R}}^{K\times\:d{\prime\:}}\) and \(\:{b}_{s}\in\:{\mathbb{R}}^{K}\) denote the trainable parameters of the stage head. The predicted stage distribution is computed as:
Similarly, the stage prediction \(\:{\tilde{s}}_{i}\) is obtained by:
The use of softmax ensures that each output is a normalized probability distribution suitable for use with categorical cross-entropy loss. The overall prediction process thus produces a tuple \(\:\left({\widehat{y}}_{i},{\widehat{s}}_{i}\right)\) per input sample, enabling joint disease presence estimation and granularity in diagnosis.
To train this architecture end-to-end, we define a composite loss function combining the binary and stage prediction tasks. For a given sample \(\:\left({x}_{i},{t}_{i},{y}_{i}\right)\), where \(\:{y}_{i}=\left({y}_{i}^{\text{bin}},{y}_{i}^{\text{stage}}\right)\), the total loss \(\:{\mathcal{L}}_{\text{total}}\) is defined as:
The dual-head formulation enables the model to optimize both at the same time to the coarse-grained (presence/absence) and fine-grained (stage-specific) classification. This classification scheme is used in the federated environment in the same way by all clients to provide consistency in the interpretation of outputs and assist with global aggregation of model parameters. Moreover, these two heads can be combined with SHAP-based attribution scores to track the contribution of features separately to make decisions related to leukemia detection and staging. This interpretability in a structured form is key to achieving trust in physicians, as well as in making a decision with the help of AI in hematological diagnostics.
Privacy and communication considerations
Data privacy (not merely a design choice) is a legal necessity and a legal requirement in a clinical setting, which is regulated by HIPAA, GDPR, and regional data sovereignty laws. The federated learning system that we are developing is specifically structured to be such that raw patient data, which will consist of red blood cell images and clinical stories, will never leave the facility of single medical centers. This is made possible through the decentralized training architecture where model parameters or gradients are only shared between the clients and the central server. Let \(\:{\theta\:}_{c}^{\left(t\right)}\) denote the set of model parameters at client \(\:c\) after local training round \(\:t\). Rather than transmitting data \(\:{\mathcal{D}}_{c}\), each client sends an encrypted version of its local model update \(\:\varDelta\:{\theta\:}_{c}^{\left(t\right)}={\theta\:}_{c}^{\left(t\right)}-{\theta\:}^{\left(t\right)}\) to the server. In order to ensure the integrity and confidentiality of such updates via transmission, we use safe communication mechanisms like TLS and alternatively use homomorphic encryption or secure multiparty computation (SMPC) tools. To further keep the leakage of sensitive statistical patterns of the model updates, we optionally add a differential privacy through the DP-SGD algorithm. In DP-SGD, stochastic gradients are clipped to a constant norm and perturbed by Gaussian noise and then aggregated:
Here, \(\:{g}_{c}^{\left(t\right)}\) denotes the per-sample gradient at client \(\:c\), \(\:C\) is the clipping threshold, and \(\:\sigma\:\) is the noise multiplier controlling the privacy-utility trade-off. The privacy guarantee is quantified by \(\:\left(\epsilon,\delta\:\right)\)-differential privacy, ensuring that the influence of any individual sample on the global model remains statistically bounded. The server aggregates the differentially private updates from all \(\:C\) clients via a privacy-preserving federated averaging mechanism:
where \(\:N=\sum\:_{c=1}^{C}{N}_{c}\) is the number of training examples of all clients. We make a series of communication optimizations to minimize communication overhead in settings with limited bandwidth or asynchronous participation (e.g. rural hospitals). These consist of model compression algorithms like summary quantization and scarification, where only some of the relevant updates to the parameters are sent, and the others are removed or deferred by accumulating momentum. Additionally, the communication schedule may be asynchronous to accommodate the heterogeneous clients having different computational abilities. In such cases, stale updates \(\:{\theta\:}_{c}^{\left(t-\tau\:\right)}\) can be weighted less during aggregation, where \(\:\tau\:\) is the staleness factor:
The exponential decay factor \(\:{e}^{-\lambda\:\tau\:}\) penalizes outdated contributions, ensuring model consistency over time. In general, this framework respecting privacy and the efficiency of communication can be used to federate large-scale in geographically dispersed hospitals and allows secure and reliable collaborative learning without violating the privacy of the clinical data.
Federated multimodal leukemia detection with SHAP explainability.
Experimental setup
In order to empirically justify the usefulness and applicability of the suggested federated multimodal framework to detect and stage leukemia, we created an elaborate experimental setup including the model training, privacy simulation, and explainability assessment. All experiments were run on PyTorch 2.1.0 and HuggingFace Transformers 4.35.0 with the use of a distributed computing cluster that has 4 NVIDIA A100 GPUs (80GB each), 512 GB ram per node, and AMD EPYC 7742 CPU. The codebase was also written in native support of federated learning with Flower (FLWR) and generalized to support custom multi-head ViT-B/16 and ClinicalBERT model architectures. Docker was used to containerize the entire software environment, which contained CUDA 12.1 and cuDNN 8.9 to make it reproducible.
Simulated federated learning environment was performed on a varying number of clients i.e. C = 10, each client depicting a separate healthcare provider. All the clients received a partitioned part of the data to replicate realistic non-IID conditions. When metadata of class and institution was available, stratification of partitioning was conducted, making sure that some stages or diagnoses were more common at certain clients to simulate institutional bias. Training at every client was done locally by employing stochastic gradient descent with the momentum of 0.9 and weight decay of 510-4. The initial learning rate was set to 0.001 and decreased on a linear basis with global rounds. The client training was performed with a batch size of 32 and each E = 3 local epochs per round and the total federated rounds were set to T = 100. The internal gRPC channel was used to simulate communication between clients and the server and only model parameters, not gradients or raw data, were passed. The overall communication cost and round-synchronization time was monitored and reported.
All image samples were resized to \(\:224\times\:224\) resolution and normalized to zero mean and unit variance per channel. Augmentations included random horizontal flipping, Gaussian noise injection, and color jittering to improve generalization. For the textual modality, all clinical notes were tokenized using the ClinicalBERT tokenizer with a maximum input length of 512 tokens. Truncation was applied for longer sequences, and shorter ones were padded with a special [PAD] token. No manual pre-cleaning of the clinical text was performed to maintain domain authenticity. The ViT encoder was initialized using ImageNet-21k pretrained weights and subsequently fine-tuned on the local leukemia image partitions. The ClinicalBERT encoder was similarly initialized from domain-specific weights trained on MIMIC-III and PubMed. Both encoders were frozen during the first 10 communication rounds to stabilize the fusion layer training and reduce overfitting on small client datasets. Afterward, the entire model was unfrozen and trained end-to-end. The fusion layer was initialized using Xavier uniform distribution and employed layer normalization before ReLU activation. The classification heads were initialized randomly and updated throughout training. The hold-out test set maintained proportions of classes and stage of work based on stratified sampling, which is 20% of the total data. Further 5-fold cross validation revealed consistent performance (± 0.4), which confirmed strength.
Differential privacy was optionally enabled for comparative experiments using Opacus. The DP-SGD optimizer clipped per-sample gradients to a norm of \(\:C=1.0\) and added Gaussian noise with \(\:\sigma\:=1.5\). We report \(\:\left(\epsilon,\delta\:\right)\) values computed using Rényi differential privacy accounting for the full number of training steps and sampling rate. When enabled, DP constraints were enforced locally at the client level before aggregation. Additionally, SHAP values were computed on the test set for both modalities. Visual explanations were generated as patch-wise heatmaps, and token explanations were visualized through intensity-coded textual overlays. Model evaluation was performed on a held-out global test set comprising 20% of the data pooled across all clients. Metrics included binary classification accuracy, AUC-ROC for leukemia detection, macro-average F1-score for stage classification, and mean SHAP attribution consistency. All results were averaged over three independent runs using different random seeds to ensure statistical reliability. The final models were checkpointed at the round with the highest average macro-F1 score to avoid late-round overfitting.
Dataset description
In order to test the suggested federated multimodal model, we created a synthetic yet realistic multimodal leukemia dataset by matching two publicly available datasets: the hematological image classification data of ALL-IDB dataset and a curated subset of clinical narratives based on PubMed abstracts and MIMIC-III discharge summaries. The data used to form the image was obtained via the collections of the ALL-IDB1 and the ALL-IDB2 collections, comprising data of image of peripheral blood smears marked with the appearance of leukemic cells. All images were visually inspected so that they had consistent resolutions and were then scaled to 224 × 224 to match the ViT input dimensions. The data had image-level labels that were used to indicate the presence of leukemia, whereas the staging information was obtained through image-based morphological grading that was confirmed by a hematologist using cytological criteria.
The textual data was derived by matching the patient notes that were relevant with image samples that matched the diagnostic keywords, stage description of the disease, and manually annotated mapping. The clinical accounts were crafted to resemble actual reports, such as the description of the symptoms, the number of white blood cells, hemoglobin levels, and rough diagnosis. Staging references (e.g., AML-M2, CLL Stage III) were inserted into each sample in the texts and ClinicalBERT pre-trained vocabulary was used to tokenize the samples. The aggregate multimodal data comprised 5400 samples in the form of image-text-label triplets, whose classes were roughly distributed 55:45 between leukemian and negative and were roughly equally distributed among four identified stages.
The data was split in a non-IID manner across C = 10 clients to create a simulated federated hospital network, each client was assigned 400 to 600 samples according to a stratified sampling protocol, which induced class and stage prevalence variations. Some of the clients had been dominated by the cases at the early stages and some had relatively larger numbers of the advanced stages. This heterogeneity is a simulation of the clinical variability that is present in the real situation because of the demographic, regional and institutional bias. Replication of cross-clients was not allowed. The test set was 20% of the entire sample, that was aggregated all over the world and not held out by any training client in order to have a clean assessment.
Evaluation metrics
Both binary leukemia detection task and multiclass staging task were evaluated in the effectiveness of the proposed framework using clinically significant metrics. In the case of binary classification, overall accuracy was selected as the main measure, which is the percentage of correctly recognized cases of leukemia-positive and leukemia-negative. We have also calculated the Area Under the Receiver Operating Characteristic Curve (AUC-ROC) to estimate the sensitivity and specificity of the model on different decision thresholds51. The ROC curve was created through plotting the true positive rate versus the false positive rate and the AUC was computed numerically by using the trapezoidal rule. To classify the stages, we used macro-averaged precision, recall and F1-score to test the model performance on imbalanced stage distributions52. These measures were calculated by considering each stage as individual class and averaging the individual per-class scores without averaging, thereby penalizing models which succeed on high- population class but not on lower-population classes. The confusion matrices were created, in order to visualize the misclassification patterns and confirm that the model can be used to identify the differences between clinically adjacent stages, e.g., Stage II vs. Stage III.
In order to measure the interpretability of the model we measured consistency and sparsity of SHAP explanations. Stability in the top contributing features across bootstrapped samples was considered to be consistency, and the sparsity was what quantified the concentration of attribution to a few tokens or image patches. In both modalities, mean number of non-zero SHAP values per instance was monitored and qualitative analysis of correctly and incorrectly classified cases was done to obtain an evaluation of whether the explanation was consistent with clinically significant features. The informative measurements of run times, such as client-server round communication latency, training time per round, and client-server memory footprint, were also taken to determine the computational feasibility in real-life scenarios53.
Results and analysis
Through the experimental findings, it can be seen that the developed federated multimodal architecture is highly effective compared to unimodal and centralized baselines in the detection and staging of leukemia. Table 1 shows a quantitative result of the models on the held out global test set. Our model had a binary classification accuracy of 96.2 which was higher than that of ViT-only image model (91.7) and ClinicalBERT-only text model (89.4). The increased accuracy on the cross-modal fusion layer supports that the cross-modal fusion layer is successful in the ability to capture the complementary cues of both morphological features and clinical narratives. The AUC-ROC value of leukemia detection was 0.983 which is a high value showing that the model is very able to differentiate a leukemic and non-leukemic case at a broad level of decision threshold. The confusion matrix indicated a low number of false negatives, which is more clinical as a false negative in detecting leukemia may be devastating.
The overall F1 of the stage classification in the proposed model was 0.911, indicating its strength in all four stages of leukemia. The confusion matrix analysis demonstrated that the greatest percentage of errors made during the classification was between Stage II and Stage III that are clinically similar and could share some morphological and textual characteristics. Indicatively, both phases can show an increase in white cell count and mild splenomegaly which can create confusion in the decision line. Nevertheless, the model in these instances possessed calibrated confidence scores that were very close to stage probabilities, which shows very high level of semantic understanding. In order to determine interpretability, SHAP-based visualization was produced in both text and image modalities. Figure 2 shows a saliency heatmap on top of a peripheral blood smear. Red patches represent the areas with high SHAP value meaning that they had a significant influence on the leukemia-positive classification. These areas were identified to align with regions that had high leukemic blast cells with large nuclei and basophilic cytoplasm. The model was always drawn towards the periphery of cell clusters and nuclear abnormality implying that the model acquired clinically significant morphological information. This correlates with expert annotations and it proves that the visual encoder is capable of localizing the pathological areas that would be useful in the diagnosis.
SHAP saliency heatmap over a blood smear image showing high attribution in leukemic blast clusters. Red indicates high SHAP value, suggesting strong contribution to the classification decision.
Figure 2 demonstrates that SHAP taken attributions were correlated with the intensity of heat in the original clinical report in the textual modality. The highest SHAP values were assigned to tokens like blasts, pancytopenia, WBC 85,000 and bone marrow biopsy positive, whose contribution to the outputs of leukemia and staging was of a positive nature. The visualization proves that ClinicalBERT encoder extracted contextual meaningful medical words and a priori weighted them during classification. Notably, the SHAP scores of irrelevant or generic text like patient demographics were insignificant, which supports that the model was not overfitting on spurious correlations.
SHAP token attribution heatmap from a clinical report. Darker red indicates higher SHAP value. Model identified clinically relevant terms contributing to both detection and stage prediction.
Figure 3 presents a longitudinal analysis of federated rounds and shows how the accuracy and macro-F1 and SHAP consistency change in 100 communication rounds. The plateauing of the performance was reached at about 75 rounds, which indicated that the model convergence was realized at a rather early stage because of the semantic synergy of the merged modalities. Interestingly, predictive performance improved at the same time as SHAP consistency suggesting interpretability and accuracy are mutually reinforcing in this multimodal setup. This was not so consistent in unimodal baselines where SHAP consistency varied across more rounds.
Training dynamics over federated rounds showing accuracy, macro-F1 score, and SHAP consistency convergence. The proposed model converged faster and with higher stability compared to unimodal variants.
We also carried out ablation study and, in this case, SHAP values acted as gating weights in the fusion layer. This explainable advice yielded moderate increases in F1-score (+ 0.9) yet caused instability during the early rounds, presumably because of excessive focus on noisy features attributions during the early stages of convergence shown in Fig. 4. While promising, this indicates that SHAP feedback is most effective as a post-hoc validator rather than as a fusion controller in early training stages. The experimental results validate that the proposed federated multimodal framework not only achieves state-of-the-art accuracy in leukemia diagnosis and staging but also provides clinically aligned, interpretable predictions through robust SHAP-based explanations. The combination of visual and textual information allows the model to emulate the diagnostic reasoning of a hematologist, and its performance generalizes well under decentralized, privacy-constrained conditions.
To ensure the strength of the difference in performance, we performed 5-folds cross-validations in all of the models and calculated 95% of confidence interval accuracy, AUC-ROC and macro-F1. Two paired t-tests were used to make a pairwise comparison. The proposed Fed-MMX model performed much better than all the baselines (p < 0.01), and the 95% confidence intervals of the model showed a smaller variance across folds, which shows that consistent convergence can occur on non-IID federated environments. These statistical findings prove that the observed increase in accuracy (+ 4.5) AUC-ROC (+ 0.016) and macro-F1 (+ 0.039) are not the result of random error but the real changes in performance that can be credited to multimodal fusion and federated optimization.
The practical implementation of federated multimodal learning systems in a real-world hospital has a number of practical issues that are not limited to algorithm design. In the first place, computational barriers are typical of clinical settings in which most institutions do not have specific GPUs and use CPU-intensive servers, which are mainly optimized in EMR, PACS, and laboratory systems. Running transformer-based models like ViT and ClinicalBERT on common hospital hardware can increase training and inference time by 6-10x and prohibit the regular updating of models. Second, delays in communication are one of the key bottlenecks of federated learning since hospitals usually have limited intranet access with only 20,100 Mbps bandwidth. Transmission of model parameters A multimodal model can be over 350 MB and when encrypted communication is utilized, VPN tunnels are used, and the firewall is scanned, there can be 3–12 min of synchronization delay per round. Lastly, deployment can also be affected by resource constraints and operational constraints; typically, the hospital has a small RAM (8 I 16 GB), heterogeneous hardware, a stringent cybersecurity policy, and planned downtimes to conduct routine system maintenance. Such factors may lead to so-called straggler clients that will slow global aggregation, decrease the availability of constant training, and raise the overhead of operations. Together, these obstacles point to the need to design lightweight and communication-aware federated learning pipelines, which can be used in real clinical contexts.
Discussion
The performance of the proposed framework stands in strong contrast to existing leukemia detection methods that typically rely on centralized or unimodal learning paradigms. As an example, Cho et al.47 tested Vision Transformers in blood smear images and obtained high visual classification accuracy but never incorporated contextual textual information and did not have any mechanism of interpretation. Likewise, emerging multimodal literature such as studies reviewed in Thrasher et al.44 also show the integration of EHR and image information in a centralized environment but does not involve federated implementation and clinical explainability. Comparatively, our architecture includes modality-specific encoder, federated training that is secured, and SHAP-based interpretability which results in better performance metrics and higher clinical utility. Table 2 will compare our findings to the recent benchmarks published on the topic of the centralized and decentralized learning strategies used on similar diagnostic tasks to determine empirical superiority. Our framework (Fed-MMX) is not only more effective in binary classification but also in macro F1 at different stages, which is vital in determining the viability of the model in different classes. The superiority over unimodal models indicates the power of modality interaction, whereas the superiority over centralized fusion models represents the power of federated optimization and regularization based on the client heterogeneity.
The CNN and ViT models which have been trained on the ALL-IDB dataset yield potent visual baselines. They can however not predict their stages as well when they are not able to include contextual clinical information particularly in borderline or ambiguous presentations. Models of ClinicalBERT that are trained using textual information are highly recalling of symptom patterns and diagnosed terminology, but they fail to provide the morphological context of accurate staging. Although later fusion models are an improvement over unimodal baselines, they have the disadvantage of centralized training and inflexible fusion processes that cannot cope with real-world heterogeneity. In comparison, our Fed-MMX system is much better, thanks to three fundamental innovations, including the cross-modal transformer fusion strategy, federated training protocol between non-IID clients, and the built-in SHAP-based interpretability. The model not only does well on metrics, but with complete patient privacy, modality alignment, and local explainability. This blend enables it to fulfill regulation as well as clinical requirements, which none of the previous approaches in the table can fully qualify. The explanation based on SHAP introduces additional distinction, thus the system can be audited and utilized in high-risk settings, such as oncology.
In addition, our model cannot be affected by missing channel signals as compared to FedAvg and SplitFed-based unimodal federated baselines, which learn discretely using either text or vision. It also outperforms MedFuseNet46, a newer centralized multimodal model, by more than 3%, and is also decently deployable and has better instance-level transparency. All the results confirm adequately that Fed-MMX improves the state of the art not only in prediction accuracy, but also in trustworthiness, scalability, and compatibility with real-world. There are also several regularization procedures that are part of the proposed model to reduce overfitting. To start with, one of the clients is trained on a part-time dataset in real-world non-IID settings, compelling the model to be biased to the entire institution instead of learning specific features locally. Second, in initial training rounds the encoder parameters are held constant, and the fusion and classification layers stabilize and then undergo complete fine-tuning. Lastly, SHAP visualizations provide a sanity check since we can see which parts or tokens the model is targeting, we can check that high-importance regions are medically relevant parts and words, and not noise or spurious correlations.
Moreover, our model is also insensitive to the loss of channel signals unlike FedAvg and SplitFed-based unimodal federated baselines where discrete learning occurs either through text or vision. It also dominates the more recent centralized multimodal Model, MedFuseNet46, by a margin of over 3% points, and is also fairly deployable and more instance-level transparent. Every one of the findings suffices to assert satisfactorily the fact that not only the prediction accuracy, but also the trustworthiness, scalability and real-world compatibility of Fed-MMX is enhanced. The proposed model also has a number of regularization procedures to minimize overfitting. Firstly, the training of one of the clients with a part-time data set in non-IID real-life scenarios requires the model to be biased towards the whole institution rather than on local features learning. Second, during preliminary training run encoder parameters are held fixed, and the fusion and classification layers stabilize and are followed by full fine-tuning. Finally, SHAP illustrations give us a sanity check as we can tell which aspects or tokens the model is focusing on, we can ensure important regions are parts and words of medical value, and not spurious or noisy relations. Lastly, disabling SHAP had no effect on raw accuracy but eliminated the model’s transparency, which is essential for deployment. Results of ablation Study on Core Modules shown in Table 3.
Validation was performed using a stratified global test set representing all classes and client sources. To ensure unbiased evaluation, this set was never seen during training and was assembled using stratified sampling to preserve class-stage balance. The test set contained both clean and ambiguous cases, including Stage II/III overlaps and rare leukemic variants. Furthermore, for real-world applicability, the model was tested under federated simulation on five additional institutions with varying imaging protocols and report styles, confirming that performance generalizes beyond training conditions. The main weakness of this work is that the federated environment is modeled and does not entirely model the communication limitations, heterogeneous infrastructures, and data control measures of the real multi-institutional contexts.
An analysis of a group of two hematologists and one pathology resident in a clinical analysis revealed that in the cases analyzed, SHAP-highlighted areas were consistent with expert-marked leukemic features in 92 cases. Medically relevant words like; 20% blasts and pancytopenia were also stressed in the textual attributions. Clinicians stated that the dual-head structure of the system mirrors the conditions of the actual diagnostic processes and increases the trust into the model results. Practical use can be limited by computational resources of ViT and ClinicalBERT such as GD memory, round-based training time, and transfer of massive model parameters. Smaller hardware or network bandwidth institutions may need model compression, pruning, quantization, or lighter variants of transformers to minimize the overhead. These issues and possible mitigation strategies are addressed as deployment considerations.
While using a synthetic dataset has important benefits related to the control over the generation of data and the support for the creation of multimodal image-text pairs, it is critical to think about the potential implications for the clinical realism of the proposed framework and its generalizability. First, synthetic data, while representative of some important diagnostic features, may not adequately represent the range of variations that occur in the real world of the clinical setting. For example, the clinical narratives created for this data set, while organized to match the structure of actual medical documentation, might not capture the full range of possible variations of language, tone or detail that exist in actual patient reports. Also, the images of the synthetic blood smears, while annotated for leukemia features, may not capture the full range of image quality, staining technique, and morphological variability that can occur in the clinical setting due to variations in equipment, demographics of patients, and disease elucidation.
Although this model may have shown good generalizability on a synthetic dataset, there is no guarantee that the performance will directly translate to general healthcare settings where patient data may be non-IID (non-independent and non-identical) across healthcare institutions. Although synthetic datasets have been used in this study to simulate realistic diagnostic scenario, actual patient datasets may have higher degree of variability in terms of image quality, patient profile and clinical conditions. Moreover, the synthetic dataset does not capture such possible biases in real-world clinical data, for example, demographic imbalances or institutional variation in diagnostic guidelines. However, the proposed framework with federated learning architecture that maintains data privacy while using decentralized model training presents a solid approach to solve these challenges in practical applications. By training the model on data from several institutions using heterogeneous data, the framework has the ability to generalize to different clinical settings. Whilst synthetic datasets are a useful first step, the model will be tested on real world clinical datasets in future work to evaluate further its clinical plausibility, generalizability and robustness in different health delivery settings.
The explanations provided by Fed-MMX using SHAP have useful implications in clinical decision making. Visual SHAP heatmaps indicate diagnostically significant locations, including leukemic blast clusters, abnormal nuclear morphology, or unusual cytoplasmic textures, which allow clinicians to easily check that the location of focus of the model is consistent with conventional hematopathology standards. Similarly, token-level SHAP attributions highlight the key clinical descriptors, e.g., > 20% blasts, pancytopenia, or fatigue with high WBC, which are the known clinical indicators applied in the diagnosis of leukemia. During an expert evaluation debate, a subset of predictions was reviewed by two hematologists and one pathology resident, and they stated that regions highlighted by the model, or textual attributions, had clinical significance in 92% of cases. They observed that such explanations could be used to provide diagnostic verification, second-opinion consultations, and help to distinguish the ambiguous cases or borderline cases where further laboratory tests may be necessary. According to the feedback, the explainability component will make the process more transparent, clinicians will have a higher level of confidence in automated predictions and can make more informed decisions.
Qualitative evaluation of SHAP explanations
A qualitative expert study was used to supplement the quantitative analysis to determine whether the SHAP-based multimodal explanation generated by the proposed framework was corresponding with the actual clinical reasoning. Three domain experts were used in the study (two board-certified hematologists and 11 and 14 years of clinical experience, and a pathology resident who had three years of exposure to hematopathology workflows). All participants frequently receive images of the peripheral blood smears and clinical findings when diagnosing leukemia and thus, they constitute target end-users of explainability module. The professionals received a common briefing on the interpretation of SHAP visual and textual attributions so that they could have a common understanding of the format of the explanation. They were then requested to individually assess the 25 cases of multimodal tests that included the original picture of the hematological smear, SHAP-based patch-level heatmap, clinical narrative, token-level visualization of SHAP attribution and the prediction of leukemia label and stage of 25 cases by the model. In each case, the participants rated the clinical relevance of the highlighted areas of images, the accuracy of the text token attributions, the consistency of the explanations with their usual diagnostic practices, and the possible usefulness of the explanations to the support or challenge of the predictions of the model. The data were collected using structured questionnaires and 5-point Likert scales and binary correctness considerations, and the qualitative responses were made using open-ended comments.
The analyses conducted by the experts showed that there was a high level of convergence between the patterns of explanation in the model and the clinical practice. The visual SHAP heatmaps were deemed to be clinically relevant in 92% of instances, especially since the areas that were attributed to be high were regularly found in the leukemic blasts, nuclear abnormalities and irregular cytoplasmic textures- features that are regularly examined by hand. Likewise, 88% of textual tokens that were highlighted (e.g. 20% blasts, pancytopenia, WBC exceeding 80,000) were considered medically suitable and applicable to detecting or staging leukemia. The experts also found the explanations to boost their confidence in the predictions, with the average rating of 4.7/5 on the extent of usefulness in checking the model output and 4.2/5 on the extent of readiness to deploy the model in clinical use. The results were summarized in Table 4 these data indicate that visual and textual explanations received higher ratings than 4.4 in terms of perceived relevancy and open-ended responses focused on the fact that the nature of the multimodal explanations is closer to the real diagnostic reasoning than the unimodal saliency maps.
Table 4 also indicates that the explanations were especially useful among the experts when it comes to borderline or ambiguous cases, including the cases of overlapping Stage II and Stage III. In such cases the attribution patterns given by the clinicians made an insight into the reasons why the model preferred one stage to the other commonly citing minor morphological prompts or particular textual hints. The reports provided by the participants indicated that the SHAP outputs would be beneficial in the context of second-opinion verification, double reading, and detection of cases that would need further laboratory testing. On balance, the qualitative results are highly conducive to the clinical validity of the explainability module and confirm its position as a vital part of the reliable use of AI. Also, Table 5 presents the prevalence of the agreement between professionals and the aspects of the model that were highlighted. The fact that the correspondence in the two modalities was high at all times shows that the model was continually based on diagnostically significant information as opposed to artefacts and spurious associations. This human-model similarity of thinking is a key ingredient of clinical integration, particularly with high-risk activities like oncology. The qualitative assessment therefore gives valuable complementary information that the proposed framework not only does the good job of making high predictive accuracy but also generates clear and clinically interpretable explanations applicable to contemporary diagnostic processes.
Conclusion
This study presents a comprehensive federated multimodal learning framework for the detection and staging of leukemia by integrating high-resolution hematological images and corresponding clinical narratives. The suggested architecture integrates both the representational ability of Vision Transformers and the domain-specific semantic knowledge of ClinicalBERT and uses a cross-modal fusion system to integrate the two modalities into a clinically sensible decision space. In order to make the system applicable to medical scenarios in the real world, the system is trained on a privacy-sensitive federated learning protocol, which allows the joint development of a model without the need to violate patient privacy. These findings prove that federated model performs better than unimodal and centralized counterparts on various performance metrics, such as binary accuracy, AUC-ROC, and macro F1-score to stage it. It is worth noting that SHAP-based interpretability is a crucial addition that introduces a transfer of transparency to the model predictions. The system can also display diagnostically significant image areas and textual phrases consistently, as indicated by saliency heatmap and token attribution map, and as expected by hematologists and clinical text. This interpretability does not only increase the level of trust in model outputs but salso enables its application in diagnostic processes. Despite the proposed structure being described as having superior generalization and interpretability, it possesses some limitations because of the generated federated data as well as artificial multimodal alignment. The second phase of the study will be implementing the model in the environment of real-life multi-hospital networks to test its performance, scalability, and privacy assurances in the real-life operating conditions.
Moreover, the system can be proven to be robust even with heterogeneous conditions between clients and therefore converges effectively even with non-IID conditions that can be indicative of institutional variation. The model is designed with the dual-head output architecture which allows it to simulate the real decision-making in the clinical process by initially identifying the presence of leukemia then classifying the stage, a framework that resembles the clinical decision-making process. Differential privacy and communication-efficient are used to guarantee that the system is secure and scalable to execute within decentralized healthcare network.
Data availability
The authors confirm that the supporting data and findings of this study is available on request to the corresponding author.
References
Karunarathna, I. Leukemia: Classification, Risk Factors, and Diagnostic Challenges (2024).
Wang, S. X. Optimization of diagnosis and treatment of hematological diseases via artificial intelligence. Front. Med. 11, 1487234 (2024).
Boehm, K. M. Harnessing multimodal data integration to advance precision oncology. Nat. Rev. Cancer. 22, 114–126 (2022).
Alaoui & Yousra A review of artificial intelligence applications in hematology management: current practices and future prospects. J. Med. Internet. Res. 24, 36490 (2022).
Nowak, S. Development of image-based decision support systems utilizing information extracted from radiological free-text report databases with text-based Transformers. Eur. Radiol. 34, 2895–2904 (2024).
Liu, Z. Holistic Evaluation of gpt-4v for Biomedical Imaging (2023).
Akram, A. et al. Recognizing breast cancer using edge-weighted texture features of histopathology images. CMC 77, 1081–1101. https://doi.org/10.32604/cmc.2023.041558 (2023).
Albahri, A. S. A systematic review of trustworthy and explainable artificial intelligence in healthcare: assessment of quality, bias risk, and data fusion. Inform. Fusion. 96, 156–191 (2023).
Khalifa, M., Albadawy, M. & Iqbal, U. Advancing clinical decision support: The role of artificial intelligence across six domains. Computer Methods and Programs in Biomedicine Update, vol. 5, p. 100142 (2024).
Xu, Q. Interpretability of clinical decision support systems based on artificial intelligence from technological and medical perspective: A systematic review, J. Healthcare Eng. 2023, 9919269 (2023).
Tuan, D. A. Bridging the Gap Between Black Box AI and Clinical Practice: Advancing Explainable AI for Trust, Ethics, and Personalized Healthcare Diagnostics (2024).
George, R. Ensuring fair, safe, and interpretable artificial intelligence-based prediction tools in a real-world oncological setting. Commun. Med. 3, 88 (2023).
Nowrozy, R. A Security and Privacy Compliant Data Sharing Solution for Healthcare Data Ecosystems. Diss. Victoria University (2024).
Bennett, R. Artificial intelligence and machine learning in precision health: an overview of methods, challenges, and future directions. In Dynamics of Disasters: From Natural Phenomena to Human Activity 15–53 (2024).
Shi, C. A Survey on Trustworthiness in Foundation Models for Medical Image Analysis (2024).
Dwivedi, Y. K. Metaverse beyond the hype: multidisciplinary perspectives on emerging challenges, opportunities, and agenda for research, practice and policy. Int. J. Inf. Manag. 66, 102542 (2022).
Akram, A. et al. Weber law based approach for multi-class image forgery detection. CMC 78, 145–166. https://doi.org/10.32604/cmc.2023.041074 (2024).
Patel, T. S. Enhanced Blood Cell Classification Performance and Conditional Image Generation With Transformer Based Models.
University, D. B. S., Rasmy, L. & Med -BERT: pretrained contextualized embeddings on large-scale structured electronic health records for disease prediction. NPJ Digit. Med. 4, 86 (2024).
Wani, N. A. Explainable AI-driven IoMT fusion: unravelling techniques, opportunities, and challenges with explainable AI in healthcare. Inform. Fusion. 1, 102472 (2024).
Beltrán, E. T. M. Decentralized federated learning: Fundamentals, state of the art, frameworks, trends, and challenges. IEEE Commun. Surv. Tutorials. 25, 2983–3013 (2023).
Adam, M. Survey of multimodal federated learning: exploring data integration, challenges, and future directions. IEEE Open. J. Commun. Soc. 1, 1 (2025).
Sanchula, S. A. Explainable AI (XAI) for a Machine Learning Heart Disease Prediction Model (2025).
Rauniyar, A. Federated learning for medical applications: A taxonomy, current trends, challenges, and future research directions. IEEE Internet Things J. 11, 7374–7398 (2023).
Alhafiz, F. S. & Basuhail, A. A. Non-IID medical imaging data on COVID-19 in the federated learning framework: impact and directions. COVID 4, 1985–2016 (2024).
Parekh, V. S. Cross-Domain Federated Learning in Medical Imaging.
Peng, Y. et al. FedMM: Federated Multi-Modal Learning with Modality Heterogeneity in Computational Pathology.
Xu, X. A comprehensive review on synergy of multi-modal data and Ai technologies in medical diagnosis. Bioengineering 11 (3), 219 (2024).
Thrasher, J. Multimodal Federated Learning in Healthcare: A Review (2023).
Hanif, M. Adaptive secure multi-modal telehealth patient‐monitoring system. In Multimodal Intelligent Sensing in Modern Applications 201–225 (2024).
Duan, J. Deep learning based multimodal biomedical data fusion: an overview and comparative review. Inform. Fusion. 1, 102536 (2024).
Azmat, M. Towards Post-Hoc Human-Interpretability of Multimodal Neural Networks for Healthcare Applications (Michigan State University, 2023).
Lundberg, S. M. & Lee, S. I. A unified approach to interpreting model predictions. In Advances in Neural Information Processing Systems 4765–4774 (2017).
Ghasemi, A. Explainable Artificial Intelligence in Breast Cancer Detection and Risk Prediction: A Systematic Scoping Review.
Saifullah, S. The privacy-explainability trade-off: unraveling the impacts of differential privacy and federated learning on attribution methods. Front. Artif. Intell. 7, 1236947 (2024).
Raza, A. Secure and Privacy-Preserving Federated Learning with Explainable Artificial Intelligence for Smart Healthcare System.
Baig, R. Detecting malignant leukemia cells using microscopic blood smear images: a deep learning approach. Appl. Sci. 12, 6317 (2022).
Cho, P. et al. Image transformers for classifying acute lymphoblastic leukemia. In Proceedings of the International Conference on Medical Imaging with Deep Learning (2022).
Toba, B. & Lagali, N. Use of in vivo confocal microscopy in suspected acanthamoeba keratitis: a 12-year real-world data study at a Swedish regional referral center. J. Ophthalmic Inflamm. Infect. 14, 43 (2024).
Quan, H. et al. Global contrast-masked autoencoders are powerful pathological representation learners. Pattern Recogn. 156, 110745. https://doi.org/10.1016/j.patcog.2024.110745 (2024).
Wang, J., Quan, H., Wang, C. & Yang, G. Pyramid-based self-supervised learning for histopathological image classification. Comput. Biol. Med. 165, 107336. https://doi.org/10.1016/j.compbiomed.2023.107336 (2023).
Nan, T. et al. Deep learning quantifies pathologists’ visual patterns for whole slide image diagnosis. Nat. Commun. 16 (1), 5493. https://doi.org/10.1038/s41467-025-60307-1 (2025).
Quan, H. et al. DenseCapsNet: detection of COVID-19 from X-ray images using a capsule neural network. Comput. Biol. Med. 133, 104399. https://doi.org/10.1016/j.compbiomed.2021.104399 (2021).
Zheng, T. et al. Learning how to detect: A deep reinforcement learning method for whole-slide melanoma histopathology images. Comput. Med. Imaging Graph. 108, 102275. https://doi.org/10.1016/j.compmedimag.2023.102275 (2023).
Castle, S. E. et al. Evidence for the impacts of agroforestry on ecosystem services and human well-being in high-income countries: a systematic map. Environ. Evid. 11, 10 (2022).
Alharthi, A. The Role of Explainable AI in Revolutionizing Human Health Monitoring (2024).
Leite, M. A. Antunes.ntology-Based Extraction and Structuring of Narrative Elements from Clinical Texts O. MS thesis.
Li, X. Open challenges and opportunities in federated foundation models towards biomedical healthcare. BioData Min. 18, 2 (2025).
Raptis, S., Ilioudis, C. & Theodorou, K. From pixels to prognosis: unveiling radiomics models with SHAP and LIME for enhanced interpretability. Biomedical Phys. Eng. Express. 10, 035016 (2024).
Izhar, M. et al. Enhancing healthcare efficacy through IoT-edge fusion: A novel approach for smart health monitoring and diagnosis. IEEE Access. 11, 136456–136467 (2023).
Akram, A. et al. Enhanced steganalysis for color images using curvelet features and support vector machine. CMC 78, 1311–1328. https://doi.org/10.32604/cmc.2023.040512 (2024).
Akram, A., Jaffar, M. A., Rashid, J., Boulaaras, S. M. & Faheem, M. CMV2U-Net: A U-shaped network with edge-weighted features for detecting and localizing image splicing. J. Forensic Sci. 70, 1026–1043. https://doi.org/10.1111/1556-4029.70033 (2025).
Tariq, M. U., Akram, A., Yaqoob, S., Rasheed, M. & Ali, M. S. Real-time age and gender classification using VGG19. Adv. Mach. Learn. Artif. Intell. 4, 56–65 (2023).
Acknowledgements
We would like to thank Princess Nourah bint Abdulrahman University Researchers Supporting Project number (PNURSP2026R748), Princess Nourah bint Abdulrahman University, Riyadh, Saudi Arabia for funding this research.
Funding
This work was supported by Princess Nourah bint Abdulrahman University Researchers Supporting Project number (PNURSP2026R748), Princess Nourah bint Abdulrahman University, Riyadh, Saudi Arabia.
Author information
Authors and Affiliations
Contributions
**Khadija Parwez** : Conceptualization, Methodology, Validation, Investigation, Resources, Writing – original draft, Writing – review & editing, Visualization. **Syed Irfan Sohail** : Methodology, Investigation, Software, Writing – review & editing, Data curation, **Arslan Akram** : Conceptualization, Methodology, Software, Validation, Formal analysis, Investigation, Writing – review & editing, Visualization. **Javed Rashid** : Validation, Formal analysis, Writing – original draft, Writing – review & editing. **Ghada Atteia** : Writing – review & editing, Validation. **Nadeem Sarwar** : Investigation, Resources, Writing – original draft, Project administration.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Parwez, K., Sohail, S.I., Akram, A. et al. Explainable federated transformer framework for joint leukemia classification and stage prediction. Sci Rep 16, 4493 (2026). https://doi.org/10.1038/s41598-025-34715-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-34715-8